Simple Summary
Quantitative real-time PCR (qRT-PCR) is necessary to select stable reference genes for normalization. In this study, the suitability of various reference genes for qRT-PCR analysis was evaluated in different developmental stages of Chilo suppressalis exposed to desiccation or rehydration stress. The results indicated that 18S rRNA was the most stable reference gene for monitoring gene expression in the third instar larvae; ACTIN, TUB, UBI, UBI, and EF1 were the optimal genes for the fifth instar larvae, male pupae, female pupae, male adults, and female adults, respectively. The optimal number of reference genes recommended by geNorm analysis indicated that two candidate reference genes were sufficient for data normalization under all experimental conditions tested. To validate these recommendations, the expression profile of Hsp60 was investigated. Hsp60 transcript levels showed significant differences when normalized to the most stable single reference gene, or combined reference genes, compared with the least stable reference gene. The reference genes identified in the present study will enhance the reliability of gene expression data for C. suppressalis under humidity stress.
Abstract
Quantitative real-time PCR (qRT-PCR) is a high-reliability, -sensitivity, and -operability technique to quantify gene expression. It is necessary to select stable reference genes for normalization. Water plays important roles in the metabolism, physiology, distribution, and so on, in insects. In this study, the suitability of various reference genes for qRT-PCR analysis was evaluated in different developmental stages of Chilo suppressalis exposed to desiccation or rehydration stress. The ∆Ct method, geNorm, NormFinder, and BestKeeper were used to evaluate the suitability of nine reference genes for normalizing gene expression in the third instar larvae, the fifth instar larvae, male pupae, female pupae, male adults, and female adults under different humidities. The results indicated that 18S rRNA was the most stable reference gene for monitoring gene expression in the third instar larvae, while ACTIN, TUB, UBI, UBI, and EF1 were the optimal genes for the fifth instar larvae, male pupae, female pupae, male adults, and female adults, respectively. The optimal number of reference genes recommended by geNorm analysis indicated that two candidate reference genes were sufficient for data normalization under all experimental conditions tested. To validate these recommendations, the expression profile of the gene encoding heat shock protein 60 (Hsp60) was investigated. Hsp60 transcript levels showed significant differences when normalized to the most stable single reference gene, or combined reference genes, compared with the least stable reference gene. The reference genes identified in the present study will enhance the reliability of gene expression data for C. suppressalis under humidity stress.
1. Introduction
Regulation of water balance is a primary component of homeostasis in all organisms. In insects, water also plays crucial roles in metabolite transport, maintenance of macromolecular structures, and determination of spatiotemporal distribution variations [1,2]. However, due to relatively small body size and high surface-to-volume ratio, insects face significant challenges in maintaining water balance [3,4]. Consequently, insects have evolved many strategies to manage water, such as an integument with abundant hydrophobic epicuticular lipids, acclimation treatments, and proteins that regulate water permeability [5,6]. With global climate change, environmental conditions including extreme temperatures, altered rainfall patterns, and droughts will occur frequently. It is very important to understand the responses of insects to water availability, because it has a pronounced influence on their survival, distribution patterns, and species richness [1,2]. Nevertheless, the molecular mechanisms underlying water rehydration and desiccation resistance in insects remain poorly understood [4,5].
The striped stem borer, Chilo suppressalis (Walker), is an important rice and water oat pest widely distributed in temperate and subtropical Asia, southern Europe, and North Africa. In China, the first instar to sixth instar larvae of C. suppressalis bore into the stems of rice and water oats, representing the main damaging stage of this pest to crops. The damage caused by C. suppressalis has spread to more than ten provinces in China and is gradually increasing [6]. The larvae prefer dry environments for pupation, while the pupae favor high-humidity conditions, but are intolerant of waterlogging. After a few days, the pupae emerge as adults, which lay eggs on host plants. All developmental stages of C. suppressalis favor high-humidity environments. In the control of C. suppressalis, humidity is often used to regulate its damage. To improve green control of C. suppressalis, clarifying the molecular mechanisms underlying its response to desiccation or rehydration stress at different developmental stages is crucial.
More accurate genomes and more transcriptomes of C. suppressalis are available [7,8]. Gene expression analyses are important for identifying the functions of different genes in various organisms. Quantitative real-time PCR (qRT-PCR) is a high-reliability, -sensitivity, and -operability technique, which is available to measure and evaluate functional gene expression [9,10]. Although qRT-PCR is often described as the gold standard for gene expression analysis, the reliability of results is directly affected by some factors, such as RNA quality and quantity, reverse transcription efficiency, and primer design [11,12]. Therefore, accurate gene expression normalization, using proper reference genes, is crucial to obtaining reliable gene expression levels. Reference genes, often called housekeeping genes, are based on their supposed constitutive expression, such as ACTIN, 18S ribosomal RNA (18S rRNA), and tubulin (TUB). However, it is noteworthy that these genes are influenced by different treatment conditions, and no universal reference gene exists [13,14]. In addition, the number of reference genes used for normalization can also affect the results of qRT-PCR [4,15]. Consequently, in order to ensure accurate results, it is crucial to validate the reference genes with respect to the specific experiment.
In this study, expression stability of nine candidate reference genes in different developmental stages of C. suppressalis under desiccation or rehydration was evaluated, including 18S rRNA, ACTIN, TUB, elongation factor 1 (EF1), ubiquitin-conjugating enzyme (UBI), nicotinamide adenine dinucleotide dehydrogenase (NADHD), histone 3 (H3), arginine kinase (AK), and ribosomal protein S11 (RPS11). To further evaluate the results, the expression profile of heat shock protein 60 gene (Hsp60) was analyzed. The findings of this study can provide support for future research on genes related to water adaptation in this important pest species.
2. Materials and Methods
2.1. Insects
C. suppressalis were collected from the rice fields of Yangzhou (32.39° N, 119.42° E) and reared on an artificial food source in an environmental chamber at 28 ± 1 °C with a 16:8 (light/dark) photoperiod and 70 ± 5% relative humidity (RH).
2.2. Sample Collection Under Various Humidity Treatments
According to the preliminary experiment, the 5th instar larvae, male and female pupae, and male and female adults, were exposed to 25% (desiccation), 50%, 75%, and 95% (rehydration) relative humidities for 24 h in the humidity chamber with ±1.5% RH accuracy (SANTN HTC-100, Santeng Instrument Co., Ltd., Shanghai, China) at 27 °C. The 3rd instar larvae were exposed to 25% (desiccation), 50%, 75%, and 95% (rehydration) relative humidities for 12 h each. The thirty individuals of C. suppressalis were included in every treatment, and surviving C. suppressalis were frozen in liquid nitrogen and stored at −80 °C immediately.
2.3. Selection of Reference Genes
ACTIN, TUB, EF1, H3, RPS11, NADHD, AK, 18S rRNA, and UBI were selected as reference genes, and the specificity and amplification efficiency of their primers were certificated (Table 1) [16].

Table 1.
Primer sequences and amplicon characteristics of the reference genes and Hsp60 from Chilo suppressalis.
2.4. RNA Isolation and Reverse Transcription
Total RNA was isolated from each sample using the SV Total RNA isolation system (Promega Z3100, Madison, WI, USA). RNA integrity was analyzed by agarose gel electrophoresis, and then RNA purity and concentration were measured by spectrophotometric measurements (Eppendorf Biophotometer plus, Hamburg, Germany). Then, 0.50 μg total RNA was used for cDNA synthesis using the Bio-Rad iScriptTM cDNA Synthesis Kit (Bio-Rad Laboratories, Berkeley, CA, USA).
2.5. Quantitative Real-Time PCR Analysis
cDNA with 1:10 dilution was used as template for qRT-PCR analysis. The PCR reaction mixtures contained 10 μL of iTaq Universal SYBR Green supermix (2×) (Bio-Rad), 1 μL of each gene-specific primer (10 μM), 2 μL of cDNA templates, and 6 μL ddH2O. All reactions were performed in CFX-96 real-time PCR system (Bio-Rad Laboratories, Berkeley, CA, USA). The specific procedure comprised 3 min of the initial denaturation at 95 °C, followed by 40 cycles of denaturation at 95 °C for 30 s, and 30 s at the Tm value of the primer pairs (Table 1) [16]. Melting curve analysis from 65 °C to 95 °C was performed to determine the specificity of the amplified PCR products. Every treatment included four replicates, and each replicate was assayed in three technical replicates.
2.6. Statistical Analyses
The threshold cycle (Ct) value is defined as the number of amplification cycles required for the fluorescent signal to reach a preset threshold, with this threshold set at a consistent fluorescence intensity across all reactions. The stability of the 9 candidate reference genes was evaluated using the ∆Ct method [17], BestKeeper [18], NormFinder version 0.953 [19], and GeNorm version 3.5 [20]. A ∆Ct method result with a lower standard deviation (SD) indicates a more stable gene. The BestKeeper creates an index, and lower index scores indicate greater stability. NormFinder estimates expression variation, in which lower values are more stable. The geNorm algorithm calculates an expression stability value (M) and compares the pair-wise variation (V) of a gene with others. The geNorm algorithm uses 0.15 as a cut-off, with a ratio of Vn/Vn + 1 exceeding 0.15 indicating the need for an additional reference gene [20].
2.7. Validation of Reference Genes
The Hsp60 in C. suppressalis was selected as a target gene to validate stable reference genes. Relative expression levels of Hsp60 in different treatments were conducted according to the 2−∆∆Ct method [17]. Hsp60 expression at 75% relative humidity was used as a control for calibration. For normalization using multiple reference genes, normalization was performed individually for each reference gene, followed by averaging the results. One-way analysis of variance (ANOVA) was performed to compare using SPSS v. 16.0.
3. Results
3.1. Total RNA Quality and PCR Amplification Specificity
The expression levels of ACTIN, TUB, EF1, H3, RPS11, NADHD, AK, 18S rRNA, UBI, and Hsp60 were investigated in different developmental stages of C. suppressalis under desiccation or rehydration by qRT-PCR (Table 2). The integrity, purity, and concentration of total RNA isolated from different samples were evaluated. The total RNA bands were intact on agarose gel electrophoresis, indicating that the RNA was of good integrity. The A260/280 ratios confirmed high purity for all RNA samples. Additionally, all ten genes produced amplicons that exhibited a single peak in melting curve analysis, demonstrating that each reaction amplified a unique product.

Table 2.
Ranking order of reference genes of Chilo suppressalis under different experimental conditions.
3.2. Expression Profiles of Reference Genes
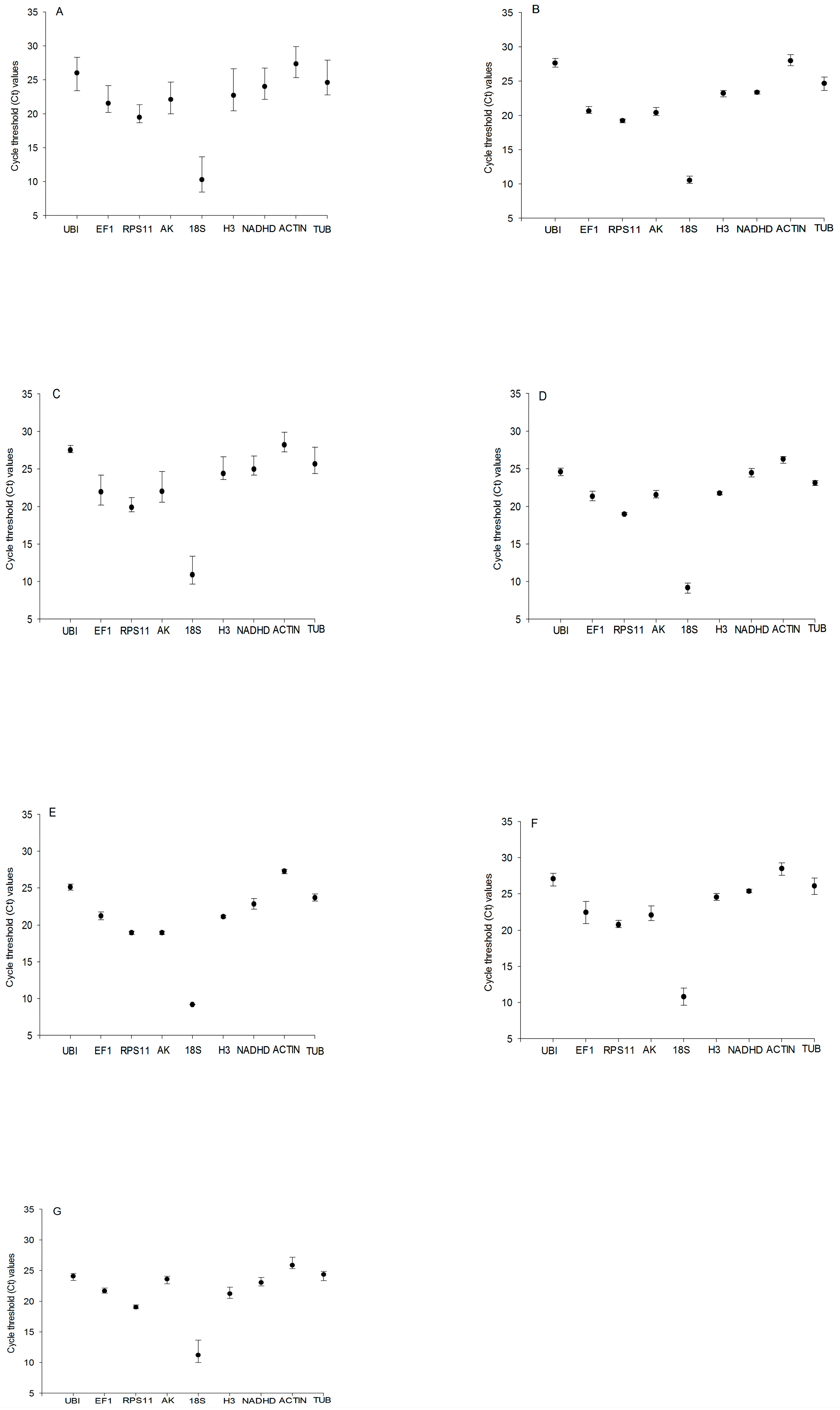
Expression levels were presented as the number of cycles required for amplification to reach a fixed threshold (threshold cycle, Ct value). The t Ct values of the nine candidate reference genes under different experimental conditions were plotted. The mean Ct values of the nine reference genes ranged from 8.45 (18S rRNA) to 29.89 (TUB). 18S rRNA (mean Ct 10.30) showed the most abundant expression levels, followed by RPS11 (mean Ct 19.47), EF1 (mean Ct 21.55), AK (mean Ct 22.12), H3 (mean Ct 22.71), NADHD (mean Ct 24.01), UBI (mean Ct 24.60), TUB (mean Ct 26.00), and ACTIN (mean Ct 27.36) (Figure 1A). However, RPS11 showed the smallest Ct variation (0.13), and TUB showed the biggest Ct variation (0.56) in samples of the third instar larvae (Figure 1B); for the fifth instar larvae, the smallest Ct variation (0.31) and the biggest Ct variation (1.32) were observed for UBI and AK, respectively (Figure 1C). RPS11 showed the smallest Ct variation (0.10) and EF1 showed the biggest Ct variation (0.52) in samples of the male pupae (Figure 1D), but, for the female pupae, the smallest Ct variation (0.03) and the biggest Ct variation (0.57) were observed for 18S rRNA and NADHD, respectively (Figure 1E). RPS11 showed the smallest Ct variation (0.15), and EF1 showed the biggest Ct variation (1.22) in samples of the male adults (Figure 1F), but, for the female adults, the smallest Ct variation (0.24) and the biggest Ct variation (1.22) were observed for RPS11 and 18S rRNA, respectively (Figure 1G). These results indicated that no single reference gene was suitable for normalizing gene expression across all developmental stages of C. suppressalis under desiccation or rehydration.
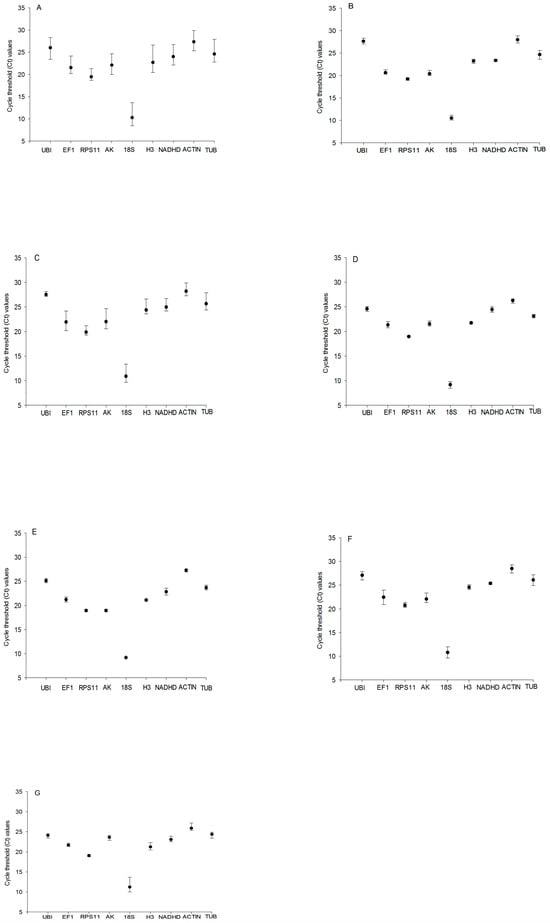
Figure 1.
Expression levels of candidate reference genes in different developmental stages of Chilo suppressalis under desiccation or rehydration. Expression stability (Ct) of reference genes in the following samples: (A) all Chilo suppressalis samples (n = 113); (B) samples of the third instar larvae (n = 36); (C) samples of the fifth instar larvae (n = 36); (D) samples of one-day-old male pupae (n = 36); (E) samples of one-day-old female pupae (n = 36); (F) samples of one-day-old male adults (n = 36); and (G) samples of one-day-old female adults (n = 36). The black circles indicate the means of samples, and the bars indicate the minimum to maximum values.
3.3. Gene Analysis of Gene Expression Stability
For the third instar larvae, under desiccation or rehydration, four methods of comprehensive analysis found that 18S rRNA and NADHD showed the highest expression stability. According to NormFinder, the stability ranking, from most to least stable, was 18S rRNA > EF1 > H3 > UBI > NADHD > RPS11 > ACTIN > TUB > AK. The comparative ∆Ct method, NormFinder, Best Keeper, and geNorm all revealed that TUB and AK were relatively unstable reference genes (Table 2). For the fifth instar larvae, under desiccation or rehydration, it was found that ACTIN and TUB showed the highest expression stability. According to NormFinder, the stability ranking, from most to least stable, was ACTIN > TUB > H3 > 18S rRNA > NADHD > RPS11 > AK > EF1 > UBI. NormFinder and geNorm both exhibited that UBI was the least stable reference gene (Table 2).
For the male pupae, under desiccation or rehydration, four methods of comprehensive analysis showed that TUB and ACTIN were most stable. According to NormFinder, the stability ranking, from most to least stable, was TUB > ACTIN > UBI > AK > H3 > RPS11 > 18S rRNA > EF1 > NADHD. NormFinder and geNorm both exhibited that NADHD was the least stable reference gene (Table 2). For the female pupae, under desiccation or rehydration, it was found that TUB, ACTIN, and UBI showed the highest expression stability. According to geNorm, the stability ranking, from most to least stable, was UBI/ACTIN > TUB > AK > 18S rRNA > EF1 > H3 > RPS11 > NADHD. The comparative ∆Ct method, NormFinder, Best Keeper, and geNorm all exhibited that NADHD was the least stable reference gene (Table 2).
For the male adults, under desiccation or rehydration, four methods of comprehensive analysis found that UBI and H3 showed the highest expression stability. According to geNorm, the stability ranking, from most to least stable, was RPS11/H3 > NADHD > ACTIN > UBI > TUB > 18S rRNA > AK > EF1. The comparative ∆Ct method, NormFinder, Best Keeper, and geNorm all exhibited that EF1 was the least stable reference gene (Table 2). For the female adults, under desiccation or rehydration, it was found that EF1, RPS11, and UBI showed the highest expression stability. NormFinder analysis revealed that the stability ranking (from most to least stable) was EF1 > NADHD > RPS11 > H3 > ACTIN > UBI > AK > TUB > 18S rRNA. The comparative ∆Ct method, NormFinder, Best Keeper, and geNorm all exhibited that 18S rRNA was least stable reference gene (Table 2).
3.4. Optimal Number of Reference Genes
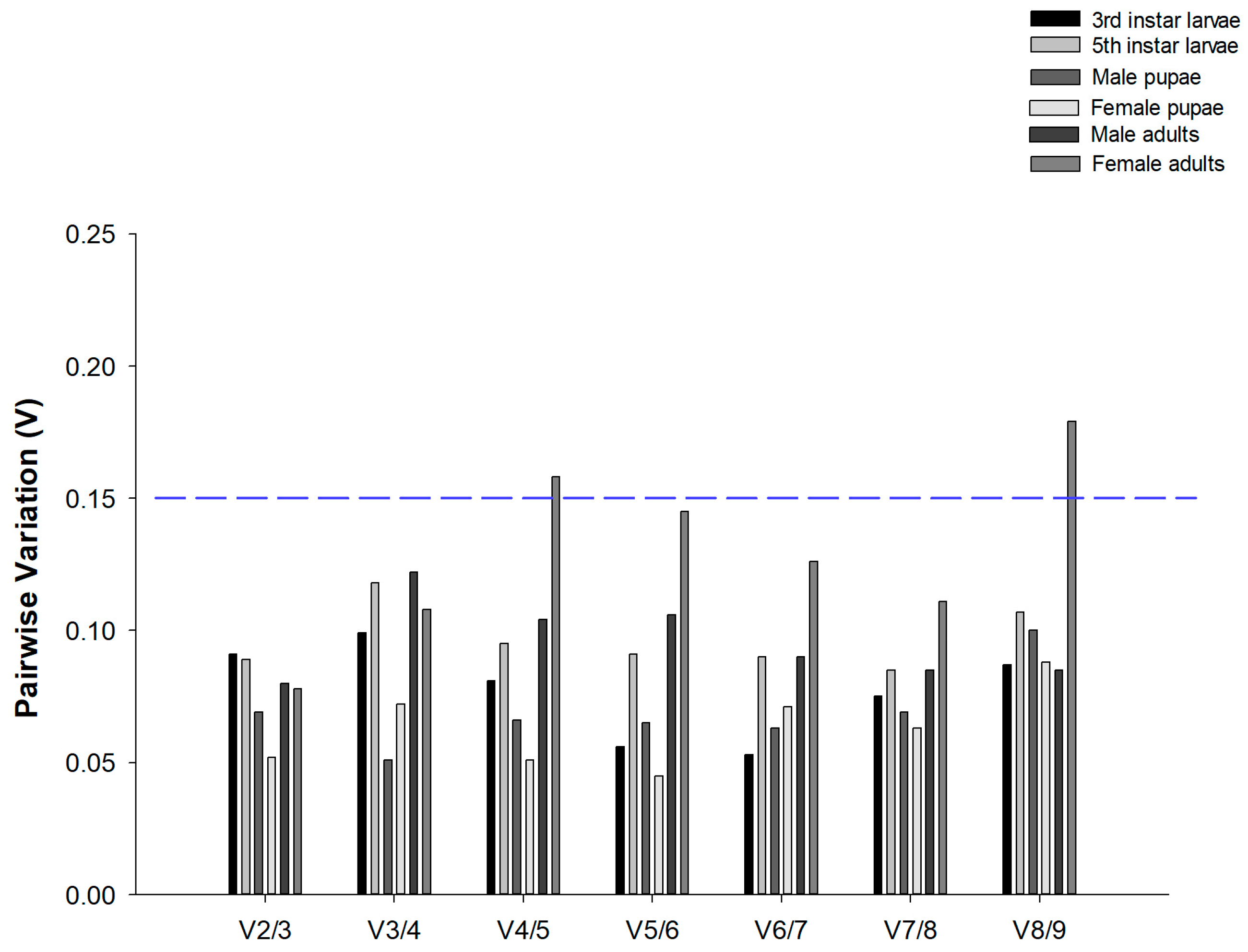
geNorm analysis also ascertained the optimal number of reference genes required for data normalization. The results revealed that pairwise variation values of V2/3 were all below the 0.15 cut-off across all developmental stages of C. suppressalis under desiccation or rehydration (Figure 2). These results indicated that two candidate reference genes were sufficient for reliable data normalization.
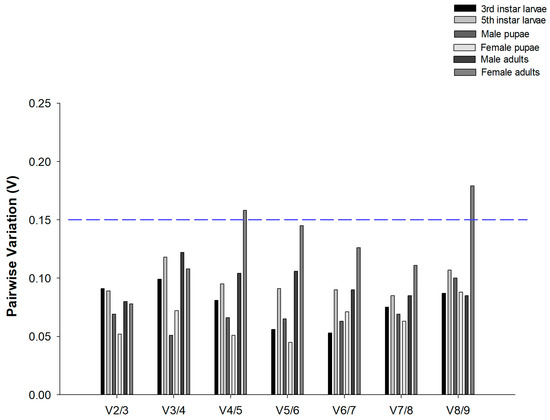
Figure 2.
Optimal number of reference genes for normalization in Chilo suppressalis. The pairwise variation (Vn/Vn + 1) was analyzed by geNorm to determine the optimal number of reference genes. When values fell below blue dashed line at 0.15, additional genes were not necessary. In total, 42 data points were subjected to analysis.
3.5. Validation of Reference Genes with Hsp60
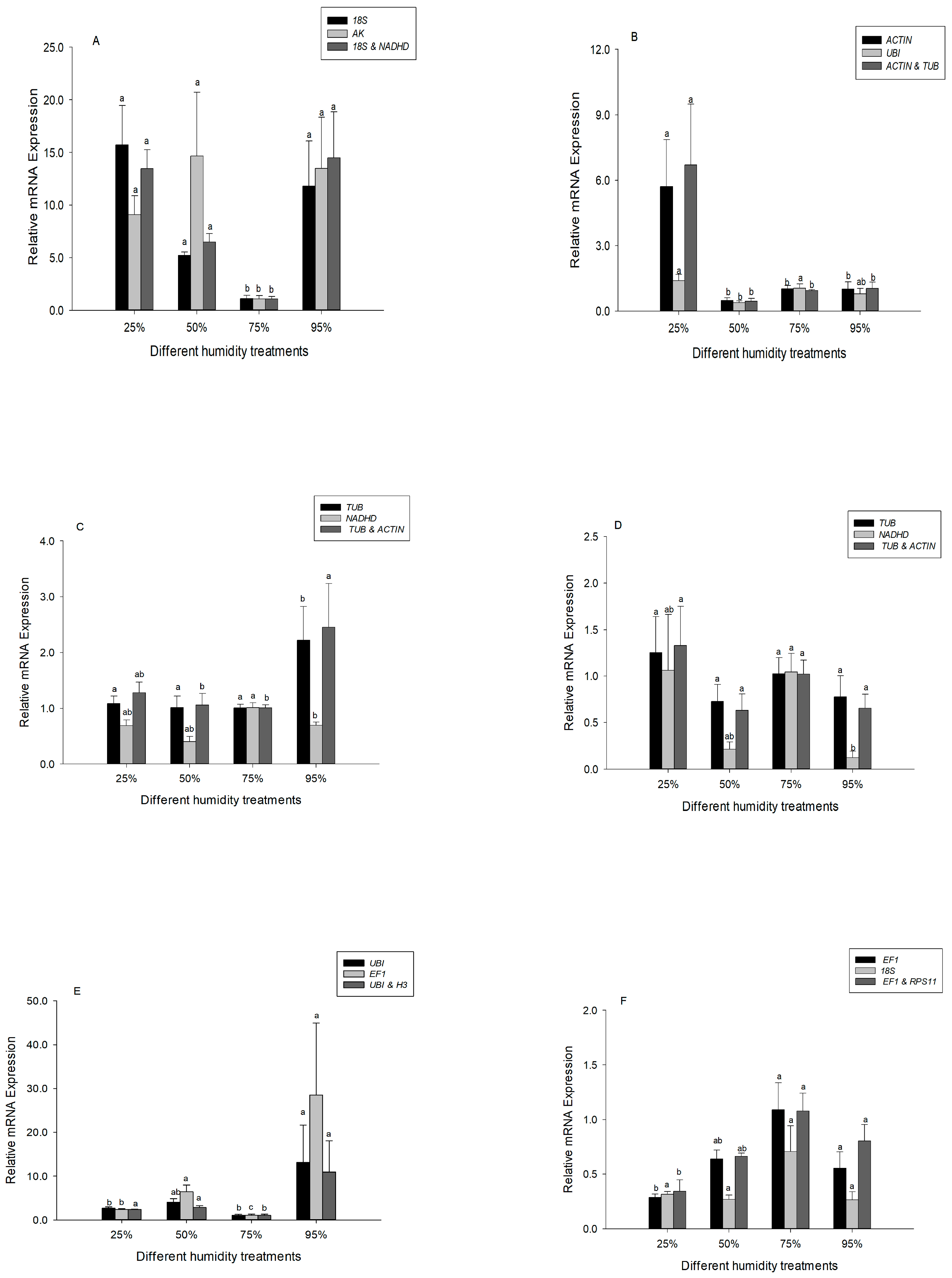
Heat shock protein 60 (Hsp60), a ubiquitous, highly abundant molecular chaperone, plays an important role in ecological adaptation. The relative expression of Hsp60 was compared with different selected reference genes under desiccation or rehydration. For the analysis of the third instar larvae, we compared expression of Hsp60 using the following selected reference genes: 18S rRNA (most stable reference gene), AK (least stable), and the two recommended reference genes (18S rRNA and NADHD). The Hsp60 expression profiles obtained using 18S rRNA, AK and the combined reference genes resulted in a similar trend (Figure 3A). However, significant differences were detected when normalizing Hsp60 expression in fifth instar larvae using ACTIN (most stable reference gene), UBI (least stable), and the combined reference genes (Figure 3B).
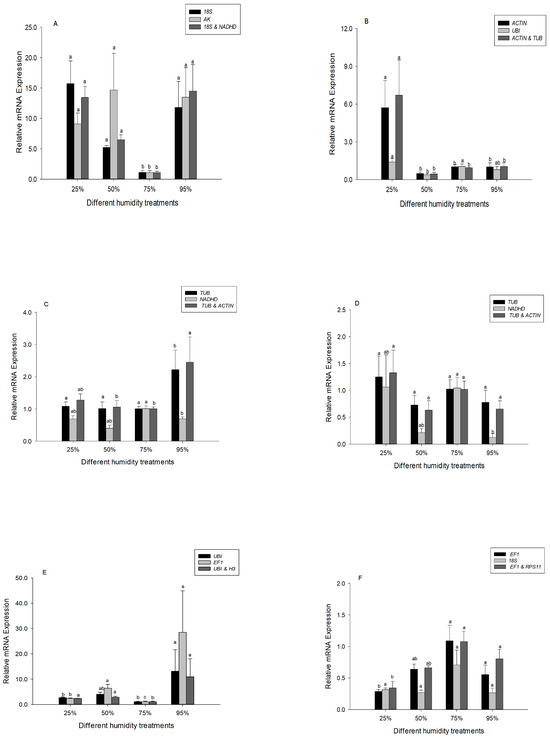
Figure 3.
Expression levels of Hsp60 in different developmental stages of Chilo suppressalis under desiccation or rehydration. (A) Expression of Hsp60 in the third instar larvae was normalized with 18S rRNA (most stable reference gene), AK (least stable), and two combined reference genes (18S rRNA and NADHD). (B) Expression of Hsp60 in the fifth instar larvae was normalized with ACTIN (most stable reference gene), UBI (least stable), and two combined reference genes (ACTIN and TUB). (C) Expression of Hsp60 in the male pupae was normalized with TUB (most stable reference gene), NADHD (least stable), and two combined reference genes (TUB and ACTIN). (D) Expression of Hsp60 in the female pupae was normalized with TUB (most stable reference gene), NADHD (least stable), and two combined reference genes (TUB and ACTIN). (E) Expression of Hsp60 in the male adults was normalized with UBI (most stable reference gene), EF1 (least stable), and two combined reference genes (UBI and H3). (F) Expression of Hsp60 in the female adults was normalized with EF1 (most stable reference gene), 18S (least stable), and two combined reference genes (EF1 and RPS11). Values are means ± SE. Columns labeled with different letters indicate significant differences in expression (p < 0.05). At least 36 data points were used for analysis in each subfigure.
For the analysis of male and female pupae, we compared expression of Hsp60 using the following selected reference genes: TUB (most stable reference gene), NADHD (least stable), and the two recommended reference genes (TUB and ACTIN). The Hsp60 expression profiles were significantly different between TUB or the combined reference genes and NADHD in male pupae (Figure 3C). In the female pupae, Hsp60 expression was not induced when TUB (most stable reference gene) or the two recommended reference genes (TUB and ACTIN) were used to normalize expression. However, when NADHD (least stable) was used for normalization, the expression profile of Hsp60 changed significantly (F3, 8 = 5.923, p = 0.020) (Figure 3D).
For the analysis of male adults, we compared expression of Hsp60 using the following selected reference genes: UBI (most stable reference gene), EF1 (least stable), and the two recommended reference genes (UBI and H3). The Hsp60 expression profiles were significantly different between the combined reference genes and EF1 (least stable) (Figure 3E). In the female adults, Hsp60 expression was not inhibited with 18S rRNA (least stable) (F3, 7 = 2.901, p = 0.111). However, when the two recommended reference genes (EF1 and RPS11) were used for normalization, the expression profile of Hsp60 was inhibited significantly (F3, 9 = 7.932, p = 0.007) (Figure 3F).
4. Discussion
qRT-PCR has become an important technique for examining gene expression levels, due to its specificity, accuracy, and reproducibility. However, the accuracy of qRT-PCR requires robust normalization with reference genes. Extensive studies indicate that there is not a single ‘universal’ reference gene across all experimental circumstances [21,22,23]. In the current study, the expression levels of nine reference genes (ACTIN, EF1, H3, 18S rRNA, AK, NADHD, TUB, RPS11, and UBI) were investigated in different developmental stages of C. suppressalis under desiccation or rehydration. These reference genes belong to different functional classes of proteins, to reduce the risk of co-regulation. Our results showed that the ∆Ct method, NormFinder, BestKeeper, and GeNorm exhibited distinct stability rankings for nine reference genes under desiccation or rehydration. Despite their differences, the four methods’ analyses resulted in similar rankings for the most and least stable reference genes under different experimental conditions; for instance, NADHD, EF1, and 18S rRNA were ranked as the least stable genes in female pupae, male adults, and female adults, exposed to desiccation or rehydration, respectively. However, most reference genes identified by the four methods of analysis were different (Table 2). Similarly, there were discrepancies in stability rankings among the different programs in previous studies, which may be attributed to distinct logical algorithms [12,23]. Therefore, in this study, a comprehensive analysis of the stability rankings of the reference genes was further used.
18S rRNA and ACTIN, involved in basic cellular processes, are most commonly used reference genes for normalizing expression. Nevertheless, some studies have demonstrated that 18S rRNA and ACTIN were not always reliable for normalizing gene expression [24,25]. In the present study, 18S rRNA and ACTIN found in the third instar larvae, as well as male or female pupae, exposed to desiccation or rehydration, are appropriate reference genes. Consistent results have been reported in other studies [26,27]. TUB plays an essential role in maintaining cell shape, regulating cell division, and facilitating substance transport. RPS11 plays an essential role in ribosomal assembly and protein translation. EF1 functions in protein translation elongation by catalyzing GTP-dependent binding of aminoacyl-tRNA. UBI is critical for protein degradation and cell cycle regulation. These genes have been increasingly used as reference genes in various insect species [12,15,28,29]. Our results also suggested that TUB, RPS11, EF1, and UBI displayed high levels of stability under different experimental conditions. The combinations of the above reference genes were the best sets of grouped genes for normalizing expression in C. suppressalis under desiccation or rehydration. To further validate the reference genes, the expression patterns of Hsp60 in different developmental stages of C. suppressalis exposed to desiccation or rehydration were analyzed. In general, the results demonstrated that the normalized expression levels of Hsp60, using the most stable reference genes or their combinations, were significantly different from those normalized using the least stable reference genes.
5. Conclusions
Insects have evolved various adaptive ecological strategies and molecular mechanisms to cope with humidity stress. However, the molecular mechanisms underlying insect adaptation to desiccation or rehydration remain poorly understood. Our study systematically investigated and validated nine potential reference genes for normalizing qRT-PCR data in C. suppressalis under desiccation or rehydration. Our findings showed that the expression stability of reference genes in C. suppressalis varied with developmental stages and sexes under humidity stress. To the best of our knowledge, this is the first study to validate reference genes in insects under desiccation or rehydration. These findings will facilitate future research on the molecular mechanisms of humidity stress adaptation in C. suppressalis.
Author Contributions
M.Z.: data curation, software, writing—original draft, funding acquisition; Y.C.: data curation, software; H.-B.Z.: data curation, software; writing—review and editing; J.-F.M.: data curation, software; Y.-J.G.: writing—review and editing. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the project of Guizhou Zhuohao Agricultural Science and Technology Co., Ltd.
Data Availability Statement
The original contributions presented in this study are included in the article. Further inquiries can be directed to the corresponding author.
Conflicts of Interest
Author Jian-Fei Mei was employed by the company Guizhou Zhuohao Agricultural Science and Technology Co., Ltd. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The authors declare that this study received funding from Guizhou Zhuohao Agricultural Science and Technology Co., Ltd. The funder had the following involvement with the study: Study on the Stability of Reference Genes and HSP60 for Expression Analysis in Chilo suppressalis in Response to Humidity Stress”.
References
- Chown, S.L.; Sørensen, J.G.; Terblanche, J.S. Water loss in insects: An environmental change perspective. J. Insect Physiol. 2011, 57, 1070–1084. [Google Scholar] [CrossRef]
- Knutelski, S.; Haranczyk, H.; Nowak, P.; Wróbel, A.; Leszczynski, B.; Okuda, T.; Strzalka, K.; Baran, E. Rehydration of the sleeping chironomid, Polypedilum vanderplanki Hinton, 1951 larvae from cryptobiotic state up to full physiological hydration (Diptera: Chironomidae). Sci. Rep. 2022, 12, 3766. [Google Scholar] [CrossRef]
- Kühsel, S.; Brückner, A.; Schmelzle, S.; Heethoff, M.; Blüthgen, N. Surface area-volume ratios in insects. Insect Sci. 2017, 24, 829–841. [Google Scholar] [CrossRef]
- Wang, Z.A.; Receveur, J.P.; Pu, J.; Cong, H.S.; Richards, C.; Liang, M.X.; Chung, H.Y. Desiccation resistance differences in Drosophila species can be largely explained by variations in cuticular hydrocarbons. eLife 2022, 11, e80859. [Google Scholar] [CrossRef]
- Chown, S.L.; Janion-Scheepers, C.; Marshall, A.; Aitkenhead, I.J.; Hallas, R.; Liu, W.P.A.; Phillips, L.M. Indigenous and introduced Collembola differ in desiccation resistance but not its plasticity in response to temperature. Curr. Res. Insect Sci. 2023, 3, 100051. [Google Scholar] [CrossRef] [PubMed]
- Lu, M.X.; He, F.J.; Xu, J.; Liu, Y.; Wang, G.R.; Du, Y.Z. Identification and physiological function of CsPrip, a new aquaporin in Chilo suppressalis. Int. J. Biol. Macromol. 2021, 184, 721–730. [Google Scholar] [CrossRef]
- Yin, C.; Liu, Y.; Liu, J.; Xiao, H.; Huang, S.; Lin, Y.; Han, Z.; Li, F. ChiloDB: A genomic and transcriptome database for an important rice insect pest Chilo suppressalis. Database 2014, 2014, bau065. [Google Scholar] [CrossRef]
- Ma, W.H.; Zhao, X.X.; Yin, C.L.; Jiang, F.; Du, X.Y.; Chen, T.Y.; Zhang, Q.H.; Lin, Q.; Xu, H.X.; Hull, J.J.; et al. A chromosome-level genome assembly reveals the genetic basis of cold tolerance in a notorious rice insect pest, Chilo suppressalis. Mol. Ecol. Resour. 2020, 20, 268–282. [Google Scholar] [CrossRef]
- Huggett, J.; Dheda, K.; Bustin, S.; Zumla, A. Real-time RT-PCR normalisation; strategies and considerations. Genes Immun. 2005, 6, 279–284. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Beaulieu, J.F.; Huggett, J.; Jaggi, R.; Kibenge, F.S.; Olsvik, P.A.; Penning, L.C.; Toegel, S. MIQE prècis: Practical implementation of minimum standard guidelines for fluorescence-based quantitative real-time PCR experiments. BMC Mol. Biol. 2010, 11, 1. [Google Scholar] [CrossRef] [PubMed]
- Bustin, S.A.; Benes, V.; Garson, J.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The need for transparency and good practices in the qPCR literature. Nat. Methods 2013, 10, 1063–1067. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.B.; Zhang, H.F.; Zhang, Z.Y.; Zhao, J.Y.; Ma, F.L.; Zheng, M.M.; Yang, M.S.; Sang, X.Y.; Ma, K.S.; Li, L.L. Selection of reference genes for normalization of qRT–PCR analysis in the soybean aphid Aphis glycines Matsumura (Hemiptera: Aphididae). J. Econ. Entomol. 2022, 115, 2083–2091. [Google Scholar] [CrossRef]
- Zhang, S.; An, S.; Li, Z.; Wu, F.; Yang, Q.; Liu, Y.; Cao, J.; Zhang, H.; Zhang, Q.; Liu, X. Identification and validation of reference genes for normalization of gene expression analysis using qRT-PCR in Helicoverpa armigera (Lepidoptera: Noctuidae). Gene 2015, 555, 393–402. [Google Scholar] [CrossRef]
- Xu, J.; Welker, D.L.; James, R.R. Variation in expression of reference genes across life stages of a bee, Megachile rotundata. Insects 2021, 12, 36. [Google Scholar] [CrossRef]
- Zheng, Y.T.; Li, H.B.; Lu, M.X.; Du, Y.Z. Evaluation and validation of reference genes for qRT-PCR normalization in Frankliniella occidentalis (Thysanoptera: Thripidae). PLoS ONE 2014, 9, e111369. [Google Scholar]
- Xu, J.; Lu, M.X.; Cui, Y.D.; Du, Y.Z. Selection and evaluation of reference genes for expression analysis using qRT-PCR in Chilo suppressalis (Lepidoptera: Pyralidae). J. Econ. Entomol. 2017, 110, 683–691. [Google Scholar]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef]
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: BestKeeper—Excel-based tool using pairwise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef]
- Andersen, C.L.; Jensen, J.L.; Ørntoft, T.F. Normalization of real-time quantitative reverse transcription-PCR data: A model-based variance estimation approach to identify genes suited for normalization, applied to bladder and colon cancer data sets. Cancer Res. 2004, 64, 5245–5250. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate normalization of real-time quantitative RT-PCR data by geometric averaging of multiple internal control genes. Genome Biol. 2002, 3, RESEARCH0034.1. [Google Scholar] [CrossRef] [PubMed]
- Li, H.B.; Dai, C.G.; Zhang, C.R.; He, Y.F.; Ran, H.Y.; Chen, S.H. Screening potential reference genes for quantitative real-time PCR analysis in the oriental armyworm, Mythimna separata. PLoS ONE 2018, 13, e0195096. [Google Scholar]
- Kong, D.; Shi, D.; Wang, C.; Zhai, R.; Lyu, L.; He, Y.; Wang, D. Identification and validation of reference genes for expression analysis using qRT-PCR in Cimex hemipterus (Hemiptera: Cimicidae). Insects 2022, 13, 784. [Google Scholar] [CrossRef]
- Yang, Q.P.; Li, Z.; Cao, J.J.; Zhang, S.D.; Zhang, H.J.; Wu, X.Y.; Zhang, Q.W.; Liu, X.X. Selection and assessment of reference genes for quantitative PCR normalization in migratory locust Locusta migratoria (Orthoptera: Acrididae). PLoS ONE 2014, 9, e98164. [Google Scholar]
- Shang, F.; Wei, D.D.; Jiang, X.Z.; Wei, D.; Shen, G.M.; Feng, Y.C.; Li, T.; Wang, J.J. Reference gene validation for quantitative PCR under various biotic and abiotic stress conditions in Toxoptera citricida (Hemiptera, Aphidiae). J. Econ. Entomol. 2015, 108, 2040–2047. [Google Scholar] [CrossRef]
- Huang, X.N.; Gao, Y.C.; Jiang, B.; Zhou, Z.C.; Zhan, A.B. Reference gene selection for quantitative gene expression studies during biological invasions: A test on multiple genes and tissues in a model ascidian Ciona savignyi. Gene 2016, 576, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Teng, X.L.; Zhang, Z.; He, G.L.; Yang, L.W.; Li, F. Validation of reference genes for quantitative expression analysis by real-time RT-PCR in four lepidopteran insects. J. Insect Sci. 2012, 12, 60. [Google Scholar] [CrossRef]
- Paim, R.M.; Pereira, M.H.; Ponzio, R.D.; Rodrigues, J.O.; Guarneri, A.A.; Gontijo, F.N.; Araújo, R.N. Validation of reference genes for expression analysis in the salivary gland and the intestine of Rhodnius prolixus (Hemiptera, Reduviidae) under different experimental conditions by quantitative real-time PCR. BMC Res. Note 2012, 5, 128. [Google Scholar] [CrossRef] [PubMed]
- Sun, M.; Lu, M.X.; Tang, X.T.; Du, Y.Z. Exploring valid reference genes for quantitative real-time PCR analysis in Sesamia inferens (Lepidoptera: Noctuidae). PLoS ONE 2015, 10, e0115979. [Google Scholar] [CrossRef]
- Yang, A.P.; Wang, Y.S.; Huang, C.; Lv, Z.C.; Liu, W.X.; Bi, S.Y.; Wan, F.H.; Wu, Q.; Zhang, G.F. Screening potential reference genes in Tuta absoluta with Real-time quantitative PCR analysis under different experimental Conditions. Genes 2021, 12, 1253. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.