Simple Summary
The brown rat, an invasive and synanthropic rodent, exhibits highly opportunistic omnivorous feeding habits and plays an important role in the transmission of a wide range of pathogens to humans and domestic animals. Sarcocystis spp. are protists that alternate between two hosts, forming sarcocysts in muscle tissue of the intermediate hosts and developing sporocysts in the intestine of definitive hosts. Information on the role of brown rats as definitive hosts of Sarcocystis parasites under natural conditions is limited, with only S. cymruensis (syn. S. rodentifelis) having been experimentally confirmed to complete its life cycle in this rodent. Intestines of 27 brown rats were collected in Lithuania and Latvia and examined for Sarcocystis spp. The results of a light microscopy examination indicated that seven rats were positive for Sarcocystis spp. sporocysts and oocysts. Based on nested PCR and sequencing of several different genetic regions, more than half of the samples were positive for these parasites. Molecular analysis revealed the presence of DNA of several different Sarcocystis species that use rodents, ungulates, and birds as intermediate hosts. Overall, the study suggests that brown rats might be involved in the transmission of Sarcocystis species in nature.
Abstract
Sarcocystis is a genus of heteroxenous, globally distributed apicomplexan parasites found in reptiles, birds, and mammals. Typically, sarcocysts develop in muscles of intermediate hosts, and oocysts sporulate in intestines of definitive hosts. The parasite’s life cycle is based on prey–predator relationships and usually involves two distinct host species. However, some Sarcocystis spp. can complete their full development within a single host species. Brown rats (Rattus norvegicus) are invasive, synanthropic, highly adaptable rodents that are true omnivores and opportunistic feeders. Therefore, it is possible that they can act as definitive hosts of Sarcocystis parasites. This study provides the first comprehensive assessment of Sarcocystis protists in brown rat intestinal samples under natural conditions, combining microscopy and molecular analyses. Of 27 brown rats investigated, 25.9% (7/27) of animals harbored oocysts/sporocysts of Sarcocystis spp. Based on nested PCR and sequencing of four genetic loci (18S rRNA, 28S rRNA, ITS1, and cox1), 59.3% of samples were positive for Sarcocystis spp. Parasites identified were genetically similar to Sarcocystis spp. using bird–bird, bird–Carnivora, rodent–Carnivora, or ungulate–Carnivora as their intermediate–definitive hosts. The present study suggests that synanthropic rodents may facilitate cross-ecosystem transmission of these parasites, increasing infection pressure on livestock, companion animals, and wildlife in human-dominated environments. Future molecular and dietary ecological studies are needed to assess the role of synanthropic and opportunistic hosts, such as the brown rat, in the transmission of Sarcocystis spp.
1. Introduction
Sarcocystis parasites, which belong to the phylum Apicomplexa, are globally distributed protists that require two hosts to complete their life cycle [1]. The life cycle of Sarcocystis species is tightly linked to predator–prey interactions, with transmission occurring between prey animals serving as intermediate hosts (IH) and predators as definitive hosts (DH) [2,3]. Sarcocystis parasites occur in a wide range of hosts, from poikilothermic reptiles to homeothermic animals, namely mammals and birds. Sarcocyst development typically occurs in muscle tissue and the central nervous system (CNS) of IH. Meanwhile, oocysts sporulate in the small intestine of the DH [3]. To date, over 220 species of Sarcocystis have been documented across reptilian, avian, and mammalian hosts. Some Sarcocystis species are zoonotic [4]. Frequent infections of livestock by these parasites are associated with recurring annual losses in animal production [5]. Additionally, several Sarcocystis species are pathogenic to their IHs, including both wildlife and farm animals [1,2,6]. The severity of disease caused by Sarcocystis is determined by parasite species, infection load, location of sarcocyst formation, immune condition of the host, and other factors.
Various combinations of IHs and DHs in the life cycle of Sarcocystis have been identified [3]. Typically, Sarcocystis exhibits a diheteroxenous life cycle, in which carnivores, omnivores, or scavengers act as DHs and prey animals serve as IHs, meaning that different animal species are required to complete the parasite’s development [1]. The diversity of Sarcocystis species can vary significantly within the same IHs. For instance, raptors typically serve as IHs for only one or two Sarcocystis species [7,8]. In contrast, the mallard (Anas platyrhynchos) may harbor up to four species [9]. Even greater diversity has been observed in other hosts, with at least 13 Sarcocystis species identified in the red deer (Cervus elaphus) [10]. Certain Sarcocystis species that use mice, rats, and lizards as IHs (S. cymruensis, S. dugesii, S. gallotiae, S. muris, S. simonyi, and S. stehlinii) have been shown to exhibit dihomoxenous life cycles, meaning that they complete their entire development within a single host species [1]. These species can be transmitted through cannibalism or coprophagic behavior, either by consuming muscle tissue containing mature sarcocysts or ingesting feces from other individuals of the same host species [11,12,13,14,15,16,17].
Traditionally, research on Sarcocystis parasites has focused mainly on IHs, with a focus on extra-intestinal tissues, especially muscles [1,2,7,8,9]. Most research attention has focused on farm animals due to their economic importance and the impact of Sarcocystis infections on livestock production [18]. Sarcocystis was first described in Switzerland by Miescher in 1843, yet it took more than a century, until 1972, for transmission experiments to clarify its life cycle and identify the DHs of various Sarcocystis species [1]. In transmission experiments, different doses of sporocysts were experimentally administered to suspected IHs, and subsequent infections were monitored [13,14,15,16]. These foundational findings have significantly contributed to the current understanding of the sources of Sarcocystis infection, transmission dynamics, species identification and classification criteria, as well as key aspects of the parasite’s biology relevant to prevention and treatment strategies. However, the application of such transmission experiments is now significantly limited worldwide due to ethical regulations [19]. Experimental studies on Sarcocystis spp. under laboratory conditions are particularly rare for wild mammals and birds. Therefore, alternative approaches to elucidating the life cycles of these protists are essential. Advancements in molecular biology now allow the identification of Sarcocystis spp. through DNA analysis of fecal or intestinal samples from DHs [19,20,21,22,23,24]. The choice of molecular markers for Sarcocystis identification depends on the certain group of parasites. For example, the internal transcribed spacer 1 (ITS1) and the 28S ribosomal RNA gene [20,21,22] are recommended for differentiating avian Sarcocystis species, while the mitochondrial cytochrome c oxidase subunit I (cox1) [19,21,23] is considered the most suitable marker for identifying Sarcocystis spp. that form sarcocysts in farmed animals and other ungulates.
Brown rats (Rattus norvegicus) serve as IHs for at least ten Sarcocystis species [1,24]. Among these, S. singaporensis and S. cymruensis are the most extensively studied [24,25,26,27,28,29,30,31,32]. The first species is important due to its pathogenicity and its potential use in rat control [26,27,28,30]. Notably, S. cymruensis forms macroscopic sarcocysts in the muscles of brown rats [24,25,29,32]. However, knowledge regarding the role of rats in transmitting these protists under natural conditions remains limited. Nevertheless, considering the dietary habits and ecological behavior of these invasive, synanthropic rodents, it is likely that they play a role in the natural transmission cycles of Sarcocystis spp. The brown rat is extremely adaptable, a true omnivore and highly opportunistic feeder [33,34]. They eat a broad spectrum of both plant and animal matter [35,36].
The brown rat inhabits a wide range of environments, especially those altered or associated with humans (e.g., urban areas, sewers, buildings, and refuse dumps) [33,34,35,36]. They also occur in rural landscapes (farmland, hedgerows, fields) and can survive in more natural settings when adequate shelter, food, and water are available [37]. As urban environments constitute a significant portion of the ecosystem encountered by human populations, pathogens carried by brown rats are potential candidates for spillover into humans [35,38]. For example, the brown rat can carry several zoonotic pathogens, including bacteria such as Leptospira [39], Salmonella [40], Listeria [41], and Yersinia pestis [42], as well as hantaviruses [43]. They are also capable of transmitting parasites like the tapeworm Hymenolepis [44] and protozoa such as Giardia [45].
The aim of this study was to assess the prevalence of Sarcocystis spp. in the intestines of brown rats using light microscopy and to identify the parasite species through PCR and sequencing.
2. Materials and Methods
2.1. Sample Collection
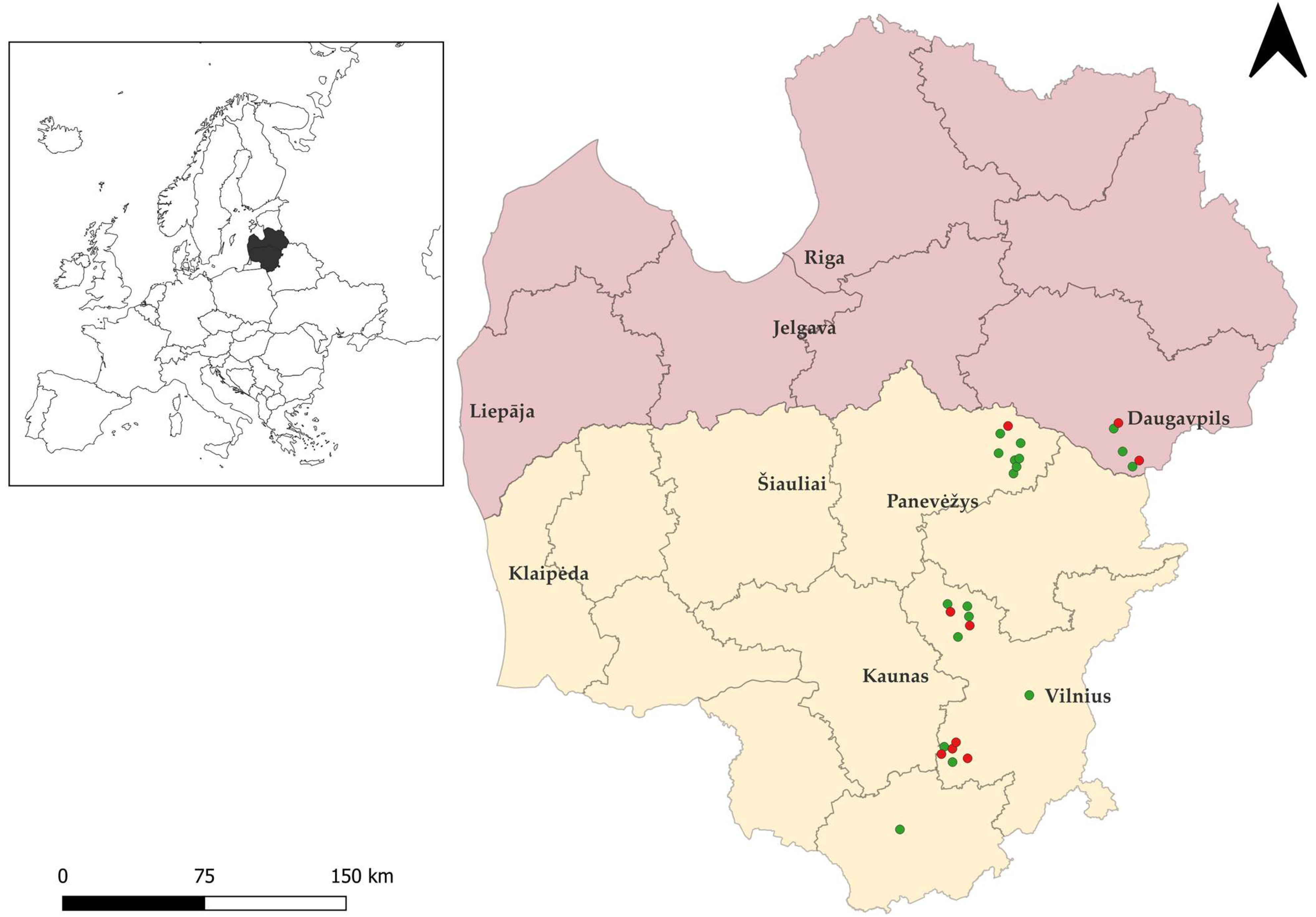
Twenty-seven brown rats were snap-trapped between 2022 and 2025 in various regions of Lithuania and in the Daugavpils municipality of Latvia (Figure 1), were necropsied. Specifically, 22 rats were collected in Lithuania, and five were obtained from Latvia. Notably, in both countries studied, brown rats can be exterminated year-round without a permit [46]. Samples of brown rats were stored at −20 °C until microscopic examination. Intestinal samples were then examined for Sarcocystis spp., and oocysts/sporocysts were recovered from the intestinal mucosa of each animal using previously described methodology [47]. The resulting sediments were examined for oocysts/sporocysts under a Nikon ECLIPSE 8oi (Nikon Corp., Tokyo, Japan) light microscope (LM) at ×400 magnification. The 400 μL of re-suspended sediments were taken from each sample and used for DNA extraction. All samples underwent molecular analysis, regardless of whether oocysts/sporocysts were detected under LM.
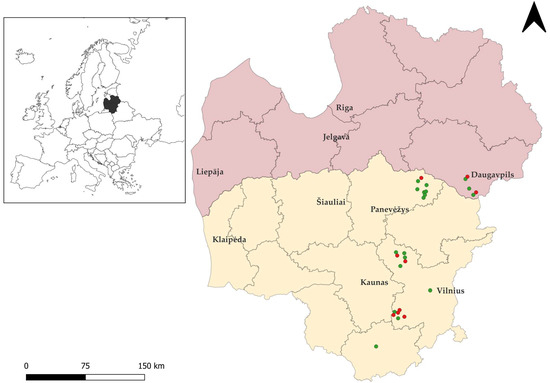
Figure 1.
Geographic distribution of rodent sampling sites in Lithuania and Latvia created with Quantum Geographic Information System (QGIS, https://www.qgis.org, accessed on 11 December 2025). The territory of Lithuania is shown in yellow and that of Latvia in pink. Green circles indicate brown rat individuals positive for Sarcocystis spp., while red circles represent negative individuals.
2.2. DNA Extraction
DNA extraction from the intestinal scrapings of brown rats was conducted with the GeneJET Genomic DNA Purification Kit (Thermo Fisher Scientific Baltics, Vilnius, Lithuania) following the manufacturer’s recommendations. DNA samples were kept frozen at −20 °C until further analysis.
2.3. Molecular Identification of Sarcocystis Species
The role of brown rats in the natural transmission of Sarcocystis spp. within their environment is unknown. Therefore, a numerous set of primers targeting several different genetic regions, i.e., 18S rRNA, 28S rRNA, ITS1 and cox1 were applied (Table 1). The specificity of primers in silico differed as some were specific to certain Sarcocystis species (S. arvalis, S. cymruensis, S. muris, S. myodes and S. ratti with rodents as their IHs), other primers were designed to amplify Sarcocystis spp. using certain groups of animals as their IHs and DHs, and lastly some primers used in the present work were genus specific. Sarcocystis species were amplified using the nested PCR (nPCR) approach. For 18S rRNA, one primer pair was used in the first round of PCR, followed by two internal primer pairs in the second round. For 28S rRNA, three primer pairs were used in the first round, and six pairs of internal primers were used in the second round, depending on the target species. For ITS1, a single primer pair was used in both the first and second rounds. For cox1, two primer pairs were used in the first round, and two internal primer pairs (one per target) were used in the second round.

Table 1.
The oligonucleotides used in this study and their characteristics.
The first round of nPCR was carried out in a 25 μL reaction containing 12.5 μL Taq Master Mix (Vazyme, Red Maple Hi-tech Industry Park, Nanjing, China), 7.5 μL nuclease-free water, 0.5 μL of each external primer, and 4 μL of extracted DNA. For the second amplification of nPCR, reactions were prepared in a 25 μL total volume, consisting of 12.5 μL Taq Master Mix (Vazyme, Red Maple Hi-tech Industry Park, Nanjing, China), 9.5 μL nuclease-free water, and 0.5 μL of each internal primer. However, instead of extracted genomic DNA, 2 μL of the first PCR product was used. Sterile water was used as the negative control in both nPCR rounds, while positive controls were included in all reactions except those for S. muris.
Both rounds of the nPCR were conducted according to the manufacturer’s protocol: beginning with the initial denaturation step at 95 °C for 3 min, followed by 35 cycles consisting of 15 s of denaturation at 95 °C, 15 s of annealing at 51–61 °C (depending on the primer pair used), and 60 s of elongation at 72 °C. A final extension was performed at 72 °C for 5 min. The efficiency of the reactions was verified using 1% agarose gel electrophoresis, and positive amplicons were enzymatically purified with ExoI and FastAP (Thermo Fisher Scientific Baltics, Vilnius, Lithuania) as per the manufacturer’s instructions.
All positive samples were subjected to direct sequencing using Big-Dye® Terminator v3.1 Cycle Sequencing Kit (Thermo Fisher Scientific, Vilnius, Lithuania) and the 3500 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). Both forward and reverse primers of the second round of nPCR were employed for Sanger sequencing. Obtained sequences were compared to most closely related Sarcocystis spp. using Nucleotide BLAST (https://blast.ncbi.nlm.nih.gov/, accessed on 10 November 2025). Phylogenetic analyses were performed in MEGA7.0.26 [54] using the Maximum Likelihood method. Bootstrap support was calculated based on 1000 replicates, and to enhance readability, values < 50% were omitted from the final phylogenetic trees. The sequences generated in the present study are available in GenBank with accession numbers PX550574–PX550577 (ITS1), PX571985–PX571989 (28S rRNA), PX571990–PX571991 (18S rRNA), PX577581–PX577584 (cox1).
3. Results
3.1. Microscopic Identification of Sarcocystis spp. in Intestine Samples
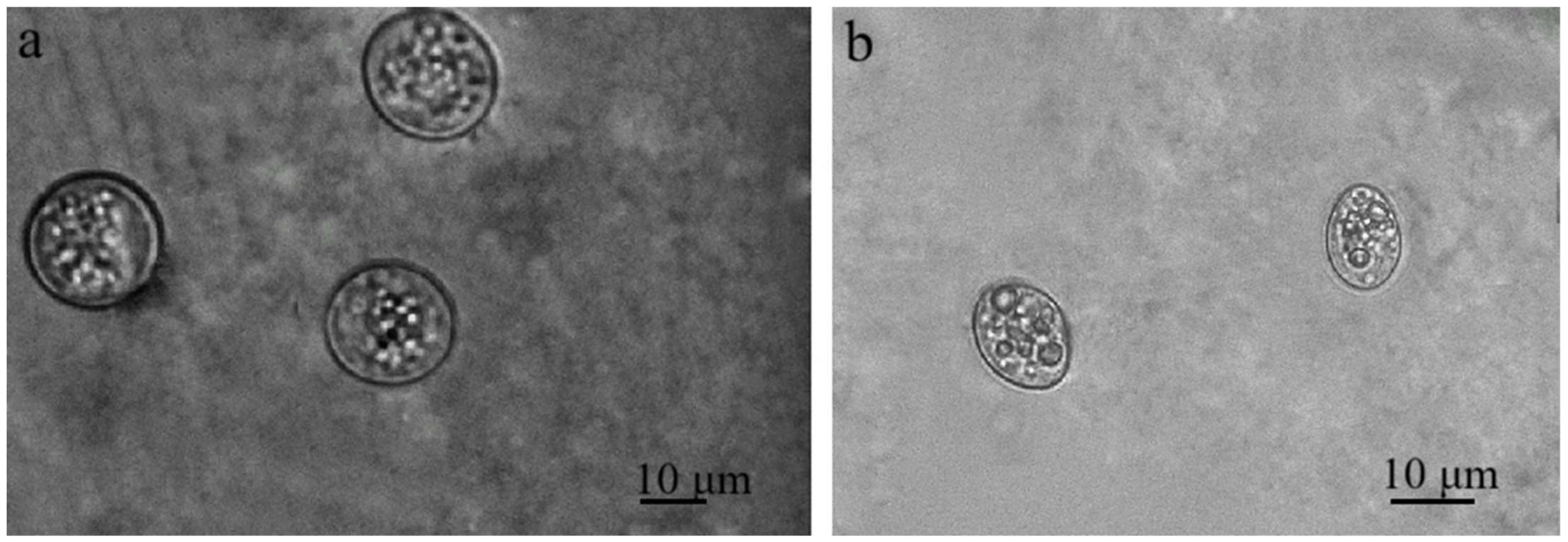
Sarcocystis spp. oocysts and/or sporocysts were observed in the intestinal epithelium under the LM (Figure 2). The number of oocysts and/or sporocysts detected in the area of the 24 × 24 mm coverslip ranged from one to five, with oocysts being more frequently observed than sporocysts. The prevalence of Sarcocystis spp. in mucosal scrapings of brown rats was 25.9% (7/27). Under LM, free sporocysts measuring 8.8–14.3 × 10.0–18.8 μm (10.9 ± 2.6 × 13.7 ± 1.7 μm, n = 24) were observed (Figure 2a). Oocysts of Sarcocystis spp. 14.0–19.5 × 15.4–23.0 μm (16.1 ± 1.7 × 19.3 ± 2.8 μm, n = 18) were also noticed (Figure 2b). It is noteworthy that oocysts and sporocysts were observed in one intestinal epithelial sample from a brown rat (RnLt25z), while in other samples, either sporocysts or oocysts were detected.
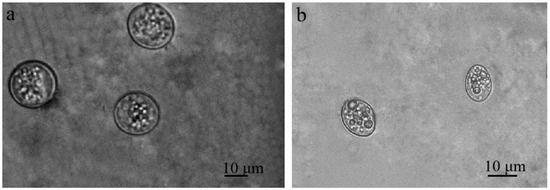
Figure 2.
Sarcocystis spp. Sporulating oocysts (a) and sporocysts (b) in the intestinal epithelium of small intestines of brown rat.
3.2. Molecular Characterization of Sarcocystis spp. in Brown Rats
Having examined the intestines of brown rats, we obtained 15 pure DNA sequences of Sarcocystis spp. (Table 2). Fragments of four different genetic loci, 18S rRNA, 28S rRNA, ITS1, and cox1, were amplified using seven different pairs of primers. BLAST analysis indicated that the sequences obtained in this study corresponded to six previously described Sarcocystis species, whereas four sequences could not be reliably assigned to species level and were therefore classified as Sarcocystis sp. In the case of S. alces, S. capreolicanis, and S. gracilis, our sequences showed 98.8–100% similarity to those of certain identified species and displayed ≤95.3% similarity to those of other Sarcocystis species. Beyond that, our 313 bp 28S rRNA sequence amplified with SgrauzinF/SspRod1R demonstrated 100% identity to S. arvalis and showed ≤97.8% similarity to those of other species. Additionally, four 282 bp ITS1 sequences generated with the primers HSpauk3/HSpauk4 displayed 96.8–99.7% similarity between themselves and showed up to 99.3–100% similarity (specifically, 97.2–100%, 95.4–99.3%, 96.8–99.7%, and 96.5–99.3% similarity) to those of S. halieti. These four sequences showed similarity of 94.7–97.2% to that of Sarcocystis sp. isolate Skua-2016-CH (MW160469). Furthermore, the 337 bp cox1 sequence amplified using SsunV1b/SsunV2b exhibited the highest values of similarity to those of S. tenella (98.8–100%) and S. capracanis (97.0–98.5%) (Table 2).

Table 2.
Genetic identification of Sarcocystis species in intestines of brown rats from Lithuania and Latvia.
Three sequences that could not be assigned to certain Sarcocystis species showed very high and comparable sequence similarity values to several Sarcocystis spp. with a rodent (Sarcocystis sp. Ziu1) or bird (Sarcocystis sp. Ziu2 and Sarcocystis sp. Ziu3) as their IHs. Finally, the 713 bp 28S rRNA sequence of Sarcocystis sp. Ziu4 displayed ≤99.0% similarity to those of other Sarcocystis spp. and demonstrated the highest values of similarity to species forming sarcocysts in muscles of birds (S. wenzeli, S. cristata, S. albifronsi, and S. anasi).
Additionally, we have obtained seven sequences that showed double peaks. These sequences were detected in seven separate brown rats (Table 2), demonstrated at least 84.9% similarity with Sarcocystis spp., and had query coverage values of from 72% to 100%. Two ITS1 sequences obtained using HSpauk3/HSpauk4 displayed the highest similarity to those of S. columbae and S. halieti parasitizing birds. A single impure 28S rRNA sequence exhibited the highest similarity to those of S. alces with moose (Alces alces) as IH. Finally, four cox1 sequences showed the highest similarity to those of Sarcocystis spp. (S. gracilis, S. linearis, and S. tenella) from domestic and wild ungulates. Out of 27 examined brown rats, oocysts and/or sporocysts were morphologically detected in seven (25.9%) samples and molecularly in 16 (59.3%) samples. Two samples were positive only by morphological examination but yielded no Sarcocystis DNA, whereas 11 samples were positive only by molecular analysis. Five samples were positive by both methods, and nine samples were negative by both morphological and molecular examinations. When only samples from animals yielding pure DNA sequences were considered, the molecular detection rate of Sarcocystis spp. was 44.4% (12/27).
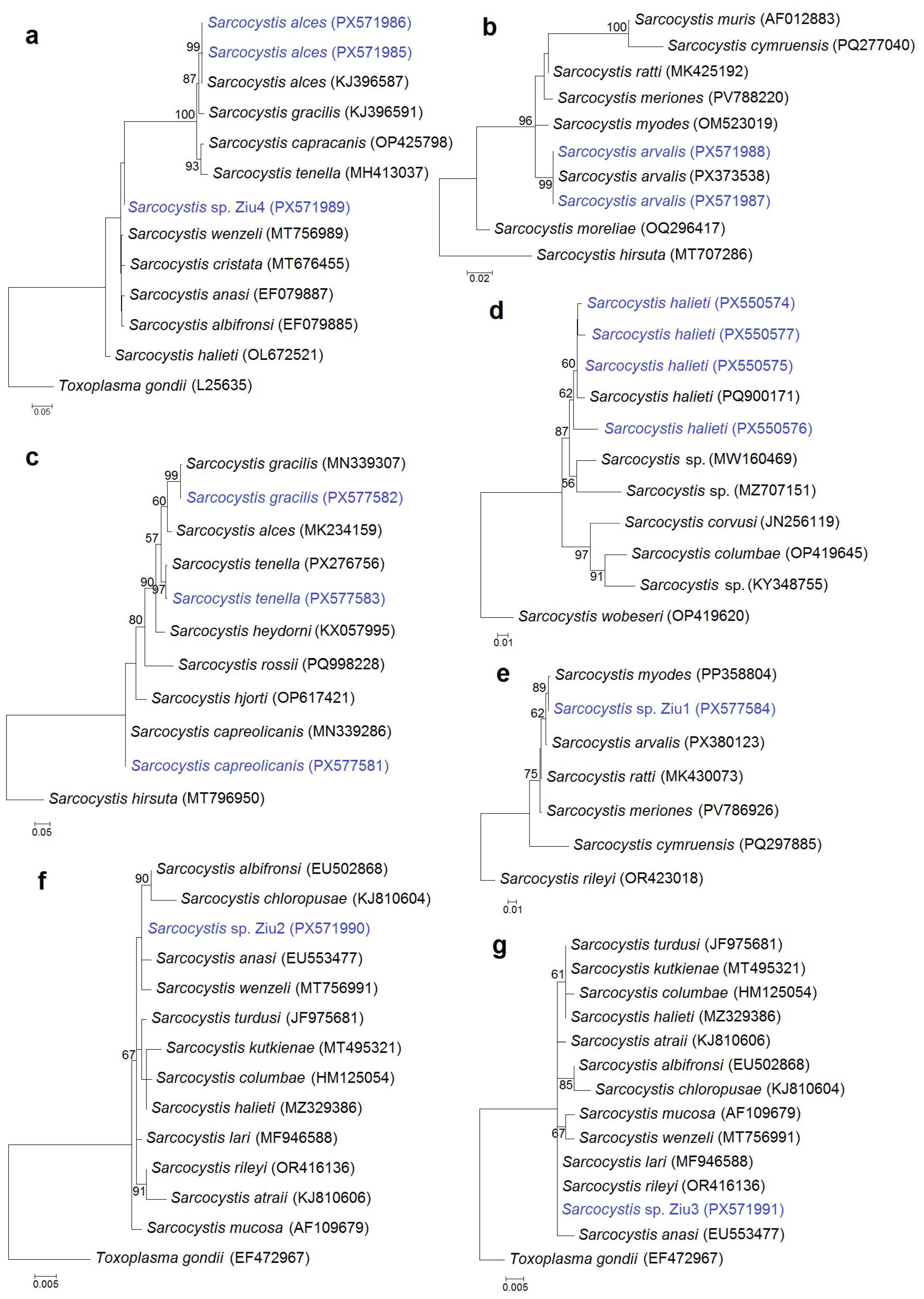
Phylogenetic analyses were performed to confirm species identification and to resolve phylogenetic relationships of the detected Sarcocystis spp. As primer sets targeting four genetic markers and different regions within the same gene were analyzed, seven separate phylogenetic trees were constructed using 28S rRNA (Figure 3a,b), cox1 (Figure 3c), ITS1 (Figure 3d), and 18S rRNA (Figure 3e,f,g) sequence datasets. Sequences of S. alces, S. arvalis, S. gracilis, S. tenella, S. capreolicanis, and S. halieti grouped with reference sequences of their respective species, supporting their molecular identification. In contrast, the variability of some genetic regions analyzed was not sufficient to reliably resolve four Sarcocystis spp., which clustered among several closely related Sarcocystis species. Sarcocystis sp. Ziu1 was grouped with species that use rodents as IHs, forming a clade with S. myodes, S. arvalis, S. ratti, and S. meriones, with a closer relationship to S. myodes. By comparison, Sarcocystis sp. Ziu2, Sarcocystis sp. Ziu3, and Sarcocystis sp. Ziu4 were placed with species that use birds as IHs and carnivores as DHs. Notably, Sarcocystis sp. Ziu4 differed by ≥1% at the conserved 28S rRNA locus from phylogenetically related taxa and formed separate branch in the phylogram (Figure 3a), indicating a genetically distinct, previously uncharacterized Sarcocystis taxon.
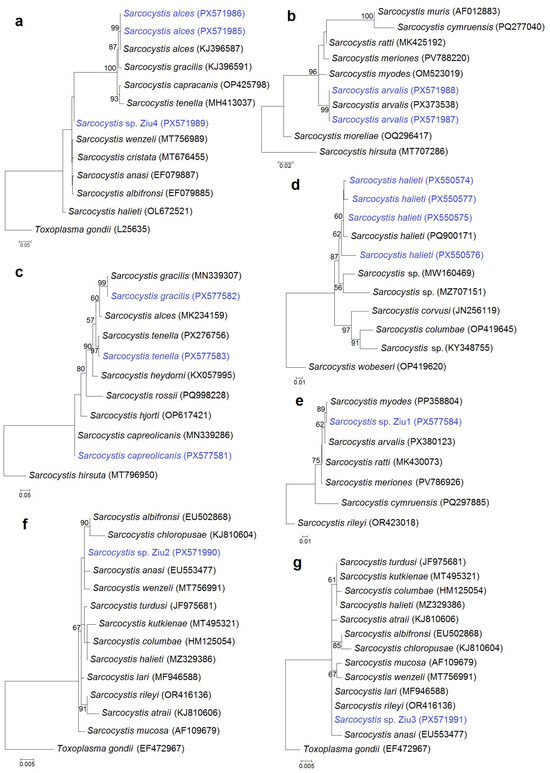
Figure 3.
Phylogenetic trees of selected Sarcocystis species based on 28S rRNA (a,b), cox1 (c,e), ITS1 (d), and 18S rRNA (f,g) gene sequences. The trees were constructed using the maximum likelihood method and rooted with Toxoplasma gondii (a,f,g), S. hirsuta (b,c), S. wobeseri (d), and S. rileyi (e). The following nucleotide substitution models were applied: T92 + G (a,e), HKY + G (b), K2 + G (c), K2 (d,g), and T92 (f). Bootstrap values are shown next to the branches. Sarcocystis spp. obtained in this study are shown in blue.
In two brown rats (RnLt24z and RnLt25z), S. arvalis, which uses rodents and carnivorous mammals as hosts [55], was identified. In addition, a sequence showing the highest similarity to Sarcocystis species using ungulates as IHs was detected in the same animal, indicating that DNA of two different Sarcocystis species was most likely present in these isolates. Likewise, S. alces, which employs moose as its IH, was identified in the brown rat RnLt21z. In this individual, an additional sequence with double peaks and the highest similarity to Sarcocystis spp. forming sarcocysts in bird muscles was also obtained, indicating a mixed infection with at least two Sarcocystis species. Furthermore, two distinct Sarcocystis species were certainly found in the brown rat RnLt14z, while three separate species were established in the rat RnLv4z. Based on 15 unambiguous sequences obtained, the detected Sarcocystis species were characterized by the following IH–DH associations: bird–bird (4 cases), bird–Carnivora (3), rodent–Carnivora (3), and ungulate–Carnivora (5). In five positive brown rats by both morphological and molecular analyses, S. arvalis, S. halieti, and Sarcocystis sp. Ziu1, which demonstrated the highest genetic similarity to Sarcocystis with rodents as IH, were identified.
4. Discussion
To our knowledge, this study provides preliminary evidence of the presence of Sarcocystis spp. in the intestinal epithelium of brown rats under natural conditions. Sarcocystis spp. oocysts/sporocysts were detected by LM in 25.9% (7/27) of brown rats examined. In comparison, the presence of Sarcocystis spp. DNA was confirmed by molecular analysis in 16 (59.3%) mucosal scrapings from brown rats. Comparative analysis of the obtained sequences revealed the highest similarity to species that employ birds, rodents, and ungulates as IHs. According to previous experimental studies, oocysts/sporocysts of S. cymruensis were detected in mucosal scrapings and within the remaining wall of the small intestine in 23 (88.5%) Wistar (Norway) laboratory rats [15]. To date, no additional data are available regarding brown rats as potential DHs for Sarcocystis spp.
Evolutionarily, parasites that have only one DH persist by using several IH species in their life cycle [56,57]. Consequently, parasite life cycles restricted to one IH and one DH species represent an evolutionary paradox, particularly when infection can occur in only one host through ingestion of sporocysts. Numerous experimental transmission attempts of Sarcocystis spp. from diverse DHs to brown rats as IHs have been unsuccessful [58,59,60], apart from the case of S. singaporensis, in which successful transmission was achieved [27]. While transmission experiments are still regarded as the gold standard for identifying the DHs of Sarcocystis species, their use has become increasingly limited due to high costs, lengthy duration, and significant ethical considerations [61]. In this study, Sarcocystis spp. were confirmed by both microscopical and molecular analyses in five brown rats. In the intestines of two brown rats (RnLt11z and RnLt12z), oocysts or sporocysts of Sarcocystis spp. were detected by light microscopy; however, no DNA sequences were obtained using the primer set applied in this study. Furthermore, molecular screening alone detected Sarcocystis DNA in the intestines of 11 additional brown rats. It is not possible to rule out pseudoparasitism (the organism is found in a host but is not actually acting as a parasite) for certain Sarcocystis species detected in the intestinal epithelium of brown rats. Additionally, the detection of Sarcocystis DNA in intestinal samples does not unequivocally demonstrate endogenous sporocyst formation, since the DNA may originate from species ingested with the rodents’ diet [62]. In this study, sporocysts characteristic of Sarcocystis were identified in three intestinal samples of brown rats (RnLt11z, RnLt21z, and RnLt25z), suggesting that these rodents may act as DHs. Additionally, oocysts were detected in the intestinal epithelium of several individuals (RnLv5z, RnLt10z, RnLt12z, RnLt15z, and RnLt25z).
Considering the ecological traits and feeding habits of invasive, synanthropic brown rats [35,36,63,64], their involvement in the natural transmission cycles of Sarcocystis spp. cannot be excluded. The brown rat represents one of the most successful commensal pests in human history, profoundly influencing numerous aspects of human life [65]. Brown rats are known to exhibit muricidal behavior, wherein they kill and devour mice [66,67]. This behavior serves dual functions, reducing interspecific competition and exploiting mice as a convenient food resource. Furthermore, brown rats display pronounced scavenging tendencies and readily consume degraded animal tissues, including fish, avian, and mammalian remains, most often sourced from human-generated waste or natural carrion [68,69].
In the present study, six Sarcocystis spp. (S. alces, S. arvalis, S. capreolicanis, S. gracilis, S. halieti, and S. tenella) were identified in the intestinal epithelium of brown rats for the first time. It is noteworthy that canids act as the DHs for S. alces [23,70], S. capreolicanis [47,71,72], S. gracilis [71,72], and S. tenella [23,72]. Meanwhile, molecular analyses indicated that three sequences of unidentified Sarcocystis species (Sarcocystis sp. Ziu2, Sarcocystis sp. Ziu3, and Sarcocystis sp. Ziu4) were most genetically similar to bird muscle–infecting species (S. albifronsi, S. anasi, S. atraii, S. cristata, S. rileyi, and S. wenzeli) transmitted by carnivores [61]. In this work, S. halieti was one of the most commonly detected species (n = 4). Molecular analyses indicate that the DHs of S. halieti are raptors belonging to the orders Accipitriformes [20,62,73,74,75] and Falconiformes [62]. Therefore, it is unlikely that brown rats serve as DHs for the above-mentioned Sarcocystis spp. Instead, these Sarcocystis species were probably associated with carrion consumption and failed to establish infection within the host.
Based on the comparison of the obtained 28S rRNA sequences, for the first time, S. arvalis has been identified in the intestinal samples of two brown rats. Recently, S. arvalis was described in the muscles of the common vole (Microtus arvalis) from Lithuania [55]. The DH of this species remains undetermined; however, based on phylogenetic relationships, predatory mammals are considered the most likely hosts. Alternative transmission routes for S. arvalis cannot be excluded, including cannibalism or coprophagy, whereby individuals consume muscle tissue harboring mature sarcocysts or feces from conspecific hosts. A similar scenario may pertain to the putative new Sarcocystis sp. Ziu1, with rodents as IHs.
Our study suggests that brown rats might be involved in the transmission of Sarcocystis species in nature. Sporocysts of Sarcocystis spp. were detected in three intestinal scrapings of brown rats, indicating that the sexual stage of the parasite may be completed in these rodents. However, the detected numbers of sporocysts and/or oocysts were low when compared with those typically found in DHs such as canids [23,47], felids [1], wild birds [22,75], and snakes [1,76]. Taken together, these results suggest that brown rats are unlikely to represent typical DHs. However, under certain ecological conditions, such as occasional coprophagy or cannibalism, they may potentially act as DHs, with sporocyst excretion occurring at low intensity but possibly over an extended period [15]. These findings suggest that brown rats may play a previously underappreciated role in maintaining Sarcocystis transmission cycles in human-dominated landscapes, potentially affecting domestic animals and wildlife. Future studies integrating large-scale molecular screening, phylogenetic analyses, dietary ecology, and histological examination of intestinal tissues will be essential to identify DHs of Sarcocystis species under natural conditions, particularly in synanthropic and opportunistic hosts such as the brown rat.
5. Conclusions
This study provides the first detailed examination of intestinal samples from brown rats for Sarcocystis protists under natural conditions using combined microscopical and molecular approaches. Molecular screening showed higher sensitivity than light microscopy (59.3% vs. 25.9%). Based on nPCR and sequencing of 18S rRNA, 28S rRNA, ITS1, and cox1 fragments, six previously described Sarcocystis species and four genetically distinct parasite lineages were detected. The identified Sarcocystis spp. were associated with bird–bird, bird–Carnivora, rodent–Carnivora, and ungulate–Carnivora IH–DH relationships, indicating that in most cases brown rats do not serve as true DHs. Nevertheless, the detection of low numbers of sporocysts and oocysts in several individuals suggests that brown rats may occasionally complete the sexual life cycle of certain Sarcocystis species, possibly through coprophagy or cannibalism. Overall, brown rats are unlikely to represent typical DHs but may play a limited, context-dependent role in Sarcocystis transmission in nature.
Author Contributions
Conceptualization, G.Š. and P.P.; methodology, E.J.-N. and P.P.; validation, D.B. and P.P.; formal analysis, G.Š., E.J.-N. and P.P.; investigation, G.Š. and E.J.-N.; resources, G.Š., E.J.-N., D.B. and P.P.; data curation, E.J.-N.; writing—original draft preparation, G.Š., E.J.-N., D.B. and P.P.; writing—review and editing, G.Š., E.J.-N., D.B. and P.P.; visualization, E.J.-N.; supervision, P.P.; project administration, P.P.; funding acquisition, G.Š. All authors have read and agreed to the published version of the manuscript.
Funding
The research was funded by the Research Council of Lithuania (grant number S-PD-24-2).
Institutional Review Board Statement
In Lithuania and Latvia, brown rats can be trapped and culled year-round without a special permit. All laboratory examinations in the present study were conducted with the approval of the Animal Welfare Committee of the State Scientific Research Institute Nature Research Centre (Protocol No. GGT-9, 12 January 2024).
Informed Consent Statement
Not applicable.
Data Availability Statement
The sequences generated in the present study have been deposited in GenBank under the accession numbers PX550574–PX550577 (ITS1), PX571985–PX571989 (28S rRNA), PX571990–PX571991 (18S rRNA), and PX577581–PX577584 (cox1).
Acknowledgments
The authors are grateful to Eglė Rudaitytė-Lukošienė for her assistance in reviewing the article and for her extremely valuable comments.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Dubey, J.P.; Calero-Bernal, R.; Rosenthal, B.M.; Speer, C.A.; Fayer, R. Sarcocystosis of Animals and Humans, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2016. [Google Scholar]
- Fayer, R. Sarcocystis spp. in Human Infections. Clin. Microbiol. Rev. 2004, 17, 894–902. [Google Scholar] [CrossRef]
- Prakas, P.; Butkauskas, D. Protozoan Parasites from Genus Sarcocystis and Their Investigations in Lithuania. Ekologija 2012, 58, 45–58. [Google Scholar] [CrossRef]
- Castro-Forero, S.P.; Bulla-Castañeda, D.M.; López Buitrago, H.A.; Díaz Anaya, A.M.; Madeira De Carvalho, L.M.; Pulido-Medellín, M.O. Sarcocystis spp., a Parasite with Zoonotic Potential. Bulg. J. Vet. Med. 2022, 25, 175–186. [Google Scholar] [CrossRef]
- Martínez-Navalón, B.; Anastasio-Giner, B.; Cano-Fructuoso, M.; Sanchez-Martínez, P.; Llopis-Morant, A.; Perez-Castarlenas, B.; Goyena, E.; Berriatua Fernández De Larrea, E. Short communication. Sarcocystis Infection: A Major Cause of Carcass Condemnation in Adult Sheep in Spain. Span. J. Agric. Res. 2012, 10, 388. [Google Scholar] [CrossRef]
- Pinto, M.D.S.; Camargo Neto, J.A.B.; Lopes, C.W.G.; Paiva, F.; Barros, L.D.; Felippelli, G.; Rodrigues, F.S.; Widmer, G.; Bresciani, K.D.S. Sarcocystosis in Farm Animals in Brazil: A One-Health Approach. Vet. Sci. 2025, 12, 842. [Google Scholar] [CrossRef]
- Prakas, P.; Bea, A.; Juozaitytė-Ngugu, E.; Olano, I.; Villanúa, D.; Švažas, S.; Butkauskas, D. Molecular Identification of Sarcocystis halieti in the Muscles of Two Species of Birds of Prey from Spain. Parasites Vectors 2021, 14, 414. [Google Scholar] [CrossRef]
- Prakas, P.; Šukytė, T.; Juozaitytė-Ngugu, E.; Butkauskas, D. Detection of Sarcocystis halieti in Muscles of Raptors from Lithuania. Front. Vet. Sci. 2025, 12, 1568013. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Zhang, M.; Wu, Z.; Zeng, H.; Tao, J. Description of Sarcocystis platyrhynchosi n. sp. (Apicomplexa: Sarcocystidae) from Domestic Ducks Anas platyrhynchos (Anseriformes: Anatidae) in China. Parasites Vectors 2023, 16, 50. [Google Scholar] [CrossRef] [PubMed]
- Rudaitytė-Lukošienė, E. Sarcocystis Species Richness in Members of the Family Cervidae. Ph.D. Thesis, State Research Institute Nature Research Centre, Vilnius, Lithuania, 2020. Available online: https://vb.gamtc.lt/object/elaba:83121806/ (accessed on 24 November 2025).
- Matuschka, F.R. Reptiles as Intermediate and/or Final Hosts of Sarcosporidia. Parasitol. Res. 1987, 73, 22–32. [Google Scholar] [CrossRef]
- Bannert, B. Sarcocystis simonyi sp. nov. (Apicomplexa: Sarcocystidae) from the Endangered Hierro giant lizard Gallotia simonyi (Reptilia: Lacertidae). Parasitol. Res. 1992, 78, 142–145. [Google Scholar] [CrossRef]
- Grikienienė, J. New Experimental Data on Development Cycle of Sarcosporidians (Sarcosporidia) of Rodents. Ekologija 1993, 1, 33–46. (In Russian) [Google Scholar]
- Grikienienė, J.; Arnastauskienė, T.; Kutkienė, L. On Some Disregarded Ways of Sarcosporidians Circulation and Remarks about Systematics of the Genus Sarcocystis Lankester, 1882 with the Description of the New Species from Rodents. Ekologija 1993, 1, 16–24. (In Russian) [Google Scholar]
- Grikienienė, J.; Kutkienė, L. New Experimental Data on the Laboratory Rat as a Definitive Host of Sarcocystis rodentifelis. Acta Zool. Litu. Parasitol. 1998, 8, 121–124. (In Russian) [Google Scholar] [CrossRef]
- Koudela, B.; Modrý, D.; Svobodová, M.; Votýpka, J.; Vávra, J.; Hudcovic, T. The Severe Combined Immunodeficient Mouse as a Definitive Host for Sarcocystis muris. Parasitol. Res. 1999, 85, 737–742. [Google Scholar] [CrossRef]
- Koudela, B.; Modrý, D. Sarcocystis muris possesses Both Diheteroxenous and Dihomoxenous Characters of Life Cycle. J. Parasitol. 2000, 86, 877–879. [Google Scholar] [CrossRef] [PubMed]
- Dubey, J.P.; Rosenthal, B.M. Bovine Sarcocystosis: Sarcocystis Species, Diagnosis, Prevalence, Economic and Public Health Considerations, and Association of Sarcocystis Species with Eosinophilic Myositis in Cattle. Int. J. Parasitol. 2022, 53, 463–475. [Google Scholar] [CrossRef] [PubMed]
- Máca, O.; Gudiškis, N.; Butkauskas, D.; González-Solís, D.; Prakas, P. Red Foxes (Vulpes vulpes) and Raccoon Dogs (Nyctereutes procyonoides) as Potential Spreaders of Sarcocystis Species. Front. Vet. Sci. 2024, 11, 1392618. [Google Scholar] [CrossRef]
- Gjerde, B.; Vikøren, T.; Hamnes, I.S. Molecular identification of Sarcocystis halieti n. sp., Sarcocystis lari and Sarcocystis truncata in the Intestine of a White-Tailed Sea Eagle (Haliaeetus albicilla) in Norway. Int. J. Parasitol. Parasites Wildl. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Juozaitytė-Ngugu, E.; Švažas, S.; Šneideris, D.; Rudaitytė-Lukošienė, E.; Butkauskas, D.; Prakas, P. The Role of Birds of the Family Corvidae in Transmitting Sarcocystis Protozoan Parasites. Animals 2021, 11, 3258. [Google Scholar] [CrossRef]
- Šukytė, T.; Butkauskas, D.; Juozaitytė-Ngugu, E.; Švažas, S.; Prakas, P. Molecular Confirmation of Accipiter Birds of Prey as Definitive Hosts of Numerous Sarcocystis Species, including Sarcocystis sp., Closely Related to Pathogenic S. calchasi. Pathogens 2023, 12, 752. [Google Scholar] [CrossRef]
- Prakas, P.; Kalashnikova, T.; Gudiškis, N.; Šneideris, D.; Juozaitytė-Ngugu, E.; Butkauskas, D. Molecular Evidence of Raccoon Dog (Nyctereutes procyonoides) as a Natural Definitive Host for Several Sarcocystis Species. Pathogens 2025, 14, 288. [Google Scholar] [CrossRef]
- Aryan, F.A.M.; Prakas, P.; Juozaitytė-Ngugu, E.; Šneideris, D.; Abd-Al-Aal, Z.; Alhoot, A.A.A.; El-Kabbany, A.I.; Tahrani, L.M.A.; El-Azazy, O.M.E. Sarcocystis cymruensis in the Brown Rat (Rattus norvegicus) from an Urban District in Kuwait: Detailed Morphologic and Molecular Characterization. Acta Parasitol. 2025, 70, 11. [Google Scholar] [CrossRef] [PubMed]
- Ashford, R.W. Sarcocystis cymruensis n. sp., a Parasite of Rats Rattus norvegicus and Cats Felis catus. Ann. Trop. Med. Parasitol. 1978, 72, 37–43. [Google Scholar] [CrossRef]
- Brehm, H.; Frank, W. Der Entwicklungskreislauf von Sarcocystis singaporensis Zaman und Colley, 1976 im End-und Zwischenwirt. Z. Parasitenkd. 1980, 62, 15–30. [Google Scholar] [CrossRef]
- Jäkel, T.; Burgstaller, H.; Frank, W. Sarcocystis singaporensis: Studies on Host Specificity, Pathogenicity, and Potential use as a Biocontrol Agent of Wild Rats. J. Parasitol. 1996, 82, 280–287. [Google Scholar] [CrossRef] [PubMed]
- Jäkel, T.; Khoprasert, Y.; Endepols, S.; Archer-Baumann, C.; Suasa-ard, K.; Promkerd, P.; Kliemt, D.; Boonsong, P.; Hongnark, S. Biological Control of Rodents using Sarcocystis singaporensis. Int. J. Parasitol. 1999, 29, 1321–1330. [Google Scholar] [CrossRef] [PubMed]
- Antunes Murata, F.H.; Cerqueira-Cézar, C.K.; Thompson, P.C.; Tiwari, K.; Mowery, J.D.; Verma, S.K.; Rosenthal, B.M.; Sharma, R.N.; Dubey, J.P. Sarcocystis cymruensis: Discovery in Western Hemisphere in the Brown rattus (Rattus norvegicus) from Grenada, West Indies: Redescription, Molecular Characterization, and Transmission to IFN-γ Gene Knockout Mice via Sporocysts from Experimentally Infected Domestic Cat (Felis catus). Parasitol. Res. 2018, 117, 1195–1204. [Google Scholar] [CrossRef] [PubMed]
- Jäkel, T.; Promkerd, P.; Sitthirath, R.; Guedant, P.; Khoprasert, Y. Biocontrol of Rats in an Urban Environment in Southeast Asia using Sarcocystis singaporensis. Pest. Manag. Sci. 2019, 75, 2148–2157. [Google Scholar] [CrossRef]
- Lee, F.C.H. Finding Sarcocystis spp. on the Tioman Island: 28S rRNA Gene Next-Generation Sequencing reveals nine New Sarcocystis Species. J. Water Health 2019, 17, 416–427. [Google Scholar] [CrossRef]
- Zeng, H.; Guo, Y.; Ma, C.; Deng, S.; Hu, J.; Zhang, Y. Redescription and Molecular Characterization of Sarcocysts of Sarcocystis cymruensis from Norway rats (Rattus norvegicus) and Sarcocystis ratti from Black rats (R. rattus) in China. Parasitol. Res. 2020, 119, 3785–3791. [Google Scholar] [CrossRef]
- Maligana, N.; Julius, R.S.; Shivambu, T.C.; Chimimba, C.T. Genetic Identification of freely Traded Synanthropic Invasive Murid Rodents in Pet Shops in Gauteng Province, South Africa. Afr. Zool. 2020, 55, 149–154. [Google Scholar] [CrossRef]
- Ringani, G.V.; Julius, R.S.; Chimimba, C.T.; Pirk, C.W.W.; Zengeya, T.A. Predicting the Potential Distribution of a Previously Undetected Cryptic Invasive Synanthropic Asian House Rat (Rattus tanezumi) in South Africa. J. Urban Ecol. 2022, 8, juac005. [Google Scholar] [CrossRef]
- Himsworth, C.G.; Parsons, K.L.; Jardine, C.; Patrick, D.M. Rats, Cities, People, and Pathogens: A Systematic Review and Narrative Synthesis of Literature Regarding the Ecology of Rat-Associated Zoonoses in Urban Centers. VBZD 2013, 13, 349–359. [Google Scholar] [CrossRef] [PubMed]
- Feng, A.Y.T.; Himsworth, C.G. The Secret Life of the City Rat: A Review of the Ecology of Urban Norway and Black Rats (Rattus norvegicus and Rattus rattus). Urban Ecosyst. 2014, 17, 149–162. [Google Scholar] [CrossRef]
- Desvars-Larrive, A.; Baldi, M.; Walter, T.; Zink, R.; Walzer, C. Brown Rats (Rattus norvegicus) in Urban Ecosystems: Are the Constraints related to Fieldwork a Limit to their Study? Urban Ecosyst. 2018, 21, 951–964. [Google Scholar] [CrossRef]
- Parsons, M.H.; Banks, P.B.; Deutsch, M.A.; Corrigan, R.F.; Munshi-South, J. Trends in Urban Rat Ecology: A Framework to define the Prevailing Knowledge Gaps and Incentives for Academia, Pest Management Professionals (PMPs) and Public Health Agencies to participate. J. Urban Ecol. 2017, 3, jux005. [Google Scholar] [CrossRef]
- Webster, J.P.; Ellis, W.A.; Macdonald, D.W. Prevalence of Leptospira spp. in Wild Brown Rats (Rattus norvegicus) on UK Farms. Epidemiol. Infect. 1995, 114, 195–201. [Google Scholar] [CrossRef]
- Hilton, A.C.; Willis, R.J.; Hickie, S.J. Isolation of Salmonella from Urban Wild Brown Rats (Rattus norvegicus) in the West Midlands, UK. Int. J. Environ. Health Res. 2002, 12, 163–168. [Google Scholar] [CrossRef]
- Webster, J.P. Wild Brown Rats (Rattus norvegicus) as a Zoonotic Risk on Farms in England and Wales. Commun. Dis. Rep. CDR Rev. 1996, 6, R46–R49. [Google Scholar]
- Anderson, D.M.; Ciletti, N.A.; Lee-Lewis, H.; Elli, D.; Segal, J.; DeBord, K.L.; Overheim, K.A.; Tretiakova, M.; Brubaker, R.R.; Schneewind, O. Pneumonic Plague Pathogenesis and Immunity in Brown Norway Rats. Am. J. Pathol. 2009, 174, 910–921. [Google Scholar] [CrossRef]
- Maas, M.; De Vries, A.; Reusken, C.; Buijs, J.; Goris, M.; Hartskeerl, R.; Ahmed, A.; Van Tulden, P.; Swart, A.; Pijnacker, R.; et al. Prevalence of Leptospira spp. and Seoul Hantavirus in Brown Rats (Rattus norvegicus) in four Regions in the Netherlands, 2011–2015. Infect. Ecol. Epidemiol. 2018, 8, 1490135. [Google Scholar] [CrossRef]
- Yang, D.; Zhao, W.; Zhang, Y.; Liu, A. Prevalence of Hymenolepis nana and H. diminuta from Brown Rats (Rattus norvegicus) in Heilongjiang Province, China. Korean J. Parasitol. 2017, 55, 351–355. [Google Scholar] [CrossRef]
- Li, J.; Qin, H.; Li, X.; Zhang, L. Role of Rodents in the Zoonotic Transmission of Giardiasis. One Health 2023, 16, 100500. [Google Scholar] [CrossRef] [PubMed]
- Antanavičius, J.; Zagrebnevienė, G.; Čaplinskas, S. Graužikai, jų Platinamų Ligų Prevencija, Graužikų Naikinimo Būdai; Užkrečiamųjų ligų ir AIDS centras: Vilnius, Lithuania, 2017. [Google Scholar]
- Šneideris, D.; Gudiškis, N.; Juozaitytė-Ngugu, E.; Kalashnikova, T.; Butkauskas, D.; Prakas, P. High Richness of Ungulate Sarcocystis Species in Intestines of the Grey Wolf (Canis lupus) from Lithuania. Vet. Res. Commun. 2025, 49, 235. [Google Scholar] [CrossRef]
- Kutkienė, L.; Prakas, P.; Sruoga, A.; Butkauskas, D. The Mallard Duck (Anas platyrhynchos) as Intermediate Host for Sarcocystis wobeseri sp. nov. from the Barnacle Goose (Branta leucopsis). Parasitol. Res. 2010, 107, 879–888. [Google Scholar] [CrossRef] [PubMed]
- Prakas, P.; Stirkė, V.; Šneideris, D.; Rakauskaitė, P.; Butkauskas, D.; Balčiauskas, L. Protozoan Parasites of Sarcocystis spp. in Rodents from Commercial Orchards. Animals 2023, 13, 2087. [Google Scholar] [CrossRef] [PubMed]
- Rudaitytė-Lukošienė, E.; Palinauskas, V.; Markakis, G.; Butkauskas, D.; Prakas, P. Detection of Sarcocystis halieti DNA in the Blood of Western House Martin (Delichon urbicum) and Barn Swallow (Hirundo rustica) from Lithuania, and in Eurasian Griffon Vulture (Gyps fulvus) from Greece. Acta Parasitol. 2025, 70, 180. [Google Scholar] [CrossRef]
- Gjerde, B. Phylogenetic Relationships among Sarcocystis Species in Cervids, Cattle and Sheep Inferred from the Mitochondrial Cytochrome C Oxidase Subunit I Gene. Int. J. Parasitol. 2013, 43, 579–591. [Google Scholar] [CrossRef]
- Prakas, P.; Rudaitytė-Lukošienė, E.; Šneideris, D.; Butkauskas, D. Invasive American Mink (Neovison vison) as Potential Definitive Host of Sarcocystis elongata, S. entzerothi, S. japonica, S. truncata and S. silva using Different Cervid Species as Intermediate Hosts. Parasitol. Res. 2021, 120, 2243–2250. [Google Scholar] [CrossRef]
- Marandykina-Prakienė, A.; Butkauskas, D.; Gudiškis, N.; Juozaitytė-Ngugu, E.; Januškevičius, V.; Rudaitytė-Lukošienė, E.; Prakas, P. Molecular Identification of Sarcocystis Species in Sheep from Lithuania. Animals 2022, 12, 2048. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Bagdonaitė, D.L.; Rudaitytė-Lukošienė, E.; Stirkė, V.; Balčiauskas, L.; Butkauskas, D.; Prakas, P. Description of Sarcocystis arvalis n. sp. from the Common Vole (Microtus arvalis) in Lithuania Using Morphological and Molecular Methods. Pathogens 2025, 14, 1086. [Google Scholar] [CrossRef]
- Choisy, M.; Brown, S.P.; Lafferty, K.D.; Thomas, F. Evolution of Trophic Transmission in Parasites: Why add Intermediate Hosts? Am. Nat. 2003, 162, 172–181. [Google Scholar] [CrossRef]
- Benesh, P.D.; Parker, G.; Chubb, C.J. Life-Cycle Complexity in Helminths: What are the Benefits? Evolution 2021, 8, 1936–1952. [Google Scholar] [CrossRef] [PubMed]
- Ruiz, A.; Frenkel, J.K. Recognition of Cyclic Transmission of Sarcocystis muris by Cats. J. Infect. Dis. 1976, 133, 409–418. [Google Scholar] [CrossRef] [PubMed]
- McKenna, P.B.; Charleston, W.A.G. Coccidia (Protozoa: Sporozoasida) of Cats and Dogs. II. Experimental Induction of Sarcocystis Infections in Mice. N. Z. Vet. J. 1980, 28, 117–119. [Google Scholar] [CrossRef] [PubMed]
- Cawthorn, R.J.; Gajadhar, A.A.; Brooks, R.J. Description of Sarcocystis rauschorum sp. n. (Protozoa: Sarcocystidae) with Experimental Cyclic Transmission between varying Lemmings (Dicrostonyx richardsoni) and Snowy Owls (Nyctea scandiaca). Can. J. Zool. 1984, 62, 217–225. [Google Scholar] [CrossRef]
- Prakas, P.; Calero-Bernal, R.; Dubey, J.P. Sarcocystis Infection in Domestic and Wild Avian Hosts: Inseparable Flight Partners. Vet. Parasitol. 2025, 335, 110413. [Google Scholar] [CrossRef]
- Juozaitytė-Ngugu, E.; Švažas, S.; Bea, A.; Šneideris, D.; Villanúa, D.; Butkauskas, D.; Prakas, P. Molecular Confirmation of Raptors from Spain as Definitive Hosts of Numerous Sarcocystis Species. Animals 2025, 15, 646. [Google Scholar] [CrossRef]
- Schein, M.W.; Orgain, H. A Preliminary Analysis of Garbage as Food for the Norway Rat. Am. J. Trop. Med. Hyg. 1953, 2, 1117–1130. [Google Scholar] [CrossRef]
- Yabe, T. The Relation of Food Habits to the Ecological Distributions of the Norway Rat (Rattus norvegicus) and the Roof Rat (R. rattus). Jpn. J. Ecol. 1979, 29, 235–244. [Google Scholar]
- Guiry, E.; Buckley, M. Urban Rats have Less Variable, Higher Protein Diets. Proc. Biol. Sci. 2018, 285, 20181441. [Google Scholar] [CrossRef] [PubMed]
- Malick, J.B. Effects of Age and Food Deprivation on the Development of Muricidal Behavior in Rats. Physiol. Behav. 1975, 14, 171–175. [Google Scholar] [CrossRef]
- Bridgman, L.J.; Innes, J.; Gillies, C.; Fitzgerald, N.B.; Miller, S.; King, C.M. Do ship Rats display Predatory Behavior towards Mice? Anim. Behav. 2013, 86, 257–268. [Google Scholar] [CrossRef]
- Schwartz, L.W.A.; Williams, F.H.; Chadwick, E.; Thomas, R.J.; Perkins, E.S. Roadkill Scavenging Behaviour in an Urban Environment. J. Urban Ecol. 2018, 1, juy006. [Google Scholar] [CrossRef]
- Muletz-Wolz, C.R.; Rankin, E.W.; McGrath-Blaser, S.; Venkatraman, M.; Maldonado, J.E.; Gruner, D.S.; Fleischer, R.C. Identification of Novel Bacterial Biomarkers to detect Bird Scavenging by Invasive Rats. Ecol. Evol. 2021, 11, 1814–1828. [Google Scholar] [CrossRef]
- Colwell, D.D.; Mahrt, J.L. Development of Sarcocystis alceslatrans Dubey, 1980, in the Small Intestine of Dogs. Am. J. Vet. Res. 1983, 44, 1813–1818. [Google Scholar] [CrossRef] [PubMed]
- Moré, G.; Maksimov, A.; Conraths, F.J.; Schares, G. Molecular Identification of Sarcocystis spp. in Foxes (Vulpes vulpes) and Raccoon Dogs (Nyctereutes procyonoides) from Germany. Vet. Parasitol. 2016, 220, 9–14. [Google Scholar] [CrossRef] [PubMed]
- Lückner, S.; Moré, G.; Marti, I.; Frey, C.F.; Fernandez, J.E.; Belhout, C.; Basso, W. High Prevalence of Sarcocystis spp. in the Eurasian Wolf (Canis lupus lupus): Third-Generation Sequencing resolves Mixed Infections. Int. J. Parasitol. Parasites Wildl. 2025, 28, 101140. [Google Scholar] [CrossRef]
- Rogers, K.H.; Arranz-Solís, D.; Saeij, J.P.J.; Lewis, S.; Mete, A. Sarcocystis calchasi and other Sarcocystidae Detected in Predatory Birds in California, USA. Int. J. Parasitol. Parasites Wildl. 2021, 17, 91–99. [Google Scholar] [CrossRef]
- Máca, O.; González-Solís, D. Role of three Bird Species in the Life Cycle of two Sarcocystis spp. (Apicomplexa, Sarcocystidae) in the Czech Republic. Int. J. Parasitol. Parasites Wildl. 2022, 17, 133–137. [Google Scholar] [CrossRef] [PubMed]
- Šukytė, T.; Juozaitytė-Ngugu, E.; Švažas, S.; Butkauskas, D.; Prakas, P. The Genetic Identification of Numerous Apicomplexan Sarcocystis Species in Intestines of Common Buzzard (Buteo buteo). Animals 2024, 14, 2391. [Google Scholar] [CrossRef] [PubMed]
- Bezerra, T.L.; de Jesus, R.F.; Soares, R.M.; Borges-Silva, W.; Gondim, L.F.P. Sarcocystis sp. shed by the Common Boa Snake (Boa constrictor) in Brazil. Int. J. Parasitol. Parasites Wildl. 2023, 22, 84–91. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.