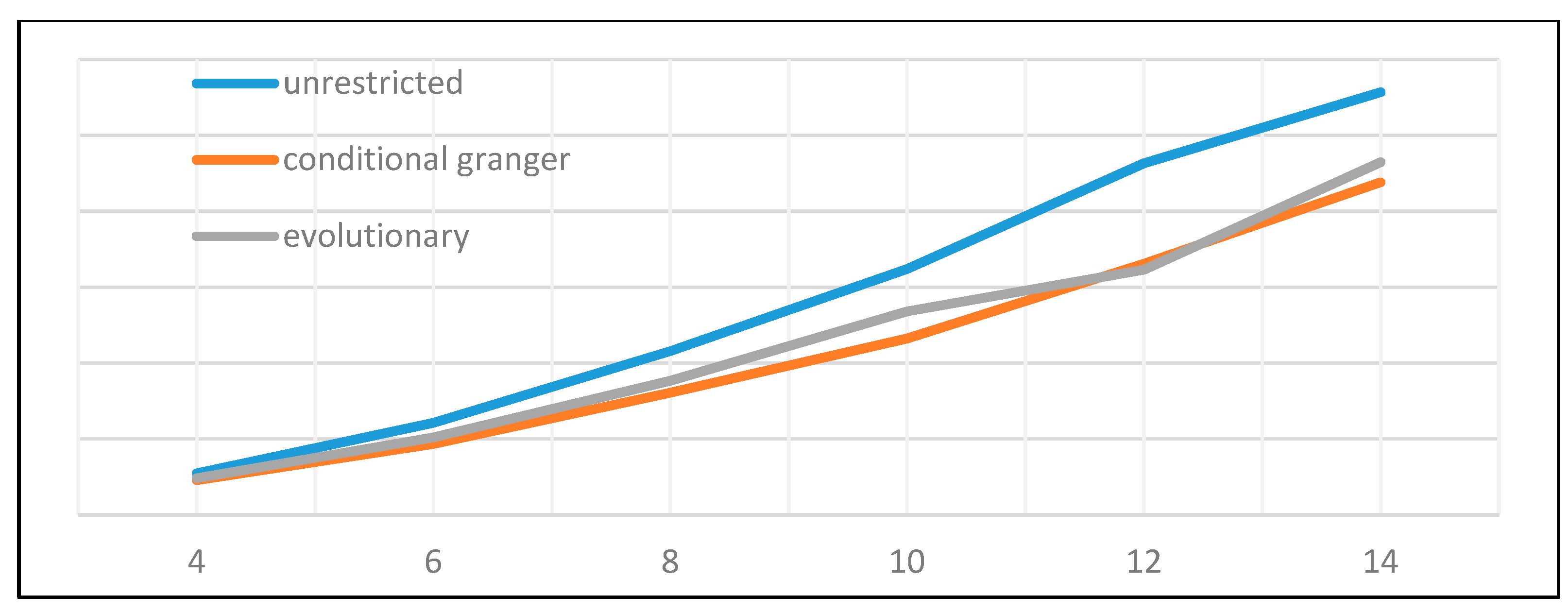
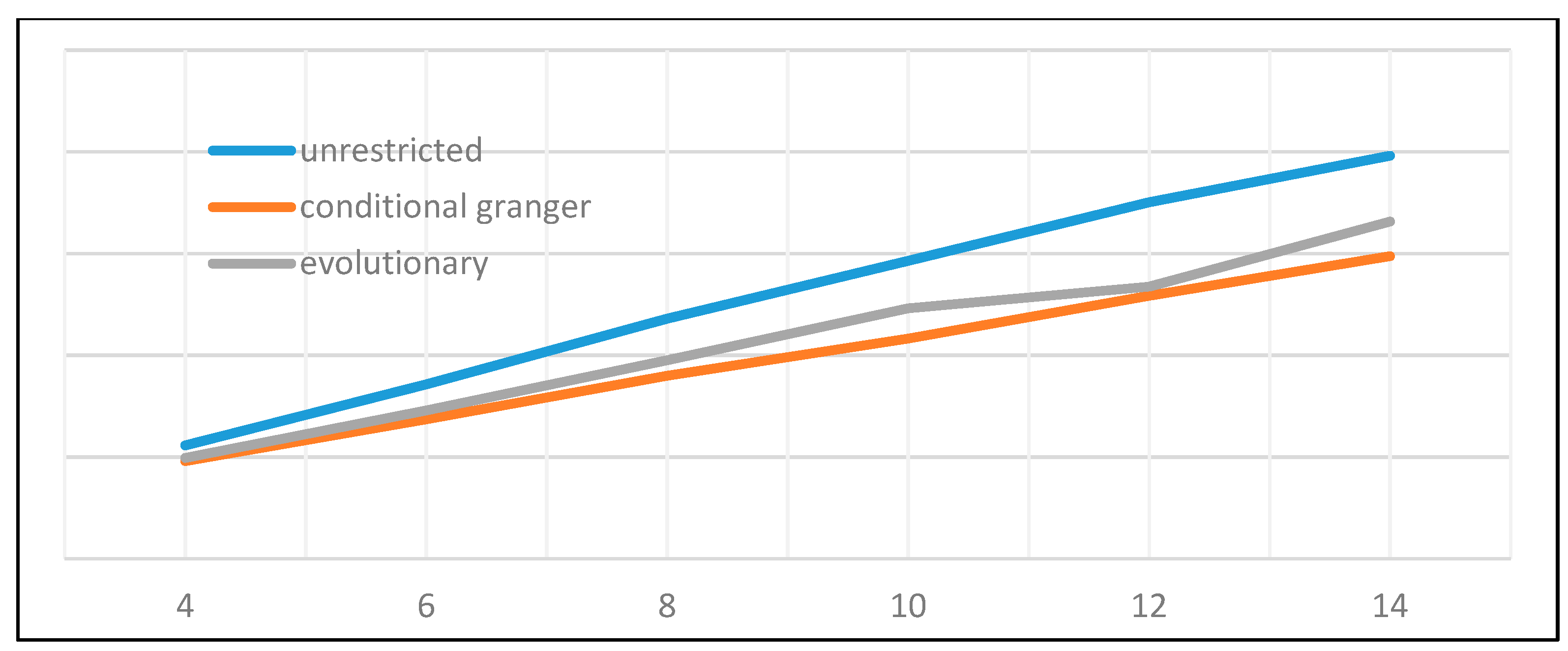
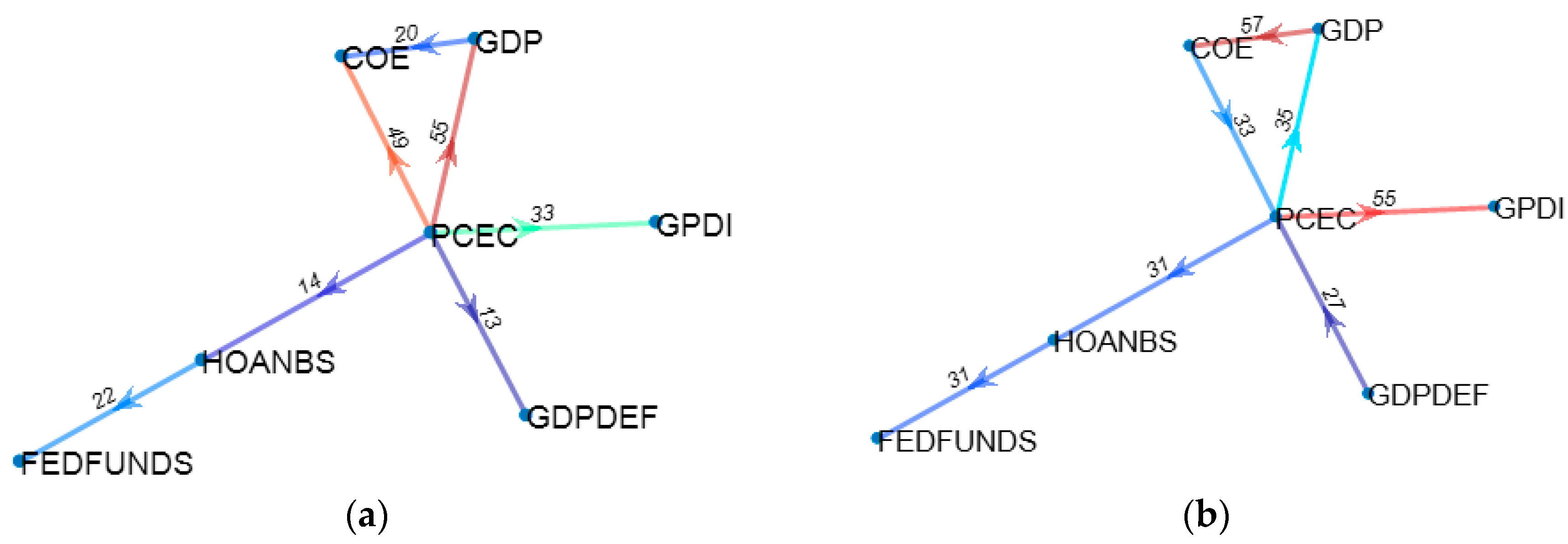
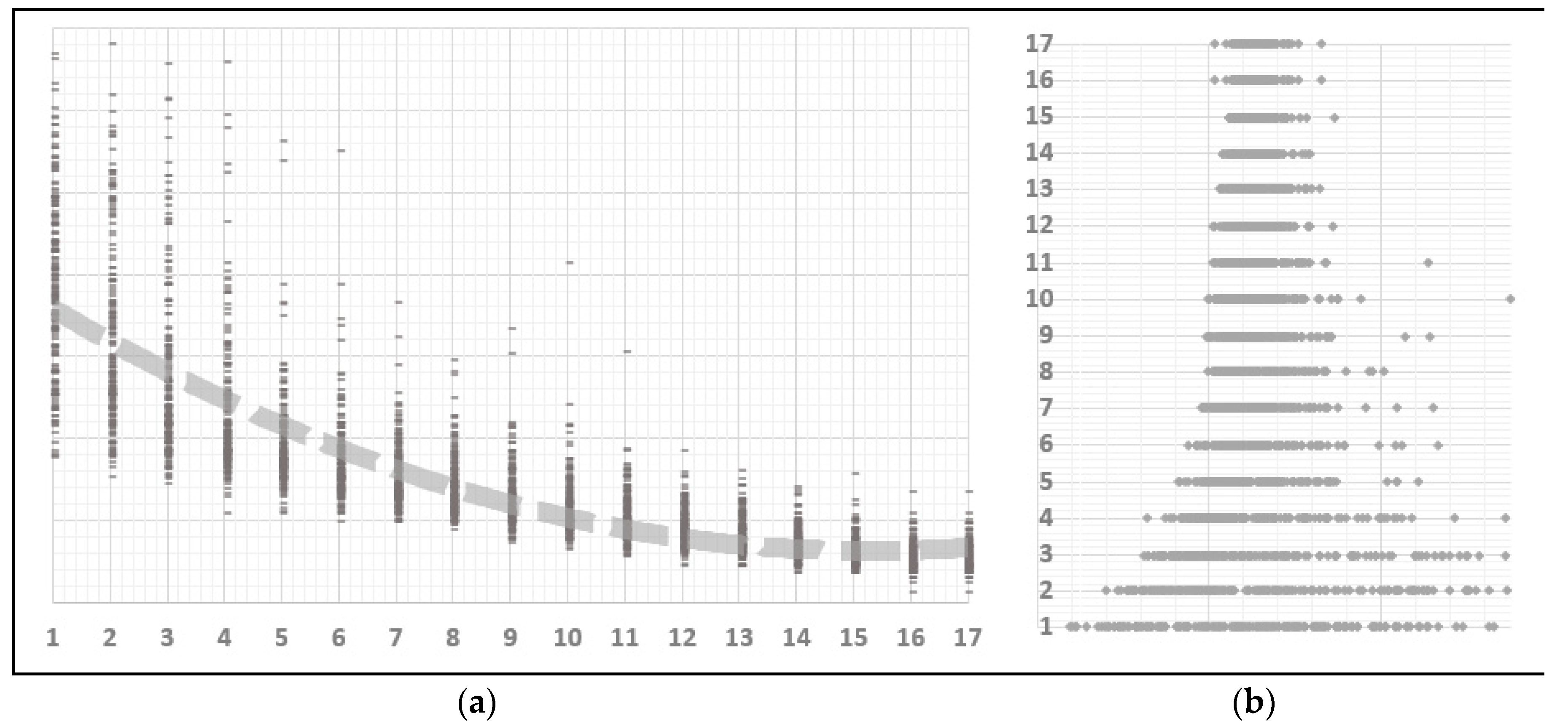
Of the many available possibilities for VAR variable and coefficient selection, four different approaches have been submitted to performance testing. The first one is represented by the unrestricted full VAR, where all positions of the matrix of coefficients are considered viable. This will be the benchmark for comparison against all other methods that attempt to eliminate variables or shrink coefficients. The second algorithm refers to a form of stepwise variable selection based not on consecutive additions or subtractions, but on the principle of conditional Granger causality. Moving forward, the third algorithm belongs to the regression regularization process. Precisely, we apply the lasso regression technique to the equation by equation process of estimating the VAR, with the objective of shrinking irrelevant coefficients towards zero. The fourth and final method focuses on genetic algorithms, with an emphasis on initial population generation using all the previously mentioned methods.
2.1.1. Unrestricted VAR and the Identification Problem
As first defined by Reference [
1], the vector autoregression represents a multivariate stochastic process or a simultaneous equation model of the following form:
Equation (2) is the structural representation of a VAR of order p where each variable y is a (k × 1) vector of historical observation of a stochastic process and εt is a (k × 1) vector of independent and standard normally distributed errors, ε~N(0,1). In the structural form of the VAR, each term of the equation has a specific form with a corresponding functional meaning; thus, B0 is a full rank (k × k) matrix with ones on the main diagonal and possible none zero off-diagonal elements. From a non-statically viewpoint, B0 describes the contemporaneous causality between variables in the system. c0 is a (k × 1) vector of constants (bias), or mean values the variables, εt is the vector of structural shock or innovations. One important feature of these error terms under the structural form of the VAR is that their covariance matrix Σ = E(εtεt’) is diagonal.
Structural VAR models depart from the assumption that all variables are endogenous to the stochastic generation process and all should be taken into consideration. The drawback lies with the presence of
B0, the contemporaneous causality effect matrix that cannot be uniquely identified, requiring
k(
k − 1)/2 zero restrictions in order to render the OLS estimation of the system possible after variable reordering. Moreover, in the structural VAR, the error terms are correlated with the regressors. To solve this issue, both sides of the equation are multiplied by the inverse of
B0, leading to the reduced form of the VAR:
Replacing
B0−1c0 with
c,
B0−1Bp with
Ap and
B0−1εt with
e, the reduced form of (3) can be reinterpreted as:
In the reduced form, Ω = E(εtεt’) is no longer diagonal. If B0 was known, then, going backwards, the system could be rewritten in its structural form and estimated. However, there are many possible values for B0 so that its inverse, multiplied with e, yields an identity matrix. If we instead aim at measuring the covariance between the error terms of Ω = E(εtεt’), the lower triangular part of the matrix has (k2/2) + k positions to estimate with k2 unknowns in the B0 matrix, so the system is undetermined. Faced with these issues, economists resort to short-run restrictions, sign restrictions, long-run restrictions, Bayesian restrictions or variable ordering depending on the degree of exogeneity.
2.1.2. Conditional Granger Causality VAR
In 1969, Clive Granger embedded the intuition of Wiener about causation into the framework of simultaneous equation models with feedback or as stated otherwise, vector autoregression models) [
12,
29]. According to Granger’s original notation, let
Pt(A|B) be the least-squares prediction of a stationary stochastic process
A at time
t, given all available information in set
B, where
B can incorporate past measurements of one or multiple stochastic variables. Given this definition, the least-squares autoregressive prediction for
A can be:
if only the past of A is considered;
if all available information in the measurable past universe () is incorporated;
if the perspective of causality from B to A is to be tested.
A later development of Granger causality [
37] tried to set a clear temporal demarcation line between different types of feedback considering that causation can be decomposed into two instantaneous components and one lagged component:
Besides conceptualizing the decomposition of causality, Geweke proposed a proper measure of “linear feedback” from
A to
B, or stated otherwise, of causality from
A to
B, can be defined as:
As explained by Reference [
37], this measure has multiple benefits when used to explain causality, like strict positiveness, invariance to variable scaling, monotony in the direction of causation, asymptotic chi-square distributional properties. With an in-depth analysis of causality measures being outside the scope of this paper, readers can find a succinct but comprehensive review about the theory behind Granger causality and its applications in Reference [
38]. Recently, it has been shown [
1] that Granger causality is equivalent to the concept of transfer entropy of Reference [
39], bridging the disciplines of autoregressive modeling and information theory.
We further proceed to detail the conditional Granger causality concept for bivariate VAR and continue to the same concept for multivariate VAR. First, consider two random variables
A and
B, defined as below:
The bivariate vector autoregression of the two variables can be represented as:
with the noise covariance matrix:
If the measure of feedback between
A and
B is null, then
and the coefficients
γ2i and
β2i are asymptotically zero. If instead the variables are not independent Equation (6) can be rewritten as
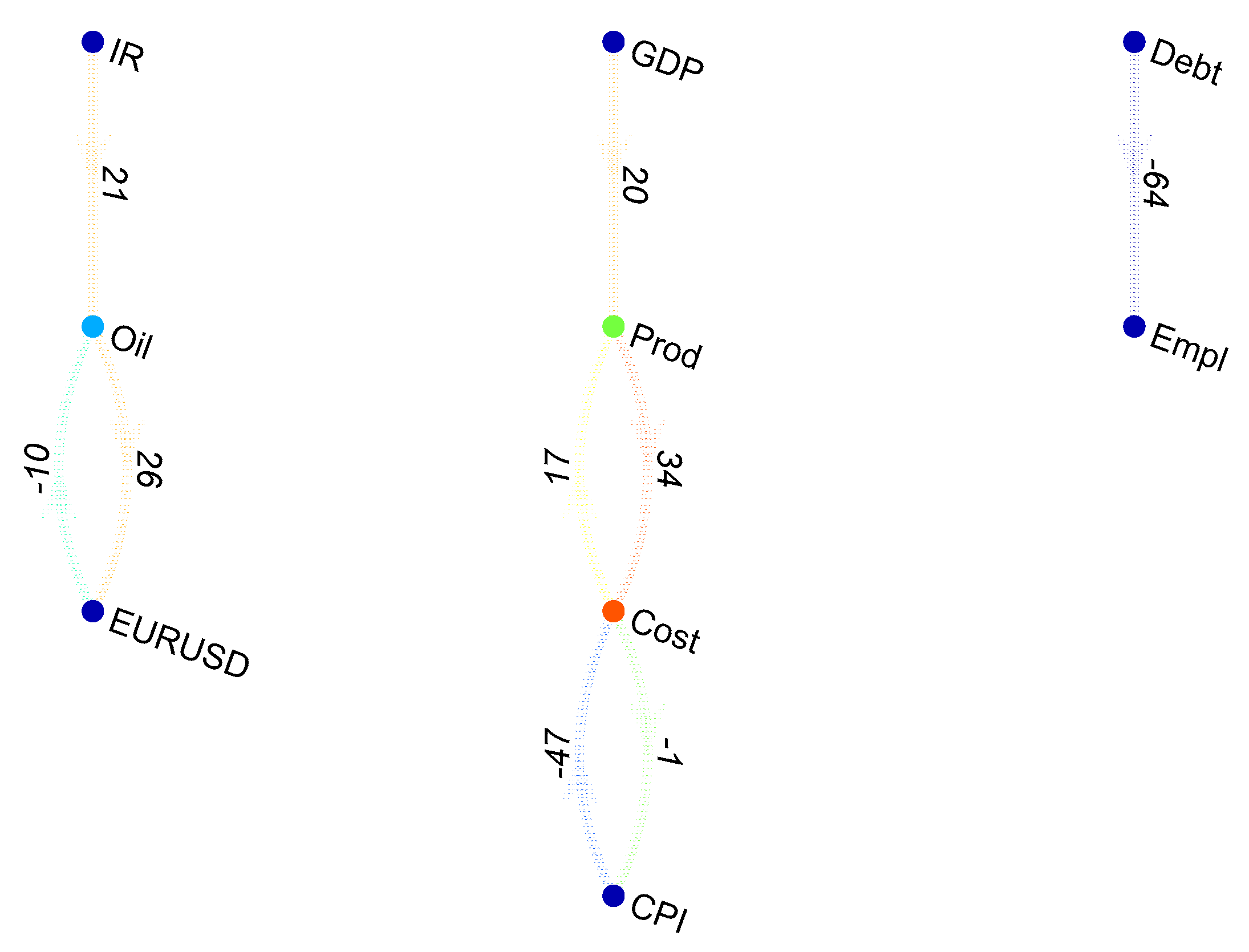
When the conditional Granger causality in a multivariate framework is considered, let us assume that variable
A causes variable
C through the intermediation of
B, as depicted in
Figure 1.
The bivariate vector autoregression of the
A and
C can be represented as
with the noise covariance matrix:
By introducing
B in the equation system, this becomes:
with the noise covariance matrix:
The Granger causality from
A to
C conditional on
B is:
If the causality chain between A and C is entirely dependent on B then the coefficients λ2i should be uniformly zero and, as a consequence, the variance of the terms ε1t and ε2t is identical and therefore is equal to 1. The above reasoning can be extended to systems of more than three variables where the causality is conditioned relative to combinations of multiple time series.
Given the theoretical basis for conditional Granger causality, it is conceivable that a recursive procedure for real causal connection identification can be devised, with the scope of selecting only those coefficients in the VAR that point to direct causation. As such we have employed a network search algorithm that iteratively tests for coefficient significance as explained by Reference [
4].
The variable selection algorithm iterations are the following:
- (1)
Identify for each variable of the dataset which are the individual lags and variables that Granger cause it. These are called “ancestors”.
- (2)
After compiling the first list of “ancestors”, each one is tested for significance in a multivariate VAR by incorporating all possible combinations of two, then three, four, etc. “ancestors”. If during this testing process the coefficient significance becomes null, the candidate “ancestor” is dropped.
- (3)
The previous procedure iterates until all possible testing is completed, leaving only the most resilient “ancestors”.
During algorithm development, variable ordering at initialization was irrelevant to the final estimation. Regardless of variable or lag shuffling, the procedure yielded the same selection results, same coefficient values. This stands to show that the principle of conditional Granger causality is consistent for linear regression.
2.1.3. Lasso VAR
As the VAR has the tendency to become over-parametrized as the number of fitted time series grows, the estimated parameters of the model have to incorporate large amounts of white noise. In theory, OLS estimators have the property of being best linear unbiased estimators under very strong assumptions, according to the Gauss-Markov theorem. The assumptions state that the expected value of the model error terms is zero, all error terms have identic finite variance and they are uncorrelated. Take for example the general case of a linear regression: where . This matrix notation can further be expanded to . The unobserved parameters βj are said to have unbiased estimators if , where , a linear combination of only observed variables. In practice, bias is always present in the estimation results due to unaccounted factors that lead to heteroskedacity of errors and correlations between error terms.
Moving forward, OLS is an optimization technique that aims to minimize a loss function, expressed as the sum of squared residuals of the estimated model with reference to the dataset:
The expected error can be decomposed into parts determined by the “quality” of the model assumptions, the variance of the data around its long term mean, and unexplained variance:
Acknowledgement of these imperfections leads to the understanding that in any estimation procedure, one has to make a tradeoff between the bias and the variance. As the model complexity increases, the bias of OLS estimators decreases. Wanting to decrease complexity, one has to accept a higher variance. In practice, this always leads to a lower in-sample goodness of fit, but better predictions out of sample. Regularization is a technique where minimization of the loss function is penalized for model complexity.
In the case of Lasso regression, the sum of the absolute values of parameters is also subject to minimization alongside the sum of squares:
Whether it is expressed in a standard form dependent on λ or in a Lagrangian form dependent on t, choosing a value for this penalty parameters determines large regression coefficients to shrink in size and thus reduce overfitting. In comparison with ridge regression, where the sum of squared coefficients is minimized, setting a finite value for the sum of absolute coefficient value determines shrinkage of some parameters to zero which is, in fact, equivalent to variable selection.
2.1.4. Genetic VAR
Genetic algorithms represent an optimization heuristic that can generate high-quality solutions by using a global search method that mimics the evolution in the real world. Just like in nature where the fittest individuals survive adverse conditions through competition and selection, genetic algorithms resort to selection, crossover and mutation to choose between possible solutions to an optimization problem. First proposed by Reference [
40], genetic algorithm procedures were formalized and tested through the work of Reference [
27]. Applications of genetic algorithms in the field of statistical analysis have been limited due to their large degrees of freedom that imply a high computational cost. It should be noted that the theoretical background of evolutionary methods is yet to be defined. Nonetheless, genetic algorithms have benefited from a proven record of identifying original solutions to classical problems [
41,
42]. With respect to regression and VAR estimation, we must mention the recent work of References [
32,
35].
In the GA framework, possible solutions to an optimization problem are referred to as “individuals” or “phenotypes”. Each individual is characterized by a set of parameters that form up the individuals’ genotype or a collection of chromosomes. Evolution towards an optimal solution thus becomes an iterative procedure composed of operations like initialization, selection, crossover, mutation and novelty search. At each iteration, the fitness of each surviving individual is assessed and the fitness function represents the objective function that is being optimized.
Just like evolved organisms have necessitated millions of years to perfect themselves, genetic algorithms require numerous trials on random individuals to select the fittest ones. Translating these requirements to variable estimation in the VAR coefficient matrix, an individual is identified by (lag number × n2) + n genes, representing the lagged coefficients and the constant terms, where n is the number of variables in the system. If one coefficient is insignificant, then its corresponding gene value should be zero, meaning that the gene is inactive; otherwise, the gene expression would equal the true value of the coefficient. The purpose of the genetic algorithm is to select those individuals who have a corresponding genotype ensuring the best fitness. Just like in classical VAR estimation, the fitness/objective function should be the maximum likelihood of the model. Other fitness functions like Akaike or Bayesian criteria could be employed, but in order to compare the GA methodology with other well-known procedures, consistency must be maintained with respect to the maximum likelihood objective.
Departing from the theoretical view of linear algebraic equations, the parameter search space of a VAR can also be regarded as an infinite population of candidate solutions. From this vast population, different samples can be extracted and tested for fitness in relation to a given objective function. Through selection, feature cross-over and mutation, a process of repetitive convergence can orient the search towards a good if not optimal solution for the model parameters. In the case of the VAR the principles of competitive evolution are described as follows:
Step 1: Initialization. When doing genetic programming, ensuring a high diversity of individuals is essential for unearthing original combinations. An individual is represented by a matrix of binary entries, equal in size to the VAR matrix of coefficients (number of variables × number of lags). For bitstring chromosomes, a standard method [
43] of uniformly sampling the search space is to toss a fair coin, though empirically better procedures have been proposed by Reference [
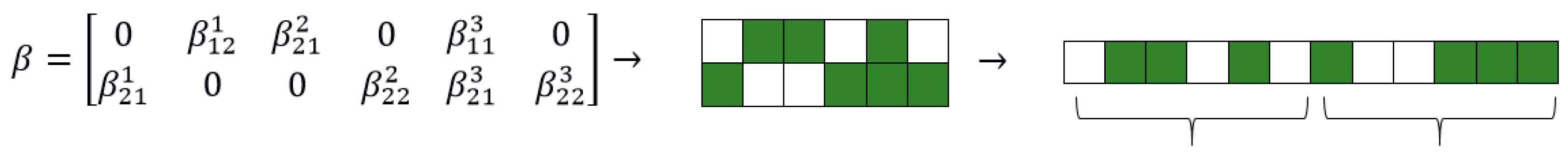
44] The matrix of coefficients is reshaped into a linear vector with the same number of elements in order to create the candidate genome required for subsequent operations of mutation and cross-over, as in
Figure 2.
A zero entry represents an inactive link between a lagged variable and the explained variable specific to each VAR equation, basically a statistically insignificant coefficient. Entries equal to one refer to coefficients that will be later estimated. One requirement for the initialization step is to propose diverse individuals. The more different the proposed genotypes are, the larger the search space covered during initialization. This insight has proven most useful in the machine learning domain where new architectures for problem solving are tedious and hard to conceive [
45].
The random generation of individuals can be very costly in terms of computational effort. Canonical genetic algorithms that perform complete random initialization of the population have been shown to never be able to reach a global optimum without the preservation of the best individual over generations [
46]. One cause of this lies in the symmetry of the search space, where quality solutions are recombined with their complementary counterparts, resulting in random drift [
47]. Moreover, to find a global solution it is necessary that the initial population set contains an individual close to the global optimum which increases time complexity, in contradiction with the optimization philosophy of quick convergence. As an example let us consider the sum of all combinations of n taken k, where n is the product (number of model variables × number of lags). According to the binomial theorem:
. Therefore, the time required to go through all possible coefficient combinations grows exponentially with the number of variables and lags as shown in
Table 1. One possible solution to overcome combinatorial randomness is the application of niching methods for the identification of local optima in the initial solution population [
48].
In order to better guide the inception process towards the zones of potential global optimum, the initial population is inseminated with the results of the simplified, conditional Granger and Lasso VAR. These three individuals will ensure that potentially beneficial genes are identified from the start and if so, are guaranteed to survive overall generation. This is our original approach that not only reduces computation time but as proved in the sections to follow, ensures a good model identification. By having incorporated in its initial genome a set of very fit individuals the VAR estimation algorithm does not stray towards potentially deceiving local optimum.
Step 2: Selection. Having an initial candidate population, the algorithm continues by selecting a small percentage of the fittest individuals. The fitness function takes in the binary coefficient matrix and estimates the VAR only for the non-zero parameters. The output is the matrix of estimators, together with its corresponding log-likelihood. As recommended in the literature a batch of random individuals is also selected, regardless of their fitness [
49,
50]. Despite being counterintuitive, experience in genetic algorithm optimization has proven that some gene combinations must survive the selection process as their utility might become relevant after a few generations [
49].
One of the advantages of genetic algorithms is that the fitness function can take any form. If for OLS the mean squared error is chosen, or the likelihood function for MLE, we chose to minimize the information criteria. To be precise, the fitness function was the product of the Akaike information criterion [
15] and the Bayesian information criterion [
16]. Such a choice was motivated by the actual purpose of the research, that of extracting structural relations from the VAR matrix of coefficients by doing variable selection. Both of the information criteria try to strike a balance between the in-sample goodness of fit and the number of parameters used in the estimation. Denoting the value of the likelihood function with
L, with
n the number of observations and with
k the total number of parameters, the formulae for the two information criteria are:
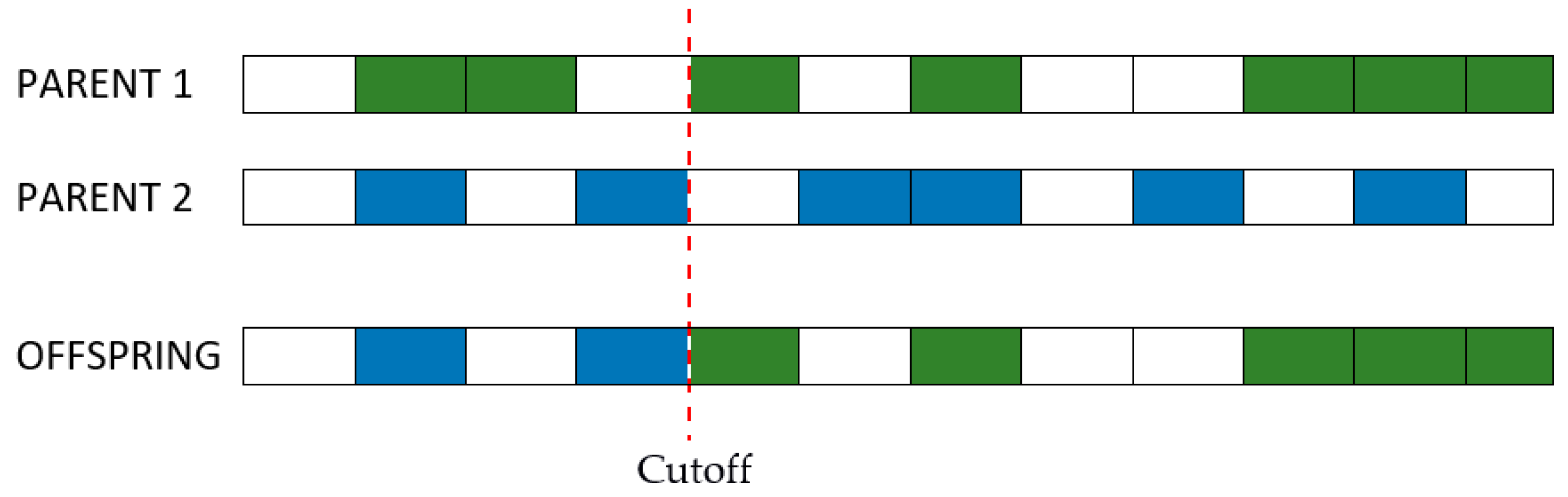
Step 3: Cross-over. Individuals that survived the first generations have their genotypes combined to a certain degree expressed by a percentage at choice. Similar to the natural laws of evolution, the purpose of selection is cross-over and passing of the strongest genes to off-springs. As the number of combinations can become very large even for a small batch of individuals, it is recommended that the cross-over be done between genotypes with a lower degree of similarity, as shown in
Figure 3 [
51].
Step 4: Mutation. The process of continuous selection and crossover may lead to a convergence of the optimization algorithm to local optima. It is then necessary to introduce random mutations to each generation so as to keep open other search paths that might have been missed during initialization. Just like in cross-over, a degree of mutation is chosen beforehand by the user. Precautions must be taken when setting the percentage of genes that can suffer mutations at each generation. A large coefficient can lead the optimization to becoming basically a random search. On the other hand, a small mutation degree can limit the search possibilities and not lead to a globally optimal solution.
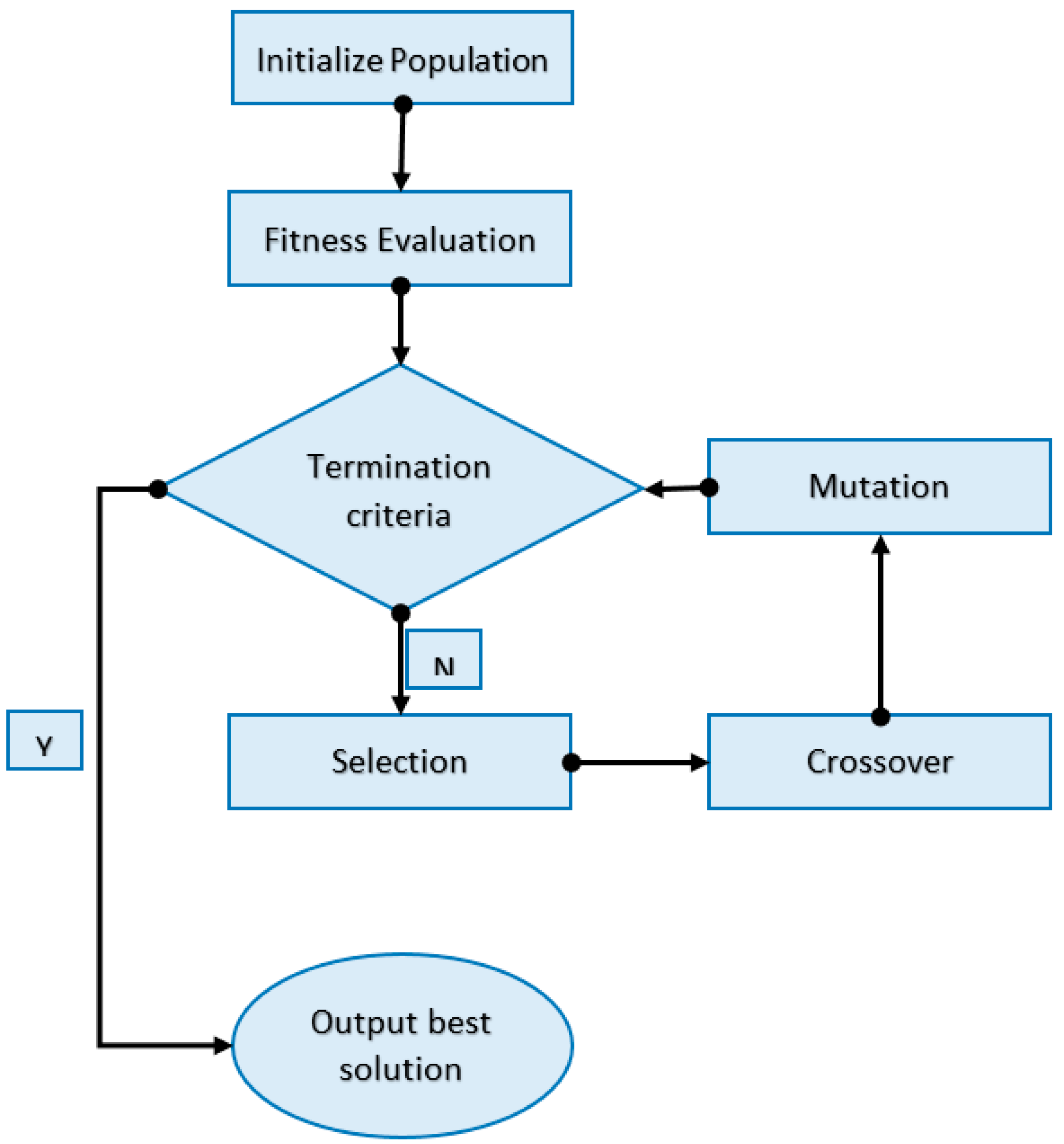
Step 5: Fitness improvement. Iterate through steps 3 and 4 (cross-over and mutation) until the average fitness of each generation cannot be improved. The entire process can be summarized by the diagram in
Figure 4.