Metabolic Profiling of Plasma in Different Calving Body Condition Score Cows Using an Untargeted Liquid Chromatography-Mass Spectrometry Metabolomics Approach
Simple Summary
Abstract
1. Introduction
2. Material and Methods
2.1. Experimental Design and Samples Collection
2.2. LC-MS Metabolomics Analysis
2.3. Measurement of Classical Metabolic Parameters
2.4. Data Processing and Analyses
3. Results
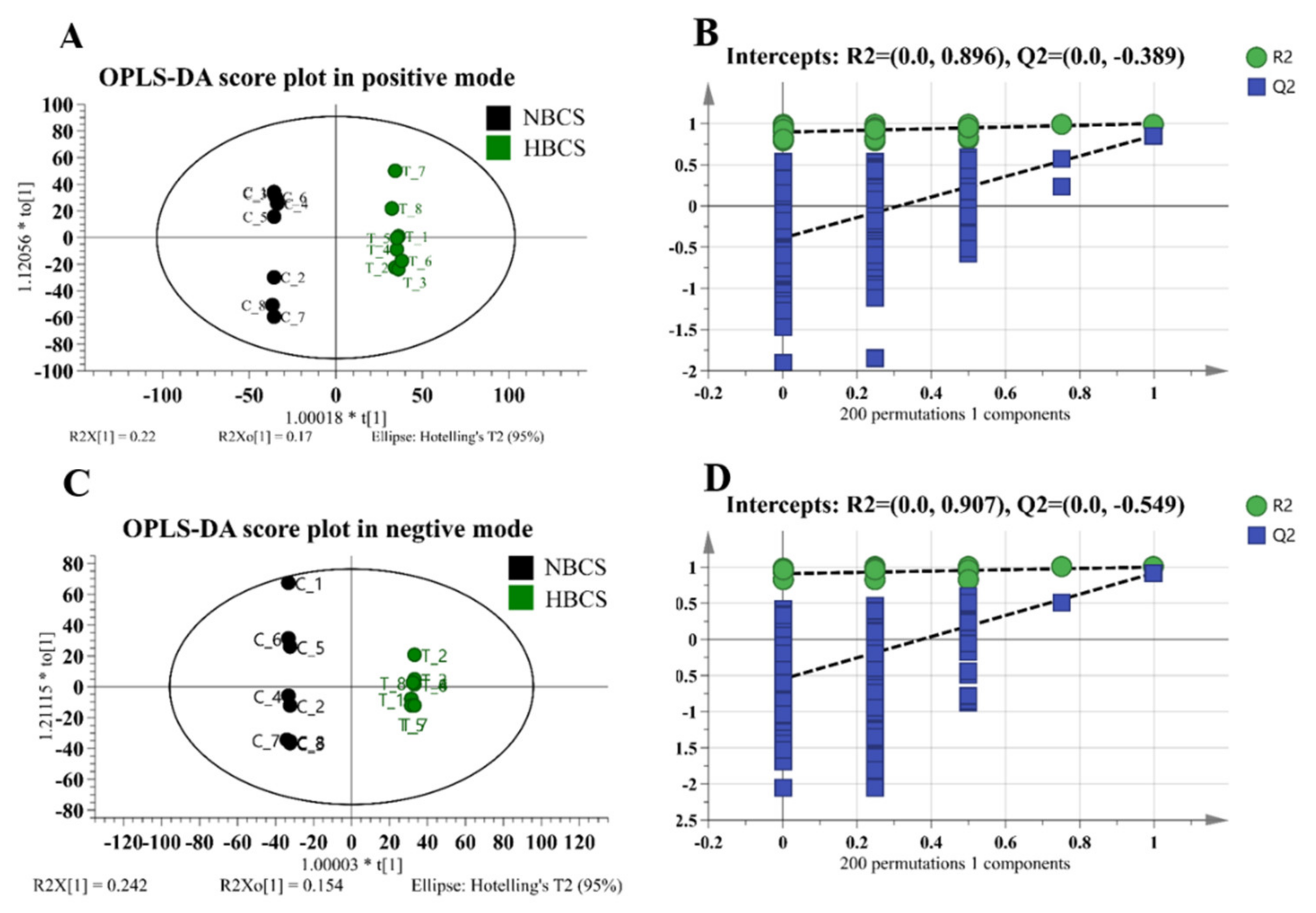
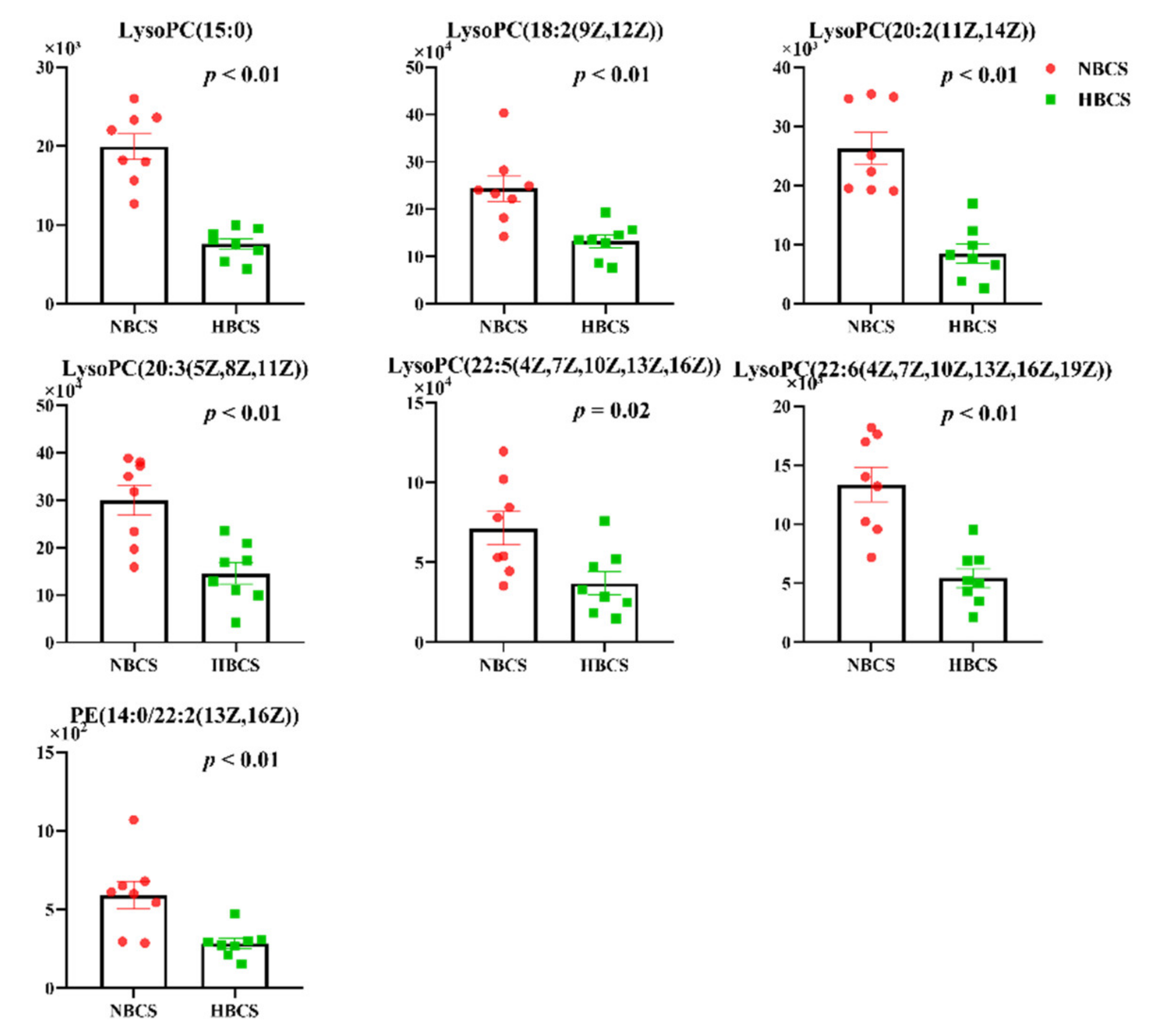
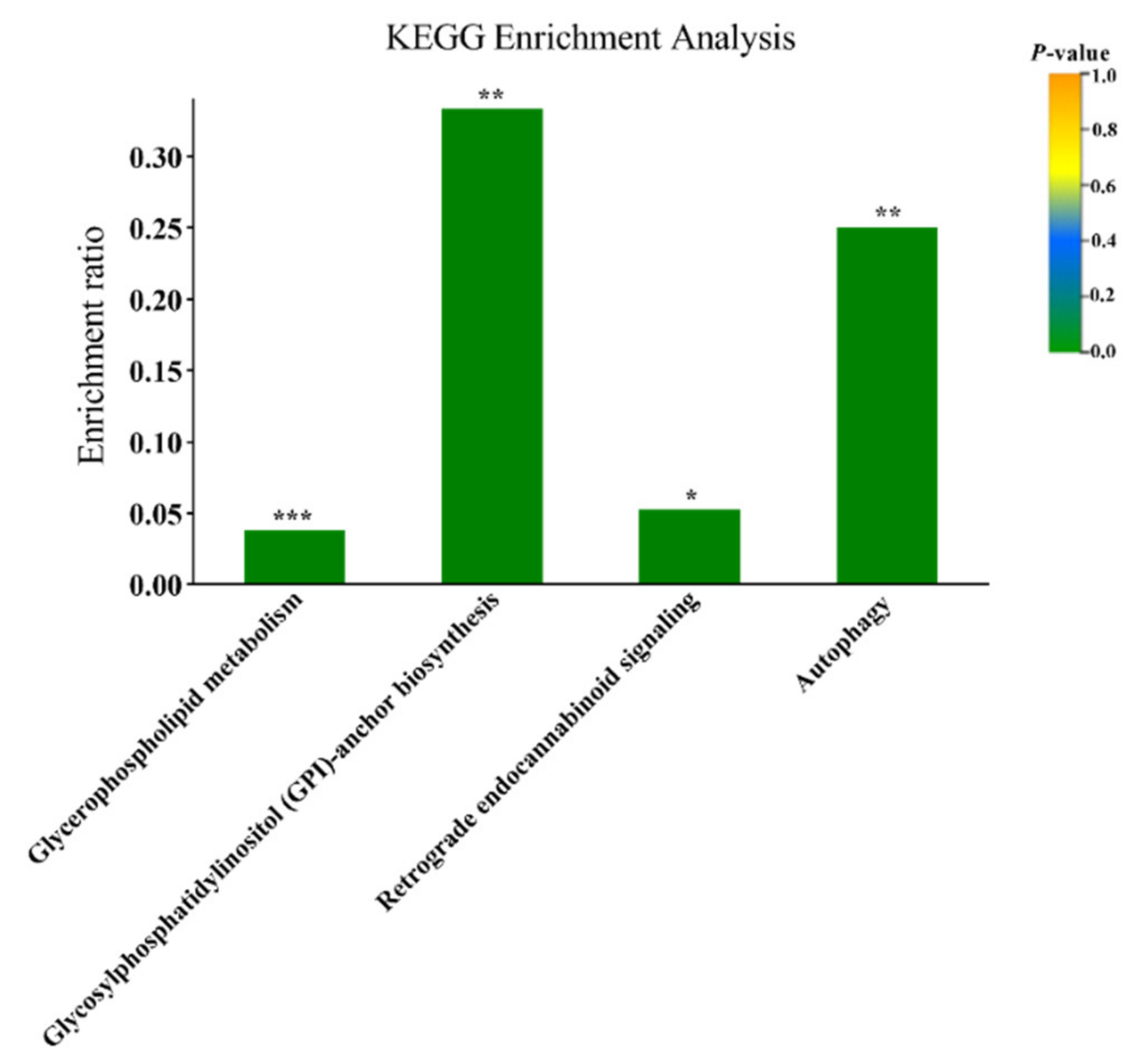
3.1. Plasma Metabolomics Profiling
3.2. Classical Metabolic Parameters
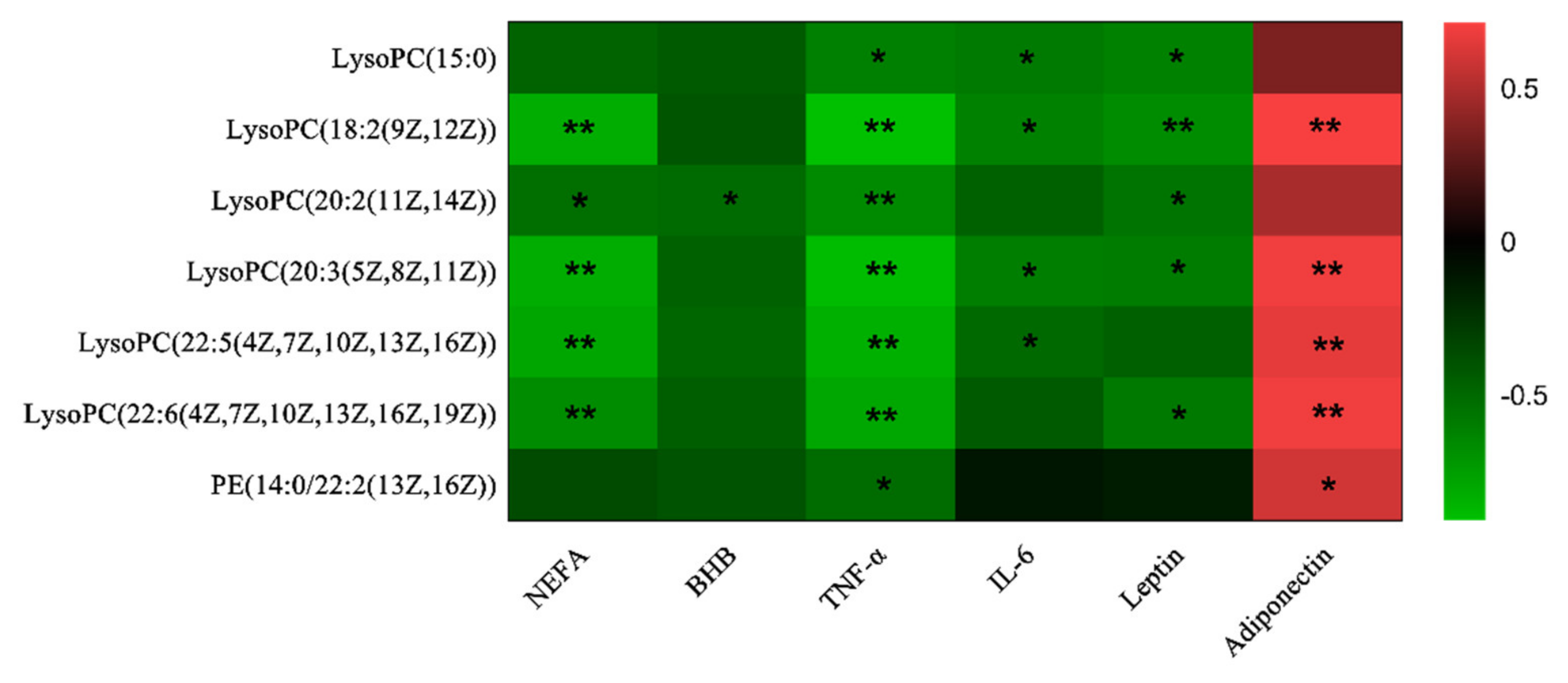
3.3. Correlations Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Roche, J.R.; Kay, J.K.; Friggens, N.C.; Loor, J.J.; Berry, D.P. Assessing and Managing Body Condition Score for the Prevention of Metabolic Disease in Dairy Cows. Veter-Clin. N. Am. Food Anim. Pr. 2013, 29, 323–336. [Google Scholar] [CrossRef]
- Edmonson, A.; Lean, I.; Weaver, L.; Farver, T.; Webster, G. A Body Condition Scoring Chart for Holstein Dairy Cows. J. Dairy Sci. 1989, 72, 68–78. [Google Scholar] [CrossRef]
- Roche, J.R.; Friggens, N.; Kay, J.; Fisher, M.; Stafford, K.; Berry, D.P. Invited review: Body condition score and its association with dairy cow productivity, health, and welfare. J. Dairy Sci. 2009, 92, 5769–5801. [Google Scholar] [CrossRef]
- Roche, J.R.; Macdonald, K.; Schütz, K.; Matthews, L.; Verkerk, G.; Meier, S.; Loor, J.J.; Rogers, A.; McGowan, J.; Morgan, S.; et al. Calving body condition score affects indicators of health in grazing dairy cows. J. Dairy Sci. 2013, 96, 5811–5825. [Google Scholar] [CrossRef] [PubMed]
- Matthews, L.; Cameron, C.; Sheahan, A.; Kolver, E.; Roche, J.R. Associations among dairy cow body condition and welfare-associated behavioral traits. J. Dairy Sci. 2012, 95, 2595–2601. [Google Scholar] [CrossRef] [PubMed]
- Ghaffari, M.H.; Jahanbekam, A.; Sadri, H.; Schuh, K.; Dusel, G.; Prehn, C.; Adamski, J.; Koch, C.; Sauerwein, H. Metabolomics meets machine learning: Longitudinal metabolite profiling in serum of normal versus overconditioned cows and pathway analysis. J. Dairy Sci. 2019, 102, 11561–11585. [Google Scholar] [CrossRef] [PubMed]
- Heimerl, S.; Fischer, M.; Baessler, A.; Liebisch, G.; Sigruener, A.; Wallner, S.; Schmitz, G. Alterations of Plasma Lysophosphatidylcholine Species in Obesity and Weight Loss. PLoS ONE 2014, 9, e111348. [Google Scholar] [CrossRef] [PubMed]
- Stevens, V.L.; Carter, B.D.; McCullough, M.L.; Campbell, P.T.; Wang, Y. Metabolomic Profiles Associated with BMI, Waist Circumference, and Diabetes and Inflammation Biomarkers in Women. Obesity 2019, 28, 187–196. [Google Scholar] [CrossRef]
- Forster, G.M.; Stockman, J.; Noyes, N.R.; Heuberger, A.L.; Broeckling, C.D.; Bantle, C.; Ryan, E.P. A Comparative Study of Serum Biochemistry, Metabolome and Microbiome Parameters of Clinically Healthy, Normal Weight, Overweight, and Obese Companion Dogs. Top. Companion Anim. Med. 2018, 33, 126–135. [Google Scholar] [CrossRef]
- Söder, J.; Höglund, K.; Dicksved, J.; Hagman, R.; Röhnisch, H.E.; Moazzami, A.A.; Wernersson, S. Plasma metabolomics reveals lower carnitine concentrations in overweight Labrador Retriever dogs. Acta Veter-Scand. 2019, 61, 10. [Google Scholar] [CrossRef]
- Pelantová, H.; Bugáňová, M.; Holubová, M.; Sediva, B.; Zemenová, J.; Sýkora, D.; Kavalkova, P.; Haluzík, M.; Železná, B.; Maletínská, L.; et al. Urinary metabolomic profiling in mice with diet-induced obesity and type 2 diabetes mellitus after treatment with metformin, vildagliptin and their combination. Mol. Cell. Endocrinol. 2016, 431, 88–100. [Google Scholar] [CrossRef] [PubMed]
- Barber, M.N.; Risis, S.; Yang, C.; Meikle, P.J.; Staples, M.; Febbraio, M.A.; Bruce, C.R. Plasma Lysophosphatidylcholine Levels Are Reduced in Obesity and Type 2 Diabetes. PLoS ONE 2012, 7, e41456. [Google Scholar] [CrossRef] [PubMed]
- Del Bas, J.M.; Caimari, A.; Rodriguez-Naranjo, M.I.; Childs, C.E.; Chavez, C.P.; West, A.L.; Miles, E.A.; Arola, L.; Calder, P.C. Impairment of lysophospholipid metabolism in obesity: Altered plasma profile and desensitization to the modulatory properties of n–3 polyunsaturated fatty acids in a randomized controlled trial. Am. J. Clin. Nutr. 2016, 104, 266–279. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Wu, H.; Liu, S.; Chai, S.; Meng, Q.; Zhou, Z. Dynamic Alterations in Yak Rumen Bacteria Community and Metabolome Characteristics in Response to Feed Type. Front. Microbiol. 2019, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Mansouryar, M.; Alamouti, H.M.; Banadaky, M.D.; Nielsen, M.O. Calving body condition score combined with milk test data and rectal tempreture improved the prognostic value of non-invasive markers for infectious diseases in holestein dairy cows. Livest. Sci. 2018, 212, 69–74. [Google Scholar] [CrossRef]
- Pires, J.A.; Delavaud, C.; Faulconnier, Y.; Pomiès, D.; Chilliard, Y. Effects of body condition score at calving on indicators of fat and protein mobilization of periparturient Holstein-Friesian cows. J. Dairy Sci. 2013, 96, 6423–6439. [Google Scholar] [CrossRef]
- Akbar, H.; Grala, T.; Riboni, M.V.; Cardoso, F.; Verkerk, G.; McGowan, J.; Macdonald, K.; Webster, J.; Schütz, K.; Meier, S.; et al. Body condition score at calving affects systemic and hepatic transcriptome indicators of inflammation and nutrient metabolism in grazing dairy cows. J. Dairy Sci. 2015, 98, 1019–1032. [Google Scholar] [CrossRef]
- Singhai, M.; Faizy, A.F.; Goyal, R.; Siddiqui, S.S. Evaluation of TNF-α and IL-6 levels in obese and non-obese diabetics: Pre- and postinsulin effects. N. Am. J. Med. Sci. 2012, 4, 180–184. [Google Scholar] [CrossRef]
- Alharthi, A.; Zhou, Z.; Lopreiato, V.; Trevisi, E.; Loor, J.J. Body condition score prior to parturition is associated with plasma and adipose tissue biomarkers of lipid metabolism and inflammation in Holstein cows. J. Anim. Sci. Biotechnol. 2018, 9, 12. [Google Scholar] [CrossRef]
- Galic, S.; Oakhill, J.S.; Steinberg, G.R. Adipose tissue as an endocrine organ. Mol. Cell. Endocrinol. 2010, 316, 129–139. [Google Scholar] [CrossRef]
- Febriza, A.; Ridwan, R.; As’Ad, S.; Kasim, V.N.; Idrus, H.H. Adiponectin and Its Role in Inflammatory Process of Obesity. Mol. Cell. Biomed. Sci. 2019, 3, 60–66. [Google Scholar] [CrossRef]
- Pietiläinen, K.H.; Sysi-Aho, M.; Rissanen, A.; Seppänen-Laakso, T.; Yki-Jarvinen, H.; Kaprio, J.; Orešič, M. Acquired Obesity Is Associated with Changes in the Serum Lipidomic Profile Independent of Genetic Effects—A Monozygotic Twin Study. PLoS ONE 2007, 2, e218. [Google Scholar] [CrossRef]
- Alarcon-Aguilar, F.; Almanza-Perez, J.; Blancas, G.; Angeles, S.; García-Macedo, R.; Román, R.; Cruz, M. Glycine regulates the production of pro-inflammatory cytokines in lean and monosodium glutamate-obese mice. Eur. J. Pharmacol. 2008, 599, 152–158. [Google Scholar] [CrossRef]
- Rico, J.; Bandaru, V.V.R.; Dorskind, J.M.; Haughey, N.J.; McFadden, J.W. Plasma ceramides are elevated in overweight Holstein dairy cows experiencing greater lipolysis and insulin resistance during the transition from late pregnancy to early lactation. J. Dairy Sci. 2015, 98, 7757–7770. [Google Scholar] [CrossRef] [PubMed]
- Hishikawa, D.; Hashidate, T.; Shimizu, T.; Shindou, H. Diversity and function of membrane glycerophospholipids generated by the remodeling pathway in mammalian cells. J. Lipid Res. 2014, 55, 799–807. [Google Scholar] [CrossRef]
- Manthei, K.A.; Ahn, J.; Glukhova, A.; Yuan, W.; Larkin, C.; Manett, T.D.; Chang, L.; Shayman, J.A.; Axley, M.J.; Schwendeman, A.; et al. A retractable lid in lecithin: Cholesterol acyltransferase provides a structural mechanism for activation by apolipoprotein A-I. J. Biol. Chem. 2017, 292, 20313–20327. [Google Scholar] [CrossRef] [PubMed]
- Wepy, J.A.; Galligan, J.; Kingsley, P.J.; Xu, S.; Goodman, M.C.; Tallman, K.A.; Rouzer, C.A.; Marnett, L.J. Lysophospholipases cooperate to mediate lipid homeostasis and lysophospholipid signaling. J. Lipid Res. 2018, 60, 360–374. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Zhu, W.; Chen, C.; Yan, B.; Zhu, L.; Chen, X.; Peng, C. The mechanisms of lysophosphatidylcholine in the development of diseases. Life Sci. 2020, 247, 117443. [Google Scholar] [CrossRef] [PubMed]
- Sevastou, I.; Kaffe, E.; Mouratis, M.-A.; Aidinis, V. Lysoglycerophospholipids in chronic inflammatory disorders: The PLA2/LPC and ATX/LPA axes. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2013, 1831, 42–60. [Google Scholar] [CrossRef]
- Hung, N.D.; Sok, D.-E.; Kim, M.R. Prevention of 1-palmitoyl lysophosphatidylcholine-induced inflammation by polyunsaturated acyl lysophosphatidylcholine. Inflamm. Res. 2012, 61, 473–483. [Google Scholar] [CrossRef]
- Jiang, X.-C. Lipid Homeostasis on Cell Membrane. In Advances in Membrane Proteins; Springer: Berlin/Heidelberg, Germany, 2019; pp. 1–30. [Google Scholar]
- Matias, I.; Gatta-Cherifi, B.; Cota, D. Obesity and the Endocannabinoid System: Circulating Endocannabinoids and Obesity. Curr. Obes. Rep. 2012, 1, 229–235. [Google Scholar] [CrossRef]
- Kovsan, J.; Blüher, M.; Tarnovscki, T.; Klöting, N.; Kirshtein, B.; Madar, L.; Shai, I.; Golan, R.; Harman-Boehm, I.; Schön, M.P.; et al. Altered Autophagy in Human Adipose Tissues in Obesity. J. Clin. Endocrinol. Metab. 2011, 96, E268–E277. [Google Scholar] [CrossRef]
- Du, X.; Liu, G.; Loor, J.J.; Fang, Z.; Bucktrout, R.; Yang, Y.; Ye, Q.; Shi, Z.; Shen, T.; Wang, X.; et al. Impaired hepatic autophagic activity in dairy cows with severe fatty liver is associated with inflammation and reduced liver function. J. Dairy Sci. 2018, 101, 11175–11185. [Google Scholar] [CrossRef]
- Abounit, K.; Scarabelli, T.M.; McCauley, R.B. Autophagy in mammalian cells. World J. Biol. Chem. 2012, 3, 1–6. [Google Scholar] [CrossRef]
- Singh, R.; Kaushik, S.; Wang, Y.; Xiang, Y.; Novak, I.; Komatsu, M.; Tanaka, K.; Cuervo, A.M.; Czaja, M.J. Autophagy regulates lipid metabolism. Nature 2009, 458, 1131–1135. [Google Scholar] [CrossRef] [PubMed]
- Koga, H.; Kaushik, S.; Cuervo, A.M. Altered lipid content inhibits autophagic vesicular fusion. FASEB J. 2010, 24, 3052–3065. [Google Scholar] [CrossRef] [PubMed]
- Ghaffari, M.; Schuh, K.; Dusel, G.; Frieten, D.; Koch, C.; Prehn, C.; Adamski, J.; Sauerwein, H.; Sadri, H. Mammalian target of rapamycin signaling and ubiquitin-proteasome-related gene expression in skeletal muscle of dairy cows with high or normal body condition score around calving. J. Dairy Sci. 2019, 102, 11544–11560. [Google Scholar] [CrossRef] [PubMed]
- Yamahara, K.; Kume, S.; Koya, D.; Tanaka, Y.; Morita, Y.; Chin-Kanasaki, M.; Araki, H.; Isshiki, K.; Araki, S.-I.; Haneda, M.; et al. Obesity-Mediated Autophagy Insufficiency Exacerbates Proteinuria-induced Tubulointerstitial Lesions. J. Am. Soc. Nephrol. 2013, 24, 1769–1781. [Google Scholar] [CrossRef]
- Mathieu, P.; Leeanna, E.-H.; Arnim, P. mTOR pathways in cancer and autophagy. Cancers 2018, 10, 18. [Google Scholar]
- Gatta-Cherifi, B.; Cota, D. New insights on the role of the endocannabinoid system in the regulation of energy balance. Int. J. Obes. 2015, 40, 210–219. [Google Scholar] [CrossRef]
- Bermúdez-Silva, F.-J.; Viveros, M.-P.; McPartland, J.; De Fonseca, F.R. The endocannabinoid system, eating behavior and energy homeostasis: The end or a new beginning? Pharmacol. Biochem. Behav. 2010, 95, 375–382. [Google Scholar] [CrossRef]
- Wang, J.; Ueda, N. Biology of endocannabinoid synthesis system. Prostaglandins Other Lipid Mediat. 2009, 89, 112–119. [Google Scholar] [CrossRef] [PubMed]
- Fu, J.; Gaetani, S.; Oveisi, F.; Verme, J.L.; Serrano, A.; De Fonseca, F.R.; Rosengarth, A.; Luecke, H.; Di Giacomo, B.; Tarzia, G.; et al. Oleylethanolamide regulates feeding and body weight through activation of the nuclear receptor PPAR-α. Nature 2003, 425, 90–93. [Google Scholar] [CrossRef] [PubMed]
- Tsuboi, K.; Uyama, T.; Okamoto, Y.; Ueda, N. Endocannabinoids and related N-acylethanolamines: Biological activities and metabolism. Inflamm. Regen. 2018, 38, 28. [Google Scholar] [CrossRef]
| Items | Normal Body Condition Score (n = 8) | High Body Condition Score (n = 8) | SEM | p-Value |
|---|---|---|---|---|
| Nonesterified fatty acid, nmol/mL | 162.6 | 171.0 | 12.77 | 0.52 |
| β-hydroxybutyrate, µg/mL | 9.97 | 11.56 | 0.916 | 0.10 |
| Leptin, pg/mL | 1840.4 | 2078.4 | 116.5 | 0.06 |
| Adiponectin, ng/mL | 10.02 | 8.67 | 0.757 | 0.09 |
| Tumor necrosis factor –α, pg/mL | 85.2 | 99.4 | 7.58 | 0.08 |
| Interleukin 6, pg/mL | 94.6 | 104.0 | 7.12 | 0.21 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, J.; Zhang, C.; Zhao, Q.; Li, C.; Jin, S.; Gu, X. Metabolic Profiling of Plasma in Different Calving Body Condition Score Cows Using an Untargeted Liquid Chromatography-Mass Spectrometry Metabolomics Approach. Animals 2020, 10, 1709. https://doi.org/10.3390/ani10091709
Wang J, Zhang C, Zhao Q, Li C, Jin S, Gu X. Metabolic Profiling of Plasma in Different Calving Body Condition Score Cows Using an Untargeted Liquid Chromatography-Mass Spectrometry Metabolomics Approach. Animals. 2020; 10(9):1709. https://doi.org/10.3390/ani10091709
Chicago/Turabian StyleWang, Jian, Chuang Zhang, Qingyao Zhao, Congcong Li, Shuang Jin, and Xianhong Gu. 2020. "Metabolic Profiling of Plasma in Different Calving Body Condition Score Cows Using an Untargeted Liquid Chromatography-Mass Spectrometry Metabolomics Approach" Animals 10, no. 9: 1709. https://doi.org/10.3390/ani10091709
APA StyleWang, J., Zhang, C., Zhao, Q., Li, C., Jin, S., & Gu, X. (2020). Metabolic Profiling of Plasma in Different Calving Body Condition Score Cows Using an Untargeted Liquid Chromatography-Mass Spectrometry Metabolomics Approach. Animals, 10(9), 1709. https://doi.org/10.3390/ani10091709





