A Novel Software and Method for the Efficient Development of Polymorphic SSR Loci Based on Transcriptome Data
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Collection
2.2. Development of a New Software
2.3. Screening and Verifying of SSR Obtained by PSSRdt
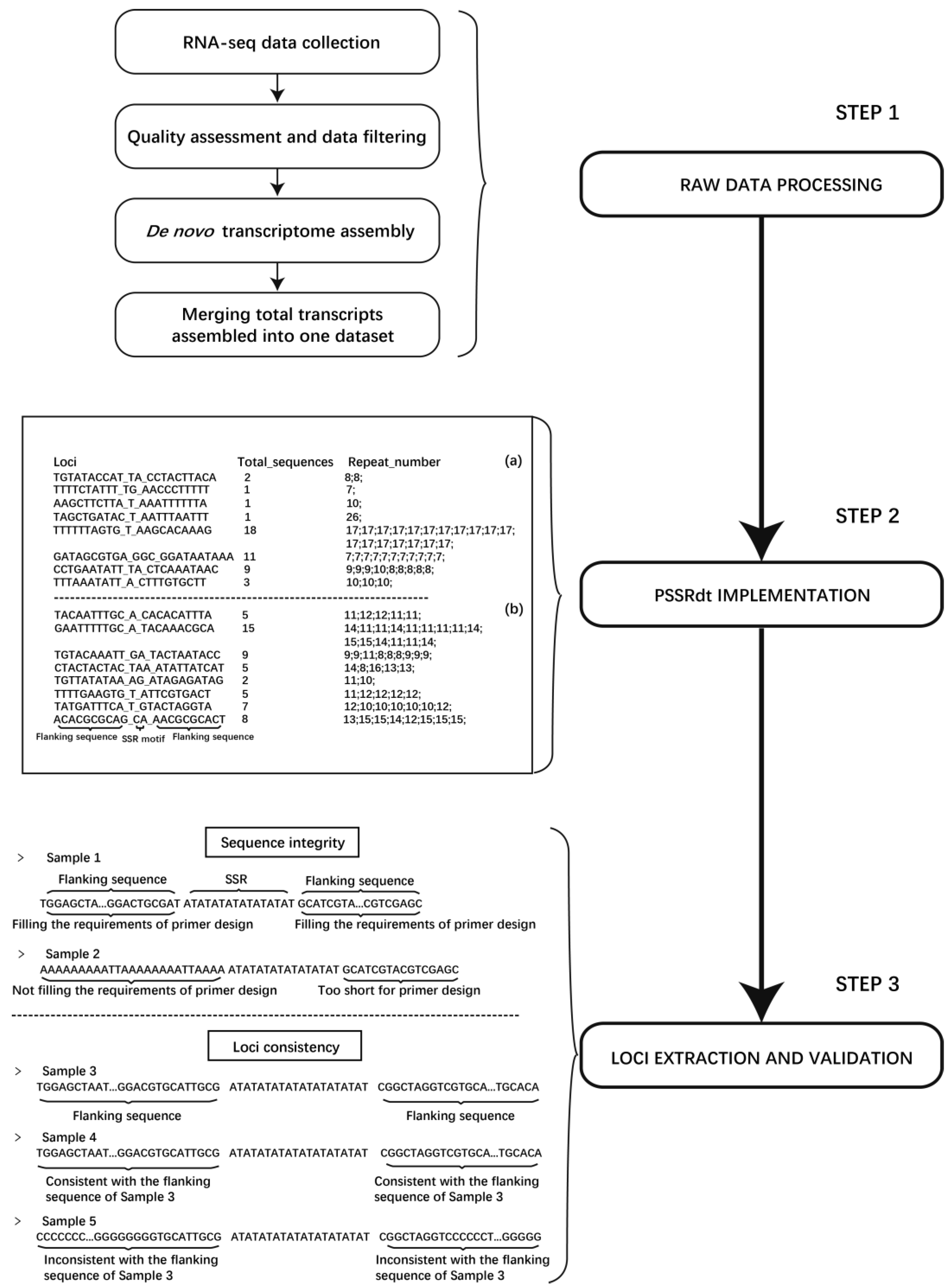
2.4. Overall Flow of the Novel Method
2.5. Experimental Validation
2.5.1. RNA-Seq Data Assembly
2.5.2. Samples and Primers
2.5.3. PCR Amplification and Statistical Analysis
3. Results
3.1. Transcriptome Data Assembly and Microsatellite Detection
3.2. PSSRdt Application
3.3. Primer Design and SSR Loci Validation
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Han, Z.; Ma, X.; Wei, M.; Zhao, T.; Zhan, R.; Chen, W. SSR marker development and intraspecific genetic divergence exploration of Chrysanthemum indicum based on transcriptome analysis. BMC Genom. 2018, 19, 291. [Google Scholar] [CrossRef] [PubMed]
- Ukoskit, K.; Posudsavang, G.; Pongsiripat, N.; Chatwachirawong, P.; Klomsa-Ard, P.; Poomipant, P.; Tragoonrung, S. Detection and validation of EST-SSR markers associated with sugar-related traits in sugarcane using linkage and association mapping. Genomics 2019, 1, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jia, H.; Yang, H.; Sun, P.; Li, J.; Zhang, J.; Guo, Y.; Han, X.; Zhang, G.; Lu, M.; Hu, J. De novo transcriptome assembly, development of EST-SSR markers and population genetic analyses for the desert biomass willow, Salix psammophila. Sci. Rep. 2016, 6, 39591. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Primavera, J.H.; Pena, L.D.D.L.; Pettit, P.; Belak, J.; Alcivar-Warren, A. Genetic diversity of wild and cultured Black Tiger Shrimp (Penaeus monodon) in the Philippines using microsatellites. Aquaculture 2001, 199, 13–40. [Google Scholar] [CrossRef]
- Lázaro-Castellanos, J.O.; Mata-Rosas, M.; González, D.; Arias, S.; Reverchon, F. In Vitro propagation of endangered Mammillaria genus (Cactaceae) species and genetic stability assessment using SSR markers. In Vitr. Cell. Dev. Biol. Plant 2018, 54, 518–529. [Google Scholar] [CrossRef]
- Yue, G.H.; Mei, Y.H.; Orban, L.; Komen, J. Microsatellites within genes and ESTs of common carp and their applicability in silver crucian carp. Aquaculture 2004, 234, 85–98. [Google Scholar] [CrossRef]
- Ding, S.; Wang, S.; Kang, H.; Jiang, M.; Fei, L. Large-scale analysis reveals that the genome features of simple sequence repeats are generally conserved at the family level in insects. BMC Genom. 2017, 18, 848. [Google Scholar] [CrossRef] [PubMed]
- Yadav, R.; Lone, S.A.; Gaikwad, K.; Singh, N.K.; Padaria, J.C. Transcriptome sequence analysis and mining of SSRs in Jhar Ber (Ziziphus nummularia (Burm. f.) Wight & Arn) under drought stress. Sci. Rep. 2018, 8, 2406. [Google Scholar] [PubMed]
- Squirrell, J.; Hollingsworth, P.M.; Woodhead, M.; Russell, J.; Lowe, A.J.; Gibby, M.; Powell, W. How much effort is required to isolate nuclear microsatellites from plants? Mol. Ecol. 2003, 12, 1339–1348. [Google Scholar] [CrossRef] [PubMed]
- Guichoux, E.; Lagache, L.; Wagner, S.; Chaumeil, P.; Léger, P.; Lepais, O.; Lepoittevin, C.; Malausa, T.; Revarel, E.; Salin, F.; et al. Current trends in microsatellite genotyping. Mol. Ecol. Resour. 2011, 11, 591–611. [Google Scholar] [CrossRef] [PubMed]
- Chamberlain, J.S.; Gibbs, R.A.; Ranier, J.E.; Nguyen, P.N.; Caskey, C.T. Deletion screening of the Duchenne muscular dystrophy locus via multiplex DNA amplification. Nucleic Acids Res. 1988, 16, 11141–11156. [Google Scholar] [CrossRef] [PubMed]
- Emese, M.; Nicolas, P.; André, G.; Vincent, D.; Pascal, H.; Aurélie, T.; Rémi, G.; Jean-François, M. QDD version 3.1: A user-friendly computer program for microsatellite selection and primer design revisited: Experimental validation of variables determining genotyping success rate. Mol. Ecol. Resour. 2015, 14, 1302–1313. [Google Scholar]
- Mudunuri, S.B.; Patnana, S.; Nagarajaram, H.A. MICdb3.0: A comprehensive resource of microsatellite repeats from prokaryotic genomes. Database 2014, 2014, bau005. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Faircloth, B. MSATCOMMANDER: Detection of microsatellite repeat arrays and automated, locus-specific primer design. Mol. Ecol. Resour. 2010, 8, 92–94. [Google Scholar] [CrossRef] [PubMed]
- Ponyared, P.; Ponsawat, J.; Tongsima, S.; Seresangtakul, P.; Akkasaeng, C.; Tantisuwichwong, N. ESAP plus: A web-based server for EST-SSR marker development. BMC Genom. 2016, 17, 163–173. [Google Scholar] [CrossRef] [PubMed]
- Christian, S. The evolution of molecular markers—Just a matter of fashion? Nat. Rev. Genet. 2004, 5, 63–69. [Google Scholar]
- Wang, Y.W.; Samuels, T.D.; Wu, Y.Q. Development of 1030 genomic SSR markers in switchgrass. Theor. Appl. Genet. 2011, 122, 677–686. [Google Scholar] [CrossRef] [PubMed]
- Contreras, J.L.; Knoppers, B.M. The genomic commons. Annu. Rev. Genom. Hum. Genet. 2018, 19, 429–453. [Google Scholar] [CrossRef] [PubMed]
- Dutta, S.; Kumawat, G.; Singh, B.P.; Gupta, D.K.; Singh, S.; Dogra, V.; Gaikwad, K.; Sharma, T.R.; Raje, R.S.; Bandhopadhya, T.K.; et al. Development of genic-SSR markers by deep transcriptome sequencing in pigeonpea [Cajanus cajan (L.) Millspaugh]. BMC Plant Biol. 2011, 11, 17. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Liang, S.; Duan, J. De novo assembly and Characterisation of the Transcriptome during seed development, and generation of genic-SSR markers in Peanut (Arachis hypogaea L.). BMC Genom. 2012, 13, 90. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Dong, W.; Lyu, T.; Lyu, Y. An RNA sequencing transcriptome analysis and development of EST-SSR markers in Chinese hawthorn through Illumina sequencing. Forests 2019, 10, 82. [Google Scholar] [CrossRef]
- Liu, C.; Fan, B.; Cao, Z.; Qiuzhu, S.U.; Wang, Y.; Zhang, Z.; Jing, W.U.; Tian, J. A deep sequencing analysis of transcriptomes and the development of EST-SSR markers in mungbean (Vigna radiata). J. Genet. 2016, 95, 527–535. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Liu, X.; Wang, J.W.; Teng, S.Y.; Shi, J.Q.; Li, Y.Y.; Huang, M.R. Transcriptome sequencing and development of novel genic SSR markers for Dendrobium officinale. Mol. Breed. 2017, 37, 18. [Google Scholar] [CrossRef]
- Eujayl, I.; Sorrells, M.E.; Baum, M.; Wolters, P.; Powell, W. Isolation of EST-derived microsatellite markers for genotyping the A and B genomes of wheat. Theor. Appl. Genet. 2002, 104, 399–407. [Google Scholar] [CrossRef] [PubMed]
- Saha, M.C.; Cooper, J.D.; Mian, M.A.R.; Konstantin, C.; May, G.D. Tall fescue genomic SSR markers: Development and transferability across multiple grass species. Theor. Appl. Genet. 2006, 113, 1449–1458. [Google Scholar] [CrossRef] [PubMed]
- Shuxian, L.I.; Yin, T.M. Map and analysis of microsatellites in the genome of Populus: The first sequenced perennial plant. Sci. China 2007, 50, 690. [Google Scholar]
- Dang, M.; Liu, Z.X.; Chen, X.; Zhang, T.; Zhou, H.J.; Hu, Y.H.; Zhao, P. Identification, development, and application of 12 polymorphic EST-SSR markers for an endemic Chinese walnut (Juglans cathayensis L.) using next-generation sequencing technology. Biochem. Syst. Ecol. 2015, 60, 74–80. [Google Scholar] [CrossRef]
- Han, X.; Ling, Q.; Li, C.; Wang, G.; Xu, Z.; Lu, G. Characterization of pikeperch (Sander lucioperca) transcriptome and development of SSR markers. Biochem. Syst. Ecol. 2016, 66, 188–195. [Google Scholar] [CrossRef]
- Li, N.; Zheng, Y.Q.; Ding, H.M.; Li, H.P.; Peng, H.Z.; Jiang, B.; Li, H.B. Development and validation of SSR markers based on transcriptome sequencing of Casuarina equisetifolia. Trees 2017, 32, 1–9. [Google Scholar] [CrossRef]
- Wong, Q.; Tanzi, A.; Ho, W.; Malla, S.; Blythe, M.; Karunaratne, A.; Massawe, F.; Mayes, S. Development of Gene-Based SSR Markers in Winged Bean (Psophocarpus tetragonolobus (L.) DC.) for Diversity Assessment. Genes 2017, 8, 100. [Google Scholar] [CrossRef] [PubMed]
- Wan, F.; Yin, C.; Tang, R.; Chen, M.; Wu, Q.; Huang, C.; Qian, W.; Rota-Stabelli, O.; Yang, N.; Wang, S.; et al. A chromosome-level genome assembly of Cydia pomonella provides insights into chemical ecology and insecticide resistance. Nat. Commun. 2019, 10, 4237. [Google Scholar] [CrossRef] [PubMed]
- Bhangale, T.R.; Stephens, M.; Nickerson, D.A. Automating resequencing-based detection of insertion-deletion polymorphisms. Nat. Genet. 2006, 38, 1457–1462. [Google Scholar] [CrossRef] [PubMed]
- Dotsev, A.V.; Sermyagin, A.A.; Gladyr′, E.A.; Deniskova, T.; Wimmers, K.; Reyer, H.; Brem, G.; Zinovieva, N.A. 163 Population structure and genetic diversity of Russian native cattle breeds. J. Anim. Sci. 2017, 95, 80. [Google Scholar] [CrossRef][Green Version]
- Farhat, M.R.; Freschi, L.; Calderon, R.; Ioerger, T.; Snyder, M.; Meehan, C.J.; de Jong, B.; Rigouts, L.; Sloutsky, A.; Kaur, D.; et al. GWAS for quantitative resistance phenotypes in Mycobacterium tuberculosis reveals resistance genes and regulatory regions. Nat. Commun. 2019, 10, 2128. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Handsaker, B.; Wysoker, A.; Fennell, T.; Ruan, J. The sequence alignment-map format and SAMtools. Bioinformatics 2009, 25, 2078–2079. [Google Scholar] [CrossRef] [PubMed]
- Wang, T.-Y.; Wang, L.; Alam, S.; Hoeppner, L.; Yang, R. ScanNeo: Identifying indel-derived neoantigens using RNA-Seq data. Bioinformatics 2019, 35, 4159–4161. [Google Scholar] [CrossRef] [PubMed]
- Hagiwara, K.; Ding, L.; Edmonson, M.N.; Rice, S.V.; Newman, S.; Meshinchi, S.; Ries, R.E.; Rusch, M.; Zhang, J. RNAIndel: A machine-learning framework for discovery of somatic coding indels using tumor RNA-Seq data. bioRxiv 2019, 10, 512749. [Google Scholar] [CrossRef]
- Burke, G.R.; Moran, N.A. Responses of the pea aphid transcriptome to infection by facultative symbionts. Insect Mol. Biol. 2011, 20, 357–365. [Google Scholar] [CrossRef] [PubMed]
- Richards, S.; Gibbs, R.A.; Gerardo, N.M.; Moran, N.; Nakabachi, A.; Richards, S.; Stern, D.; Tagu, D.; Wilson, A.C.; Muzny, D. Genome sequence of the pea aphid Acyrthosiphon pisum. PLoS Biol. 2015, 8, e1000313. [Google Scholar]
- Hansen, A.K.; Moran, N.A. Aphid genome expression reveals host-symbiont cooperation in the production of amino acids. Proc. Natl. Acad. Sci. USA 2011, 108, 2849–2854. [Google Scholar] [CrossRef] [PubMed]
- Purandare, S.R.; Bickel, R.D.; Jaquiery, J.; Rispe, C.; Brisson, J.A. Accelerated evolution of morph-biased genes in pea aphids. Mol. Biol. Evol. 2015, 31, 2073. [Google Scholar] [CrossRef] [PubMed]
- da Maia, L.C.; Palmieri, D.A.; de Souza, V.Q.; Kopp, M.M.; de Carvalho, F.I.; de Oliveira, A.C. SSR Locator: Tool for simple sequence repeat discovery integrated with primer design and PCR simulation. Int. J. Plant Genom. 2008, 2008, 412696. [Google Scholar] [CrossRef] [PubMed]
- Urquhart, A.; Kimpton, C.P.; Downes, T.J.; Gill, P. Variation in short tandem repeat sequences—A survey of twelve microsatellite loci for use as forensic identification markers. Int. J. Leg. Med. 1994, 107, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Grabherr, M.G.; Haas, B.J.; Moran, Y.; Levin, J.Z.; Thompson, D.A.; Ido, A.; Xian, A.; Lin, F.; Raktima, R.; Qiandong, Z. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat. Biotechnol. 2011, 29, 644. [Google Scholar] [CrossRef] [PubMed]
- Rozen, S.; Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 2000, 132, 365–386. [Google Scholar] [PubMed]
- Schuelke, M. An economic method for the fluorescent labeling of PCR fragments. Nat. Biotechnol. 2000, 18, 233–234. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Muse, S. PowerMarker: An integrated analysis environment for genetic marker analysis. Bioinformatics 2005, 21, 2128–2129. [Google Scholar] [CrossRef] [PubMed]
- Yin, C.; Shen, G.; Guo, D.; Wang, S.; Li, F. InsectBase: A resource for insect genomes and transcriptomes. Nucleic Acids Res. 2016, 44, 801–807. [Google Scholar] [CrossRef] [PubMed]
- Ip, J.C.H.; Mu, H.; Chen, Q.; Sun, J.; Ituarte, S.; Heras, H.; Van Bocxlaer, B.; Ganmanee, M.; Huang, X.; Qiu, J.-W. AmpuBase: A transcriptome database for eight species of apple snails (Gastropoda: Ampullariidae). BMC Genom. 2018, 19, 179. [Google Scholar] [CrossRef] [PubMed]
- Kandpal, R.P.; Kandpal, G.; Weissman, S.M. Construction of libraries enriched for sequence repeats and jumping clones, and hybridization selection for region-specific markers. Proc. Natl. Acad. Sci. USA 1994, 91, 88–92. [Google Scholar] [CrossRef] [PubMed]
- Fisher, P. Single locus microsatellites isolated using 5′ anchored PCR. Nucleic Acids Res. 1996, 24, 4369–4371. [Google Scholar] [CrossRef] [PubMed]
- Mudunuri, S.B.; Nagarajaram, H.A. IMEx: Imperfect microsatellite extractor. Bioinformatics 2007, 23, 1181–1187. [Google Scholar] [CrossRef] [PubMed]
- Alam, C.M.; Singh, A.K.; Sharfuddin, C.; Ali, S. Genome-wide scan for analysis of simple and imperfect microsatellites in diverse carlaviruses. Infect. Genet. Evol. 2014, 21, 287–294. [Google Scholar] [CrossRef] [PubMed]
- Jo, W.-S.; Kim, H.-Y.; Kim, K.-M. Development and characterization of polymorphic EST based SSR markers in barley (Hordeum vulgare). 3 Biotech 2017, 7, 265. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; An, Y.; Li, F.; Li, S.; Wei, C. Genome-wide identification of simple sequence repeats and development of polymorphic SSR markers for genetic studies in tea plant (Camellia sinensis). Mol. Breed. 2018, 38, 59. [Google Scholar] [CrossRef]
- Tiwari, J.K.; Ali, N.; Devi, S.; Zinta, R.; Chakrabarti, S.K. Analysis of allelic variation in wild potato (Solanum) species by simple sequence repeat (SSR) markers. 3 Biotech 2019, 7, 262. [Google Scholar] [CrossRef] [PubMed]
- Vinces, M.D.; Legendre, M.; Caldara, M.; Hagihara, M.; Verstrepen, K.J. Unstable tandem repeats in promoters confer transcriptional evolvability. Science 2009, 324, 1213–1216. [Google Scholar] [CrossRef] [PubMed]

| Accession ID | Total Sequences a | Total Size (bp) | Sequences with SSRs b | Total SSRs | 1 c | 2 | 3 | 4 | 5 | 6 | Submission Institution |
|---|---|---|---|---|---|---|---|---|---|---|---|
| SRR063706 | 1584,7 | 2811,5408 | 7193 | 1499,2 | 9979 | 1860 | 3011 | 102 | 31 | 9 | The University of Arizona |
| SRR063707 | 2667,3 | 9125,155 | 3736 | 5370 | 2947 | 1034 | 1351 | 21 | 14 | 3 | The University of Arizona |
| SRR064408 | 8548 | 3035,495 | 1493 | 2138 | 985 | 508 | 630 | 9 | 4 | 2 | Yale University |
| SRR064409 | 2422,2 | 3274,3820 | 7230 | 1175,3 | 6877 | 1456 | 3337 | 63 | 13 | 7 | Yale University |
| SRR071347 | 9558 | 3108,732 | 1196 | 1653 | 663 | 300 | 680 | 6 | 2 | 2 | Baylor College of Medicine |
| SRR073136 | 1014,5 | 3367,825 | 1481 | 2082 | 1122 | 376 | 574 | 7 | 2 | 1 | University of Nebraska-Lincoln |
| SRR073272 | 1888,5 | 2141,4328 | 5184 | 8737 | 5423 | 1090 | 2153 | 52 | 13 | 6 | University of Nebraska-Lincoln |
| SRR073274 | 7634 | 2076,016 | 407 | 514 | 151 | 73 | 283 | 6 | 1 | 0 | University of Nebraska-Lincoln |
| SRR073276 | 3775,9 | 1020,0134 | 2122 | 2766 | 1143 | 375 | 1223 | 16 | 5 | 4 | University of Nebraska-Lincoln |
| SRR353539 | 2789,8 | 4192,9144 | 1014,8 | 2241,8 | 1363,7 | 2943 | 5646 | 143 | 40 | 9 | University of Nebraska-Lincoln |
| SRR073426 | 4643,7 | 1260,2772 | 2662 | 3487 | 1597 | 452 | 1410 | 19 | 5 | 4 | Cornell university |
| SRR073573 | 2073,0 | 1504,9227 | 3212 | 4541 | 2745 | 593 | 1171 | 18 | 10 | 4 | National Institute for Basic Biology |
| SRR073574 | 2164,6 | 8890,341 | 2478 | 3269 | 1405 | 447 | 1386 | 22 | 5 | 4 | National Institute for Basic Biology |
| SRR073575 | 2001,8 | 2192,9991 | 4421 | 7262 | 3453 | 997 | 2758 | 40 | 9 | 5 | National Institute for Basic Biology |
| SRR073576 | 1979,1 | 1799,6584 | 3611 | 5772 | 2199 | 872 | 2646 | 40 | 10 | 5 | National Institute for Basic Biology |
| SRR073588 | 1668,3 | 2225,0609 | 6410 | 1291,7 | 7607 | 1742 | 3472 | 73 | 20 | 3 | National Institute for Basic Biology |
| SRR074231 | 2336,2 | 4099,6212 | 1021,9 | 2242,0 | 1476,3 | 2745 | 4715 | 148 | 41 | 8 | University of Nebraska-Lincoln |
| SRR074233 | 2133,8 | 2352,6029 | 7417 | 1472,3 | 9237 | 1897 | 3463 | 93 | 28 | 5 | University of Nebraska-Lincoln |
| SRR075802 | 2376,3 | 3008,1669 | 8363 | 1792,7 | 1085,3 | 2391 | 4544 | 108 | 25 | 6 | INRA d |
| SRR075803 | 3108,7 | 3924,7189 | 1047,2 | 2068,1 | 1362,0 | 2470 | 4414 | 127 | 43 | 7 | INRA |
| SRR097896 | 3299,3 | 3999,7626 | 9993 | 1783,0 | 1139,9 | 2139 | 4150 | 97 | 36 | 9 | Centro Nacional de Análisis Genómico |
| SRR098330 | 3110,8 | 3628,8769 | 9981 | 1905,8 | 1276,0 | 2254 | 3898 | 104 | 35 | 7 | Centro Nacional de Análisis Genómico |
| SRR1239439 | 3280,9 | 3742,6415 | 1015,9 | 1920,7 | 1267,7 | 2272 | 4090 | 119 | 40 | 9 | Gene Expression Omnibus |
| SRR1239440 | 2037,3 | 3664,0425 | 7828 | 1561,6 | 8987 | 2073 | 4421 | 103 | 24 | 8 | Gene Expression Omnibus |
| SRR1239441 | 1687,1 | 1151,2399 | 2089 | 2859 | 1654 | 365 | 820 | 15 | 2 | 3 | Gene Expression Omnibus |
| SRR1239442 | 1581,1 | 2480,9531 | 6033 | 1218,1 | 7532 | 1470 | 3084 | 74 | 16 | 5 | Gene Expression Omnibus |
| SRR1239443 | 1571,6 | 2352,5854 | 6212 | 1250,6 | 7943 | 1556 | 2918 | 68 | 18 | 3 | Gene Expression Omnibus |
| SRR1239444 | 1232,7 | 1797,2772 | 4418 | 7931 | 5175 | 913 | 1783 | 45 | 11 | 4 | Gene Expression Omnibus |
| SRR1239445 | 1376,8 | 2219,2206 | 5276 | 9848 | 6445 | 1139 | 2186 | 63 | 10 | 5 | Gene Expression Omnibus |
| SRR1239446 | 6832,1 | 2486,0272 | 7185 | 1092,4 | 6648 | 1428 | 2759 | 68 | 15 | 6 | Gene Expression Omnibus |
| SRR1239448 | 6799,5 | 2526,1149 | 7454 | 1140,1 | 7018 | 1473 | 2821 | 66 | 16 | 7 | Gene Expression Omnibus |
| SRR1239449 | 2097,3 | 2681,8239 | 5932 | 9427 | 5914 | 1066 | 2369 | 58 | 13 | 7 | Gene Expression Omnibus |
| SRR1239450 | 6334,7 | 1765,8260 | 3957 | 5229 | 2464 | 770 | 1954 | 28 | 9 | 4 | Gene Expression Omnibus |
| SRR1239451 | 3222,4 | 8346,763 | 1515 | 1913 | 786 | 256 | 856 | 10 | 3 | 2 | Gene Expression Omnibus |
| SRR1239452 | 2073,0 | 15049,227 | 3212 | 4541 | 2745 | 593 | 1171 | 18 | 10 | 4 | Gene Expression Omnibus |
| SRR1239453 | 2080,9 | 8469,795 | 2382 | 3158 | 1270 | 441 | 1416 | 23 | 4 | 4 | Gene Expression Omnibus |
| SRR1793299 | 2384,4 | 4240,2758 | 1037,1 | 2327,6 | 1495,4 | 2898 | 5216 | 155 | 39 | 14 | Cornell university |
| SRR1793300 | 2042,4 | 3121,8422 | 7938 | 1492,9 | 9771 | 1845 | 3179 | 100 | 27 | 7 | Cornell university |
| SRR924106 | 3001,7 | 3667,7810 | 9772 | 1859,3 | 1207,1 | 2275 | 4082 | 115 | 44 | 6 | INRA |
| SRR924118 | 2568,1 | 3458,2798 | 8421 | 1534,3 | 9796 | 1877 | 3552 | 84 | 26 | 8 | INRA |
| SRR924119 | 2478,0 | 3590,0468 | 8810 | 1695,9 | 1087,9 | 2045 | 3888 | 108 | 31 | 8 | INRA |
| SRR924120 | 1589,6 | 2747,1266 | 5755 | 1011,2 | 6294 | 1202 | 2535 | 63 | 11 | 7 | INRA |
| SRR924121 | 1600,2 | 2609,7341 | 6519 | 1377,2 | 8406 | 1733 | 3511 | 93 | 23 | 6 | INRA |
| SRR924122 | 1445,5 | 2071,9694 | 5758 | 1126,0 | 7336 | 1348 | 2497 | 60 | 16 | 3 | INRA |
| Locus | N | NA | FM | PIC |
|---|---|---|---|---|
| 3 | 12 | 5 | 0.4167 | 0.5748 |
| 4 | 18 | 6 | 0.5278 | 0.6194 |
| 5 | 20 | 7 | 0.4250 | 0.7164 |
| 6 | 18 | 6 | 0.3333 | 0.7444 |
| 7 | 14 | 3 | 0.6786 | 0.4090 |
| 8 | 17 | 9 | 0.2059 | 0.8313 |
| 9 | 21 | 6 | 0.4286 | 0.7006 |
| 10 | 12 | 6 | 0.2917 | 0.7517 |
| 13 | 16 | 9 | 0.2813 | 0.8122 |
| 14 | 18 | 8 | 0.4444 | 0.7118 |
| 15 | 16 | 1 | 1.0000 | 0.0000 |
| 16 | 16 | 7 | 0.4375 | 0.7081 |
| 17 | 15 | 3 | 0.8333 | 0.2604 |
| 18 | 21 | 8 | 0.2381 | 0.8207 |
| 19 | 14 | 5 | 0.4643 | 0.6469 |
| 21 | 16 | 2 | 0.6250 | 0.3589 |
| 22 | 11 | 5 | 0.5000 | 0.6257 |
| 23 | 11 | 6 | 0.3182 | 0.7436 |
| 27 | 17 | 4 | 0.6176 | 0.5239 |
| 29 | 13 | 6 | 0.3462 | 0.6874 |
| 31 | 11 | 11 | 0.2273 | 0.8595 |
| 33 | 16 | 2 | 0.9375 | 0.1103 |
| 34 | 12 | 3 | 0.4583 | 0.5697 |
| 35 | 18 | 4 | 0.4722 | 0.5851 |
| 38 | 17 | 4 | 0.6176 | 0.5269 |
| 39 | 17 | 8 | 0.2647 | 0.7888 |
| 40 | 20 | 1 | 1.0000 | 0.0000 |
| 41 | 10 | 7 | 0.3500 | 0.7700 |
| 43 | 12 | 8 | 0.2917 | 0.8013 |
| 46 | 17 | 5 | 0.3235 | 0.7130 |
| 47 | 16 | 1 | 1.0000 | 0.0000 |
| 48 | 13 | 7 | 0.4231 | 0.6867 |
| 49 | 16 | 2 | 0.8750 | 0.1948 |
| 51 | 14 | 5 | 0.3214 | 0.7248 |
| 52 | 20 | 4 | 0.4500 | 0.5249 |
| 53 | 14 | 6 | 0.3929 | 0.7072 |
| 101 | 15 | 3 | 0.7667 | 0.3227 |
| 102 | 15 | 12 | 0.2000 | 0.8685 |
| 108 | 12 | 7 | 0.2917 | 0.7614 |
| 109 | 16 | 3 | 0.8438 | 0.2478 |
| 110 | 15 | 6 | 0.4333 | 0.6675 |
| 112 | 17 | 9 | 0.2353 | 0.8319 |
| 113 | 15 | 3 | 0.8333 | 0.2710 |
| 114 | 15 | 9 | 0.3667 | 0.7762 |
| 116 | 9 | 3 | 0.7222 | 0.3709 |
| 117 | 9 | 7 | 0.2778 | 0.8053 |
| 119 | 12 | 8 | 0.2500 | 0.7957 |
| 121 | 18 | 5 | 0.2778 | 0.7429 |
| 122 | 17 | 2 | 0.8824 | 0.1861 |
| 128 | 11 | 5 | 0.3182 | 0.7319 |
| 131 | 18 | 5 | 0.3056 | 0.7165 |
| 132 | 18 | 1 | 1.0000 | 0.0000 |
| Mean | 15.2115 | 5.3462 | 0.4966 | 0.5751 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tian, R.; Zhang, C.; Huang, Y.; Guo, X.; Chen, M. A Novel Software and Method for the Efficient Development of Polymorphic SSR Loci Based on Transcriptome Data. Genes 2019, 10, 917. https://doi.org/10.3390/genes10110917
Tian R, Zhang C, Huang Y, Guo X, Chen M. A Novel Software and Method for the Efficient Development of Polymorphic SSR Loci Based on Transcriptome Data. Genes. 2019; 10(11):917. https://doi.org/10.3390/genes10110917
Chicago/Turabian StyleTian, Ruizheng, Cunhuan Zhang, Yixiao Huang, Xin Guo, and Maohua Chen. 2019. "A Novel Software and Method for the Efficient Development of Polymorphic SSR Loci Based on Transcriptome Data" Genes 10, no. 11: 917. https://doi.org/10.3390/genes10110917
APA StyleTian, R., Zhang, C., Huang, Y., Guo, X., & Chen, M. (2019). A Novel Software and Method for the Efficient Development of Polymorphic SSR Loci Based on Transcriptome Data. Genes, 10(11), 917. https://doi.org/10.3390/genes10110917





