Meta-Analysis of Grainyhead-Like Dependent Transcriptional Networks: A Roadmap for Identifying Novel Conserved Genetic Pathways
Abstract
1. Introduction
2. Methods
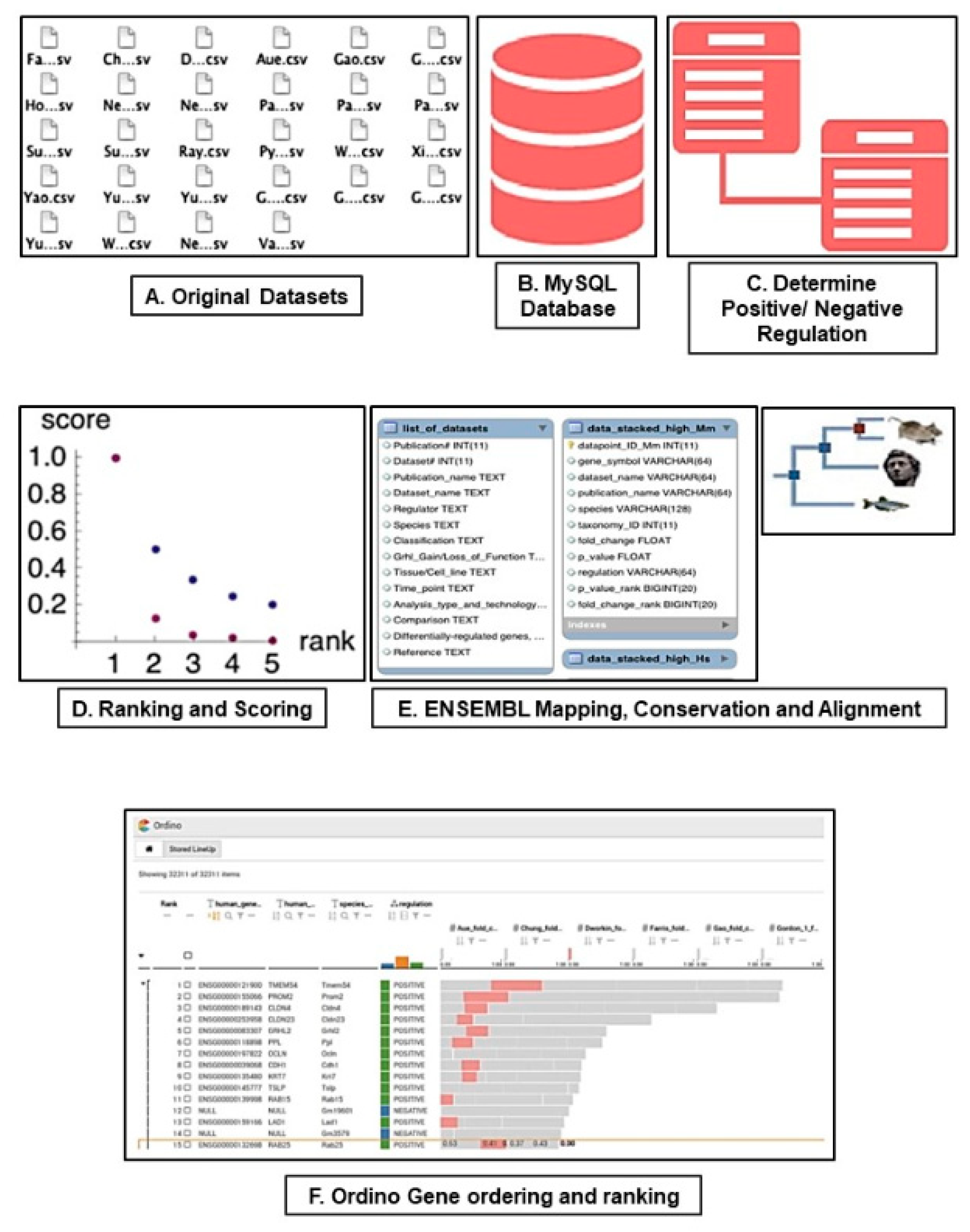
2.1. RNA-SEQ and Microarray Meta-Analysis and Database Construction
2.2. Zebrafish Experimentation
2.3. RNA Extraction and cDNA Synthesis
3. Results
3.1. Meta-Analysis Reveals the Most Consistently Differentially-Regulated Genes Following grh/Grhl Misexpression
3.2. Biological Pathways Regulated by grhl in Epithelia as Identified by GO Analysis
3.3. Grhl-Dependent Target Genes are Well-Conserved between Mouse and Human; Less so between Mouse and Drosophila
3.4. Refining Meta-Analyses to Encompass Only Large-Scale Datasets Generated from Epithelial Tissue or Cancer Cell Lines Reveals grh/Grhl-dependent Transcriptome Specificity
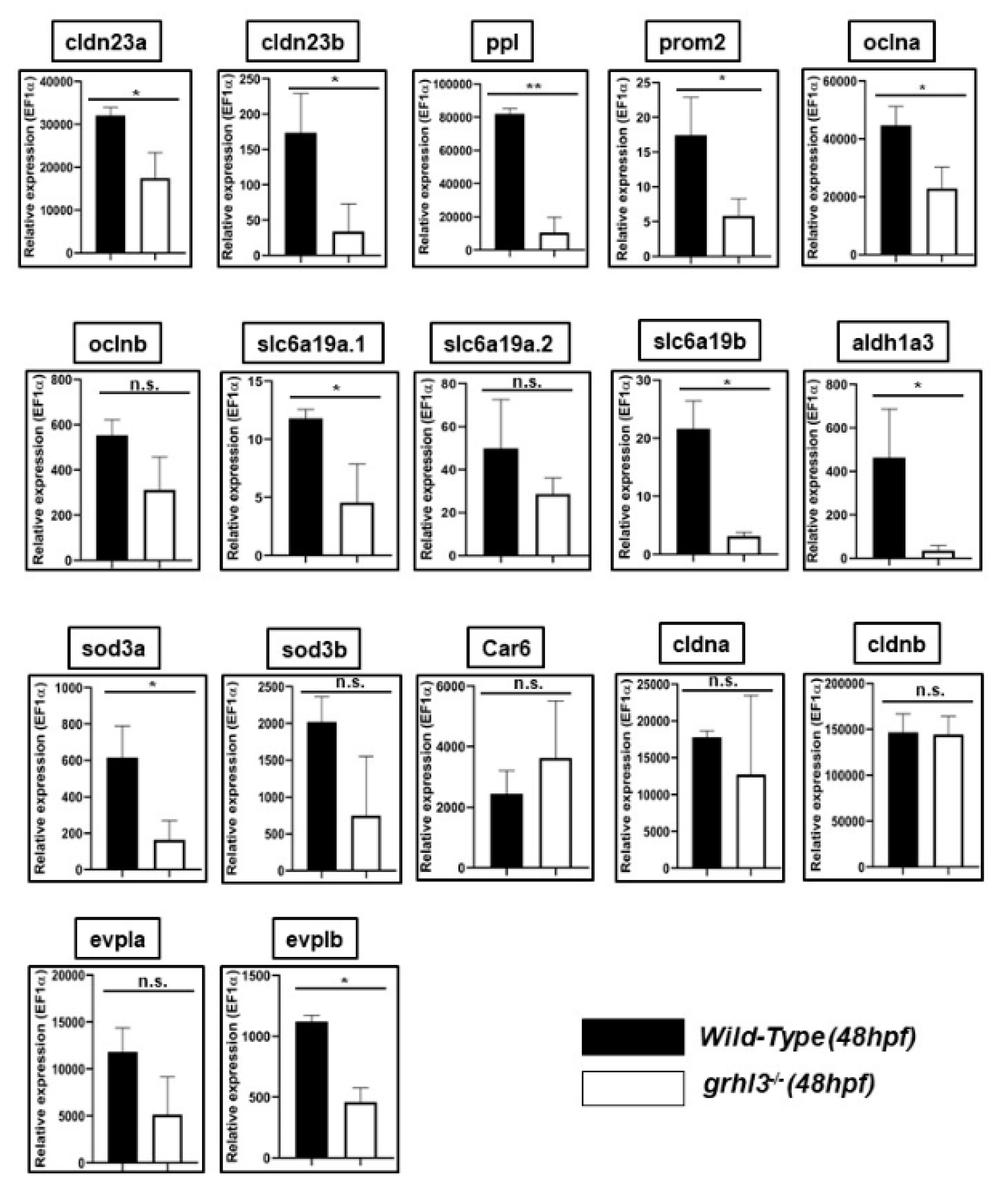
3.5. Expression of Differentially-Regulated Epithelial Genes in a Novel Vertebrate Model
4. Discussion
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bray, S.J.; Kafatos, F.C. Developmental function of Elf-1: An essential transcription factor during embryogenesis in Drosophila. Genes Dev. 1991, 5, 1672–1683. [Google Scholar] [CrossRef] [PubMed]
- Ting, S.B.; Caddy, J.; Hislop, N.; Wilanowski, T.; Auden, A.; Zhao, L.L.; Ellis, S.; Kaur, P.; Uchida, Y.; Holleran, W.M.; et al. A homolog of Drosophila grainy head is essential for epidermal integrity in mice. Science 2005, 308, 411–413. [Google Scholar] [CrossRef] [PubMed]
- Dworkin, S.; Jane, S.M.; Darido, C. The planar cell polarity pathway in vertebrate epidermal development, homeostasis and repair. Organogenesis 2011, 7, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Pare, A.; Kim, M.; Juarez, M.T.; Brody, S.; McGinnis, W. The functions of grainy head-like proteins in animals and fungi and the evolution of apical extracellular barriers. PLoS ONE 2012, 7, e36254. [Google Scholar] [CrossRef]
- Ting, S.B.; Wilanowski, T.; Auden, A.; Hall, M.; Voss, A.K.; Thomas, T.; Parekh, V.; Cunningham, J.M.; Jane, S.M. Inositol- and folate-resistant neural tube defects in mice lacking the epithelial-specific factor Grhl-3. Nat. Med. 2003, 9, 1513–1519. [Google Scholar] [CrossRef]
- Tao, J.; Kuliyev, E.; Wang, X.; Li, X.; Wilanowski, T.; Jane, S.M.; Mead, P.E.; Cunningham, J.M. BMP4-dependent expression of Xenopus Grainyhead-like 1 is essential for epidermal differentiation. Development 2005, 132, 1021–1034. [Google Scholar] [CrossRef]
- Miles, L.B.; Darido, C.; Kaslin, J.; Heath, J.K.; Jane, S.M.; Dworkin, S. Mis-expression of grainyhead-like transcription factors in zebrafish leads to defects in enveloping layer (EVL) integrity, cellular morphogenesis and axial extension. Sci. Rep. 2017, 7, 17607. [Google Scholar] [CrossRef]
- Wang, S.; Samakovlis, C. Grainy head and its target genes in epithelial morphogenesis and wound healing. Curr. Top. Dev. Biol. 2012, 98, 35–63. [Google Scholar]
- Mlacki, M.; Darido, C.; Jane, S.M.; Wilanowski, T. Loss of Grainy head-like 1 is associated with disruption of the epidermal barrier and squamous cell carcinoma of the skin. PLoS ONE 2014, 9, e89247. [Google Scholar] [CrossRef]
- Yu, Z.; Lin, K.K.; Bhandari, A.; Spencer, J.A.; Xu, X.; Wang, N.; Lu, Z.; Gill, G.N.; Roop, D.R.; Wertz, P.; et al. The Grainyhead-like epithelial transactivator Get-1/Grhl3 regulates epidermal terminal differentiation and interacts functionally with LMO4. Dev. Biol. 2006, 299, 122–136. [Google Scholar] [CrossRef]
- Goldie, S.J.; Cottle, D.L.; Tan, F.H.; Roslan, S.; Srivastava, S.; Brady, R.; Partridge, D.D.; Auden, A.; Smyth, I.M.; Jane, S.M.; et al. Loss of GRHL3 leads to TARC/CCL17-mediated keratinocyte proliferation in the epidermis. Cell Death Dis. 2018, 9, 1072. [Google Scholar] [CrossRef]
- Rifat, Y.; Parekh, V.; Wilanowski, T.; Hislop, N.R.; Auden, A.; Ting, S.B.; Cunningham, J.M.; Jane, S.M. Regional neural tube closure defined by the Grainy head-like transcription factors. Dev. Biol. 2010, 345, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Kimura-Yoshida, C.; Mochida, K.; Ellwanger, K.; Niehrs, C.; Matsuo, I. Fate specification of neural plate border by canonical wnt signaling and Grhl3 is crucial for neural tube closure. EBioMedicine 2015, 2, 513–527. [Google Scholar] [CrossRef] [PubMed]
- De Castro, S.C.P.; Gustavsson, P.; Marshall, A.R.; Gordon, W.M.; Galea, G.; Nikolopoulou, E.; Savery, D.; Rolo, A.; Stanier, P.; Andersen, B.; et al. Overexpression of Grainyhead-like 3 causes spina bifida and interacts genetically with mutant alleles of Grhl2 and Vangl2 in mice. Hum. Mol. Genet. 2018, 27, 4218–4230. [Google Scholar] [CrossRef] [PubMed]
- Kousa, Y.A.; Zhu, H.; Fakhouri, W.D.; Lei, Y.; Kinoshita, A.; Roushangar, R.R.; Patel, N.K.; Agopian, A.J.; Yang, W.; Leslie, E.J.; et al. The TFAP2A-IRF6-GRHL3 genetic pathway is conserved in neurulation. Hum. Mol. Genet. 2019, 28, 1726–1737. [Google Scholar] [CrossRef]
- Dworkin, S.; Darido, C.; Georgy, S.R.; Wilanowski, T.; Srivastava, S.; Ellett, F.; Pase, L.; Han, Y.; Meng, A.; Heath, J.K.; et al. Midbrain-hindbrain boundary patterning and morphogenesis are regulated by diverse grainy head-like 2-dependent pathways. Development 2012, 139, 525–536. [Google Scholar] [CrossRef]
- Dworkin, S.; Simkin, J.; Darido, C.; Partridge, D.D.; Georgy, S.R.; Caddy, J.; Wilanowski, T.; Lieschke, G.J.; Doggett, K.; Heath, J.K.; et al. Grainyhead-like 3 regulation of endothelin-1 in the pharyngeal endoderm is critical for growth and development of the craniofacial skeleton. Mech. Dev. 2014, 133, 77–90. [Google Scholar] [CrossRef]
- Goldie, S.J.; Arhatari, B.D.; Anderson, P.; Auden, A.; Partridge, D.D.; Jane, S.M.; Dworkin, S. Mice lacking the conserved transcription factor Grainyhead-like 3 (Grhl3) display increased apposition of the frontal and parietal bones during embryonic development. BMC Dev. Biol. 2016, 16, 37. [Google Scholar] [CrossRef]
- Peyrard-Janvid, M.; Leslie, E.J.; Kousa, Y.A.; Smith, T.L.; Dunnwald, M.; Magnusson, M.; Lentz, B.A.; Unneberg, P.; Fransson, I.; Koillinen, H.K. Dominant mutations in GRHL3 cause Van der Woude Syndrome and disrupt oral periderm development. Am. J. Hum. Genet. 2014, 94, 23–32. [Google Scholar] [CrossRef]
- Mangold, E.; Bohmer, A.C.; Ishorst, N.; Hoebel, A.K.; Gultepe, P.; Schuenke, H.; Klamt, J.; Hofmann, A.; Golz, L.; Raff, R.; et al. Sequencing the GRHL3 Coding Region Reveals Rare Truncating Mutations and a Common Susceptibility Variant for Nonsyndromic Cleft Palate. Am. J. Hum. Genet. 2016, 98, 755–762. [Google Scholar] [CrossRef]
- Lemay, P.; De Marco, P.; Emond, A.; Spiegelman, D.; Dionne-Laporte, A.; Laurent, S.; Merello, E.; Accogli, A.; Rouleau, G.A.; Capra, V.; et al. Rare deleterious variants in GRHL3 are associated with human spina bifida. Hum. Mutat. 2017, 38, 716–724. [Google Scholar] [CrossRef] [PubMed]
- Hosoya, M.; Fujioka, M.; Ogawa, K.; Okano, H. Distinct expression patterns of causative genes responsible for hereditary progressive hearing loss in non-human primate cochlea. Sci. Rep. 2016, 6, 22250. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Mu, Y.; Li, X.; Xu, P.; Tong, J.; Liu, Z.; Ma, T.; Zeng, G.; Yang, S.; Du, J.; et al. Grhl2 deficiency impairs otic development and hearing ability in a zebrafish model of the progressive dominant hearing loss DFNA28. Hum. Mol. Genet. 2011, 20, 3213–3226. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Liu, C.; Zhao, Z.; Gao, N.; Chen, G.; Wang, Y.; Cui, J. Clinical implications of GRHL3 protein expression in breast cancer. Tumour. Biol. 2014, 35, 1827–1831. [Google Scholar] [CrossRef]
- Xiang, J.; Fu, X.; Ran, W.; Chen, X.; Hang, Z.; Mao, H.; Wang, Z. Expression and role of grainyhead-like 2 in gastric cancer. Med. Oncol. 2013, 30, 714. [Google Scholar] [CrossRef]
- Chen, W.; Yi, J.K.; Shimane, T.; Mehrazarin, S.; Lin, Y.-L.; Shin, K.-H.; Kim, R.H.; Park, N.-H.; Kang, M.K. Grainyhead-like 2 regulates epithelial plasticity and stemness in oral cancer cells. Carcinogenesis 2016, 37, 500–510. [Google Scholar] [CrossRef]
- Wang, Y.; Sun, Y.; Huang, Y.; Pan, Y.; Jia, Z.; Ma, L.; Ma, L.; Lan, F.; Zhou, Y.; Shi, J.; et al. Association study between Van der Woude Syndrome causative gene GRHL3 and nonsyndromic cleft lip with or without cleft palate in a Chinese cohort. Gene 2016, 588, 69–73. [Google Scholar] [CrossRef]
- Petrof, G.; Nanda, A.; Howden, J.; Takeichi, T.; McMillan, J.R.; Aristodemou, S.; Ozoemena, L.; Liu, L.; South, A.P.; Pourreyron, C.; et al. Mutations in GRHL2 result in an autosomal-recessive ectodermal Dysplasia syndrome. Am. J. Hum. Genet. 2014, 95, 308–314. [Google Scholar] [CrossRef]
- Brouns, M.R.; De Castro, S.C.; Terwindt-Rouwenhorst, E.A.; Massa, V.; Hekking, J.W.; Hirst, C.S.; Savery, D.; Munts, C.; Partridge, D.; Lamers, W.; et al. Over-expression of Grhl2 causes spina bifida in the Axial defects mutant mouse. Hum. Mol. Genet. 2011, 20, 1536–1546. [Google Scholar] [CrossRef]
- Darido, C.; Georgy, S.R.; Wilanowski, T.; Dworkin, S.; Auden, A.; Zhao, Q.; Rank, G.; Srivastava, S.; Finlay, M.J.; Papenfuss, A.T.; et al. Targeting of the tumor suppressor GRHL3 by a miR-21-dependent proto-oncogenic network results in PTEN loss and tumorigenesis. Cancer Cell 2011, 20, 635–648. [Google Scholar] [CrossRef]
- Gao, X.; Vockley, C.M.; Pauli, F.; Newberry, K.M.; Xue, Y.; Randell, S.H.; Reddy, T.E.; Hogan, B.L. Evidence for multiple roles for grainyheadlike 2 in the establishment and maintenance of human mucociliary airway epithelium. Proc. Natl. Acad. Sci. USA 2013, 110, 9356–9361. [Google Scholar] [CrossRef] [PubMed]
- Hemphala, J.; Uv, A.; Cantera, R.; Bray, S.; Samakovlis, C. Grainy head controls apical membrane growth and tube elongation in response to Branchless/FGF signalling. Development 2003, 130, 249–258. [Google Scholar] [CrossRef] [PubMed]
- Mace, K.A.; Pearson, J.C.; McGinnis, W. An epidermal barrier wound repair pathway in Drosophila is mediated by grainy head. Science 2005, 308, 381–385. [Google Scholar] [CrossRef] [PubMed]
- Hislop, N.R.; Caddy, J.; Ting, S.B.; Auden, A.; Vasudevan, S.; King, S.L.; Lindeman, G.J.; Visvader, J.E.; Cunningham, J.M.; Jane, S.M. Grhl3 and Lmo4 play coordinate roles in epidermal migration. Dev. Biol. 2008, 321, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Cangkrama, M.; Darido, C.; Georgy, S.R.; Partridge, D.; Auden, A.; Srivastava, S.; Wilanowski, T.; Jane, S.M. Two ancient gene families are critical for maintenance of the mammalian skin barrier in postnatal life. J. Investig. Dermatol. 2016, 136, 1438–1448. [Google Scholar] [CrossRef]
- Caddy, J.; Wilanowski, T.; Darido, C.; Dworkin, S.; Ting, S.B.; Zhao, Q.; Rank, G.; Auden, A.; Srivastava, S.; Papenfuss, T.A.; et al. Epidermal wound repair is regulated by the planar cell polarity signaling pathway. Dev. Cell 2010, 19, 138–147. [Google Scholar] [CrossRef]
- Wilanowski, T.; Caddy, J.; Ting, S.B.; Hislop, N.R.; Cerruti, L.; Auden, A.; Zhao, L.L.; Asquith, S.; Ellis, S.; Sinclair, R.; et al. Perturbed desmosomal cadherin expression in grainy head-like 1-null mice. EMBO J. 2008, 27, 886–897. [Google Scholar] [CrossRef]
- Dworkin, S.; Heath, J.K.; de Jong-Curtain, T.A.; Hogan, B.M.; Lieschke, G.J.; Malaterre, J.; Ramsay, R.G.; Mantamadiotis, T. CREB activity modulates neural cell proliferation, midbrain-hindbrain organization and patterning in zebrafish. Dev. Biol. 2007, 307, 127–141. [Google Scholar] [CrossRef]
- Nevil, M.; Bondra, E.R.; Schulz, K.N.; Kaplan, T.; Harrison, M.M. Stable binding of the conserved transcription factor grainy head to its target genes throughout drosophila melanogaster development. Genetics 2017, 205, 605–620. [Google Scholar] [CrossRef]
- Tuckfield, A.; Clouston, D.R.; Wilanowski, T.M.; Zhao, L.L.; Cunningham, J.M.; Jane, S.M. Binding of the RING polycomb proteins to specific target genes in complex with the grainyhead-like family of developmental transcription factors. Mol. Cell. Biol. 2002, 22, 1936–1946. [Google Scholar] [CrossRef]
- Yu, Z.; Mannik, J.; Soto, A.; Lin, K.K.; Andersen, B. The epidermal differentiation-associated Grainyhead gene Get1/Grhl3 also regulates urothelial differentiation. EMBO J. 2009, 28, 1890–1903. [Google Scholar] [CrossRef] [PubMed]
- Ray, J.H.; Niswander, L.A. Grainyhead-like 2 downstream targets act to suppress epithelial-to-mesenchymal transition during neural tube closure. Development 2016, 143, 1192–1204. [Google Scholar] [CrossRef] [PubMed]
- Kersbergen, A.; Best, S.A.; Dworkin, S.; Ah-Cann, C.; de Vries, M.E.; Asselin-Labat, M.L.; Ritchie, M.E.; Jane, S.M.; Sutherland, K.D. Lung morphogenesis is orchestrated through Grainyhead-like 2 (Grhl2) transcriptional programs. Dev. Biol. 2018, 443, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Hopkin, A.S.; Gordon, W.; Klein, R.H.; Espitia, F.; Daily, K.; Zeller, M.; Baldi, P.; Andersen, B. GRHL3/GET1 and trithorax group members collaborate to activate the epidermal progenitor differentiation program. PLoS Genet. 2012, 8, e1002829. [Google Scholar] [CrossRef] [PubMed]
- Gordon, W.M.; Zeller, M.D.; Klein, R.H.; Swindell, W.R.; Ho, H.; Espetia, F.; Gudjonsson, J.E.; Baldi, P.F.; Andersen, B. A GRHL3-regulated repair pathway suppresses immune-mediated epidermal hyperplasia. J. Clin. Investig. 2014, 124, 5205–5218. [Google Scholar] [CrossRef]
- Xiang, X.; Deng, Z.; Zhuang, X.; Ju, S.; Mu, J.; Jiang, H.; Zhang, L.; Yan, J.; Miller, D.; Zhang, H.G. Grhl2 determines the epithelial phenotype of breast cancers and promotes tumor progression. PLoS ONE 2012, 7, e50781. [Google Scholar] [CrossRef]
- Cieply, B.; Riley, P.; Pifer, P.M.; Widmeyer, J.; Addison, J.B.; Ivanov, A.V.; Denvir, J.; Frisch, S.M. Suppression of the Epithelial-Mesenchymal Transition by Grainyhead-Like-2. Cancer Res. 2012, 72, 2440–2453. [Google Scholar] [CrossRef]
- Farris, J.C.; Pifer, P.M.; Zheng, L.; Gottlieb, E.; Denvir, J.; Frisch, S.M. Grainyhead-like 2 reverses the metabolic changes induced by the oncogenic epithelial-mesenchymal transition: Effects on anoikis. Mol. Cancer Res. 2016, 14, 528–538. [Google Scholar] [CrossRef]
- Chung, V.Y.; Tan, T.Z.; Tan, M.; Wong, M.K.; Kuay, K.T.; Yang, Z.; Ye, J.; Muller, J.; Koh, C.M.; Guccione, E.; et al. GRHL2-miR-200-ZEB1 maintains the epithelial status of ovarian cancer through transcriptional regulation and histone modification. Sci. Rep. 2016, 6, 19943. [Google Scholar] [CrossRef]
- Paltoglou, S.; Das, R.; Townley, S.L.; Hickey, T.E.; Tarulli, G.A.; Coutinho, I.; Fernandes, R.; Hanson, A.R.; Denis, I.; Carroll, J.S.; et al. Novel androgen receptor coregulator GRHL2 exerts both oncogenic and antimetastatic functions in prostate cancer. Cancer Res. 2017, 77, 3417–3430. [Google Scholar] [CrossRef]
- Boglev, Y.; Wilanowski, T.; Caddy, J.; Parekh, V.; Auden, A.; Darido, C.; Hislop, N.R.; Cangkrama, M.; Ting, S.B.; Jane, S.M. The unique and cooperative roles of the Grainy head-like transcription factors in epidermal development reflect unexpected target gene specificity. Dev. Biol. 2011, 349, 512–522. [Google Scholar] [CrossRef]
- Pyrgaki, C.; Liu, A.; Niswander, L. Grainyhead-like 2 regulates neural tube closure and adhesion molecule expression during neural fold fusion. Dev. Biol. 2011, 353, 38–49. [Google Scholar] [CrossRef] [PubMed]
- Ingraham, C.R.; Kinoshita, A.; Kondo, S.; Yang, B.; Sajan, S.; Trout, K.J.; Malik, M.I.; Dunnwald, M.; Goudy, S.L.; Lovett, M.; et al. Abnormal skin, limb and craniofacial morphogenesis in mice deficient for interferon regulatory factor 6 (Irf6). Nat. Genet. 2006, 38, 1335–1340. [Google Scholar] [CrossRef] [PubMed]
- Georgy, S.R.; Cangkrama, M.; Srivastava, S.; Partridge, D.; Auden, A.; Dworkin, S.; McLean, C.A.; Jane, S.M.; Darido, C. Identification of a novel proto-oncogenic network in head and neck squamous cell carcinoma. J. Natl. Cancer Inst. 2015, 107, pii: djv152. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.; McGinnis, W. Phosphorylation of Grainy head by ERK is essential for wound-dependent regeneration but not for development of an epidermal barrier. Proc. Natl. Acad. Sci. USA 2011, 108, 650–655. [Google Scholar] [CrossRef] [PubMed]

| Gene | Species | Dataset | Tissue/Cell Line | Classification/Origin | Reference |
| grh | Drosophila melanogaster | 1 | 2–3 hour whole embryos | Other - non-mammalian | Nevil et al., Genetics, 2017 |
| 2 | 11–12 hour whole embryos | Other - non-mammalian | Nevil et al., Genetics, 2017 | ||
| 3 | 15–16 hour whole embryos | Other - non-mammalian | Nevil et al., Genetics, 2017 | ||
| 4 | 13–16 hour whole embryos | Other - non-mammalian | Yao et al., Development, 2017 | ||
| 5 | 13–16 hour whole embryos | Other - non-mammalian | Yao et al., Development, 2017 | ||
| 6 | Stage 16–17 whole embryos | Other - non-mammalian | Yao et al., Development, 2017 | ||
| Grhl | Neurospora crassa | 7 | 48 hour aerial hyphae and conidia | Other - non-mammalian | Pare et al., PLoS ONE, 2012 |
| Grhl2 | Mus musculus | 8 | E9.5 Non-neural ectoderm | Primary Epithelia | Ray et al., Development,2016 |
| 9 | E16.5 Lung epithelium | Primary Epithelia | Kersbergen et al., Dev. Biol., 2018 | ||
| 10 | E16.5 Lung epithelium | Primary Epithelia | Kersbergen et al., Dev. Biol., 2018 | ||
| 11 | E9.5 Cranial tissue | Other - mammalian | Pyrgaki et al., Dev. Biol., 2011 | ||
| 12 | Extraembryonic trophectoderm-derived tissues at E7.5 | Other - mammalian | Walentin et al., Development, 2015 | ||
| 13 | E9.5 Placenta | Other - mammalian | Walentin et al., Development, 2015 | ||
| 14 | E9.5 First pharyngeal arch (PA1) | Other - mammalian | Carpinelli et al., submitted, 2019 | ||
| 15 | Immortalised Mouse Lung Epithelial Cells (MLE15) | Cell line - Non-Cancer | Varma et al., J. Biol. Chem., 2012 | ||
| 16 | NIH3T3 fibroblast cells | Cell line - Non-Cancer | Werner et al., J. Biol. Chem., 2013 | ||
| 17 | IMCD-3 (inner medullary collecting duct) cell line | Cell line - Non-Cancer | Aue et al., J AmSoc Nephrol, 2015 | ||
| Grhl2 | Canis familiaris | 18 | Madin-Darby Canine Kidney (MDCK) cells | Cell line - Non-Cancer | Pifer et al., Mol. Biol. Cell, 2016 |
| 19 | Madin-Darby Canine Kidney (MDCK) cells | Cell line - Non-Cancer | Pifer et al., Mol. Biol. Cell, 2016 | ||
| Grhl2 | Homo Sapiens | 20 | "Early Passage" Primary Normal Human Epidermal Keratinocytes (NHEK) | Primary Epithelia | Chen et al., Cell Death Dis., 2012 |
| 21 | "Late passage" NHEK | Primary Epithelia | Chen et al., Cell Death Dis., 2012 | ||
| 22 | Undifferentiated primary human bronchial epithelial cells | Primary Epithelia | Gao et al., PNAS, 2014 | ||
| 23 | 4T1 breast tumour cells | Cell line - Cancer | Xiang et al., PLOS ONE, 2012 | ||
| 24 | HMLE-Twist-ER cells | Cell line - Cancer | Cieply et al., Cancer Res., 2012 | ||
| 25 | MSP (mesenchymal sub-population) cells obtained from HMLE cells | Cell line - Cancer | Farris et al., Mol Cancer Res. 2016 | ||
| 26 | OVCA429 (ovarian cystadenocarcinoma; intermediate epithelial [IE] phenotype) | Cell line - Cancer | Chung et al., Scientific Reports, 2016 | ||
| 27 | LNCaP (human prostate carcinoma cells) | Cell line - Cancer | Paltoglou et al., Cancer Res. 2017 | ||
| Gene | Species | Dataset | Tissue/Cell line | Classification/Origin | Reference |
| Grhl3 | Mus musculus | 28 | Embryonic day 18.5 (E18.5) embryo backskin | Primary Epithelia | Yu et al., Dev. Biol., 2006 |
| 29 | E14.5 Bladder | Primary Epithelia | Yu et al., EMBO J, 2009 | ||
| 30 | E16.5 Bladder | Primary Epithelia | Yu et al., EMBO J, 2009 | ||
| 31 | E18.5 Bladder | Primary Epithelia | Yu et al., EMBO J, 2009 | ||
| 32 | E14.5 Skin | Primary Epithelia | Gordon et al., J Clin. Invest., 2014 | ||
| 33 | E16.5 Skin | Primary Epithelia | Gordon et al., J Clin. Invest., 2014 | ||
| 34 | E18.5 Skin | Primary Epithelia | Gordon et al., J Clin. Invest., 2014 | ||
| 35 | Adult Skin | Primary Epithelia | Gordon et al., J Clin. Invest., 2014 | ||
| 36 | Adult Skin | Primary Epithelia | Gordon et al., J Clin. Invest., 2014 | ||
| 37 | Adult Skin | Primary Epithelia | Gordon et al., J Clin. Invest., 2014 | ||
| Grhl3 | Homo sapiens | 38 | NHEK (Neonatal Human Epidermal Keratinocytes) cells | Primary Epithelia | Hopkin et al., PLoS Genet. 2012 |
| 39 | NHEKs and adult psoriasis skin | Primary Epithelia | Gordon et al., J Clin. Invest., 2014 | ||
| Grhl3-Isoform 1 | Homo Sapiens | 40 | Human Embryonic Kidney neuroepithelial cell line (HEK-293) | Cell line - Non-Cancer | Haendeler et al., Arter. Thromb. Vasc. Biol., 2013 |
| Grhl3-Isoform 3 | Homo Sapiens | 41 | HEK-293 | Cell line - Non-Cancer | Haendeler et al., Arter. Thromb. Vasc. Biol., 2013 |
| GRHL Targets | |||||
|---|---|---|---|---|---|
| Rank | Gene | Regulation | Normalised Score | Known relationships with Grhl Transcription Factors | |
| Locus bound by GRHL (ChIP and ChIP-SEQ) | Experimental validation | ||||
| 1 | Tmem54 | POSITIVE | 0.184 | Aue, et al. 2015,Walentin et al. 2015 | |
| 2 | Prom2 | POSITIVE | 0.182 | Aue, et al. 2015,Walentin et al. 2015 | |
| 3 | Cldn4 | POSITIVE | 0.148 | Chung, et al. 2016, Werth et al., 2010. Varma et al., 2012 | Senga, et al. 2012, Varma et al. 2012, Tanimizu and Mitaka 2013, Aue et al. 2015 |
| 4 | Cldn23 | POSITIVE | 0.113 | ||
| 5 | Grhl2 | POSITIVE | 0.089 | ||
| 6 | Ppl | POSITIVE | 0.0868 | Aue, Hinze et al. 2015 | |
| 7 | Ocln | POSITIVE | 0.0777 | ||
| 8 | Cdh1 | POSITIVE | 0.0754 | Aue, et al. 2015, Chung, et al. 2016, Werth et al. 2010. Varma et al., 2012 | Werth, et al. 2010. Pyrgaki, et al. 2011, Xiang, et al. 2012, Gao, et al. 2013, Tanimizu and Mitaka 2013, Chung, et al. 2016, Nishino, Takano et al. 2017, Pan, et al. 2017 |
| 9 | Krt7 | POSITIVE | 0.0752 | ||
| 10 | Tslp | POSITIVE | 0.0741 | ||
| 11 | Rab15 | POSITIVE | 0.0708 | Walentin et al. 2015 | |
| 12 | Gm19601 | NEGATIVE | 0.069 | ||
| 13 | Lad1 | POSITIVE | 0.0661 | Walentin et al. 2015 | |
| 14 | Gm3579 | NEGATIVE | 0.065 | ||
| 15 | Rab25 | POSITIVE | 0.0631 | Aue, et al. 2015 | Senga, et al. 2012 |
| 16 | Epcam | POSITIVE | 0.0618 | Chung, et al. 2016 | Xiang, et al. 2012 |
| 17 | Snx31 | POSITIVE | 0.0607 | ||
| 18 | Slc6a19 | NEGATIVE | 0.0579 | ||
| 19 | Zasp66 | NEGATIVE | 0.0578 | ||
| 20 | Ap1m2 | POSITIVE | 0.0576 | ||
| 21 | Sfn | POSITIVE | 0.0565 | ||
| 22 | IL36RN | POSITIVE | 0.0562 | ||
| 23 | 2310001H18Rik | POSITIVE | 0.0562 | ||
| 24 | Il17re | POSITIVE | 0.0558 | ||
| 25 | Macc1 | POSITIVE | 0.0558 | ||
| 26 | Evpl | POSITIVE | 0.055 | ||
| 27 | Gm20305 | NEGATIVE | 0.0547 | ||
| 28 | Aldh1a3 | POSITIVE | 0.0538 | ||
| 29 | Tacstd2 | POSITIVE | 0.0531 | Chung, et al. 2016 | |
| 30 | Sprr1a | POSITIVE | 0.0528 | ||
| 31 | St14 | POSITIVE | 0.0525 | Chung, et al. 2016 | |
| 32 | Rbbp8 | POSITIVE | 0.0521 | ||
| 33 | Tmprss11b | POSITIVE | 0.0519 | ||
| 34 | Cldn7 | POSITIVE | 0.0515 | ||
| 35 | CG10345 | NEGATIVE | 0.0501 | ||
| 36 | Sod3 | NEGATIVE | 0.05 | ||
| 37 | CG12480 | NEGATIVE | 0.0499 | ||
| 38 | Esrp1 | POSITIVE | 0.0499 | Chung, et al. 2016 | Xiang, et al. 2012 |
| 39 | Prss22 | POSITIVE | 0.0497 | Aue, et al. 2015 | |
| 40 | Cyp2b19 | NEGATIVE | 0.0497 | ||
| 41 | Car6 | NEGATIVE | 0.0497 | ||
| 42 | Car2 | POSITIVE | 0.0495 | ||
| 43 | Fmo2 | NEGATIVE | 0.0493 | ||
| 44 | Elovl1 | NEGATIVE | 0.0492 | ||
| 45 | Elovl7 | NEGATIVE | 0.0492 | ||
| 46 | Defb3 | POSITIVE | 0.0488 | ||
| 47 | Spint1 | POSITIVE | 0.0481 | Chung, et al. 2016, Walentin et al. 2015 | Walentin, et al. 2015, Matsushita, et al. 2018 |
| 48 | Prss8 | POSITIVE | 0.0481 | Chung, et al. 2016 | |
| 49 | Cldn6 | POSITIVE | 0.0481 | ||
| 50 | Gsta2 | NEGATIVE | 0.0479 | ||
| Source | Term Name | Term ID | n of Term Genes | n of Query Genes | n of Common Genes | Corrected p Value |
|---|---|---|---|---|---|---|
| Gene Ontology (Biological Process) | ||||||
| BP | biological adhesion | GO:0022610 | 1351 | 80 | 18 | 1.69 × 10−2 |
| BP | cell adhesion | GO:007155 | 1343 | 80 | 18 | 1.56 × 10−2 |
| BP | cell-cell adhesion | GO:0098609 | 776 | 79 | 14 | 6.29 × 10−3 |
| BP | cell-cell adhesion via plasma membrane adhesion molecules | GO:0098742 | 245 | 79 | 10 | 1.43 × 10−4 |
| BP | calcium-independent cell-cell adhesion via plasma membrane adhesion molecules | GO:0016338 | 23 | 79 | 6 | 9.48 × 10−7 |
| BP | developmental process | GO:0032502 | 6212 | 78 | 45 | 3.91 × 10−3 |
| BP | anatomical structure development | GO:0048856 | 7793 | 78 | 44 | 1.43 × 10−3 |
| BP | tissue development | GO:0009888 | 1926 | 66 | 27 | 6.12 × 10−8 |
| BP | epithelium development | GO:0060429 | 1218 | 77 | 26 | 8.22 × 10−10 |
| BP | epidermis development | GO:0008544 | 459 | 77 | 19 | 2.62 × 10−11 |
| BP | multicellular organismal process | GO:0032501 | 7414 | 74 | 48 | 4.19 × 10−3 |
| BP | multicellular organism development | GO:0007275 | 5321 | 78 | 41 | 3.72 × 10−3 |
| BP | system development | GO:0048731 | 4760 | 78 | 40 | 5.09 × 10−4 |
| BP | animal organ development | GO:0048513 | 3428 | 77 | 35 | 2.65 × 10−5 |
| BP | skin development | GO:0043588 | 412 | 77 | 21 | 1.18 × 10−14 |
| BP | water homeostasis | GO:0030104 | 69 | 77 | 5 | 2.14 × 10−2 |
| BP | epithelial cell differentiation | GO:0030855 | 759 | 77 | 19 | 1.83 × 10−7 |
| BP | epithelial cell differentiation involved in embryonic placenta development | GO:0060671 | 3 | 25 | 2 | 1.39 × 10−2 |
| BP | epidermal cell differentiation | GO:0009913 | 353 | 77 | 16 | 1.13 × 10−9 |
| BP | keratinocyte differentiation | GO:0030216 | 299 | 77 | 16 | 8.72 × 10−11 |
| BP | keratinization | GO:0031424 | 224 | 77 | 15 | 2.25 × 10−11 |
| BP | cornification | GO:0070268 | 111 | 73 | 11 | 8.16 × 10−10 |
| BP | multicellular organismal water homeostasis | GO:0050891 | 64 | 77 | 5 | 1.48 × 10−2 |
| BP | regulation of water loss via skin | GO:0033561 | 22 | 77 | 5 | 5.79 × 10−5 |
| BP | establishment of skin barrier | GO:0061436 | 20 | 77 | 5 | 3.43 × 10−5 |
| BP | epithelial cell morphogenesis | GO:0003382 | 32 | 25 | 3 | 2.65 × 10−2 |
| BP | embryonic placenta morphogenesis | GO:0060669 | 25 | 31 | 3 | 2.17 × 10−2 |
| BP | labyrinthine layer morphogenesis | GO:0060713 | 21 | 31 | 3 | 1.26 × 10−2 |
| BP | branching involved in labyrinthine layer morphogenesis | GO:0060670 | 12 | 31 | 3 | 2.11 × 10−3 |
| BP | epithelial cell morphogenesis involved in placental branching | GO:0060672 | 3 | 25 | 2 | 1.39 × 10−2 |
| BP | peptide cross-linking | GO:0018149 | 59 | 64 | 5 | 3.86 × 10−3 |
| Grhl Targets - Mus Musculus | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Rank | Mus musculus Gene Symbol | Regulation | Average Normalised Score | Significant Differential Regulation in Drosophila | Significant Differential Regulation in in Homo sapiens | ||||||
| Orthologue | Rank | Regulation | Average Normalised Score | Orthologue | Rank | Regulation | Average Normalised Score | ||||
| 1 | Prom2 | Positive | 0.163 | prominin-like | PROM2 | 361 | Positive | 0.0116 | |||
| 2 | Tmem54 | Positive | 0.154 | No known orthologue | TMEM54 | 1062 | Positive | 0.0178 | |||
| 3 | Cldn4 | Positive | 0.116 | No known orthologue | CLDN4 | 28 | Positive | 0.0264 | |||
| 4 | Cldn23 | Positive | 0.106 | No known orthologue | CLDN23 | ||||||
| 5 | Ppl | Positive | 0.0812 | shot | PPL | 110 | Positive | 0.0187 | |||
| 6 | Grhl2 | Positive | 0.0752 | grh | 577 | Positive | 0.0323 | GRHL2 | 159 | Positive | 0.0165 |
| 7 | Rab15 | Positive | 0.0663 | RabX4 | RAB15 | ||||||
| 8 | Ocln | Positive | 0.0655 | Su(Tpl) | OCLN | 1141 | Positive | 0.00717 | |||
| 9 | Gm19601 | Negative | 0.0645 | No known orthologue | POLR2K * | ||||||
| 10 | Tslp | Positive | 0.064 | No known orthologue | TSLP | 1957 | Positive | 0.00534 | |||
| 11 | Gm3579 | Negative | 0.0608 | No known orthologue | No known orthologue | ||||||
| 12 | Cdh1 | Positive | 0.0592 | CadN2 | CDH1 | 138 | Positive | 0.0177 | |||
| 13 | Snx31 | Positive | 0.0568 | CG5734 | SNX31 | ||||||
| 14 | Slc6a19 | Negative | 0.0542 | CG43066 | SLC6A19 | ||||||
| 15 | 2310001H18Rik | Positive | 0.0526 | No known orthologue | No known orthologue | ||||||
| 16 | Il17re | Positive | 0.0522 | No known orthologue | IL17RE | ||||||
| 17 | Evpl | Positive | 0.0514 | shot | EVPL | 2179 | Positive | 0.00501 | |||
| 18 | Gm20305 | Negative | 0.0512 | No known orthologue | No known orthologue | ||||||
| 19 | Lad1 | Positive | 0.0508 | Nuak1 | LAD1 | 412 | Positive | 0.0102 | |||
| 20 | Aldh1a3 | Positive | 0.0503 | CG31075 | ALDH1A3 | 1528 | Positive | 0.00614 | |||
| 21 | Sprr1a | Positive | 0.0494 | CG17377 | SPR1A | ||||||
| 22 | Rbbp8 | Positive | 0.0487 | No known orthologue | RBBP8NL | ||||||
| 23 | Tmprss11b | Positive | 0.0485 | CG11836 | TMPRSS2 | ||||||
| 24 | Sod3 | Negative | 0.0468 | CG5948 | SOD3 | ||||||
| 25 | Cyp2b19 | Negative | 0.0465 | Cyp18a1 | CYP2B6 | ||||||
| 26 | Car6 | Negative | 0.0464 | CAH7 | 582 | Positive | 0.0323 | CA6 | |||
| 27 | Car2 | Positive | 0.0463 | CAH1 | 590 | Positive | 0.0323 | CA2 | |||
| 28 | Fmo2 | Negative | 0.0461 | Fmo-2 | FMO2 | ||||||
| 29 | Defb3 | Positive | 0.0456 | No known orthologue | HBETAD3 | ||||||
| 30 | Rab25 | Positive | 0.0453 | Rab11 | RAB25 | 241 | Positive | 0.0137 | |||
| 31 | Prss8 | Positive | 0.045 | CG16749 | PRSS8 | ||||||
| 32 | Cldn6 | Positive | 0.045 | No known orthologue | CLDN6 | ||||||
| 33 | Gsta2 | Negative | 0.0448 | GstS1 | GSTA1/2/3/5 | ||||||
| 34 | Aox4 | Negative | 0.0446 | ry | AOX1 | ||||||
| 35 | Tmprss13 | Positive | 0.0441 | Sb | TMPRSS13 | 5 | Positive | 0.0346 | |||
| 36 | Smpdl3b | Positive | 0.0437 | CG32052 | SMPDL3B | ||||||
| 37 | Ap1m2 | Positive | 0.0434 | AP-1mu | AP1M2 | 464 | Positive | 0.0105 | |||
| 38 | Pax8 | Negative | 0.0433 | sv | PAX8 | ||||||
| 39 | Klk6 | Positive | 0.0433 | CG12951 | KLK6 | 176 | Positive | 0.0157 | |||
| 40 | Il33 | Positive | 0.0425 | No known orthologue | IL33 | ||||||
| 41 | Fabp5 | Positive | 0.0424 | fabp | FABP5 | ||||||
| 42 | Sfn | Positive | 0.0419 | 14-3-3zeta | SFN | 418 | Positive | 0.0109 | |||
| 43 | Cldn1 | Positive | 0.0417 | No known orthologue | CLDN1 | 442 | Negative | 0.0108 | |||
| 44 | Slpi | Positive | 0.041 | CG5639 | SLPI | ||||||
| 45 | Unc93a | Negative | 0.0409 | CG4928 | 1219 | Negative | 0.041 | UNC93A | |||
| 46 | Krt6b | Positive | 0.0408 | LamC | KRT6B | ||||||
| 47 | Macc1 | Positive | 0.0399 | No known orthologue | MACC1 | ||||||
| 48 | Rptn | Positive | 0.0393 | No known orthologue | RPTN | 79 | Positive | 0.0203 | |||
| 49 | Gsta1 | Negative | 0.039 | No known orthologue | GSTA1 | ||||||
| 50 | 9930013L23Rik | Positive | 0.0387 | No known orthologue | CEMIP | ||||||
| Rank | Gene | Regulation | Normalised Score |
|---|---|---|---|
| 1 | Cldn23 | POSITIVE | 0.0954 |
| 2 | Ppl | POSITIVE | 0.0693 |
| 3 | Prom2 | POSITIVE | 0.0689 |
| 4 | Gm19601 | NEGATIVE | 0.0689 |
| 5 | Tslp | POSITIVE | 0.0684 |
| 6 | Gm3579 | NEGATIVE | 0.065 |
| 7 | Ocln | POSITIVE | 0.0633 |
| 8 | Slc6a19 | NEGATIVE | 0.0579 |
| 9 | 2310001H18Rik | POSITIVE | 0.0562 |
| 10 | Gm20305 | NEGATIVE | 0.0547 |
| 11 | Aldh1a3 | POSITIVE | 0.0538 |
| 12 | Tmprss11bnl | POSITIVE | 0.0519 |
| 13 | Snx31 | POSITIVE | 0.0517 |
| 14 | Sod3 | NEGATIVE | 0.05 |
| 15 | Cyp2b19 | NEGATIVE | 0.0496 |
| 16 | Car6 | NEGATIVE | 0.0496 |
| 17 | Cldn4 | POSITIVE | 0.0494 |
| 18 | Fmo2 | NEGATIVE | 0.0493 |
| 19 | Defb3 | POSITIVE | 0.0488 |
| 20 | Gsta2 | NEGATIVE | 0.0479 |
| 21 | Aox4 | NEGATIVE | 0.0476 |
| 22 | Evpl | POSITIVE | 0.047 |
| 23 | Klk6 | POSITIVE | 0.0462 |
| 24 | Il33 | POSITIVE | 0.0455 |
| 25 | Cldn1 | POSITIVE | 0.0446 |
| 26 | Slpi | POSITIVE | 0.106 |
| 27 | Unc93a | NEGATIVE | 0.106 |
| 28 | Krt6b | POSITIVE | 0.105 |
| 29 | Sprr1a | POSITIVE | 0.105 |
| 30 | Rptn | POSITIVE | 0.102 |
| 31 | Gsta1 | NEGATIVE | 0.101 |
| 32 | Cemip | POSITIVE | 0.0999 |
| 33 | Fabp7 | POSITIVE | 0.0991 |
| 34 | Adh6a | NEGATIVE | 0.0985 |
| 35 | Wfdc12 | POSITIVE | 0.0961 |
| 36 | Fxyd4 | POSITIVE | 0.096 |
| 37 | Krt78 | NEGATIVE | 0.0956 |
| 38 | Tmem54 | POSITIVE | 0.0953 |
| 39 | Gldc | NEGATIVE | 0.0951 |
| 40 | Gdpd1 | NEGATIVE | 0.0945 |
| 41 | Slurp1 | POSITIVE | 0.0939 |
| 42 | Hyal4 | NEGATIVE | 0.0934 |
| 43 | Lce3f | POSITIVE | 0.0933 |
| 44 | Unc93a2 | NEGATIVE | 0.0932 |
| 45 | Sptlc3 | POSITIVE | 0.0926 |
| 46 | Pglyrp1 | NEGATIVE | 0.0924 |
| 47 | Gm10639 | NEGATIVE | 0.0915 |
| 48 | Gdpd3 | NEGATIVE | 0.0912 |
| 49 | Cldnb8 | POSITIVE | 0.0912 |
| 50 | Tmprss4 | POSITIVE | 0.0894 |
| Rank | Gene | Regulation | Normalised Score |
|---|---|---|---|
| 1 | ALOX15 | POSITIVE | 0.045 |
| 2 | CCDC88A | NEGATIVE | 0.0405 |
| 3 | FAT4 | NEGATIVE | 0.0345 |
| 4 | EPCAM | POSITIVE | 0.0345 |
| 5 | IL36RN | POSITIVE | 0.0345 |
| 6 | AHSG | NEGATIVE | 0.0345 |
| 7 | Krt7 | POSITIVE | 0.0345 |
| 8 | Sele | NEGATIVE | 0.0345 |
| 9 | AIM1 | POSITIVE | 0.0285 |
| 10 | CD24 | POSITIVE | 0.0277 |
| 11 | ALOXE3 | POSITIVE | 0.0274 |
| 12 | SOD1 | NEGATIVE | 0.0274 |
| 13 | ITGB6 | POSITIVE | 0.0274 |
| 14 | SERPINE1 | NEGATIVE | 0.0274 |
| 15 | NOX5 | POSITIVE | 0.0274 |
| 16 | ALB | NEGATIVE | 0.0274 |
| 17 | Epcam | POSITIVE | 0.0274 |
| 18 | Ptges | NEGATIVE | 0.0274 |
| 19 | KRT19 | POSITIVE | 0.0263 |
| 20 | TYMS | NEGATIVE | 0.0248 |
| 21 | ALOX12P2 | POSITIVE | 0.0239 |
| 22 | Cat | NEGATIVE | 0.0239 |
| 23 | FGF2 | NEGATIVE | 0.0239 |
| 24 | CRISP3 | POSITIVE | 0.0239 |
| 25 | SYN3 | NEGATIVE | 0.0239 |
| 26 | Cldn4 | POSITIVE | 0.0239 |
| 27 | Otop1 | NEGATIVE | 0.0239 |
| 28 | SLC1A6 | POSITIVE | 0.0217 |
| 29 | NQO1 | NEGATIVE | 0.0217 |
| 30 | TMPRSS13 | POSITIVE | 0.0217 |
| 31 | ATOH8 | NEGATIVE | 0.0217 |
| 32 | Cldn7 | POSITIVE | 0.0217 |
| 33 | Xdh | NEGATIVE | 0.0217 |
| 34 | NCF2 (p67-PHOX) | POSITIVE | 0.0202 |
| 35 | GLUD1 | NEGATIVE | 0.0202 |
| 36 | ITGA2 | POSITIVE | 0.0202 |
| 37 | SMARCA1 | NEGATIVE | 0.0202 |
| 38 | ALDH3B2 | POSITIVE | 0.0202 |
| 39 | BRSK1 | NEGATIVE | 0.0202 |
| 40 | Lamc2 | POSITIVE | 0.0202 |
| 41 | Olfr1372-ps1 | NEGATIVE | 0.0202 |
| 42 | SOD2 | POSITIVE | 0.019 |
| 43 | ITGB8 | POSITIVE | 0.019 |
| 44 | CDH2 | NEGATIVE | 0.019 |
| 45 | A2ML1 | POSITIVE | 0.019 |
| 46 | TMEM45A | NEGATIVE | 0.019 |
| 47 | Tmem54 | POSITIVE | 0.019 |
| 48 | Dnahc6 | NEGATIVE | 0.019 |
| 49 | CDH1 | POSITIVE | 0.0189 |
| 50 | GALNT3 | POSITIVE | 0.0188 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mathiyalagan, N.; Miles, L.B.; Anderson, P.J.; Wilanowski, T.; Grills, B.L.; McDonald, S.J.; Keightley, M.C.; Charzynska, A.; Dabrowski, M.; Dworkin, S. Meta-Analysis of Grainyhead-Like Dependent Transcriptional Networks: A Roadmap for Identifying Novel Conserved Genetic Pathways. Genes 2019, 10, 876. https://doi.org/10.3390/genes10110876
Mathiyalagan N, Miles LB, Anderson PJ, Wilanowski T, Grills BL, McDonald SJ, Keightley MC, Charzynska A, Dabrowski M, Dworkin S. Meta-Analysis of Grainyhead-Like Dependent Transcriptional Networks: A Roadmap for Identifying Novel Conserved Genetic Pathways. Genes. 2019; 10(11):876. https://doi.org/10.3390/genes10110876
Chicago/Turabian StyleMathiyalagan, Nishanthi, Lee B. Miles, Peter J. Anderson, Tomasz Wilanowski, Brian L. Grills, Stuart J. McDonald, M. Cristina Keightley, Agata Charzynska, Michal Dabrowski, and Sebastian Dworkin. 2019. "Meta-Analysis of Grainyhead-Like Dependent Transcriptional Networks: A Roadmap for Identifying Novel Conserved Genetic Pathways" Genes 10, no. 11: 876. https://doi.org/10.3390/genes10110876
APA StyleMathiyalagan, N., Miles, L. B., Anderson, P. J., Wilanowski, T., Grills, B. L., McDonald, S. J., Keightley, M. C., Charzynska, A., Dabrowski, M., & Dworkin, S. (2019). Meta-Analysis of Grainyhead-Like Dependent Transcriptional Networks: A Roadmap for Identifying Novel Conserved Genetic Pathways. Genes, 10(11), 876. https://doi.org/10.3390/genes10110876





