Abstract
The dynamic and never exactly repeatable tumor transcriptomic profile of people affected by the same form of cancer requires a personalized and time-sensitive approach of the gene therapy. The Gene Master Regulators (GMRs) were defined as genes whose highly controlled expression by the homeostatic mechanisms commands the cell phenotype by modulating major functional pathways through expression correlation with their genes. The Gene Commanding Height (GCH), a measure that combines the expression control and expression correlation with all other genes, is used to establish the gene hierarchy in each cell phenotype. We developed the experimental protocol, the mathematical algorithm and the computer software to identify the GMRs from transcriptomic data in surgically removed tumors, biopsies or blood from cancer patients. The GMR approach is illustrated with applications to our microarray data on human kidney, thyroid and prostate cancer samples, and on thyroid, prostate and blood cancer cell lines. We proved experimentally that each patient has his/her own GMRs, that cancer nuclei and surrounding normal tissue are governed by different GMRs, and that manipulating the expression has larger consequences for genes with higher GCH. Therefore, we launch the hypothesis that silencing the GMR may selectively kill the cancer cells from a tissue.
1. Introduction
A very rich literature compared gene expression profiles in tissues collected from healthy and cancer donors to identify the transcriptomic signatures of various cancer phenotypes (e.g., references [1,2,3,4,5]) that are periodically organized in the atlas form (e.g., references [6,7]). Nanostring launched recently a panel claiming to categorize the disease heterogeneity using 32 biological signatures involving 770 genes across 23 key breast cancer pathways (https://www.nanostring.com/products/gene-expression-panels/gene-expression-panels-overview/ncounter-breast-cancer-360-panel). There are also available platforms to compare the gene expression profiles of surgically removed tumors with publicly available transcriptomes of cancer standard samples (e.g.: https://www.origene.com/products/tissues/tissuescan).
However, comparing samples collected from different persons may not be such a good idea, because in addition to the disease itself, the gene expressions depend on several other risk factors, making each human unique and with a unique life pathway. The never repeatable combination of factors affecting the gene expression profile is related to the person’s race, sex, age, genetic background, diet (affecting the microbiome), environment (exposure to ionizing radiation, carcinogenic toxins, stress), bad habits (smocking, drugs, alcohol), medial history, etc. Our gene expression studies on tissues from humans and animal models proved the transcriptomic profile dependence on strain and genetic background [8], sex [9], age [10], exposure to stress [11] and carcinogenic toxins [12], medical history and treatments [13]. This is why numerous investigators (e.g., reference [14] on papillary thyroid cancer) started to pair the transcriptomes of the cancer region with the cancer free adjacent tissue of the same patient.
The legitimate question in the transcriptomic signature quest is how many of the tested genes should be found as regulated (and how many of them up and how many down) to assess the designated form of cancer. Since there are 1.9 × 1022 distinct sets of genes if “only” 10 hits are needed from the 770 candidates of the nanostring nCounter® breast cancer 360TM), there is no way to determine the predictive value of each of these sets from metadata. Moreover, in addition to the checked biomarkers, hundreds other genes are regulated and their (never repeatable) contributions to the cancer phenotype are neglected without knowing whether they are really negligible.
Still, let us suppose that a particular cancer form does have a transcriptomic signature, as was indicated by the meta-analysis of gene expression data from a large population of cancer patients. Are the signature genes valuable targets for the cancer gene therapy or are they good only for diagnosis (if the above supposition is true)? Being selected from the most frequently regulated genes in the population of cancer patients, the signature genes appeared as little protected by the cellular homeostatic mechanisms as are minor players. Therefore, restoring their right expression level may be of little consequence for the cell.
Instead of genes whose altered sequence or expression allegedly triggers a particular form of cancer in everybody [15,16,17,18,19,20], we proposed [21,22] that the most legitimate targets for cancer gene therapy are what we call “gene master regulators” (GMRs) of cancer nuclei. We defined the GMR as the gene whose highly protected expression level by the cellular homeostatic mechanisms sets up the cell phenotype by controlling major functional pathways through expression correlation with their genes. The high protection makes the GMR less sensitive to the environmental oscillations and therefore less variably expressed among biological replicas. However, small oscillations of the GMR expression are amplified by in-phase (positive) or anti-phase (negative) oscillations of the expression of many other genes. The composite metric termed Gene Commanding Height (GCH) was introduced by us [21,22] to establish the gene hierarchy in each cell phenotype, with the GMR having the top GCH.
The idea of “master regulators” was floating in genomics for a long time, most investigators looking for transcription factors whose regulation might have large downstream effects on the expression of many genes (e.g., references [23,24]). In addition to defining the GMRs in quantitative terms (by the GCH), our procedure does not restrict the GMR’s quest to transcription factors. Instead, we rank with respect to the GCH scores all coding AND non-coding RNAs whose abundance was adequately quantified via the used (RNA next generation sequence or microarray) platform. Moreover, in a recent paper [22], we have shown how five non-coding RNAs (ANKRD36BP2, FAM86B3P, H19, HCG11 and PMS2L2) regulate apoptosis in a surgically removed papillary thyroid cancer via expression correlation with apoptotic genes. Thus, our results are in line with other studies reporting the involvement of the non-coding RNAs in cancer development (e.g., references [25,26,27]) and therapy (e.g., references [28]).
The GMR approach is based on our Genomic Fabric Paradigm (e.g., references [29,30]) and runs on the computer software package CANCER-GMR (coded in Python v.3) with statistical and graphical user interface packages SciPy (https://www.scipy.org/) and Tkinter GUI. Our procedure can be applied to the four quarters of > 1 mm diameter cancer nucleus identified in a biopsy or surgically removed tumor, or to four dishes with the same cancer cell line.
The GMR targeting would be effective in selectively destroying the cancer cells from a tissue if: (1) cancer nuclei and surrounding quasi-normal tissues are governed by different GCH hierarchies, (2) expression manipulation of a gene has larger consequences in cells where that gene has higher GCH and (3) the GCH of the GMR is well above the GCHs of the next genes in the hierarchy. In this report, the GMR approach is illustrated with applications to our microarray data on human kidney, thyroid and prostate cancer samples, and on thyroid, prostate and blood cancer cell lines.
2. Materials and Methods
2.1. Tumor Samples
We have profiled the cancer nuclei (labeled CANCER1 and CANCER2, both Gleason score 4 + 5 = 9) and the surrounding normal tissues (NORM1 and NORM2) from the frozen, surgically removed prostate tumors of two men. The study was part of Dr. D.A. Iacobas’ project approved by the Institutional Review Boards (IRB) of the New York Medical College’s (NYMC) and Westchester Medical Center (WMC) Committees for Protection of Human Subjects. The approved IRB (L11,376 from 2 October 2015) granted access to frozen cancer specimens from the WMC Pathology Archives and depersonalized pathology reports, waiving patient’s informed consent. Four 2–8 mm 3 samples were collected from the cancer nuclei and normal tissues of each tumor. Although the selected regions were as homogeneous as possible, cells of different phenotypes were not completely eliminated, and expression of their genes affected the reported results.
In this report, we reprocess also our previously published microarray data from surgically removed, frozen preserved kidney (CCRCC—clear cell renal cell carcinoma) and papillary thyroid cancer tumors. Two primary cancer nuclei (labeled as PTA and PTB) and the cancer free (NOR) tissue from the right kidney, together with a chest metastasis (MET) were profiled from a 74 years old man with metastatic CCRCC, Fuhrman grade 3 [21]. The unilateral, single, papillary carcinoma (PAP-C), pathological stage pT3NOMx and the cancer free surrounding tissue were collected from a 33-year-old woman [22].
No treatment was undertaken by the patients whose tumors were profiled and no result of our investigation was revealed to them or to their doctors to influence the therapeutic procedures. The purpose of the study was only to test the GMR approach and show the research community that there are other potentially valuable avenues in cancer therapy that deserve to be explored.
2.2. Cell Lines
The results for the surgically removed thyroid and prostate tumors were compared with those obtained from the commercially available standard human cancer cell lines: BCPAP, 8505C, LNCaP and DU145. We have also determined the GCH hierarchy in the human leukemia cell line HL-60. The HL-60 cell line was originally obtained at MD Anderson Cancer Center from a 36-year-old woman with acute promyelocytic leukemia [31].
The BCPAP cell line is a papillary thyroid carcinoma cell line isolated from a female patient, with a TP53 mutation in the codon 278 in heterozygosity (Pro→Leu) [32]. The 8505C cell line was established from undifferentiated thyroid carcinomas of a 78-year-old-female patient. Her tumor also contained residual well differentiated components, suggesting “well differentiated to undifferentiated carcinoma progression” [33]. We used both BCPAP and 8505C cell lines to test whether manipulation of the expression of a gene has larger transcriptomic consequences if that gene has a higher GCH. As mentioned in a previous publication [22], both cell lines were certified by the Genomic Facility of the Albert Einstein College of Medicine (Bonx, NY 10461, USA) (https://www.einstein.yu.edu/research/shared-facilities/cores/46/genomics/).
The LNCaP cells (Lymph Node Carcinoma of the Prostate) are androgen-sensitive adherent epithelial cells, obtained from a 50-year-old white male in 1977 [34]. The DU145 hormone insensitive cells were derived in 1976 from prostate adenocarcinoma metastatic to the brain of a 70-year-old white male [35].
2.3. Biological Replicas
The biological replicas (the quarters of a quadrisected homogeneous region of a tumor or four cell culture dishes of a cell line) can be considered as being the same system but subjected to slightly different environmental conditions. As such, the transcriptomic data provide valuable information on how much the genes resist or adapt to the external influences and how the variations of their expression levels are correlated to optimize the functional pathways. From the expression values in biological replicas we derive three independent measures to be used in subsequent analyses for each coding and non-coding RNA: (1) average level, (2) coefficient of variation and (3) expression correlation with each other RNA. The average expression level is used to identify what gene is up/down-regulated when compared two conditions. The coefficient of variation (CV) is used to estimate the control of the transcript abundance in each condition, and the expression correlation to identify and quantify the transcriptomic networks.
2.4. Microarray
We have used our standard protocol [36] for RNA extraction, purification, reverse transcription and fluorescent labeling, and hybridization with Agilent human 4 × 44 k gene expression two-color G2519F microarrays. (Full experimental details can be found in the genomic datasets cited at Section 3.1 below, deposited by us in https://www.ncbi.nlm.nih.gov/gds/?term=iacobas). The chips were scanned with an Agilent G2539A dual laser scanner and primary analysis performed with (Agilent) feature extraction v.12.0 software. All Agilent products (equipment, consumables and software were purchased from Agilent Technologies, Santa Clara, CA 95051, USA (https://www.agilent.com/cs/agilent/en/contact-us/united-states) All control spots as well as spots with corrupted or saturated pixels, or with forward fluorescence less than twice the background one in any of the four profiled biological replicas were removed from the analysis of that type of samples.
2.5. Relative Expression Variation (REV) and Relative Expression Stability (RES)
All microarray platforms probe transcripts redundantly by several (unfortunately not uniform numbers of) spots. Therefore, instead of the coefficient of variation for the expression level in biological replicas as determined by one spot we use the Relative Expression Variation (REV, [37]) that takes into account all spots Ri probing redundantly the same transcript i. REV is the mid-interval chi-square estimate of the pooled expression level CVs in biological replicates of that condition (cancer or normal) with a pre-established probability ε and number of degrees of freedom derived from the number of spots probing the same transcript:
where: ski and μki are respectively the standard deviation and the average expression level of the gene i probed by spot k, ri = λRi−1 is the number of degrees of freedom, λ (≥4) is the number of biological replicas and Ri is the number of spots probibng redundantly transcript i.
In our experiments with Agilent 4 × 44 k microarrays the number of spots probing redundantly the same transcript ranges from 1 to 28, so that r = 3, 4, …, 71. Our Cancer GMR software (presented in Section 2.10 below) has uploaded the chi-square values for all these values of r and for ε = 0.010, 0.025, 0.050, 0.100, values less than 0.010 being useless because of the technical noise affecting the gene expression levels. However, in the applications presented here, we used λ = 4 and ε = 0.05 for which the correction coefficient of the CV ranges from 1.566 (R = 1) to 0.960 (R = 28). Therefore, λ and ε will be omitted from the next equations.
Relative Expression Stability (RES) is a measure we introduce to rank the priorities of the cell homeostatic mechanisms in controlling the right abundance of a particular transcript. RES is applied to all transcripts regardless of them translating into proteins or having only regulatory roles for the expression of other transcripts.
The log form was selected to assign positive values to the more stably expressed and negative ones to the less stably expressed genes than the median one.
2.6. Expression Regulation
Instead of an arbitrarily introduced (e.g., 1.5× or 2.0×) absolute fold-change, we consider a gene as significantly regulated in cancer with respect to the normal counterpart if the absolute expression ratio |x| exceeds the cut-off calculated for that gene.
CUT observes the uncertainty about expression regulation by taking into account the contributions of both biological variability and technical noise. CUT is not uniform among the quantified transcripts and takes >1 values that may be even smaller than 1.5.
However, rather than the popular percentage of the regulated out of quantified genes, we measure the change in the transcriptional profiles by the Weighted Pathway Regulation (WPR). WPR is not restricted to the regulated but ponders all quantified genes. We have previously used WPR to quantify the remodeling and recovery of functional genomic fabrics in heart [38], hypothalamus [13] and hippocampus [39]:
The percentage of the regulated genes regards all regulated genes as equal contributors regardless of the fold-change and p-value. In contrast, WPR weights the genes by considering the expression ratio (x), the p-value of the t-test of equal expression, and the average expression level in the normal tissue (µ(normal)).
2.7. Expression Correlation
Pearson pair-wise correlation coefficient ρij was computed for the log2 expressions of all N(N − 1)/2 pairs that can be formed with the N adequately quantified distinct transcripts in all biological replicas. The correlation coefficient takes values from −1 to +1. Close to positive and negative unit ρ’s indicate that the expression of one gene of the pair has strong synergistic or antagonistic consequences on the expression of the other, without specifying what gene comes first. Expressions of synergistic partners fluctuate in phase, those of antagonistic partners fluctuate in antiphase. We believe that strong synergism and antagonism occur when the expressing genes are linked in a functional pathway, providing the “transcriptomic stoichiometry” that rules the expression levels of the involved proteins [40]. Close to zero correlation coefficient means either that the two genes are independently expressed (not networked in any functional pathway) or (very unlikely) that their synergistic correlation in some pathways is balanced by their antagonistic correlation in other pathways.
2.8. Gene Commanding Height (GCH)
The Gene Commanding Height (GCH) was introduced by us [21,22] to quantify the importance of each gene for the cell phenotype. We consider that the expression level of a critical gene for the cell phenotype should be under a stricter control/protection of the homeostatic mechanisms and therefore should have higher expression stability among biological replicas. The same gene should also have a major regulatory role by coordinating the expression of many other genes. If we are allowed a comparison, the most protected persons in the UK are the Queen and the Prime Minister. However, the Prime Minister is the Master Regulator but the Queen is not, owing to the power of the office to oversee all major sectors in the UK policy and economy.
2.9. Gene Ontology and Functional Pathways
Whenever available through Gene Ontology Consortium [41,42], (www.geneontology.org) and/or Kyoto Encyclopedia of Genes and Genomes (KEGG, [43], (http://www.genome.jp)) functional pathways are assigned to the genes. This can be done either by using the KEGG Ontology number as in GAEV [44], (https://github.com/UtaDaphniaLab/Gene_Annotation_Easy_Viewer), either by the gene symbol as in our GMR-Pathway (see below).
In this report we considered the following KEGG pathways: APO (map hsa4210 apoptosis), BTF (hsa03022 basal transcription factors), CCY (hsa04110 cell cycle), CSP (hsa04062 chemokine signaling), OXP (hsa00190 oxidative phosphorylation), RCC (hsa05211 renal cell carcinoma) and RPO (hsa03020 RNA polymerase).
2.10. CANCER-GMR Software
Programs of our CANCER-GMR software package were designed using the Anaconda distribution of Python 3 with statistical and graphical user interface packages such as SciPy (https://www.scipy.org/) and Tkinter GUI (Graphical User Interface). The GMR Software Package includes executable programs to determine the absolute fold-change cut-offs (CUT) when comparing gene expression average levels in cancer and healthy tissues, identify functional pathways within KEGG dbase (PATHWAY), the Weighted Pathway Regulation (WPR), the Pearson correlation coefficients between the expression levels of all gene pairs (CORRELATION) and the Gene Commanding Height (GCH).
#CUT#
REV1 = CORRECTION * std1/mean1
REVV2 = CORRECTION * std2/mean2
CUT = 1 + np.sqrt(2 * (np.square(df[“REV1”]) + np.square(df[“REV2”])))
#PATHWAY#
gene = REST.kegg_get(species+’:’+gene).read()
### find all pathways in KEGG
if current_section == “PATHWAY”:
gene_identifiers = line[1 2:].split(“;”) #Splits each line based on ‘;’
#WPR#
avg_gch = avg_gch/length #avg of GCH of the genes in a certain pathway
wpr = np.mean(avg_gch * (abs(fc)-1) * (1-p_val))
#CORRELATION#
df = pd.read_csv(df, header = 0, na_values = “NaN”)
data = pd.concat([df[condition_input+”1”],df[condition_input+”2”],
df[condition_input+”3”], df[condition_input+”4”]], axis = 1)
logvalues = np.log2(data)
results = (logvalues.T).corr(method = ‘pearson’) #transpose and find correlation
pearsons_df = pd.DataFrame(results.values, columns = df[‘GeneName’], index =
df[GeneName’].values) ## change to dataframe
#GENE COMMANDING HEIGHT#
expcon = median/cov #expression control
results = (logvalues.T).corr(method = ’pearson’) #transpose and find correlation
thesum = (results * results).sum(axis = 0)
### gene commanding height calculations
controlcoor = np.exp((4*thesum-1)/len(pearsons_df.index)-1)
gch = expcon.values * np.exp((4*thesum-1)/len(pearsons_df.index)-1).values
gch = pd.DataFrame(gch, dtype = ’float’)
2.11. Experimental Design to Validate the GMR Theory
One way to validate the GMR theory is to establish the GCH hierarchy in two cell lines, transfect each line with genes having same expression level but different GCHs and compare the transcriptomic alterations (Figure 1). The theory is validated if the alterations of the same gene transfection are higher in the cells where that gene has a higher GCH. We tested the usefulness of the GMR approach for cancer gene therapy by stable lentiviral transfecting the human BCPAP (papillary) and 8505C (anaplastic) thyroid cancer cell lines with four genes NEMP1, PANK2, DDX19B and UBALD1. The selected genes have similar average expression levels (AVE, normalized to the median of all quantified transcripts) but significantly different GCH scores in the two cell lines.
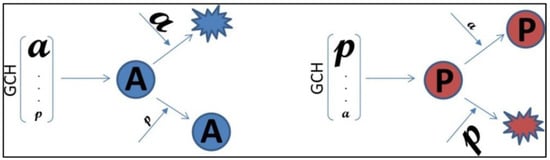
Figure 1.
Experimental design to validate the Gene Master Regulators (GMR) theory. a is a gene with higher Gene Commanding Height (GCH) in the A-cells (here the 8505C anaplastic thyroid cancer cell line) than in the P-cells (here the BCPAP papillary thyroid cancer cell line), while p is a gene with higher GCH in the P-cells than in the A-cells. The theory is verified if transfection of a will induce significantly larger Weighted Pathway Regulation (WPR) in the A-cells than in the P-cells and transfection of p will induce larger WPR in the P-calls than in the A-cells
3. Results
3.1. Experimental Data
The GMR approach is illustrated here with applications to our gene expression datasets from surgically removed human (kidney, thyroid, prostate) cancer tissues and commercially available human cancer cell lines deposited by us in the Gene Expression Omnibus of the National Center for Biotechnology Information (www.ncbi.nlm.nih.gov/gds). Data used in this report were from GSE72304 (a case of metastatic clear cell renal cell carcinoma (CCRCC), GSE97001 (a case of papillary thyroid cancer), GSE97002 (BCPAP papillary and 8505C anaplastic thyroid cancer cell lines) and from two cases of prostate cancer (GSE133891, GSE133906). Expression data for the GMR Theory validation on thyroid cancer cell lines BCPAP and 8505C were collected from GSE97031 (transfection with NEMP1), GSE97028 (DDX19B), GSE97030 (PANK2) and GSE97427 (UBALD1). Other expression data from cancer cell lines were collected from GSE72333 (DU145), GSE72414 (LNCaP) and GSE72415 (HL-60).
3.2. Expression Stability, Expression Correlation and Weighted Pathway Regulation
The Relative Expression Stability (RES) can be used not only to establish the hierarchy of individual genes but also the hierarchy of functional pathways. Figure 2A presents the average RES scores of several pathways analyzed in each of the four regions profiled from the CCRCC samples. The averages were determined for 107 APO, 37 BTF, 131 CSP, 91 CCY, 100 OXP, 54 RCC and 31 RPO genes out of the 12610 distinct genes whose expression was adequately quantified in all four regions of the CCRCC samples. WPR analysis of the CCRCC samples for the same pathways returned the results from Figure 2B. Note that (by far) the most affected pathway was (as expected) RCC (the renal cell carcinoma) followed by OXP (oxidative phosphorylation).
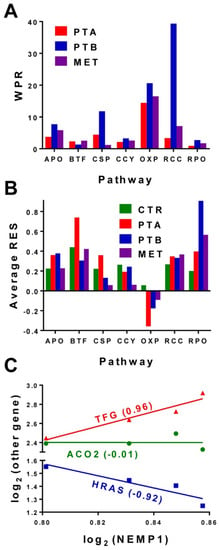
Figure 2.
Expression data from [21]: (A). Average Relative Expression Stability (RES) of several functional pathways in the four regions profiled from the clear cell renal cell (CCRCC) samples. Note that the RNA polymerase (RPO) and basal transcription factors (BTF) are the most protected pathways, while the control of the oxidative phosphorylation (OXP) is relaxed, presumably to allow the cells to adapt the environmental conditions. (B). Weighted Pathway Regulation (WPR) analysis of several functional pathways. Pathways: APO = apoptosis, BTF = basal transcription factors, CCY = cell cycle, CSP = chemokine signaling, OXP = oxidative phosphorylation, RCC = renal cell carcinoma, RPO = RNA polymerase. (C). Example of the correlation analysis in BCPAP cells (expression data from [22]). NEMP1 is synergistically expressed with TFG (trafficking from ER to golgi regulator), antagonistically expressed with HRAS (Harvey rat sarcoma viral oncogene homolog) and independently expressed with ACO2 (aconitase 2, mitochondrial) in BCPAP cells.
Figure 2C illustrates the correlation analysis with examples of synergistically, antagonistically and independently expressed partners of NEMP1 (nuclear envelope integral membrane protein 1) in the BCPAP cells. In our opinion, the strong correlations of the (not yet assigned to a pathway) NEMP1 gene with the oncogenes TFG and HRAS indicate its potential role in the papillary thyroid cancer. HRAS is among the most documented genes whose mutations have been associated with thyroid cancer [45]. TFG gene is described in https://www.ncbi.nlm.nih.gov/gene/10342 as partially encoding several fusion oncoproteins and participating in several “oncogenic rearrangements resulting in anaplastic lymphoma and mixoid chondrosarcoma”. TFG-MET (MET proto-oncogene receptor tyrosine-kinase) translocation was reported in a follicular variant of the papillary thyroid carcinoma [46]. Interestingly, NEMP1 was recently shown as promoting tamoxifen resistance in breast cancer cells [47]. Thus, the correlation analysis may be used to refine the maps of the functional pathways by determining the gene pairs whose correlated expression may result from a functional relationship between their encoded proteins.
3.3. Cancer Nuclei and Surrounding Normal Tissue Are Governed by Distinct GMRs
Table 1, Table 2 and Table 3 present the GCH scores of the top 3 genes in cancer nuclei and surrounding normal tissue in a case of metastatic clear cell renal cell carcinoma, a case of papillary thyroid cancer and two cases of prostate cancer.

Table 1.
Expression data from GSE72304, [21].

Table 2.
Expression data from GSE97001, GSE97002, [22].

Table 3.
Data from GSE133891 and GSE133906.
In Table 1, only ALG13 is an actionable GMR (for MET region) owing to its significantly higher GCH (82.95) with respect to the second gene, NUDT18 (GCH = 48.40). Interestingly, ALG13 (an early target of miR-34a) was reported as correlated with worse clinical outcomes for neuroblastoma [48]. Results in Table 1 also indicate that distinct cancer nuclei of the same tumor (PTA, PTB) may have a distinct gene hierarchy that explains their phenotypic diversity. Note also that MIR1915HG, previous names: chromosome 10 open reading frame 114, cancer susceptibility candidate 10 and cancer susceptibility 10, is a long non-coding RNA.
In a previous publication [22], we have presented the GCH scores of 78 cancer biomarkers, 44 oncogenes, 55 apoptosis genes and 120 ncRNAs in the cancer and normal areas of a surgically removed papillary thyroid tumor. In that selection of 297 genes, all but RAB15 (GCH(cancer) = 26.14) had GCH scores below 20. The complete GCH analysis of the expression data in the same tumor sample revealed that the GMR of the cancer area is SPINT2 (GCH(cancer) = 54.97). Interestingly, SPINT2 a transmembrane protein that inhibits serine proteases implicated in cancer progression [49], acts as a putative tumor suppressor when hypermethylated [50].
It is notable that WFDC3, the far above GMR of the androgen-sensitive prostate cancer LNCaP cells, was also reported as one of the most down-regulated gene in the ventral prostate of aged (18 moths) estrogen receptor β−/− mouse [51]. Interestingly, the GMR of the cancer nucleus of the second man (LOC145474) is a non-coding RNA, confirming that both coding and non-coding RNAs may play dominant roles in prostate tumorigenesis [52]. To our knowledge, this is the first time that LOC145474 is reported as related to a prostate cancer.
3.4. Experimental Validation of the GMR Theory
Our experimental results (summarized in Figure 3) on the 8505C (anaplastic, 3A & 3C) and BCPAP (papillary, 3B & 3D) human thyroid cancer cell lines stably transfected with DDX19B, NEMP1, PANK2 or UBALD1 (characteristics in Figure 1) indicate that:
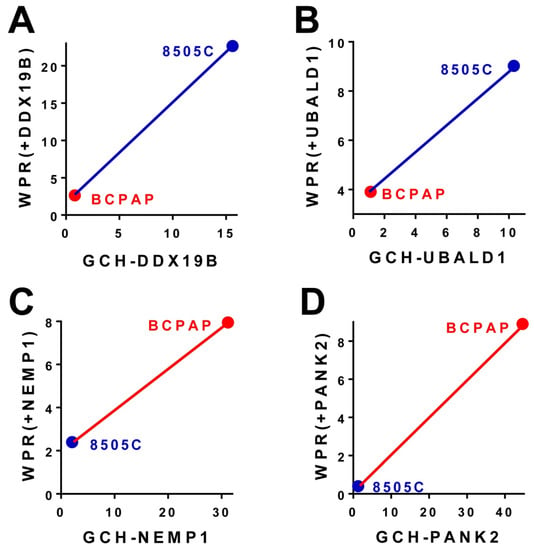
Figure 3.
Data from GSE97031, GSE97028, GSE97030, GSE97427, [22]: Validation of the GMR Theory. (A,B). Stable transfection of genes with higher GCH in the 8505C cells than in BCAP cells had larger transcriptomic effects in 8505C cells as measured by the Weighted Pathway Regulation (WPR). (C,D). Stable transfection of genes with higher GCH in the BCPAP cells than in 8505C cells had larger transcriptomic effects in BCPAP cells as measured by the Weighted Pathway Regulation (WPR).
We have also observed that transfections of NEMP1 and PANK2 significantly slowed down multiplication of BCPAP cells, while transfection of DDX19B or UBALD1 had little effect on these cells. By contrast, both transfections of DDX19B and UBALD1 significantly slowed down grow of 8505C cells, while transfection of NEMP1 and PANK2 had little effect. Together, these observations confirm that expression manipulation of a gene has larger consequences in cells where that gene has higher GCH.
3.5. Predicted Transcriptomic Alteration by GMR Manipulation
At the time, we had no possibility to alter the expression of the GMRs identified in the thyroid cancer cell lines 8505C and BCPAP. Figure 4 presents the predicted Weighted Pathway Regulation if significantly altering the expressions of the top three genes in the 8505C and BCPAP cells.
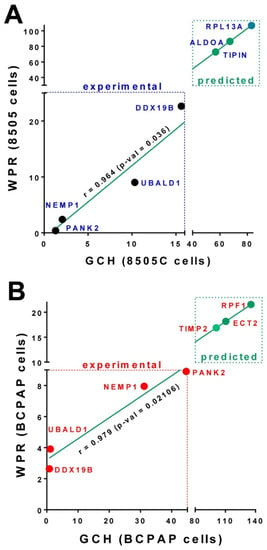
Figure 4.
(A). Experimentally measured and theoretically predicted effects of stably transfecting PANK2, NEMP1, UBALD1, DDX19B and the top three genes (RPL13A, ALDOA and TIPIN) in the 8505C. (B). Experimentally measured and theoretically predicted effects of stably transfecting PANK2, NEMP1, UBALD1, DDX19B and the top three genes (RPF1, ECT2 and TIMP2) in the BCPAP cells.
Interestingly, the GMR of the anaplastic cell line (RPL13A) was found as the most stably expressed gene in two ovarian cancer cell lines (UACC–1598 and SKOV3) subjected to two widely used anticancer treatment [53], confirming the major role played by this gene in stabilizing the cancer phenotype.
3.6. Ribosomal Genes Top the Hierarchy in the Acute Promyelocytic Leukemia HL-60 Cell Line
An interesting gene hierarchy was obtained for the HL-60 cells (Figure 5), where most of the ribosomal genes top all KEGG identified genes as associated with acute myeloid leukemia (AML, https://www.kegg.jp/kegg-bin/show_pathway?map=has05221&show_description=show). Thus, the top two AML genes, MPO (myeloperoxidase) and PML (promyelocytic leukemia) have the GCHs 22.96 and 21.95, below those of the 56th and the 59th ranked ribosomal proteins RPL29 (23.64) and RPL13 (22.12). As presented in Figure 5C, genes from both large (RPL) and small (RPS) ribosomal subunits are among the highest ranked genes in the HL-60 cells. According to our results, certain ribosomal genes (RPL13A, RPS5) are more influential in dictating/preserving the HL-60 phenotype than RARA (retinoic acid receptor alpha, GCH = 19.67), the gene whose translocation t (15;17) [54,55] is associated with 98% of acute promyelocytic leukemia cases. Interestingly, RPL13A was also found to be the most influential gene in 8505C (anaplastic thyroid cancer) cells and to have a high GCH (63.26) in the BCPAP cells (Table 2 above). For comparison, Figure 5 also presents the GCH scores for the genes associated with the immune system KEGG pathway of leukocyte transendothelial migration (https://www.kegg.jp/kegg-bin/show_pathway?map=hsa04670&show_description=show).
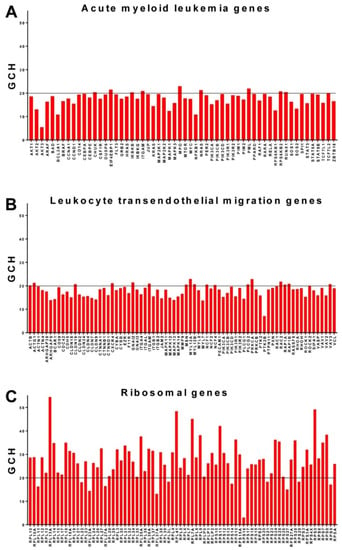
Figure 5.
Data from GSE72415: Gene Commanding Height (GCH) scores in the acute promyelocytic leukemia cell line HL-60. (A). GCH scores of genes associated with acute myeloid leukemia. (B). GHC scores of genes associated with KEGG pathway of leukocyte transendothelial migration. (C). GCH scores of the ribosomal genes.
4. Discussions
In this report, we propose that the most legitimate targets for the cancer gene therapy are the genes whose highly controlled expressions by the homeostatic mechanisms are the most influential by being correlated with expressions of many other genes. We termed these targets Gene Master Regulators (GMRs), and developed and used the necessary experimental protocol, mathematical algorithm and computer software to identify them from gene expression data. Owing to the astronomic number of possible combinations of risk factors (race, sex, age, medical history, environment, diet, bad habits etc.), each cancer patient has his/her unique gene hierarchy and therefore the GMRs should be identified for each patient separately. Most likely, the gene hierarchy is not rigid but changes in time with the progression of the disease or/and in response to a treatment and environmental changes. Therefore, the gene therapy should be both personalized and timely applied.
Our approach is a significant departure from the identification of aberrantly expressed tumor specific genes that may (or may not) have caused cancerization. Instead, we are looking for the genes whose targeting would selectively kill the cancer cells from the tissue. Such genes have the expression so strongly correlated with that of many other genes that their forced alteration triggers unbearable perturbations of major functional pathways. In tens of previous papers on a large variety of cells and tissues from humans [21,22] and animal (e.g., mouse [10,11,29,38], rat [13,30,39], rabbit [56], dog, chicken embryo [12]) models of human diseases, we proved that major functional pathways remodel in disease, with certain genes being promoted to command positions. We know that a gene is critical for a particular phenotype when its expression is strongly controlled by the homeostatic mechanisms, making it resistant to the small environmental fluctuations and thus very stably expressed across biological replicas.
The GMR approach was applied to our microarray data on five standard (anaplastic thyroid, papillary thyroid, prostate androgen-sensitive and insensitive, and blood) cancer cell lines and ten profiled regions from surgically removed CCRCC (4), prostate (4) papillary thyroid (2) cancers. In each tumor sample, the fully profiled transcriptome(s) of the cancer nodule(s) was(were) compared to that of the cancer-free resection margins. No particular criterion was used in selecting the tumors excepting their availability and to provide enough diversity for the GMR proof of concept. At this stage of the research, the types of cancer are less important. Although the tested population was very small, increasing its size most likely would not provide common targets for different patients but will increase the confidence in the approach. GCH analysis is highly sensitive and reveals that, although similar, humans are not identical. Nor are the gene hierarchies, even in the normal regions or in cancer nodules with the same Gleason score (as in Table 3). Anyhow, collaborative experiments with medical centers are under way to test our approach on several other cancers.
Even though we did our best to select as histo-pathologically homogeneous as possible the regions to profile, the samples were still heterogeneous and cells of other than the dominant phenotype (normal or cancer) affected the results. However, our manuscript does not intend to provide definite results for kidney, papillary and anaplastic thyroid, androgen-sensitive and androgen-insensitive prostate or blood cancers, but to show a way to identify potential targets for gene therapy of any cancer type. Nonetheless, next round of experiments will include separation of the tumor cells into distinct phenotypes with our Sony SH8005 sorter and then profile separately their transcriptomes.
The present studies confirmed that the GMRs may differ even for patients with the same form of cancer (like in the above two prostate cases and in comparisons with corresponding cell lines), justifying the necessity of a personalized approach of cancer gene therapy. We found that the GMRs can be located in any chromosome, that their transcripts can be both coding and non-coding RNAs and that the encoded proteins may be involved in a wide diversity of biological processes. Although we don’t have yet the experimental evidence, most likely the gene hierarchy changes in time, so that the GMRs should be targeted as soon as possible after their identification.
Importantly, we found that the cancer nuclei and the surrounding normal tissues are governed by different GMRs and that manipulation of the expression of a gene has consequences in line with its Gene Commanding Height. Based on these findings, we provide the hypothesis that silencing the GMR (using CRISPR or shRNA) may selectively kill the cancer cells with little effect on the normal cells of the tissue. However, we generally we found the GMR to be well above the rest of the genes, with the most notable exception in this report being WFDC3 (GCH = 173.58) in LNCaP cells (next gene is RPL31 at 39.11). Therefore, this hypothesis would work much better for cases where the GMR has a significantly dominant GCH over the other genes in the cancer nucleus and a very low GCH in the normal cells of the tissue. If the GMR transcript is coding, then its forced alteration affects directly the encoded protein, although coding or non-coding, through the strong expression coordination and manipulation of the GMR should affect the levels of numerous proteins involved in major functional pathways.
At this moment, we do not know whether targeting cancer GMRs may improve the results of the chemo- or radiation therapy, but combining gene manipulation with traditional treatment (as suggested by one of the reviewers) appears to be a very interesting idea.
Author Contributions
Conceptualization, D.A.I.; Methodology, D.A.I., S.I.; Software, N.E.; Validation, S.I., D.A.I.; Formal Analysis, S.I., D.A.I.; Investigation, S.I., D.A.I.; Resources, D.A.I.; Data Curation, D.A.I.; Writing—Original Draft Preparation, D.A.I.; Writing—Review and Editing, D.A.I.; Visualization, D.A.I.; Supervision, D.A.I.; Project Administration, D.A.I.
Acknowledgments
This research was funded in part by the Texas A&M University System Chancellor’s Research Initiative for Center for Computational Systems Biology at the Prairie View A&M University (D.A.I.) and by the Prairie View A&M University Undergraduate Scholarly Research Award (N.E.).
Conflicts of Interest
The authors declare no conflict of interest.
References
- Erstad, D.J.; Fuchs, B.C.; Tanabe, K.K. Molecular signatures in hepatocellular carcinoma: A step toward rationally designed cancer therapy. Cancer 2018, 124, 3084–3104. [Google Scholar] [CrossRef] [PubMed]
- Kretschmer, A.; Tilki, D. Biomarkers in prostate cancer—Current clinical utility and future perspectives. Crit. Rev. Oncol. Hematol. 2017, 120, 180–193. [Google Scholar] [CrossRef] [PubMed]
- Lam, M.; Roszik, J.; Kanikarla-Marie, P.; Davis, J.S.; Morris, J.; Kopetz, S.; Menter, D.G. The potential role of platelets in the consensus molecular subtypes of colorectal cancer. Cancer Metastasis Rev. 2017, 36, 273–288. [Google Scholar] [CrossRef] [PubMed]
- Sacco, A.; Fenotti, A.; Affò, L.; Bazzana, S.; Russo, D.; Presta, M.; Malagola, M.; Anastasia, A.; Motta, M.; Patterson, C.J.; et al. The importance of the genomic landscape in Waldenström’s Macroglobulinemia for targeted therapeutical interventions. Oncotarget 2017, 8, 35435–35444. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Santosh, A.B.; Jones, T.; Harvey, J. A review on oral cancer biomarkers: Understanding the past and learning from the present. J. Cancer Res. Ther. 2016, 12, 486–492. [Google Scholar] [CrossRef]
- The Cancer Genome Atlas (TCGA) Research Network; Weinstein, J.N.; Collisson, E.A.; Mills, G.B.; Mills Shaw, K.R.; Ozenberger, B.A.; Ellrott, K.; Shmulevich, I.; Sander, C.; Stuart, J.M. The Cancer Genome Atlas Pan-Cancer analysis project. Nat. Genet. 2013, 45, 1113–1120. [Google Scholar] [CrossRef]
- Uhlen, M.; Zhang, C.; Lee, S.; Sjöstedt, E.; Fagerberg, L.; Bidkhori, G.; Benfeitas, R.; Arif, M.; Liu, Z.; Edfors, F.; et al. A pathology atlas of the human cancer transcriptome. Science 2017, 357, eaan2507. [Google Scholar] [CrossRef]
- Iacobas, S.; Iacobas, D.A.; Spray, D.C.; Scemes, E. The connexin43 transcriptome during brain development: Importance of genetic background. Brain Res. 2012, 1487, 131–139. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Iacobas, S.; Thomas, N.; Spray, D.C. Sex-dependent gene regulatory networks of the heart rhythm. Funct. Integr. Genomics 2010, 10, 73–86. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Fan, C.; Iacobas, S.; Spray, D.C.; Haddad, G.G. Transcriptomic changes in developing kidney exposed to chronic hypoxia. Biochem. Biophys. Res. Comm. 2006, 349, 329–338. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Fan, C.; Iacobas, S.; Haddad, G.G. Integrated transcriptomic response to cardiac chronic hypoxia: Translation regulators and response to stress in cell survival. Funct. Integr. Genomics 2008, 8, 265–275. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kobets, T.; Iatropoulos, M.J.; Duan, J.D.; Brunnemann, K.D.; Iacobas, D.A.; Iacobas, S.; Vock, E.; Deschl, U.; Williams, G.M. Effects of Nitrosamines on the Expression of Genes Involved in Xenobiotic Metabolism in the Chicken Egg Alternative Genotoxicity Model. Toxicol. Sci. 2018, 166, 82–96. [Google Scholar] [CrossRef] [PubMed]
- Iacobas, D.A.; Chachua, T.; Iacobas, S.; Benson, M.J.; Borges, K.; Veliskova, J.; Velisek, L. ACTH and PMX53 recover the normal synaptic transcriptome in a rat model of infantile spasms. Sci. Rep. 2018. [Google Scholar] [CrossRef]
- Teng, H.; Mao, F.; Liang, J.; Xue, M.; Wei, W.; Li, X.; Zhang, K.; Feng, D.; Liu, B.; Sun, Z. Transcriptomic signature associated with carcinogenesis and aggressiveness of papillary thyroid carcinoma. Theranostics 2018, 8, 4345–4358. [Google Scholar] [CrossRef] [PubMed]
- Coghlin, C.; Murray, G.I. Biomarkers of colorectal cancer: Recent advances and future challenges. Proteomics Clin. Appl. 2015, 9, 64–71. [Google Scholar] [CrossRef]
- Vogelstein, B.; Papadopoulos, N.; Velculescu, V.E.; Zhou, S.; Diaz, L.A., Jr.; Kinzler, K.W. Cancer genome landscapes. Science 2013, 339, 1546–1558. [Google Scholar] [CrossRef] [PubMed]
- Estevez-Garcia, P.; Rivera, F.; Molina-Pinelo, S.; Benavent, M.; Gómez, J.; Limón, M.L.; Pastor, M.D.; Martinez-Perez, J.; Paz-Ares, L.; Carnero, A.; et al. Gene expression profile predictive of response to chemotherapy in metastatic colorectal cancer. Oncotarget 2015, 6, 6151–6159. [Google Scholar] [CrossRef]
- Li, H.; Samawi, H.; Heng, D.Y. The use of prognostic factors in metastatic renal cell carcinoma. Urol. Oncol 2015, 33, 509–516. [Google Scholar] [CrossRef]
- Martínez-Bosch, N.; Rodriguez-Vida, A.; Juanpere, N.; Lloreta, J.; Rovira, A.; Albanell, J.; Bellmunt, J.; Navarro, P. Galectins in prostate and bladder cancer: Tumorigenic roles and clinical opportunities. Nat. Rev. Urol. 2019, 16, 433–445. [Google Scholar] [CrossRef]
- Tang, K.; Xu, H. Prognostic value of meta-signature miRNAs in renal cell carcinoma: An integrated miRNA expression profiling analysis. Sci. Rep. 2015, 14, 10272. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Iacobas, S. Towards a personalized cancer gene therapy: A case of clear cell renal cell carcinoma. Cancer Oncol. Res. 2017, 5. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Tuli, N.; Iacobas, S.; Rasamny, J.K.; Moscatello, A.; Geliebter, J.; Tiwari, R.J. Gene master regulators of papillary and anaplastic thyroid cancer phenotypes. Oncotarget 2018, 9, 2410–2424. [Google Scholar] [CrossRef] [PubMed]
- Clevers, H.C.; Owen, M.J. Towards a molecular understanding of T-cell differentiation. Immunol. Today 1991, 12, 86–92. [Google Scholar] [CrossRef][Green Version]
- Doevendans, P.A.; van Bilsen, M. Transcription factors and the cardiac gene programme. Int. J. Biochem. Cell Biol. 1996, 28, 387–403. [Google Scholar] [CrossRef]
- Liyanarachchi, S.; Li, W.; Yan, P.; Bundschuh, R.; Brock, P.; Senter, L.; Ringel, M.D.; de la Chapelle, A.; He, H. Genome-Wide Expression Screening Discloses Long Noncoding RNAs Involved in Thyroid Carcinogenesis. J. Clin. Endocrinol. Metab. 2016, 101, 4005–4013. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.Y.; Lin, X.D.; Fu, X.H.; Yan, W.; Lin, F.S.; Kuang, P.H.; Luo, Y.; Lin, E.; Hong, X.; Wu, G. Long non-coding RNA BANCR regulates cancer stem cell markers in papillary thyroid cancer via the RAF/MEK/ERK signaling pathway. Oncol. Rep. 2018, 40, 859–866. [Google Scholar] [CrossRef]
- Braicu, C.; Zimta, A.A.; Harangus, A.; Iurca, I.; Irimie, A.; Coza, O.; Berindan-Neagoe, I. The Function of Non-Coding RNAs in Lung Cancer Tumorigenesis. Cancers 2019, 11, 605. [Google Scholar] [CrossRef]
- Braicu, C.; Gulei, D.; Cojocneanu, R.; Raduly, L.; Jurj, A.; Knutsen, E.; Knutsen, E.; Calin, G.A.; Berindan-Neagoe, I. miR-181a/b therapy in lung cancer: Reality or myth? Mol. Oncol. 2019, 13, 9–25. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Iacobas, S.; Haddad, G.G. Heart rhythm genomic fabric in hypoxia. Biochem. Biophys. Res. Commun. 2010, 391, 1769–1774. [Google Scholar] [CrossRef]
- Iacobas, D.A. The Genomic Fabric Perspective on the transcriptome between universal quantifiers and personalized genomic medicine. Biol. Theory 2016, 11, 123–137. [Google Scholar] [CrossRef]
- Gallagher, R.; Collins, S.; Trujillo, J.; McCredie, K.; Ahearn, M.; Tsai, S.; Metzgar, R.; Aulakh, G.; Ting, R.; Ruscetti, F.; et al. Characterization of the continuous, differentiating myeloid cell line (HL-60) from a patient with acute promyelocytic leukemia. Blood 1979, 54, 713–733. [Google Scholar] [PubMed]
- Corso, C.; Ulucan, H.; Parry, E.M.; Parry, J.M. Comparative analysis of two thyroid tumor cell lines by fluorescence in situ hybridization and comparative genomic hybridization. Cancer Genet. Cytogenet. 2002, 137, 108–118. [Google Scholar] [CrossRef]
- Ito, T.; Seyama, T.; Hayashi, Y.; Hayashi, T.; Dohi, K.; Mizuno, T.; Iwamoto, K.; Tsuyama, N.; Nakamura, N.; Akiyama, M. Establishment of 2 human thyroid-carcinoma cell-lines (8305c, 8505c) bearing p53 gene-mutations. Int. J. Oncol. 1994, 4, 583–586. [Google Scholar] [CrossRef] [PubMed]
- Horoszewicz, J.S.; Leong, S.S.; Chu, T.M.; Wajsman, Z.L.; Friedman, M.; Papsidero, L.; Kim, U.; Chai, L.S.; Kakati, S.; Arya, S.K. The LNCaP cell line—A new model for studies on human prostatic carcinoma. Prog. Clin. Biol. Res. 1980, 37, 115–132. [Google Scholar] [PubMed]
- Stone, K.R.; Mickey, D.D.; Wunderli, H.; Mickey, G.H.; Paulson, D.F. Isolation of a human prostate carcinoma cell line (DU 145). Int. J. Cancer 1978, 21, 274–281. [Google Scholar] [CrossRef]
- Kravchick, D.O.; Hrdinka, M.; Iacobas, S.; Iacobas, D.A.; Kreutz, M.R.; Jordan, B.A. Synaptonuclear messenger PRR7 inhibits c-Jun ubiquitination and regulates NMDA mediated excitotoxicity. EMBOJ 2016, 35, 1923–1934. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Iacobas, S.; Urban-Maldonado, M.; Spray, D.C. Sensitivity of the brain transcriptome to connexin ablation. Biochim. Biophys. Acta 2005, 1711, 183–196. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Iacobas, S.; Tanowitz, H.B.; deCarvalho, A.C.; Spray, D.C. Functional genomic fabrics are remodeled in a mouse model of Chagasic cardiomyopathy and restored following cell therapy. Microbes Infect. 2018, 20, 185–195. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Iacobas, S.; Nebieridze, N.; Velisek, L.; Veliskova, J. Estrogen protects neurotransmission transcriptome during status epilepticus. Front Neurosci. 2018, 12. [Google Scholar] [CrossRef]
- Iacobas, D.A.; Iacobas, S.; Spray, D.C. Connexin43 and the brain transcriptome of the newborn mice. Genomics 2007, 89, 113–123. [Google Scholar] [CrossRef][Green Version]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- The Gene Ontology Consortium. Expansion of the Gene Ontology knowledgebase and resources. Nucleic Acids Res. 2017, 45, D331–D338. [Google Scholar] [CrossRef]
- Kanehisa, M.; Furumichi, M.; Tanabe, M.; Sato, Y.; Morishima, K. KEGG: New perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017, 45, D353–D361. [Google Scholar] [CrossRef]
- Huynh, T.; Xu, S. Gene Annotation Easy Viewer (GAEV): Integrating KEGG’s Gene Function Annotations and Associated Molecular Pathways. F1000Research 2018. [Google Scholar] [CrossRef]
- Dou, R.; Zhang, L.; Lu, T.; Liu, D.; Mei, F.; Huang, J.; Qian, L. Identification of a novel HRAS variant and its association with papillary thyroid carcinoma. Oncol. Lett. 2018, 15, 4511–4516. [Google Scholar] [CrossRef]
- Cipriani, N.A.; Agarwal, S.; Dias-Santagata, D.; Faquin, W.C.; Sadow, P.M. Clear Cell Change in Thyroid Carcinoma: A Clinicopathologic and Molecular Study with Identification of Variable Genetic Anomalies. Thyroid 2017, 27, 819–824. [Google Scholar] [CrossRef]
- Liu, Y.; Tong, C.; Cao, J.; Xiong, M. NEMP1 Promotes Tamoxifen Resistance in Breast Cancer Cells. Biochem. Genet. 2019. [Google Scholar] [CrossRef]
- De Antonellis, P.; Carotenuto, M.; Vandenbussche, J.; De Vita, G.; Ferrucci, V.; Medaglia, C.; Boffa, I.; Galiero, A.; Di Somma, S.; Magliulo, D.; et al. Early targets of miR-34a in neuroblastoma. Mol. Cell Proteomics 2014, 13, 2114–2131. [Google Scholar] [CrossRef]
- Pereira, M.S.; de Almeida, G.C.; Pinto, F.; Viana-Pereira, M.; Reis, R.M. SPINT2 Deregulation in Prostate Carcinoma. J. Histochem. Cytochem. 2016, 64, 32–41. [Google Scholar] [CrossRef]
- Liu, F.; Cox, C.D.; Chowdhury, R.; Dovek, L.; Nguyen, H.; Li, T.; Li, S.; Ozer, B.; Chou, A.; Nguyen, N.; et al. SPINT2 is hypermethylated in both IDH1 mutated and wild-type glioblastomas, and exerts tumor suppression via reduction of c-Met activation. J. Neurooncol. 2019, 142, 423–434. [Google Scholar] [CrossRef]
- Wu, W.F.; Maneix, L.; Insunza, J.; Nalvarte, I.; Antonson, P.; Kere, J.; Yu, N.Y.; Tohonen, V.; Katayama, S.; Einarsdottir, E.; et al. Estrogen receptor β, a regulator of androgen receptor signaling in the mouse ventral prostate. Proc. Natl. Acad. Sci. USA 2017, 114, E3816–E3822. [Google Scholar] [CrossRef]
- Wang, X.F.; Chen, J.; Gong, Y.B.; Qin, Y.C.; Wang, L.; Li, N.C. Long non-coding RNA DUXAP10 promotes the proliferation, migration, and inhibits apoptosis of prostate cancer cells. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 3699–3708. [Google Scholar] [CrossRef]
- Bian, Z.; Yu, Y.; Quan, C.; Guan, R.; Jin, Y.; Wu, J.; Xu, L.; Chen, F.; Bai, J.; Sun, W.; et al. RPL13A as a reference gene for normalizing mRNA transcription of ovarian cancer cells with paclitaxel and 10-hydroxycamptothecin treatments. Mol. Med. Rep. 2015, 11, 3188–3194. [Google Scholar] [CrossRef]
- De Thé, H.; Lavau, C.; Marchio, A.; Chomienne, C.; Degos, L.; Dejean, A. The PML-RAR alpha fusion mRNA generated by the t(15;17) translocation in acute promyelocytic leukemia encodes a functionally altered RAR. Cell 1991, 66, 675–684. [Google Scholar]
- Zelent, A.; Guidez, F.; Melnick, A.; Waxman, S.; Licht, J.D. Translocations of the RARalpha gene in acute promyelocytic leukemia. Oncogene 2001, 20, 7186–7203. [Google Scholar] [CrossRef]
- Zia, M.T.K.; Vinukonda, G.; Vose, L.; Bhimavarapu, B.B.R.; Iacobas, S.; Pandey, N.K.; Beall, A.M.; Dohare, P.; LaGamma, E.F.; Iacobas, D.A.; et al. Postnatal glucocorticoid-induced hypomyelination, gliosis, neurologic deficits are dose-dependent, preparation-specific, and reversible. Exp. Neurol. 2014, 263, 200–213. [Google Scholar] [CrossRef]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).