Effect of Transmission and Vaccination on Time to Dominance of Emerging Viral Strains: A Simulation-Based Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Stochastic Multi-Strain SIR Model with Cross-Immunity
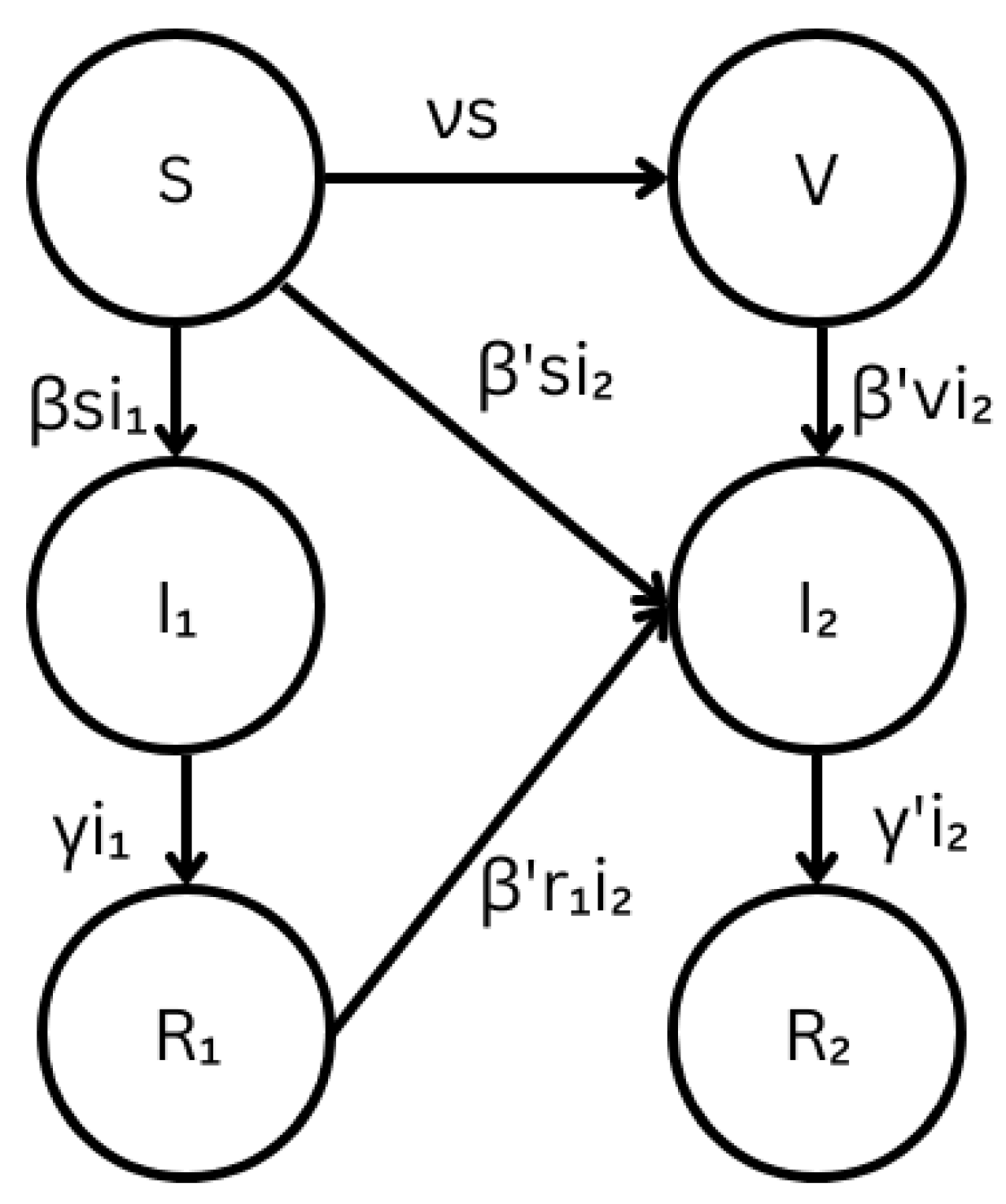
2.1.1. Modified Multi-Strain SIR Model
2.1.2. Continuous-Time Markov Chain Multi-Strain SIR Model
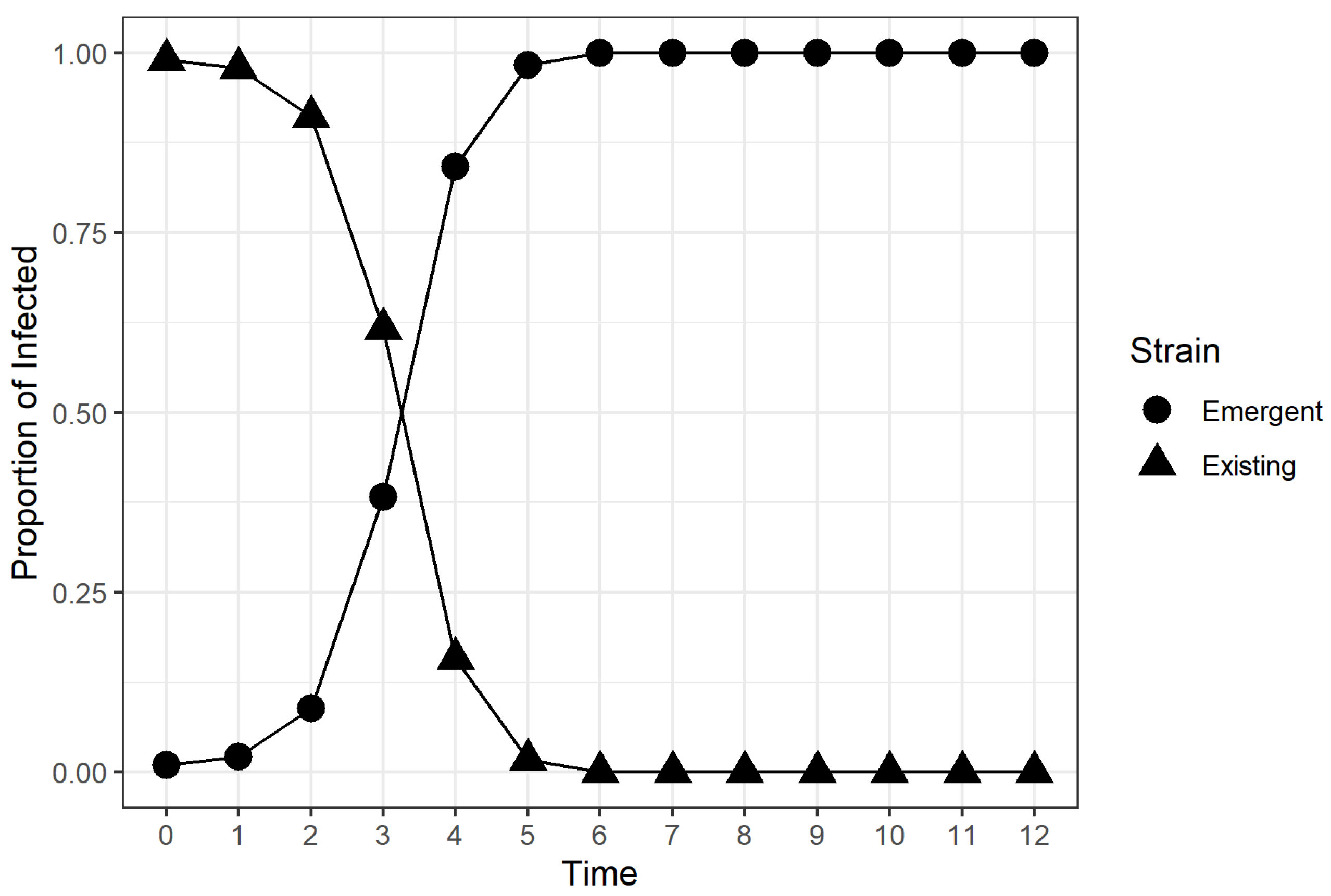
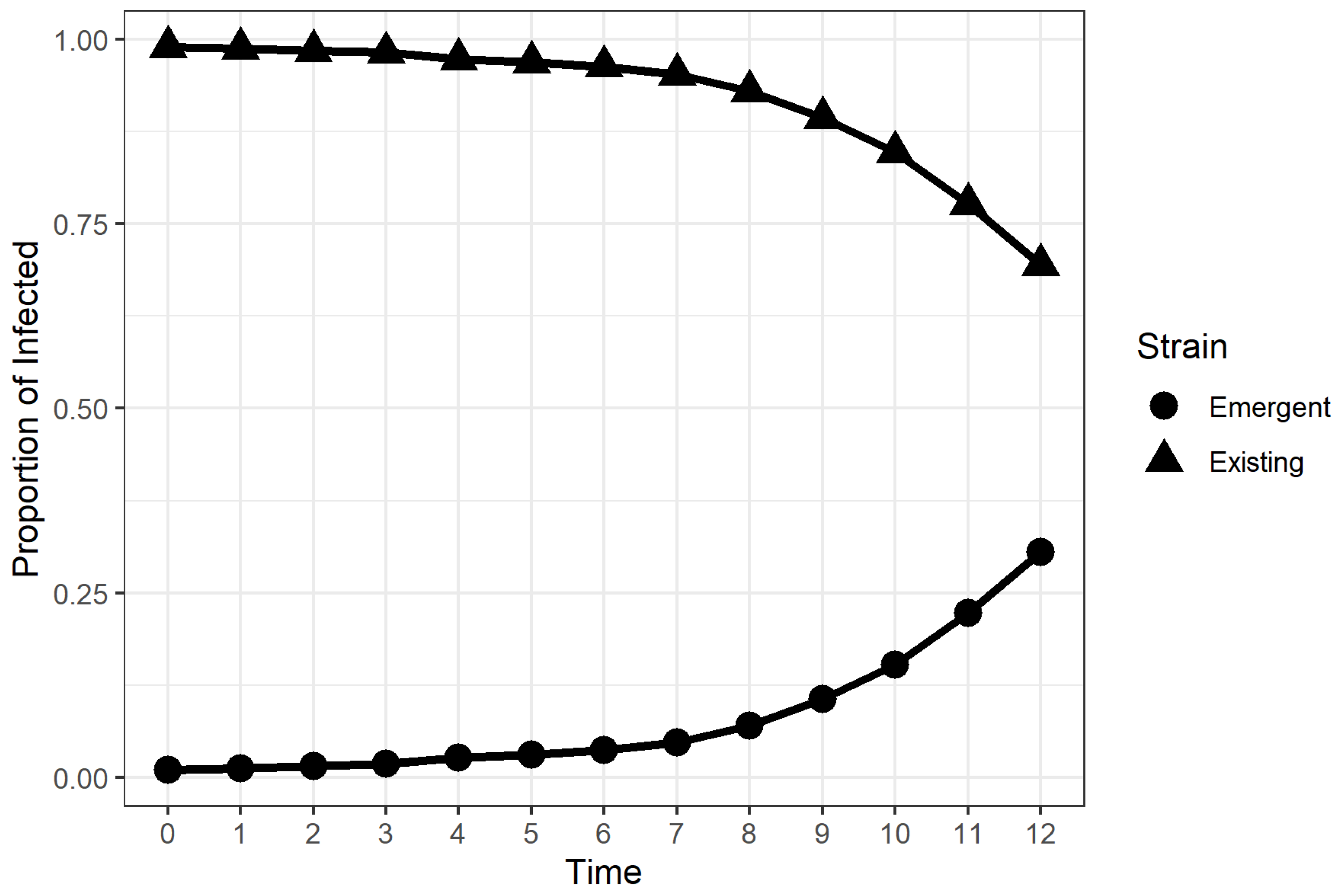
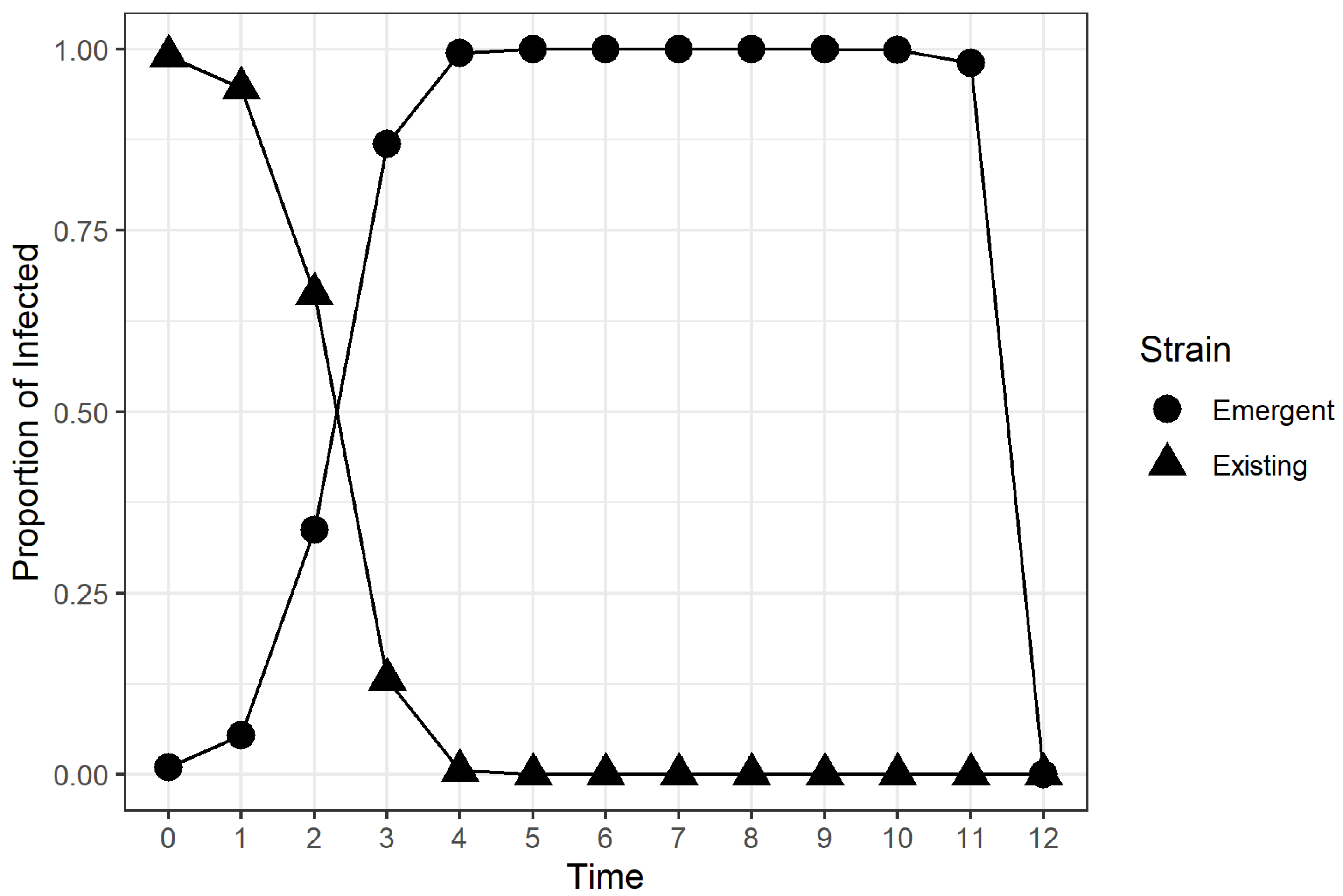
2.2. Time to Dominance (TTD)
2.3. Simulations and Factorial Experiment
3. Results
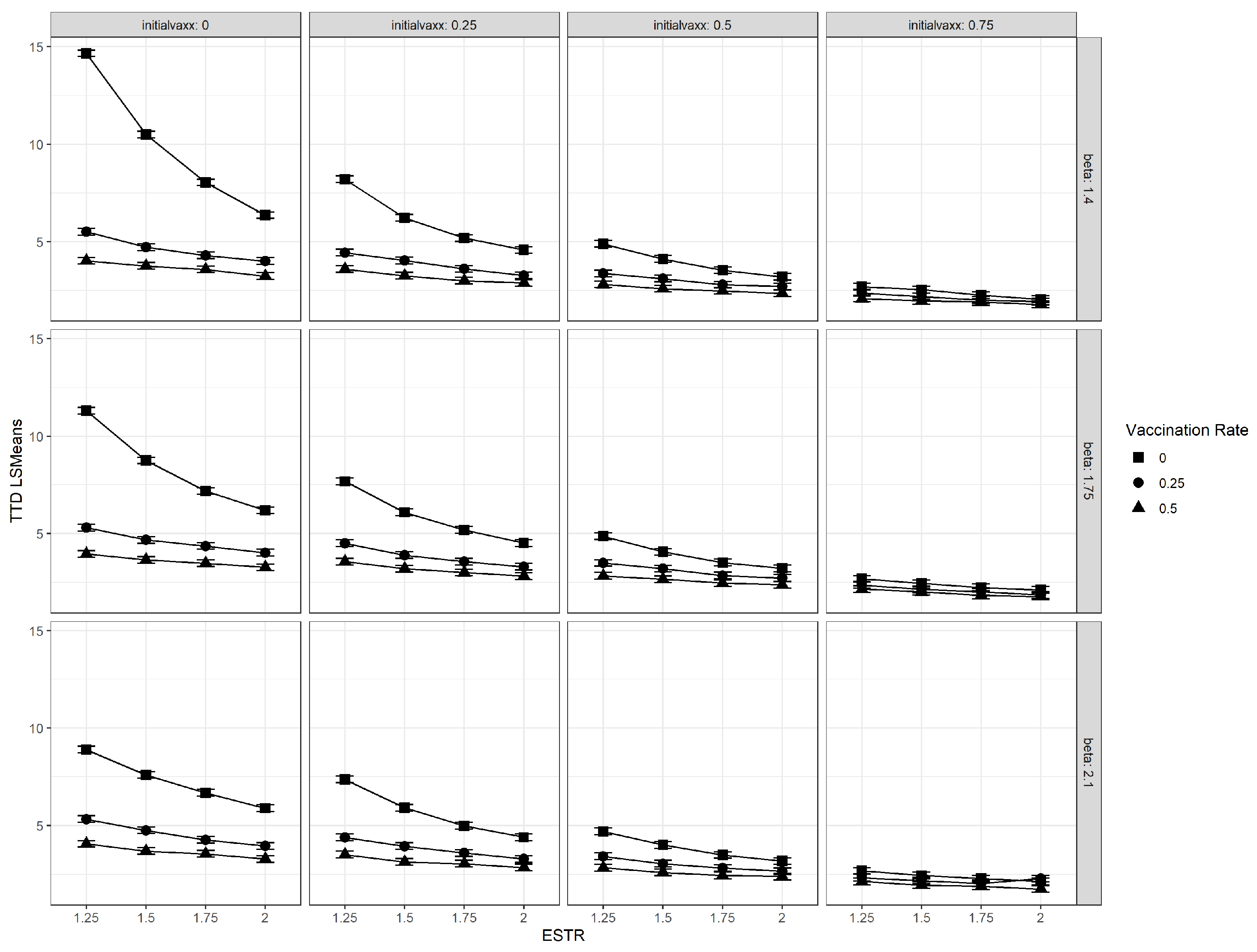
3.1. Interaction Plots
3.2. Vaccination Rate
3.3. Initial Vaccination Coverage
3.4. Relative Transmissibility of Emergent Strain (ESTR)
4. Discussion
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| SIR Model | Susceptible-Infected-Recovered Model |
| TTD | Time to Dominance |
| ESTR | Emergent Strain Transmission Ratio |
Appendix A
| Coefficient | Definition |
|---|---|
| s | Proportion of susceptible individuals |
| v | Proportion of vaccinated individuals |
| Proportion of individuals infected by the existing strain | |
| Proportion of individuals infected by the emergent strain | |
| Proportion of individuals that recovered from the existing strain | |
| Proportion of individuals that recovered from the emergent strain | |
| Transmission coefficient of the existing strain | |
| Transmission coefficient of the emergent strain | |
| Recovery coefficient of the existing strain | |
| Recovery coefficient of the existing strain | |
| Vaccination coefficient |
References
- Hui, E.K.-W. Reasons for the increase in emerging and re-emerging viral infectious diseases. Microbes Infect. 2006, 8, 905–916. [Google Scholar] [CrossRef]
- Howard, C.R.; Fletcher, N.F. Emerging virus diseases: Can we ever expect the unexpected? Emerg. Microbes Infect. 2012, 1, e46. [Google Scholar] [CrossRef]
- Pekosz, A.; Glass, G.E. Emerging Viral Diseases. Md. Med. 2008, 9, 11–16. [Google Scholar]
- Nichol, S.T.; Arikawa, J.; Kawaoka, Y. Emerging viral diseases. Proc. Natl. Acad. Sci. USA 2000, 97, 12411–12412. [Google Scholar] [CrossRef]
- Marston, H.D.; Folkers, G.K.; Morens, D.M.; Fauci, A.S. Emerging Viral Diseases: Confronting Threats with New Technologies. Sci. Transl. Med. 2014, 6, 253ps10. [Google Scholar] [CrossRef]
- Burt, F.J.; Rolph, M.S.; Rulli, N.E.; Mahalingam, S.; Heise, M.T. Chikungunya: A re-emerging virus. Lancet 2012, 379, 662–671. [Google Scholar] [CrossRef]
- Acosta-Ampudia, Y.; Monsalve, D.M.; Rodríguez, Y.; Pacheco, Y.; Anaya, J.M.; Ramírez-Santana, C. Mayaro: An emerging viral threat? Emerg. Microbes Infect. 2018, 7, 163. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. SARS-CoV-2 Variant Classifications and Definitions. Available online: https://www.cdc.gov/coronavirus/2019-ncov/variants/variant-classifications.html#anchor_1632154493691 (accessed on 1 February 2023).
- Khyar, O.; Allali, K. Global dynamics of a multi-strain SEIR epidemic model with general incidence rates: Application to COVID-19 pandemic. Nonlinear Dyn. 2020, 102, 489–509. [Google Scholar] [CrossRef]
- Fudolig, M.; Howard, R. The local stability of a modified multi-strain SIR model for emerging viral strains. PLoS ONE 2020, 15, e0243408. [Google Scholar] [CrossRef]
- Otunuga, O.M. Analysis of multi-strain infection of vaccinated and recovered population through epidemic model: Application to COVID-19. PLoS ONE 2022, 17, e0271446. [Google Scholar] [CrossRef]
- Massard, M.; Eftimie, R.; Perasso, A.; Saussereau, B. A multi-strain epidemic model for COVID-19 with infected and asymptomatic cases: Application to French data. J. Theor. Biol. 2022, 545, 111117. [Google Scholar] [CrossRef]
- Li, C.L.; Cheng, C.Y.; Li, C.H. Global dynamics of two-strain epidemic model with single-strain vaccination in complex networks. Nonlinear Anal. Real World Appl. 2023, 69, 103738. [Google Scholar] [CrossRef]
- Chang, Y.C.; Liu, C.T. A Stochastic Multi-Strain SIR Model with Two-Dose Vaccination Rate. Mathematics 2022, 10, 1804. [Google Scholar] [CrossRef]
- Mu, X.; Zhang, Q. Near-optimal control for a stochastic multi-strain epidemic model with age structure and Markovian switching. Int. J. Control 2022, 95, 1191–1205. [Google Scholar] [CrossRef]
- World Health Organization. Tracking SARS-CoV-2 variants. Available online: https://www.who.int/activities/tracking-SARS-CoV-2-variants (accessed on 5 February 2023).
- Chauhan, S.; Misra, O.P.; Dhar, J. Stability analysis of SIR model with vaccination. Am. J. Comput. Appl. Math. 2014, 4, 17–23. [Google Scholar]
- d’Onofrio, A.; Manfredi, P.; Salinelli, E. Vaccinating behaviour, information, and the dynamics of SIR vaccine preventable diseases. Theor. Popul. Biol. 2007, 71, 301–317. [Google Scholar] [CrossRef]
- Zaman, G.; Kang, Y.H.; Jung, I.H. Stability analysis and optimal vaccination of an SIR epidemic model. BioSystems 2008, 93, 240–249. [Google Scholar] [CrossRef]
- Kar, T.K.; Batabyal, A. Stability analysis and optimal control of an SIR epidemic model with vaccination. Biosystems 2011, 104, 127–135. [Google Scholar] [CrossRef]
- Buonomo, B.; d’Onofrio, A.; Lacitignola, D. Global stability of an SIR epidemic model with information dependent vaccination. Math. Biosci. 2008, 216, 9–16. [Google Scholar] [CrossRef]
- Ehrhardt, M.; Gašper, J.; Kilianová, S. SIR-based mathematical modeling of infectious diseases with vaccination and waning immunity. J. Comput. Sci. 2019, 37, 101027. [Google Scholar] [CrossRef]
- Stone, L.; Shulgin, B.; Agur, Z. Theoretical examination of the pulse vaccination policy in the SIR epidemic model. Math. Comput. Model. 2000, 31, 207–215. [Google Scholar] [CrossRef]
- Shulgin, B.; Stone, L.; Agur, Z. Pulse vaccination strategy in the SIR epidemic model. Bull. Math. Biol. 1998, 60, 1123–1148. [Google Scholar] [CrossRef]
- Meng, X.; Chen, L. The dynamics of a new SIR epidemic model concerning pulse vaccination strategy. Appl. Math. Comput. 2008, 197, 582–597. [Google Scholar] [CrossRef]
- Bretó, C.; He, D.; Ionides, E.L.; King, A.A. Time series analysis via mechanistic models. Ann. Appl. Stat. 2009, 3, 319–348. [Google Scholar] [CrossRef]
- Allen, L.J. An introduction to stochastic epidemic models. In Mathematical Epidemiology; Springer: Berlin/Heidelberg, Germany, 2008; pp. 81–130. [Google Scholar]
- Zhou, Y.; Zhang, W. Threshold of a stochastic SIR epidemic model with Lévy jumps. Phys. A Stat. Mech. Its Appl. 2016, 446, 204–216. [Google Scholar] [CrossRef]
- Maki, Y.; Hirose, H. Infectious disease spread analysis using stochastic differential equations for SIR model. In Proceedings of the 2013 4th International Conference on Intelligent Systems, Modelling and Simulation, Bangkok, Thailand, 29–31 January 2013; pp. 152–156. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2022. [Google Scholar]
- Bolze, A.; Basler, T.; White, S.; Dei Rossi, A.; Wyman, D.; Dai, H.; Roychoudhury, P.; Greninger, A.L.; Hayashibara, K.; Beatty, M.; et al. Evidence for SARS-CoV-2 Delta and Omicron co-infections and recombination. Med 2022, 3, 848–859.e4. [Google Scholar] [CrossRef]
- Wertheim, J.O.; Wang, J.C.; Leelawong, M.; Martin, D.P.; Havens, J.L.; Chowdhury, M.A.; Pekar, J.E.; Amin, H.; Arroyo, A.; Awandare, G.A.; et al. Detection of SARS-CoV-2 intra-host recombination during superinfection with Alpha and Epsilon variants in New York City. Nat. Commun. 2022, 13, 3645. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Gong, Y.; Zhang, C.; Sun, J.; Wong, G.; Shi, W.; Liu, W.; Gao, G.F.; Bi, Y. Co-existence and co-infection of influenza A viruses and coronaviruses: Public health challenges. Innovation 2022, 3, 100306. [Google Scholar] [CrossRef]
- Pérez-García, F.; Vásquez, V.; de Egea, V.; Catalán, P.; Rodríguez-Sánchez, B.; Bouza, E. Influenza A and B co-infection: A case–control study and review of the literature. Eur. J. Clin. Microbiol. Infect. Dis. 2016, 35, 941–946. [Google Scholar] [CrossRef]
- Cai, X.; Xu, Z. K-leap method for accelerating stochastic simulation of coupled chemical reactions. J. Chem. Phys. 2007, 126, 074102. [Google Scholar] [CrossRef]
- Tian, T.; Burrage, K. Binomial leap methods for simulating stochastic chemical kinetics. J. Chem. Phys. 2004, 121, 10356–10364. [Google Scholar] [CrossRef] [PubMed]
- King, A.A.; Ionides, E.L. Simulation of Stochastic Dynamic Models. Available online: https://kingaa.github.io/short-course/stochsim/stochsim.html#the-simple-continuous-time-markov-chain-version-of-the-sir-model (accessed on 1 February 2023).
- UK Health Security Agency. Coronavirus (COVID-19) in the UK. Available online: https://coronavirus.data.gov.uk/ (accessed on 1 February 2023).
- Centers for Disease Control and Prevention. COVID-19 Vaccinations in the United States. Available online: https://covid.cdc.gov/covid-data-tracker/#vaccinations_vacc-people-booster-percent-total (accessed on 18 March 2023).
- Centers for Disease Control and Prevention. Weekly Flu Vaccination Dashboard. Available online: https://www.cdc.gov/flu/fluvaxview/dashboard/vaccination-dashboard.html (accessed on 18 March 2023).
- Lenth, R.V. emmeans: Estimated Marginal Means, aka Least-Squares Means; R Package Version 1.8.4-1; R Foundation: Vienna, Austria, 2023. [Google Scholar]
- Bowick, G.C.; McAuley, A.J. Vaccine and adjuvant design for emerging viruses: Mutations, deletions, segments and signaling. Bioeng. Bugs 2011, 2, 129–135. [Google Scholar] [CrossRef]
- García-Sastre, A.; Mena, I. Novel vaccine strategies against emerging viruses. Curr. Opin. Virol. 2013, 3, 210–216. [Google Scholar] [CrossRef]
- Graham, R.L.; Donaldson, E.F.; Baric, R.S. A decade after SARS: Strategies for controlling emerging coronaviruses. Nat. Rev. Microbiol. 2013, 11, 836–848. [Google Scholar] [CrossRef]
- Agnihothram, S.; Gopal, R.; Yount, B.L., Jr.; Donaldson, E.F.; Menachery, V.D.; Graham, R.L.; Scobey, T.D.; Gralinski, L.E.; Denison, M.R.; Zambon, M. Evaluation of serologic and antigenic relationships between middle eastern respiratory syndrome coronavirus and other coronaviruses to develop vaccine platforms for the rapid response to emerging coronaviruses. J. Infect. Dis. 2014, 209, 995–1006. [Google Scholar] [CrossRef] [PubMed]
- Tse, L.V.; Meganck, R.M.; Graham, R.L.; Baric, R.S. The Current and Future State of Vaccines, Antivirals and Gene Therapies Against Emerging Coronaviruses. Front. Microbiol. 2020, 11, 658. [Google Scholar] [CrossRef] [PubMed]
| Factor | Simulation Parameter | Values |
|---|---|---|
| Transmission coefficient of existing strain | 1.4, 1.75, 2.1 | |
| Emergent strain transmission ratio (ESTR) | 1.25, 1.5, 1.75, 2 | |
| Vaccination coefficient | 0, 0.25, 0.5 | |
| Initial vaccination coverage | 0, 0.25, 0.5, 0.75 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fudolig, M. Effect of Transmission and Vaccination on Time to Dominance of Emerging Viral Strains: A Simulation-Based Study. Microorganisms 2023, 11, 860. https://doi.org/10.3390/microorganisms11040860
Fudolig M. Effect of Transmission and Vaccination on Time to Dominance of Emerging Viral Strains: A Simulation-Based Study. Microorganisms. 2023; 11(4):860. https://doi.org/10.3390/microorganisms11040860
Chicago/Turabian StyleFudolig, Miguel. 2023. "Effect of Transmission and Vaccination on Time to Dominance of Emerging Viral Strains: A Simulation-Based Study" Microorganisms 11, no. 4: 860. https://doi.org/10.3390/microorganisms11040860
APA StyleFudolig, M. (2023). Effect of Transmission and Vaccination on Time to Dominance of Emerging Viral Strains: A Simulation-Based Study. Microorganisms, 11(4), 860. https://doi.org/10.3390/microorganisms11040860






