Abstract
Background: Aeromonas hydrophila is a widely recognized broad-spectrum pathogen that primarily targets the gastrointestinal tract. Type IV pili (T4P) are proteinaceous nano-machines located on the bacterial cell surface, playing a crucial role in host colonization and infection. Regrettably, the T4P systems of A. hydrophila remain largely underexplored. Methods: A. hydrophila genomes with complete genome assembly and annotation reports up to 31 March 2023, were obtained from the NCBI Genome database or KEGG genome database, followed by a global search for T4P secretion system genes. Protein sequences of these manually curetted genes were used as secondary quarry for Synteny analysis. Protein–protein interaction analysis was performed by string analysis and in silico study of genomic islands. Results: We identified 27 orthologs of type IV pili (T4P) nano-machine components in A. hydrophila. These orthologs are primarily distributed across three operons: pilABCD, pilMNOPQ, and pilVWXY. While the first two operons are commonly found in all experimental genomes, the presence of the pilVWXY operon, coding for 11 orthologs, is reported here for the first time in A. hydrophila. Notably, the complete pilVWXY operon is absent in nonvirulent strains. A genomic islands study between a nonvirulent and hypervirulent strain also confirms absence of most of the genes coded by pilVWXY in nonvirulent strain. Interestingly, among the 51 experimental genomes analyzed, the pilVWXY operon was completely absent in 10 strains, most of which are categorized as nonvirulent; Conclusions: The distribution of two major type IV pili (T4P) nano-machines, PilABCDMNOPQ and PilVWXY, is reported here for the first time in A. hydrophila. Additionally, this study suggests a potential role for the PilVWXY nano-machine in establishing human disease.
1. Introduction
Aeromonas hydrophila is a ubiquitous, Gram-negative, facultative anaerobic bacterium commonly found in water sources, with a broad host range spanning protozoa [1], mollusks [2], arthropods [3], fish [4], amphibians [5], reptiles [6], birds [7], and mammals [8]. It is known to cause zoonotic infections, transmitted from diseased fish, contaminated water, and partially cooked seafood to humans [9]. These infections can manifest as septic arthritis, gastroenteritis with diarrhea, skin and soft tissue infections, meningitis, and bacteremia [10]. A. hydrophila poses a serious threat, particularly to children [11] and immunocompromised individuals [12].
Colonization within the intestinal mucosa is a multistep process facilitated by pili, proteinaceous filamentous appendages found on the surface of many bacteria responsible for host cell adherence. Type IV pili (T4P), a multimeric proteinaceous nano-machine, are expressed by a majority of Gram-negative pathogenic bacteria, including Vibrio cholera, enterotoxigenic Escherichia coli, and enteropathogenic E. coli. T4P play a critical role in adhesion [13], as they undergo force-driven contraction to enhance attachment to the target surface and facilitate colonization through rapid cycles of extension and retraction, a process known as twitching motility [14]. Additionally, T4P promote surface sensing, motility, biofilm formation, and DNA uptake. They also play a pivotal role in attachment to epithelial cells and GM-1 and GM-2 gangliosides in Pseudomonas aeruginosa [15], facilitating movement across solid surfaces without the involvement of flagella [16]. Mutant strains lacking functional T4P often display impaired attachment and colonization capabilities in P. aeruginosa [16,17].
Type IV pili (T4P) in Aeromonas are recognized as intestinal colonization factors associated with gastrointestinal infections and diarrhea [18]. Aeromonas species possess complete T4P biogenesis gene clusters, such as tapABCD, which are homologous to the T4P gene clusters (pilABCD) [19] found in Pseudomonas aeruginosa. Previous studies, such as the work by Boyd et al., have reported the presence of tapMNOPQ in A. salmonicida subsp. salmonicida [20]. However, limited information is available regarding T4P nano-machine components and their function in A. hydrophila. To address this gap, we conducted a comprehensive global search across 51 known strains of A. hydrophila using a novel protocol developed for this study. The nomenclature of T4P components or its orthologs across different bacterial genera is not uniform [21]. For example, in P. aeruginosa PilV is thought to be part of the tip complex, but in Neisseria meningitidis, PilV is an unrelated protein which is incorporated throughout the length of the pilus. Similarly, PilX is a putative tip adhesin in P. aeruginosa, homologous to PilK in N. meningitidis [21]. To resolve this problem, we used gene ortholog databases (KEGG-KO) to identify the orthologs of T4P components unanimously. To avoid any confusion related to the nomenclature of these orthologs, we use the nomenclature from Pseudomonas T4P throughout the manuscript. This protocol integrates KEGG-KO, genome, and proteome databases to investigate T4P components and their potential functions in A. hydrophila.
2. Materials and Methods
2.1. Data Mining
A. hydrophila genomes with complete genome assembly and annotation reports up to 31 March 2023, were obtained from the NCBI Genome database (https://www.ncbi.nlm.nih.gov/genome/browse/#!/prokaryotes/1422/) or from the KEGG genome database (https://www.genome.jp/dbget-bin/www_bfind_sub?mode=bfind&max_hit=1000&dbkey=genome&keywords=Aeromonas+hydrophila). This dataset comprises 51 genomes (refer to Table 1). Sequences of stress-responsive enzymes and stress regulators were retrieved from the UniProt database (https://www.uniprot.org/), while protein structures were obtained from the RCSB PDB database (https://www.rcsb.org/). All these databases were accessed on or before 31 March 2023.

Table 1.
A. hydrophila genomes with complete genome assembly and annotation report used for this study (accessed till 31 March 2023).
2.2. Searching for T4P Secretion System Genes
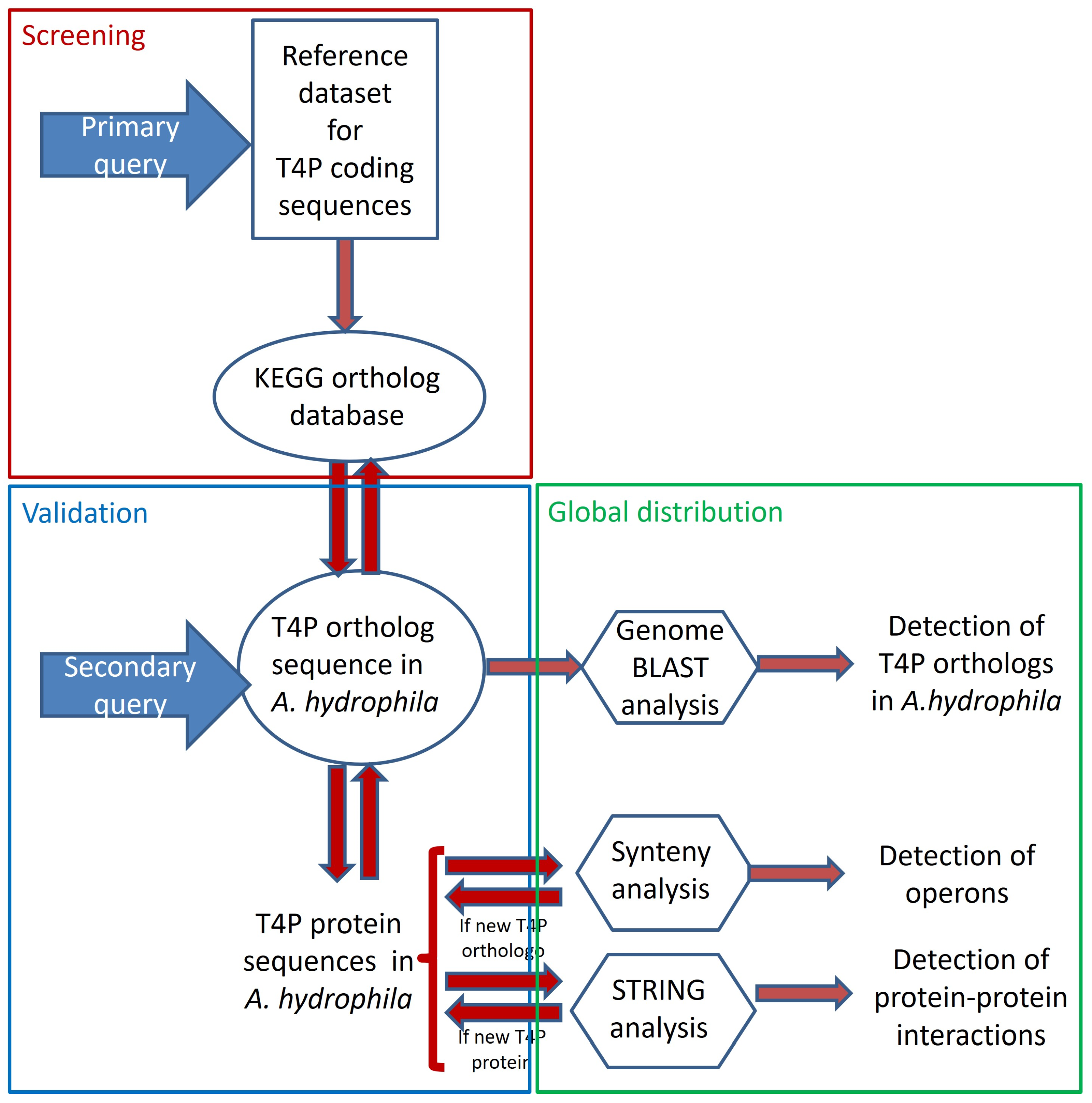
In this study, we developed a protocol aimed at maximizing the identification of candidate genes related to type IV pili (T4P) secretion systems (see Figure 1). Information regarding the components of T4P secretion systems was gathered from original data previously published by various researchers to create a reference dataset, which served as the primary resource (refer to Table S1). The protein sequences of these manually curated genes were then used as the secondary resource for synteny analysis, leading to the discovery of previously unidentified T4P components in A. hydrophila. The results were validated using the KEGG-KO database and further confirmed through STRING analysis (see Table S2). To investigate the distribution of these T4P components across the experimental genomes, a genome blast approach was employed (refer to Table S3).
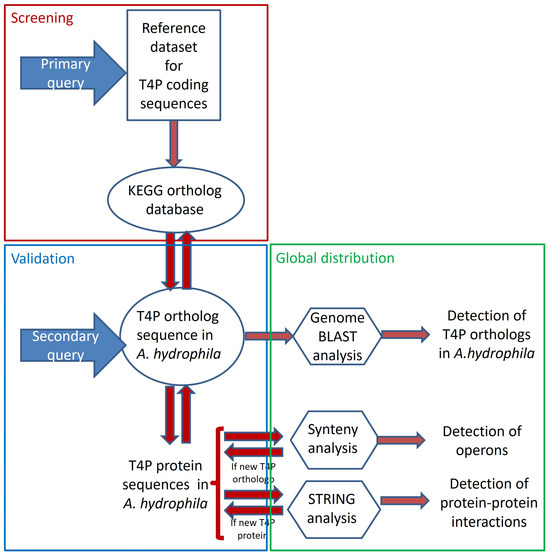
Figure 1.
Detailed experimental design and step-by-step workflow.
2.3. Synteny Analysis
Synteny analysis is crucial for examining the conservation of gene order, especially in bacterial secretion systems [22]. To investigate synteny among bacterial secretion systems across selected A. hydrophila genomes, we utilized the SyntTax server (https://archaea.i2bc.paris-saclay.fr/SyntTax/, accessed on 31 March 2023). According to its creators, SyntTax is a web service tailored to harness the vast array of prokaryotic genomes by connecting them through taxonomic relationships [23]. The functional methodology of SyntTax primarily relies on the Absynte algorithm [24].
2.4. String Analysis
STRING is one of the best platforms to construct and visualize protein interaction networks (PINs) [25]. The STRING v11.5 (http://string-db.org) (last accessed on 31 March 2023) server is used for predicting interactions among components of the T4P nano-machine. During STRING analysis, T4P components were aligned with A. hydrophila subsp. hydrophila ATCC 7966. The first aligned protein with an E-value below 1 × 10−10 was considered a homologous protein. Then, these homologous proteins and their corresponding interactions were extracted from the whole interaction dataset of the related organism to compose the model organism-based protein–protein interaction sub-network. To obtain high-quality PINs, we considered the interactions with the highest confidence limits (0.9) from STRING.
2.5. Comparing Genomic Islands
Genetic islands encode factors that enhance the adaptability and competitiveness of the bacteria within a niche, including virulence factors and other medically or environmentally important adaptations [26]. Here, we conducted a comparative analysis of genomes of two A. hydrophila strains using IslandViewer 4 (http://www.pathogenomics.sfu.ca/islandviewer/, accessed on 31 March 2023) to elucidate the presence and absence of specific genes.
2.6. Statistical Analyses
The max-score of all T4P secretion system-related genes was recorded and used to generate a data matrix, which was then utilized to create a heat map visualizing the diversity of T4P components across A. hydrophila genomes. The heat map was generated using an online tool called Morpheus (https://software.broadinstitute.org/morpheus/), last accessed on 31 March 2023. Additionally, the same data matrix was employed for multivariate analysis, including Principal Component Analysis (PCA) and cluster analysis (CA), using Past v4.13 software (https://www.nhm.uio.no/english/research/resources/past/, accessed on 31 March 2023). Binary scattered plots were presented following PCA analysis. Furthermore, a cluster dendrogram was constructed via PCA using the p-distance method of neighbor joining with 10,000 bootstraps. Lastly, Venn diagrams were created using an online tool provided by Bioinformatics & Evolutionary Genomics (available at: http://bioinformatics.psb.ugent.be/webtools/Venn/), last accessed on 31 March 2023.
3. Results
3.1. Searching for Components of A. hydrophila T4P System
A new protocol was developed for identifying components of the A. hydrophila type IV pili (T4P) system (refer to Figure 1). Initially, 19 query candidates for bacterial T4P secretion system components were identified based on experimental records (Table S1). Additionally, one additional paralog each for FimT, PilE, and PilZ was confirmed from the KEGG KO database (Table S2). The protein sequences of these 22 T4P components were used as a secondary query for synteny analysis across available annotated A. hydrophila genomes (Table S3).
During the synteny analysis, three new candidates, PilT, and two paralogs of PilU (totaling 27) were observed. These were initially verified using the KEGG KO database and then re-confirmed using protein–protein interaction network (PIN) analysis. For the reference strain A. hydrophila ATCC 7966, the locus tags for these three new components were identified as AHA_0694, AHA_0695, and AHA_0696 (Table S4). Ultimately, employing this protocol, 27 T4P components were identified in all A. hydrophila strains (Table 2).

Table 2.
Orthologs of T4P nano-machine components in A. hydrophila.
3.2. Synteny among A. hydrophila T4P Systems
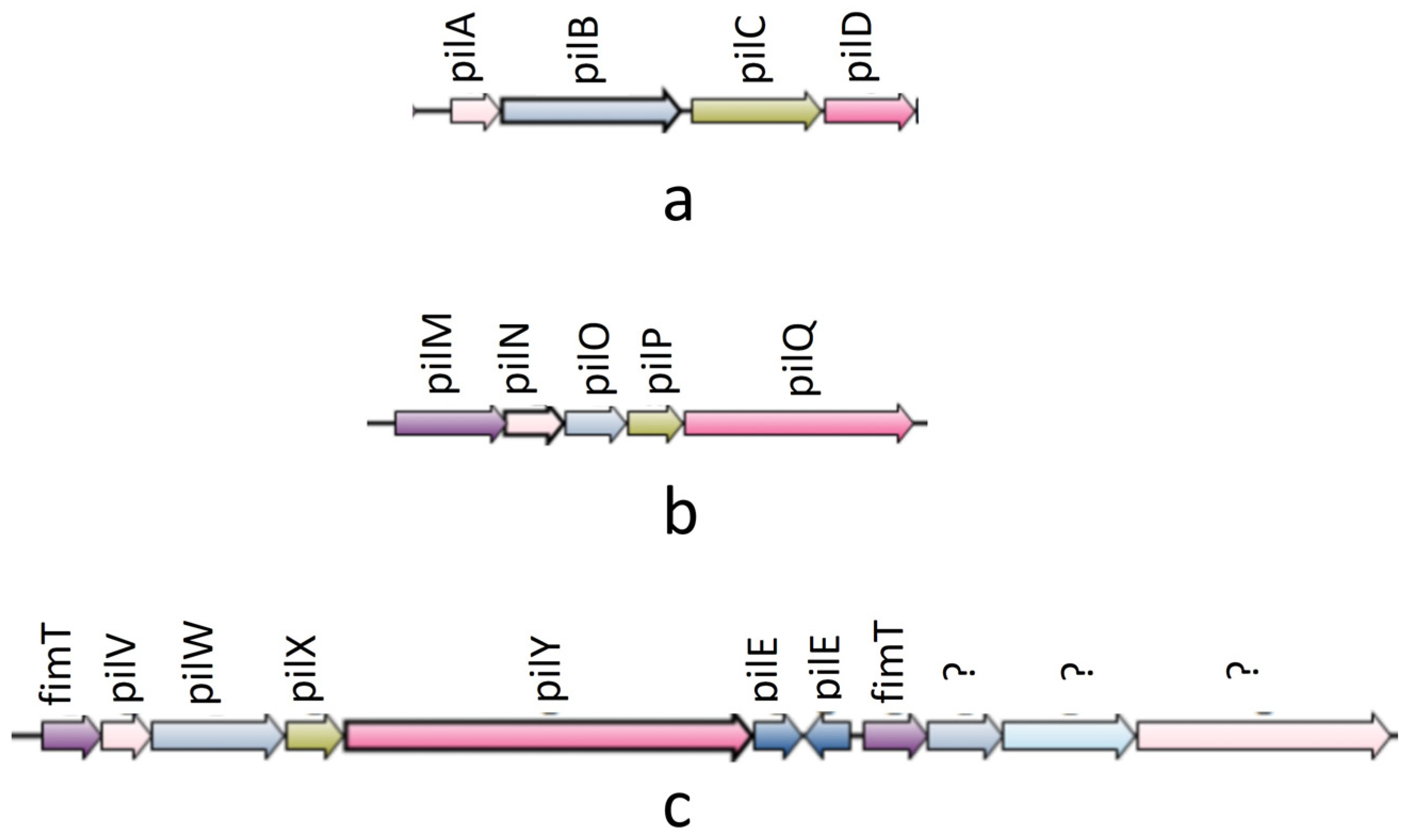
In the synteny analysis, 27 T4P components were predominantly found to be organized into three operons: pilABCD; pilMNOPQ; and pilVWXY (see Figure 2). The first two operons were identified in all experimental genomes. Notably, the pilVWXY operon emerged as a major one, encoding 11 T4P orthologs. Interestingly, among the 51 experimental genomes, synteny for the pilVWXY operon components was not observed in 10 genomes, namely Ah2111, AC185, WP7 S18 ESBL 06, AH10, CSUSB2, 71339, A008N2, MX16A, NEB7245, and 4AK4.
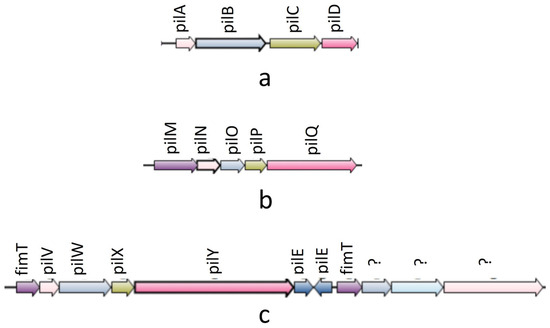
Figure 2.
Synteny diagram showing three operons of T4P nano-machine components: (a) pilABCD operon: synteny found in all genomes; (b) pilMNOPQ: synteny found in all genomes; (c) pilVWXY operon(‘?’ marks were put in case of un-identified ortholgs): synteny found in all genomes except Ah2111, AC185, WP7 S18 ESBL 06, AH10, CSUSB2, 71339, MX16A, NEB724, and 4AK4.
3.3. Protein–Protein Interaction Network Study
To validate our results, all 27 components (Table 2) underwent a protein–protein interaction (PIN) study within the A. hydrophila genome using STRING v11.5. In this analysis, T4P components were aligned with A. hydrophila subsp. hydrophila ATCC 7966. Proteins with an E-value below 1 × 10−10 were considered homologous proteins. Homologous proteins and their corresponding interactions were extracted from the entire interaction dataset of the related organism to construct the model organism-based protein–protein interaction sub-network. To ensure high-quality PINs, interactions with confidence limits above 0.9 were considered from STRING.
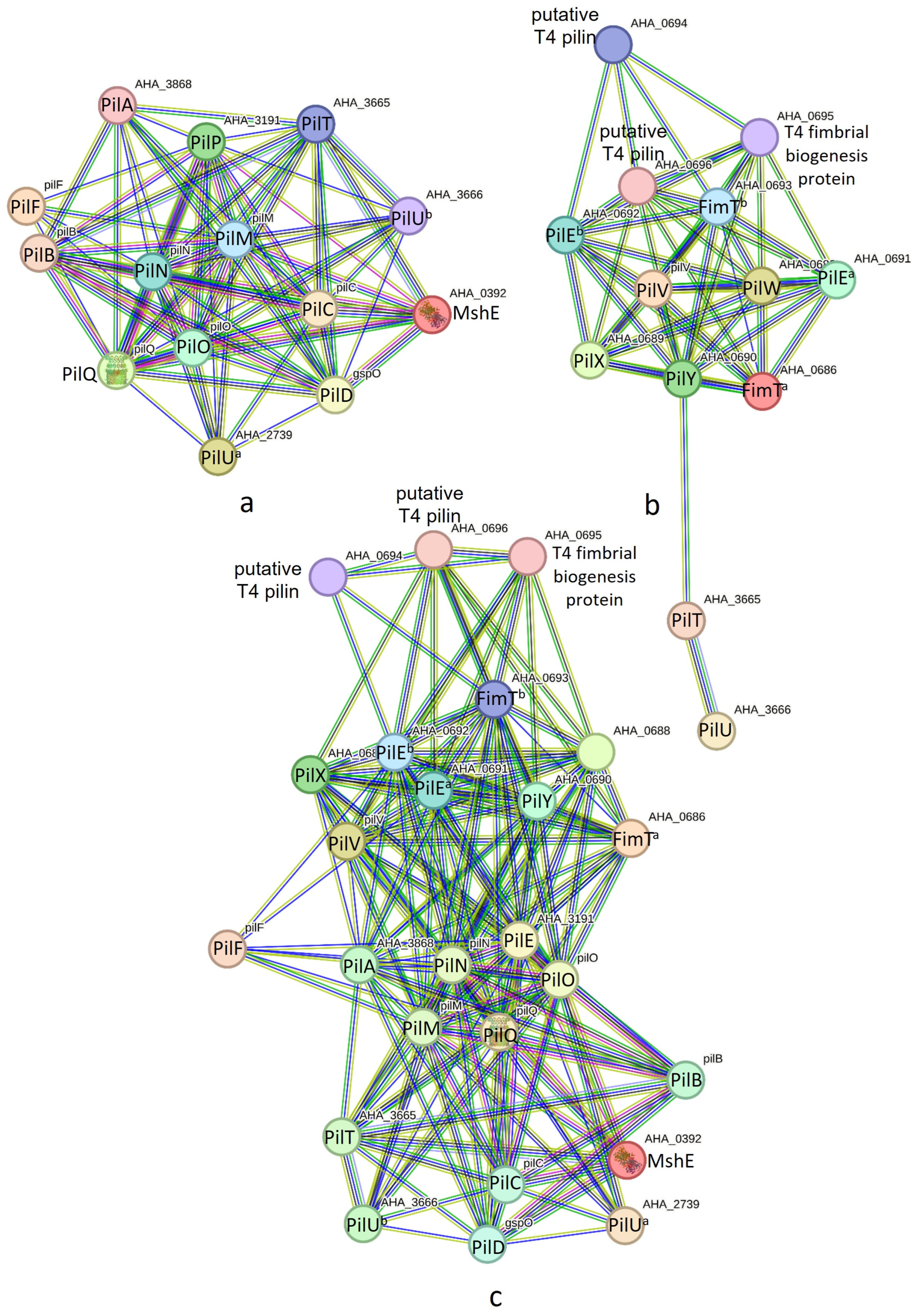
T4P nano-machine components encoded in the pilABCD and pilMNOPQ operons were found to interact, forming the PilABCDMNOPQ nano-machine (see Figure 3a). For the PilABCDMNOPQ nano-machine, we identified two separate retraction ATPases encoded by PilT and PilU. These genes were found to be distinct from the pilABCD and pilMNOPQ operons. T4P nano-machine components encoded in the pilVWXY operon exhibited strong PINs (see Figure 3b), comprising 13 nodes (T4P components) and 47 edges (interactions), with an average node degree of 7.23 and a local clustering coefficient of 0.797. In addition to 11 T4P components coded by pilVWXY, two retraction ATPases (PilT and PilU) coded outside the operon may play a crucial role here. The concurrence of PilY and PilT across bacterial genomes may indicate some level of interaction between them (Figure S1). The network within this nano-machine displayed significantly more interactions than expected. Otherwise, T4P components encoded in the pilABCD, pilMNOPQ, and pilVWXY operons all together may form a nano-machine as they exhibited strong PINs (Figure 3c), comprising 25 nodes (T4P components) and 169 edges (interactions), with an average node degree of 13.5 and a local clustering coefficient of 0.8217.
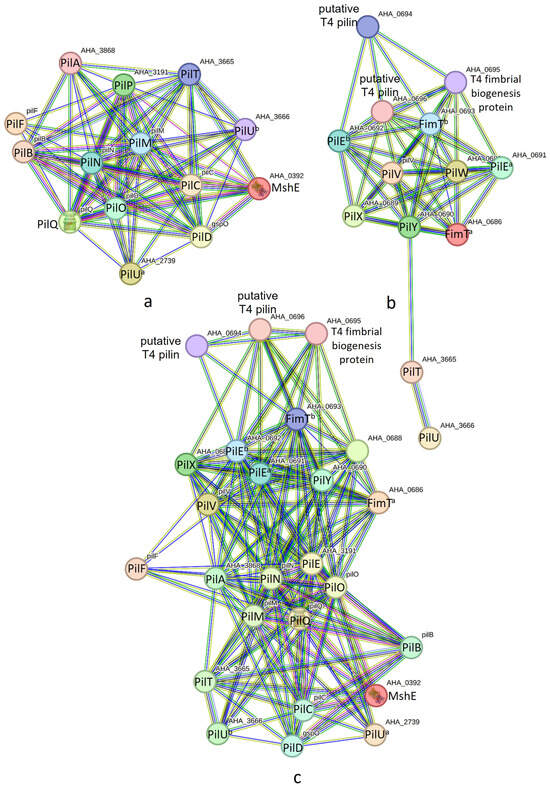
Figure 3.
(a) Protein interaction networks of PilABCDMNOPQ nano-machine components. (b) Protein interaction networks of PilVWXY nano-machine components. (c) Protein interaction networks of PilABCDMNOPQVWXY nano-machine components.
3.4. Global Survey of T4P Components within A. hydrophila Genomes
The nucleotide sequences of these 27 T4P components were utilized as tertiary queries for genome BLAST across available complete annotated A. hydrophila genomes (see Table S4). In the genome BLAST analysis, pilVWXY operon components did not return any results for the 10 experimental genomes where synteny was also absent (namely Ah2111, AC185, WP7 S18 ESBL 06, AH10, CSUSB2, 71339, A008N2, MX16A, NEB7245, and 4AK4). The differential distribution of the 27 T4P components in A. hydrophila genomes was illustrated as a heat map (refer to Figure S2).
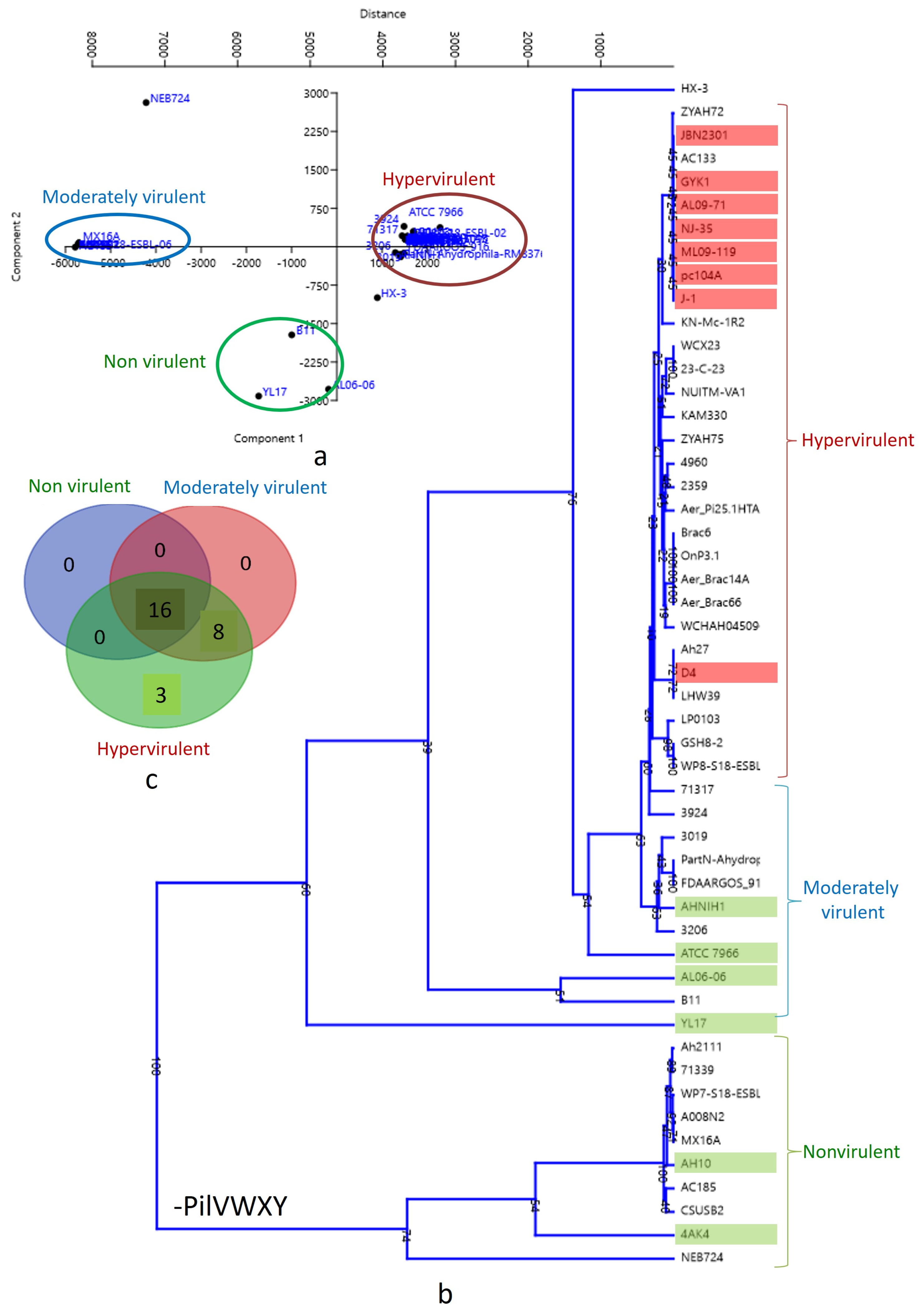
A biplot with PCA 1 and PCA 2 categorized the strains into three clusters (Figure 4a). The lower left quadrant represents nonvirulent strains, while the upper left quadrant represents low-virulent strains. Hypervirulent strains, mostly human pathogens, were clustered in the middle (just above and beneath the Y axis) of the right quadrant. A similar pattern was observed in the dendrogram analysis (Figure 4b), where the 10 strains lacking the pilVWXY operon clustered together at the bottom of the tree. The correlation between virulence and T4P components was illustrated through a Venn diagram (Figure 4c). According to the present analysis, the number of common T4P orthologs among nonvirulent, moderately virulent, and hypervirulent clusters is 16, indicating core T4P components (Table 2). Non-core T4P orthologs were primarily encoded by the pilVWXY operon only. The moderately virulent and hypervirulent clusters shared eight T4P orthologs. The hypervirulent cluster, mainly representing human pathogens, exhibited three unique T4P orthologs, including PilV and two unidentified components (Table 2).
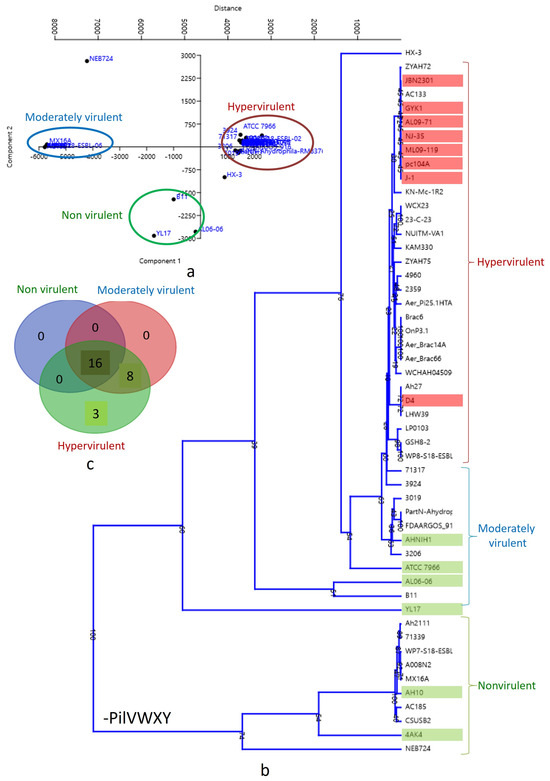
Figure 4.
(a) Biplot with strains distributed into three clusters by PCA 1 and PCA 2: nonvirulent strains: lower left quadrant; low-virulent strains: upper left quadrant; hypervirulent strains: middle (just above and beneath Y axis) of the right quadrant. (b) Dendrogram showing hypervirulent, low-virulent, and nonvirulent Aeromonas hydrophila strains in three different clusters. (c) Venn diagram showing the correlation of virulence and T4P components in Aeromonas hydrophila.
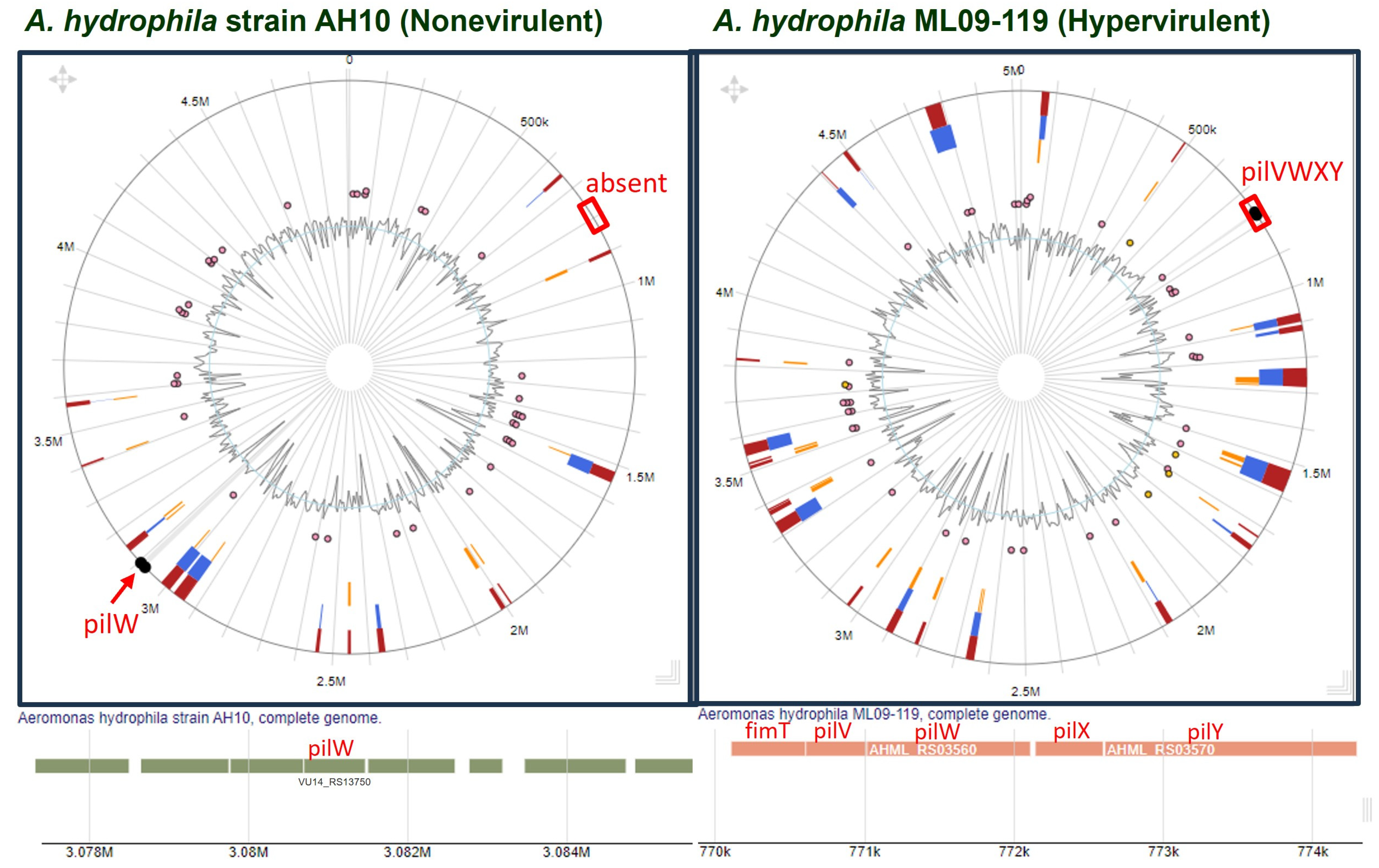
Synteny analysis indicated the absence of the PilVWXY operon in nonvirulent strains. However, due to recombination, these genes might have changed their position. Thus, to confirm, we compared the genomic region of a nonvirulent strain (A. hydrophila AH10) and that of a hypervirulent strain (A. hydrophila ML09-119) using IslandViewer 4. Interestingly, the result of the comparative study made it clear that PilV and 10 other components (except PilW) coded by pilVWXY were absent from the nonvirulent strain (Figure 5).
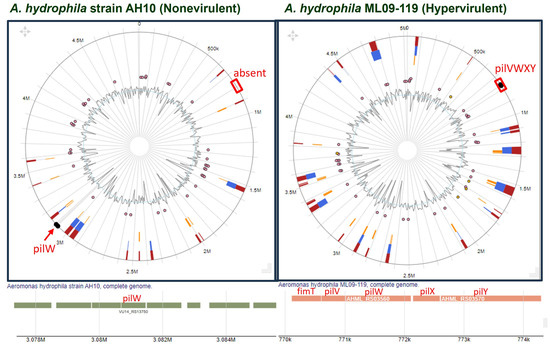
Figure 5.
Comparison of genomic islands between a nonvirulent (A. hydrophila AH10) and a hypervirulent (A. hydrophila ML09-119) strain with special emphasis on pilVWXY.
4. Discussion
4.1. A. hydrophila as a Model
As a pathogen, A. hydrophila offers a unique opportunity to study bacterial infections across various animal models due to its broad host range, spanning from protozoa to mammals. A. hydrophila has already been proposed as a model for studying infections [1]. The genomes of several strains are readily available in public databases [27]. In our exploration of a specific secretion system, we observed significant divergence across bacterial species, potentially contributing to virulence (Figure 4c). Therefore, this bacterial species has already demonstrated its potential as a model for studying bacterial infections and the underlying molecular machinery.
4.2. Emerging New Protocol to Study the T4P Secretion System
There are various standalone programs and server-based applications available for determining type IV pili (T4P), including TXSScan by Abby et al., 2016 [28]. TXSScan identified only one T4P profile in A. hydrophila ATCC 7966 and A. hydrophila ML09-119 [28]. However, in the present study, we identified 25 T4P components distributed into three operons for both strains. In our previous study, we developed a protocol for identifying all major secretion systems of Bradyrhizobium spp., which resulted in better resolution than the standalone program TXSScan [22]. However, type II secretion (T2S) systems are very similar to T4P systems, including all components except the retraction ATPase, and, in some cases, components are shared between the two [29]. For example, Acinetobacter baumannii has both T4P and TIIS and only one prepilin peptidase. Therefore, specific protocols for mining specialized systems are crucial. In the present investigation, we identified two potential retraction ATPases (PilU and PilT) for the PilABCDMNOPQ nano-machine. However, the same retraction ATPases may also be shared by the PilVWXY nano-machine. We believe that the present protocol will aid researchers in better mining T4P components in other bacterial systems as well.
4.3. Discovery of a New T4P System in A. hydrophila
This marks the first report on the distribution of pilVWXY in A. hydrophila. PilV is recognized for promoting bacterial adhesion to human epithelial cells [30], directly interacting with the beta-adrenergic receptor of human endothelial cells [31]. PilV-mediated inter-bacterial adhesion has also been documented [32]. Hence, the presence of PilV may indeed be a determining factor in the virulence of A. hydrophila (Figure 4c). In the Venn diagram, it is evident that PilV is a unique T4P component found exclusively in the hypervirulent cluster of A. hydrophila (Figure 4c). PilW-mediated active electron transfer has also been proposed for Acidithiobacillus ferrooxidans [33]. PilX was detected via immuno-electron microscopy and was associated with type IV pili [34]. Recent evidence suggests that PilX has a global effect on the conformation of pili, implying a potential indirect impact on pilus function [35].
The PilABCDMNOPQ nano-machine (Figure 3a) may adequately fit into the T4P architecture depicted by Craig et al. for P. aeruginosa [21]. However, there is also a probability that the T4P nano-machine components encoded in the pilVWXY operon may contribute to the development of a new nano-machine (Figure 3b). This nano-machine may adequately fit into the T4P assembly model proposed by Nguyen et al. for P. aeruginosa [36]. There is also a probability that the pilVWXY nano-machine may interact with the PilABCDMNOPQ nano-machine to form PilABCDMNOPQVWXY. This probability is supported by the schematic representation of T4P in Xylella fastidiosa by Merfa et al. [37].
4.4. Contribution of pilVWXY to Virulence
Based on T4P components, all the experimental genomes were clustered into three groups, i.e., hypervirulent, low-virulent, and nonvirulent, as outlined in the PCA (Figure 4a) and dendrogram analyses (Figure 4b). However, the absence of pilVWXY is the distinguishing feature of the nonvirulent cluster (Figure 4a,c). A. hydrophila AH10 and 4AK4 were already reported as nonvirulent strains by Abdella et al. [28]. Therefore, the nonvirulent cluster is well supported by our investigation and by the experimental data of previous workers. A whole-genome comparison of A. hydrophila AH10 with A. hydrophila ML09-119 (hypervirulent) clearly demonstrated the absence of all or most T4P components of the PilVWXY nano-machine from the nonvirulent strain of A. hydrophila (Figure 5).
Though A. hydrophila AHNIH1, ATCC 7966, AL0606, and YL17 were reported as nonvirulent by Abdella et al. [28], these strains were reported to cause infection by other researchers. As for example A. hydrophila AHNIH1 was reportedly isolated from an infected patient by Rodrigues et al. [38]. Similarly, A. hydrophila ATCC 7966 infection in Carassius auratus was reported by Peng et al. [39]. Therefore, these strains may be re-classified as low-virulent strains, as evident in the PCA (Figure 4a) and dendrogram (Figure 4b) in our study. Interestingly, both the low-virulent and hypervirulent clusters share components of pilVWXY (Figure 4c). The primary distinction between these two clusters is the presence of PilV, which is unique to the hypervirulent cluster (Figure 4c) and is mainly composed of human pathogenic strains. The adhesive capability of T4P is attributed to the presence of PilY as an adhesin [40]. PilY in P. aeruginosa has been shown to be vital for T4P assembly [41] and functions as a mechanosensor, facilitating surface contact-dependent activation of virulence gene expression [42]. Hence, our investigation strongly suggests the role of the newly described T4P system (pilVWXY) as a virulence determinant in A. hydrophila.
5. Conclusions
Here, we reported the distribution of 27 T4P components in A. hydrophila. Our study also suggests the potential role of the PilVWXY nano-machine in establishing human disease. Interestingly, this nano-machine was completely absent from 10 strains, most of which are considered nonvirulent or non-human pathogens. PilV, along with two other orthologs (bearing the locus tags AHA_0694 and AHA_0696 in A. hydrophila ATCC 7966), was found to be unique to hypervirulent or human-pathogenic strains. Hence, there is a strong possibility that PilVWXY serves as an important virulence determinant for A. hydrophila, contributing to its pathogenicity in humans.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/pathogens13050365/s1, Table S1: Reference dataset for components of T4P secretion system from previous publications used as primary quarry; Table S2: Reference dataset used for identification of orthologs for components of T4P secretion system; Table S3: Reference dataset for protein components of T4P secretion system used as secondary quarry; Table S4: Max score of Genome Blast for T4P components across A. hydrophila genomes; Figure S1: Co-occurrence of PilY and PilT across genomes indicating probable interaction between them; Figure S2: Heatmap showing distribution of T4P components within 51 A. hydrophila genome [30,34,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58].
Author Contributions
Conceptualization, P.C.; methodology, P.C., A.B. and G.B.; formal analysis, P.C. and A.B.; data mining, A.B.; writing—original draft preparation, A.B., P.C. and G.B.; software, P.C. and G.B.; writing—review and editing, P.C. and G.B.; supervision, P.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
This article does not require an Institutional Review Board statement.
Informed Consent Statement
This article does not contain any studies with human participants and/or animals performed by any of the authors. Formal consent was not required in this study.
Data Availability Statement
A. hydrophila genomes with whole-genome assembly and annotation report up to 31.03.2023 were downloaded from the NCBI Genome database (https://www.ncbi.nlm.nih.gov/genome/browse/#!/prokaryotes/1422/ accessed on 31 March 2023) or from the KEGG genome database (https://www.genome.jp/dbget-bin/www_bfind_sub?mode=bfind&max_hit=1000&dbkey=genome&keywords=Aeromonas+hydrophila accessed on 31 March 2023), which includes 51 genomes (Table 1). Sequences of stress-responsive enzymes and stress regulators were retrieved from the UniPort database (https://www.uniprot.org/ accessed on 31 March 2023). Protein structures were retrieved from the RCSB PDB database (https://www.rcsb.org/ accessed on 31 March 2023).
Acknowledgments
We would like to express our sincere gratitude for the invaluable support extended by Raja Rammohun Roy Mahavidyalaya and M.U.C. Women’s College. We are also very thankful to the University of Illinois, Urbana-Champaign, Illinois, USA for providing the necessary support.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Li, J.; Zhang, X.-L.; Liu, Y.-J.; Lu, C.-P. Development of an Aeromonas hydrophila infection model using the protozoan Tetrahymena thermophila. FEMS Microbiol. Lett. 2011, 316, 160–168. [Google Scholar] [CrossRef] [PubMed]
- Kodjo, A.; Haond, F.; Richard, Y. Molecular and phenotypic features of aeromonads isolated from snails (Helix aspersa) affected with a new summer disease. J. Vet. Med. Ser. B 1997, 44, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.; Li, X.; Gao, X.; Jiang, Q.; Chen, Q.; Zhang, S.; Tong, S.; Liu, X.; Zhu, J.; Zhang, X. Pathogenicity of Aeromonas hydrophila causing mass mortalities of Procambarus clarkia and its induced host immune response. Microb. Pathog. 2020, 147, 104376. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, H.A.; Mohamed, M.E.; Rezk, M.M.; Gharieb, R.M.; Abdel-Maksoud, S.A. Aeromonas hydrophila in fish and humans; Prevalence, virulotyping and antimicrobial resistance. Slov. Vet. Res. 2018, 55, 113–124. [Google Scholar]
- Neto, P.G.G.; Titon, S.C.; Muxel, S.M.; Titon, B., Jr.; de Figueiredo, A.C.; Floreste, F.R.; Lima, A.S.; Assis, V.R.; Gomes, F.R. Immune and endocrine alterations at the early stage of inflammatory assemblage in toads after stimulation with heat-killed bacteria (Aeromonas hydrophila). Comp. Biochem. Physiol. Part A Mol. Integr. Physiol. 2024, 111606. [Google Scholar]
- Kwon, J.; Kim, S.G.; Kim, S.W.; Yun, S.; Kim, H.J.; Giri, S.S.; Han, S.J.; Oh, W.T.; Park, S.C. A case of mortality caused by Aeromonas hydrophila in wild-caught red-eyed crocodile skinks (Tribolonotus gracilis). Vet. Sci. 2019, 7, 4. [Google Scholar] [CrossRef] [PubMed]
- Awaad, M.; Hatem, M.; El-Ghany, W.; El-Sawy, A.; Fathi, A. Certain epidemiological aspects of Aeromonas hydrophila infection in chickens. J. Am. Sci. 2011, 7, 761–770. [Google Scholar]
- Pérez, D.A.M.; Baez, A.; Venegas, B.; de Lourdes Reyes-Luna, R.M.; Molina-Romero, D. Resistance to antibiotics of the genus Aeromonas, as an emerging opportunistic pathogen in fish and humans. Rev. Biomédica 2023, 34, 191–207. [Google Scholar]
- Sorsa, M.; Mamo, G.; Abera, L. Major fish-borne bacterial and parasitic zoonoses in Ethiopia: A review. Int. J. Fauna Biol. Stud. 2019, 6, 50–58. [Google Scholar]
- El-Hossary, D.; Mahdy, A.; Elariny, E.Y.; Askora, A.; Merwad, A.M.; Saber, T.; Dahshan, H.; Hakami, N.Y.; Ibrahim, R.A. Antibiotic Resistance, Virulence Gene Detection, and Biofilm Formation in Aeromonas spp. Isolated from Fish and Humans in Egypt. Biology 2023, 12, 421. [Google Scholar] [CrossRef]
- Kali, A.; Kalaivani, R.; Charles, P.; Seetha, K. Aeromonas hydrophila meningitis and fulminant sepsis in preterm newborn: A case report and review of literature. Indian J. Med. Microbiol. 2016, 34. [Google Scholar] [CrossRef]
- Janda, J.M.; Abbott, S.L. Evolving concepts regarding the genus Aeromonas: An expanding panorama of species, disease presentations, and unanswered questions. Clin. Infect. Dis. 1998, 27, 332–344. [Google Scholar] [CrossRef] [PubMed]
- Raynaud, C.; Sheppard, D.; Berry, J.-L.; Gurung, I.; Pelicic, V. PilB from Streptococcus sanguinis is a bimodular type IV pilin with a direct role in adhesion. Proc. Natl. Acad. Sci. USA 2021, 118, e2102092118. [Google Scholar] [CrossRef] [PubMed]
- Mattick, J.S. Type IV pili and twitching motility. Annu. Rev. Microbiol. 2002, 56, 289–314. [Google Scholar] [CrossRef] [PubMed]
- Suh, J.-Y.; Spyracopoulos, L.; Keizer, D.W.; Irvin, R.T.; Sykes, B.D. Backbone dynamics of receptor binding and antigenic regions of a Pseudomonas aeruginosa pilin monomer. Biochemistry 2001, 40, 3985–3995. [Google Scholar] [CrossRef]
- Tennent, J.M.; Mattick, J.S. Type 4 Fimbriae. In Fimbriae Adhesion, Genetics, Biogenesis, and Vaccines; CRC Press: Boca Raton, FL, USA, 2020; pp. 127–146. [Google Scholar]
- Nieto, V.; Kroken, A.R.; Grosser, M.R.; Smith, B.E.; Metruccio, M.M.; Hagan, P.; Hallsten, M.E.; Evans, D.J.; Fleiszig, S.M. Type IV pili can mediate bacterial motility within epithelial cells. mBio 2019, 10. [Google Scholar] [CrossRef]
- Kirov, S.M.; Barnett, T.C.; Pepe, C.M.; Strom, M.S.; Albert, M.J. Investigation of the role of type IV Aeromonas pilus (Tap) in the pathogenesis of Aeromonas gastrointestinal infection. Infect. Immun. 2000, 68, 4040–4048. [Google Scholar] [CrossRef]
- Pepe, C.M.; Eklund, M.W.; Strom, M.S. Cloning of an Aeromonas hydrophila type IV pilus biogenesis gene cluster: Complementation of pilus assembly functions and characterization of a type IV leader peptidase/N-methyltransferase required for extracellular protein secretion. Mol. Microbiol. 1996, 19, 857–869. [Google Scholar] [CrossRef] [PubMed]
- Boyd, J.M.; Dacanay, A.; Knickle, L.C.; Touhami, A.; Brown, L.L.; Jericho, M.H.; Johnson, S.C.; Reith, M. Contribution of type IV pili to the virulence of Aeromonas salmonicida subsp. salmonicida in Atlantic salmon (Salmo salar L.). Infect. Immun. 2008, 76, 1445–1455. [Google Scholar] [CrossRef]
- Craig, L.; Forest, K.T.; Maier, B. Type IV pili: Dynamics, biophysics and functional consequences. Nat. Rev. Microbiol. 2019, 17, 429–440. [Google Scholar] [CrossRef]
- Banerjee, G.; Basak, S.; Roy, T.; Chattopadhyay, P. Intrinsic role of bacterial secretion systems in phylogenetic niche conservation of Bradyrhizobium spp. FEMS Microbiol. Ecol. 2019, 95, fiz165. [Google Scholar] [CrossRef] [PubMed]
- Oberto, J. SyntTax: A web server linking synteny to prokaryotic taxonomy. BMC Bioinform. 2013, 14, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Despalins, A.; Marsit, S.; Oberto, J. Absynte: A web tool to analyze the evolution of orthologous archaeal and bacterial gene clusters. Bioinformatics 2011, 27, 2905–2906. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Bertelli, C.; Laird, M.R.; Williams, K.P.; Group, S.F.U.R.C.; Lau, B.Y.; Hoad, G.; Winsor, G.L.; Brinkman, F.S. IslandViewer 4: Expanded prediction of genomic islands for larger-scale datasets. Nucleic Acids Res. 2017, 45, W30–W35. [Google Scholar] [CrossRef] [PubMed]
- Abdella, B.; Abozahra, N.A.; Shokrak, N.M.; Mohamed, R.A.; El-Helow, E.R. Whole spectrum of Aeromonas hydrophila virulence determinants and the identification of novel SNPs using comparative pathogenomics. Sci. Rep. 2023, 13, 7712. [Google Scholar] [CrossRef] [PubMed]
- Abby, S.S.; Cury, J.; Guglielmini, J.; Néron, B.; Touchon, M.; Rocha, E.P. Identification of protein secretion systems in bacterial genomes. Sci. Rep. 2016, 6, 23080. [Google Scholar] [CrossRef] [PubMed]
- Ayers, M.; Howell, P.L.; Burrows, L.L. Architecture of the type II secretion and type IV pilus machineries. Future Microbiol. 2010, 5, 1203–1218. [Google Scholar] [CrossRef] [PubMed]
- Winther-Larsen, H.C.; Hegge, F.T.; Wolfgang, M.; Hayes, S.F.; Van Putten, J.P.; Koomey, M. Neisseria gonorrhoeae PilV, a type IV pilus-associated protein essential to human epithelial cell adherence. Proc. Natl. Acad. Sci. USA 2001, 98, 15276–15281. [Google Scholar] [CrossRef]
- Barnier, J.-P.; Meyer, J.; Kolappan, S.; Bouzinba-Ségard, H.; Gesbert, G.; Jamet, A.; Frapy, E.; Schönherr-Hellec, S.; Capel, E.; Virion, Z. The minor pilin PilV provides a conserved adhesion site throughout the antigenically variable meningococcal type IV pilus. Proc. Natl. Acad. Sci. USA 2021, 118, e2109364118. [Google Scholar] [CrossRef]
- Ishiwa, A.; Komano, T. PilV adhesins of plasmid R64 thin pili specifically bind to the lipopolysaccharides of recipient cells. J. Mol. Biol. 2004, 343, 615–625. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, H. Type IV pili of Acidithiobacillus ferrooxidans can transfer electrons from extracellular electron donors. J. Basic Microbiol. 2014, 54, 226–231. [Google Scholar] [CrossRef] [PubMed]
- Helaine, S.; Dyer, D.H.; Nassif, X.; Pelicic, V.; Forest, K.T. 3D structure/function analysis of PilX reveals how minor pilins can modulate the virulence properties of type IV pili. Proc. Natl. Acad. Sci. USA 2007, 104, 15888–15893. [Google Scholar] [CrossRef] [PubMed]
- Brissac, T.; Mikaty, G.; Duménil, G.; Coureuil, M.; Nassif, X. The meningococcal minor pilin PilX is responsible for type IV pilus conformational changes associated with signaling to endothelial cells. Infect. Immun. 2012, 80, 3297–3306. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, Y.; Sugiman-Marangos, S.; Harvey, H.; Bell, S.D.; Charlton, C.L.; Junop, M.S.; Burrows, L.L. Pseudomonas aeruginosa minor pilins prime type IVa pilus assembly and promote surface display of the PilY1 adhesin. J. Biol. Chem. 2015, 290, 601–611. [Google Scholar] [CrossRef] [PubMed]
- Merfa, M.V.; Zhu, X.; Shantharaj, D.; Gomez, L.M.; Naranjo, E.; Potnis, N.; Cobine, P.A.; De La Fuente, L. Complete functional analysis of type IV pilus components of a reemergent plant pathogen reveals neofunctionalization of paralog genes. PLoS Pathog. 2023, 19, e1011154. [Google Scholar] [CrossRef] [PubMed]
- Rodrigues, M.V.; Dias, M.F.F.; Francisco, C.J.; David, G.S.; da Silva, R.J.; Junior, J.P.A. Aeromonas hydrophila in Nile tilapia (Oreochromis niloticus) from Brazilian aquaculture: A public health problem. Emergent Life Sci. Res. 2019, 5, 48–55. [Google Scholar] [CrossRef]
- Peng, M.; Tong, W.; Zhao, Z.; Xiao, L.; Wang, Z.; Liu, X.; He, X.; Song, Z. Attenuation of Aeromonas hydrophila Infection in Carassius auratus by YtnP, a N-acyl Homoserine Lactonase from Bacillus licheniformis T-1. Antibiotics 2021, 10, 631. [Google Scholar] [CrossRef]
- Berry, J.-L.; Pelicic, V. Exceptionally widespread nanomachines composed of type IV pilins: The prokaryotic Swiss Army knives. FEMS Microbiol. Rev. 2015, 39, 134–154. [Google Scholar] [CrossRef]
- Heiniger, R.W.; Winther-Larsen, H.C.; Pickles, R.J.; Koomey, M.; Wolfgang, M.C. Infection of human mucosal tissue by Pseudomonas aeruginosa requires sequential and mutually dependent virulence factors and a novel pilus-associated adhesin. Cell. Microbiol. 2010, 12, 1158–1173. [Google Scholar] [CrossRef]
- Webster, S.; Mathelie-Guinlet, M.; Verissimo, A.; Schultz, D.; Viljoen, A.; Lee, C. Force-Induced Changes of PilY1 Drive Surface Sensing by Pseudomonas aeruginosa. mBio 2022, 13, e0375421. [Google Scholar] [CrossRef] [PubMed]
- Averhoff, B.; Kirchner, L.; Pfefferle, K.; Yaman, D. Natural transformation in Gram-negative bacteria thriving in extreme environments: From genes and genomes to proteins, structures and regulation. Extremophiles 2021, 25, 425–436. [Google Scholar] [CrossRef] [PubMed]
- Potapova, A.; Carreira, L.A.M.; Søgaard-Andersen, L. The small GTPase MglA together with the TPR domain protein SgmX stimulates type IV pili formation in M. xanthus. Proc. Natl. Acad. Sci. USA 2020, 117, 23859–23868. [Google Scholar] [CrossRef] [PubMed]
- Vahvelainen, N.; Bozkurt, E.; Maula, T.; Johansson, A.; Pöllänen, M.T.; Ihalin, R. Pilus PilA of the naturally competent HACEK group pathogen Aggregatibacter actinomycetemcomitans stimulates human leukocytes and interacts with both DNA and proinflammatory cytokines. Microb. Pathog. 2022, 173, 105843. [Google Scholar] [CrossRef] [PubMed]
- Dye, K.J.; Vogelaar, N.J.; Sobrado, P.; Yang, Z. High-throughput screen for inhibitors of the type IV pilus assembly ATPase PilB. mSphere 2021, 6, 121. [Google Scholar] [CrossRef]
- Rudel, T.; Scheuerpflug, I.; Meyer, T.F. Neisseria PilC protein identified as type-4 pilus tip-located adhesin. Nature 1995, 373, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Strom, M.S.; Nunn, D.N.; Lory, S. Posttranslational processing of type IV prepilin and homologs by PilD of Pseudomonas aeruginosa. Methods Enzymol. 1994, 235, 527–540. [Google Scholar] [PubMed]
- Darzins, A.; Russell, M.A. Molecular genetic analysis of type-4 pilus biogenesis and twitching motility using Pseudomonas aeruginosa as a model system–a review. Gene 1997, 192, 109–115. [Google Scholar] [CrossRef] [PubMed]
- Koo, J.; Tammam, S.; Ku, S.-Y.; Sampaleanu, L.M.; Burrows, L.L.; Howell, P.L. PilF is an outer membrane lipoprotein required for multimerization and localization of the Pseudomonas aeruginosa type IV pilus secretin. J. Bacteriol. 2008, 190, 6961–6969. [Google Scholar] [CrossRef]
- Liu, J.; Hu, L.; Xu, Z.; Tan, C.; Yuan, F.; Fu, S.; Cheng, H.; Chen, H.; Bei, W. Actinobacillus pleuropneumoniae two-component system QseB/QseC regulates the transcription of PilM, an important determinant of bacterial adherence and virulence. Vet. Microbiol. 2015, 177, 184–192. [Google Scholar] [CrossRef]
- McCallum, M.; Tammam, S.; Little, D.J.; Robinson, H.; Koo, J.; Shah, M.; Calmettes, C.; Moraes, T.F.; Burrows, L.L.; Howell, P.L. PilN binding modulates the structure and binding partners of the Pseudomonas aeruginosa type IVa pilus protein PilM. J. Biol. Chem. 2016, 291, 11003–11015. [Google Scholar] [CrossRef] [PubMed]
- Ozanic, M.; Marecic, V.; Knezevic, M.; Kelava, I.; Stojková, P.; Lindgren, L.; Bröms, J.E.; Sjöstedt, A.; Abu Kwaik, Y.; Santic, M. The type IV pili component PilO is a virulence determinant of Francisella novicida. PLoS ONE 2022, 17, e0261938. [Google Scholar] [CrossRef] [PubMed]
- Balasingham, S.V.; Collins, R.F.; Assalkhou, R.; Homberset, H.v.; Frye, S.A.; Derrick, J.P.; Tønjum, T. Interactions between the lipoprotein PilP and the secretin PilQ in Neisseria meningitidis. J. Bacteriol. 2007, 189, 5716–5727. [Google Scholar] [CrossRef] [PubMed]
- Narulita, E.; Addy, H.S.; Kawasaki, T.; Fujie, M.; Yamada, T. The involvement of the PilQ secretin of type IV pili in phage infection in Ralstonia solanacearum. Biochem. Biophys. Res. Commun. 2016, 469, 868–872. [Google Scholar] [CrossRef] [PubMed]
- Hughes, H.Q.; Christman, N.D.; Dalia, T.N.; Ellison, C.K.; Dalia, A.B. The PilT retraction ATPase promotes both extension and retraction of the MSHA type IVa pilus in Vibrio cholerae. PLoS Genet. 2022, 18, e1010561. [Google Scholar] [CrossRef] [PubMed]
- Carbonnelle, E.; Hélaine, S.; Prouvensier, L.; Nassif, X.; Pelicic, V. Type IV pilus biogenesis in Neisseria meningitidis: PilW is involved in a step occurring after pilus assembly, essential for fibre stability and function. Mol. Microbiol. 2005, 55, 54–64. [Google Scholar] [CrossRef]
- Guzzo, C.R.; Salinas, R.K.; Andrade, M.O.; Farah, C.S. PILZ protein structure and interactions with PILB and the FIMX EAL domain: Implications for control of type IV pilus biogenesis. J. Mol. Biol. 2009, 393, 848–866. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).