Abstract
Root architectural traits at the seedling stage have been demonstrated to be crucial for the efficient uptake of nutrients and drought tolerance in wheat. To dissect the genetic basis of these traits from the D genome, 182 recombinant inbred lines (RILs) derived from the common wheat TAA10 crossed with resynthesized allohexaploid wheat XX329 possessed similar AABB genomes were used for QTL mapping of five root traits in hydroponic-cultured seedlings, including lateral root number (LRN), seminal root number (SRN), root hair length (RHL), root diameter (RD), and total root volume (TRV). A total of seven QTLs were identified for the five root traits, with six possible novel QTLs for LRN, RHL, RD and TRV, accounting for 4.98–12.17% of phenotypic variation. One QTL (QLrn.qau-5D.2), controlling lateral root number, was fine mapped an approximate 5.0-Mb interval harboring 80 annotated genes, including five auxin-related genes. We further validated that QLrn.qau-5D.2 in NILTAA10 significantly enhanced yield-related traits, such as plant height, spike length, spike compactness, tiller number per plant and grain yield per plant, as comparison with NILXX329. Collectively, these results provide vital insights for fine-mapping QTLs associated with LRN, SRN, RHL, RD and TRV and facilitate the root morphologic designs for enhancing yield performance.
1. Introduction
Bread wheat (Triticum aestivum L.) is one of the earliest crop species to undergo domestication about 10,000 years ago in a core area of the Fertile Crescent to global environments, serving as more than 20% of human dietary calories and proteins [1,2]. Root system architecture (RSA, the spatial and temporal colonization of root types), encompassing the number, length, orientation, and branching of several root classes, plays a pivotal role in both nutrients and water absorption, soil anchorage, nutrient storage and rhizosphere microbial interaction, and serves as the primary plant organ responding to environmental abiotic stresses, such as drought, salinity, waterlogging, heavy metal toxicity and nutrient deficiencies [3,4,5]. Consequently, RSA directly influences the development and production of wheat [5,6]. The domestication and subsequent evolutionary process of wheat has led to a great deal of phenotypic variability in root systems morphology, such as longer root hair, more lateral root number and seminal root. These changes contribute to enhanced agronomic performance and a wider range of adaptability [7,8,9]. Historically, the primary emphasis of wheat selection and breeding programs has been on above-ground traits and yield. However, recent advancements in high-throughput root phenotyping and marker-assisted selections have made it increasingly feasible to genetically improve root system architecture in wheat breeding [10,11,12]. As such, the identification and use of the crucial genes or quantitative trait loci (QTL) that control root architectural traits are vital to expedite molecular improvements in wheat breeding program.
The root architectural traits of wheat which are controlled by complexes of polygenes with quantitative effects. Measuring and assessing them is challenging due to their susceptibility to environmental factors in field settings [13,14]. Traditionally, root architecture is described by multiple root parameters, such as root length, root number, root hair length, root diameter, root surface area, and root volume [15,16,17]. Previous researches have identified hundreds of QTL associated with wheat RSA traits in variable mapping populations with various types of markers, and these findings are summarized in Soriano and Alvaro (2019). Among them, several studies have indicated that QTLs of wheat root traits overlapped with various previously reported QTLs for nutrient uptake, drought tolerance and production in wheat [16,18,19]. However, little is known about the underlying genes controlling root traits in wheat, except two formally identified genes controlling wheat RSA traits through the forward-genetics approach. The first one was VERNALIZATION1 (VRN1) gene, a key regulator of flowering behavior in cereals, which has been demonstrated to modulate root angle at all growth stages of wheat [20]. More recently, the ENHANCED GRAVITROPISM 2 (EGT2) gene, an evolutionary conserved regulator of root growth angle in barley and wheat, is proposed to be a valuable target for root-based crop improvement [21]. Meanwhile, a few genes controlling root traits in wheat have been discovered by reverse genetics approaches, focusing on wheat orthologues of genes known to affect RSA in other species, such as TaWRKY51 [22] and TaLBD16 [8] regulating lateral root formation, TaRSL4 [9] and TaRSL2 [23] associated with the elongation of root hair, the LOB family member TaMOR [24] controlling number of roots, and transcription factors TaNAC2-5A, TaNAC69-1, and TaRNAC1 related to increased root length [24,25,26]. Hence, pinpointing QTLs/genes controlling root traits is of great interest in improving root function and contributing to increased wheat yields via these valuable genetic information and beneficial resources.
Bread wheat (2n = 6x = 42, AABBDD) is a typical allohexaploid species, derived from the hybridization of allotetraploid wheat (Triticum turgidum L., AABB) and diploid Aegilops tauschii (DD) [27,28]. Compared to the A and B sub-genomes, the D sub-genome exhibits limited genetic diversity in allohexaploid wheat, which is partly due to the involvement of only a few accessions of Ae. tauschii in the formation of allohexaploid wheat. This suggests that the D sub-genome holds significant potential for wheat improvement [29,30]. However, the genetic diversity of Ae. tauschii accessions has been found to be markedly higher than that of the D sub-genome from allohexaploid Triticum aestivum [31,32]. Thus, integrating alleles underlying beneficial traits from Ae. tauschii into allohexaploid wheat using synthetic wheat and its introgression lines might be an efficient strategy to expand D sub-genome diversity [33,34]. Specifically, extensive variation of seedling lateral root number observed among Ae. tauschii accessions is retained in synthetic allohexaploid wheat lines [8]. However, there have been relatively fewstudies focused on identifying beneficial QTL for root morphological traits from the diploid D donor of bread wheat. For instance, three QTL for root diameter were found to be associated with P-deficiency tolerance on chromosomes (Chr.) 1D, 3D and 7D, with desirable alleles contributed by synthetic allohexaploid wheat [35]. Positive alleles for two QTL controlling the fresh weight of roots and number of root tips associated with drought tolerance on Chr. 7D and 3D were derived from synthetic allohexaploid wheat [36]. Similarly, the synthetic allohexaploid line (XX329) used in the current study is comprised of A and B sub-genomes from the common allohexaploid bread wheat TAA10 (cv Canthach) and D sub-genome from the Ae. tauschii subspecies strangulate with accession number of TQ18 [37]. Since XX329 and TAA10 share almost identical AABB sub-genomes but have diversified D sub-genome, the phenotypic variations between them are primarily due to the different genetic origin of the D sub-genome [38,39]. A recombinant inbred lines (RILs) population derived from a cross between XX329 and TAA10 has been utilized to identify QTLs for various traits, including grain size and shape [38], spike morphological traits, plant height and heading date [40], as well as heat stress tolerance [41]. In this study, to unravel the underlying genetic basis on the variations of root traits between XX329 and TAA10, we conducted QTL analysis for lateral root number (LRN), seminal root number (SRN), root hair length (RHL), root diameter (RD), and total root volume (TRV) with this RILs population. Moreover, we developed near isogenic line (NIL) populations to validate the genetic effect of QLrn.qau-5D.2 and identified possible candidate genes within the targeted QTL region. This research can offer crucial insights into the genetic basis of root traits in wheat throughout its evolution and domestication, thus laying the groundwork for future targeted selection of root traits in wheat improvement and breeding programs.
2. Materials and Methods
2.1. Plant Materials
A population of 182 F8 RILs was used, following the protocol as described by Xu [40]. The RIL population was developed from the cross of common wheat TAA10 and resynthesized allohexaploid wheat XX329 with similar AABB sub-genomes using a single seed descent method. One line (RIL131) carrying heterozygous segment within the candidate interval of QLrn.qau-5D.2 were selected out from RIL population. It was then self-pollinated twice to the F10 generation. Afterwards, selfing of these plants was carried out to obtain nine sets of NIL pairs (L1–L9) with overlapping heterozygous fragments genotyped with QLrn.qau-5D.2 flanking markers. Each NIL set consisted of 30–107 TAA10-type homozygotes (NILTAA10) and 33–98 XX329-type homozygotes (NILXX329) to evaluating the effect of QLrn.qau-5D.2.
2.2. Root System Architecture Phenotyping
Seminal RSA traits except for root hair length were evaluated according to the protocol described by Cane [42] and were further modified in the present work. Seeds for each genotype were surface sterilized in 1% NaClO for 10 min, rinsed five times with distilled water and placed in Petri dishes with one sheet of germinating paper moistened with 5 mL of distilled water. The dishes were kept in the dark at 4 °C for 2 days and then cultured at 20 °C until germination. Seeds with uniform germination were cultivated for 2 days in a glasshouse under controlled conditions (55% humidity; 16-h light at 24 °C and 8-h dark at 18 °C). Next, the seedlings were transferred to a culture box with distilled water and were further cultured for an additional 8 days. The distilled water was refreshed every 2 days. Each genotype was performed with three biological replicates and each replicate comprised at least 8 plants. In addition, root hair phenotyping was conducted following the method described by Han [9]. In brief, seeds with uniform germination were sown on a medium containing 1% agar and were cultivated for 2 days in the above glasshouse. For each genotype, we performed three biological replicates, each of which consisted of at least five plants.
Seminal RSA traits were investigated on a single-plant basis: Lateral root number (LRN) on the primary root and seminal root number (SRN) were determined by an Epson Perfection LA2400 scanner (EPSON, Nagano, Japan) and analyzed using Adobe Photoshop CS5 (Adobe Systems Inc., San Jose, CA, USA). Root hair length (RHL) was recorded with a fluorescent stereomicroscope (SZX16; Olympus, Tokyo, Japan) attached to a digital imaging system (DP72; Olympus, Tokyo, Japan) and measured by Adobe PHOTOSHOP CS5 (Adobe Systems Inc., San Jose, CA, USA). Root diameter (RD) and total root volume (TRV) were measured using a WinRHIZO image analyzing system (Zeal quest Scientific Technology Co., Ltd., Shanghai, China). The phenotype for seminal RSA traits of RIL population was displayed in Table S1.
2.3. Field Evaluation of Yield-Related Traits
Phenotypic data for182 RILs as reported by Xu [40] including spike length (SL), fertile spikelet number per spike (FSN), sterile spikelet number per spike (SSN), total spikelet number per spike (TSN), spike compactness (SC), plant height (PH) and heading date (HD) were used to investigate the relationships between root traits and yield-related traits. The NIL pair (NIL-IITAA10 and NIL-IIXX329) from NIL pair L6 were planted in the field at Qingdao (36.59° N, 120.12° E) in Shandong province in China during the 2021–2022 growing season. The field trials were conducted in randomized complete block design with three replications. Each plot consisted of two rows that were 1.5 m long and 0.3 m apart with a sowing rate of 15 seeds per row. For each replicate, 6 to 10 representative plants or spikes were selected for phenotypic measurement of yield-related traits, i.e., PH, SL, FSN, SSN, TSN, SC, thousand grain weight (TGW), grain length (GL), grain width (GW), tiller number per plant (TN), and grain yield per plant (GYP).
2.4. QTL Analysis
The TAA10/XX329 genetic linkage map for the D genome used here has been reported by Xu [40]. For the analysis of QTLs associated with seminal RSA traits, we employed the composite interval mapping (CIM) method using Windows QTL Cartographer software version 2.5 [43]. In this method, model 6 (standard model) with forward and backward regression, five control markers as cofactors, a 10 cM scanning window size and a walk speed of 1 cM were determined as QTL detection parameters.
The empirical genome-wide LOD threshold values were estimated by conducting 1000 random permutation tests at a significance level of p ≤ 0.05. Confidence intervals were calculated based on position ±2 LOD away from the peak of likelihood ratios. All QTLs were named following the recommendations of McIntosh [44].
2.5. InDel Markers Development
Whole genome re-sequencing of TAA10 and XX329 was conducted to obtain a polymorphism data set [40]. Polymorphic nucleotide sequences between TAA10 and XX329 within the QTL interval were utilized to develop InDel markers in Primer3 v0.4.0, available at: http://bioinfo.ut.ee/primer3-0.4.0/ (accessed on 20 October 2020). The Polymerase Chain Reaction (PCR) amplification was performed in a volume of 10 μL including 5 μL, 2 × Taq PCR StarMix, 2 μL primer (mixture of forward and reverse primers, 2 μM), 1.5 μL DNA template and 1.5 μL sterilized ddH2O. The PCR program was carried out at 94 °C 5 min, followed by 32 cycles of 94 °C 30 s, 55–60 °C 30 s, 72 °C 30 s and 72 °C 5 min. The PCR products were separated using 2% agarose gel or 8% non-denatured polyacrylamide gel electrophoresis (PAGE) gels. The primer sequences of InDel markers used in this study were listed in the Supplementary Table S2.
2.6. Prediction of Candidate Genes
To predict candidate/flanking genes, the interval flanking marker sequences were blasted against the International Wheat Genome Sequencing Consortium and Ensembl Plants databases to determine the physical position with the highest identity of the identified QTL. To further obtain candidate genes among the target interval of QLrn.qau-7D.2, predicted genes of high-confidence were extracted from the JBrowse website, available at: https://urgi.versailles.inra.fr/jbrowseiwgsc/gmod_jbrowse/ (accessed on 6 April 2023). The gene expression pattern was determined in the Wheat eFP Browser, available at: http://bar.utoronto.ca/efp_wheat/cgi-bin/efpWeb.cgi (accessed on 15 April 2023). The whole-genome resequencing data of TAA10 and XX329 were used to retrieve the sequences of candidate genes, including the 2-kb upstream region of the translation start codon, exons, introns and the 2-kb downstream region of the translation termination codon.
2.7. Auxin Treatment
The seeds of NILTAA10 and NILXX329 from NIL pair L6 were surface sterilized with 1% NaClO for 10 min and were then incubated in Petri dishes at 4 °C for 2 d in the dark and exposed to white light for 1 d. The 2-day-old seedlings of NILTAA10 and NILXX329 were transferred to a culture box filled with different concentrations of 2,4-D (0.01, 0.1, 1, and 10 μM) and 1/5th strength Hoagland solution for 10 days in the greenhouse. Lateral root number on the primary root was measured as described previously Wang [8]. The experiment was repeated three times, with at least eight plants per treatment group.
2.8. Detection of Candidate Gene Expression
The 2-day-old seedlings of NILTAA10 and NILXX329 were transferred to a culture box filled with distilled water for 4 days in the greenhouse. The culture solution was changed every 2 days. Then, the 6-day-old uniform seedlings were treated with 0.1 μM IAA. Total RNA was extracted from wheat roots with SteadyPure Plant RNA Extraction Kit (Accurate Biotechnology, Changsha, China). The RNA samples were digested with purified DNase I, and first-strand cDNA synthesis was performed using HiScript II One-Step RT-PCR Kit (Vazyme Biotech, Nanjing, China). SYBR Color qPCR Master Mix (Vazyme Biotech, Nanjing, China) was used to perform reverse transcription qPCR (RT-qPCR) assays to detect gene transcript levels. The RT-qPCR conditions and analytical methods were the same as those described by Wang [8]. The wheat Actin gene was used as a standard control. The accession numbers of TaActin homoeologous genes were TraesCS1A02G274400, TraesCS1B02G283900, and TraesCS1D02G274400 (Wang [45]). Three independent biological replicates were applied to each sample for RT-qPCR. A description of the RT-qPCR primers is given in Table S2.
2.9. Statistical Analysis
Basic phenotypic statistical analyses, correlation coefficients among traits and Shapiro-Wilk test for testing departures from normal distribution were conducted using IBM SPSS Statistics 26.0 (SPSS, Chicago, IL, USA). The broad-sense heritability (HB2) based on the mean values of each trait among the three replications was calculated using IciMapping software v4.0 following the formula: HB2 = σg2/(σg2 + σe2/r), where σg2 is the genetic variance, σe2 is the residual variance, and r is the number of replications. In the progeny test, the significance of the phenotypic variation between the NIL pairs was determined by Student’s t test.
3. Results
3.1. Phenotypic Evaluation of Seminal Root Traits
The phenotypic performance and broad-sense heritability (HB2) of five seminal RSA traits (LRN, SRN, RHL, RD and TRV) from the RIL population and their parents are shown in Table 1. Notably, XX329 had more LRN, SRN, RD and TRV compared to TAA10, while TAA10 had longer RHL than XX329. Significant differences in root traits were observed among the RILs. Specifically, LRN ranged from 36.98 to 2.35 roots per plant with a mean value of 8.84, SRN varied from 4.22 to 6.00 roots per plant with an average of 5.18, RHL spanned from 0.53 to 1.41 mm with a mean value of 0.96 mm, RD was between 0.20 and 0.13 mm with a mean value of 0.17 mm, and TRV varied from 0.14 to 0.02 cm3 with a mean value of 0.05 cm3. The coefficient of variance (CV) of seminal RSA traits varied from 6.37% for SRN to 52.37% for LRN. The estimated HB2 of these five root traits ranged from 76.25 to 98.49%, among which LRN, SRN and RHL exhibited higher heritability (98.01, 91.39 and 98.49%, respectively) compared to RD and TRV, which were 76.25% and 84.48% respectively. The Shapiro-Wilk test for normality distribution showed that RD displayed normal distributions (p > 0.05), while the distribution of LRN, SRN, RHL and TRV exhibited deviations from normality (p < 0.05) (Figure 1). The frequency distribution histograms showed pronounced transgressive segregation for each trait, indicating that TAA10 and XX329 contributed positive alleles for these traits.

Table 1.
Two parental and RIL population means, ranges and broad-sense heritability for LRN, SRN, RHL, RD and TRV.
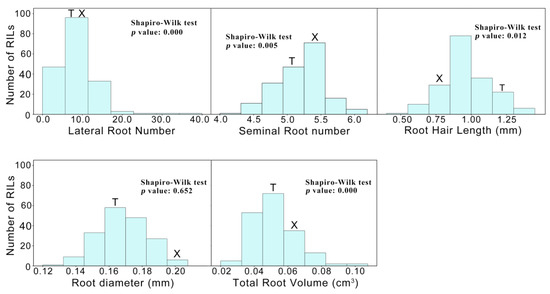
Figure 1.
The frequency distribution histograms of lateral root number (LRN), seminal root number (SRN), root hair length (RHL), root diameter (RD), and total root volume (TRV) for the TAA10 (T) × XX329 (X) recombinant inbred population. p < 0.05 indicates a significant departure from the normal distribution (Shapiro-Wilk test).
3.2. Correlation among Seminal Root Traits and Yield Components
The BLUP values of yield-related traits including SL, FSN, SSN, TSN, PHT, HD from the RIL population have been reported in Xu [40]. These values were utilized in the current study to calculate correlation coefficients between seminal root traits and yield components. Of all root traits, LRN exhibited a significantly positive correlation with HD (p < 0.05). SRN showed a positive correlation with FSN and TSN (p < 0.01) (Table 2). Additionally, RD and TRV were notably positively correlated with GL, GW and TGW (p < 0.05) (Table 2).

Table 2.
Phenotypic correlations seedling root traits with yield components in the TAA10/XX329 RIL population.
3.3. QTL Analysis
In this study, we detected seven QTL associated with LRN, SRN, RHL, RD and TRV on chromosomes 2D, 4D, 5D and 7D (Table 3). Of these identified QTL, three QTL associated with LRN (QLrn.qau-5D.1, QLrn.qau-5D.2 and QLrn.qau-7D) were identified on chromosome arms 5DS, 5DL and 7DS, respectively. QLrn.qau-5D.1 and QLrn.qau-5D.2 carried the favorable alleles contributed by TAA10, while QLrn.qau-7D had the favorable allele contributed by XX329. QLrn.qau-5D.1 had a LOD value of 2.55 and accounted for 4.98% of LRN variation. QLrn.qau-5D.2 had a LOD value of 5.78 and contributed to 12.17% of phenotypic variation. QLrn.qau-7D had a LOD value of 2.84 and accounted for 5.35% of LRN variation. For SRN, a major QTL (QSrn.qau-2D) with a LOD of 7.30 was detected on chromosome arm 2DS to control SRN. This QTL explained 14.73% of the phenotypic variance for SRN and was associated with positive alleles from XX329. For RHL, a major QTL located in a region on chromosome 4DL (QRhl.qau-4D) explained 9.32% of the observed RHL variation, with the positive allele contributed by TAA10. Additionally, a minor QTL (QRd.qau-2D) with LOD of 2.86 was detected on chromosome arm 2DS, controlling RD. This QTL explained 7.16% of the phenotypic variance for RD and was associated with positive alleles from XX329. A QTL located on chromosome 5DL (QTrv.qau-5D) with a LOD score of 2.95 explained 6.97% of phenotypic variation in TRV. The positive allele for QTrv.qau-5D was contributed by XX329.

Table 3.
The QTL regions associated with LRN, SRN, RHL, RD and TRV in the TAA10/XX329 RIL population.
3.4. Fine Mapping and Verification of QLrn.qau-5D.2
Residual heterozygous line (RHL) is an efficient tool for QTL fine-mapping without extensive backcrossing [46,47]. Fortunately, we successfully identified a RHL from the RIL population that carries a heterozygous segment in the target region of QLrn.qau-5D.2. Hence, we chose to focus on fine mapping the major QTL QLrn.qau-5D.2. The confidence interval of this QTL was determined to be between marker 5D432 and 5D483 (Figure 2A,B). To fine mapping QLrn.qau-5D.2, six new polymorphic InDel markers between TAA10 and XX329 were developed to genotype the segregating populations the segregating progenies of RHL (Table S2). Based on genotype of these markers, nine sets of NILs with overlapping recombinant segments were developed (L1–L9) (Figure 2C). The phenotyping analysis on LRN was applied among these homozygous non-recombinant lines. In the progeny test, NILTAA10 (with TAA10 allele) exhibited a significantly larger lateral root number than that of NILXX329 (with XX329 allele) within the populations L1, L2, L4, L6 and L8 (p < 0.01), while no significant differences were observed between NILTAA10 and NILXX329 within the populations L3, L5, L7 and L9. Taken together, QLrn.qau-5D.2 was precisely mapped to an approximate 5.0 Mb interval flanked by markers of 5D1204 and 5D122, corresponding to the genomic region from 386.3 Mb to 391.3 Mb in the IWGSC RefSeq v1.1. (Figure 2C).
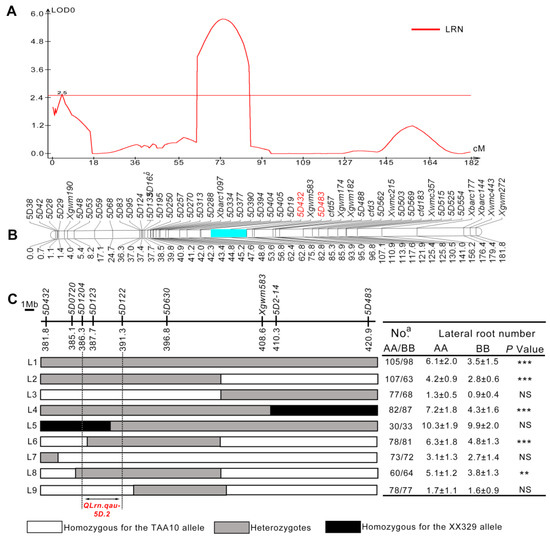
Figure 2.
Remapped QLrn.qau-5D used saturated genetic linkage map on chromosome 5D. (A) QTL mapping results for LRN on chromosome 5D. (B) Genetic map of chromosome 5D in the RIL population. The genetic region of QTL is labeled in blue and flanked by markers 5D432 and 5D483 in red. (C) Physical mapping for QLrn.qau-5D.2. The chromosomal locations of markers were defined according to Chinese Spring reference genome. Left side is the nine markers used to screen recombinants and the graphical genotypes of nine recombinants Arrows represent the 5.0 Mb interval of fine mapping. Right side is the comparisons of LRN between NILTAA10 (AA) and NILXX329 (BB) within each family. The values of LRN are the means (mean ± SD) of the homozygous plants in NIL families. a Represents the number of homozygous plants. White, gray and black bars represent AA (TAA10), heterozygous haplotypes and BB (XX329), respectively. Significant differences are indicated by ** (p < 0.01), *** (p < 0.001). “NS” indicates no significance (Student’s t test).
To further validate and assess the effect of QLrn.qau-5D.2 on LRN and yield component traits, we evaluated the effect of a set of NIL pair (L6) under field conditions. The results revealed that the average LRN of NILTAA10 was 33.33% higher than that of NILXX329 in the NIL pair derived from L6 (Figure 2C). In addition, an analysis of RIL lines possessing TAA10 or XX329 homozygous allele across the interval of QLrn.qau-5D.2 between markers 5D1204 and 5D122 revealed that the lines with the TAA10 allele exhibited a higher LRN than those with the TAA10 allele, which suggested that the 5D1204-5D122 interval contained a functional unit that controls LRN (Figure S1). To determine the possible pleiotropic effects of QLrn.qau-5D.2 associated with the effect on LRN, we measured a series of other yield-related traits in the set of NIL pair. Notably, NILTAA10 displayed significant increases in PHT, SL, SC, TN and GYP, with improvements of 5.7%, 9.0%, 12.0%, 23.96% and 24.44%, respectively (p < 0.05), when compared to NILXX329 in the L6-derived pair (Table S3).
3.5. Analysis of Candidate Genes
To identify the candidate genes in the 5.0-Mb interval of QLrn.qau-5D.2 (5D1204-5D122), we searched the annotated genes based on the gene annotations of the Chinese Spring reference genome (IWGSC RefSeq v1.1). This interval contained 80 high-confidence genes (Table S4). Utilizing the Wheat eFP Browser database, we analyzed 39 genes primarily expressed in root tissues (Table S5). It is well-established that Auxin plays a pivotal role in the formation of lateral and adventitious roots [48]. The orthologs of the above 39 genes in Arabidopsis and rice were analyzed by using Triticeae-GeneTribe (Table S6). Interestingly, five predicted genes (TraesCS5D02G286000, TraesCS5D02G286100, TraesCS5D02G288000, TraesCS5D02G291800 and TraesCS5D02G293100) have been identified to be associated with the auxin-related pathway, suggesting that they might be the candidate genes (Table S7). DNA sequence analysis uncovered SNP variants of TraesCS5D02G286000 between TAA10 and XX329. These variants were found in the upstream region of the translation start codon and the downstream region of the termination codon, respectively. For TraesCS5D02G286100, only one single InDel was found in the upstream region of translation start codon and downstream region of the termination codon between TAA10 and XX329, respectively. As for TraesCS5D02G288000, 6 SNPs were observed between two parent lines. In particular, one SNP in the exon 3 was found in XX329, as compared with TAA10, which led to the substitution of amino acid (Asn/Ile). For TraesCS5D02G291800, a total of 17 SNPs and 11 InDels were found between TAA10 and XX329. These variants included 3 SNPs in the upstream region of translation start codon, 1 SNP in the intron, as well as 13 SNPs and 11 InDels in the downstream region of the termination codon. For TraesCS5D02G293100, altogether 17 SNPs and 3 InDels were identified between two parents. Notably, 4 SNPs were located in the exon 1 and exon 3, resulting in 3 amino acid substitutions.
Given the missense mutations in TraesCS5D02G288000 and TraesCS5D02G293100, which involved in auxin-associated pathways, between TAA10 and XX329, we hypothesize that these genes may play a role in auxin-induced lateral root development. We first examined the expression patterns of these two candidate genes in roots of NILTAA10 and NILXX329 from one NIL pair of L6 under 0.1 μM IAA treatment. The results showed that the accumulation of TraesCS5D02G28800 mRNA continued to increase in NILTAA10 after 3 h of IAA treatment. In contrast, there was no dramatic change in the transcript level of this gene in NILXX329 following IAA treatment (Figure 3A). Compared with NILXX329, the transcript level of TraesCS5D02G28800 remained consistently higher in NILTAA10 throughout the 3 h to 12 h period of IAA treatment (Figure 3A). As for TraesCS5D02G293100 mRNA, there was a significant increase in the transcript level of this gene in NILTAA10 and NILXX329 after a 6-h IAA treatment (Figure 3B). Moreover, the relative expression level of TraesCS5D02G293100 was significantly higher under IAA treatment for different durations in NILTAA10 than that of NILXX329 (Figure 3B). Next, we examined how NILTAA10 and NILXX329 plants response to different concentrations of IAA treatments. There was a noticeable difference in lateral root number between NILTAA10 and NILXX329 in response to increasing IAA concentrations from 0.01 to 1 μM. The NILTAA10 plants exhibited more lateral root number compared to the NILXX329 plants (Figure 3C). These data indicated that TraesCS5D02G288000 and TraesCS5D02G293100 may be involved in the development of lateral roots, partially in an auxin-associated manner.
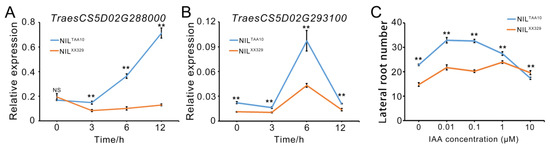
Figure 3.
Effects of TraesCS5D02G288000 and TraesCS5D02G293100 in NILTAA10 and NILXX329 from one NIL pair L6 in response to IAA treatment. Expression analysis of TraesCS5D02G288000 (A) and TraesCS5D02G293100 (B) by RT-qPCR in roots of NILTAA10 and NILXX329 under 0.1 μM IAA treatment. TaActin was used as internal control. (C) Comparison of lateral root number between NILTAA10 and NILXX329 with different concentrations of IAA at the seedling stage. The IAA concentrations were 0, 0.01, 0.1, 1, and 10 μM, respectively. Values are indicated as mean ± SD. Statistical differences between NILTAA10 and NILXX329 are indicated by ** (p < 0.01). “NS” indicates no significance (Student’s t test).
4. Discussion
4.1. Novel QTLs for Root-Related Traits in the D Genome of Wheat
Allohexaploid common wheat is a product of hybridization between Triticum turgidum and a single lineage of Ae. tauschii only once or at best a few times [32,49]. Possibly because this, the genetic diversity of the D genome is far less than that of the A and B genomes [50,51]. Synthetic allohexaploid wheats (AABBDD) provide promising resources to widen the genetic diversity associated with the D genome of Ae. tauschii [52]. To date, a number of genes/QTL controlling important agronomic traits have been found in the D genome of synthetic allohexaploid wheat, such as pest and disease resistance, stress tolerance, quality and yield [33,53,54,55]. Plant roots are important organs in determining grain productivity. Early vigorous growth of roots can improve the efficient acquisition of water and nutrients, and is closely related to plant adaptation and yield [56,57]. Although root traits in the field are difficult to be directly selected by wheat breeders, identification of QTLs for root traits and development of marker-assisted selection (MAS) can help breeders to select wheat root traits desirable for improving wheat water-nutrient use efficiencies [58,59,60]. Especially, some QTLs for root characteristics were co-located with loci affecting plant height, grain-related traits or drought tolerance [36,61,62]. Thus, the loci for root traits could be selected to enhance the potential and stability of yield and drought tolerance by MAS.
Since the genome (AABB) of the resynthesized allohexaploid wheat XX329 is near-identical to that of the allohexaploid common wheat TAA10 [38], the observed phenotypic variation of root-related traits in this study could primarily be attributed to differences on the D genome between TAA10 and XX329. Here, we identified 7 QTL for root traits (LRN, SRN, RHL, RD and TRV) using the TAA10/XX329 RIL population, and these QTLs were primarily located on chromosomes 2D, 4D, 5D and 7D (Table 3). Previous studies showed that QTL for root traits (LRN, SRN, RHL, RD and TRV) have been detected on various chromosomes [63,64]. To the best of our knowledge, only one of the seven QTLs (QTrv.qau-5D) coincided with that of a previously reported QTL controlling root volume [65], and the remaining six QTLs identified in this study, namely QLrn.qau-5D.1, QLrn.qau-5D.2, QLrn.qau-7D, QSrn.qau-2D, QRhl.qau-4D and QRd.qau-2D, appear to be novel. These findings provide valuable insights into the underlying mechanisms of root traits [62]. Among these QTLs, three novel ones (QLrn.qau-5D.1, QLrn.qau-5D.2 and QRhl.qau-4D) had positive alleles originating from TAA10, while QLrn.qau-7D, QSrn.qau-2D, and QRd.qau-2D were also novel QTLs with the increasing allele from XX329 (Table 3). Similarly, in a study by [36], two novel QTLs for root fresh weight and root diameter, along with six novel QTLs for root fresh weight, the ratio of root water loss, total root surface area, number of root tips, and number of root forks were detected in a synthetic allohexaploid wheat line. Among these QTLs, four novel QTLs contributed by positive alleles from the synthetic allohexaploid wheat line were mapped to the D genome [36]. Therefore, it is necessary to further develop molecular markers linked to new QTL/genes controlling root traits obtained from Ae. tauschii, which can provide more desirable targets for improving wheat root traits through molecular breeding.
4.2. Candidate Genes of QLrn.qau-5D.2
To date, several wheat genes controlling lateral root number have been characterized using a homology-based approach, such as TaWRKY51, TaLBD16, TaSNAC8 and TaMOR [8,22,24,66]. To identify the candidate genes for LRN, we conducted an analysis of the annotated genes within the confidence interval of QLrn.qau-5D.2 which controls lateral root number. Given that many genetic studies highlighted the pivotal role of the plant hormone auxin in regulating lateral root development [67,68,69], this study focused on auxin-related genes that might be the candidate genes of QLrn.qau-5D.2. Based on functional annotations and transcriptome analyses of 80 high confidence candidate genes in the genetic interval of QLrn.qau-5D.2 (Table S4), this study identified five genes (TraesCS5D02G286000, TraesCS5D02G286100, TraesCS5D02G288000, TraesCS5D02G291800, and TraesCS5D02G293100 that have the potential to affect lateral root development. These genes encode proteins involved in the auxin-related pathway (Tables S4 and S6). Of these, two genes (TraesCS5D02G286000 and TraesCS5D02G286100) encode glycosylphosphatidylinositol tail anchored proteins related to root growth, which were both orthologs of Arabidopsis AUXIN INDUCED IN ROOTS12 (AtAIR12). In Arabidopsis, the auxin-responsive gene AtAIR12 plays a key role in regulating the formation of lateral roots [70,71]. Additionally, TraesCS5D02G288000 encodes a TRANSPORT INHIBITOR RESISTANT 1 (TIR1)/AUXIN SIGNALLING F-BOX (AFB) protein. The sequence variation of TaTIR1/AFB (TraesCS5D02G288000) including one SNP in the exon and 4 SNPs in the upstream region of the translation start codon was detected between two parents. Notably, the SNP in the exon caused an amino acid substitution (Asn/Ile). Recent studies have demonstrated that the TIR1/AFB proteins are essential in very rapid auxin responses in root growth, such as lateral root formation [72,73]. In rice, downregulation of the auxin receptor genes OsTIR1 and OsAFB2 by overexpression of OsMIR393 affects plant height, tillering, grain yield, and primary and crown root growth [74,75]. In wheat, the wheat auxin receptor TaTIR1 (TraesCS1A02G091300) negatively regulates defense against Fusarium graminearum [76], but the function of this gene (TraesCS5D02G288000) has not been reported in wheat. Interestingly, our results revealed that the positive allele of QLrn.qau-5D.2 increased not only lateral root number, but also plant height, tillering number and grain yield per plant (Table S3), suggesting that TraesCS5D02G288000 (TaTIR1/AFB) may play a similar role in wheat, root system, plant height, tillering, and grain yield. Analysis of another two candidate genes (TraesCS5D02G291800 and TraesCS5D02G293100) showed that both genes encode an ER-localized protein, and its homologous gene OsPIN5b in rice. OsPIN5b participates in auxin homeostasis and regulates plant architecture and yield. Specifically, overexpression of the OsPIN5b leads to reduced plant height, tiller number, yield, root length and root tips number. In contrast, reduced expression of OsPIN5b results in higher tiller number, more vigorous root system and increased yield [77]. Moreover, both TraesCS5D02G291800 and TraesCS5D02G293100 have several sequence variations between TAA10 and XX329, but only TraesCS5D02G293100 has 4 SNPs in exon that caused the substitution of 3 amino acids (Table S7). Notably, TraesCS5D02G288000 (CTIR1/AFB) and TraesCS5D02G293100 (TaPIN5b) have missense mutations between TAA10 and XX329 (Supplementary Table S7). The transcript level of TraesCS5D02G288000 (TaTIR1/AFB) continued to increase in NILTAA10 after 3 h IAA treatment, whereas no drastic change in NILXX329 was detected in the transcript level of this gene after IAA treatment (Figure 3A). The relative expression level of TraesCS5D02G288000 (TaTIR1/AFB) at peaks was significantly higher throughout the period of 3–12 h IAA treatment in NILTAA10 than that of NILXX329 (Figure 3A). TraesCS5D02G293100 (TaPIN5b) mRNA showed that there was a remarkable increase in the transcript level of this gene in NILTAA10 and NILXX329 after a 6-h IAA treatment, respectively (Figure 3B). The transcript level of this gene was significantly higher under IAA treatment for different durations in NILTAA10 than that of NILXX329 (Figure 3B). Interestingly, there was a noticeable difference in lateral root number between NILTAA10 and NILXX329 in response to increasing IAA concentrations from 0.01 to 1 μM, and The NILTAA10 plants exhibited more lateral root number compared to the NILXX329 plants (Figure 3C). This indicates that TraesCS5D02G288000 (TaTIR1/AFB) and TraesCS5D02G293100 (TaPIN5b) may be involved in lateral root development partly in an auxin-associated manner. More further experiments, such as more in-depth map, quantitative PCR, targeted gene sequencing, and genetic transformation of wheat, is needed to validate which one is the causal gene for QLrn.qau-5D.2.
5. Conclusions
In this study, we identified seven root QTLs including six novel loci for LRN, RHL, RD and TRV. One QTL (QLrn.qau-5D.2) for LRN on chromosome 5DL was fine mapped an approximate 5.0-Mb interval harboring 80 annotated genes. Field trials showed that the allele of QLrn.qau-5D.2 from TAA10 significantly increased enhanced yield-related traits, such as plant height (PH), spike length (SL), spike compactness (SC), tiller number per plant (TN) and grain yield per plant (GYP), as comparison to the allele from XX329. These QTLs facilitate the identification of candidate genes and root morphologic designs for enhancing yield performance.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/agronomy14030608/s1, Figure S1: Difference in LRN between two alleles within the genetic interval flanking by the molecular markers 5D1204 and 5D122 in the TAA10/XX329 RIL population; Table S1: Phenotypic data in the TAA10/XX329 recombinant inbred line (RIL) population; Table S2: Sequences of developed markers used in this study; Table S3: Statistical analysis for plant height (PH), spike length (SL), fertile spikelet number per spike (FSN), sterile spikelet number per spike (SSN), total spikelet number per spike (TSN), spike compactness (SC), grain length (GL), grain width (GW), thousand grain weight (TGW), tiller number per plant (TN) and grain yield per plant (GYP) between the two homozygous genotypes from NIL pairs L6; Table S4: Annotated genes in the interval between the molecular markers 5D1204 and 5D122; Table S5: Expression profile of 39 high confidence genes based on the wheat expression atlas; Table S6: The orthologs of 39 genes in Arabidopsis and rice; Table S7: Sequence variation analysis of the candidate genes.
Author Contributions
H.W. participated in writing the first draft. B.Y., X.Z., H.C., F.L., Y.R., X.W. and X.F. carried out measurements of the phenotypes and data analysis. W.G., X.L. and N.G. revised the manuscript. Y.Z. designed the research, wrote and extensively revised this manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by grants from the Key Research and Development Project of Shandong Province (2021LZGC025, 2023LZGC002 and 2021LZG013), and the Foundation for High-level Talents of Qingdao Agriculture University (6631119057).
Data Availability Statement
The original contribution presented in the study are publicly available.
Acknowledgments
We are grateful to Zhongfu Ni providing the recombinant inbred line population and genotyping, Shandong Provincial Key Laboratory of Dryland Farming Technology, and Shandong Engineering Research Center of Germplasm Innovation and Utilization of Salt-Tolerant Crops for their technical support for this research.
Conflicts of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
- Salamini, F.; Özkan, H.; Brandolini, A.; Schäfer-Pregl, R.; Martin, W. Genetics and geography of wild cereal domestication in the near east. Nat. Rev. Genet. 2002, 3, 429–441. [Google Scholar] [CrossRef]
- Zhou, Y.; Zhao, X.; Li, Y.; Xu, J.; Bi, A.; Kang, L.; Xu, D.; Chen, H.; Wang, Y.; Wang, Y.-g.; et al. Triticum population sequencing provides insights into wheat adaptation. Nat. Genet. 2020, 52, 1412–1422. [Google Scholar] [PubMed]
- Trinidad, J.L.; Grajo, H.L.; Abucay, J.B.; Kohli, A. Cereal root proteomics for complementing the mechanistic understanding of plant abiotic stress tolerance. Agric. Proteom. 2016, 2, 19–51. [Google Scholar]
- Atkinson, J.A.; Wingen, L.U.; Griffiths, M.; Pound, M.P.; Gaju, O.; Foulkes, M.J.; Le Gouis, J.; Griffiths, S.; Bennett, M.J.; King, J.; et al. Phenotyping pipeline reveals major seedling root growth QTL in hexaploid wheat. J. Exp. Bot. 2015, 66, 2283–2292. [Google Scholar] [CrossRef] [PubMed]
- Manschadi, A.; Manske, G.; Vlek, P. Root architecture and resource acquisition: Wheat as a model plant. In Plant Roots-The Hidden Half; Routledge: London, UK, 2013; pp. 1–22. [Google Scholar]
- Bai, C.; Ge, Y.; Ashton, R.; Evans, J.; Milne, A.; Hawkesford, M.; Whalley, W.; Parry, M.; Melichar, J.; Feuerhelm, D.; et al. The relationships between seedling root screens, root growth in the field and grain yield for wheat. Plant Soil 2019, 440, 311–326. [Google Scholar] [CrossRef]
- Golan, G.; Hendel, E.; Méndez Espitia, G.E.; Schwartz, N.; Peleg, Z. Activation of seminal root primordia during wheat domestication reveals underlying mechanisms of plant resilience. Plant Cell Environ. 2018, 41, 755–766. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Hu, Z.; Huang, K.; Han, Y.; Zhao, A.; Han, H.; Song, L.; Fan, C.; Li, R.; Xin, M.; et al. Three genomes differentially contribute to the seedling lateral root number in allohexaploid wheat: Evidence from phenotype evolution and gene expression. Plant J. 2018, 95, 976–987. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Xin, M.; Huang, K.; Xu, Y.; Liu, Z.; Hu, Z.; Yao, Y.; Peng, H.; Ni, Z.; Sun, Q. Altered expression of TaRSL4 gene by genome interplay shapes root hair length in allopolyploid wheat. New Phytol. 2016, 209, 721–732. [Google Scholar] [CrossRef] [PubMed]
- Munns, R.; James, R.A.; Xu, B.; Athman, A.; Conn, S.J.; Jordans, C.; Byrt, C.S.; Hare, R.A.; Tyerman, S.D.; Tester, M.; et al. Wheat grain yield on saline soils is improved by an ancestral Na+ transporter gene. Nat. Biotechnol. 2012, 30, 360–364. [Google Scholar] [CrossRef]
- Wasson, A.P.; Nagel, K.A.; Tracy, S.; Watt, M. Beyond digging: Noninvasive root and rhizosphere phenotyping. Trends Plant Sci. 2020, 25, 119–120. [Google Scholar] [CrossRef]
- Steele, K.; Price, A.; Witcombe, J.; Shrestha, R.; Singh, B.; Gibbons, J.; Virk, D. QTLs associated with root traits increase yield in upland rice when transferred through marker-assisted selection. Theor. Appl. Genet. 2013, 126, 101–108. [Google Scholar] [CrossRef]
- Malamy, J. Intrinsic and environmental response pathways that regulate root system architecture. Plant Cell Environ. 2005, 28, 67–77. [Google Scholar] [CrossRef]
- Wasson, A.P.; Richards, R.; Chatrath, R.; Misra, S.; Prasad, S.S.; Rebetzke, G.; Kirkegaard, J.; Christopher, J.; Watt, M. Traits and selection strategies to improve root systems and water uptake in water-limited wheat crops. J. Exp. Bot. 2012, 63, 3485–3498. [Google Scholar] [CrossRef] [PubMed]
- Maccaferri, M.; El-Feki, W.; Nazemi, G.; Salvi, S.; Canè, M.A.; Colalongo, M.C.; Stefanelli, S.; Tuberosa, R. Prioritizing quantitative trait loci for root system architecture in tetraploid wheat. J. Exp. Bot. 2016, 67, 1161–1178. [Google Scholar] [CrossRef] [PubMed]
- Manschadi, A.M.; Hammer, G.L.; Christopher, J.T.; Devoil, P. Genotypic variation in seedling root architectural traits and implications for drought adaptation in wheat (Triticum aestivum L.). Plant Soil 2008, 303, 115–129. [Google Scholar] [CrossRef]
- Chen, X.; Ding, Q.; Błaszkiewicz, Z.; Sun, J.; Sun, Q.; He, R.; Li, Y. Phenotyping for the dynamics of field wheat root system architecture. Sci. Rep. 2017, 7, 37649. [Google Scholar] [CrossRef] [PubMed]
- Xie, Q.; Fernando, K.M.; Mayes, S.; Sparkes, D.L. Identifying seedling root architectural traits associated with yield and yield components in wheat. Ann. Bot. 2017, 119, 1115–1129. [Google Scholar] [CrossRef] [PubMed]
- An, D.; Su, J.; Liu, Q.; Zhu, Y.; Tong, Y.; Li, J.; Jing, R.; Li, B.; Li, Z. Mapping QTLs for nitrogen uptake in relation to the early growth of wheat (Triticum aestivum L.). Plant Soil 2006, 284, 73–84. [Google Scholar] [CrossRef]
- Voss-Fels, K.P.; Robinson, H.; Mudge, S.R.; Richard, C.; Newman, S.; Wittkop, B.; Stahl, A.; Friedt, W.; Frisch, M.; Gabur, I.; et al. VERNALIZATION1 modulates root system architecture in wheat and barley. Mol. Plant 2018, 11, 226–229. [Google Scholar] [CrossRef] [PubMed]
- Kirschner, G.K.; Rosignoli, S.; Guo, L.; Vardanega, I.; Imani, J.; Altmüller, J.; Milner, S.G.; Balzano, R.; Nagel, K.A.; Pflugfelder, D.; et al. ENHANCED GRAVITROPISM 2 encodes a STERILE ALPHA MOTIF–containing protein that controls root growth angle in barley and wheat. Proc. Natl. Acad. Sci. USA 2021, 118, e2101526118. [Google Scholar] [CrossRef]
- Hu, Z.; Wang, R.; Zheng, M.; Liu, X.; Meng, F.; Wu, H.; Yao, Y.; Xin, M.; Peng, H.; Ni, Z. TaWRKY51 promotes lateral root formation through negative regulation of ethylene biosynthesis in wheat (Triticum aestivum L.). Plant J. 2018, 96, 372–388. [Google Scholar] [CrossRef] [PubMed]
- Han, H.; Wang, H.; Han, Y.; Hu, Z.; Xin, M.; Peng, H.; Yao, Y.; Sun, Q.; Ni, Z. Altered expression of the TaRSL2 gene contributed to variation in root hair length during allopolyploid wheat evolution. Planta 2017, 246, 1019–1028. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Liu, D.; Li, Q.; Mao, X.; Li, A.; Wang, J.; Chang, X.; Jing, R. Overexpression of wheat gene TaMOR improves root system architecture and grain yield in Oryza sativa. J. Exp. Bot. 2016, 67, 4155–4167. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Chai, S.; McIntyre, C.L.; Xue, G.P. Overexpression of a predominantly root-expressed NAC transcription factor in wheat roots enhances root length, biomass and drought tolerance. Plant Cell Rep. 2018, 37, 225–237. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Richardson, T.; Chai, S.; Lynne McIntyre, C.; Rae, A.L.; Xue, G.P. Drought-up-regulated TaNAC69-1 is a transcriptional repressor of TaSHY2 and TaIAA7, and enhances root length and biomass in wheat. Plant Cell Physiol. 2016, 57, 2076–2090. [Google Scholar] [CrossRef] [PubMed]
- Petersen, G.; Seberg, O.; Yde, M.; Berthelsen, K. Phylogenetic relationships of Triticum and Aegilops and evidence for the origin of the A, B, and D genomes of common wheat (Triticum aestivum). Mol. Phylogenetics Evol. 2006, 39, 70–82. [Google Scholar] [CrossRef] [PubMed]
- The International Wheat Genome Sequencing Consortium (IWGSC); Mayer, K.F.; Rogers, J.; Doležel, J.; Pozniak, C.; Eversole, K.; Feuillet, C.; Gill, B.; Friebe, B.; Lukaszewski, A.J. A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 2014, 345, 1251788. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Börner, A.; Röder, M.; Ganal, M. Assessing genetic diversity of wheat (Triticum aestivum L.) germplasm using microsatellite markers. Theor. Appl. Genet. 2002, 105, 699–707. [Google Scholar] [CrossRef]
- Hao, C.; Jiao, C.; Hou, J.; Li, T.; Liu, H.; Wang, Y.; Zheng, J.; Liu, H.; Bi, Z.; Xu, F.; et al. Resequencing of 145 landmark cultivars reveals asymmetric sub-genome selection and strong founder genotype effects on wheat breeding in China. Mol. Plant 2020, 13, 1733–1751. [Google Scholar] [CrossRef]
- Caldwell, K.S.; Dvorak, J.; Lagudah, E.S.; Akhunov, E.; Luo, M.-C.; Wolters, P.; Powell, W. Sequence polymorphism in polyploid wheat and their D-genome diploid ancestor. Genetics 2004, 167, 941–947. [Google Scholar] [CrossRef]
- Gaurav, K.; Arora, S.; Silva, P.; Sánchez-Martín, J.; Horsnell, R.; Gao, L.; Brar, G.S.; Widrig, V.; John Raupp, W.; Singh, N.; et al. Population genomic analysis of Aegilops tauschii identifies targets for bread wheat improvement. Nat. Biotechnol. 2022, 40, 422–431. [Google Scholar] [CrossRef]
- Okamoto, Y.; Nguyen, A.T.; Yoshioka, M.; Iehisa, J.C.; Takumi, S. Identification of quantitative trait loci controlling grain size and shape in the D genome of synthetic hexaploid wheat lines. Breed. Sci. 2013, 63, 423–429. [Google Scholar] [CrossRef]
- Gao, L.; Zhao, G.; Huang, D.; Jia, J. Candidate loci involved in domestication and improvement detected by a published 90K wheat SNP array. Sci. Rep. 2017, 7, 44530. [Google Scholar] [CrossRef]
- Wu, F.; Yang, X.; Wang, Z.; Deng, M.; Ma, J.; Chen, G.; Wei, Y.; Liu, Y. Identification of major quantitative trait loci for root diameter in synthetic hexaploid wheat under phosphorus-deficient conditions. J. Appl. Genet. 2017, 58, 437–447. [Google Scholar] [CrossRef] [PubMed]
- Liu, R.-x; Wu, F.-k; Xin, Y.; Yu, L.; Wang, Z.-q; Liu, S.-h; Mei, D.; Jian, M.; Wei, Y.-m; Zheng, Y.; et al. Quantitative trait loci analysis for root traits in synthetic hexaploid wheat under drought stress conditions. J. Integr. Agric. 2020, 19, 1947–1960. [Google Scholar] [CrossRef]
- Kerber, E. Wheat: Reconstitution of the tetraploid component (AABB) of hexaploids. Science 1964, 143, 253–255. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Liang, F.; Xu, H.; Zhang, X.; Zhai, H.; Sun, Q.; Ni, Z. Identification of QTL for grain size and shape on the D genome of natural and synthetic allohexaploid wheats with near-identical AABB genomes. Front. Plant Sci. 2017, 8, 1705. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhu, B.; Qi, B.; Gou, X.; Dong, Y.; Xu, C.; Zhang, B.; Huang, W.; Liu, C.; Wang, X. Evolution of the BBAA component of bread wheat during its history at the allohexaploid level. Plant Cell 2014, 26, 2761–2776. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Zhang, R.; Wang, M.; Li, L.; Yan, L.; Wang, Z.; Zhu, J.; Chen, X.; Zhao, A.; Su, Z.; et al. Identification and characterization of QTL for spike morphological traits, plant height and heading date derived from the D genome of natural and resynthetic allohexaploid wheat. Theor. Appl. Genet. 2022, 135, 389–403. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R.; Liu, G.; Xu, H.; Lou, H.; Zhai, S.; Chen, A.; Hao, S.; Xing, J.; Liu, J.; You, M.; et al. Heat Stress Tolerance 2 confers basal heat stress tolerance in allohexaploid wheat (Triticum aestivum L.). J. Exp. Bot. 2022, 73, 6600–6614. [Google Scholar] [CrossRef] [PubMed]
- Cane, M.A.; Maccaferri, M.; Nazemi, G.; Salvi, S.; Francia, R.; Colalongo, C.; Tuberosa, R. Association mapping for root architectural traits in durum wheat seedlings as related to agronomic performance. Mol. Breed. 2014, 34, 1629–1645. [Google Scholar] [CrossRef]
- Wang, S.; Basten, C.; Zeng, Z. Windows QTL Cartographer 2.5; Department of Statistics, North Carolina State University: Raleigh, NC, USA, 2010. [Google Scholar]
- Mcintosh, R.A.; Dubcovsky, J.; Rogers, W.J.; Morris, C.; Appels, R.; Xia, X. Catalogue of Gene Symbols for Wheat: 2017 Supplement. 2017. Available online: https://shigen.nig.ac.jp/wheat/komugi/genes/macgene/supplement2017.pdf (accessed on 5 March 2019).
- Wang, H.; Han, X.; Fu, X.; Chen, H.; Wei, X.; Cui, S.; Liu, Y.; Guo, W.; Li, X.; Xing, J.; et al. Overexpression of TaLBD16-4D alters plant architecture and heading date in transgenic wheat. Front. Plant Sci. 2022, 13, 911993. [Google Scholar] [CrossRef] [PubMed]
- Liu, N.; Liu, J.; Li, W.; Pan, Q.; Liu, J.; Yang, X.; Yan, J.; Xiao, Y. Intraspecific variation of residual heterozygosity and its utility for quantitative genetic studies in maize. BMC Plant Biol. 2018, 18, 66. [Google Scholar] [CrossRef]
- Tuinstra, M.; Ejeta, G.; Goldsbrough, P. Heterogeneous inbred family (HIF) analysis: A method for developing near-isogenic lines that differ at quantitative trait loci. Theor. Appl. Genet. 1997, 95, 1005–1011. [Google Scholar] [CrossRef]
- Lavenus, J.; Goh, T.; Roberts, I.; Guyomarc’h, S.; Lucas, M.; De Smet, I.; Fukaki, H.; Beeckman, T.; Bennett, M.; Laplaze, L. Lateral root development in Arabidopsis: Fifty shades of auxin. Trends Plant Sci 2013, 18, 450–458. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Luo, M.C.; Chen, Z.; You, F.M.; Wei, Y.; Zheng, Y.; Dvorak, J.J. Aegilops tauschii single nucleotide polymorphisms shed light on the origins of wheat D-genome genetic diversity and pinpoint the geographic origin of hexaploid wheat. New Phytol. 2013, 198, 925–937. [Google Scholar] [CrossRef] [PubMed]
- Chao, S.; Dubcovsky, J.; Dvorak, J.; Luo, M.-C.; Baenziger, S.P.; Matnyazov, R.; Clark, D.R.; Talbert, L.E.; Anderson, J.A.; Dreisigacker, S.; et al. Population- and genome-specific patterns of linkage disequilibrium and SNP variation in spring and winter wheat (Triticum aestivum L.). BMC Genom. 2010, 11, 727. [Google Scholar] [CrossRef]
- Dubcovsky, J.; Dvorak, J.J.S. Genome plasticity a key factor in the success of polyploid wheat under domestication. Science 2007, 316, 1862–1866. [Google Scholar] [CrossRef]
- Mares, D.; Mrva, K. Genetic variation for quality traits in synthetic wheat germplasm. BMC Genom. 2008, 59, 406–412. [Google Scholar] [CrossRef]
- Tadesse, W.; Schmolke, M.; Hsam, S.; Mohler, V.; Wenzel, G.; Zeller, F.J.T.; Genetics, A. Molecular mapping of resistance genes to tan spot [Pyrenophora tritici-repentis race 1] in synthetic wheat lines. Theor. Appl. Genet. 2007, 114, 855–862. [Google Scholar] [CrossRef]
- Li, Y.; Zhou, R.; Wang, J.; Liao, X.; Branlard, G.; Jia, J. Novel and favorable QTL allele clusters for end-use quality revealed by introgression lines derived from synthetic wheat. Mol. Breed. 2012, 29, 627–643. [Google Scholar] [CrossRef]
- Singh, S.; Franks, C.; Huang, L.; Brown-Guedira, G.; Marshall, D.; Gill, B.; Fritz, A.J.T.; Genetics, A. Lr41, Lr39, and a leaf rust resistance gene from Aegilops cylindrica may be allelic and are located on wheat chromosome 2DS. Theor. Appl. Genet. 2004, 108, 586–591. [Google Scholar] [CrossRef]
- Liao, M.; Fillery, I.R.; Palta, J. Early vigorous growth is a major factor influencing nitrogen uptake in wheat. Plant. Biol. 2004, 31, 121–129. [Google Scholar] [CrossRef]
- Liu, X.; Li, R.; Chang, X.; Jing, R.J.E. Mapping QTLs for seedling root traits in a doubled haploid wheat population under different water regimes. Euphytica 2013, 189, 51–66. [Google Scholar] [CrossRef]
- Den Herder, G.; Van Isterdael, G.; Beeckman, T.; De Smet, I. The roots of a new green revolution. Trends Plant Sci. 2010, 15, 600–607. [Google Scholar] [CrossRef]
- Xu, Y.; Yang, Y.; Wu, S.; Liu, D.; Ren, Y. QTL mapping for root traits and their effects on nutrient uptake and yield performance in common wheat (Triticum aestivum L.). Agriculture 2023, 13, 210. [Google Scholar] [CrossRef]
- Wachsman, G.; Sparks, E.E.; Benfey, P.N. Genes and networks regulating root anatomy and architecture. New Phytol. 2015, 208, 26–38. [Google Scholar] [CrossRef]
- Bai, C.; Liang, Y.; Hawkesford, M.J. Identification of QTLs associated with seedling root traits and their correlation with plant height in wheat. J. Exp. Bot. 2013, 64, 1745–1753. [Google Scholar] [CrossRef]
- Liu, P.; Jin, Y.; Liu, J.; Liu, C.; Yao, H.; Luo, F.; Guo, Z.; Xia, X.; He, Z.J.E. Genome-wide association mapping of root system architecture traits in common wheat (Triticum aestivum L.). Euphytica 2019, 215, 121. [Google Scholar] [CrossRef]
- Soriano, J.M.; Alvaro, F.J.S.R. Discovering consensus genomic regions in wheat for root-related traits by QTL meta-analysis. Sci. Rep. 2019, 9, 10537. [Google Scholar] [CrossRef] [PubMed]
- Zheng, X.; Wen, X.; Qiao, L.; Zhao, J.; Zhang, X.; Li, X.; Zhang, S.; Yang, Z.; Chang, Z.; Chen, J.; et al. A novel QTL QTrl.saw-2D.2 associated with the total root length identified by linkage and association analyses in wheat (Triticum aestivum L.). Planta 2019, 250, 129–143. [Google Scholar] [CrossRef]
- Ibrahim, S.E.; Schubert, A.; Pillen, K.; Lèon, J. QTL analysis of drought tolerance for seedling root morphological traits in an advanced backcross population of spring wheat. Biol. Sci. 2012, 2, 619–626. [Google Scholar]
- Mao, H.; Li, S.; Wang, Z.; Cheng, X.; Li, F.; Mei, F.; Chen, N.; Kang, Z. Regulatory changes in TaSNAC8-6A are associated with drought tolerance in wheat seedlings. Plant Biotechnol. J. 2020, 18, 1078–1092. [Google Scholar] [CrossRef]
- Tian, Q.; Reed, J.W.J.D. Control of auxin-regulated root development by the Arabidopsis thaliana SHY2/IAA3 gene. Development 1999, 126, 711–721. [Google Scholar] [CrossRef]
- Rogg, L.E.; Bartel, L. A gain-of-function mutation in IAA28 suppresses lateral root development. Plant Cell 2001, 13, 465–480. [Google Scholar] [CrossRef]
- Wilmoth, J.C.; Wang, S.; Tiwari, S.B.; Joshi, A.D.; Hagen, G.; Guilfoyle, T.J.; Alonso, J.M.; Ecker, J.R.; Reed, J.W. NPH4/ARF7 and ARF19 promote leaf expansion and auxin-induced lateral root formation. Plant J. 2010, 43, 118–130. [Google Scholar] [CrossRef]
- Gibson, S.W.; Todd, C.D. Arabidopsis AIR12 influences root development. Physiol. Mol. Biol. Plants 2015, 21, 479–489. [Google Scholar] [CrossRef]
- Neuteboom, L.W.; Ng, J.M.; Kuyper, M.; Clijdesdale, O.R.; Hooykaas, P.J.; Van Der Zaal, B.J. Isolation and characterization of cDNA clones corresponding with mRNAs that accumulate during auxin-induced lateral root formation. Plant Mol. Biol. 1999, 39, 273–287. [Google Scholar] [CrossRef]
- Fendrych, M.; Akhmanova, M.; Merrin, J.; Glanc, M.; Hagihara, S.; Takahashi, K.; Uchida, N.; Torii, K.U.; Friml, J.J. Rapid and reversible root growth inhibition by TIR1 auxin signalling. Nat. Plants 2018, 4, 453–459. [Google Scholar] [CrossRef]
- Prigge, M.J.; Platre, M.; Kadakia, N.; Zhang, Y.; Greenham, K.; Szutu, W.; Pandey, B.K.; Bhosale, R.A.; Bennett, M.J.; Busch, W.J.; et al. Genetic analysis of the Arabidopsis TIR1/AFB auxin receptors reveals both overlapping and specialized functions. Elife 2020, 9, e54740. [Google Scholar] [CrossRef]
- Bian, H.; Xie, Y.; Guo, F.; Han, N.; Ma, S.; Zeng, Z.; Wang, J.; Yang, Y.; Zhu, M.J. Distinctive expression patterns and roles of the miRNA393/TIR1 homolog module in regulating flag leaf inclination and primary and crown root growth in rice (Oryza sativa). New Phytol. 2012, 196, 149–161. [Google Scholar] [CrossRef]
- Guo, F.; Huang, Y.; Qi, P.; Lian, G.; Hu, X.; Han, N.; Wang, J.; Zhu, M.; Qian, Q.; Bian, H.J. Functional analysis of auxin receptor OsTIR1/OsAFB family members in rice grain yield, tillering, plant height, root system, germination, and auxinic herbicide resistance. New Phytol. 2021, 229, 2676–2692. [Google Scholar] [CrossRef]
- Su, P.; Zhao, L.; Li, W.; Zhao, J.; Yan, J.; Ma, X.; Li, A.; Wang, H.; Kong, L.J. Integrated metabolo-transcriptomics and functional characterization reveals that the wheat auxin receptor TIR1 negatively regulates defense against Fusarium graminearum. J. Integr. Plant. Biol. 2021, 63, 340–352. [Google Scholar] [CrossRef]
- Lu, G.; Coneva, V.; Casaretto, J.A.; Ying, S.; Mahmood, K.; Liu, F.; Nambara, E.; Bi, Y.M.; Rothstein, S.J. OsPIN5b modulates rice (Oryza sativa) plant architecture and yield by changing auxin homeostasis, transport and distribution. Plant J. 2015, 83, 913–925. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).