Appendix A
Table A1.
The table shows the metrics 1 for all the encoder and architecture combinations trained for segmentation in the study. The highest value for each is in bold.
Table A1.
The table shows the metrics 1 for all the encoder and architecture combinations trained for segmentation in the study. The highest value for each is in bold.
| Architecture | Encoder | Threshold | Precision | Recall | F1-Score | Specificity | IoU |
|---|
| MA-Net | EfficientNet-b2 | 0.25 | 0.257 | 0.629 | 0.365 | 0.968 | 0.223 |
| MA-Net | EfficientNet-b2 | 0.5 | 0.296 | 0.568 | 0.390 | 0.976 | 0.242 |
| MA-Net | EfficientNet-b2 | 0.75 | 0.336 | 0.501 | 0.402 | 0.982 | 0.251 |
| MA-Net | EfficientNet-b3 | 0.25 | 0.302 | 0.494 | 0.375 | 0.980 | 0.230 |
| MA-Net | EfficientNet-b3 | 0.5 | 0.347 | 0.428 | 0.383 | 0.986 | 0.237 |
| MA-Net | EfficientNet-b3 | 0.75 | 0.391 | 0.364 | 0.377 | 0.990 | 0.232 |
| MA-Net | EfficientNet-b4 | 0.25 | 0.295 | 0.593 | 0.394 | 0.975 | 0.245 |
| MA-Net | EfficientNet-b4 | 0.5 | 0.336 | 0.534 | 0.412 | 0.981 | 0.260 |
| MA-Net | EfficientNet-b4 | 0.75 | 0.377 | 0.470 | 0.418 | 0.986 | 0.264 |
| MA-Net | EfficientNet-b5 | 0.25 | 0.341 | 0.518 | 0.411 | 0.982 | 0.259 |
| MA-Net | EfficientNet-b5 | 0.5 | 0.383 | 0.461 | 0.419 | 0.987 | 0.265 |
| MA-Net | EfficientNet-b5 | 0.75 | 0.422 | 0.402 | 0.412 | 0.990 | 0.259 |
| PA-Net | EfficientNet-b2 | 0.25 | 0.239 | 0.690 | 0.355 | 0.961 | 0.216 |
| PA-Net | EfficientNet-b2 | 0.5 | 0.289 | 0.603 | 0.391 | 0.974 | 0.243 |
| PA-Net | EfficientNet-b2 | 0.75 | 0.337 | 0.505 | 0.405 | 0.982 | 0.254 |
| PA-Net | EfficientNet-b3 | 0.25 | 0.242 | 0.587 | 0.342 | 0.967 | 0.207 |
| PA-Net | EfficientNet-b3 | 0.5 | 0.290 | 0.501 | 0.367 | 0.978 | 0.225 |
| PA-Net | EfficientNet-b3 | 0.75 | 0.336 | 0.414 | 0.371 | 0.985 | 0.227 |
| PA-Net | EfficientNet-b4 | 0.25 | 0.263 | 0.631 | 0.372 | 0.969 | 0.228 |
| PA-Net | EfficientNet-b4 | 0.5 | 0.312 | 0.549 | 0.398 | 0.979 | 0.248 |
| PA-Net | EfficientNet-b4 | 0.75 | 0.359 | 0.461 | 0.404 | 0.985 | 0.253 |
| PA-Net | EfficientNet-b5 | 0.25 | 0.291 | 0.589 | 0.390 | 0.975 | 0.242 |
| PA-Net | EfficientNet-b5 | 0.5 | 0.345 | 0.501 | 0.408 | 0.983 | 0.257 |
| PA-Net | EfficientNet-b5 | 0.75 | 0.396 | 0.409 | 0.402 | 0.989 | 0.252 |
| U-Net | EfficientNet-b2 | 0.25 | 0.280 | 0.572 | 0.376 | 0.974 | 0.232 |
| U-Net | EfficientNet-b2 | 0.5 | 0.325 | 0.498 | 0.394 | 0.982 | 0.245 |
| U-Net | EfficientNet-b2 | 0.75 | 0.368 | 0.422 | 0.394 | 0.987 | 0.245 |
| U-Net | EfficientNet-b3 | 0.25 | 0.341 | 0.537 | 0.417 | 0.982 | 0.263 |
| U-Net | EfficientNet-b3 | 0.5 | 0.385 | 0.476 | 0.426 | 0.987 | 0.271 |
| U-Net | EfficientNet-b3 | 0.75 | 0.427 | 0.415 | 0.421 | 0.990 | 0.267 |
| U-Net | EfficientNet-b4 | 0.25 | 0.319 | 0.586 | 0.413 | 0.978 | 0.261 |
| U-Net | EfficientNet-b4 | 0.5 | 0.363 | 0.523 | 0.428 | 0.984 | 0.273 |
| U-Net | EfficientNet-b4 | 0.75 | 0.404 | 0.458 | 0.429 | 0.988 | 0.273 |
| U-Net | EfficientNet-b5 | 0.25 | 0.348 | 0.485 | 0.405 | 0.984 | 0.254 |
| U-Net | EfficientNet-b5 | 0.5 | 0.394 | 0.420 | 0.407 | 0.989 | 0.255 |
| U-Net | EfficientNet-b5 | 0.75 | 0.438 | 0.356 | 0.393 | 0.992 | 0.244 |
| U-Net++ | EfficientNet-b2 | 0.25 | 0.282 | 0.657 | 0.395 | 0.970 | 0.246 |
| U-Net++ | EfficientNet-b2 | 0.5 | 0.325 | 0.589 | 0.419 | 0.978 | 0.265 |
| U-Net++ | EfficientNet-b2 | 0.75 | 0.365 | 0.517 | 0.428 | 0.984 | 0.272 |
| U-Net++ | EfficientNet-b3 | 0.25 | 0.321 | 0.566 | 0.409 | 0.979 | 0.257 |
| U-Net++ | EfficientNet-b3 | 0.5 | 0.364 | 0.507 | 0.423 | 0.984 | 0.269 |
| U-Net++ | EfficientNet-b3 | 0.75 | 0.404 | 0.447 | 0.424 | 0.988 | 0.269 |
| U-Net++ | EfficientNet-b4 | 0.25 | 0.315 | 0.622 | 0.418 | 0.976 | 0.265 |
| U-Net++ | EfficientNet-b4 | 0.5 | 0.358 | 0.562 | 0.438 | 0.982 | 0.280 |
| U-Net++ | EfficientNet-b4 | 0.75 | 0.400 | 0.499 | 0.444 | 0.987 | 0.285 |
| U-Net++ | EfficientNet-b5 | 0.25 | 0.347 | 0.535 | 0.421 | 0.982 | 0.267 |
| U-Net++ | EfficientNet-b5 | 0.5 | 0.393 | 0.473 | 0.429 | 0.987 | 0.273 |
| U-Net++ | EfficientNet-b5 | 0.75 | 0.435 | 0.410 | 0.422 | 0.991 | 0.268 |
Figure A1.
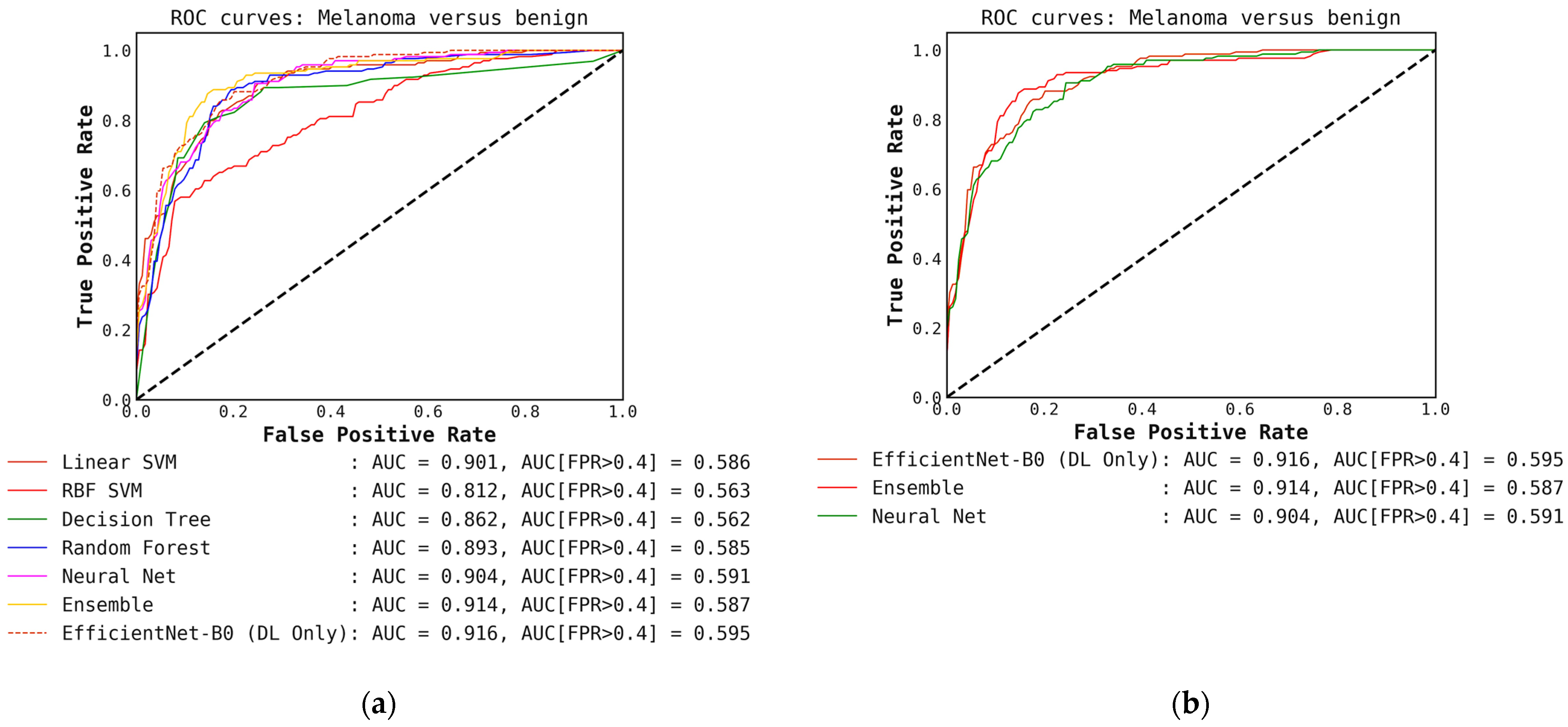
Comparison of ROC curves for conventional classifiers, DL classifiers, and their respective ensembles obtained by averaging the output probabilities. (a) ROC curves for conventional classifiers applied directly to hand-crafted features, without cascade generalization; (b) ROC curves for the deep learning (DL) models only and their ensemble. The area under the curve (AUC) and the AUC for a false-positive rate higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Figure A1.
Comparison of ROC curves for conventional classifiers, DL classifiers, and their respective ensembles obtained by averaging the output probabilities. (a) ROC curves for conventional classifiers applied directly to hand-crafted features, without cascade generalization; (b) ROC curves for the deep learning (DL) models only and their ensemble. The area under the curve (AUC) and the AUC for a false-positive rate higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Figure A2.
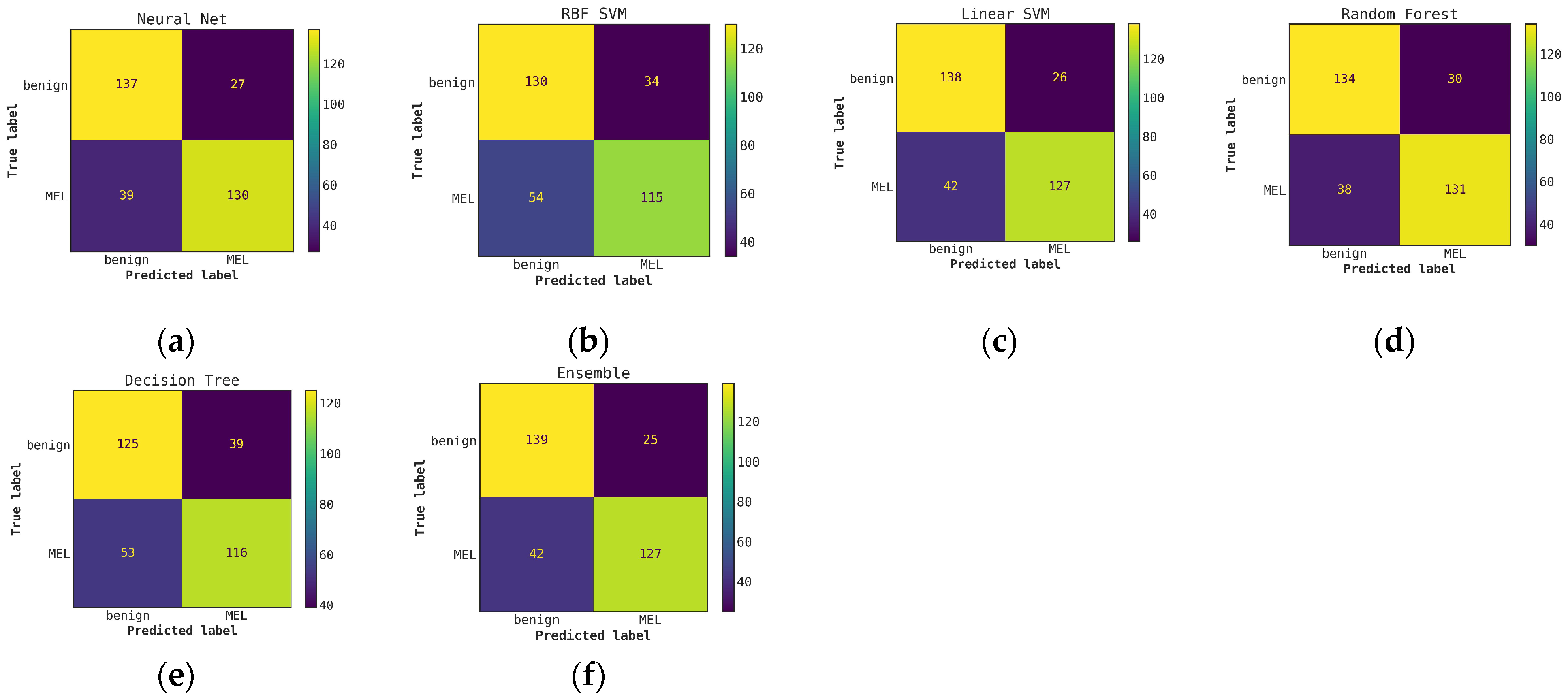
Comparison of confusion matrices for conventional classifiers using only hand-crafted features for classification (without Cascade generalization) on the classification hold-out test set. (a) Neural network; (b) RBF SVM; (c) Linear SVM; (d) Random Forest; (e) Decision Tree; (f) Ensemble of conventional models.
Figure A2.
Comparison of confusion matrices for conventional classifiers using only hand-crafted features for classification (without Cascade generalization) on the classification hold-out test set. (a) Neural network; (b) RBF SVM; (c) Linear SVM; (d) Random Forest; (e) Decision Tree; (f) Ensemble of conventional models.
Figure A3.
Comparison of confusion matrices for DL classifiers without cascade generalization on the classification hold-out test set. (a) ResNet50; (b) EfficientNet-B0; (c) EfficientNet-B1; (d) Ensemble of DL classifier models.
Figure A3.
Comparison of confusion matrices for DL classifiers without cascade generalization on the classification hold-out test set. (a) ResNet50; (b) EfficientNet-B0; (c) EfficientNet-B1; (d) Ensemble of DL classifier models.
Figure A4.
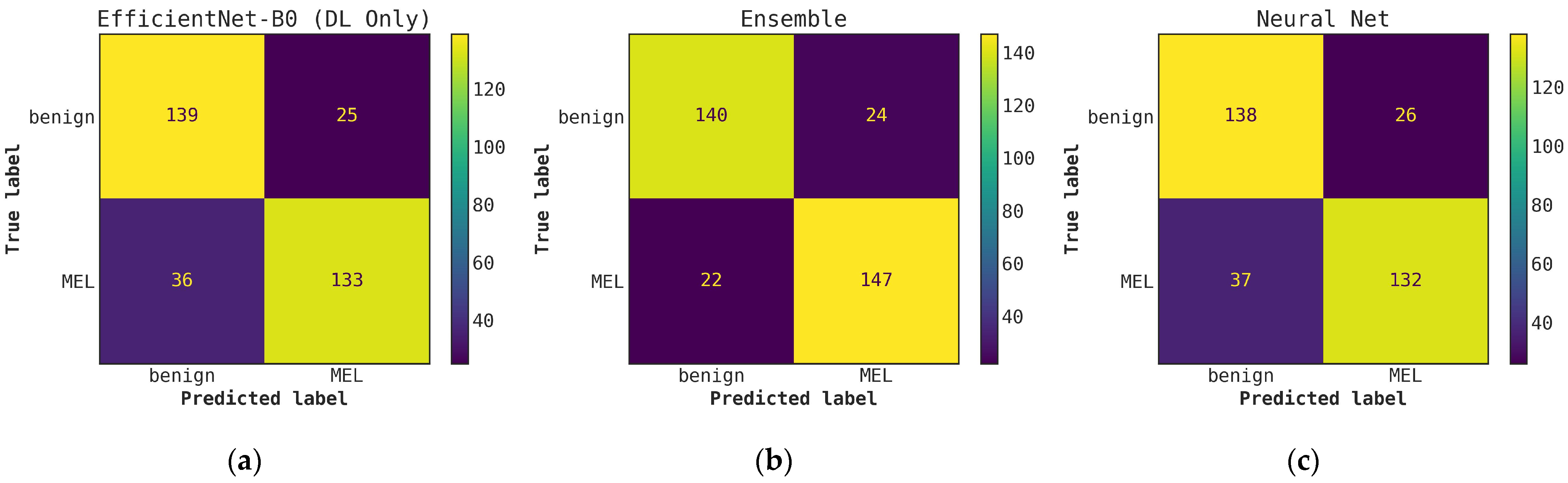
Illustrates a comparison of confusion matrices obtained using cascade generalization, utilizing the probability outputs of the EfficientNet-B0 model. (a) Confusion matrix for classification using only the EfficientNet-B0 model (DL-only, level-0); (b) Confusion matrix for classification utilizing the EfficientNet-B0 probability outputs and hand-crafted features, obtained through an ensemble of conventional classifiers (averaging); (c) Confusion matrix for classification using EfficientNet-B0 (level-0) probability outputs and hand-crafted features for the best conventional classifier, Neural networks. All confusion matrices are generated after applying a threshold of 0.5 to the models probability outputs to obtain the final predictions.
Figure A4.
Illustrates a comparison of confusion matrices obtained using cascade generalization, utilizing the probability outputs of the EfficientNet-B0 model. (a) Confusion matrix for classification using only the EfficientNet-B0 model (DL-only, level-0); (b) Confusion matrix for classification utilizing the EfficientNet-B0 probability outputs and hand-crafted features, obtained through an ensemble of conventional classifiers (averaging); (c) Confusion matrix for classification using EfficientNet-B0 (level-0) probability outputs and hand-crafted features for the best conventional classifier, Neural networks. All confusion matrices are generated after applying a threshold of 0.5 to the models probability outputs to obtain the final predictions.
Figure A5.
(a) ROC curves for conventional classifiers (level-1) used with the DL output probabilities from the EfficientNet-B0 (level-0) model and the hand-crafted features (cascade generalization). (b) Figure showing ROC curves only for the EfficientNet-B0 (level-0) model, the ensemble of conventional classifiers, and the best conventional level-1 classifier using the EfficientNet-B0 probability outputs. The AUC and the AUC for FPR higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Figure A5.
(a) ROC curves for conventional classifiers (level-1) used with the DL output probabilities from the EfficientNet-B0 (level-0) model and the hand-crafted features (cascade generalization). (b) Figure showing ROC curves only for the EfficientNet-B0 (level-0) model, the ensemble of conventional classifiers, and the best conventional level-1 classifier using the EfficientNet-B0 probability outputs. The AUC and the AUC for FPR higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Figure A6.
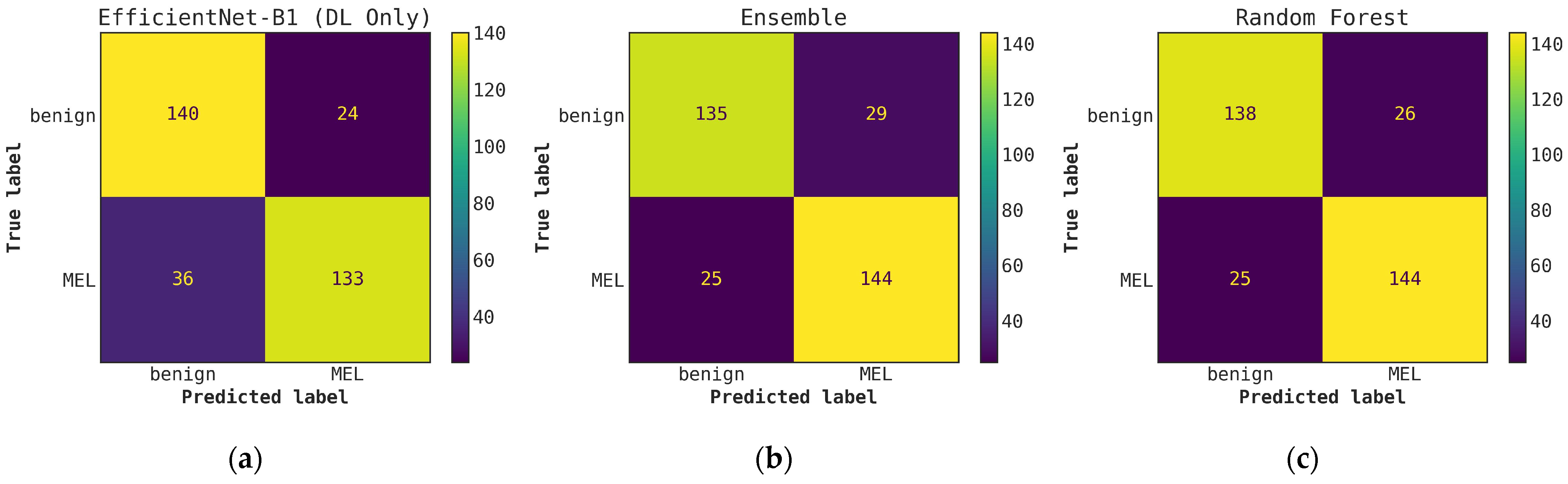
Illustrates a comparison of confusion matrices obtained using cascade generalization, utilizing the probability outputs of the EfficientNet-B1 model. (a) Confusion matrix for classification using only the EfficientNet-B1 model (DL-only, level-0); (b) Confusion matrix for classification utilizing the EfficientNet-B0 probability outputs and hand-crafted features, obtained through an ensemble of conventional classifiers (averaging); (c) Confusion matrix for classification using EfficientNet-B1 (level-0) probability outputs and hand-crafted features for the best conventional classifier, random forest. All confusion matrices are generated after applying a threshold of 0.5 to the model’s probability outputs to obtain the final predictions.
Figure A6.
Illustrates a comparison of confusion matrices obtained using cascade generalization, utilizing the probability outputs of the EfficientNet-B1 model. (a) Confusion matrix for classification using only the EfficientNet-B1 model (DL-only, level-0); (b) Confusion matrix for classification utilizing the EfficientNet-B0 probability outputs and hand-crafted features, obtained through an ensemble of conventional classifiers (averaging); (c) Confusion matrix for classification using EfficientNet-B1 (level-0) probability outputs and hand-crafted features for the best conventional classifier, random forest. All confusion matrices are generated after applying a threshold of 0.5 to the model’s probability outputs to obtain the final predictions.
Figure A7.
(a) ROC curves for conventional classifiers (level-1) used with the DL output probabilities from the EfficientNet-B1 (level-0) model and the hand-crafted features (cascade generalization); (b) Figure showing ROC curves only for the EfficientNet-B1 (level-0) model, the ensemble of conventional classifiers, and the best conventional level-1 classifier using the EfficientNet-B1 probability outputs. The AUC and the AUC for FPR higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Figure A7.
(a) ROC curves for conventional classifiers (level-1) used with the DL output probabilities from the EfficientNet-B1 (level-0) model and the hand-crafted features (cascade generalization); (b) Figure showing ROC curves only for the EfficientNet-B1 (level-0) model, the ensemble of conventional classifiers, and the best conventional level-1 classifier using the EfficientNet-B1 probability outputs. The AUC and the AUC for FPR higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Figure A8.
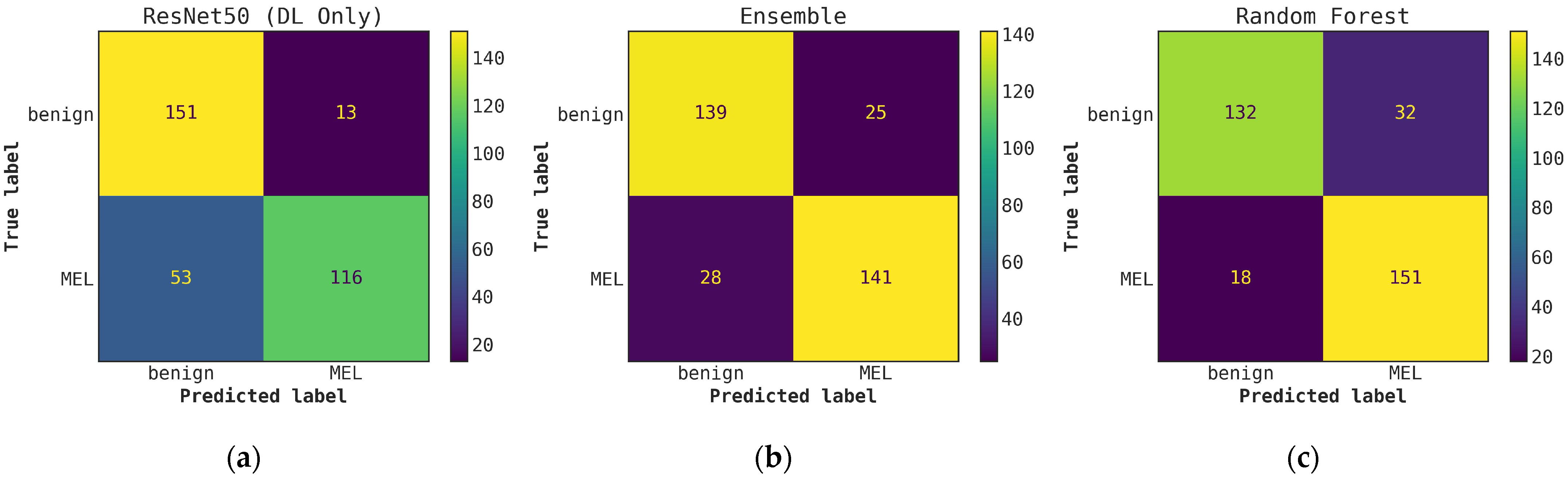
Illustrates a comparison of confusion matrices obtained through cascade generalization, utilizing the probability outputs of the ResNet50 model. (a) Confusion matrix for classification using only the ResNet50 model (DL-only, level-0); (b) Confusion matrix for classification utilizing the ResNet50 probability outputs and hand-crafted features, obtained through an ensemble of conventional classifiers (averaging); (c) Confusion matrix for classification using the ResNet50 (level-0) probability outputs and hand-crafted features for the best conventional classifier, random forest. All confusion matrices are generated after applying a threshold of 0.5 to the model’s probability outputs to obtain the final predictions.
Figure A8.
Illustrates a comparison of confusion matrices obtained through cascade generalization, utilizing the probability outputs of the ResNet50 model. (a) Confusion matrix for classification using only the ResNet50 model (DL-only, level-0); (b) Confusion matrix for classification utilizing the ResNet50 probability outputs and hand-crafted features, obtained through an ensemble of conventional classifiers (averaging); (c) Confusion matrix for classification using the ResNet50 (level-0) probability outputs and hand-crafted features for the best conventional classifier, random forest. All confusion matrices are generated after applying a threshold of 0.5 to the model’s probability outputs to obtain the final predictions.
Figure A9.
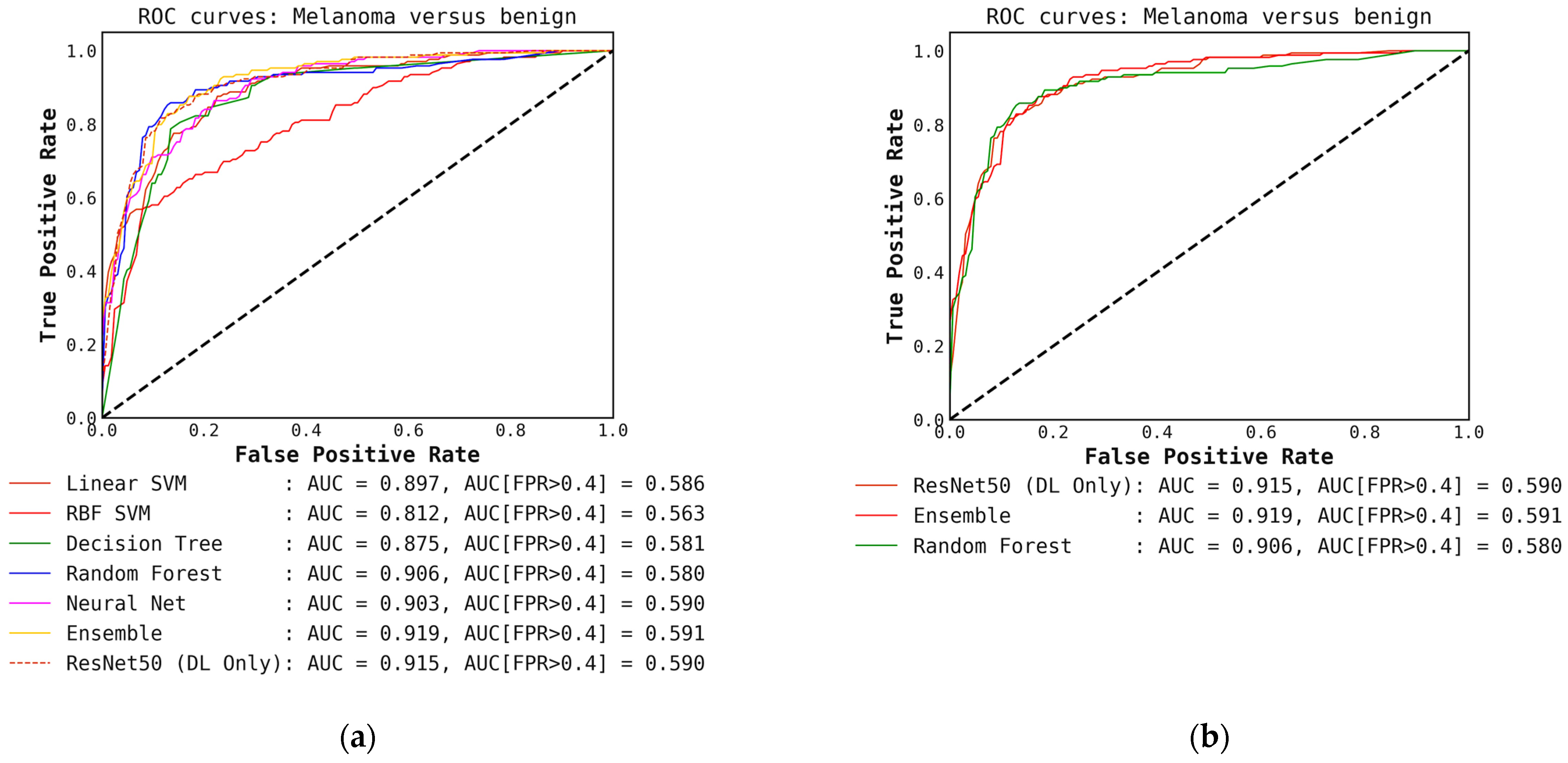
(a) ROC curves for conventional classifiers (level-1) used with the DL output probabilities from the ResNet50 (level-0) model and the hand-crafted features (cascade generalization); (b) Figure showing ROC curves only for the ResNet50 (level-0) model, the ensemble of conventional classifiers, and the best conventional level-1 classifier using the ResNet50 probability outputs. The AUC and the AUC for FPR higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Figure A9.
(a) ROC curves for conventional classifiers (level-1) used with the DL output probabilities from the ResNet50 (level-0) model and the hand-crafted features (cascade generalization); (b) Figure showing ROC curves only for the ResNet50 (level-0) model, the ensemble of conventional classifiers, and the best conventional level-1 classifier using the ResNet50 probability outputs. The AUC and the AUC for FPR higher than 0.4 are both presented for comparison. All values are rounded to three significant digits.
Table A2.
Table shows the 20 most important features based on the mean accuracy decrease after feature permutation. The scores are the averaged mean importance scores of all conventional classifiers without cascade generalization. Note that objects are non-overlapping distinct contours (blobs) in the irregular network binary mask generated by the segmentation model.
Table A2.
Table shows the 20 most important features based on the mean accuracy decrease after feature permutation. The scores are the averaged mean importance scores of all conventional classifiers without cascade generalization. Note that objects are non-overlapping distinct contours (blobs) in the irregular network binary mask generated by the segmentation model.
| Feature | Importance Score |
|---|
| Standard deviation of object’s color in L-plane inside lesion | 0.036 |
| Mean of object’s color in L-plane inside lesion | 0.026 |
| Standard deviation of object’s color in B-plane inside lesion | 0.026 |
| Total number of objects inside lesion | 0.021 |
| Mean of skin color in A-plane (excluding both lesion and irregular networks) | 0.019 |
| Mean of object’s color in B-plane inside lesion | 0.018 |
| Standard deviation of skin color in L-plane (excluding both lesion and irregular networks) | 0.016 |
| Maximum width for all objects | 0.014 |
| Standard deviation of skin color in B-plane (excluding both lesion and irregular networks) | 0.012 |
| Standard deviation of eccentricity for all objects | 0.011 |
| Standard deviation of width for all objects | 0.010 |
| Density of objects inside lesion (objects’ area inside lesion/lesion area) | 0.010 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 7 | 0.007 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 9 | 0.006 |
| Standard deviation of object’s color in L-plane outside lesion | 0.006 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 5 | 0.006 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 8 | 0.005 |
| Mean of skin color in B-plane (excluding both lesion and irregular networks) | 0.005 |
| Mean of object’s color in A-plane inside lesion | 0.004 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 3 | 0.004 |
Table A3.
Table shows the 20 most important features based on the mean accuracy decrease after feature permutation. The scores are the averaged mean importance scores for all conventional classifiers with cascade generalization. Note that objects are non-overlapping distinct contours (blobs) in the irregular network binary mask generated by the segmentation model.
Table A3.
Table shows the 20 most important features based on the mean accuracy decrease after feature permutation. The scores are the averaged mean importance scores for all conventional classifiers with cascade generalization. Note that objects are non-overlapping distinct contours (blobs) in the irregular network binary mask generated by the segmentation model.
| Feature | Importance Score |
|---|
| Standard deviation of object’s color in L-plane inside lesion | 0.037 |
| Deep Learning probability output | 0.030 |
| Mean of skin color in A-plane (excluding both lesion and irregular networks) | 0.030 |
| Total number of objects inside lesion | 0.021 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 3 | 0.015 |
| Mean of object’s color in L-plane inside lesion | 0.014 |
| Standard deviation of skin color in B-plane (excluding both lesion and irregular networks) | 0.014 |
| Standard deviation of objects color in B-plane inside lesion | 0.014 |
| Standard deviation of skin color in L-plane (excluding both lesion and irregular networks) | 0.013 |
| Mean of object’s color in B-plane inside lesion | 0.012 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 2 | 0.011 |
| Standard deviation of object’s color in A-plane inside lesion | 0.010 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 1 | 0.010 |
| Density of objects inside lesion (objects’ area inside lesion/lesion area) | 0.009 |
| Standard deviation of eccentricity of objects | 0.009 |
| Maximum width for all objects | 0.009 |
| Mean of object’s color in A-plane inside lesion | 0.007 |
| Total mask area after applying erosion with circular structuring element of radius 1 | 0.006 |
| Total number of objects remaining after applying erosion with circular structuring element of radius 4 | 0.006 |
| Mean of objects color in L-plane outside lesion | 0.006 |