An Insight into the microRNAs Associated with Arteriovenous and Cavernous Malformations of the Brain
Abstract
1. Introduction
2. Methods
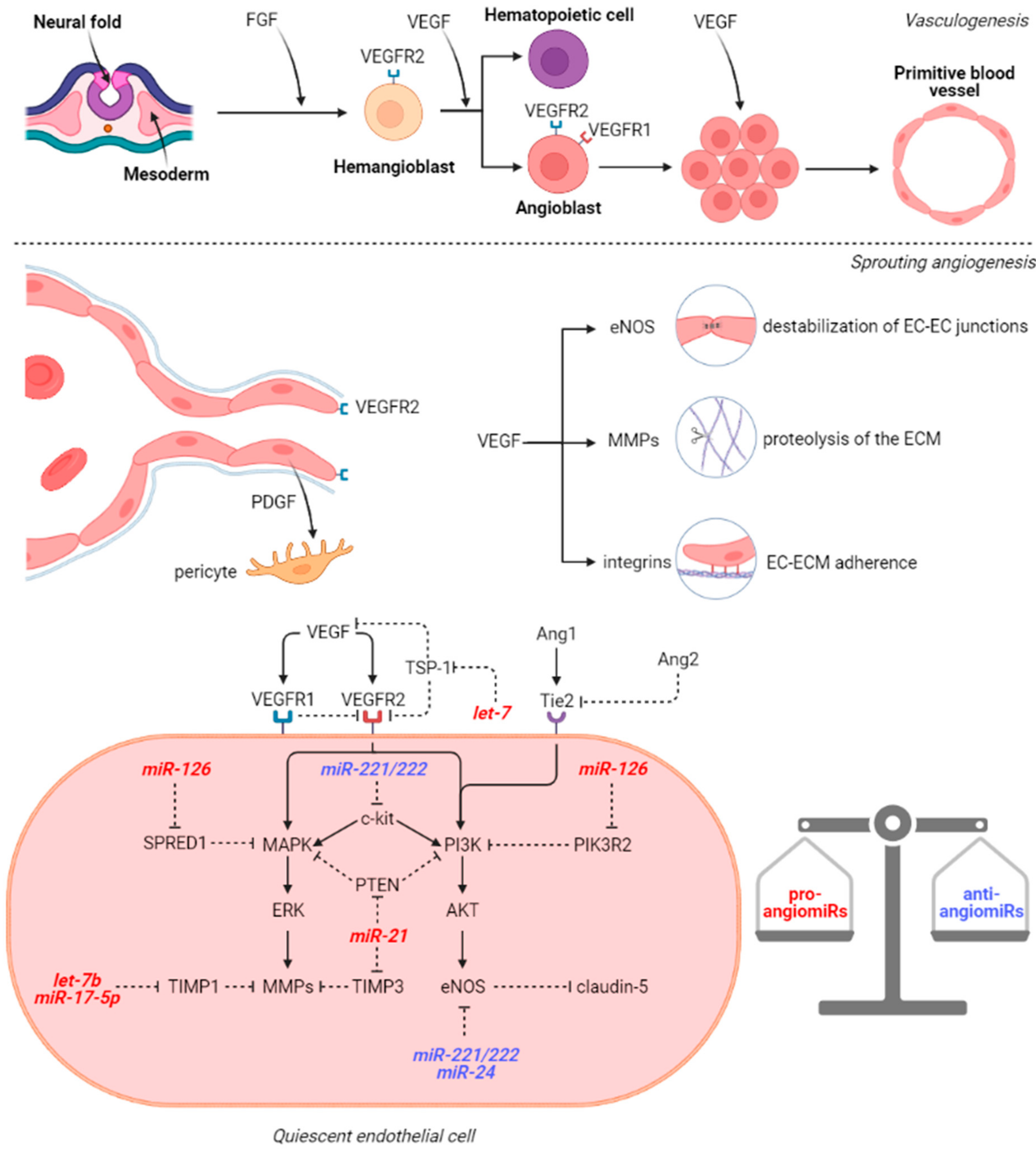
3. microRNAs Involved in Cerebral Vasculogenesis and Angiogenesis
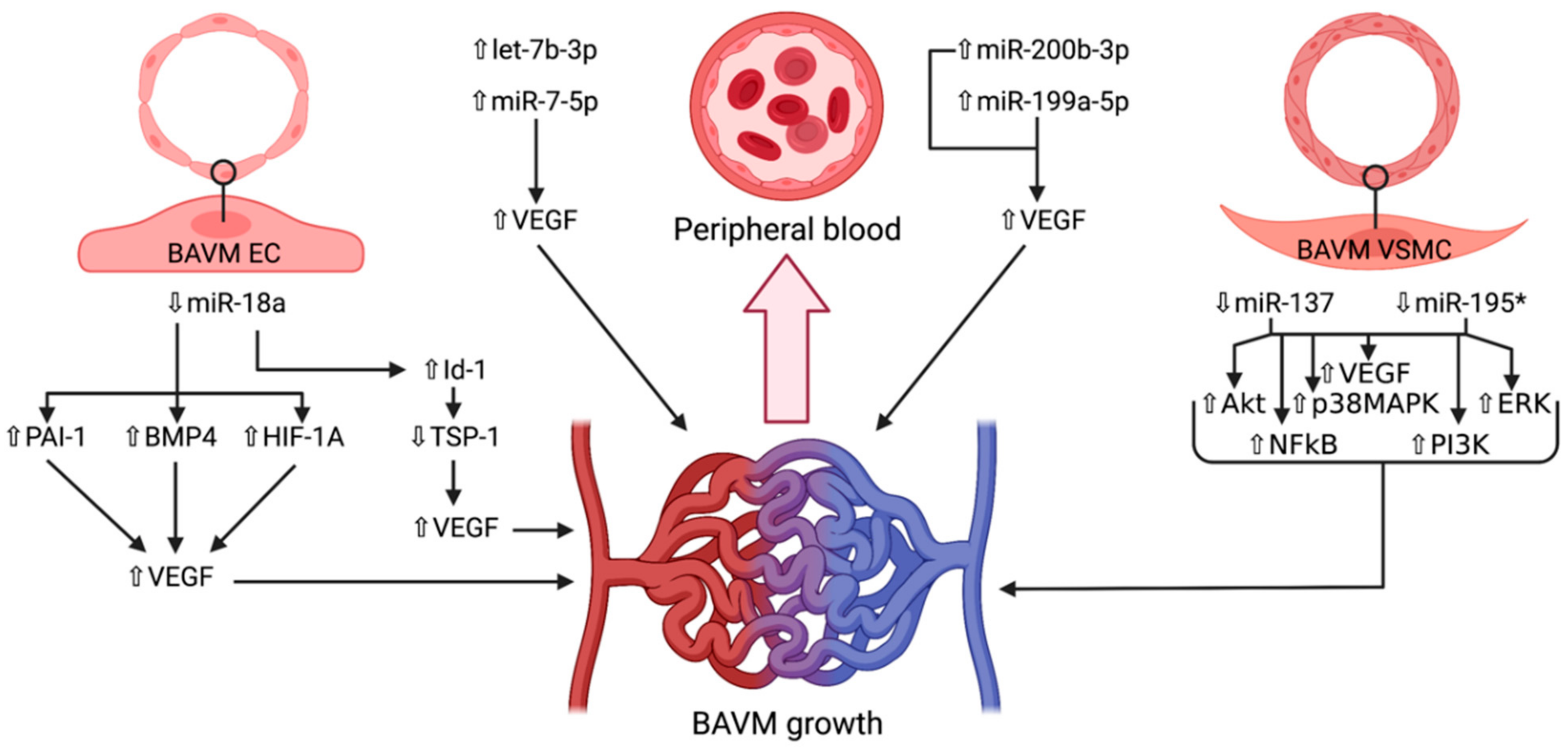
4. microRNAs in Brain Arteriovenous Malformations
4.1. miR-7-5p
4.2. miR-18a
4.3. miR-137 and miR-195*
4.4. miR-199a-5p
4.5. miR-200b-3p
4.6. let-7b-3p
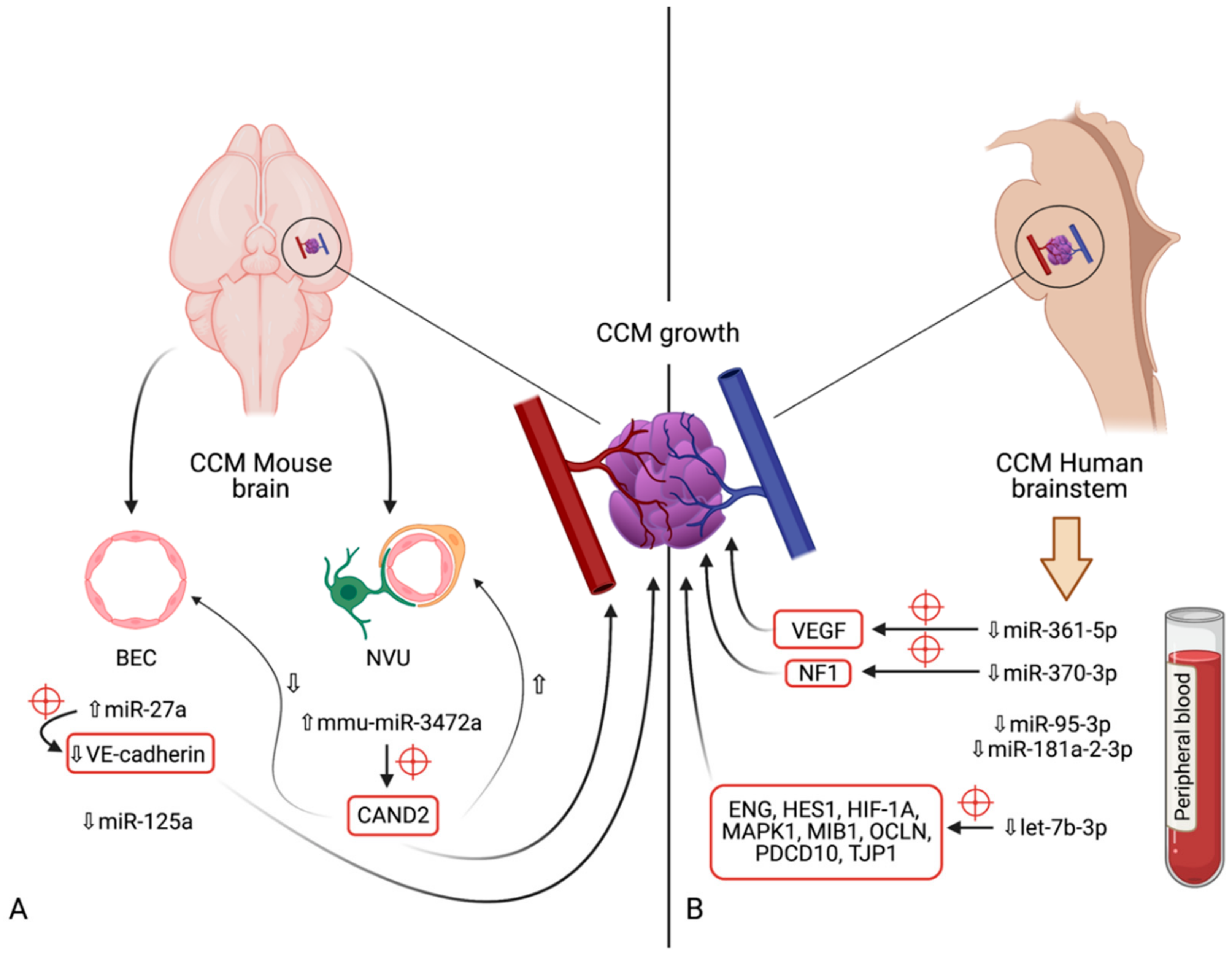
5. microRNAs in Cerebral Cavernous Malformations
5.1. miR-27a
5.2. miR-125a
5.3. mmu-miR-3472a
5.4. miR-361-5p
5.5. miR-370-3p
5.6. miR-181a-2-3p
5.7. miR-95-3p
6. Limitations
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| BAVM | brain arteriovenous malformation |
| CCM | cerebral cavernous malformation |
| miRNA | microRNA |
References
- Abecassis, I.; Xu, D.; Batjer, H.; Al, E. Natural history of brain arteriovenous malformations: A systematic review. Neurosurg Focus. 2014, 37, E7. [Google Scholar] [CrossRef]
- Ion, G.; Chiriac, A.; Dobrin, N.; Poeată, I. Multimodality treatment of brain arteriovenous malformation. Case report. Rom. Neurosurg. 2016, 30, 70–76. [Google Scholar] [CrossRef]
- Kalb, S.; Gross, B.A.; Nakaji, P. Vascular Malformations (Arteriovenous Malformations and Dural Arteriovenous Fistulas). In Principles of Neurological Surgery, 4th ed.; Ellenbogen, R.G., Sekhar, L.N., Kitchen, N.D., Eds.; Elsevier: Amsterdam, The Netherlands, 2018; Volume 20, pp. 313–324. [Google Scholar]
- Florian, I.S.; Barițchii, A.; Trifoi, S.V. Arterio-venous Mallformations. In Tratat de Chirurgie, 2nd ed.; Popescu, I., Ciuce, C., Eds.; Editura Academiei Române: București, Romania, 2014; Volume VI, pp. 402–410. (In Romanian) [Google Scholar]
- Florian, I.S.; Perju-Dumbravă, L. Therapeutic Options in Hemorrhagic Strokes; Editura Medicală Universitară, Iuliu Hațieganu: Cluj-Napoca, Romania, 2007; pp. 331–346. (In Romanian) [Google Scholar]
- Shetty, R.; Kato, Y.; Watabe, T.; Oguri, D. Unruptured Arteriovenous Malformations of Brain An overview. Rom. Neurosurg. 2010, 17, 34–45. [Google Scholar]
- Florian, I.; Beni, L.; Moisoiu, V.; Timis, T.; Florian, I.; Balașa, A.; Berindan-Neagoe, I. ‘De Novo’ Brain AVMs—Hypotheses for Development and a Systematic Review of Reported Cases. Medicina 2021, 57, 201. [Google Scholar] [CrossRef] [PubMed]
- Park, H.; Koh, E.J.; Lee, E.J.; Cheon, J.-E.; Kim, S.-K. An acquired cerebral arteriovenous malformation after brain abscess treatment: Case report and a review of the literature. Child’s Nerv. Syst. 2021, 1–4. [Google Scholar] [CrossRef]
- Bertalanffy, H.; Florian, I.A.; Timiș, T.L. Cavernous Malformations of the Pineal Region: Overview, Management, and Controversies. In Pineal Region Lesions; Florian, I.S., Ed.; Springer: Cham, Switzerland, 2020. [Google Scholar] [CrossRef]
- Choudhri, O.; Chen, R.P.; Bulsara, K. Cavernous Malformations of the Brain and Spinal Cord, 4th ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2018. [Google Scholar] [CrossRef]
- Iliescu, B.F.; Poeata, I. Cerebral Cavernomas. In Tratat de Chirurgie, 2nd ed.; Popescu, I., Ciuce, C., Eds.; Editura Academiei Române: București, Romania, 2014; Volume VI, pp. 411–414. (In Romanian) [Google Scholar]
- Bozinov, O.; Hatano, T.; Sarnthein, J.; Burkhardt, J.-K.; Bertalanffy, H. Current clinical management of brainstem cavernomas. Swiss Med. Wkly. 2010, 140, w13120. [Google Scholar] [CrossRef]
- Akers, A.; Salman, R.A.-S.; Awad, I.A.; Dahlem, K.; Flemming, K.; Hart, B.; Kim, H.; Jusue-Torres, I.; Kondziolka, D.; Lee, C.; et al. Synopsis of Guidelines for the Clinical Management of Cerebral Cavernous Malformations: Consensus Recommendations Based on Systematic Literature Review by the Angioma Alliance Scientific Advisory Board Clinical Experts Panel. Neurosurgery 2017, 80, 665–680. [Google Scholar] [CrossRef] [PubMed]
- A Winkler, E.; Rutledge, C.; Ward, M.; Tihan, T.; Sneed, P.K.; Barbaro, N.; Garcia, P.; McDermott, M.; Chang, E.F. Radiation-induced Cavernous Malformation as a Late Sequelae of Stereotactic Radiosurgery for Epilepsy. Cureus 2018, 10, e2308. [Google Scholar] [CrossRef]
- Hui, X.; Wang, Q.; Zhang, S. Cavernous malformation induced by stereotactic radiosurgery: A report and literature review. Neurol. India 2018, 66, 515. [Google Scholar] [CrossRef]
- Yu, Z.; Huang, B.; Liang, R. Radiation-induced cavernous malformation after stereotactic radiosurgery for cavernous sinus meningioma: A case report. BMC Neurol. 2020, 20, 422. [Google Scholar] [CrossRef]
- Araldi, E.; Suárez, Y. MicroRNAs as regulators of endothelial cell functions in cardiometabolic diseases. Biochim. Biophys. Acta BBA Mol. Cell Biol. Lipids 2016, 1861, 2094–2103. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Li, Z.; Shi, Y.; Huang, G.; Chen, L.; Tan, H.; Wang, Z.; Yin, C.; Hu, J. Deep Sequencing of Small RNAs in Blood of Patients with Brain Arteriovenous Malformations. World Neurosurg. 2018, 115, e570–e579. [Google Scholar] [CrossRef] [PubMed]
- Ferreira, R.; Santos, T.; Amar, A.; Tahara, S.M.; Chen, T.C.; Giannotta, S.L.; Hofman, F.M. MicroRNA-18a improves human cerebral arteriovenous malformation endothelial cell function. Stroke 2014, 45, 293–297. [Google Scholar] [CrossRef]
- Huang, J.; Song, J.; Qu, M.; Wang, Y.; An, Q.; Song, Y.; Yan, W.; Wang, B.; Wang, X.; Zhang, S.; et al. MicroRNA-137 and -195* inhibit vasculogenesis in brain arteriovenous malformations. Ann. Neurol. 2017, 82, 371–384. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Dai, Y.-X.; Wang, S.-Q.; Qiu, M.-K.; Quan, Z.-W.; Liu, Y.-B.; Ou, J.-M. miR-199a-5p inhibits proliferation and induces apoptosis in hemangioma cells through targeting HIF1A. Int. J. Immunopathol. Pharmacol. 2018, 31, 0394632017749357. [Google Scholar] [CrossRef]
- Zammar, S.G.; El Tecle, N.E.; El Ahmadieh, T.Y.; McClendon, J.; Comair, Y.G.; Bendok, B.R. A Biological Approach to Treating Brain Arteriovenous Malformations. Neurosurgery 2014, 74, N15–N17. [Google Scholar] [CrossRef]
- Fong, G.-H.; Rossant, J.; Gertsenstein, M.; Breitman, M.L. Role of the Flt-1 receptor tyrosine kinase in regulating the assembly of vascular endothelium. Nature 1995, 376, 66–70. [Google Scholar] [CrossRef]
- Shalaby, F.; Rossant, J.; Yamaguchi, T.P.; Gertsenstein, M.; Wu, X.-F.; Breitman, M.L.; Schuh, A.C. Failure of blood-island formation and vasculogenesis in Flk-1-deficient mice. Nature 1995, 376, 62–66. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.J.; Yang, D.D.; Na, S.; Sandusky, G.E.; Zhang, Q.; Zhao, G. Dicer Is Required for Embryonic Angiogenesis during Mouse Development. J. Biol. Chem. 2005, 280, 9330–9335. [Google Scholar] [CrossRef]
- Thomas, K.A. Angiogenesis. In Encyclopedia of Cell Biology; Elsevier: Amsterdam, The Netherlands, 2016; pp. 102–116. Available online: https://linkinghub.elsevier.com/retrieve/pii/B9780123944474400192 (accessed on 30 April 2021).
- Taddei, A.; Giampietro, C.; Conti, A.; Orsenigo, F.; Breviario, F.; Pirazzoli, V.; Potente, M.; Daly, C.; Dimmeler, S.; Dejana, E. Endothelial adherens junctions control tight junctions by VE-cadherin-mediated upregulation of claudin-5. Nat. Cell Biol. 2008, 10, 923–934. [Google Scholar] [CrossRef]
- Hogan, K.A.; Ambler, C.A.; Chapman, D.; Bautch, V.L. The neural tube patterns vessels developmentally using the VEGF signaling pathway. Development 2004, 131, 1503–1513. [Google Scholar] [CrossRef]
- Rundhaug, J.E. Matrix metalloproteinases and angiogenesis. J. Cell. Mol. Med. 2005, 9, 267–285. [Google Scholar] [CrossRef]
- Sato, T.N.; Tozawa, Y.; Deutsch, U.; Wolburg-Buchholz, K.; Fujiwara, Y.; Gendron-Maguire, M.; Gridley, T.; Wolburg, H.; Risau, W.; Qin, Y. Distinct roles of the receptor tyrosine kinases Tie-1 and Tie-2 in blood vessel formation. Nature 1995, 376, 70–74. [Google Scholar] [CrossRef] [PubMed]
- Seegar, T.C.; Eller, B.; Tzvetkova-Robev, D.; Kolev, M.V.; Henderson, S.C.; Nikolov, D.B.; Barton, W.A. Tie1-Tie2 Interactions Mediate Functional Differences between Angiopoietin Ligands. Mol. Cell 2010, 37, 643–655. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Olson, E.N. AngiomiRs-Key regulators of angiogenesis. Curr. Opin. Genet. Dev. 2009, 19, 205–211. [Google Scholar] [CrossRef] [PubMed]
- Lawler, P.; Lawler, J. Molecular Basis for the Regulation of Angiogenesis by Thrombospondin-1 and -2. Cold Spring Harb. Perspect. Med. 2012, 2, a006627. [Google Scholar] [CrossRef] [PubMed]
- Kuehbacher, A.; Urbich, C.; Zeiher, A.M.; Dimmeler, S. Role of Dicer and Drosha for Endothelial MicroRNA Expression and Angiogenesis. Circ. Res. 2007, 101, 59–68. [Google Scholar] [CrossRef] [PubMed]
- Otsuka, M.; Zheng, M.; Hayashi, M.; Lee, J.D.; Yoshino, O.; Lin, S.; Han, J. Impaired microRNA processing causes corpus luteum insufficiency and infertility in mice. J. Clin. Investig. 2008, 118, 1944–1954. [Google Scholar] [CrossRef] [PubMed]
- Dews, M.; Homayouni, A.; Yu, D.; Murphy, D.; Sevignani, C.; Wentzel, E.; Furth, E.E.; Lee, W.M.; Enders, G.H.; Mendell, J.T.; et al. Augmentation of tumor angiogenesis by a Myc-activated microRNA cluster. Nat. Genet. 2006, 38, 1060–1065. [Google Scholar] [CrossRef]
- Liu, L.-Z.; Li, C.; Chen, Q.; Jing, Y.; Carpenter, R.; Jiang, Y.; Kung, H.-F.; Lai, L.; Jiang, B.-H. MiR-21 Induced Angiogenesis through AKT and ERK Activation and HIF-1α Expression. PLoS ONE 2011, 6, e19139. [Google Scholar]
- Hu, J.; Ni, S.; Cao, Y.; Zhang, T.; Wu, T.; Yin, X.; Lang, Y.; Lu, H. The Angiogenic Effect of microRNA-21 Targeting TIMP3 through the Regulation of MMP2 and MMP9. PLoS ONE 2016, 11, e0149537. [Google Scholar] [CrossRef]
- Chistiakov, D.A.; Sobenin, I.A.; Orekhov, A.N.; Bobryshev, Y.V. Human miR-221/222 in Physiological and Atherosclerotic Vascular Remodeling. BioMed Res. Int. 2015, 2015, 354517. [Google Scholar] [CrossRef] [PubMed]
- Marchetti, M.; Meloni, M.; Anwar, M.; Al-Haj-Zen, A.; Sala-Newby, G.; Slater, S.; Ford, K.; Caporali, A.; Emanueli, C. MicroRNA-24-3p Targets Notch and Other Vascular Morphogens to Regulate Post-ischemic Microvascular Responses in Limb Muscles. Int. J. Mol. Sci. 2020, 21, 1733. [Google Scholar] [CrossRef] [PubMed]
- Fasanaro, P.; D’Alessandra, Y.; Di Stefano, V.; Melchionna, R.; Romani, S.; Pompilio, G.; Capogrossi, M.C.; Martelli, F. MicroRNA-210 Modulates Endothelial Cell Response to Hypoxia and Inhibits the Receptor Tyrosine Kinase Ligand Ephrin-A3. J. Biol. Chem. 2008, 283, 15878–15883. [Google Scholar] [CrossRef]
- Chen, Z.; Lai, T.-C.; Jan, Y.-H.; Lin, F.-M.; Wang, W.-C.; Xiao, H.; Wang, Y.-T.; Sun, W.; Cui, X.; Li, Y.-S.; et al. Hypoxia-responsive miRNAs target argonaute 1 to promote angiogenesis. J. Clin. Investig. 2013, 123, 1057–1067. [Google Scholar] [CrossRef]
- Jeyapalan, Z.; Deng, Z.; Shatseva, T.; Fang, L.; He, C.; Yang, B.B. Expression of CD44 3′-untranslated region regulates endogenous microRNA functions in tumorigenesis and angiogenesis. Nucleic Acids Res. 2011, 39, 3026–3041. [Google Scholar] [CrossRef]
- Suárez, Y.; Fernández-Hernando, C.; Yu, J.; Gerber, S.A.; Harrison, K.D.; Pober, J.S.; Iruela-Arispe, M.L.; Merkenschlager, M.; Sessa, W.C. Dicer-dependent endothelial microRNAs are necessary for postnatal angiogenesis. Proc. Natl. Acad. Sci. USA 2008, 105, 14082–14087. [Google Scholar] [CrossRef]
- Dong, L.; Li, Y.; Han, C.; Wang, X.; She, L.; Zhang, H. miRNA microarray reveals specific expression in the peripheral blood of glioblastoma patients. Int. J. Oncol. 2014, 45, 746–756. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Huang, M.; Cai, Y.; Yang, Y.; Sun, X.; Ke, Y. Circ-U2AF1 promotes human glioma via derepressing neuro-oncological ventral antigen 2 by sponging hsa-miR-7-5p. J. Cell. Physiol. 2019, 234, 9144–9155. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Niu, W.; Mu, M.; Hu, S.; Niu, C. Long non-coding RNA LPP-AS2 promotes glioma tumorigenesis via miR-7-5p/EGFR/PI3K/AKT/c-MYC feedback loop. J. Exp. Clin. Cancer Res. 2020, 39, 1–20. [Google Scholar] [CrossRef]
- Liu, Z.; Liu, Y.; Li, L.; Xu, Z.; Bi, B.; Wang, Y.; Li, J.Y. MiR-7-5p is frequently downregulated in glioblastoma microvasculature and inhibits vascular endothelial cell proliferation by targeting RAF1. Tumor Biol. 2014, 35, 10177–10184. [Google Scholar] [CrossRef]
- Xu, H.; Nie, B.; Liu, L.; Zhang, C.; Zhang, Z.; Xu, M.; Mei, Y. Curcumin Prevents Brain Damage and Cognitive Dysfunction During Ischemic-reperfusion Through the Regulation of miR-7-5p. Curr. Neurovascular Res. 2020, 16, 441–454. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Wang, B. MiR-7-5p Enhances Cerebral Ischemia–Reperfusion Injury by Degrading sirt1 mRNA. J. Cardiovasc. Pharmacol. 2020, 76, 227–236. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Deng, S.; Lei, Q.; He, Q.; Ren, Y.; Zhang, Y.; Nie, J.; Lu, W. miR-7-5p Affects Brain Edema After Intracerebral Hemorrhage and Its Possible Mechanism. Front. Cell Dev. Biol. 2020, 8, 598020. [Google Scholar] [CrossRef]
- Xu, M.; Xu, H.; Qin, Z.; Zhang, J.; Yang, X.; Xu, F. Increased Expression of Angiogenic Factors in Cultured Human Brain Arteriovenous Malformation Endothelial Cells. Cell Biophys. 2014, 70, 443–447. [Google Scholar] [CrossRef]
- Rangel-Castilla, L.; Russin, J.J.; Martinez-Del-Campo, E.; Soriano-Baron, H.; Spetzler, R.F.; Nakaji, P. Molecular and cellular biology of cerebral arteriovenous malformations: A review of current concepts and future trends in treatment. Neurosurg. Focus 2014, 37, E1. [Google Scholar] [CrossRef]
- Kolenda, T.; Guglas, K.; Kopczyńska, M.; Sobocińska, J.; Teresiak, A.; Bliźniak, R.; Lamperska, K. Good or not good: Role of miR-18a in cancer biology. Rep. Pract. Oncol. Radiother. 2020, 25, 808–819. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Wang, M.; Deng, D. c-Fos/microRNA-18a feedback loop modulates the tumor growth via HMBOX1 in human gliomas. Biomed. Pharmacother. 2018, 107, 1705–1711. [Google Scholar] [CrossRef]
- Gruszka, R.; Zakrzewski, K.; Liberski, P.P.; Zakrzewska, M. mRNA and miRNA Expression Analyses of the MYC/E2F/miR-17-92 Network in the Most Common Pediatric Brain Tumors. Int. J. Mol. Sci. 2021, 22, 543. [Google Scholar] [CrossRef]
- Miao, Y.-S.; Zhao, Y.-Y.; Zhao, L.-N.; Wang, P.; Liu, Y.-H.; Ma, J.; Xue, Y.-X. MiR-18a increased the permeability of BTB via RUNX1 mediated down-regulation of ZO-1, occludin and claudin-5. Cell. Signal. 2015, 27, 156–167. [Google Scholar] [CrossRef]
- Song, Y.; Wang, P.; Zhao, W.; Yao, Y.; Liu, X.; Ma, J.; Xue, Y.; Liu, Y. MiR-18a regulates the proliferation, migration and invasion of human glioblastoma cell by targeting neogenin. Exp. Cell Res. 2014, 324, 54–64. [Google Scholar] [CrossRef]
- Wilson, N.H.; Key, B. Neogenin: One receptor, many functions. Int. J. Biochem. Cell Biol. 2007, 39, 874–878. [Google Scholar] [CrossRef]
- Cole, S.J.; Bradford, D.; Cooper, H.M. Neogenin: A multi-functional receptor regulating diverse developmental processes. Int. J. Biochem. Cell Biol. 2007, 39, 1569–1575. [Google Scholar] [CrossRef]
- De Vries, M.; Cooper, H.M. Emerging roles for neogenin and its ligands in CNS development. J. Neurochem. 2008, 106, 1483–1492. [Google Scholar] [CrossRef]
- Rodrigues, S.P.; De Wever, O.; Bruyneel, E.; Rooney, R.J.; Gespach, C. Opposing roles of netrin-1 and the dependence receptor DCC in cancer cell invasion, tumor growth and metastasis. Oncogene 2007, 26, 5615–5625. [Google Scholar] [CrossRef]
- Wang, Y.; Tian, Y.; Li, Z.; Zheng, Z.; Zhu, L. miR-92 regulates the proliferation, migration, invasion and apoptosis of glioma cells by targeting neogenin. Open Med. 2020, 15, 283–291. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Li, Y.; Wan, X.; Kayira, T.M.; Cao, R.; Ju, X.; Zhu, X.; Zhao, G. Down-Regulation of Neogenin Accelerated Glioma Progression through Promoter Methylation and Its Overexpression in SHG-44 Induced Apoptosis. PLoS ONE 2012, 7, e38074. [Google Scholar] [CrossRef]
- Sesen, J.; Driscoll, J.; Shah, N.; Moses-Gardner, A.; Luiselli, G.; Alexandrescu, S.; Zurakowski, D.; Baxter, P.A.; Su, J.M.; Fehnel, K.P.; et al. Neogenin is highly expressed in diffuse intrinsic pontine glioma and influences tumor invasion. Brain Res. 2021, 1762, 147348. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.; Yao, L.-L.; Pan, J.-X.; Zhang, J.-S.; Mei, L.; Wang, Y.-G.; Xiong, W.-C. Linking cortical astrocytic neogenin deficiency to the development of Moyamoya disease–like vasculopathy. Neurobiol. Dis. 2021, 154, 105339. [Google Scholar] [CrossRef]
- Yao, L.-L.; Hu, J.-X.; Li, Q.; Lee, D.; Ren, X.; Zhang, J.-S.; Sun, D.; Zhang, H.-S.; Wang, Y.-G.; Mei, L.; et al. Astrocytic neogenin/netrin-1 pathway promotes blood vessel homeostasis and function in mouse cortex. J. Clin. Investig. 2020, 130, 6490–6509. [Google Scholar] [CrossRef] [PubMed]
- Marín-Ramos, N.I.; Thein, T.Z.; Ghaghada, K.B.; Chen, T.C.; Giannotta, S.L.; Hofman, F.M. miR-18a Inhibits BMP4 and HIF-1α Normalizing Brain Arteriovenous Malformations. Circ. Res. 2020, 127. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Cai, Q.; Lin, S.; Chen, B.; Jia, B.; Ye, R.; Weygant, N.; Chu, J.; Peng, J. Qingda granule exerts neuroprotective effects against ischemia/reperfusion-induced cerebral injury via lncRNA GAS5/miR-137 signaling pathway. Int. J. Med. Sci. 2021, 18, 1687–1698. [Google Scholar] [CrossRef]
- Zhang, M.; Ge, D.; Su, Z.; Qi, B. miR-137 alleviates focal cerebral ischemic injury in rats by regulating JAK1/STAT1 signaling pathway. Hum. Exp. Toxicol. 2020, 39, 816–827. [Google Scholar] [CrossRef]
- Tian, R.; Wu, B.; Fu, C.; Guo, K. miR-137 prevents inflammatory response, oxidative stress, neuronal injury and cognitive impairment via blockade of Src-mediated MAPK signaling pathway in ischemic stroke. Aging 2020, 12, 10873–10895. [Google Scholar] [CrossRef] [PubMed]
- Pacheco, A.; Berger, R.; Freedman, R.; Law, A.J. A VNTR Regulates miR-137 Expression Through Novel Alternative Splicing and Contributes to Risk for Schizophrenia. Sci. Rep. 2019, 9, 11793. [Google Scholar] [CrossRef]
- Arakawa, Y.; Yokoyama, K.; Tasaki, S.; Kato, J.; Nakashima, K.; Takeyama, M.; Nakatani, A.; Suzuki, M. Transgenic mice overexpressing miR-137 in the brain show schizophrenia-associated behavioral deficits and transcriptome profiles. PLoS ONE 2019, 14, e0220389. [Google Scholar] [CrossRef] [PubMed]
- Kandratsenka, H.; Nestsiarovich, A.; Goloenko, I.; Danilenko, N.; Makarevich, A.; Obyedkov, V.; Davydenko, O.; Waszkiewicz, N. Association of MIR137 With Symptom Severity and Cognitive Functioning in Belarusian Schizophrenia Patients. Front. Psychiatry 2018, 9, 295. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.; Shang, S.; Wang, J.; Zhang, T.; Nie, F.; Song, X.; Zhao, H.; Zhu, C.; Zhang, R.; Hao, D. Identification of miR-22-3p, miR-92a-3p, and miR-137 in peripheral blood as biomarker for schizophrenia. Psychiatry Res. 2018, 265, 70–76. [Google Scholar] [CrossRef]
- Sudesh, R.; Thalamuthu, A.; John, S.; Thara, R.; Mowry, B.; Munirajan, A.K. Replication of GWAS identified miR-137 and its target gene polymorphisms in Schizophrenia of South Indian population and meta-analysis with Psychiatric Genomics Consortium. Schizophr. Res. 2018, 199, 189–194. [Google Scholar] [CrossRef]
- Sakamoto, K.; Crowley, J.J. A comprehensive review of the genetic and biological evidence supports a role for MicroRNA-137 in the etiology of schizophrenia. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 2018, 177, 242–256. [Google Scholar] [CrossRef]
- Bier, A.; Giladi, N.; Kronfeld, N.; Lee, H.K.; Cazacu, S.; Finniss, S.; Xiang, C.; Poisson, L.; Decarvalho, A.C.; Slavin, S.; et al. MicroRNA-137 is downregulated in glioblastoma and inhibits the stemness of glioma stem cells by targeting RTVP-1. Oncotarget 2013, 4, 665–676. [Google Scholar] [CrossRef]
- Liang, M.-L.; Hsieh, T.-H.; Ng, K.-H.; Tsai, Y.-N.; Tsai, C.-F.; Chao, M.-E.; Liu, D.-J.; Chu, S.-S.; Chen, W.; Liu, Y.-R.; et al. Downregulation of miR-137 and miR-6500-3p promotes cell proliferation in pediatric high-grade gliomas. Oncotarget 2016, 7, 19723–19737. [Google Scholar] [CrossRef] [PubMed]
- Sun, G.; Cao, Y.; Shi, L.; Sun, L.; Wang, Y.; Chen, C.; Wan, Z.; Fu, L.; You, Y. Overexpressed miRNA-137 Inhibits Human Glioma Cells Growth by Targeting Rac1. Cancer Biother. Radiopharm. 2013, 28, 327–334. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Wang, X.; Wang, H.; Li, Y.; Yan, W.; Han, L.; Zhang, K.; Zhang, J.; Wang, Y.; Feng, Y.; et al. miR-137 is frequently down-regulated in glioblastoma and is a negative regulator of Cox-2. Eur. J. Cancer 2012, 48, 3104–3111. [Google Scholar] [CrossRef] [PubMed]
- Li, K.K.-W.; Yang, L.; Pang, J.C.-S.; Chan, A.K.-Y.; Zhou, L.; Mao, Y.; Wang, Y.; Lau, K.-M.; Poon, W.S.; Shi, Z.; et al. MIR-137 Suppresses Growth and Invasion, is Downregulated in Oligodendroglial Tumors and Targets CSE1L. Brain Pathol. 2012, 23, 426–439. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Mi, Y.; Guo, N.; Xu, H.; Jiang, P.; Zhang, R.; Xu, L. Glioma in Schizophrenia: Is the Risk Higher or Lower? Front. Cell. Neurosci. 2018, 12, 289. [Google Scholar] [CrossRef]
- Zhao, W.-J.; Zhang, H.-F.; Su, J.-Y. Downregulation of microRNA-195 promotes angiogenesis induced by cerebral infarction via targeting VEGFA. Mol. Med. Rep. 2017, 16, 5434–5440. [Google Scholar] [CrossRef]
- Pan, S.; Feng, W.; Li, Y.; Huang, J.; Chen, S.; Cui, Y.; Tian, B.; Tan, S.; Wang, Z.; Yao, S.; et al. The microRNA-195-BDNF pathway and cognitive deficits in schizophrenia patients with minimal antipsychotic medication exposure. Transl. Psychiatry 2021, 11, 117. [Google Scholar] [CrossRef]
- Huang, X.; Bao, C.; Lv, Q.; Zhao, J.; Wang, Y.; Lang, X.; Li, Z.; Yi, Z. Sex difference in cognitive impairment in drug-free schizophrenia: Association with miR-195 levels. Psychoneuroendocrinology 2020, 119, 104748. [Google Scholar] [CrossRef]
- Cao, J.; Huang, M.; Guo, L.; Zhu, L.; Hou, J.; Zhang, L.; Pero, A.; Ng, S.; El Gaamouch, F.; Elder, G.; et al. MicroRNA-195 rescues ApoE4-induced cognitive deficits and lysosomal defects in Alzheimer’s disease pathogenesis. Mol. Psychiatry 2020. [Google Scholar] [CrossRef]
- Zhu, H.-C.; Wang, L.-M.; Wang, M.; Song, B.; Tan, S.; Teng, J.-F.; Duan, D.-X. MicroRNA-195 downregulates Alzheimer’s disease amyloid-β production by targeting BACE1. Brain Res. Bull. 2012, 88, 596–601. [Google Scholar] [CrossRef]
- Ai, J.; Sun, L.-H.; Che, H.; Zhang, R.; Zhang, T.-Z.; Wu, W.-C.; Su, X.-L.; Chen, X.; Yang, G.; Li, K.; et al. MicroRNA-195 Protects Against Dementia Induced by Chronic Brain Hypoperfusion via Its Anti-Amyloidogenic Effect in Rats. J. Neurosci. 2013, 33, 3989–4001. [Google Scholar] [CrossRef]
- Chen, X.; Jiang, X.-M.; Zhao, L.-J.; Sun, L.-L.; Yan, M.-L.; Tian, Y.; Zhang, S.; Duan, M.-J.; Zhao, H.-M.; Li, W.-R.; et al. MicroRNA-195 prevents dendritic degeneration and neuron death in rats following chronic brain hypoperfusion. Cell Death Dis. 2017, 8, e2850. [Google Scholar] [CrossRef] [PubMed]
- Chang, L.; Zhang, W.; Shi, S.; Peng, Y.; Wang, D.; Zhang, L.; Zhang, J. microRNA-195 attenuates neuronal apoptosis in rats with ischemic stroke through inhibiting KLF5-mediated activation of the JNK signaling pathway. Mol. Med. 2020, 26, 31. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.-Y.; Wang, Y.-S.; Hsu, P.-Y.; Chen, C.-Y.; Liao, Y.-C.; Juo, S.-H.H. miR-195 Has a Potential to Treat Ischemic and Hemorrhagic Stroke through Neurovascular Protection and Neurogenesis. Mol. Ther. Methods Clin. Dev. 2019, 13, 121–132. [Google Scholar] [CrossRef] [PubMed]
- Song, L.-R.; Li, D.; Weng, J.-C.; Li, C.-B.; Wang, L.; Wu, Z.; Zhang, J.-T. MicroRNA-195 Functions as a Tumor Suppressor by Directly Targeting Fatty Acid Synthase in Malignant Meningioma. World Neurosurg. 2020, 136, e355–e364. [Google Scholar] [CrossRef]
- Jia, Y.; Tian, Y.; An, S.; Yang, D. Effects of microRNA-195 on the Prognosis of Glioma Patients and the Proliferation and Apoptosis of Human Glioma Cells. Pathol. Oncol. Res. 2020, 26, 753–763. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-P.; Zhang, N.-N.; Ren, X.-Q.; He, J.; Li, Y. miR-103/miR-195/miR-15b Regulate SALL4 and Inhibit Proliferation and Migration in Glioma. Molecules 2018, 23, 2938. [Google Scholar] [CrossRef]
- Gu, S.; Cheung, H.H.; Lee, T.L.; Lu, G.; Poon, W.S.; Chan, W.Y. Molecular Mechanisms of Regulation and Action of microRNA-199a in Testicular Germ Cell Tumor and Glioblastomas. PLoS ONE 2013, 8, e83980. [Google Scholar] [CrossRef]
- Li, Z.; Gu, X.-D.; Fang, Y.; Xiang, J.; Chen, Z. microRNA expression profiles in human colorectal cancers with brain metastases. Oncol. Lett. 2012, 3, 346–350. [Google Scholar] [CrossRef]
- Wang, G.; Li, Y.; Li, J.; Zhang, D.; Luo, C.; Zhang, B.; Sun, X. microRNA-199a-5p suppresses glioma progression by inhibiting MAGT1. J. Cell. Biochem. 2019, 120, 15248–15254. [Google Scholar] [CrossRef]
- Zhang, C.; Chen, Q.; Zhu, J.-W.; Liu, Z.-F. MicroRNA-199a-5p regulates glioma progression via targeting MARCH8. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7482–7487. [Google Scholar] [CrossRef]
- Chi, G.; Yang, F.; Xu, D.; Liu, W. Silencing hsa_circ_PVT1 (circPVT1) suppresses the growth and metastasis of glioblastoma multiforme cells by up-regulation of miR-199a-5p. Artif. Cells Nanomed. Biotechnol. 2020, 48, 188–196. [Google Scholar] [CrossRef] [PubMed]
- Zhong, W.; Li, Y.-C.; Huang, Q.-Y.; Tang, X.-Q. lncRNA ANRIL Ameliorates Oxygen and Glucose Deprivation (OGD) Induced Injury in Neuron Cells via miR-199a-5p/CAV-1 Axis. Neurochem. Res. 2020, 45, 772–782. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Luan, L.; Liu, Q.; Liu, Y.; Lan, X.; Li, Z.; Liu, W. MiRNA-199a-5p Protects Against Cerebral Ischemic Injury by Down-Regulating DDR1 in Rats. World Neurosurg. 2019, 131, e486–e494. [Google Scholar] [CrossRef]
- Bao, N.; Fang, B.; Lv, H.; Jiang, Y.; Chen, F.; Wang, Z.; Ma, H. Upregulation of miR-199a-5p Protects Spinal Cord Against Ischemia/Reperfusion-Induced Injury via Downregulation of ECE1 in Rat. Cell. Mol. Neurobiol. 2018, 38, 1293–1303. [Google Scholar] [CrossRef] [PubMed]
- Hou, J.; Wang, L.; Wu, Q.; Zheng, G.; Long, H.; Wu, H.; Zhou, C.; Guo, T.; Zhong, T.; Wang, L.; et al. Long noncoding RNA H19 upregulates vascular endothelial growth factor A to enhance mesenchymal stem cells survival and angiogenic capacity by inhibiting miR-199a-5p. Stem Cell Res. Ther. 2018, 9, 109. [Google Scholar] [CrossRef]
- Heuslein, J.L.; Gorick, C.M.; McDonnell, S.P.; Song, J.; Annex, B.H.; Price, R.J. Exposure of Endothelium to Biomimetic Flow Waveforms Yields Identification of miR-199a-5p as a Potent Regulator of Arteriogenesis. Mol. Ther. Nucleic Acids 2018, 12, 829–844. [Google Scholar] [CrossRef]
- Zhang, N.; Yang, L.; Meng, L.; Cui, H. Inhibition of miR-200b-3p alleviates hypoxia-ischemic brain damage via targeting Slit2 in neonatal rats. Biochem. Biophys. Res. Commun. 2020, 523, 931–938. [Google Scholar] [CrossRef]
- Visani, M.; Marucci, G.; De Biase, D.; Giangaspero, F.; Buttarelli, F.R.; Brandes, A.A.; Franceschi, E.; Acquaviva, G.; Ciarrocchi, A.; Rhoden, K.J.; et al. miR-196B-5P and miR-200B-3P Are Differentially Expressed in Medulloblastomas of Adults and Children. Diagnostics 2020, 10, 265. [Google Scholar] [CrossRef]
- Minn, Y.-K.; Lee, D.; Hyung, W.; Kim, J.; Choi, J.; Yang, S.-H.; Song, H.; Lim, B.; Kim, S.H. MicroRNA-200 family members and ZEB2 are associated with brain metastasis in gastric adenocarcinoma. Int. J. Oncol. 2014, 45, 2403–2410. [Google Scholar] [CrossRef]
- Comijn, J.; Berx, G.; Vermassen, P.; Verschueren, K.; van Grunsven, L.; Bruyneel, E.; Mareel, M.; Huylebroeck, D.; van Roy, F. The Two-Handed E Box Binding Zinc Finger Protein SIP1 Downregulates E-Cadherin and Induces Invasion. Mol. Cell 2001, 7, 1267–1278. [Google Scholar] [CrossRef]
- Li, J.; Yuan, J.; Yuan, X.; Zhao, J.; Zhang, Z.; Weng, L.; Liu, J. MicroRNA-200b inhibits the growth and metastasis of glioma cells via targeting ZEB2. Int. J. Oncol. 2016, 48, 541–550. [Google Scholar] [CrossRef] [PubMed]
- Wu, J.; Cui, H.; Zhu, Z.; Wang, L. MicroRNA-200b-3p suppresses epithelial-mesenchymal transition and inhibits tumor growth of glioma through down-regulation of ERK5. Biochem. Biophys. Res. Commun. 2016, 478, 1158–1164. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Jiang, Q.; Luo, D.; Zhao, L.; Fu, X.; Chen, Y.; Song, X.; Li, L.; Zhao, H.; He, Y.; et al. miR-200b is a key regulator of tumor progression and metabolism targeting lactate dehydrogenase A in human malignant glioma. Oncotarget 2016, 7, 48423–48431. [Google Scholar] [CrossRef] [PubMed]
- Kong, X.; Gong, S.; Yan, T.; Yang, Y. MicroRNA-200b expression level is negatively associated with pathological grading in human gliomas. Cancer Manag. Res. 2018, 10, 2825–2834. [Google Scholar] [CrossRef]
- Liu, H.; Tian, T.; Qin, S.; Li, W.; Zhang, X.; Wang, X.; Gao, Y.; Huang, G. Folic acid deficiency enhances abeta accumulation in APP/PS1 mice brain and decreases amyloid-associated miRNAs expression. J. Nutr. Biochem. 2015, 26, 1502–1508. [Google Scholar] [CrossRef]
- Higaki, S.; Muramatsu, M.; Matsuda, A.; Matsumoto, K.; Satoh, J.-I.; Michikawa, M.; Niida, S. Defensive effect of microRNA-200b/c against amyloid-beta peptide-induced toxicity in Alzheimer’s disease models. PLoS ONE 2018, 13, e0196929. [Google Scholar] [CrossRef]
- Boese, A.S.; Saba, R.; Campbell, K.; Majer, A.; Medina, S.; Burton, L.; Booth, T.F.; Chong, P.; Westmacott, G.R.; Dutta, S.M.; et al. MicroRNA abundance is altered in synaptoneurosomes during prion disease. Mol. Cell. Neurosci. 2016, 71, 13–24. [Google Scholar] [CrossRef]
- Osei, J.; Kelly, W.; Toffolo, K.; Donahue, K.; Levy, B.; Bard, J.; Wang, J.; Levy, E.; Nowak, N.; Poulsen, D. Thymosin beta 4 induces significant changes in the plasma miRNA profile following severe traumatic brain injury in the rat lateral fluid percussion injury model. Expert Opin. Biol. Ther. 2018, 18, 159–164. [Google Scholar] [CrossRef]
- Lee, K.H.; Lim, B.J.; Ferreira, V.H.; Min, S.Y.; Hong, Y.-M.; Jo, J.-H.; Han, S.H. Expression of human miR-200b-3p and -200c-3p in cytomegalovirus-infected tissues. Biosci. Rep. 2018, 38, BSR20180961. [Google Scholar] [CrossRef]
- Kar, S.; Bali, K.K.; Baisantry, A.; Geffers, R.; Samii, A.; Bertalanffy, H. Genome-Wide Sequencing Reveals MicroRNAs Downregulated in Cerebral Cavernous Malformations. J. Mol. Neurosci. 2017, 61, 178–188. [Google Scholar] [CrossRef] [PubMed]
- Alsharafi, W.A.; Xiao, B.; Abuhamed, M.M.; Luo, Z. miRNAs: Biological and clinical determinants in epilepsy. Front. Mol. Neurosci. 2015, 8, 59. [Google Scholar] [CrossRef]
- Srivastava, P.K.; Roncon, P.; Lukasiuk, K.; Gorter, J.A.; Aronica, E.; Pitkänen, A.; Petretto, E.; Johnson, M.R.; Simonato, M. Meta-Analysis of MicroRNAs Dysregulated in the Hippocampal Dentate Gyrus of Animal Models of Epilepsy. Eneuro 2017, 4. [Google Scholar] [CrossRef] [PubMed]
- Mills, J.D.; van Vliet, E.; Chen, B.J.; Janitz, M.; Anink, J.J.; Baayen, J.C.; Idema, S.; Devore, S.; Friedman, D.; Diehl, B.; et al. Coding and non-coding transcriptome of mesial temporal lobe epilepsy: Critical role of small non-coding RNAs. Neurobiol. Dis. 2020, 134, 104612. [Google Scholar] [CrossRef]
- Li, S.; Chen, L.; Zhou, X.; Li, J.; Liu, J. miRNA-223-3p and let-7b-3p as potential blood biomarkers associated with the ischemic penumbra in rats. Acta Neurobiol. Exp. 2019, 79, 205–216. [Google Scholar] [CrossRef]
- Tian, Y.; Hao, S.; Ye, M.; Zhang, A.; Nan, Y.; Wang, G.; Jia, Z.; Yuan, T.; Guo, L.; Pu, P.; et al. MicroRNAs let-7b/i suppress human glioma cell invasion and migration by targeting IKBKE directly. Biochem. Biophys. Res. Commun. 2015, 458, 307–312. [Google Scholar] [CrossRef]
- Song, H.; Zhang, Y.; Liu, N.; Zhang, D.; Wan, C.; Zhao, S.; Kong, Y.; Yuan, L. Let-7b inhibits the malignant behavior of glioma cells and glioma stem-like cells via downregulation of E2F2. J. Physiol. Biochem. 2016, 72, 733–744. [Google Scholar] [CrossRef] [PubMed]
- Zakrzewska, M.; Gruszka, R.; Stawiski, K.; Fendler, W.; Kordacka, J.; Grajkowska, W.; Daszkiewicz, P.; Liberski, P.P.; Zakrzewski, K. Expression-based decision tree model reveals distinct microRNA expression pattern in pediatric neuronal and mixed neuronal-glial tumors. BMC Cancer 2019, 19, 544. [Google Scholar] [CrossRef]
- Li, J.; Zhao, Y.; Choi, J.; Ting, K.K.; Coleman, P.; Chen, J.; Cogger, V.C.; Wan, L.; Shi, Z.; Moller, T.; et al. Targeting miR-27a/VE-cadherin interactions rescues cerebral cavernous malformations in mice. PLoS Biol. 2020, 18, e3000734. [Google Scholar] [CrossRef]
- Zhou, Z.; Tang, A.T.; Wong, W.-Y.; Bamezai, S.; Goddard, L.M.; Shenkar, R.; Zhou, S.; Yang, J.; Wright, A.C.; Foley, M.; et al. Cerebral cavernous malformations arise from endothelial gain of MEKK3–KLF2/4 signalling. Nature 2016, 532, 122–126. [Google Scholar] [CrossRef]
- Cuttano, R.; Rudini, N.; Bravi, L.; Corada, M.; Giampietro, C.; Papa, E.; Morini, M.F.; Maddaluno, L.; Baeyens, N.; Adams, R.H.; et al. KLF 4 is a key determinant in the development and progression of cerebral cavernous malformations. EMBO Mol. Med. 2016, 8, 6–24. [Google Scholar] [CrossRef]
- Lu, J.; Zhou, N.; Yang, P.; Deng, L.; Liu, G. MicroRNA-27a-3p Downregulation Inhibits Inflammatory Response and Hippocampal Neuronal Cell Apoptosis by Upregulating Mitogen-Activated Protein Kinase 4 (MAP2K4) Expression in Epilepsy: In Vivo and In Vitro Studies. Med. Sci. Monit. 2019, 25, 8499–8508. [Google Scholar] [CrossRef] [PubMed]
- Xi, T.; Jin, F.; Zhu, Y.; Wang, J.; Tang, L.; Wang, Y.; Liebeskind, D.S.; Scalzo, F.; He, Z. miR-27a-3p protects against blood–brain barrier disruption and brain injury after intracerebral hemorrhage by targeting endothelial aquaporin-11. J. Biol. Chem. 2018, 293, 20041–20050. [Google Scholar] [CrossRef]
- Sun, L.; Zhao, M.; Wang, Y.; Liu, A.; Lv, M.; Li, Y.; Yang, X.; Wu, Z. Neuroprotective effects of miR-27a against traumatic brain injury via suppressing FoxO3a-mediated neuronal autophagy. Biochem. Biophys. Res. Commun. 2017, 482, 1141–1147. [Google Scholar] [CrossRef]
- Sabirzhanov, B.; Zhao, Z.; Stoica, B.A.; Loane, D.J.; Wu, J.; Borroto, C.; Dorsey, S.G.; Faden, A.I. Downregulation of miR-23a and miR-27a following Experimental Traumatic Brain Injury Induces Neuronal Cell Death through Activation of Proapoptotic Bcl-2 Proteins. J. Neurosci. 2014, 34, 10055–10071. [Google Scholar] [CrossRef] [PubMed]
- Raoof, R.; Bauer, S.; El Naggar, H.; Connolly, N.; Brennan, G.P.; Brindley, E.; Hill, T.; McArdle, H.; Spain, E.; Forster, R.J.; et al. Dual-center, dual-platform microRNA profiling identifies potential plasma biomarkers of adult temporal lobe epilepsy. EBioMedicine 2018, 38, 127–141. [Google Scholar] [CrossRef] [PubMed]
- Yasmeen, S.; Kaur, S.; Mirza, A.H.; Brodin, B.; Pociot, F.; Kruuse, C. miRNA-27a-3p and miRNA-222-3p as Novel Modulators of Phosphodiesterase 3a (PDE3A) in Cerebral Microvascular Endothelial Cells. Mol. Neurobiol. 2019, 56, 5304–5314. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, A.; Volsko, C.; Garcia, J.P.; Agirre, E.; Allan, K.C.; Tesar, P.; Trapp, B.D.; Castelo-Branco, G.; Sim, F.J.; Dutta, R. Oligodendrocyte Intrinsic miR-27a Controls Myelination and Remyelination. Cell Rep. 2019, 29, 904–919.e9. [Google Scholar] [CrossRef]
- Abdel, R.H.; Kholoussi, N.M.; Eissa, E.; El Nady, H.G.; Fayed, D.B.; Abdelkawy, R.F.M. MicroRNAs as Immune Regulators of Inflammation in Children with Epilepsy. Int. J. Mol. Cell Med. 2020, 9, 188–197. [Google Scholar] [CrossRef]
- Liu, D.-Z.; Tian, Y.; Ander, B.P.; Xu, H.; Stamova, B.S.; Zhan, X.; Turner, R.; Jickling, G.; Sharp, F.R. Brain and blood microRNA expression profiling of ischemic stroke, intracerebral hemorrhage, and kainate seizures. J. Cereb. Blood Flow Metab. 2010, 30, 92–101. [Google Scholar] [CrossRef]
- Liu, Q.; Wang, L.; Yan, G.; Zhang, W.; Huan, Z.; Li, J. MiR-125a-5p Alleviates Dysfunction and Inflammation of Pentylenetetrazol- induced Epilepsy Through Targeting Calmodulin-dependent Protein Kinase IV (CAMK4). Curr. Neurovascular Res. 2019, 16, 365–372. [Google Scholar] [CrossRef]
- Tiedt, S.; Prestel, M.; Malik, R.; Schieferdecker, N.; Duering, M.; Kautzky, V.; Stoycheva, I.; Böck, J.; Northoff, B.; Klein, M.; et al. RNA-Seq Identifies Circulating miR-125a-5p, miR-125b-5p, and miR-143-3p as Potential Biomarkers for Acute Ischemic Stroke. Circ. Res. 2017, 121, 970–980. [Google Scholar] [CrossRef] [PubMed]
- Pan, Q.; Ma, C.; Wang, Y.; Wang, J.; Zheng, J.; Du, D.; Liao, X.; Chen, Y.; Chen, Y.; Bihl, J.; et al. Microvesicles-mediated communication between endothelial cells modulates, endothelial survival, and angiogenic function via transferring of miR-125a-5p. J. Cell. Biochem. 2019, 120, 3160–3172. [Google Scholar] [CrossRef] [PubMed]
- Pan, Q.; Liao, X.; Liu, H.; Wang, Y.; Chen, Y.; Zhao, B.; Lazartigues, E.; Yang, Y.; Ma, X. MicroRNA-125a-5p alleviates the deleterious effects of ox-LDL on multiple functions of human brain microvessel endothelial cells. Am. J. Physiol. Cell Physiol. 2017, 312, C119–C130. [Google Scholar] [CrossRef]
- Koskimäki, J.; Zhang, D.; Li, Y.; Saadat, L.; Moore, T.; Lightle, R.; Polster, S.; Carrión-Penagos, J.; Lyne, S.B.; Zeineddine, H.A.; et al. Transcriptome clarifies mechanisms of lesion genesis versus progression in models of Ccm3 cerebral cavernous malformations. Acta Neuropathol. Commun. 2019, 7, 132. [Google Scholar] [CrossRef]
- Wei, T.; Song, J.; Xu, M.; Lv, L.; Liu, C.; Shen, J.; Huang, Y. NEURL rs6584555 and CAND2 rs4642101 contribute to postoperative atrial fibrillation: A prospective study among Chinese population. Oncotarget 2016, 7, 42617–42624. [Google Scholar] [CrossRef]
- Shiraishi, S.; Zhou, C.; Aoki, T.; Sato, N.; Chiba, T.; Tanaka, K.; Yoshida, S.; Nabeshima, Y.; Nabeshima, Y.-I.; Tamura, T.-A. TBP-interacting Protein 120B (TIP120B)/Cullin-associated and Neddylation-dissociated 2 (CAND2) Inhibits SCF-dependent Ubiquitination of Myogenin and Accelerates Myogenic Differentiation. J. Biol. Chem. 2007, 282, 9017–9028. [Google Scholar] [CrossRef]
- Goldenberg, S.J.; Cascio, T.C.; Shumway, S.D.; Garbutt, K.C.; Liu, J.; Xiong, Y.; Zheng, N. Structure of the Cand1-Cul1-Roc1 Complex Reveals Regulatory Mechanisms for the Assembly of the Multisubunit Cullin-Dependent Ubiquitin Ligases. Cell 2004, 119, 517–528. [Google Scholar] [CrossRef] [PubMed]
- Sakaue, T.; Sakakibara, I.; Uesugi, T.; Fujisaki, A.; Nakashiro, K.-I.; Hamakawa, H.; Kubota, E.; Joh, T.; Imai, Y.; Izutani, H.; et al. The CUL3-SPOP-DAXX axis is a novel regulator of VEGFR2 expression in vascular endothelial cells. Sci. Rep. 2017, 7, 42845. [Google Scholar] [CrossRef]
- Sakaue, T.; Fujisaki, A.; Nakayama, H.; Maekawa, M.; Hiyoshi, H.; Kubota, E.; Joh, T.; Izutani, H.; Higashiyama, S. Neddylated Cullin 3 is required for vascular endothelial-cadherin-mediated endothelial barrier function. Cancer Sci. 2017, 108, 208–215. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.-W.; Lo, H.-H.; Chiu, Y.-L.; Chang, S.-J.; Huang, P.-H.; Liao, K.-H.; Tasi, C.-F.; Wu, C.-H.; Tsai, T.-N.; Cheng, C.-C.; et al. Dysregulated miR-361-5p/VEGF Axis in the Plasma and Endothelial Progenitor Cells of Patients with Coronary Artery Disease. PLoS ONE 2014, 9, e98070. [Google Scholar] [CrossRef]
- Hemond, C.C.; Healy, B.C.; Tauhid, S.; Mazzola, M.A.; Quintana, F.J.; Gandhi, R.; Weiner, H.L.; Bakshi, R. MRI phenotypes in MS. Neurol. Neuroimmunol. Neuroinflamm. 2019, 6, e530. [Google Scholar] [CrossRef]
- Wang, F.; Long, G.; Zhao, C.; Li, H.; Chaugai, S.; Wang, Y.; Chen, C.; Wang, D.W. Atherosclerosis-Related Circulating miRNAs as Novel and Sensitive Predictors for Acute Myocardial Infarction. PLoS ONE 2014, 9, e105734. [Google Scholar] [CrossRef] [PubMed]
- Gonçalves, T.F.; Piergiorge, R.M.; Dos Santos, J.M.; Gusmao, J.; Pimentel, M.; Santos-Rebouças, C.B. Network Profiling of Brain-Expressed X-Chromosomal MicroRNA Genes Implicates Shared Key MicroRNAs in Intellectual Disability. J. Mol. Neurosci. 2019, 67, 295–304. [Google Scholar] [CrossRef]
- Lulli, V.; Buccarelli, M.; Ilari, R.; Castellani, G.; De Dominicis, C.; Di Giamberardino, A.; D′alessandris, Q.G.; Giannetti, S.; Martini, M.; Stumpo, V.; et al. Mir-370-3p Impairs Glioblastoma Stem-Like Cell Malignancy Regulating a Complex Interplay between HMGA2/HIF1A and the Oncogenic Long Non-Coding RNA (lncRNA) NEAT1. Int. J. Mol. Sci. 2020, 21, 3610. [Google Scholar] [CrossRef] [PubMed]
- Peng, Z.; Wu, T.; Li, Y.; Xu, Z.; Zhang, S.; Liu, B.; Chen, Q.; Tian, D. MicroRNA-370-3p inhibits human glioma cell proliferation and induces cell cycle arrest by directly targeting β-catenin. Brain Res. 2016, 1644, 53–61. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zong, J.; Zhao, C. lncRNA CTBP1-AS2 promotes proliferation and migration of glioma by modulating miR-370-3p–Wnt7a-mediated epithelial–mesenchymal transition. Biochem. Cell Biol. 2020, 98, 661–668. [Google Scholar] [CrossRef] [PubMed]
- Visitchanakun, P.; Tangtanatakul, P.; Trithiphen, O.; Soonthornchai, W.; Wongphoom, J.; Tachaboon, S.; Srisawat, N.; Leelahavanichkul, A. Plasma miR-370-3P as a Biomarker of Sepsis-Associated Encephalopathy, the Transcriptomic Profiling Analysis of Microrna-Arrays from Mouse Brains. Shock 2019, 54, 347–357. [Google Scholar] [CrossRef]
- Ebrahimkhani, S.; Vafaee, F.; Young, P.E.; Hur, S.S.J.; Hawke, S.; Devenney, E.; Beadnall, H.; Barnett, M.H.; Suter, C.M.; Buckland, M.E. Exosomal microRNA signatures in multiple sclerosis reflect disease status. Sci. Rep. 2017, 7, 14293. [Google Scholar] [CrossRef]
- Plata-Bello, J.; Fariña-Jerónimo, H.; Betancor, I.; Salido, E. High Expression of FOXP2 Is Associated with Worse Prognosis in Glioblastoma. World Neurosurg. 2021, in press. [Google Scholar] [CrossRef]
- RahulAgrawal, R.; Pandey, P.; Jha, P.; Dwivedi, V.; Sarkar, C.; Kulshreshtha, R. Hypoxic signature of microRNAs in glioblastoma: Insights from small RNA deep sequencing. BMC Genom. 2014, 15, 1–16. [Google Scholar] [CrossRef]
- Piwecka, M.; Rolle, K.; Belter, A.; Barciszewska, A.M.; Żywicki, M.; Michalak, M.; Nowak, S.; Naskręt-Barciszewska, M.Z.; Barciszewski, J. Comprehensive analysis of microRNA expression profile in malignant glioma tissues. Mol. Oncol. 2015, 9, 1324–1340. [Google Scholar] [CrossRef] [PubMed]
- Fan, B.; Jiao, B.-H.; Fan, F.-S.; Lu, S.-K.; Song, J.; Guo, C.-Y.; Yang, J.-K.; Yang, L. Downregulation of miR-95-3p inhibits proliferation, and invasion promoting apoptosis of glioma cells by targeting CELF2. Int. J. Oncol. 2015, 47, 1025–1033. [Google Scholar] [CrossRef] [PubMed]
- Hwang, S.J.; Lee, H.W.; Kim, H.R.; Song, H.J.; Lee, D.H.; Lee, H.; Shin, C.H.; Joung, J.-G.; Kim, D.-H.; Joo, K.M.; et al. Overexpression of microRNA-95-3p suppresses brain metastasis of lung adenocarcinoma through downregulation of cyclin D1. Oncotarget 2015, 6, 20434–20448. [Google Scholar] [CrossRef] [PubMed]
| Author | miRNA | Location | Expression in BAVM | Putative Targets (BAVM) | Known Cerebral Pathologies Associated (Expression)—Respective Putative Target(s) |
|---|---|---|---|---|---|
| Chen et al., 2018 [18] | miR-7-5p | Peripheral blood | Upregulated | VEGF | Glioma, glioblastoma (downregulated)—RAF Ischemia-reperfusion injury Hemorrhagic stroke (downregulated)—PI3K/AKT pathway |
| Ferreira et al., 2014 [19] | miR-18a | BAVM BEC | Downregulated | Id-1, TSP-1 (indirect) | Pediatric medulloblastoma, ependymoma, astrocytoma (upregulated)—RUNX1 Glioblastoma (upregulated)—neogenin Moyamoya-like vasculopathy (downregulated) |
| Marin-Ramos et al., 2020 [46] | PAI-1, BMP4, HIF-1A | ||||
| Huang et al., 2017 [20] | miR-137 | VSMC | Downregulated | Akt, p38MAPK, PI3K, ERK, VEGF, NFkB | Ischemic stroke—lncRNA GAS5//JAK1 and STAT 1//Src and MAPK Schizophrenia (upregulated) Gliomas (downregulated)—Rac1 or Cox-2 Oligodendrogliomas (downregulated)—CSE1L |
| Huang et al., 2017 [20] | miR-195* | VSMC | Downregulated | Akt, p38MAPK, PI3K, ERK, VEGF, NFkB | Ischemic stroke—VEGFA schizophrenia (upregulated)—BDNF Alzheimer’s disease (downregulated)—BACE1//KLF5 signalling//NFkB Gliomas (downregulated) Malignant meningioma (downregulated)—FASN |
| Chen et al., 2018 [18] | miR-199a-5p | Peripheral blood | Upregulated | VEGF | Glioma (upregulated)—MAGT1//MARCH8 Testicular germ cell tumors (downregulated) Ischemic stroke (upregulated)—CAV-1 mediated MEK/ERK pathway//DDR1//ECE1 Hemangioma—HIF-1A |
| Chen et al., 2018 [18] | miR-200b-3p | Peripheral blood | Upregulated | VEGF | Ischemic stroke (upregulated) Medulloblastoma (upregulated) Gastric adenocrcinoma metastasis (upregulated)—ZEB2 Glioma (downregulated)—ERK5 Alzheimer’s disease mouse models (downregulated)—APP Cytomegalus infection (upregulated) |
| Chen et al., 2018 [18] | let-7b-5p | Peripheral blood | Upregulated | NS | CCM (downregulated) Epilepsy (downregulated) Ischemic stroke models (downregulated) Glioma (downregulated)—IKBKE, E2F2, oncogenes KRAS, HMGA2, and MYC |
| Author | miRNA | Location | Expression in CCM | Putative Targets | Known Cerebral Pathologies Associated (Expression)—Respective Putative Target(s) |
|---|---|---|---|---|---|
| Li et al., 2020 [105] | miR-27a | BEC (mouse model) | Upregulated | VE-cadherin | Temporal lobe epilepsy (upregulated)—GLRA2 Neuroprotection in rat models after TBI—Bax, Noxa, and Puma//FoxO3a |
| Li et al., 2020 [105] | miR-125a | Downregulated | NS | Epilepsy—CAMK4 Ischemic stroke | |
| Koskimäki et al., 2019 [121] | mmu-miR-3472a | Mouse NVU | Dysregulated | CAND2 | NS |
| Kar et al., 2017 [97] | miR-361-5p | Brainstem CCM | Downregulated | VEGF, EGFL7 | Acute myocardial infarction Ischemic stroke Pulmonary embolism Multiple sclerosis Intellectual disability |
| Kar et al., 2017 [97] | miR-370-3p | Downregulated | NF1 | Glioma (downregulated)—HMGA2, HIF-1A, NEAT1//CTBP1-AS2 Sepsis associated encepahlopathy in mouse models | |
| Kar et al., 2017 [97] | miR-181a-2-3p | Downregulated | NS | Glioblastoma (downregulated)—FOXP2 | |
| Kar et al., 2017 [97] | miR-95-3p | Downregulated | NS | Glioma (upregulated)—CELF2 Lung cell adenocarcinoma metastasis (downregulated)—cyclin D1 | |
| Kar et al., 2017 [97] | let-7b-5p | Downregulated | ENG, HES1, HIF-1A, MAPK1, MIB1, OCLN, PDCD10, TJP1 | BAVM (upregulated) Epilepsy (downregulated) Ischemic stroke models (downregulated) Glioma (downregulated)—IKBKE, E2F2, oncogenes KRAS, HMGA2 and MYC |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Florian, I.A.; Buruiana, A.; Timis, T.L.; Susman, S.; Florian, I.S.; Balasa, A.; Berindan-Neagoe, I. An Insight into the microRNAs Associated with Arteriovenous and Cavernous Malformations of the Brain. Cells 2021, 10, 1373. https://doi.org/10.3390/cells10061373
Florian IA, Buruiana A, Timis TL, Susman S, Florian IS, Balasa A, Berindan-Neagoe I. An Insight into the microRNAs Associated with Arteriovenous and Cavernous Malformations of the Brain. Cells. 2021; 10(6):1373. https://doi.org/10.3390/cells10061373
Chicago/Turabian StyleFlorian, Ioan Alexandru, Andrei Buruiana, Teodora Larisa Timis, Sergiu Susman, Ioan Stefan Florian, Adrian Balasa, and Ioana Berindan-Neagoe. 2021. "An Insight into the microRNAs Associated with Arteriovenous and Cavernous Malformations of the Brain" Cells 10, no. 6: 1373. https://doi.org/10.3390/cells10061373
APA StyleFlorian, I. A., Buruiana, A., Timis, T. L., Susman, S., Florian, I. S., Balasa, A., & Berindan-Neagoe, I. (2021). An Insight into the microRNAs Associated with Arteriovenous and Cavernous Malformations of the Brain. Cells, 10(6), 1373. https://doi.org/10.3390/cells10061373






