Comprehensive Analysis of Feature Extraction Methods for Emotion Recognition from Multichannel EEG Recordings
Abstract
1. Introduction
2. Materials and Methods
2.1. Emotion-Related EEG Datasets
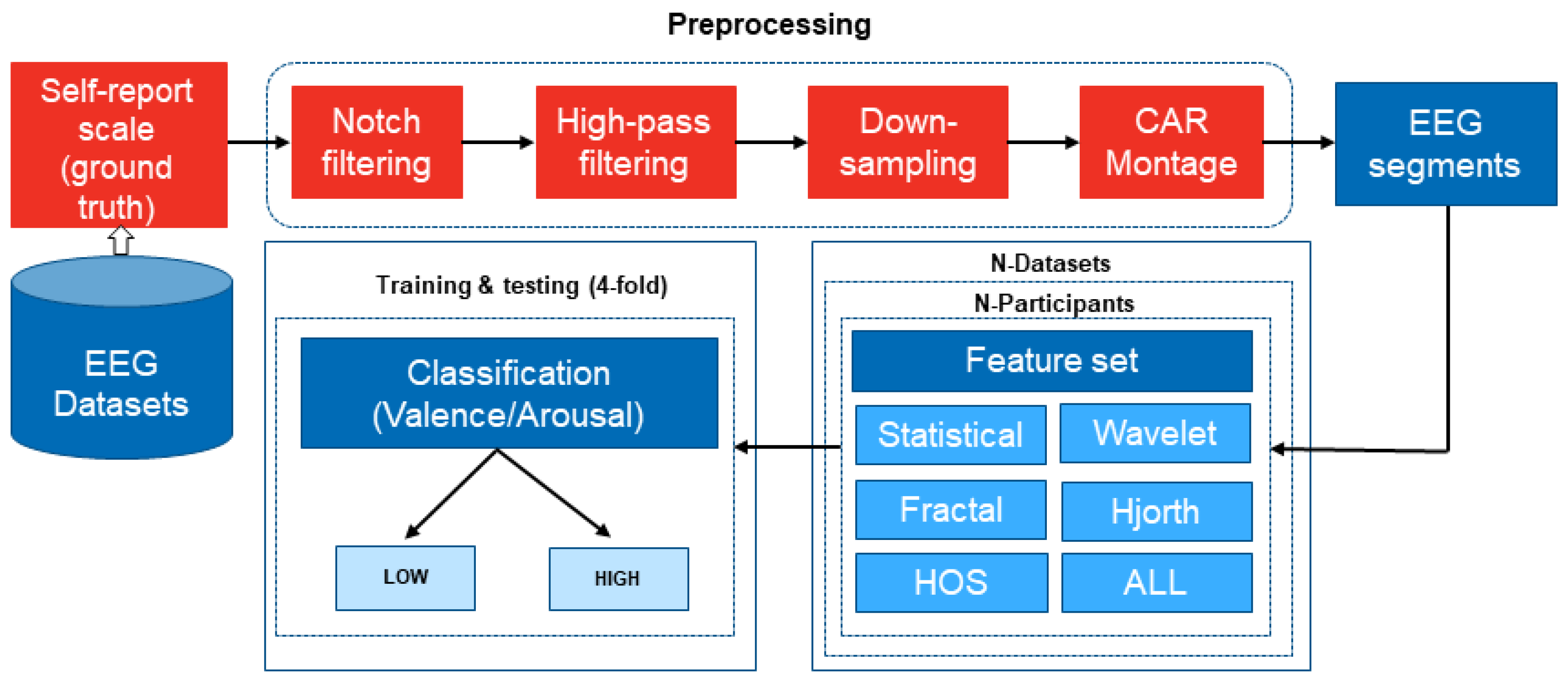
2.2. EEG Signal Preprocessing
2.3. EEG Feature Extraction
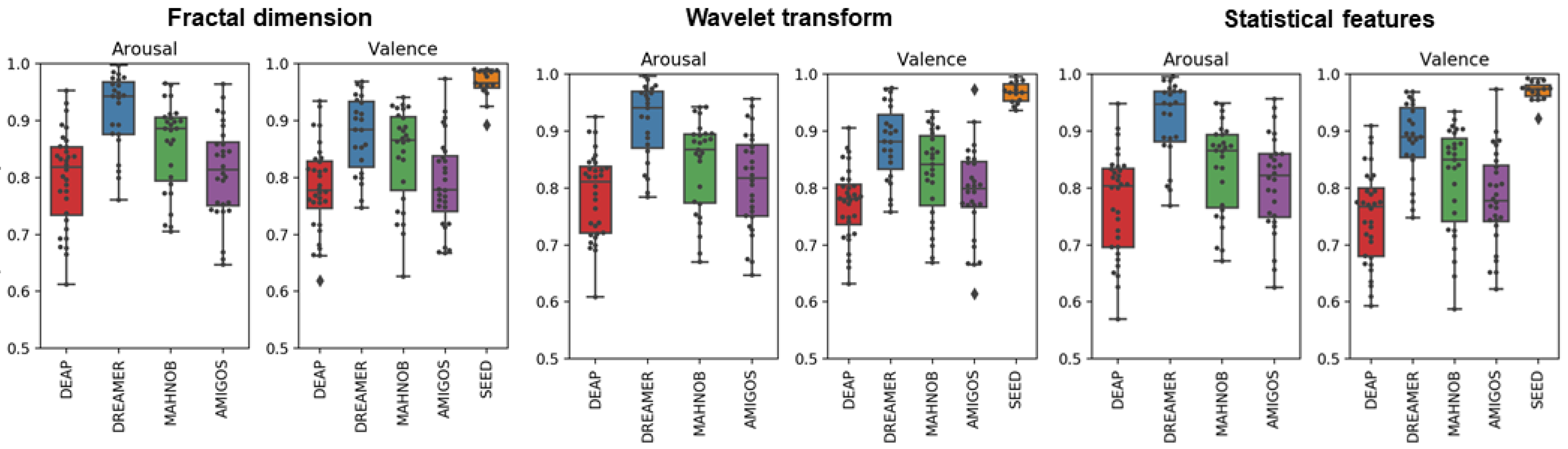
2.3.1. Statistical Features
2.3.2. Wavelet Analysis
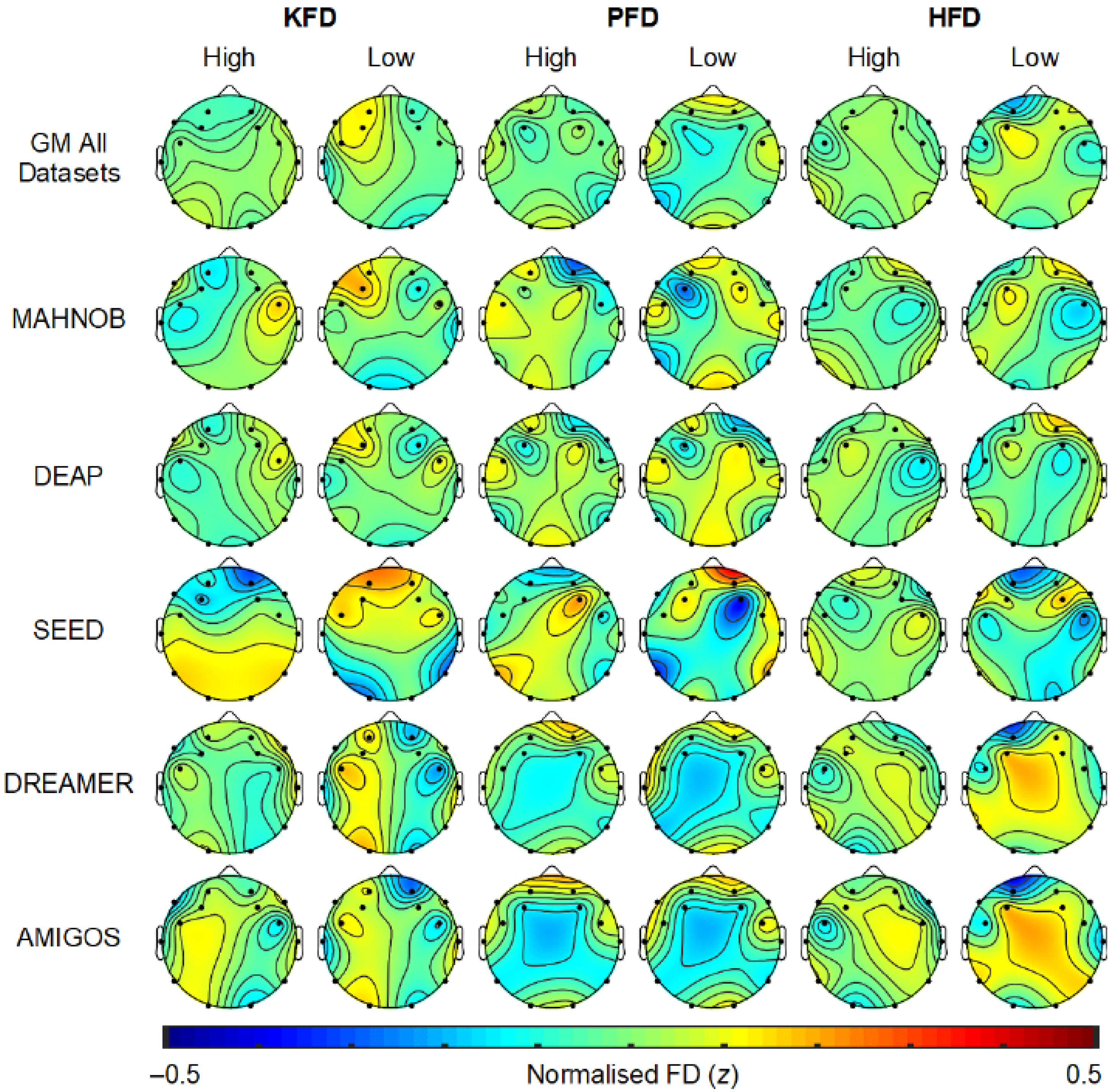
2.3.3. Fractal Dimension
2.3.4. Hjorth Parameters
2.3.5. Higher Order Spectra
2.4. Emotion and EEG Feature-Classification Techniques
2.5. EEG Feature-Classification Accuracy
2.6. Statistical Analysis: Comparing Feature-Classification Performance between EEG Feature Sets
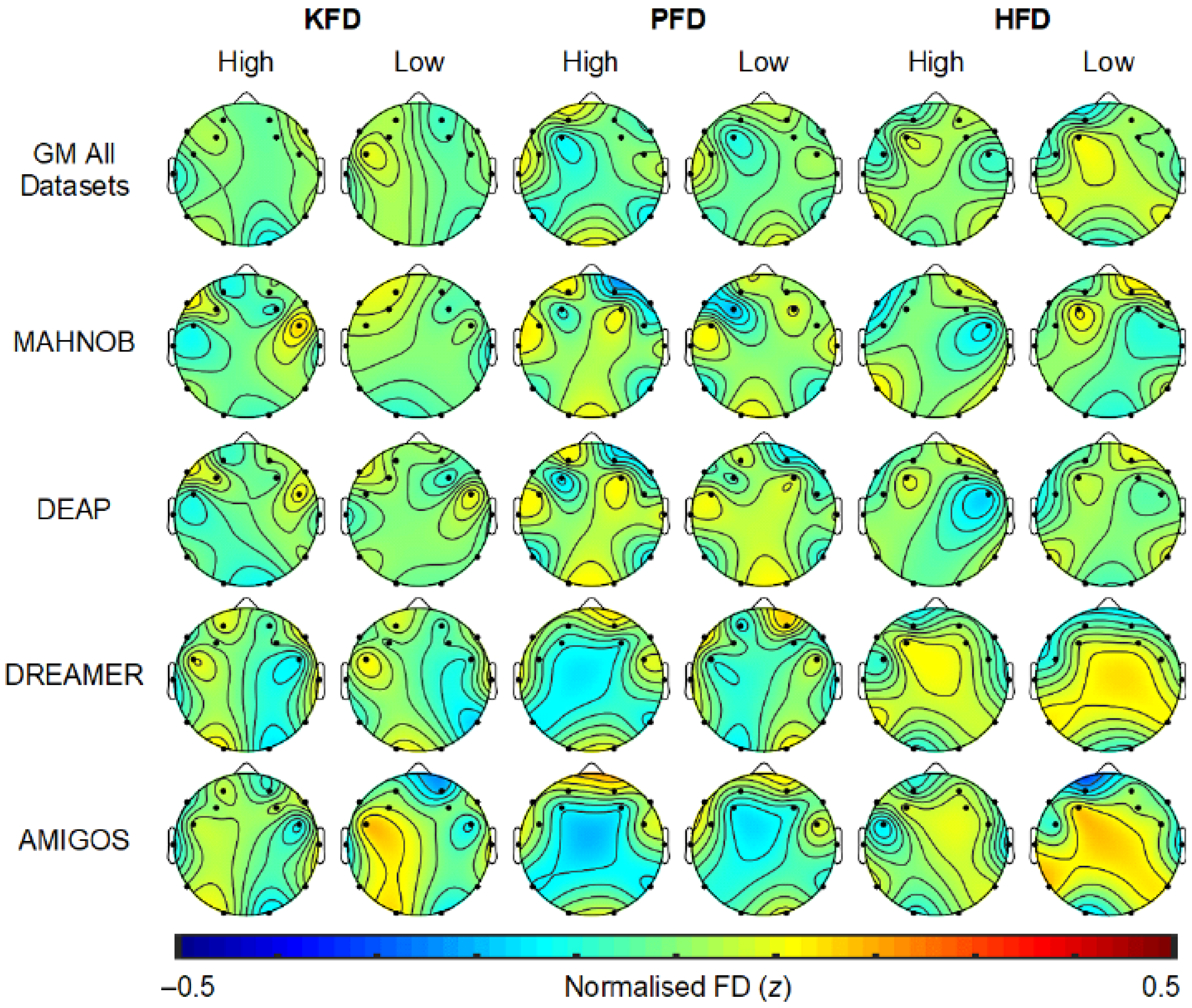
2.7. EEG Scalp Topography Related to Emotion Processing
3. Experimental Results and Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CART | Classification and Regression Tree |
| DWT | Discrete Wavelet Transform |
| DEAP | Dataset for Emotion Analysis using Physiological signals |
| ECG | Electrocardiogram |
| EEG | Electroencephalogram |
| EDA | Electrodermal Activity |
| EMG | electromyogram |
| ERP | Event Related Potential |
| ERO | Event Related Oscillation |
| FD | Fractal Dimension |
| GM | Grand Mean |
| GSVM | Gaussian Radial Basis Fucntion Support Vector Machine |
| GSR | Galvanic Skin Resistance |
| HFD | Higuchi’s fractal dimension |
| HOC | Higher Order Crossing |
| HOS | Higher Order Spectra |
| KFD | Katz’s Fractal Dimension |
| KNN | K-Nearest Neighbor |
| PFD | Petrosian fractal dimension |
| PNS | Peripheral Nervous System |
| RF | Random Forest |
| RR | Respiration Rate |
| SD | Standard Deviation |
| SVM | Support Vector Machine |
References
- Shu, L.; Xie, J.; Yang, M.; Li, Z.; Li, Z.; Liao, D.; Xu, X.; Yang, X. A Review of Emotion Recognition Using Physiological Signals. Sensors 2018, 18, 2074. [Google Scholar] [CrossRef] [PubMed]
- Egger, M.; Ley, M.; Hanke, S. Emotion recognition from physiological signal analysis: A review. Electron. Notes Theor. Comput. Sci. 2019, 343, 35–55. [Google Scholar] [CrossRef]
- Ko, B.C. A brief review of facial emotion recognition based on visual information. Sensors 2018, 18, 401. [Google Scholar] [CrossRef] [PubMed]
- Wani, T.M.; Gunawan, T.S.; Qadri, S.A.A.; Kartiwi, M.; Ambikairajah, E. A comprehensive review of speech emotion recognition systems. IEEE Access 2021, 9, 47795–47814. [Google Scholar] [CrossRef]
- Liu, H.; Zhang, Y.; Li, Y.; Kong, X. Review on emotion recognition based on electroencephalography. Front. Comput. Neurosci. 2021, 15, 758212. [Google Scholar] [CrossRef]
- Schaaff, K.; Schultz, T. Towards emotion recognition from electroencephalographic signals. In Proceedings of the 3rd International Conference on Affective Computing and Intelligent Interaction and Workshops (ACII 2009), Amsterdam, The Netherlands, 10–12 September 2009. [Google Scholar]
- Petrantonakis, P.C.; Hadjileontiadis, L.J. Emotion recogntion from brain signals using hybrid adaptive filtering and higher order crossings analysis. IEEE Trans. Affect. Comput. 2010, 1, 81–96. [Google Scholar] [CrossRef]
- Frantzidis, C.A.; Bratsas, C.; Papadelis, C.L.; Konstantinidis, E.; Pappas, C.; Bamidis, P.D. Toward emotion aware computing:An integrated approach using multichannel neurophysiological recordings and affective visual stimuli. IEEE Trans. Inf. Technol. Biomed. 2010, 14, 589–597. [Google Scholar] [CrossRef]
- Hadjidimitriou, S.K.; Hadjileontiadis, L.J. Toward an EEG-based recognition of music liking using time-frequency analysis. IEEE Trans. Biomed. Eng. 2012, 59, 3498–3510. [Google Scholar] [CrossRef]
- Jenke, R.; Peer, A.; Buss, M. Feature extraction and selection for emotion recogntion from EEG. IEEE Trans. Affect. Comput. 2014, 5, 327–339. [Google Scholar] [CrossRef]
- Liu, Y.; Sourina, O. Real-time subject-dependent eeg-based emotion recognition algorithm. In Transactions on Computational Science XXIII; Springer: Berlin/Heidelberg, Germany, 2014; pp. 199–223. [Google Scholar]
- Yuvaraj, R.; Murugappan, M.; Norlinah, M.I.; Sundaraj, K.; Omar, M.I.; Khairiyah, M.; Palaniappan, R. Optimal set of EEG features for emotional state classification and trajectory visualization in Parkinsosn’s disease. Int. J. Psychophysiol. 2014, 94, 482–495. [Google Scholar] [CrossRef]
- Nawaz, R.; Cheah, K.H.; Nisar, H.; Yap, V.V. Comparison of different feature extraction methods for EEG-based emotion recognition. Biocybern. Biomed. Eng. 2020, 40, 910–926. [Google Scholar] [CrossRef]
- Koelstra, S.; Soleymani, M.; Lee, J.-S.; Yazdani, A.; Ebrahimi, T.; Pun, T.; Nijholt, A.; Patras, I. DEAP: A Database for Emotion Analysis ;Using Physiological Signals. IEEE Trans. Affect. Comput. 2012, 3, 18–31. [Google Scholar] [CrossRef]
- Katsigiannis, S.; Ramzan, N. DREAMER: A Database for Emotion Recognition Through EEG and ECG Signals From Wireless Low-cost Off-the-Shelf Devices. IEEE J. Biomed. Health Inform. 2018, 22, 98–107. [Google Scholar] [CrossRef]
- Soleymani, M.; Lichtenauer, J.; Pun, T.; Pantic, M. A Multimodal Database for Affect Recognition and Implicit Tagging. IEEE Trans. Affect. Comput. 2012, 3, 42–55. [Google Scholar] [CrossRef]
- Miranda-Correa, J.A.; Abadi, M.K.; Sebe, N.; Patras, I. AMIGOS: A Dataset for Affect, Personality and Mood Research on Individuals and Groups. IEEE Trans. Affect. Comput. 2017, 12, 479–493. [Google Scholar] [CrossRef]
- Zheng, W.L.; Lu, B.L. Investigating critical frequency bands and channels for EEG-based emotion recognition with deep neural networks. IEEE Trans. Auton. Ment. Dev. 2015, 7, 162–175. [Google Scholar] [CrossRef]
- Topic, A.; Russo, M. Emotion recognition based on EEG feature maps through deep learning network. Int. J Eng. Sci. Technol. 2021, 24, 1441–1454. [Google Scholar] [CrossRef]
- Li, X.; Song, D.; Zhang, P.; Zhang, Y.; Hou, Y.; Hu, B. Exploring EEG features in cross-subject emotion recognition. Front. Neurosci. 2018, 12, 162. [Google Scholar] [CrossRef]
- Lan, Z.; Sourina, O.; Wang, L.; Scherer, R.; Müller-Putz, G.R. Domain adaptation techniques for EEG-based emotion recognition: A comparative study on two public datasets. IEEE Trans. Cogn. Dev. Syst. 2018, 11, 85–94. [Google Scholar] [CrossRef]
- Posner, J.; Russell, J.A.; Peterson, B.S. The circumplex model of affect: An integrative approach to affective neuroscience, cognitive development, and psychopathology. Dev. Psychopathol. 2005, 17, 715–734. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Meng, H.; Li, M.; Zhang, F.; Qin, R.; Nandi, A.K. Emotion detection from EEG recordings based on supervised and unsupervised dimension reduction. Concurr. Comput. 2018, 30, 1–13. [Google Scholar] [CrossRef]
- Piho, L.; Tjahjadi, T. A mutual information based adaptive windowing of informative EEG for emotion recognition. IEEE Trans. Affect. Comput. 2018, 11, 722–735. [Google Scholar] [CrossRef]
- Garg, A.; Kapoor, A.; Bedi, A.K.; Sunkaria, R.K. Merged LSTM Model for emotion classification using EEG signals. In Proceedings of the International Conference on Data Science and Engineering (ICDSE), Patna, India, 26–28 September 2019. [Google Scholar]
- Putra, A.E.; Atmaji, C.; Ghaleb, F. EEG-Based Emotion Classification Using Wavelet Decomposition and K-Nearest Neighbor. In Proceedings of the 4th International Conference on Science and Technology (ICST), Yogyakarta, Indonesia, 7–8 August 2018. [Google Scholar]
- Wang, X.W.; Nie, D.; Lu, B.L. Emotional state classification from EEG data using machine learning approach. Neurocomputing 2013, 129, 94–106. [Google Scholar] [CrossRef]
- Taran, S.; Bajaj, V. Emotion recognition from single-channel EEG signals using a two-stage correlation and instantaneous frequency-based filtering method. Comput. Programs Biomed. 2019, 173, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Bajaj, V.; Pachori, R.B. Human Emotion Classification from EEG Signals Using Multiwavelet Transform. In Proceedings of the International Conference on Medical Biometrics, Shenzhen, China, 30 May–1 June 2014; pp. 125–130. [Google Scholar]
- Avramidis, K.; Zlatintsi, A.; Garoufis, C.; Maragos, P. Multiscale Fractal Analysis on EEG Signals for Music-Induced Emotion Recognition. Available online: https://arxiv.org/abs/2010.16310 (accessed on 20 January 2022).
- Liu, Y.; Sourina, O. Real-Time Fractal-Based Valence Level Recognition from EEG. In Transactions on Computational Science XVIII; Gavrilova, M.L., Tan, C.J.K., Kuijper, A., Eds.; Lecture Notes in Computer Science; Springer: Berlin/Heidelberg, Germany, 2013; p. 7848. [Google Scholar]
- Katz, M.J. Fractals and the analysis of waveforms. Comput. Biol. Med. 1998, 18, 145–156. [Google Scholar] [CrossRef]
- Hatamikia, S.; Nasrabadi, A.M. Recognition of emotional states induced by music videos based on nonlinear feature extraction and SOM classification. In Proceedings of the 21st Iranian Conference on Biomedical Engineering (ICBME), Tehran, Iran, 26–28 November 2014; pp. 333–337. [Google Scholar]
- Higuchi, T. Approach to an irregular time series on the basis of the fractal theory. Phys. D 1988, 31, 277–283. [Google Scholar] [CrossRef]
- Garcia-Martinez, B.; Martinez-Rodrigo, A.; Alcaraz, R.; Fernandez-Caballero, A. A Review on nonlinear methods using electroencephalographic recordings for emotion recognition. IEEE Trans. Affect. Comput. 2019, 10, 1–20. [Google Scholar] [CrossRef]
- Mehmood, R.M.; Du, R.; Lee, H.J. Optimal feature selection and deep learning ensembles method for emotion recognition from human brain EEG sensors. IEEE Access. 2017, 5, 14797–14806. [Google Scholar] [CrossRef]
- Hjorth, B. EEG analysis based on time domain properties. Electroencephalogr. Clin. Neurophysiol. 1970, 29, 306–310. [Google Scholar] [CrossRef]
- Hjorth, B. The physical significance of time domain descriptors in EEG analysis. Electroencephalogr. Clin. Neurophysiol. 1973, 34, 312–325. [Google Scholar]
- Hosseini, S.A. Classification of brain activity in emotional states using HOS analysis. Int. J Image Graph. Signal Process. 2012, 1, 21–27. [Google Scholar] [CrossRef]
- Yuvaraj, R.; Acharya, U.; Hagiwara, Y. A novel Parkinson’s Disease Diagnosis Index using higher-order spectra features in EEG signals. Neural Comput. Appl. 2016, 30, 1225–1235. [Google Scholar] [CrossRef]
- Yang, J.; Wu, Z.; Peng, K.; Okolo, P.N.; Zhang, W.; Zhao, H.; Sun, J. Parameter selection of Gaussian kernel SVM based on local density of training set. Inverse. Probl. Sci. Eng. 2021, 29, 536–548. [Google Scholar] [CrossRef]
- Mehmood, R.M.; Lee, H.J. Emotion classification of EEG brain signal using SVM and KNN. In Proceedings of the International Conference on Multimedia & Expo Workshops (ICMEW), Turin, Italy, 29 June–3 July 2015. [Google Scholar]
- Satyanarayana, K.N.V.; Shankar, T.; Poojita, G.; Vinay, G.; Amaranadh, H.N.S.V.l.S.; Babu, A.G. An Approach to EEG based Emotion Identification by SVM classifier. In Proceedings of the 6th International Conference on Computing Methodologies and Communication (ICCMC), Erode, India, 29–31 March 2022. [Google Scholar]
- Witten, I.H.; Frank, E.; Hall, M.A. Data Mining: Practical Machine Learning Tools and Techniques; Elsevier: Burlington, NJ, USA, 2011. [Google Scholar]
- Liu, Y.; Sourina, O.; Nguyen, M.K. Real time EEG-based emotion recognition and its applications. Trans. Comput. Sci. 2011, 6670, 256–277. [Google Scholar]
- Lan, Z.; Sourina, O.; Wang, L.; Liu, Y. Real-time EEGbased emotion monitoring using stable features. Visual Comput. 2016, 32, 347–358. [Google Scholar] [CrossRef]
- Thomas, J.; Jin, J.; Thangavel, P.; Bagheri, E.; Yuvaraj, R.; Dauwels, J.; Rathakrishnan, R.; Halford, J.J.; Cash, S.S.; Westover, B. Automated Detection of Interictal Epileptiform Discharges from Scalp Electroencephalograms by Convolutional Neural Networks. Int. J. Neural Syst. 2020, 30, 2050030. [Google Scholar] [CrossRef]
- Jukic, S.; Saracevic, M.; Subasi, A.; Kevric, J. Comparison of Ensemble Machine Learning Methods for Automated Classification of Focal and Non-Focal Epileptic EEG Signals. Mathematics 2020, 8, 1481. [Google Scholar] [CrossRef]
- Ruiz-Padial, E.; Ibáñez-Molina, A.J. Fractal dimesion of EEG signals and heart dynamics in discrete emotional states. Biol. Psychol. 2018, 137, 42–48. [Google Scholar] [CrossRef]
- Zappasodi, F.; Olejarczyk, E.; Marzetti, L.; Assenza, G.; Pizzella, V.; Tecchio, F. Fractal dimension of EEG activity senses neuronal impairment in acute stroke. PLoS ONE 2014, 9, e100199. [Google Scholar] [CrossRef]
- Plessow, F.; Fischer, R.; Kirschbaum, C.; Goschke, T. Inflexibly focused under stress: Acute psychosocial stress increases shielding of action goals at the expense of reduced cognitive flexibility with increasing time lag to the stressor. J. Cogn. Neurosci. 2011, 23, 3218–3227. [Google Scholar] [CrossRef]
- Siddharth, S.; Jung, T.-P.; Sejnowski, T.J. Utilizing deep learning towards multi-modal bio-sensing and vision-based affective computing. arXiv 2019, arXiv:1905.07039. [Google Scholar] [CrossRef]
- Li, P.; Liu, H.; Si, Y.; Li, C.; Li, F.; Zhu, X.; Huang, X.; Zeng, Y.; Yao, D.; Zhang, Y.; et al. EEG based emotion recognition by combining functional connectivity network and local activations. IEEE Trans. Biomed. Eng. 2019, 66, 2869–28881. [Google Scholar] [CrossRef]
- Cui, H.; Liu, A.; Zhang, X.; Chen, X.; Wang, K.; Chen, X. EEG-based Emotion Recognition using an End-to-End Regional-Asymmetric Convolutional Neural Network. Knowl. Based. Systs. 2020, 205, 106243. [Google Scholar] [CrossRef]
- Han, Z.; Chang, H.; Zhou, X.; Wang, J.; Wang, L.; Shao, Y. E2ENNet: An End-to-End Neural Network for Emotional Brain-Computer Interface. Font. Comput. Neurosci. 2022, 16, 942979. [Google Scholar] [CrossRef] [PubMed]
| Public Dataset Name | Pub.
Year | Sample Size (N) | Gender Ratio (Mean Age ± SD) | Total Trials or Videos | Trial/ Video Dura. | Rec. Ses. | # EEG Channels /Device /Fs | Emotional States | Rating Scale Ranges (Thres.) |
|---|---|---|---|---|---|---|---|---|---|
| MAHNOB -HCI | 2011 | 27 | 11M
/16F (NS ± NS) | 20 | 34.92 to 117 s | 1 | 32/ BioSemi Active II /256 Hz | Valence & Arousal | 1–9 (4.5) |
| DEAP | 2012 | 32 | 16M/16F (26.9 ± NS) | 40 | 60 s | 1 | 32/ BioSemi Active II /512 Hz | Valence & Arousal | 1–9 (4.5) |
| SEED | 2015 | 15 | 7M/8F (23.27 ± 2.37) | 10 | ∼240 s | 3 | 62/ ESI Neuro Scan /1000 Hz | Positive & Negative | −1, 0, & 1 (NA) |
| AMIGOS | 2018 | 40 | 27M/13F (28.3 ± NS) | 16 | <250 s | 2 | 14/ Emotive EPOC /128 Hz | Valence & Arousal | 1–9 (4.5) |
| DREAMER | 2018 | 23 | 14M/9F (26.6 ± 2.7) | 18 | 65–393 s | 1 | 14/ Emotiv EPO /128 Hz | Valence & Arousal | 1–5 (2.5) |
| Feature Set | Features | No. of Features |
|---|---|---|
| Statistical | Mean (), Median (), Standard deviation (), Skewness, Kurtosis, Mean of absolute values of 1st difference (), Mean of absolute values of 2nd difference (), Normalized 1st difference (), and Normalized 2nd difference () | 9 |
| Wavelet | Mean and standard deviation of the absolute values of the coefficients in each of the 12 scales (with Morlet as mother wavelet). | 24 |
| Fractal dimension (FD) | Katz’s fractal dimension (KFD), Petrosian fractal dimension (PFD), and Higuchi’s fractal dimension (HFD). | 3 |
| Hjorth parameters | Mobility (), and Complexity (). | 2 |
| Higher order spectra (HOS) | Bispectrum magnitude (), Sum of logarithmic amplitudes of Bispectrum (), Sum of logarithmic amplitudes of diagonal elements in the bispectrum (), and 1st-order spectral moment of amplitudes of diagonal elements of the bispectrum (). | 4 |
| Feature Set | Classifier | Dataset Name | Average | ||||
|---|---|---|---|---|---|---|---|
| DEAP | DREAMER | MAHNOB | AMIGOS | SEED | |||
| Combined-ALL | GSVM | 73.09 ± 0.060 | 86.56 ± 0.063 | 78.23 ± 0.080 | 76.94 ± 0.076 | 96.73 ± 0.024 | 82.31 ± 0.084 |
| CART | 76.38 ± 0.072 | 88.44 ± 0.066 | 82.08 ± 0.081 | 78.47 ± 0.087 | 97.08 ± 0.022 | 84.49 ± 0.075 | |
| Statistical | GSVM | 69.62 ± 0.066 | 83.74 ± 0.064 | 75.47 ± 0.077 | 74.47 ± 0.082 | 96.14 ± 0.035 | 79.89 ± 0.093 |
| CART | 75.02 ± 0.086 | 88.26 ± 0.067 | 81.67 ± 0.097 | 78.19 ± 0.087 | 97.01 ± 0.020 | 84.03 ± 0.078 | |
| Wavelet | GSVM | 69.11 ± 0.064 | 82.10 ± 0.075 | 71.94 ± 0.079 | 72.34 ± 0.070 | 93.39 ± 0.047 | 77.78 ± 0.089 |
| CART | 77.34 ± 0.066 | 87.99 ± 0.066 | 82.57 ± 0.082 | 79.12 ± 0.084 | 96.78 ± 0.023 | 84.76 ± 0.070 | |
| Fractal dimension | GSVM | 75.69 ± 0.065 | 83.94 ± 0.065 | 80.91 ± 0.078 | 76.83 ± 0.084 | 96.40 ± 0.030 | 82.75 ± 0.074 |
| CART | 78.18 ± 0.079 | 87.59 ± 0.067 | 83.98 ± 0.087 | 79.07 ± 0.084 | 96.50 ± 0.030 | 85.06 ± 0.066 | |
| Hjorth parameters | GSVM | 73.23 ± 0.068 | 82.20 ± 0.066 | 78.82 ± 0.069 | 71.57 ± 0.067 | 96.32 ± 0.023 | 80.43 ± 0.088 |
| CART | 70.52 ± 0.063 | 80.86 ± 0.065 | 75.33 ± 0.071 | 70.21 ± 0.070 | 94.25 ± 0.037 | 78.23 ± 0.089 | |
| Higher order spectra | GSVM | 73.78 ± 0.073 | 83.31 ± 0.076 | 79.39 ± 0.081 | 72.23 ± 0.086 | 96.83 ± 0.028 | 81.11 ± 0.088 |
| CART | 72.18 ± 0.076 | 83.68 ± 0.069 | 78.27 ± 0.087 | 72.98 ± 0.082 | 95.66 ± 0.037 | 80.56 ± 0.086 | |
| Feature Set | Classifier | Dataset Name | Average | |||
|---|---|---|---|---|---|---|
| DEAP | DREAMER | MAHNOB | AMIGOS | |||
| Combined-ALL | GSVM | 75.83 ± 0.072 | 90.35 ± 0.072 | 80.55 ± 0.081 | 79.49 ± 0.093 | 81.56 ± 0.053 |
| CART | 78.82 ± 0.076 | 92.02 ± 0.065 | 83.21 ± 0.082 | 81.00 ± 0.092 | 83.76 ± 0.050 | |
| Statistical | GSVM | 72.51 ± 0.085 | 88.92 ± 0.072 | 77.46 ± 0.084 | 77.10 ± 0.103 | 79.00 ± 0.060 |
| CART | 77.38 ± 0.092 | 91.76 ± 0.068 | 83.72 ± 0.084 | 80.94 ± 0.087 | 83.45 ± 0.053 | |
| Wavelet | GSVM | 71.60 ± 0.079 | 87.62 ± 0.089 | 75.65 ± 0.095 | 77.15 ± 0.096 | 78.01 ± 0.059 |
| CART | 78.83 ± 0.077 | 91.60 ± 0.067 | 84.14 ± 0.082 | 81.20 ± 0.087 | 83.94 ± 0.048 | |
| Fractal dimension | GSVM | 77.48 ± 0.072 | 88.99 ± 0.074 | 82.80 ± 0.079 | 79.10 ± 0.088 | 82.09 ± 0.044 |
| CART | 79.90 ± 0.086 | 91.60 ± 0.067 | 85.58 ± 0.085 | 81.11 ± 0.087 | 84.55 ± 0.045 | |
| Hjorth parameters | GSVM | 75.62 ± 0.069 | 87.02 ± 0.083 | 80.70 ± 0.075 | 75.28 ± 0.099 | 79.66 ± 0.047 |
| CART | 73.14 ± 0.071 | 86.07 ± 0.085 | 76.94 ± 0.080 | 74.21 ± 0.107 | 77.59 ± 0.050 | |
| Higher order spectra | GSVM | 75.83 ± 0.079 | 88.35 ± 0.082 | 81.13 ± 0.083 | 76.77 ± 0.091 | 80.52 ± 0.049 |
| CART | 75.36 ± 0.081 | 88.69 ± 0.081 | 79.62 ± 0.086 | 76.79 ± 0.098 | 80.11 ± 0.051 | |
| Condition | p-Value | Cliff’s Delta Effect Size | ||
|---|---|---|---|---|
| Arousal | Valence | Arousal | Valence | |
| FD vs. Wavelet | 3.31 × 10−3 | 3.53 × 10−2 | 0.045 | 0.022 |
| FD vs. Statistical | 6.85 × 10−8 | 1.31 × 10−9 | 0.063 | 0.061 |
| FD vs. Hjorth Parameters | 2.13 × 10−19 | 1.22 × 10−21 | 0.397 | 0.420 |
| FD vs. Higher order spectra | 3.01 × 10−19 | 1.53 × 10−21 | 0.261 | 0.277 |
| Combined-ALL | 3.93 × 10−4 | 1.25 × 10−3 | 0.058 | 0.045 |
| Research Study | Features Employed Classification Method | Best Accuracy (%) Achieved | ||||
|---|---|---|---|---|---|---|
| DEAP | DREAMER | MAHNOB | AMIGOS | SEED | ||
| Topic and Russo, [19] | HOLOfm CNN-SVM | V:76.61 A:77.72 | V:88.20 A:90.43 | - | V:87.39 A:90.54 | V:88.45 A: - |
| Topic and Russo, [19] | TOPOfm CNN-SVM | V:76.30 A:76.54 | V:81.96 A:84.92 | - | V:80.63 A:85.75 | V:70.37 A: - |
| Siddharth et al. [52] | RGB colored image CNN-ELM | V:71.09 A:72.58 | V:78.99 A:79.23 | V:80.77 A:80.42 | V:83.02 A:79.13 | - |
| Li et al. [53] | Spatial map GRELM-SVM | V:62.00 A: - | - | - | - | V:88.00 A: - |
| Katsigiannis et al. [15] | PSD SVM | - | V: 62.49 A:62.17 | - | - | - |
| Miranda et al. [17] | PSD, SPA SVM | - | - | - | V:57.60 A:59.20 | - |
| This study | FD-CART | V:78.18 A:79.90 | V:87.59 A:91.60 | V:83.98 A:85.58 | V:79.07 A:81.11 | V:96.50 A: - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yuvaraj, R.; Thagavel, P.; Thomas, J.; Fogarty, J.; Ali, F. Comprehensive Analysis of Feature Extraction Methods for Emotion Recognition from Multichannel EEG Recordings. Sensors 2023, 23, 915. https://doi.org/10.3390/s23020915
Yuvaraj R, Thagavel P, Thomas J, Fogarty J, Ali F. Comprehensive Analysis of Feature Extraction Methods for Emotion Recognition from Multichannel EEG Recordings. Sensors. 2023; 23(2):915. https://doi.org/10.3390/s23020915
Chicago/Turabian StyleYuvaraj, Rajamanickam, Prasanth Thagavel, John Thomas, Jack Fogarty, and Farhan Ali. 2023. "Comprehensive Analysis of Feature Extraction Methods for Emotion Recognition from Multichannel EEG Recordings" Sensors 23, no. 2: 915. https://doi.org/10.3390/s23020915
APA StyleYuvaraj, R., Thagavel, P., Thomas, J., Fogarty, J., & Ali, F. (2023). Comprehensive Analysis of Feature Extraction Methods for Emotion Recognition from Multichannel EEG Recordings. Sensors, 23(2), 915. https://doi.org/10.3390/s23020915







