Simple Summary
Spodoptera frugiperda, a significant global agricultural pest, poses a substantial threat to maize and other crops, resulting in considerable yield losses. To control this pest, CRISPR/Cas9 is considered a state-of-the-art genetic control strategy, as it significantly disrupts S. frugiperda mating communication and volatile sensing mechanisms. After overcoming the challenges associated with CRISPR/Cas9 application, it remains a powerful, new, environmentally friendly, and revolutionary platform for precise, targeted pest management in S. frugiperda.
Abstract
Globally, Spodoptera frugiperda is a major threat to many important crops, including maize, rice, and cotton, causing significant economic damage. To control this invasive pest, environmentally friendly pest control techniques, including pheromone detection and identification of potential molecular targets to disrupt S. frugiperda mating communication, are needed. Female moths biosynthesize pheromones and emit them from the pheromone gland, which significantly depends on the intrinsic factors of the moth. Male S. frugiperda have a sophisticated olfactory circuit on their antennae that recognizes pheromone blends via olfactory receptor neurons (ORNs). With its potential to significantly modify the insect genome, CRISPR/Cas9 offers a revolutionary strategy to control this insect pest. The impairing physiological behaviors and disrupting the S. frugiperda volatile-sensing mechanism are the main potential applications of CRISPR/Ca9 explored in this review. Furthermore, the release of mutant S. frugiperda for their long-term persistence must be integral to the adoption of this technology. Looking forward, CRISPR/Cas9-based gene drive systems have the potential to synergistically target pheromone signaling pathways in S. frugiperda by disrupting pheromone receptors and key biosynthesis genes, thereby effectively blocking intraspecific communication and reproductive success. In conclusion, CRISPR/Cas9 provides an environmentally friendly and revolutionary platform for precise, targeted pest management in S. frugiperda.
1. Introduction
Genome editing, commonly known as gene editing, involves inserting, deleting, labeling, or reordering genetic material to produce a desired trait through genetic manipulation [1,2]. The Clustered Regularly Interspaced Short Palindromic Repeats (CRISPR)/CRISPR-associated protein 9 (Cas9) originates from the archaeal and bacterial immune systems and is the most advanced and superior genome editing technology [3,4]. The CRISPR/Cas9 system consists of the Cas9 protein, which cleaves double-stranded DNA (dsDNA) to facilitate genome modification, and a chimeric 20 bp single-guide RNA (sgRNA) that instructs Cas9 to target the desired sequence [5]. The mechanism of CRISPR/Cas9 genome editing comprises three steps: recognition, cleavage, and repair [6]. Over the past ten years, the insect-resistant plants and various insects have been modified using CRISPR-Cas gene editing for agricultural insect pest management [7]. This gene-editing tool has altered the genomes of many insects, affecting biological and physiological traits such as pigmentation, reproduction, development, insecticide resistance, metabolism, olfaction, and body segmentation [8]. A previous study conducted by Sun et al. [9] also highlighted the application of CRISPR-Cas9 in insects and non-insect arthropods. Additionally, this technology is pivotal in advancing economic insect breeding by being applied to Bombyx mori Linnaeus, 1758, thereby enhancing disease resistance and improving silk quality [10]. In addition, CRISPR-Cas9 can disrupt pheromone recognition in insects by targeting and altering genes involved in pheromone recognition, which is recognized as the most efficient and promising, eco-friendly alternative for the sustainable management of major insect pests worldwide [11,12].
Globally, Spodoptera frugiperda is an invasive and one of the major pest insects in diverse crop plants, including maize, rice, and cotton [13], that causes significant economic damage [14]. It is native to the tropical and subtropical regions of the Americas, where it accounted for about 36% of the annual loss of maize production in sub-Saharan Africa, reducing maize production by 0.67 million tons between 2017 and 2019 [15,16]. The extensive and indiscriminate use of insecticides to control this insect pest has led to the evolution of insecticide resistance and also caused ecotoxicological effects on non-target insects [17,18]. Therefore, manipulating the olfactory-guided behaviors of insects provides new avenues for controlling insect pests [19]. Currently, disrupting pheromone recognition gene in S. frugiperda using CRISPR-Cas9-mediated genome editing is the most promising and effective pest management technique [11,12]. In S. frugiperda, the 8th and 9th intersegmental membranes produce sex pheromones, which the insect uses to find mates, identify species, and select mates [20]. Furthermore, insect pest physiological parameters, such as age and mating status, can affect pheromone production, providing valuable information for pest management [21].
A pheromone communication system is where male insects recognize the pheromone signals to identify a potential mate, ensuring reproductive success [22]. The (Z)-9-tetradecenyl acetate (Z9–14:OAc), a crucial sexual communication signal produced by S. frugiperda females [23,24,25,26], can be detected by males even at minute concentrations [27]. Olfactory receptor neurons (ORNs) are dispersed across several appendages on the antennae and express obligate odorant receptor co-receptors (Orco) [28,29] coupled with a “tuning” odorant receptor (OR) to form heteromeric, odor-gated ion channels in the membranes of these neurons to facilitate the chemosensory process [30,31,32]. These results provide a solid foundation for understanding advances in the molecular mechanisms of S. frugiperda sex pheromone biosynthesis and identify new targets for developing novel pest control methods that disrupt sexual communication. Currently, the olfactory-based pest control strategies are considered most promising and effective, in which various behavioral processes were impaired by mutagenesis of Orco across multiple insect pests such as World screwworm, Cochliomyia hominivorax, Coquerel [33], cotton bollworm, Helicoverpa armigera Hubner [34] hawkmoth, Manduca sexta Linnaeus [35]. However, research on Orco mutagenesis in S. frugiperda remains limited; therefore, this review focuses on the biosynthesis, release, and recognition of sex pheromones in S. frugiperda, elucidating the role of the olfactory system in mediating critical behaviors such as mating, foraging, and oviposition. Furthermore, we have evaluated the potential of CRISPR/Cas9-mediated gene editing, particularly Orco disruption, in functionally characterizing pheromone receptor pathways and altering behavioral response.
2. Pheromone Dynamics and Recognition in S. frugiperda
2.1. Biosynthesis of Sexual Pheromone in S. frugiperda
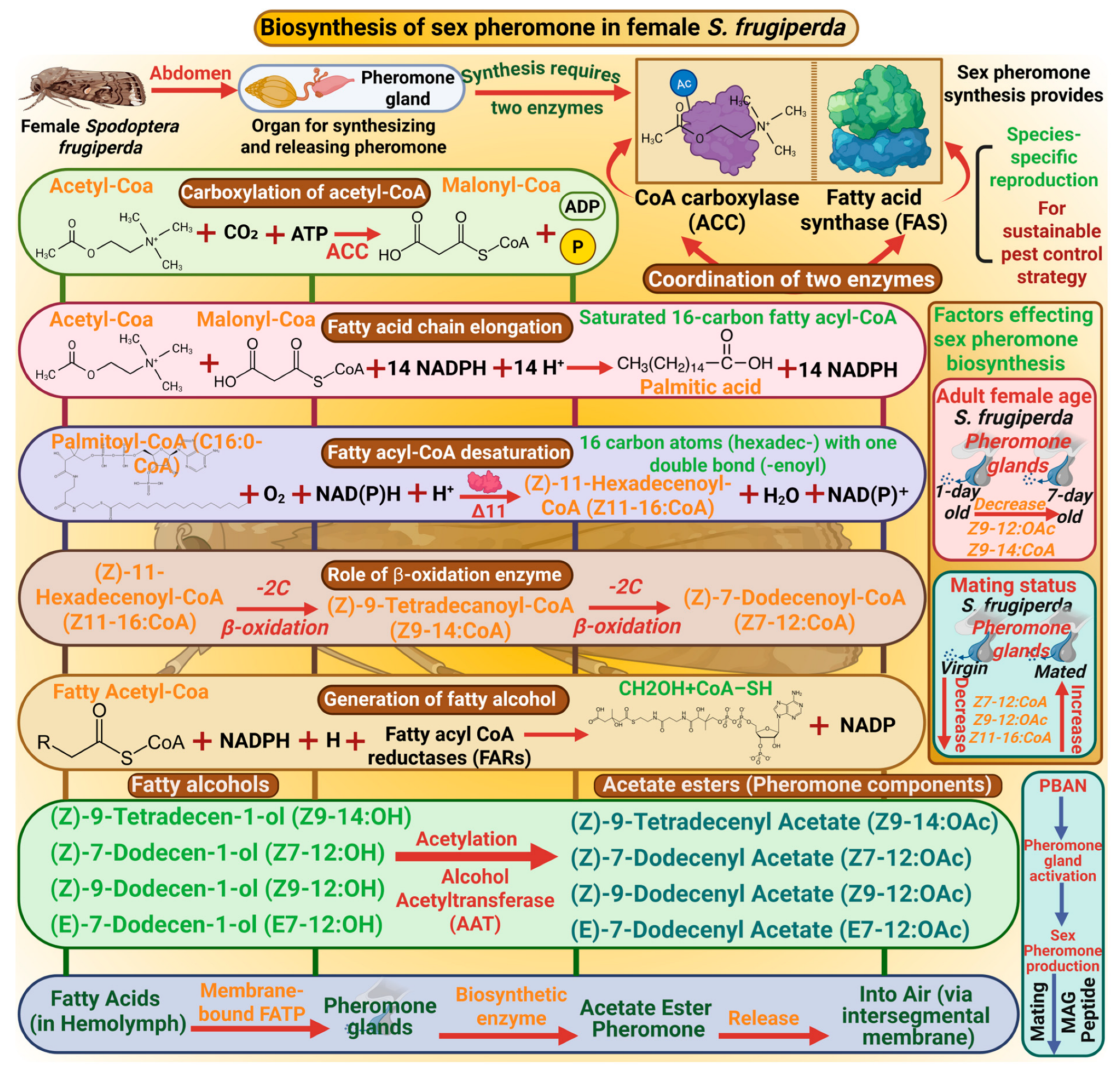
The biosynthesis of sexual pheromones in S. frugiperda is central to understanding species-specific reproduction and offers a vital basis for sustainable pest control strategies. In S. frugiperda, pheromone glands are present in the abdomen, essential organs for synthesizing and releasing pheromones [36]. The synthesis of pheromones requires the coordination of different types of enzymes, including acetyl-CoA carboxylase (ACC) and fatty acid synthase (FAS). ACC catalyzes the carboxylation of acetyl-CoA to malonyl-CoA [37]. Then FAS catalyzes the condensation of malonyl CoA units with NADPH as a reducing agent to elongate the fatty acid chain and produces the palmitic acid (C16:0-CoA), a saturated 16-carbon fatty acyl-CoA [38]. Thus, fatty acyl-CoA desaturation occurred by enzyme Δ11 desaturase to generate unsaturated fatty acids such as (Z)-11-hexadecenoyl-CoA (Z11–16-CoA) by introducing a double bond at the 11th carbon of palmitic acid. After this desaturation, beta-oxidation enzymes shorten the fatty acyl-CoA chain by removing two carbon units, producing intermediate molecules such as Z9–14:OAc and (Z)-7-dodecenyl acetate (Z7–12:CoA). These precursors are then reduced by fatty acyl CoA reductases (FARs) to generate fatty alcohols, which are important intermediates in pheromone formation. Furthermore, Alcohol Acetyltransferase (AAT) enzyme catalyzes acetylation to produce acetate esters such as (Z)-9-tetradecenyl acetate (Z9–14:OAc), (Z)-7-dodecenyl acetate (Z7–12:OAc), (Z)-9-dodecenyl acetate (Z9–12:OAc), and (E)-7-dodecenyl acetate (E7–12:OAc), which are ultimate pheromone components [39]. A membrane-bound fatty acid transport protein carries fatty acids through hemolymph to the pheromone gland, synthesizing pheromones [24]. The female moth emits pheromone from the pheromone gland which is located at the intersegmental membrane between the 8th and 9th abdominal segments [40] (Figure 1).
2.2. Factors Affecting Sex Pheromone Release from Female S. frugiperda
Age and mating status are the most promising factors influencing the release of sex pheromones from female S. frugiperda, and these factors are explained in detail.
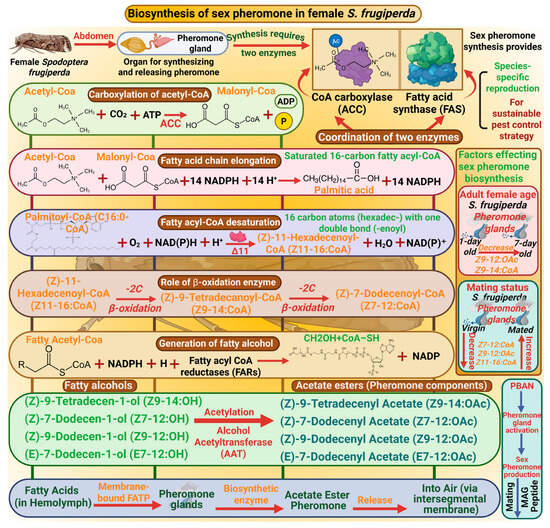
Figure 1.
Biosynthesis of the sex pheromone in female Spodoptera frugiperda involves key enzymes, such as CoA carboxylase and fatty acid synthase, which coordinate fatty acid synthesis. These fatty acids are desaturated and oxidized, then converted into fatty alcohols and acetate esters, which are the main pheromone components. This process is regulated by PBAN (pheromone biosynthesis-activating neuropeptide), which activates the pheromone production and attracts males for mating. “Figure created using Biorender.com”.
Figure 1.
Biosynthesis of the sex pheromone in female Spodoptera frugiperda involves key enzymes, such as CoA carboxylase and fatty acid synthase, which coordinate fatty acid synthesis. These fatty acids are desaturated and oxidized, then converted into fatty alcohols and acetate esters, which are the main pheromone components. This process is regulated by PBAN (pheromone biosynthesis-activating neuropeptide), which activates the pheromone production and attracts males for mating. “Figure created using Biorender.com”.
2.2.1. Adult Female Age
Insect aging is the crucial factor that influences the release of pheromone. In this context, Akter et al. [41] investigated the pheromone glands of females S. frugiperda between the 4th and 6th hours of the scotophase. After extracting glands of varying female ages, it was found that the release of volatile compounds such as Z9–12:OAc and Z9–14:CoA was higher in one-day-old females compared with four-day and seven-day-old females (p < 0.001 and p < 0.01, respectively). Therefore, aging has a negative impact on the pheromone release from S. frugiperda females (Figure 1). Furthermore, while collecting headspace volatiles from female moths of various ages also indicated that younger females released and produced higher amounts of compounds than older ones [41]. Similarly, Domínguez et al. [21] reported that pheromone release from glands is significantly reduced as the female age increases. Because pheromone production is governed by the endocrine system, aging in insects negatively influences the endocrine system by decreasing juvenile hormone production from the corpora allata, reducing the sensitivity of both endocrine tissues and pheromone receptor cells, diminishing ovarian activity and weekend neural regulation of endocrine glands, and ultimately reducing pheromone production [42,43]. Consequently, one-day-old females release pheromones with enhanced mate-attraction potential and achieve higher mating success rates than older individuals. Therefore, these findings suggest that younger females produce more pheromones to boost reproductive fitness and mating chances.
2.2.2. Mating Status
Mating dramatically alters the pheromone release, often reducing or stopping sexual activity in females to avoid further mate attention and focused reproduction. Therefore, Akter et al. [41] revealed that mated females had retained significantly higher concentrations of headspace volatiles, including Z7–12:CoA, Z9–12:OAc, and Z11–16-CoA, in their glands compared to virgin females. The production of these volatile sex pheromones is regulated by the pheromone biosynthesis-activating neuropeptide (PBAN), which is predominantly produced by neurosecretory cells of the subesophageal ganglion and transported to the corpora cardiaca before its release into the hemolymph [44]. Mating triggers the transfer of a sex peptide, a pheromone depressant peptide, from the male’s accessory gland (MAG) to the female genital tract, ultimately triggering juvenile hormone production in the corpus allatum in female S. frugiperda [45,46]. The production of juvenile hormone inhibits the release of PBAN, resulting in decreased sex pheromone production in females and suppressing female receptivity, suggesting that mating significantly affects the release of pheromone compounds from glands by suppressing it, possibly as a mechanism to prevent attracting additional males after mating [47]. On the other hand, virgin S. frugiperda releases significantly higher quantities of pheromone compounds than mated females to secure mating first, suggesting that mating status directly modulates both the production and release of sex pheromones in S. frugiperda (Figure 1).
2.3. Mechanism of Pheromone Recognition by Male S. frugiperda
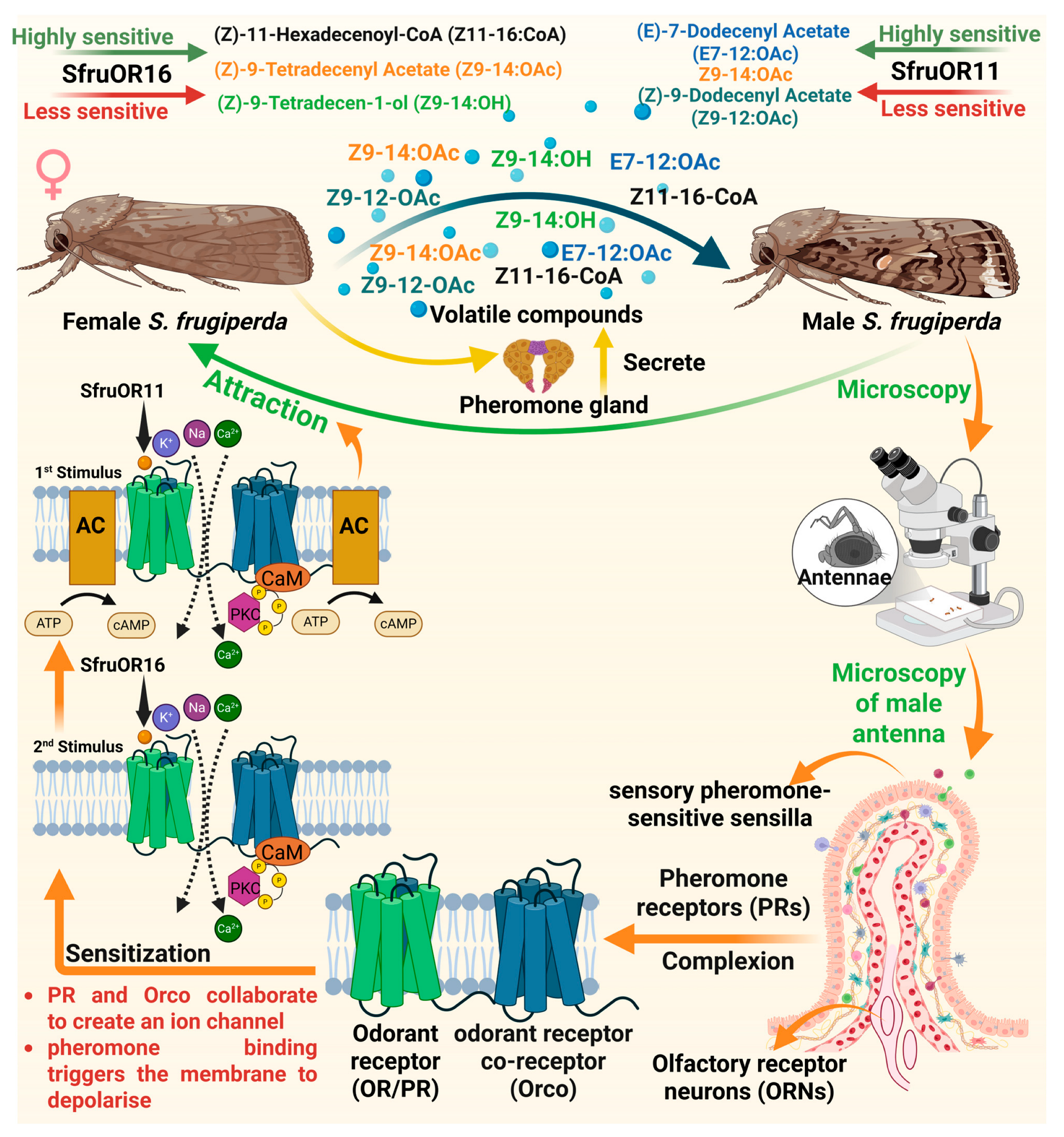
Pheromones consist of blend of several volatile compounds which are species-specific and secreted by the female S. frugiperda [48]. The female S. frugiperda produces Z11–16-OAc, Z9–12:OAc, Z9–14:OAc, Z9–14:OH and E7–12:OAc in their pheromone gland that are attractive to the male. These pheromone components are detected by the ORNs located in the sensory pheromone-sensitive sensilla, carried by the male antennae [49]. The ORNs detect pheromone components through a specific class of ORs known as pheromone receptors (PRs) in S. frugiperda [50]. Furthermore, ORs are seven-transmembrane receptors located on the dendrites of ORNs in S. frugiperda, and they function by forming complexes with a co-receptor called Orco [51,52]. PR and Orco collaborate to create an ion channel that opens when pheromones bind to it, triggering the membrane to depolarize [53,54]. Resultantly, the pheromone signal is transformed into an electrical signal that is transmitted to the brain for processing and integration, which then leads to the attraction behavior [48,49,50,51,52,53,54,55]. The PR, Spodoptera frugiperda odorant receptor 11 (SfruOR11), is highly sensitive to Z9–14:OAc and E7–12:OAc, which are the major sex pheromone components released by the female S. frugiperda [56]. Whereas less sensitive to Z9–12:OAc, which is the minor pheromone component [56]. Spodoptera frugiperda odorant receptor 16 (SfruOR16) responds weakly to pheromone components, Z9–14:OAc and Z11–16-CoA, but strongly to pheromone analog, Z9–14:OH. When these pheromone components bind to PRs, they trigger a large inward ion current, indicating strong receptor activation [56]. This receptor activation initiates the conversion of chemical signals into electrical signals, which are essential for accurate detection and the subsequent attraction behavior in male S. frugiperda (Figure 2).
Dose-Dependent Response of Male S. frugiperda to Sex Pheromones
The response of male S. frugiperda to sex pheromones is significantly influenced by the dose of pheromone components. In this context, Malo et al. [57] determined the antennal response of male S. frugiperda to female sex pheromone using electroantennography and observed varying antennal activity. The lower doses (0.01 µg) of sex pheromone components such as Z9–14:OAc and Z9, E12–14:OAc produce substantially weaker responses than the higher dose (10 µg), which produces the strongest response (Figure 2), indicating the optimal biosensing ability of male antennae. Furthermore, this differential responsiveness suggests a highly tuned olfactory system capable of detecting minute variations in pheromone concentration, presumably enabling males to locate conspecific females more precisely under varying signal intensity levels.
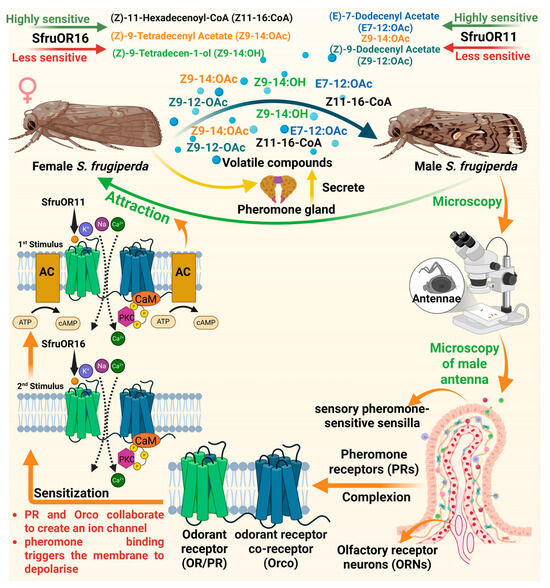
Figure 2.
Mechanism of pheromone recognition by male Spodoptera frugiperda. The volatile compounds such as Z11–16:CoA, Z9–14:OAc, Z9–14:OH, E7–12:OAc, Z914:OAc and Z9–12:OAc emitted by female moths attract male moths. These are detected by pheromone receptors (PRs) on the antennae, where ion channels such as SfruOR11 and SfruOR16 trigger a dose-dependent response, sensitized by Orco, leading to membrane depolarization and behavioral attraction. “Figure created using Biorender.com”.
Figure 2.
Mechanism of pheromone recognition by male Spodoptera frugiperda. The volatile compounds such as Z11–16:CoA, Z9–14:OAc, Z9–14:OH, E7–12:OAc, Z914:OAc and Z9–12:OAc emitted by female moths attract male moths. These are detected by pheromone receptors (PRs) on the antennae, where ion channels such as SfruOR11 and SfruOR16 trigger a dose-dependent response, sensitized by Orco, leading to membrane depolarization and behavioral attraction. “Figure created using Biorender.com”.
3. Mechanisms of CRISPR/CAS Editing in Insects
3.1. Basic Mechanism of CRISPR/Cas System
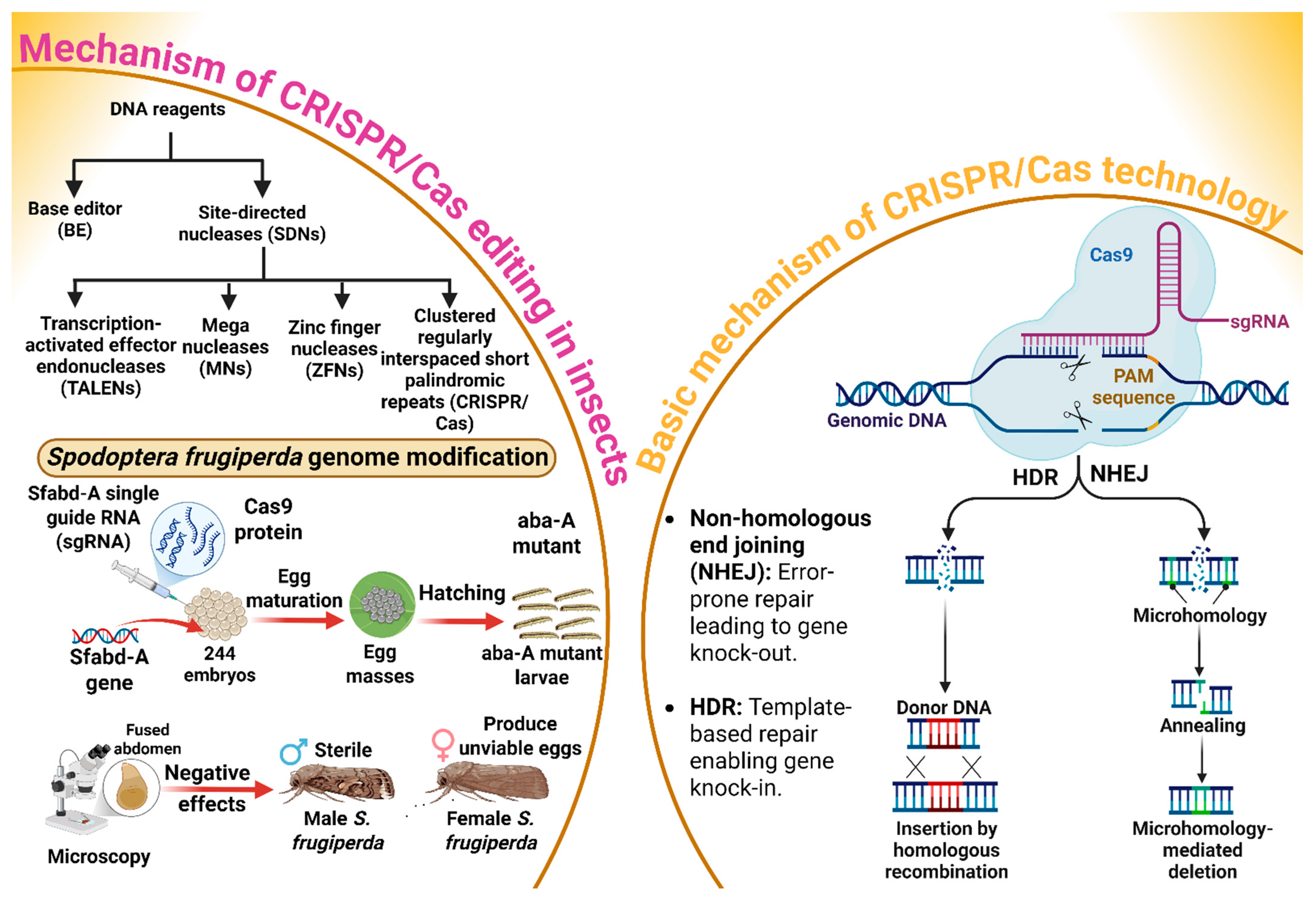
CRISPR/Cas genome editing has evolutionary roots in the adaptive immune system of bacteria and archaea, where it functions as a defense mechanism against foreign genetic elements [58]. The basic mechanism of gene editing through the CRISPR/Cas system is explored in Figure 3. Base editors (BE) and site-directed nucleases (SDNs) are major proteins or DNA reagents that cause gene editing [59]. An SDN typically consists of a nuclease domain and the DNA-binding domain, which work together to create targeted breaks in DNA [58]. The DNA-binding domain identifies and binds to a specific DNA sequence, while the nuclease domain, often from the Fokl enzyme, cuts the DNA at that site [58]. Furthermore, transcription-activated effector endonucleases (TALENs), mega nucleases (MNs), zinc finger nucleases (ZFNs), and CRISPR/Cas are the four main types of SDNs used for gene editing [60]. The CRISPR/Cas9 genome-editing system relies on two components: the Cas9 protein, an endonuclease that introduces precise double-stranded breaks in DNA, and synthetic (sgRNA), which is engineered to direct Cas9 to specific sites or loci [61]. Two major pathways are used in this process. One is non-homologous end joining (NHEJ), which introduces the insertion or deletion of a targeted gene at the cleavage site, potentially disrupting the coding sequence of the targeted gene and leading to gene knock-out. Conversely, homology-directed repair (HDR) offers a precise alternative by utilizing a homologous DNA template to repair the break, thereby enabling targeted insertion called knock-in [61]. An extension of this technology is Co-CRISPR/Cas, in which multiple sgRNAs are required concurrently to target several loci within the genome. This multiplexed approach not only increases the throughput of gene editing but also accelerates the functional annotation of genes by facilitating simultaneous edits [62,63]. In this context, Wu et al. [64] explored the possibility of using the CRISPR/Cas9 system to modify the genome of S. frugiperda through the procedure mentioned in Figure 3. The results indicated that embryos injected with the Spodoptera frugiperda abdominal-A homeotic (Sfabd-A) sgRNA developed into larvae that displayed typical aba-A mutant phenotypes, including fused segments. Surprisingly, besides immatures, the adult S. frugiperda were also prone to the negative effects of mutation, so that males were sterile and females produced unviable eggs. These results suggest that the S. frugiperda genome can be effectively modified through the CRISPR/Cas system, demonstrating its potential as a powerful tool for inducing precise mutations (Figure 3). The successful generation of aba-A mutant phenotypes in larvae, coupled with the observed detrimental effects on adult fertility, highlights the system’s significant promise for advancing functional genomics. Moving forward, research should prioritize broadening the application of CRISPR/Cas9 across a wider array of insect species, particularly by adopting multiplexed CRISPR strategies, such as Co-CRISPR, to accelerate the functional annotation of genomes. Moreover, optimizing the system’s efficiency and precision while thoroughly investigating the long-term consequences of induced mutations will be crucial for fully realizing the potential of CRISPR-based technologies for agricultural innovation and pest control.
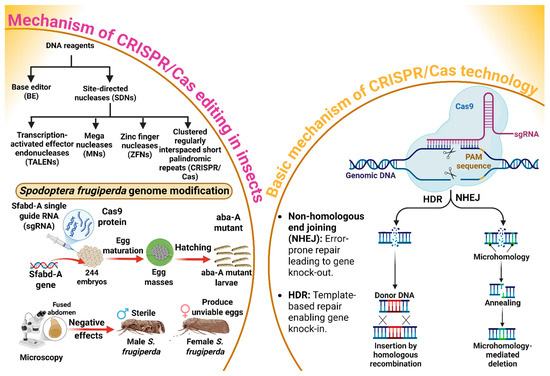
Figure 3.
CRISPR/Cas9 gene editing in Spodoptera frugiperda involves using specific DNA reagents, such as base editors and site-directed nucleases, to modify the genome. The process focuses on editing the Sfabd-A gene, leading to the production of sterile male moths and unviable eggs. This mutation impacts the insect’s reproductive capabilities, which is key in pest control strategies. The CRISPR/Cas9 system operates through two primary DNA repair mechanisms: Non-homologous end joining (NHEJ) for gene knockouts, and homology-directed repair (HDR) for precise gene knockins. “Figure created using Biorender.com”.
3.2. Application in Insects
CRISPR/Cas system has modified the genetic engineering process in biotechnology in various organisms and established genetically modified animal and cell models of many human diseases [65]. This technology provides high precision, versatility and efficiency in genome editing for functional genomics and genetic engineering in insects. This tool can alter the function of heritable genes by targeting the domains of the white gene (Ex2Ex5) in spotted wing Drosophila, Drosophila suzukii (Matsumura), resulting in disruption of pigmentation and visible phenotypic changes, especially in eye color [66]. Moreover, another study conducted by Lamb et al. [67] revealed that gene editing in D. suzukii can cause alterations in various characteristics, including body pigmentation, eye color, and wing fluorescence. Besides providing insights into gene function, CRISPR/Cas technology is also helpful for precisely editing the DNA of insect pests to suppress their populations or for developing novel insect genetic control strategies that cause developmental abnormalities [68]. Structural abnormalities in abdominal segmentations of tobacco cutworm, Spodoptera litura, Fabricius, were observed when manipulating the function of Slabd-A, a homolog of the abdominal-A gene [69]. Moreover, Zuo et al. [70] used the CRISPR/Cas9 as a tool to functionally validate resistance mutations in Beet armyworm, Spodopera exigua, Hubner by introducing a specific mutation in the ryanodine receptor gene, and this mutation confers high resistance to diamide insecticides, specifically chlorantraniliprole, cyantraniliprole and flubendiamide, demonstrating how the CRISPR/Cas9 system has revolutionized genetic engineering in insects by enabling precise modifications that not only advance our understanding of gene function but also hold promise for developing novel insect pest control strategies.
4. CRISPR/Cas-Mediated Disruption of the Olfactory System in Spodoptera frugiperda
4.1. Identification and Function of Key Olfactory Genes in Spodoptera frugiperda
PRs refer to the members of the OR superfamily that facilitate intraspecific sexual communication between males and females, S. frugiperda [71]. The identification of functionally active PRs provides a key step in mate location and reproductive success (Figure 4) [72]. In this context, Guo et al. [56] functionally characterize the PRs in male antennae of S. frugiperda and found six candidate PRs (Spodoptera frugiperda odorant receptor 6 (SfruOR6), SfruOR11, Spodoptera frugiperda odorant receptor 13 (SfruOR13), SfruOR16, Spodoptera frugiperda odorant receptor 56 (SfruOR56), and Spodoptera frugiperda odorant receptor 62 (SfruOR62). Furthermore, Zhang et al. [71] first identified novel ORs (Spodoptera frugiperda odorant receptor 5 (SfruOR5), Spodoptera frugiperda odorant receptor 18 (SfruOR18), Spodoptera frugiperda odorant receptor 22 (SfruOR22), Spodoptera frugiperda odorant receptor 23 (SfruOR23), Spodoptera frugiperda odorant receptor 30 (SfruOR30), Spodoptera frugiperda odorant receptor 36 (SfruOR36), Spodoptera frugiperda odorant receptor 38 (SfruOR38), Spodoptera frugiperda odorant receptor 41 (SfruOR41), Spodoptera frugiperda odorant receptor 49 (SfruOR49), and Spodoptera frugiperda odorant receptor 52 (SfruOR52) responding to the sex pheromones in S. frugiperda because PRs fall into a single monophyletic clade in the OR family. These PRs are not exclusively found in male insect antennae; while they are more abundant and specialized in males, females also possess PRs for autodetection of their pheromones [73]. However, the expression of PR genes varies depending on the gender of the insect in such a way that expression of SfruOR6, SfruOR11, SfruOR13, SfruOR16, SfruOR56, and SfruOR62 genes was higher in male antennae than female antennae because male antennae have more sensilla than female antennae [74,75]. The functional characterization of sex-biased PRs in S. frugiperda, with predominant expression in male antennae, provides promising targets for blocking the signal and its perception, thereby enhancing the efficiency of genetic pest management.
The ionotropic receptors (IRs) are also the most important family of insect olfactory receptors, which are widely expressed in both olfactory organs of insects [76]. IRs are derived from ionotropic glutamate receptors (iGluRs), are composed of 600–1000 amino acids, and have an architectural arrangement of two extracellular ligand-binding domains and three transmembrane domains [76]. In S. frugiperda, Sun et al. [77] identified expression of 10 IRs across different anatomical structures in male and female individuals, involved in sensory and reproductive functions. These IRs includes Spodoptera frugiperda ionotropic receptor 25a (SfruIR25a), Spodoptera frugiperda ionotropic receptor 60a (SfruIR60a), Spodoptera frugiperda ionotropic receptor 75d (SfruIR75d), Spodoptera frugiperda ionotropic receptor 64a (SfruIR64a), Spodoptera frugiperda ionotropic receptor 40a (SfruIR40a), Spodoptera frugiperda ionotropic receptor 100 (SfruIR100), Spodoptera frugiperda ionotropic receptor 41a (SfruIR41a), Spodoptera frugiperda ionotropic receptor 75p (SfruIR75p), Spodoptera frugiperda ionotropic receptor 76b (SfruIR76b), Spodoptera frugiperda ionotropic receptor 75q.1 (SfruIR75q.1). Furthermore, all these IRs majorly contribute to olfaction due to the expression of these IRs (except SfruIR60a) in male and female antennae rather than male and female proboscises, male and female tarsi, and female pheromone gland ovipositor. The gustatory receptors (GRs) are also the most important family of insect olfactory receptors, crucial for the peripheral coding of non-volatile compounds and essential for multiple behaviors including feeding [78]. For feeding S. frugiperda adults taste food by utilizing the GRs Spodoptera frugiperda gustatory receptor 1 (SfruGR1) and Spodoptera frugiperda ionotropic receptor 64a (SfruGR2), which are mainly expressed in male and female maxillae. In contrast, the Spodoptera frugiperda ionotropic receptor 64a (SfruGR3) and Spodoptera frugiperda ionotropic receptor 64a (SfruGR9) are involved in larval gustation [79]. The general odorant-binding proteins (GOBPs) are another class of olfactory proteins that serve as the link between external odorant molecules and ORs [80]. The insect GOBPs are small in size, with 15–17 kDa soluble proteins and concentrated in the lymph of chemosensory sensilla [81]. In S. frugiperda, the GOBPs are expressed at higher levels in adults than in other developmental stages and bind general plant volatiles and insecticides, ultimately providing a basis for controlling this insect pest. This knowledge can help explore plant volatile-sensing mechanisms and improve resistance to insecticides [82]. Furthermore, the Spodoptera frugiperda general odorant-binding protein 1 (SfruGOBP1) exhibits a lower binding affinity than Spodoptera frugiperda general odorant-binding protein 2 (SfruGOBP2), because the former binds to 4 of 38 general tested plant volatiles and 3 of 7 insecticides. In contrast, the latter binds to 21 volatiles and 4 insecticides. SfruGOBP2 has more amino acid residues (E28, E31, E42, H7, S56, S79) compared to SfruGOBP1 (F33, S56, and S65) showed a potentially strong binding affinity [82]. Besides the role of GOBPs in olfaction, the non-sensory OBP, odorant-binding protein 27 (OBP27), is highly expressed in the S. frugiperda fat body and mediates the transport of lipid metabolites during the eclosion process. Furthermore, it also supports the physiological function of S. frugieda due to its abundance in male reproductive organs, ultimately promoting mating [83]. Similarly to the OBPs, the chemosensory proteins (CSPs) are water-soluble, acidic proteins of about with a lenfht of 13–17kDa of approximately 120 to 170 amino acid residues [84,85], with an α-helical domain that forms a hydrophobic cavity [86]. In S. frugiperda, twenty-two Spodoptera frugiperda chemosensory proteins (SfruCSPs) with amino acid lengths between 107 and 233 and highly expressed in the larval or egg stage, indicating their functional complementation [87]. The larval inner endocuticle and outer epicuticle of S. frugiperda are also enriched with Spodoptera frugiperda chemosensory protein 1 (SfruCSP1) and Spodoptera frugiperda chemosensory protein 2 (SfruCSP2), which exhibited a high binding affinity with insecticides, chlorfenapyr, chlorpyrifos, and indoxacarb. Therefore, knocking down these proteins will significantly enhance the penetration of insecticides and increase S. frugiperda’s susceptibility to them [88].
4.2. Practice and Effects of CRISPR/Cas9-Mediated Orco Knockout
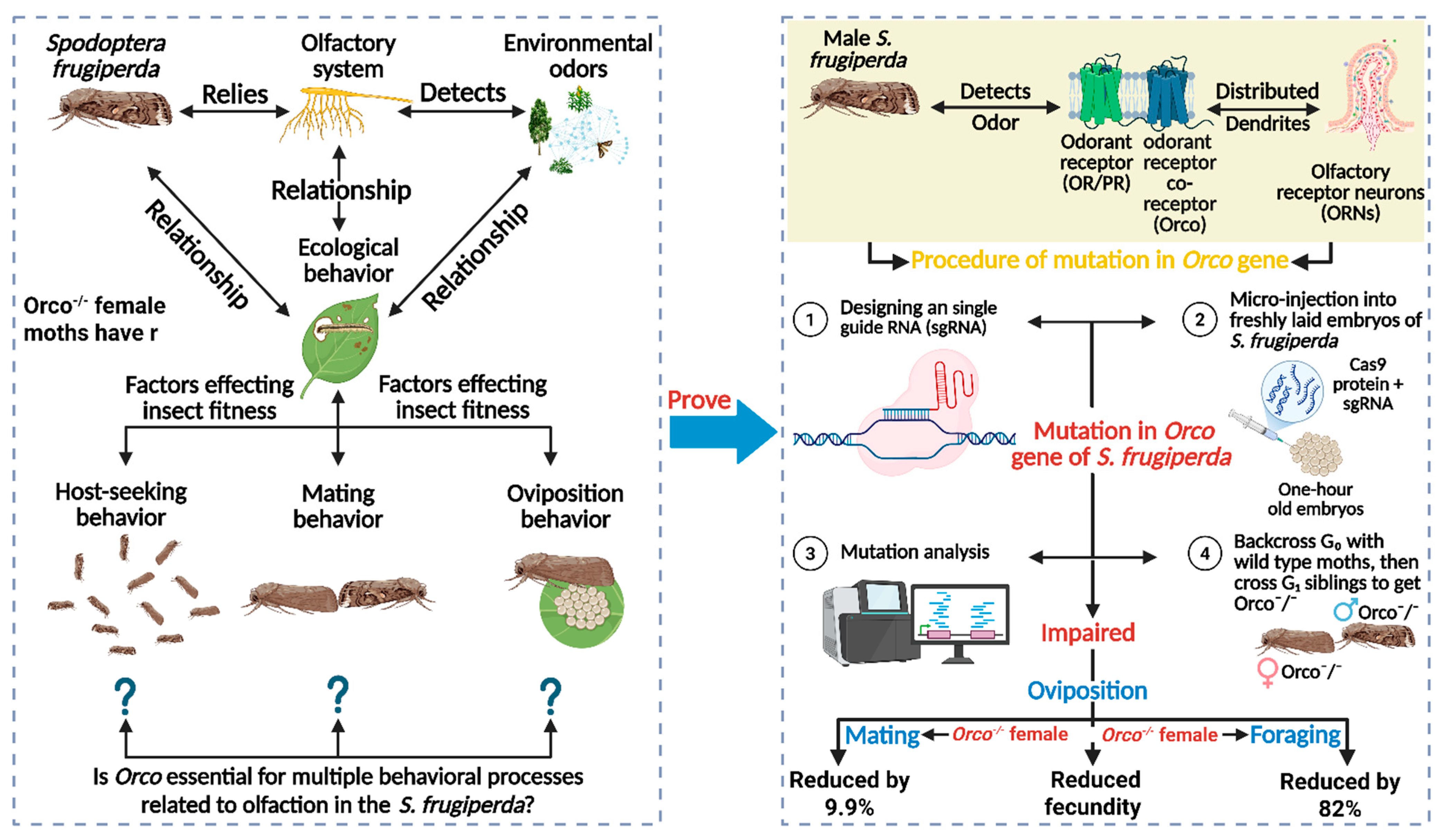
The S. frugiperda relies exclusively on its olfactory system to detect the environmental odors, necessary for its ecological behavior, such as host-seeking, mating, and egg laying [89,90,91]. For the detection of odors from the environment, ORs in the olfactory system play a significant role, which are distributed on the dendrites of ORNs [92]. The Orco is an obligatory component that is required for dimerization with ORs through an intracellular loop to form a ligand-gated ion channel, which together is activated by odor molecules [93,94]. For mutating this Orco gene, Sun et al. [95] applied the CRISPR/Cas9 genome editing tool by designing an sgRNA targeting exon 2 using the ZiFit Tool. After combining the synthesized sgRNA with Cas9 protein, they were microinjected into freshly laid one-hour-old embryos of S. frugiperda. Then, injected embryos were incubated for hatching, and genomic DNA from larvae was screened for mutations using PCR and sequencing. A 4 bp deletion was identified, causing a frameshift and an early stop codon, resulting in a non-functional Orco protein. Mutant lines were established by backcrossing G0 individuals with wild-type moths, followed by screening G1 and developing homozygous Orco−/− mutants through G1 sibling crosses. Orco gene was successfully knocked out, confirmed by quantitative polymerase chain reaction (qPCR), showing near-absent Orco mRNA expression in mutants [95]. Resultantly, the physiological behavior, such as mating, oviposition and foraging behaviors of S. frugiperda was significantly impaired in such a way that the response of Orco−/− male moths to two sex pheromones, Z9–14: Ac and Z7–12: Ac, were abolished and therefore, Orco−/− male and female moths reduced the mating frequency by 100% as compared to wild type. Furthermore, Orco−/− female moths have reduced fecundity on the maize host plant as compared to wildtype, as well as Orco−/− larvae locate the food source 12.3 min longer than wild type larvae (2.2 min). Overall, the mutation in Orco causes impaired behavior in both larval and adult S. frugiperda (Figure 5), offering a precise, eco-friendly pest control strategy and reducing reliance on conventional insecticides.
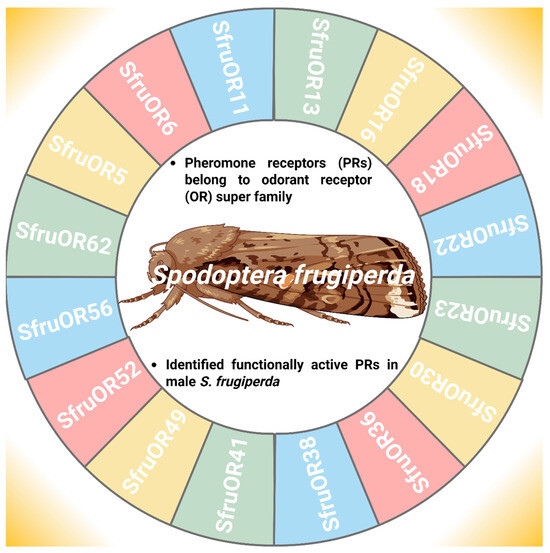
Figure 4.
Odorant receptor genes involved in pheromone detection in Spodoptera frugiperda. “Figure created using Biorender.com”.
Figure 4.
Odorant receptor genes involved in pheromone detection in Spodoptera frugiperda. “Figure created using Biorender.com”.
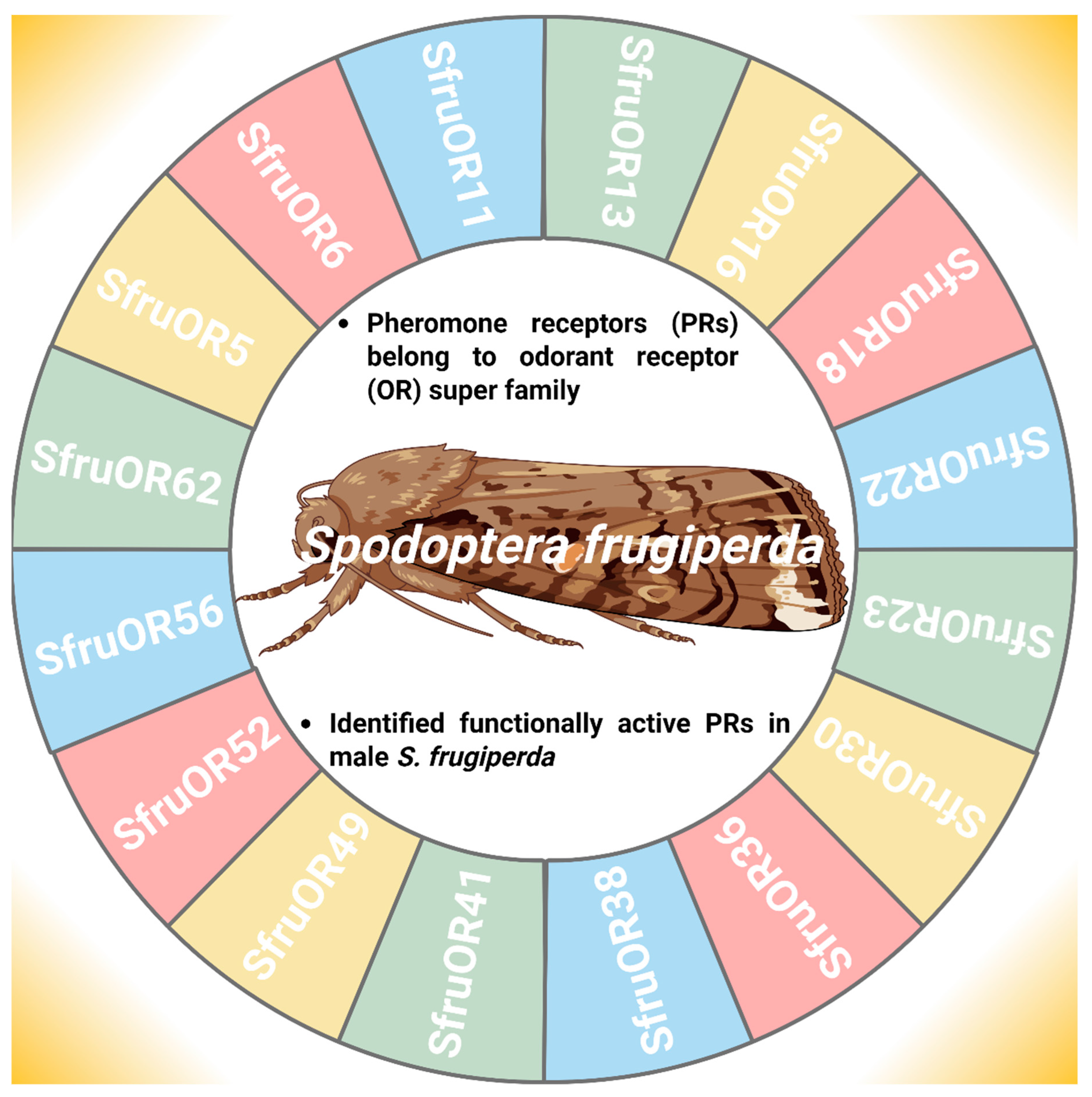

Figure 5.
The role of the olfactory receptor co-receptor (Orco) gene in the olfactory system of Spodoptera frugiperda and its impact on behavioral processes, including host-seeking, mating, and oviposition. The mutation of the Orco gene using CRISPR technology impairs these behaviors, with significant reductions in mating, fecundity, and foraging efficiency. “Figure created using Biorender.com”.
Figure 5.
The role of the olfactory receptor co-receptor (Orco) gene in the olfactory system of Spodoptera frugiperda and its impact on behavioral processes, including host-seeking, mating, and oviposition. The mutation of the Orco gene using CRISPR technology impairs these behaviors, with significant reductions in mating, fecundity, and foraging efficiency. “Figure created using Biorender.com”.
5. Effects of Orco Knockout on Host Plant Volatile Perception in Spodoptera frugiperda
Insect pests use variety of plant volatiles to locate hosts for phytophagy and reproduction [96,97]. The perception of plant volatiles in S. frugiperda depends on ORs, with Orco being essential for the formation of functional ion channels in the membrane of ORNs located in the middle receptors of the male antennae [98,99,100]. The ORNa and ORNb in the male antennae of S. frugiperda females utilized specific maize volatiles, p-xylene and (+)-camphor, to attract the maize plant for oviposition and phytophagy [101]. However, these findings provide a foundation for sustainable pest management by introducing odor-based traps that effectively control S. frugiperda in maize fields. In addition, silencing the Orco gene in insects that perceive plant volatile odors is a promising insect control strategy [100,101,102]. To utilize this tool, Song et al. [103] knocked out the Orco gene in S. frugiperda using the CRISPR/Cas9 gene-editing system. It impaired the ability of Orco knock-out S. frugiperda to perceive plant volatiles, including benzaldehyde, β-pinene, cis-2-hexen-1-ol, jasmone, myrcene, and salicylaldehyde, suggesting that Orco is an excellent target for disrupting S. frugiperda’s normal behavior and providing a feasible pest control approach [103].
The Orco gene constitutes a 1422 bp coding sequence and both 5′ and 3′ untranslated regions, which play a critical role in gene expression. Furthermore, this gene encodes a protein of 473 amino acids, with a molecular weight of 5.3 kDa [95]. The CRISPR/Cas9 system specifically alters the 20 bp target sequence in exon 3 of the Orco gene, causing a 4 bp deletion that ultimately results in a frameshift and a truncated 72-amino acid protein, ultimately disrupting the formation of the functional Orco/OR heteromeric complex and impairing insect olfactory detection of plant volatiles [95].
6. Self-Limiting Spodoptera frugiperda: A Genetic Control Strategy
Until 2024, it was considered that achieving large-scale, effective CRISPR/Cas delivery systems for gene editing remained a critical bottleneck, since embryo microinjection is time-consuming, labor-intensive, and technically demanding, and is the main limitation making it unsuitable for field applications [104]. Furthermore, this approach has been successfully employed only in the laboratory, and large-scale field trials are necessary to validate the laboratory findings [63,105]. However, the introduction of self-limiting transgenic insects carrying lethal genes is the most promising genetic strategy for insect pest management [106]. In this context, a recent study by Reavey et al. [107] investigated the long-term effectiveness and practicality of releasing a genetically engineered, self-limiting strain of S. frugiperda (OX5382G) in a real-world agricultural environment in Brazil as a novel plant protection tool. To develop this strain, the embryos of S. frugiperda “Starkville” wild-type were injected with the pOX5382 plasmid DNA and piggyBac mRNA encoding the transposase. The survivors were screened for the presence of the DsRed2 marker to identify transgenic larvae (OX5382G). The OX5382G strain is self-limiting due to its female-specific mortality, which was conditional on the presence of deoxycycline in the larval diet. Therefore, males are released in the field, and this self-limiting factor reduces the female population [108]. After the release of laboratory-edited male moths into a maize field in Brazil, they showed no significant fitness penalties compared with wild-type males. They were able to disperse and mate effectively with wild females, and their survival rates were similar to those of their wild counterparts in terms of movement and mating ability. Furthermore, the OX5382G males possess a self-limiting gene that, upon mating with wild-type female moths in the field, causes the female offspring to die prematurely, thereby preventing their survival and reproduction [107]. Therefore, genetically modified insect release is emerging as a promising tool to suppress insect pest populations without contaminating the environment with insecticides [109,110,111].
The persistence of laboratory-edited insects in the field is a crucial factor in the success of field release. The lower persistence of the genetically modified strain of S. frugiperda (OX5382G) was observed, with no mutant moths detected in the corn field 25 days post-release, indicating that its self-limiting trait led to population collapse (die-off) shortly after release, preventing long-term persistence [107]. Because in a self-limiting strategy, the genetic modification is programmed to disappear from the population after several generations. However, a self-sustaining strategy is utilized to design a genetic element for enhancing frequency over generations, even when associated with a fitness cost, and seeded at a low proportion in the target population [112]. Regarding biosafety concerns, Reavey et al. [107] also stated that OX5382G S. frugiperda poses no significant risk to the environment or to human or animal health. Because the inserted proteins, such as tetracycline repressor protein (tTAV) and discosoma red fluorescent protein 2 (DsRed2), are non-toxic, non-allergenic, and do not adversely affect predators or parasitoids, the Comissão Técnica Nacional de Biossegurança (CTNBio) approved the commercial deployment of this strain for operational use in the agricultural land in Brazil [107]. Furthermore, to achieve an overflooding ratio (OX5382G males to wild males) in a 200-acre area, release rates of 27–200 males per acre per week were required. When considering multiple release points, this rate was reduced to 14–38 males per acre per week, accounting for incoming OX5382G males from neighboring areas, which is more practical for large-scale corn fields in Brazil [107]. More research is needed to develop this new approach for sustainable crop protection and resistance management across different countries and to assess its potential in the field. However, the large-scale release of mutant insects presents significant cost challenges, including high production costs, logistical complexities, and the need for continuous monitoring [107].
7. Practical Advantages and Limitations of CRISPR/Cas9
The CRISPR/Cas system achieves remarkable precision in targeting DNA by using sgRNAs to cleave DNA at specific locations, resulting in highly accurate genetic modifications [113]. First, it was applied in D. melanogaster and then used in silkworms, mosquitoes, moths, and other insect species [114]. Currently, this tool is widely used and highly effective in manipulating many biological functions of S. frugiperda, including growth and development, reproduction, sex determination, communication, and physiological functions [11,63,115,116,117,118,119]. Moreover, this technology is lower-cost, user-friendly, straightforward, and simplifies the editing process compared to more advanced gene-editing tools [119]. In genome editing, Cas enzymes, including CRISPR-associated protein 9 (Cas9), CRISPR-associated protein 12a (Cas12a), CRISPR-associated protein 12b (Cas12b), CRISPR-associated protein 13 (Cas13), and CRISPR-associated protein 7–11 (Cas7–11), are widely used and offer distinct advantages [120]. Cas12a and Cas12b identify T-rich protospacer adjacent motif (PAM) sites (TTTVs), which increase target-site flexibility and genome-editing effectiveness [121], whereas Cas13 cleaves RNA rather than DNA and does not require a PAM site [122]. Generally, Cas enzymes other than Cas9 generate fewer off-target effects, exhibit lower cytotoxicity, and are smaller, thereby facilitating a more efficient and effective delivery system [120]. For enhancing the efficiency of CRISPR/Cas9 in S. frugiperda, the strong tissue-specific active promoters are required for effective expression of Cas9 and sgRNAs, such as Spodoptera frugiperda heatshock protein 20.15 (SfHsp20.15), Spodoptera frugiperda heatshock protein 20.71 (SfHsp20.71), Spodoptera frugiperda heatshock protein 20D (SfHsp20D), Spodoptera frugiperda heatshock protein 70D (SfHsp70D) from S. frugiperda have been identified and characterized [123]. Regarding the limitations of this system, off-target effects are the most common challenge in CRISPR/Cas9 applications, as the Cas9 enzyme can cleave DNA at unintended locations [124]. For mitigating this issue, the case enzyme variants, including Streptococcus pyogenes Cas9-High fidelity variant 1 (SpCas9-HF1) and xCas9, have high specificity and edit target gene sequences precisely without introducing mutations at off-target sites [125]. Microinjection is another significant limitation on the efficiency of the CRISPR/Cas9 delivery system, due to its time-consuming, labor-intensive, and technically demanding nature, as well as its confinement to the laboratory [104]. To address this issue, a revolutionary technique, nanoparticle-mediated CRISPR/Cas9 delivery, for accurate gene editing of Spodoptera frugiperda 9 (Sf9) cell lines, is crucial for the long-term success of this technology-based insect pest control [126,127].
8. Conclusions and Future Perspectives
Volatile compounds play a crucial role in sexual communication between conspecific S. frugiperda, and their emission is highly age- and mating-status-dependent. In male S. frugiperda, the olfactory system is diversified due to containing olfactory receptor families, including PR, IRs, GRs, GOBPs, and CSPs, which play a significant functional role, including mating, feeding, and providing resistance. Among these receptor families, Orco is the obligatory component of ORs, is essential for OR localization to dendritic membranes, and thus is necessary for odorant detection. Concerning the importance of Orco in the olfactory system, a CRISPR/Cas9-based genetic control strategy was employed to knock out the Orco gene in S. frugiperda, ultimately impairing ecological behaviors. Despite its limitations, this CRISPR/Cas9 system also has advantages and should be explored for large-scale mutagenesis. A novel self-limiting genetic control strategy for S. frugiperda has been practically investigated at a large scale in Brazil by introducing transgenic S. frugiperda with no significant environmental risk. Looking ahead, the behavioral responses of S. frugiperda mutants generated using CRIAPR/Cas9 and self-limitation techniques should be investigated across multiple countries.
Author Contributions
Conceptualization, Y.W., C.Z., B.-M.Y. and L.J.; Investigation, M.-J.L. and A.I. (Asim Iqbal); Visualization, K.S.A., B.-M.Y. and L.J.; Writing—Original Draft Preparation, Y.W., C.Z., M.-J.L., A.I. (Asim Iqbal), B.-M.Y. and L.J.; Writing—Review and Editing, K.S.A., H. and A.I. (Atif Idrees); Funding Acquisition, Y.W., C.Z., B.-M.Y. and L.J. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Innovation Capacity Building Project of the Jilin Provincial Development and Reform Commission (Grant No. 2024C014-2).
Data Availability Statement
No new data were created or analyzed in this study. Data sharing is not applicable to this article.
Acknowledgments
The authors of this manuscript declare that the ideas, content, and theories presented were not generated through AI-assisted technologies. However, AI tools, including ChatGPT, were utilized solely to enhance readability and refine the language. The use of AI was strictly supervised, and the authors thoroughly reviewed and edited the manuscript to ensure its accuracy, coherence, and adherence to the intended message.
Conflicts of Interest
Authors Asim Iqbal and Kanwer Shahzad Ahmed are employed by companies (Imdaad: Integrated Facilities Management Company and Mastermind Scientific Consultants (SMC-Private) Limited, respectively). The remaining authors declare that the research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Abbreviations
| AAT | Alcohol acetyltransferase |
| ACC | Acetyl-CoA carboxylase |
| BE | Base editors |
| C16:0-CoA | Saturated 16-carbon fatty acyl-CoA |
| Cas12a | CRISPR-associated protein 12a |
| Cas12b | CRISPR-associated protein 12b |
| Cas13 | CRISPR-associated protein 13 |
| Cas7–11 | CRISPR-associated protein 7–11 |
| Cas9 | CRISPR-associated protein 9 |
| CRISPR-Cas9 | Clustered Regularly Interspaced Short Palindromic Repeats |
| CTNBio | Comissão Técnica Nacional de Biossegurança |
| CTNBio | Comissão Técnica Nacional de Biossegurança |
| DNA | Deoxyribonucleic acid |
| DsRed2 | Discosoma red fluorescent protein 2 |
| E7–12:OAc | (E)-7-dodecenyl acetate |
| FARs | Fatty acyl CoA reductases |
| FAS | Fatty acid synthase |
| HDR | Homology-directed repair |
| MAG | Male’s accessory gland |
| MNs | Meganucleases |
| mRNA | Messenger RNA |
| NHEJ | Non-homologous end joining |
| Orco | Odorant receptor co-receptor |
| ORNs | Olfactory receptor neurons |
| ORs | Odorant receptors |
| PAM | Protospacer adjacent motif |
| PBAN | Pheromone biosynthesis-activating neuropeptide |
| PRs | Pheromone receptors |
| qPCR | Quantitative polymerase chain reaction |
| RNA | Ribonucleic acid |
| SDNs | Site-directed nucleases |
| Sfabd-A | Spodoptera frugiperda abdominal-A homeotic |
| SfHsp20.15 | Spodoptera frugiperda heatshock protein 20.15 |
| SfHsp20.71 | Spodoptera frugiperda heatshock protein 20.71 |
| SfHsp20D | Spodoptera frugiperda heatshock protein 20D |
| SfHsp70D | Spodoptera frugiperda heatshock protein 70D |
| SfruOR11 | Spodoptera frugiperda odorant receptor 11 |
| SfruOR13 | Spodoptera frugiperda odorant receptor 13 |
| SfruOR16 | Spodoptera frugiperda odorant receptor 16 |
| SfruOR18 | Spodoptera frugiperda odorant receptor 18 |
| SfruOR22 | Spodoptera frugiperda odorant receptor 22 |
| SfruOR23 | Spodoptera frugiperda odorant receptor 23 |
| SfruOR30 | Spodoptera frugiperda odorant receptor 30 |
| SfruOR36 | Spodoptera frugiperda odorant receptor 36 |
| SfruOR38 | Spodoptera frugiperda odorant receptor 38 |
| SfruOR41 | Spodoptera frugiperda odorant receptor 41 |
| SfruOR49 | Spodoptera frugiperda odorant receptor 49 |
| SfruOR5 | Spodoptera frugiperda odorant receptor 5 |
| SfruOR52 | Spodoptera frugiperda odorant receptor 52 |
| SfruOR56 | Spodoptera frugiperda odorant receptor 56 |
| SfruOR6 | Spodoptera frugiperda odorant receptor 6 |
| SfruOR62 | Spodoptera frugiperda odorant receptor 62 |
| sgRNA | Single-guide RNA |
| SpCas9-HF1 | Streptococcus pyogenes Cas9-High-fidelity variant 1 |
| TALENs | Transcription activator-like effector nucleases |
| tTAV | Tetracycline repressor protein |
| Z11–16-CoA | (Z)-11-hexadecenoyl-CoA |
| Z7–12:CoA | (Z)-7-dodecenyl acetate |
| Z7–12:OAc | (Z)-7-dodecenyl acetate |
| Z9–12:OAc | (Z)-9-dodecenyl acetate |
| Z9–14:OAc | (Z)-9-tetradecenyl acetate |
| Z9–14:OAc | (Z)-9-tetradecenyl acetate |
| ZFNs | Zinc finger nucleases |
References
- Maeder, M.L.; Gersbach, C.A. Genome-editing technologies for gene and cell therapy. Mol. Ther. 2016, 24, 430–446. [Google Scholar] [CrossRef] [PubMed]
- Khalil, A.M. The genome editing revolution. J. Genet. Eng. Biotechnol. 2020, 18, 68. [Google Scholar] [CrossRef]
- Li, J.; Wu, S.; Zhang, K.; Sun, X.; Lin, W.; Wang, C.; Lin, S. Clustered regularly interspaced short palindromic repeat/crispr-associated protein and its utility all at sea: Status, challenges, and prospects. Microorganisms 2024, 12, 118. [Google Scholar] [CrossRef] [PubMed]
- Ali, A.; Zafar, M.M.; Farooq, Z.; Ahmed, S.R.; Ijaz, A.; Anwar, Z.; Abbas, H.; Tariq, M.S.; Tariq, H.; Mustafa, M.; et al. Breakthrough in CRISPR/Cas system: Current and future directions and challenges. Biotechnol. J. 2023, 18, 2200642. [Google Scholar] [CrossRef] [PubMed]
- Gupta, D.; Bhattacharjee, O.; Mandal, D.; Sen, M.K.; Dey, D.; Dasgupta, A.; Kazi, T.A.; Gupta, R.; Sinharoy, S.; Acharya, K.; et al. CRISPR-Cas9 system: A new-fangled dawn in gene editing. Life Sci. 2019, 232, 116636. [Google Scholar] [CrossRef]
- Asmamaw, M.; Zawdie, B. Mechanism and applications of CRISPR/Cas-9-mediated genome editing. Biol. Targets Ther. 2021, 15, 353–361. [Google Scholar]
- Komal, J.; Desai, H.R.; Samal, I.; Mastinu, A.; Patel, R.D.; Kumar, P.D.; Majhi, P.K.; Mahanta, D.K.; Bhoi, T.K. Unveiling the genetic symphony: Harnessing CRISPR-Cas genome editing for effective insect pest management. Plants 2023, 12, 3961. [Google Scholar] [CrossRef]
- Singh, S.; Rahangdale, S.; Pandita, S.; Saxena, G.; Upadhyay, S.K.; Mishra, G.; Verma, P.C. CRISPR/Cas9 for insect pests management: A comprehensive review of advances and applications. Agriculture 2022, 12, 1896. [Google Scholar] [CrossRef]
- Sun, D.; Guo, Z.; Liu, Y.; Zhang, Y. Progress and prospects of CRISPR/Cas systems in insects and other arthropods. Front. Physiol. 2017, 8, 608. [Google Scholar] [CrossRef]
- Li, J.J.; Shi, Y.; Wu, J.N.; Li, H.; Smagghe, G.; Liu, T.X. CRISPR/Cas9 in lepidopteran insects: Progress, application and prospects. J. Insect Physiol. 2021, 135, 104325. [Google Scholar] [CrossRef]
- Ashok, K.; Bhargava, C.N.; Asokan, R.; Pradeep, C.; Kennedy, J.S.; Manamohan, M.; Rai, A. CRISPR/Cas9 mediated mutagenesis of the major sex pheromone gene, acyl-CoA delta-9 desaturase (DES9) in fall armyworm Spodoptera frugiperda (JE Smith) (Lepidoptera: Noctuidae). Int. J. Biol. Macromol. 2023, 253, 126557. [Google Scholar] [CrossRef]
- Ashok, K.; Bhargava, C.N.; Asokan, R.; Pradeep, C.; Pradhan, S.K.; Kennedy, J.S.; Balasubramani, V.; Murugan, M.; Jayakanthan, M.; Geethalakshmi, V.; et al. CRISPR/Cas9 mediated editing of pheromone biosynthesis activating neuropeptide (PBAN) gene disrupts mating in the Fall armyworm, Spodoptera frugiperda (JE Smith) (Lepidoptera: Noctuidae). 3 Biotech 2023, 13, 370. [Google Scholar] [CrossRef]
- Durand, K.; An, H.; Nam, K. Invasive fall armyworms are corn strain. Sci. Rep. 2024, 14, 5696. [Google Scholar] [CrossRef]
- Nurkomar, I.; Putra, I.L.; Buchori, D.; Setiawan, F. Association of a global invasive pest Spodoptera frugiperda (Lepidoptera: Noctuidae) with local parasitoids: Prospects for a new approach in selecting biological control agents. Insects 2024, 15, 205. [Google Scholar] [CrossRef]
- Montezano, D.G.; Specht, A.; Soja, E.; Sosa-Gomez, D.R.; Roque-Specht, V.F.; Malaquias, J.V.; Paula-Moraes, S.V.; Peterson, J.A.; Hunt, T.E. Biotic potential and reproductive parameters of Spodoptera frugiperda (JE Smith, 1797) (Lepidoptera: Noctuidae). J. Agric. Sci. 2019, 11, 240. [Google Scholar]
- Abro, Z.; Kimathi, E.; De Groote, H.; Tefera, T.; Sevgan, S.; Niassy, S.; Kassie, M. Socioeconomic and health impacts of fall armyworm in Ethiopia. PLoS ONE 2021, 16, e0257736. [Google Scholar] [CrossRef]
- Zhang, D.D. Insecticide resistance monitoring for the invasive populations of fall armyworm, Spodoptera frugiperda in China. J. Integr. Agric. 2021, 20, 783–791. [Google Scholar] [CrossRef]
- Sánchez-Bayo, F. Insecticides mode of action in relation to their toxicity to non-target organisms. J. Environ. Anal. Toxicol. 2012, S4, 002. [Google Scholar]
- Reisenman, C.E.; Lei, H.; Guerenstein, P.G. Neuroethology of olfactory-guided behavior and its potential application in the control of harmful insects. Front. Physiol. 2016, 7, 271. [Google Scholar] [CrossRef] [PubMed]
- Shi, L.; Liu, X.; Liu, H.; Shan, S.; Shen, S.; Bai, M.; Lan, H.; Khashaveh, A.; Gu, S.; Zhang, Y. Knockout of the delta11-desaturase Sfrudes1 disrupts sex pheromone biosynthesis, mating and oviposition in the fall armyworm, Spodoptera frugiperda. Pestic. Biochem. Physiol. 2024, 200, 105832. [Google Scholar] [CrossRef] [PubMed]
- Domínguez, A.; López, S.; Bernabé, A.; Guerrero, Á.; Quero, C. Influence of age, host plant and mating status in pheromone production and new insights on perception plasticity in Tuta absoluta. Insects 2019, 10, 256. [Google Scholar] [CrossRef] [PubMed]
- Fleischer, J.; Krieger, J. Insect pheromone receptors–key elements in sensing intraspecific chemical signals. Front. Cell. Neurosci. 2018, 12, 425. [Google Scholar] [CrossRef]
- Bratovich, C.; Saluso, A.; Murua, M.G.; Guerenstein, P.G. Evaluation of sex pheromone formulations to attract Spodoptera frugiperda (Lepidoptera: Noctuidae) adult males in Argentina. Rev. De La Soc. Entomológica Argent. 2019, 78, 7–14. [Google Scholar] [CrossRef]
- Qu, C.; Kang, Z.; Zhang, B.; Fang, Y.; Wang, R.; Li, F.; Zhao, H.; Luo, C. Genome-wide identification and expression profiling of candidate sex pheromone biosynthesis genes in the fall armyworm (Spodoptera frugiperda). Insects 2022, 13, 1078. [Google Scholar] [CrossRef]
- Muthukumar, M.; Ashok, K.; Kennedy, J.S. Study of sexual behaviour and characterization of female sex pheromone of fall armyworm Spodoptera frugiperda (JE Smith) (Lepidoptera: Noctuidae) collected from different locations of Tamil Nadu, India. J. Asia-Pac. Entomol. 2023, 26, 102150. [Google Scholar] [CrossRef]
- Phuong, T.T.; Fujii, T.; Ishikawa, Y.; Tung, N.D.; Giang, H.T. Female sex pheromone of Spodoptera frugiperda Vietnam population and a selected lure for adult monitoring in maize fields. J. Asia-Pac. Entomol. 2024, 27, 102343. [Google Scholar] [CrossRef]
- Renou, M. Pheromones and general odor perception in insects. Neurobiol. Chem. Commun. 2014, 1, 23–56. [Google Scholar]
- Stengl, M.; Funk, N.W. The role of the coreceptor Orco in insect olfactory transduction. J. Comp. Physiol. A 2013, 199, 897–909. [Google Scholar] [CrossRef]
- Yan, H.; Jafari, S.; Pask, G.; Zhou, X.; Reinberg, D.; Desplan, C. Evolution, developmental expression and function of odorant receptors in insects. J. Exp. Biol. 2020, 223, jeb208215. [Google Scholar] [CrossRef]
- Wicher, D.; Miazzi, F. Functional properties of insect olfactory receptors: Ionotropic receptors and odorant receptors. Cell Tissue Res. 2021, 383, 7–19. [Google Scholar] [CrossRef] [PubMed]
- Wicher, D. Olfactory signaling in insects. Prog. Mol. Biol. Transl. Sci. 2015, 130, 37–54. [Google Scholar]
- Zufall, F.; Domingos, A.I. The structure of Orco and its impact on our understanding of olfaction. J. Gen. Physiol. 2018, 150, 1602–1605. [Google Scholar] [CrossRef]
- Paulo, D.F.; Junqueira, A.C.; Arp, A.P.; Vieira, A.S.; Ceballos, J.; Skoda, S.R.; Pérez-de-León, A.A.; Sagel, A.; McMillan, W.O.; Scott, M.J.; et al. Disruption of the odorant coreceptor Orco impairs foraging and host finding behaviors in the New World screwworm fly. Sci. Rep. 2021, 11, 11379. [Google Scholar] [CrossRef] [PubMed]
- Fan, X.B.; Mo, B.T.; Li, G.C.; Huang, L.Q.; Guo, H.; Gong, X.L.; Wang, C.Z. Mutagenesis of the odorant receptor co-receptor (Orco) reveals severe olfactory defects in the crop pest moth Helicoverpa armigera. BMC Biol. 2022, 20, 214. [Google Scholar] [CrossRef]
- Fandino, R.A.; Haverkamp, A.; Bisch-Knaden, S.; Zhang, J.; Bucks, S.; Nguyen, T.A.; Schröder, K.; Werckenthin, A.; Rybak, J.; Stengl, M.; et al. Mutagenesis of odorant coreceptor Orco fully disrupts foraging but not oviposition behaviors in the hawkmoth Manduca sexta. Proc. Natl. Acad. Sci. USA 2019, 116, 15677–15685. [Google Scholar] [CrossRef] [PubMed]
- Saveer, A.M.; Hatano, E.; Wada-Katsumata, A.; Meagher, R.L.; Schal, C. Nonanal, a new fall armyworm sex pheromone component, significantly increases the efficacy of pheromone lures. Pest Manag. Sci. 2023, 79, 2831–2839. [Google Scholar] [CrossRef] [PubMed]
- Alabaster, A.; Isoe, J.; Zhou, G.; Lee, A.; Murphy, A.; Day, W.A.; Miesfeld, R.L. Deficiencies in acetyl-CoA carboxylase and fatty acid synthase 1 differentially affect eggshell formation and blood meal digestion in Aedes aegypti. Insect Biochem. Mol. Biol. 2011, 41, 946–955. [Google Scholar] [CrossRef]
- Choi, M.Y.; Jurenka, R.A. C75, a fatty acid synthase inhibitor, inhibits feeding activity and pheromone production in a moth, Helicoverpa zea. J. Asia-Pac. Entomol. 2006, 9, 43–48. [Google Scholar] [CrossRef]
- Groot, A.T.; Marr, M.; Schöfl, G.; Lorenz, S.; Svatos, A.; Heckel, D.G. Host strain specific sex pheromone variation in Spodoptera frugiperda. Front. Zool. 2008, 5, 20. [Google Scholar] [CrossRef]
- Ma, P.W.; Ramaswamy, S.B. Biology and ultrastructure of sex pheromone-producing tissue. In Insect Pheromone Biochemistry and Molecular Biology; Academic Press: Cambridge, MA, USA, 2003; pp. 19–51. [Google Scholar]
- Akter, S.; Hossain, M.S.; Ali, R.; Regmi, R.; Park, S.J.; Mainali, B. Variations in Sex Pheromone of the Australian Population of Fall Armyworm: Influence of Age and Mating Status. J. Chem. Ecol. 2025, 51, 55. [Google Scholar] [CrossRef]
- Smith, A.F.; Schal, C. Corpus allatum control of sex pheromone production and calling in the female brown-banded cockroach, Supella longipalpa (F.) (Dictyoptera: Blattellidae). J. Insect Physiol. 1990, 36, 251–257. [Google Scholar] [CrossRef]
- Toivonen, J.M.; Partridge, L. Endocrine regulation of aging and reproduction in Drosophila. Mol. Cell. Endocrinol. 2009, 299, 39–50. [Google Scholar] [CrossRef]
- Force, E.; Debernard, S. Endocrine regulation of reproductive behaviors in insects: A comprehensive review. Curr. Opin. Insect Sci. 2025, 69, 101360. [Google Scholar] [CrossRef]
- Liu, Y.; Zou, K.; Wang, T.; Guan, M.; Duan, H.; Yu, H.; Wu, D.; Du, J. Genome-Wide Identification and Analysis of Family Members with Juvenile Hormone Binding Protein Domains in Spodoptera frugiperda. Insects 2024, 15, 573. [Google Scholar] [CrossRef] [PubMed]
- Hassanien, I.T.; Grötzner, M.; Meyering-Vos, M.; Hoffmann, K.H. Neuropeptides affecting the transfer of juvenile hormones from males to females during mating in Spodoptera frugiperda. J. Insect Physiol. 2014, 66, 45–52. [Google Scholar] [CrossRef]
- Li, C.; Yu, J.F.; Lu, Q.; Xu, J.; Liu, J.H.; Ye, H. Molecular characterization and functional analysis of a putative sex-peptide receptor in the tobacco cutworm Spodoptera litura (F abricius, 1775) (Lepidoptera: N octuidae). Austral Entomol. 2014, 53, 424–431. [Google Scholar] [CrossRef]
- Comte, A.; Delarue, A.; François, M.C.; Monsempes, C.; Meslin, C.; Montagné, N.; Jacquin-Joly, E. Functional Investigation of the Receptor to the Major Pheromone Component in the C-Strain and the R-Strain of the Fall Armyworm Spodoptera frugiperda. Insects 2025, 16, 304. [Google Scholar] [CrossRef]
- Wu, H.; Xu, M.; Hou, C.; Huang, L.Q.; Dong, J.F.; Wang, C.Z. Specific olfactory neurons and glomeruli are associated to differences in behavioral responses to pheromone components between two Helicoverpa species. Front. Behav. Neurosci. 2015, 9, 206. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Gong, X.L.; Li, G.C.; Mo, B.T.; Jiang, N.J.; Huang, L.Q.; Wang, C.Z. Functional analysis of pheromone receptor repertoire in the fall armyworm, Spodoptera frugiperda. Pest Manag. Sci. 2022, 78, 2052–2064. [Google Scholar] [CrossRef]
- Qiu, L.; He, L.; Tan, X.; Zhang, Z.; Wang, Y.; Li, X.; He, H.; Ding, W.; Li, Y. Identification and phylogenetics of Spodoptera frugiperda chemosensory proteins based on antennal transcriptome data. Comp. Biochem. Physiol. Part D Genom. Proteom. 2020, 34, 100680. [Google Scholar] [CrossRef]
- Liu, X.; Shi, L.; Khashaveh, A.; Shan, S.; Lv, B.; Gu, S.; Zhang, Y. Loss of binding capabilities in an ecologically important odorant receptor of the fall armyworm, Spodoptera frugiperda, by a single point mutation. J. Agric. Food Chem. 2023, 71, 13003–13013. [Google Scholar] [CrossRef]
- Wang, Y.; Qiu, L.; Wang, B.; Guan, Z.; Dong, Z.; Zhang, J.; Cao, S.; Yang, L.; Wang, B.; Gong, Z.; et al. Structural basis for odorant recognition of the insect odorant receptor OR-Orco heterocomplex. Science 2024, 384, 1453–1460. [Google Scholar] [CrossRef]
- Ray, I.; Mohanty, S. Pheromone-binding proteins in pest control: From molecular insights to real-world applications. J. Agric. Food Chem. 2025, 73, 21701–21727. [Google Scholar] [CrossRef]
- Zhang, D.D.; Löfstedt, C. Moth pheromone receptors: Gene sequences, function, and evolution. Front. Ecol. Evol. 2015, 3, 105. [Google Scholar] [CrossRef]
- Guo, J.M.; Liu, X.L.; Liu, S.R.; Wei, Z.Q.; Han, W.K.; Guo, Y.; Dong, S.L. Functional characterization of sex pheromone receptors in the fall armyworm (Spodoptera frugiperda). Insects 2020, 11, 193. [Google Scholar] [CrossRef] [PubMed]
- Malo, E.A.; Castrejón-Gómez, V.R.; Cruz-López, L.; Rojas, J.C. Antennal sensilla and electrophysiological response of male and female Spodoptera frugiperda (Lepidoptera: Noctuidae) to conspecific sex pheromone and plant odors. Ann. Entomol. Soc. Am. 2004, 97, 1273–1284. [Google Scholar] [CrossRef]
- Malzahn, A.; Lowder, L.; Qi, Y. Plant genome editing with TALEN and CRISPR. Cell Biosci. 2017, 7, 21. [Google Scholar] [CrossRef] [PubMed]
- Cardi, T.; Murovec, J.; Bakhsh, A.; Boniecka, J.; Bruegmann, T.; Bull, S.E.; Eeckhaut, T.; Fladung, M.; Galovic, V.; Linkiewicz, A.; et al. CRISPR/Cas-mediated plant genome editing: Outstanding challenges a decade after implementation. Trends Plant Sci. 2023, 28, 1144–1165. [Google Scholar] [CrossRef]
- Redman, M.; King, A.; Watson, C.; King, D. What is CRISPR/Cas9? Arch. Dis. Child. Educ. Pract. 2016, 101, 213–215. [Google Scholar] [CrossRef] [PubMed]
- Drury, D.W.; Dapper, A.L.; Siniard, D.J.; Zentner, G.E.; Wade, M.J. CRISPR/Cas9 gene drives in genetically variable and nonrandomly mating wild populations. Sci. Adv. 2017, 3, e1601910. [Google Scholar] [CrossRef]
- Kim, H.; Ishidate, T.; Ghanta, K.S.; Seth, M.; Conte, D., Jr.; Shirayama, M.; Mello, C.C. A co-CRISPR strategy for efficient genome editing in Caenorhabditis elegans. Genetics 2014, 197, 1069–1080. [Google Scholar] [CrossRef] [PubMed]
- Salum, Y.M.; Yin, A.; Zaheer, U.; Liu, Y.; Guo, Y.; He, W. CRISPR/Cas9-based genome editing of fall armyworm (Spodoptera frugiperda): Progress and prospects. Biomolecules 2024, 14, 1074. [Google Scholar] [CrossRef]
- Wu, K.; Shirk, P.D.; Taylor, C.E.; Furlong, R.B.; Shirk, B.D.; Pinheiro, D.H.; Siegfried, B.D. CRISPR/Cas9 mediated knockout of the abdominal-A homeotic gene in fall armyworm moth (Spodoptera frugiperda). PLoS ONE 2018, 13, e0208647. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Li, Z. CRISPR-Cas systems: Overview, innovations and applications in human disease research and gene therapy. Comput. Struct. Biotechnol. J. 2020, 18, 2401–2415. [Google Scholar] [CrossRef] [PubMed]
- Kalajdzic, P.; Schetelig, M.F. CRISPR/Cas-mediated gene editing using purified protein in Drosophila suzukii. Entomol. Exp. Appl. 2017, 164, 350–362. [Google Scholar] [CrossRef]
- Lamb, A.M.; Walker, E.A.; Wittkopp, P.J. Tools and strategies for scarless allele replacement in Drosophila using CRISPR/Cas9. Fly 2017, 11, 53–64. [Google Scholar] [CrossRef]
- El-Awaad, E.; Merzendorfer, H. CRISPR/Cas: An emerging molecular technology for biological control of fall armyworm. New Plant Prot. 2025, 2, e26. [Google Scholar] [CrossRef]
- Bi, H.L.; Xu, J.; Tan, A.J.; Huang, Y.P. CRISPR/Cas9-mediated targeted gene mutagenesis in Spodoptera litura. Insect Sci. 2016, 23, 469–477. [Google Scholar] [CrossRef]
- Zuo, Y.; Wang, H.; Xu, Y.; Huang, J.; Wu, S.; Wu, Y.; Yang, Y. CRISPR/Cas9 mediated G4946E substitution in the ryanodine receptor of Spodoptera exigua confers high levels of resistance to diamide insecticides. Insect Biochem. Mol. Biol. 2017, 89, 79–85. [Google Scholar] [CrossRef]
- Zhang, S.; Jacquin-Joly, E.; Montagné, N.; Liu, F.; Liu, Y.; Wang, G. Identification of an odorant receptor responding to sex pheromones in Spodoptera frugiperda extends the novel type-I PR lineage in moths. Insect Sci. 2024, 31, 489–502. [Google Scholar] [CrossRef]
- Zhang, S.; Yan, S.; Mei, X.; Wang, G.; Liu, Y. Identification of a new lineage of pheromone receptors in mirid bugs (Heteroptera: Miridae). Pestic. Biochem. Physiol. 2025, 208, 106277. [Google Scholar] [CrossRef] [PubMed]
- Holdcraft, R.; Rodriguez-Saona, C.; Stelinski, L.L. Pheromone autodetection: Evidence and implications. Insects 2016, 7, 17. [Google Scholar] [CrossRef]
- Costa, C.G.; Rodrigues, S.R.; Fuhrmann, J. Morphology of the antennal sensilla of two species of Hoplopyga Thomson, 1880 (Coleoptera, Scarabaeidae, Cetoniinae). Rev. Bras. De Entomol. 2021, 65, e20200078. [Google Scholar] [CrossRef]
- Wang, W.; He, P.; Liu, T.; Jing, X.; Zhang, S. Morphology and distribution of antennal sensilla on Spodoptera frugiperda (Lepidoptera: Noctuidae) larvae and adults. Diversity 2023, 15, 992. [Google Scholar] [CrossRef]
- Benton, R.; Vannice, K.S.; Gomez-Diaz, C.; Vosshall, L.B. Variant ionotropic glutamate receptors as chemosensory receptors in Drosophila. Cell 2009, 136, 149–162. [Google Scholar] [CrossRef]
- Sun, Y.L.; Dong, J.F.; Yang, H.B.; Li, D.X.; Tian, C.H. Identification and characterization of chemosensory receptors in the pheromone gland-ovipositor of Spodoptera frugiperda (JE Smith). Insects 2022, 13, 481. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.L.; Yan, Q.; Yang, Y.L.; Hou, W.; Miao, C.L.; Peng, Y.C.; Dong, S.L. A gustatory receptor GR8 tunes specifically to D-fructose in the common cutworm Spodoptera litura. Insects 2019, 10, 272. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.L.; Jiang, P.S.; Dong, B.X.; Tian, C.H.; Dong, J.F. Candidate chemosensory receptors in the antennae and maxillae of Spodoptera frugiperda (JE Smith) larvae. Front. Physiol. 2022, 13, 970915. [Google Scholar] [CrossRef]
- Zhang, R.; Wang, B.; Grossi, G.; Falabella, P.; Liu, Y.; Yan, S.; Lu, J.; Xi, J.; Wang, G. Molecular basis of alarm pheromone detection in aphids. Curr. Biol. 2017, 27, 55–61. [Google Scholar] [CrossRef]
- Pelosi, P.; Zhou, J.J.; Ban, L.P.; Calvello, M. Soluble proteins in insect chemical communication. Cell. Mol. Life Sci. CMLS 2006, 63, 1658–1676. [Google Scholar] [CrossRef]
- Liu, X.L.; Wu, Z.R.; Liao, W.; Zhang, X.Q.; Pei, Y.W.; Lu, M. The binding affinity of two general odorant binding proteins in Spodoptera frugiperda to general volatiles and insecticides. Int. J. Biol. Macromol. 2023, 252, 126338. [Google Scholar] [CrossRef] [PubMed]
- Han, W.K.; Tang, F.X.; Yu, N.; Zhang, Y.X.; Liu, Z.W. A nonsensory odorant-binding protein plays an important role in the larval development and adult mating of Spodoptera frugiperda. Insect Sci. 2023, 30, 1325–1336. [Google Scholar] [CrossRef] [PubMed]
- Lagarde, A.; Spinelli, S.; Tegoni, M.; He, X.; Field, L.; Zhou, J.J.; Cambillau, C. The crystal structure of odorant binding protein 7 from Anopheles gambiae exhibits an outstanding adaptability of its binding site. J. Mol. Biol. 2011, 414, 401–412. [Google Scholar] [CrossRef]
- Leal, W.S.; Nikonova, L.; Peng, G. Disulfide structure of the pheromone binding protein from the silkworm moth, Bombyx mori. FEBS Lett. 1999, 464, 85–90. [Google Scholar] [CrossRef]
- Tsouri, A.; Douris, V. The Role of Chemosensory Proteins in Insecticide Resistance: A Review. Insects 2025, 16, 496. [Google Scholar] [CrossRef]
- Jia, C.; Mohamed, A.; Cattaneo, A.M.; Huang, X.; Keyhani, N.O.; Gu, M.; Zang, L.; Zhang, W. Odorant-binding proteins and chemosensory proteins in Spodoptera frugiperda: From genome-wide identification and developmental stage-related expression analysis to the perception of host plant odors, sex pheromones, and insecticides. Int. J. Mol. Sci. 2023, 24, 5595. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Zhao, R.; Gao, J.; Xiao, X.; Yin, X.; Hu, S.; Zhang, Y.; Liang, P.; Gu, S. Two cuticle-enriched chemosensory proteins confer multi-insecticide resistance in Spodoptera frugiperda. Int. J. Biol. Macromol. 2024, 266, 130941. [Google Scholar] [CrossRef]
- Chen, Q.; Liu, X.; Cao, S.; Ma, B.; Guo, M.; Shen, J.; Wang, G. Fine structure and olfactory reception of the labial palps of Spodoptera frugiperda. Front. Physiol. 2021, 12, 680697. [Google Scholar] [CrossRef]
- Peter, E.; Tamiru, A.; Sevgan, S.; Dubois, T.; Kelemu, S.; Kruger, K.; Torto, B.; Yusuf, A. Companion crops alter olfactory responses of the fall armyworm (Spodoptera frugiperda) and its larval endoparasitoid (Cotesia icipe). Chem. Biol. Technol. Agric. 2023, 10, 61. [Google Scholar] [CrossRef]
- Chen, X.; Yao, S.; Xie, L.; Li, J.; Xiong, L.; Yang, X.; Chen, Y.; Cao, F.; Hou, Q.; You, M.; et al. Disruption of the odorant receptor co-receptor (Orco) reveals its critical role in multiple olfactory behaviors of a cosmopolitan pest. Insect Biochem. Mol. Biol. 2025, 177, 104248. [Google Scholar] [CrossRef]
- Boronat-Garcia, A.; Iben, J.; Dominguez-Martin, E.; Stopfer, M. Identification and analysis of odorant receptors expressed in the two main olfactory organs, antennae and palps, of Schistocerca americana. Sci. Rep. 2022, 12, 22628. [Google Scholar] [CrossRef] [PubMed]
- Mukunda, L.; Lavista-Llanos, S.; Hansson, B.S.; Wicher, D. Dimerisation of the Drosophila odorant coreceptor Orco. Front. Cell. Neurosci. 2014, 8, 261. [Google Scholar] [CrossRef]
- Tiwari, V.; Sowdhamini, R. Structure modelling of odorant receptor from Aedes aegypti and identification of potential repellent molecules. Comput. Struct. Biotechnol. J. 2023, 21, 2204–2214. [Google Scholar] [CrossRef]
- Sun, H.; Bu, L.A.; Su, S.C.; Guo, D.; Gao, C.F.; Wu, S.F. Knockout of the odorant receptor co-receptor, orco, impairs feeding, mating and egg-laying behavior in the fall armyworm Spodoptera frugiperda. Insect Biochem. Mol. Biol. 2023, 152, 103889. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, I. Attracting carnivorous arthropods with plant volatiles: The future of biocontrol or playing with fire? Biol. Control 2012, 60, 77–89. [Google Scholar] [CrossRef]
- Darathi, D.; Santa, K. Volatile mediated plant-insect interactions: A review. Int. J. Zool. Investig. 2022, 8, 291–304. [Google Scholar] [CrossRef]
- Jones, W.D.; Nguyen, T.A.; Kloss, B.; Lee, K.J.; Vosshall, L.B. Functional conservation of an insect odorant receptor gene across 250 million years of evolution. Curr. Biol. 2005, 15, R119–R121. [Google Scholar] [CrossRef]
- Wicher, D.; Schäfer, R.; Bauernfeind, R.; Stensmyr, M.C.; Heller, R.; Heinemann, S.H.; Hansson, B.S. Drosophila odorant receptors are both ligand-gated and cyclic-nucleotide-activated cation channels. Nature 2008, 452, 1007–1011. [Google Scholar] [CrossRef]
- Wang, Q.; Jia, Y.; Smid, H.M.; Weldegergis, B.T.; Greenberg, L.O.; Jongsma, M.; Dicke, M.; Haverkamp, A. Loss of olfaction reduces caterpillar performance and increases susceptibility to a natural enemy. elife 2025, 14, RP105585. [Google Scholar] [CrossRef]
- Li, H.; Wan, P.; Zhu, Z.; Xu, D.; Cong, S.; Xu, M.; Yin, H. Spodoptera frugiperda Uses Specific Volatiles to Assess Maize Development for Optimal Offspring Survival. Insects 2025, 16, 592. [Google Scholar] [CrossRef]
- Zhang, C.; Tang, B.; Tan, H.; Wang, X.; Dai, W. The Orco gene involved in recognition of host plant volatiles and sex pheromone in the chive maggot Bradysia odoriphaga. Pestic. Biochem. Physiol. 2023, 197, 105709. [Google Scholar] [CrossRef] [PubMed]
- Song, C.A.; Yang, L.I.; Qing, Y.A.; Gui-rong, W.A. Mutagenesis of odorant coreceptor Orco reveals the distinct role of olfaction between sexes in Spodoptera frugiperda. J. Integr. Agric. 2023, 22, 2162–2172. [Google Scholar]
- Zhao, Y.; Li, L.; Wei, L.; Wang, Y.; Han, Z. Advancements and Future Prospects of CRISPR-Cas-Based Population Replacement Strategies in Insect Pest Management. Insects 2024, 15, 653. [Google Scholar] [CrossRef]
- Yadav, S.; Sarangi, S.; Parmar, S.; Thodusu, M.; Soni, S.; Pandey, K. Genetic engineering in insect management: New frontiers in pest control. Microbiol. Res. J. Int. 2024, 34, 106–121. [Google Scholar] [CrossRef]
- Zhou, L.; Alphey, N.; Walker, A.S.; Travers, L.M.; Morrison, N.I.; Bonsall, M.B.; Raymond, B. The application of self-limiting transgenic insects in managing resistance in experimental metapopulations. J. Appl. Ecol. 2019, 56, 688–698. [Google Scholar] [CrossRef]
- Reavey, C.E.; Domingues, F.A.; Ercit, K.; Pinto, R.L.; Naish, N.; Yadav, M.; Frazon, T.; Cabala, K.; Sulston, E.; Pickl-Herk, A.; et al. Field Performance of a Self-Limiting, Genetically Engineered Fall Armyworm for Biological Pest Management. Front. Agron. 2025, 7, 1613689. [Google Scholar] [CrossRef]
- Reavey, C.E.; Walker, A.S.; Joyce, S.P.; Broom, L.; Willse, A.; Ercit, K.; Poletto, M.; Barnes, Z.H.; Marubbi, T.; Troczka, B.J.; et al. Self-limiting fall armyworm: A new approach in development for sustainable crop protection and resistance management. BMC Biotechnol. 2022, 22, 5. [Google Scholar] [CrossRef]
- Reeves, R.G.; Phillipson, M. Mass releases of genetically modified insects in area-wide pest control programs and their impact on organic farmers. Sustainability 2017, 9, 59. [Google Scholar] [CrossRef]
- Shelton, A.M.; Long, S.J.; Walker, A.S.; Bolton, M.; Collins, H.L.; Revuelta, L.; Johnson, L.M.; Morrison, N.I. First field release of a genetically engineered, self-limiting agricultural pest insect: Evaluating its potential for future crop protection. Front. Bioeng. Biotechnol. 2020, 7, 482. [Google Scholar] [CrossRef] [PubMed]
- Grilli, S.; Galizi, R.; Taxiarchi, C. Genetic technologies for sustainable management of insect pests and disease vectors. Sustainability 2021, 13, 5653. [Google Scholar] [CrossRef]
- Burt, A. Heritable strategies for controlling insect vectors of disease. Philos. Trans. R. Soc. B Biol. Sci. 2014, 369, 20130432. [Google Scholar] [CrossRef]
- Khalil, A.M. Traditional Strategies and Cutting-Edge Technologies Used for Animal Breeding: A Comprehensive Review. Sustain. Food Connect 2025, 1, 2025.0002. [Google Scholar] [CrossRef]
- Lim, L.W. CRISPR-Cas application in insect: Food, medicine, pest control, disease resistance and textile. Food Biosci. 2025, 171, 07292. [Google Scholar] [CrossRef]
- Gu, J.; Wang, J.; Bi, H.; Li, X.; Merchant, A.; Zhang, P.; Zhang, Q.; Zhou, X. CRISPR/Cas9-mediated mutagenesis of sex-specific doublesex splicing variants leads to sterility in Spodoptera frugiperda, a global invasive pest. Cells 2022, 11, 3557. [Google Scholar] [CrossRef]
- Yogi, D.; Ashok, K.; Anu, C.N.; Shashikala, T.; Pradeep, C.; Bhargava, C.N.; Parvathy, M.S.; Jithesh, M.N.; Manamohan, M.; Jha, G.K.; et al. CRISPR/Cas12a ribonucleoprotein mediated editing of tryptophan 2, 3-dioxygenase of Spodoptera frugiperda. Transgenic Res. 2024, 33, 369–381. [Google Scholar] [CrossRef] [PubMed]
- Zhu, G.H.; Chereddy, S.C.; Howell, J.L.; Palli, S.R. Genome editing in the fall armyworm, Spodoptera frugiperda: Multiple sgRNA/Cas9 method for identification of knockouts in one generation. Insect Biochem. Mol. Biol. 2020, 122, 103373. [Google Scholar] [CrossRef] [PubMed]
- Basit, A.; Mobarak, S.H.; Liu, T.X. CRISPR/Cas9-based strategies for effective control of Spodoptera frugiperda in maize: A review of precision gene editing approaches. Entomol. Gen. 2025, 45, 651–666. [Google Scholar] [CrossRef]
- Tanny, T.; Sallam, M.; Soda, N.; Nguyen, N.T.; Alam, M.; Shiddiky, M.J. CRISPR/Cas-based diagnostics in agricultural applications. J. Agric. Food Chem. 2023, 71, 11765–11788. [Google Scholar] [CrossRef]
- Koonin, E.V.; Gootenberg, J.S.; Abudayyeh, O.O. Discovery of diverse CRISPR-Cas systems and expansion of the genome engineering toolbox. Biochemistry 2023, 62, 3465–3487. [Google Scholar] [CrossRef] [PubMed]
- Bryson, J.W.; Auxillos, J.Y.; Rosser, S.J. Multiplexed activation in mammalian cells using a split-intein CRISPR/Cas12a based synthetic transcription factor. Nucleic Acids Res. 2022, 50, 549–560. [Google Scholar] [CrossRef]
- Abudayyeh, O.O.; Gootenberg, J.S.; Konermann, S.; Joung, J.; Slaymaker, I.M.; Cox, D.B.; Shmakov, S.; Makarova, K.S.; Semenova, E.; Minakhin, L.; et al. C2c2 is a single-component programmable RNA-guided RNA-targeting CRISPR effector. Science 2016, 353, aaf5573. [Google Scholar] [CrossRef]
- Chen, X.; Tan, A.; Palli, S.R. Identification and functional analysis of promoters of heat-shock genes from the fall armyworm, Spodoptera frugiperda. Sci. Rep. 2020, 10, 2363. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y. Advances in CRISPR/Cas9. BioMed Res. Int. 2022, 2022, 9978571. [Google Scholar] [CrossRef] [PubMed]
- Matsumoto, D.; Matsugi, E.; Kishi, K.; Inoue, Y.; Nigorikawa, K.; Nomura, W. SpCas9-HF1 enhances accuracy of cell cycle-dependent genome editing by increasing HDR efficiency, and by reducing off-target effects and indel rates. Mol. Ther. -Nucleic Acids 2024, 35, 102124. [Google Scholar] [CrossRef] [PubMed]
- Dhandapani, R.K.; Gurusamy, D.; Palli, S.R. Protamine–lipid–dsRNA nanoparticles improve RNAi efficiency in the fall armyworm, Spodoptera frugiperda. J. Agric. Food Chem. 2022, 70, 6634–6643. [Google Scholar] [CrossRef]
- Aghamiri, S.; Talaei, S.; Ghavidel, A.A.; Zandsalimi, F.; Masoumi, S.; Hafshejani, N.H.; Jajarmi, V. Nanoparticles-mediated CRISPR/Cas9 delivery: Recent advances in cancer treatment. J. Drug Deliv. Sci. Technol. 2020, 56, 101533. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.