Computer-Aided Drug Design Across Breast Cancer Subtypes: Methods, Applications and Translational Outlook
Abstract
1. Introduction
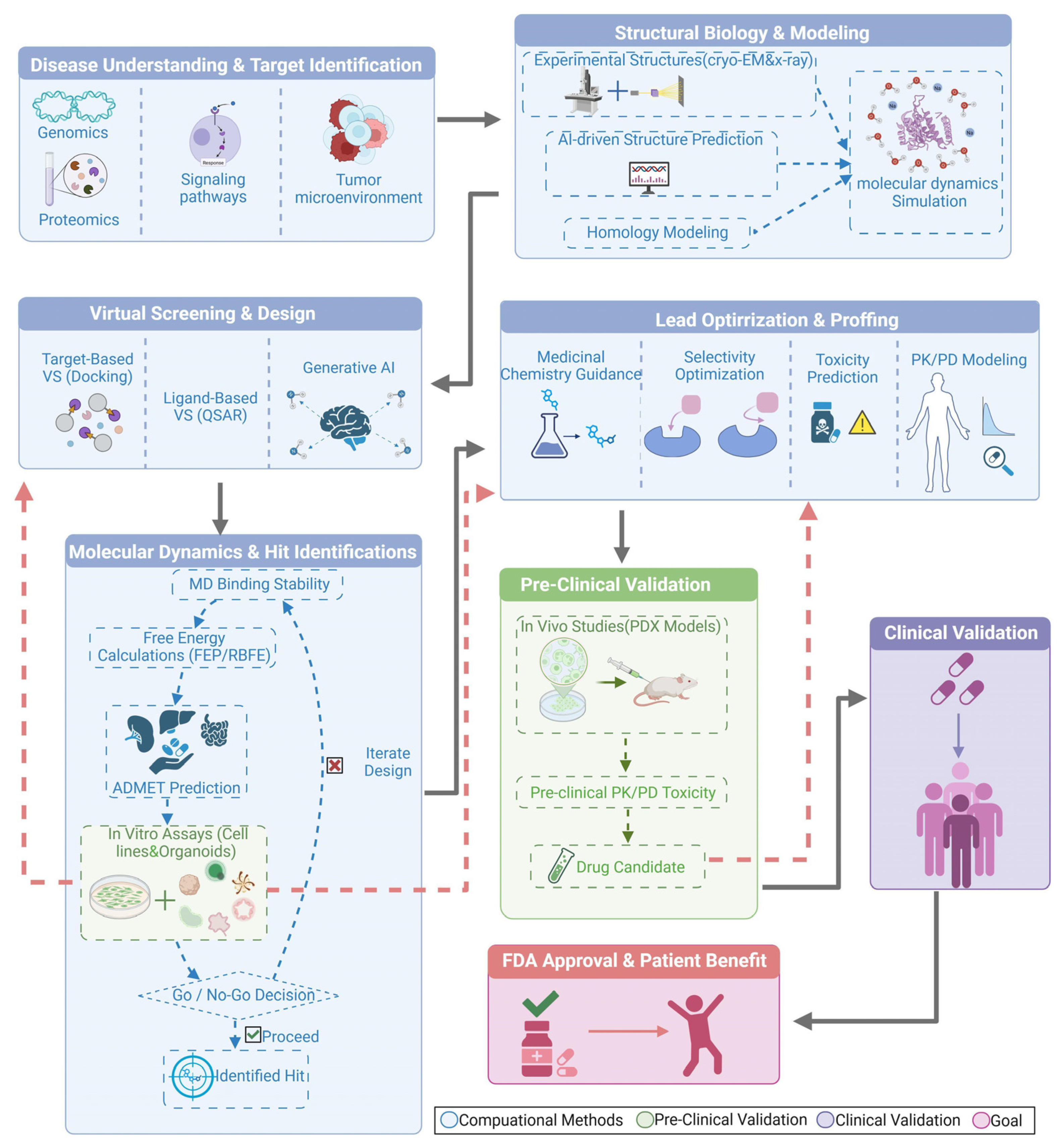
2. Foundations and Workflow of Computer-Aided Drug Design (CADD) in Cancer Therapeutics
2.1. Structural Foundations and Computational Methods
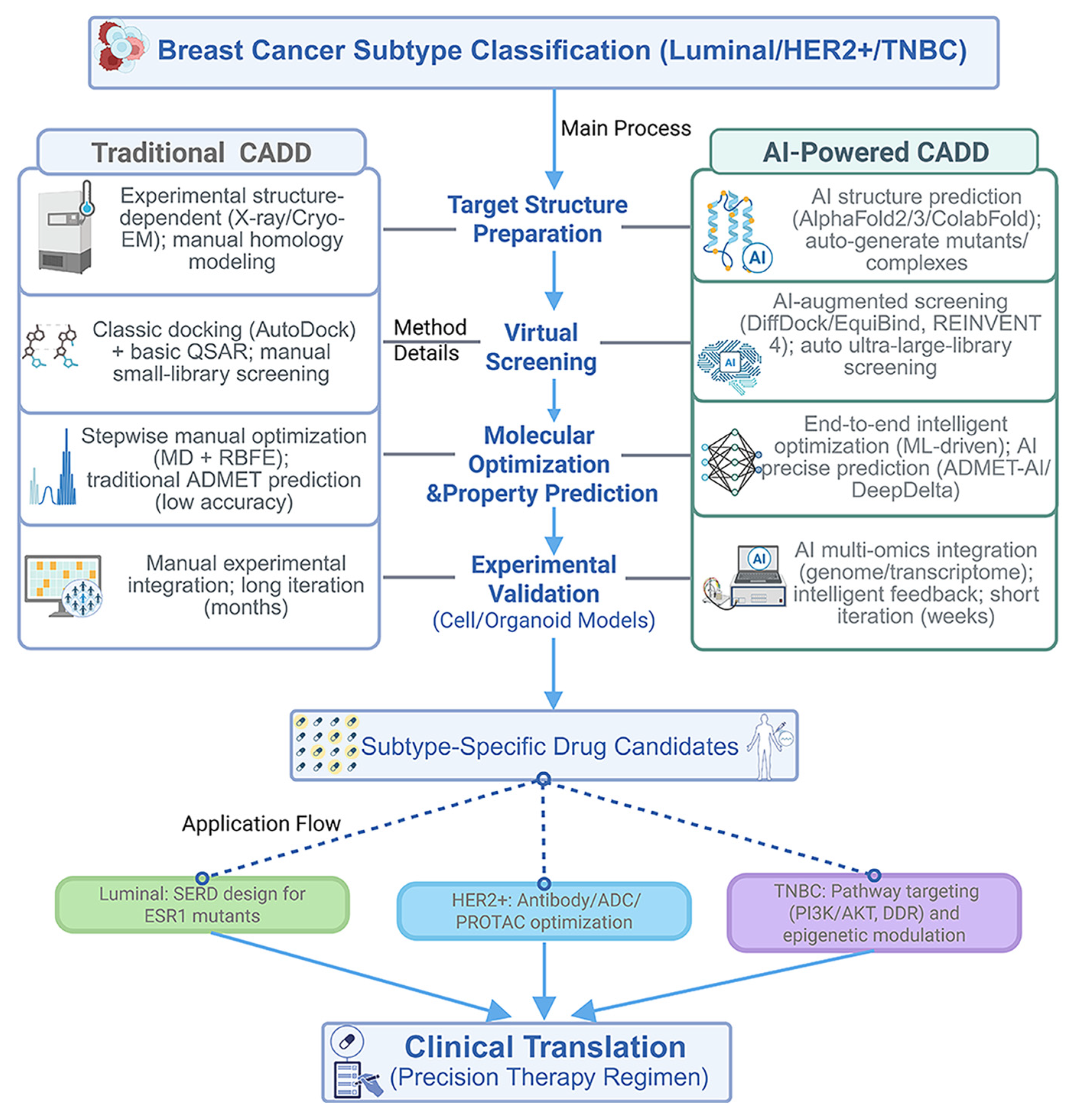
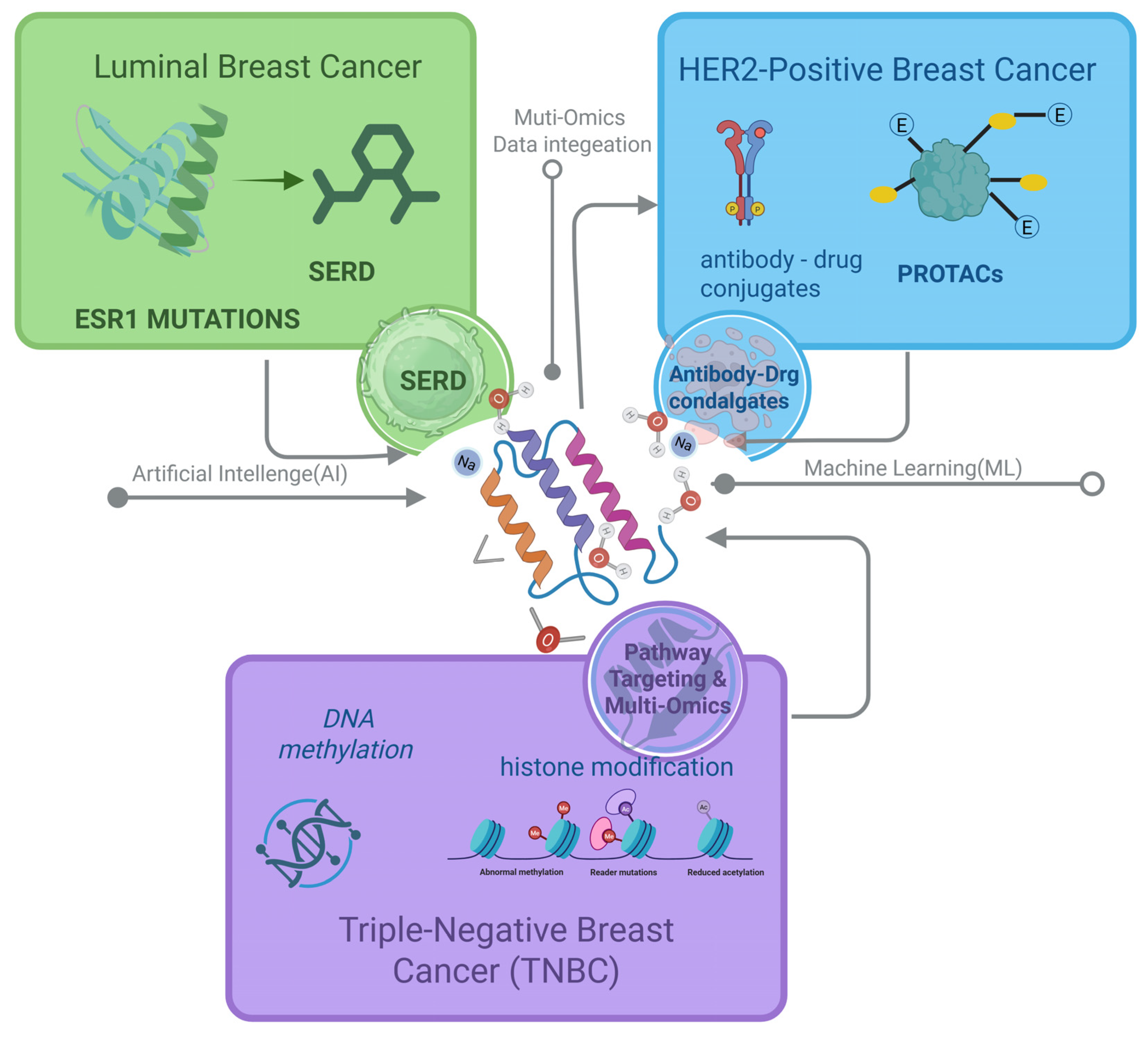
2.2. Applications of CADD Across Breast Cancer Subtypes
3. CADD Strategies for Luminal Breast Cancer
3.1. Targeting Estrogen Receptor Degradation
3.2. Structural Insights and Advanced Design Strategies
3.3. Overcoming Pharmacokinetic Challenges Through CADD
3.4. Clinical Applications and Translational Challenges of CADD-Designed SERDs
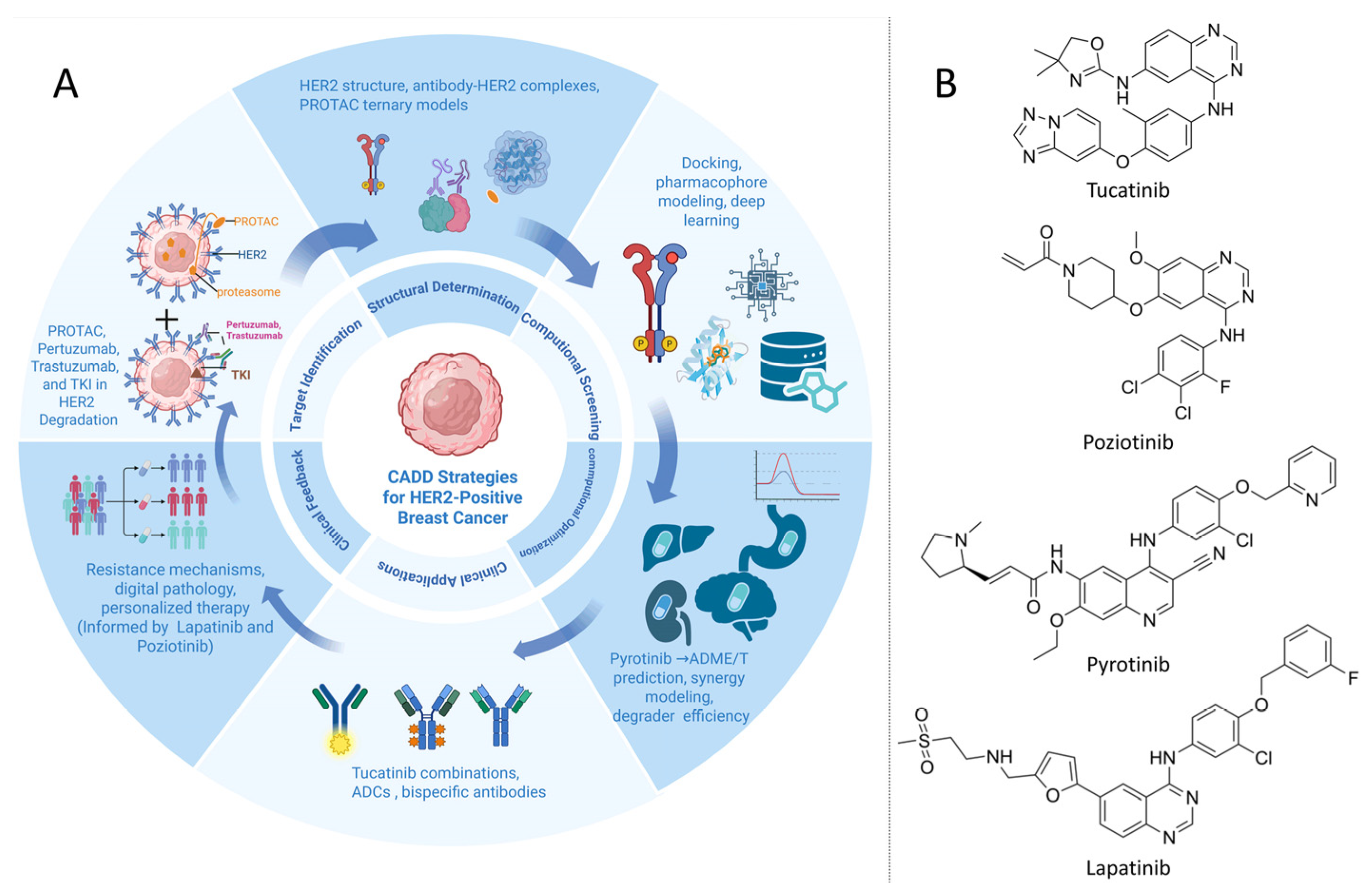
4. CADD Strategies for HER2+ Breast Cancer
4.1. Targeting HER2 with Antibodies and Small Molecules
4.2. PROTACs for HER2 Degradation
4.3. Combining CADD with Multi-Omics Data for Personalized Therapy
4.4. Resistance Mechanisms and CADD-Guided Counterstrategies in HER2+ Breast Cancer
5. Multi-Dimensional CADD Strategies for Triple-Negative Breast Cancer (TNBC)
5.1. Computational Targeting of Molecular Pathways in TNBC
5.2. PROTACs Targeting Transcriptional Regulators in TNBC
5.3. Multi-Omics Integration for Personalized TNBC Therapy
5.4. Resistance Mechanisms and CADD Counterstrategies in TNBC
6. CADD Key Technologies and Limitations
7. Conclusions and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer Statistics, 2024. CA Cancer J. Clin. 2024, 74, 8–31. [Google Scholar] [CrossRef]
- Curigliano, G.; Mueller, V.; Borges, V.F.; Hamilton, E.P.; Hurvitz, S.A.; Loi, S.; Murthy, R.K.; Okines, A.F.C.; Paplomata, E.; Cameron, D.A.; et al. Updated results of tucatinib versus placebo added to trastuzumab and capecitabine for patients with pretreated HER2+ metastatic breast cancer with and without brain metastases (HER2CLIMB). J. Clin. Oncol. 2021, 39, 1043. [Google Scholar] [CrossRef]
- Zheng, Y.; Kim, R.; Yu, T.; Gayle, J.A.; Wassel, C.L.; Dreyfus, J.; Phatak, H.; George, S. Real-World Clinical and Economic Outcomes in Selected Immune-Related Adverse Events Among Patients with Cancer Receiving Immune Checkpoint Inhibitors. Oncologist 2021, 26, e2002–e2012. [Google Scholar] [CrossRef]
- Abelman, R.O.; Spring, L.; Fell, G.G.; Ryan, P.; Vidula, N.; Medford, A.J.; Shin, J.; Abraham, E.; Wander, S.A.; Isakoff, S.J.; et al. Sequential Use of Antibody-Drug Conjugate After Antibody-Drug Conjugate for Patients with Metastatic Breast Cancer: ADC After ADC (A3) Study. J. Clin. Oncol. 2023, 41, 1022. [Google Scholar] [CrossRef]
- Turner, N.C.; Mayer, E.L.; Park, Y.H.; Janni, W.; Ma, C.X.; Cristofanilli, M.; Bianchini, G.; Kalinsky, K.; Iwata, H.; Chia, S.K.L.; et al. Camizestrant + CDK4/6 inhibitor for emergent ESR1 mutations during first-line endocrine-based therapy in HR+/HER2− advanced breast cancer: Phase 3, double-blind, ctDNA-guided SERENA-6 trial. J. Clin. Oncol. 2025, 43 (Suppl. 17), LBA4. [Google Scholar] [CrossRef]
- Garg, P.; Singhal, G.; Kulkarni, P.; Horne, D.; Salgia, R.; Singhal, S.S. Artificial intelligence-driven computational approaches in the development of anticancer drugs. Cancers 2024, 16, 3884. [Google Scholar] [CrossRef]
- De Gracia Triviño, J.A.; Delcey, M.G.; Wendin, G. Complete active space methods for NISQ devices: The importance of canonical orbital optimization for accuracy and noise resilience. J. Chem. Theory Comput. 2023, 19, 2863–2872. [Google Scholar] [CrossRef] [PubMed]
- Hossain, M.S.; Alom, M.S.; Kader, M.S.; Hossain, M.A.; Halim, M.A. Structure-guided antiviral peptides identification targeting the HIV-1 integrase. ACS Phys. Chem. Au. 2024, 4, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Tropsha, A.; Isayev, O.; Varnek, A.; Schneider, G.; Cherkasov, A. Integrating QSAR Modelling and Deep Learning in Drug Discovery: The Emergence of Deep QSAR. Nat. Rev. Drug Discov. 2024, 23, 141–155. [Google Scholar] [CrossRef]
- Swanson, K.; Walther, P.; Leitz, J.; Mukherjee, S.; Wu, J.C.; Shivnaraine, R.V.; Zou, J. ADMET-AI: A Machine Learning Platform for Fast and Accurate ADMET Property Prediction. Bioinformatics 2024, 40, btae416. [Google Scholar] [CrossRef] [PubMed]
- Vandenbossche, J.; Yogaratnam, J.; Hillewaert, V.; Rasschaert, F.; Talloen, W.; Biewenga, J.; Snoeys, J.; Kakuda, T.N.; Palmer, M.; Nangosyah, J.; et al. Drug-Drug Interactions with the Hepatitis B Virus Capsid Assembly Modulator JNJ-56136379 (Bersacapavir). Clin. Pharmacol. Drug Dev. 2022, 11, 1419–1429. [Google Scholar] [CrossRef]
- Chen, B.; Liu, S.-L.; Li, Q.; Liu, M.; Yang, H.; Xi, M. Predicting Role of Circulating Tumor DNA and Blood-Based Tumor Mutational Burden in Esophageal Squamous Cell Carcinoma Receiving Chemoradiotherapy Combined with Toripalimab: Exploratory Analyses from a Phase II Trial (EC-CRT-001). J. Clin. Oncol. 2023, 41, 4056. [Google Scholar] [CrossRef]
- Hosseini, S.-R.; Zhou, X. CCSynergy: An integrative deep-learning framework enabling context-aware prediction of anti-cancer drug synergy. Brief. Bioinform. 2023, 24, bbac588. [Google Scholar] [CrossRef]
- Zingg, D.; Bhin, J.; Yemelyanenko, J.; Kas, S.M.; Rolfs, F.; Lutz, C.; Lee, J.K.; Klarenbeek, S.; Silverman, I.M.; Annunziato, S.; et al. Truncated FGFR2 Is a Clinically Actionable Oncogene in Multiple Cancers. Nature 2022, 608, 609–617. [Google Scholar] [CrossRef] [PubMed]
- Menden, M.P.; Wang, D.; Mason, M.J.; Szalai, B.; Bulusu, K.C.; Guan, Y.; Yu, T.; Kang, J.; Jeon, M.; Wolfinger, R.; et al. Community Assessment to Advance Computational Prediction of Cancer Drug Combinations in a Pharmacogenomic Screen. Nat. Commun. 2019, 10, 2674. [Google Scholar] [CrossRef]
- Uludoğan, G.; Ozkirimli, E.; Ulgen, K.O.; Karalı, N.; Özgür, A. Exploiting Pretrained Biochemical Language Models for Targeted Drug Design. Bioinformatics 2022, 38, ii155–ii161. [Google Scholar] [CrossRef]
- Sharaf, B.; Hajahjeh, A.; Bani Hani, H.; Abdel-Razeq, H. Next generation selective estrogen receptor degraders in postmenopausal women with advanced-stage hormone receptors-positive, HER2-negative breast cancer. Front. Oncol. 2024, 14, 1385577. [Google Scholar] [CrossRef]
- Bilodeau, C.; Jin, W.; Jaakkola, T.; Barzilay, R.; Jensen, K.F. Generative Models for Molecular Discovery: Recent Advances and Challenges. WIREs Comput. Mol. Sci. 2022, 12, e1608. [Google Scholar] [CrossRef]
- Garioni, M.; Tschan, V.J.; Blukacz, L.; Nuciforo, S.; Parmentier, R.; Roma, L.; Coto-Llerena, M.; Pueschel, H.; Piscuoglio, S.; Vlajnic, T.; et al. Patient-derived organoids identify tailored therapeutic options and determinants of plasticity in sarcomatoid urothelial bladder cancer. NPJ Precis. Oncol. 2023, 7, 112. [Google Scholar] [CrossRef]
- Hu, M.; Li, Y.; Li, J.; Zhou, H.; Liu, C.; Liu, Z.; Gong, Y.; Ying, B.; Xie, Y. Discovery of potent and selective HER2 PROTAC degrader based on tucatinib with improved efficacy against HER2-positive cancers. Eur. J. Med. Chem. 2022, 244, 114775. [Google Scholar] [CrossRef]
- Zhang, C.; Condon, A.; Dao Duc, K. CryoSAMU: Enhancing 3D Cryo-EM Density Maps of Protein Structures at Intermediate Resolution with Structure-Aware Multimodal U-Nets. arXiv 2025. [Google Scholar] [CrossRef]
- Wu, L.; Wen, Y.; Leng, D.; Zhang, Q.; Dai, C.; Wang, Z.; Liu, Z.; Yan, B.; Zhang, Y.; Wang, J.; et al. Machine Learning Methods, Databases and Tools for Drug Combination Prediction. Brief. Bioinform. 2022, 23, bbab355. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Callaway, E. ‘The Entire Protein Universe’: AlphaFold Predictions for 200 Million Proteins. Nature 2022, 608, 15–16. [Google Scholar] [CrossRef]
- Mirdita, M.; Schütze, K.; Moriwaki, Y.; Heo, L.; Ovchinnikov, S.; Steinegger, M. ColabFold: Making Protein Folding Accessible to All. Nat. Methods 2022, 19, 679–682. [Google Scholar] [CrossRef]
- Varadi, M.; Anyango, S.; Deshpande, M.; Nair, S.; Natassia, C.; Yordanova, G.; Yuan, D.; Stroe, O.; Wood, G.; Laydon, A.; et al. AlphaFold Protein Structure Database: Massively expanding the structural coverage of protein-sequence space with high-accuracy models. Nucleic Acids Res. 2022, 50, D439–D444. [Google Scholar] [CrossRef]
- Zhu, W.; Shenoy, A.; Kundrotas, P.; Elofsson, A. Evaluation of AlphaFold-Multimer Prediction on Multi-Chain Protein Complexes. Bioinformatics 2023, 39, btad424. [Google Scholar] [CrossRef]
- Ge, J.; Hsieh, C.-Y.; Fang, M.; Sun, H.; Hou, T. Development of PROTACs Using Computational Approaches. Trends Pharmacol. Sci. 2024, 45, 1162–1174. [Google Scholar] [CrossRef]
- Hu, X.; Liu, G.; Chen, C.; Zhao, Y.; Zhang, H.; Liu, X. TransDiffSBDD: Causality-Aware Multi-Modal Structure-Based Drug Design. arXiv 2025, arXiv:2503.20913. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef] [PubMed]
- Corso, G.; Stärk, H.; Jing, B.; Barzilay, R.; Jaakkola, T. DiffDock: Diffusion steps, twists, and turns for molecular docking. arXiv 2023, arXiv:2210.01776. [Google Scholar] [CrossRef]
- Dupont, M.; Didier, N.; Hodson, M.J.; Moore, J.E.; Reagor, M.J. Entanglement Perspective on the Quantum Approximate Optimization Algorithm. Phys. Rev. A 2022, 106, 022423. [Google Scholar] [CrossRef]
- Jafarnejad, S.; Baradaran, A.; Zhang, P.; Kumar, A.; Li, Y.; Wang, J.; Patel, R.; Johnson, M.; Chen, L.; Anderson, K.; et al. Cryo-electron microscopy-based drug design. Front. Mol. Biosci. 2024, 11, 1342179. [Google Scholar] [CrossRef]
- Robo, M.T.; Hayes, R.L.; Ding, X.; Pulawski, B.; Vilseck, J.Z. Fast Free Energy Estimates from λ-Dynamics with Bias for Relative Binding Free Energies. Nat. Commun. 2023, 14, 8515. [Google Scholar] [CrossRef]
- Du, Y.; Jamasb, A.R.; Guo, J.; Fu, T.; Harris, C.; Wang, Y.; Duan, C.; Liò, P.; Schwaller, P.; Blundell, T. Machine learning-aided generative molecular design. Nat. Mach. Intell. 2024, 6, 589–604. [Google Scholar] [CrossRef]
- Stärk, H.; Ganea, O.E.; Pattanaik, L.; Barzilay, R.; Jaakkola, T. EquiBind: Geometric deep learning for drug binding structure prediction. In Proceedings of the 39th International Conference on Machine Learning, Baltimore, MD, USA, 17–23 July 2022; Volume 162, pp. 20503–20521. [Google Scholar]
- Mehla, K.; Singh, P.K. Metabolic Regulation of Macrophage Polarization in Cancer. Trends Cancer 2019, 5, 822–834. [Google Scholar] [CrossRef] [PubMed]
- Jhaveri, K.L.; Neven, P.; Casalnuovo, M.L.; Kim, S.-B.; Tokunaga, E.; Aftimos, P.; Saura, C.; O’Shaughnessy, J.; Harbeck, N.; Carey, L.C.; et al. Imlunestrant with or without Abemaciclib in Advanced Breast Cancer. N. Engl. J. Med. 2025, 392, 1189–1202. [Google Scholar] [CrossRef]
- Huang, J.S.; Huang, S.T.; Sun, K.P.; Hong, Z.N.; Chen, L.W.; Kuo, Y.R.; Chen, Q. Health-Related Quality of Life in Children and Adolescents Undergoing Intraoperative Device Closure of Isolated Perimembranous Ventricular Septal Defects in Southeastern China. J. Cardiothorac. Surg. 2019, 14, 218. [Google Scholar] [CrossRef]
- Martin, T.; Usmani, S.Z.; Berdeja, J.G.; Agha, M.; Cohen, A.D.; Hari, P.; Avigan, D.; Deol, A.; Htut, M.; Lesokhin, A.; et al. Ciltacabtagene Autoleucel, an Anti-B-Cell Maturation Antigen Chimeric Antigen Receptor T-Cell Therapy, for Relapsed/Refractory Multiple Myeloma: CARTITUDE-1 2-Year Follow-Up.J. Clin. Oncol. 2023, 41, 1265–1274. [Google Scholar] [CrossRef]
- Zhang, Q.; Shao, B.; Tong, Z.; Ouyang, Q.; Wang, Y.; Xu, G.; Li, S.; Li, H. Phase Ib study of camrelizumab combined with apatinib and fuzuloparib in recurrent/metastatic triple-negative breast cancer. BMC Med. 2022, 20, 321. [Google Scholar] [CrossRef]
- Li, Z.; Zhang, Z. Histone Chaperones: A Multinodal Highway Network Inside the Cell. Mol. Cell 2023, 83, 1024–1026. [Google Scholar] [CrossRef]
- Hernando, C.; Ortega-Morillo, B.; Tapia, M.; Moragón, S.; Martínez, M.T.; Eroles, P.; Garrido-Cano, I.; Adam-Artigues, A.; Lluch, A.; Bermejo, B.; et al. Oral Selective Estrogen Receptor Degraders (SERDs) as a Novel Breast Cancer Therapy: Present and Future from a Clinical Perspective. Int. J. Mol. Sci. 2021, 22, 7812. [Google Scholar] [CrossRef]
- Wang, G. Fulvestrant as a reference antiestrogen and estrogen receptor degrader in preclinical studies: Treatment dosage, efficacy, and implications on development of new ER-targeting agents. Transl. Cancer Res. 2020, 9, 4464–4468. [Google Scholar] [CrossRef] [PubMed]
- Pappalardo, M.; Sipala, F.M.; Nicolosi, M.C.; Guccione, S.; Ronsisvalle, S. Recent Applications ofIn SilicoApproaches for Studying Receptor Mutations Associated with Human Pathologies. Molecules 2024, 29, 5349. [Google Scholar] [CrossRef] [PubMed]
- Bardia, A.; Cortés, J.; Bidard, F.C.; Neven, P.; Garcia-Sáenz, J.; Aftimos, P.; O’Shaughnessy, J.; Lu, J.; Tonini, G.; Scartoni, S.; et al. Elacestrant in ER+, HER2− metastatic breast cancer with ESR1-mutated tumors: Subgroup analyses from the phase III EMERALD trial by prior duration of endocrine therapy plus CDK4/6 inhibitor and in clinical subgroups. Clin. Cancer Res. 2024, 30, 4299–4309. [Google Scholar] [CrossRef]
- Sarfraz, A.; Sarfraz, M.; Javad, F.; Khalid, M.; Shah, B.; Gul, A.; Ganiyani, M.A.; Ismail, A.; Cheema, K. Elacestrant in hormone receptor-positive metastatic breast cancer: A post-hoc analysis. Explor. Target Antitumor Ther. 2025, 6, 1002293. [Google Scholar] [CrossRef]
- Cortés, J.; Bidard, F.C.; Bardia, A.; Kaklamani, V.G.; Vlachaki, I.; Tonini, G.; Habboubi, N.; Aftimos, P.G. 188O EMERALD trial analysis of patient-reported outcomes (PROs) in patients with ER+/HER2− advanced or metastatic breast cancer (mBC) comparing oral elacestrant vs standard of care (SoC) endocrine therapy. ESMO Open 2023, 8, 101377. [Google Scholar] [CrossRef]
- Novak, B.J.; Fraser, D.; Maloney, T.H. Transforming ocean conservation: Applying the genetic rescue toolkit. Genes 2020, 11, 209. [Google Scholar] [CrossRef]
- Donohue, J.K.; Calcaterra, M.J.; Fowler, J.R. Surgical Management of Hook of Hamate Fractures: A Systematic Review of Outcomes. J. Hand Surg. Glob. Online 2023, 6, 183–187. [Google Scholar] [CrossRef]
- Safwan, N.; Saadedine, M.; Shufelt, C.L.; Kapoor, E.; Kling, J.M.; Chaudhry, R.; Faubion, S.S. Menopause in the Workplace: Challenges, Impact, and Next Steps. Maturitas 2024, 185, 107983. [Google Scholar] [CrossRef]
- Ren, Z.; Liu, Y.; Cai, A.; Yu, Y.; Wang, X.; Lan, L.; Guo, X.; Yan, H.; Gao, X.; Li, H.; et al. Cannabidiol Represses miR-143 to Promote Cardiomyocyte Proliferation and Heart Regeneration After Myocardial Infarction. Eur. J. Pharmacol. 2024, 963, 176245. [Google Scholar] [CrossRef] [PubMed]
- Tebben, K.; Dia, A.; Serre, D. Determination of the Stage Composition of Plasmodium Infections from Bulk Gene Expression Data. mSystems 2022, 7, e0025822. [Google Scholar] [CrossRef]
- Zhang, W.; Zhang, P.; Wang, H.; Li, J.; Dai, S.Y. Design of Biomass-Based Renewable Materials for Environmental Remediation. Trends Biotechnol. 2022, 40, 1519–1534. [Google Scholar] [CrossRef]
- Fanucci, K.; Mayer, E.L. The State of the Science of Oral Selective Oestrogen Receptor Degraders. Lancet Oncol. 2024, 25, 1388–1389. [Google Scholar] [CrossRef]
- Robinson, D.R.; Wu, Y.M.; Vats, P.; Su, F.; Lonigro, R.J.; Cao, X.; Kalyana-Sundaram, S.; Wang, R.; Ning, Y.; Hodges, L.; et al. ActivatingESR1Mutations in Hormone-Resistant Metastatic Breast Cancer. Nat. Genet. 2013, 45, 1446–1451. [Google Scholar] [CrossRef] [PubMed]
- Young, K.S.; Hancock, G.R.; Fink, E.C.; Zigrossi, A.; Flowers, B.; Cooper, D.A.; Nguyen, V.T.; Martinez, M.C.; Mon, K.S.; Bosland, M.; et al. Targeting unique ligand binding domain structural features downregulates DKK1 in Y537S ESR1 mutant breast cancer cells. Breast Cancer Res. 2025, 27, 10. [Google Scholar] [CrossRef] [PubMed]
- Downton, T.; Zhou, F.; Segara, D.; Jeselsohn, R.; Lim, E. Oral selective estrogen receptor degraders (SERDs) in breast cancer: Advances, challenges, and current status. Drug Des. Dev. Ther. 2022, 16, 2933–2948. [Google Scholar] [CrossRef]
- Robertson, J.F.R.; Evans, A.; Henschen, S.; Kirwan, C.C.; Jahan, A.; Kenny, L.M.; Dixon, J.M.; Schmid, P.; Kothari, A.; Mohamed, O.; et al. A randomized, open-label, presurgical, window-of-opportunity study comparing the pharmacodynamic effects of the novel oral SERD AZD9496 with fulvestrant in patients with newly diagnosed ER+ HER2− primary breast cancer. Clin. Cancer Res. 2020, 26, 4242–4249. [Google Scholar] [CrossRef]
- Sabit, H.; Abbas, S.; El-Safoury, M.T.; Madkour, E.M.; Mahmoud, S.; Abdel-Ghany, S.; Albrahim, Y.; Al-Dhuayan, I.S.; Rashwan, S.; El-Hashash, A.; et al. Antibody-drug conjugates in breast cancer: Navigating innovations, overcoming resistance, and shaping future therapies. Biomedicines 2025, 13, 2227. [Google Scholar] [CrossRef]
- Hummer, A.M.; Abanades, B.; Deane, C.M. Advances in Computational Structure-Based Antibody Design. Curr. Opin. Struct. Biol. 2022, 74, 102379. [Google Scholar] [CrossRef]
- Lin, C.-J.; Xiao, W.-X.; Fu, T.; Jin, X.; Shao, Z.-M.; Di, G.-H. Calcifications in triple-negative breast cancer: Molecular features and treatment strategies. NPJ Breast Cancer 2023, 9, 26. [Google Scholar] [CrossRef] [PubMed]
- Jin, R.; Ye, Q.; Wang, J.; Cao, Z.; Jiang, D.; Wang, T.; Kang, Y.; Xu, W.; Hsieh, C.; Hou, T. AttABseq: An Attention-Based Deep Learning Prediction Method for Antigen-Antibody Binding Affinity Changes Based on Protein Sequences. Brief. Bioinform. 2024, 25, bbae304. [Google Scholar] [CrossRef] [PubMed]
- Mohite, P.; Yadav, V.; Pandhare, R.; Maitra, S.; Saleh, F.M.; Saleem, R.M.; Al-Malky, H.S.; Kumarasamy, V.; Subramaniyan, V.; Abdel-Daim, M.M.; et al. Revolutionizing Cancer Treatment: Unleashing the Power of Viral Vaccines, Monoclonal Antibodies, and Proteolysis-Targeting Chimeras in the New Era of Immunotherapy. ACS Omega 2024, 9, 7277–7295. [Google Scholar] [CrossRef]
- Chen, J.; Hao, L.; Qian, X.; Lin, L.; Pan, Y.; Han, X. Machine Learning Models Based on Immunological Genes to Predict the Response to Neoadjuvant Therapy in Breast Cancer Patients. Front. Immunol. 2022, 13, 948601. [Google Scholar] [CrossRef]
- Huang, Z.; Shao, W.; Han, Z.; Alkashash, A.M.; De la Sancha, C.; Parwani, A.V.; Nitta, H.; Hou, Y.; Wang, T.; Salama, P.; et al. Artificial Intelligence Reveals Features Associated with Breast Cancer Neoadjuvant Chemotherapy Responses from Multi-Stain Histopathologic Images. NPJ Precis. Oncol. 2023, 7, 14. [Google Scholar] [CrossRef]
- La, Z.; Chen, J.; Lu, X.; Lei, C.; Li, F.; Zhao, L.; Yi, Y. AI Microscope Facilitates Accurate Interpretation of HER2 Immunohistochemical Scores 0 and 1+ in Invasive Breast Cancer. Sci. Rep. 2025, 15, 13820. [Google Scholar] [CrossRef]
- Wang, Z.-H.; Li, Y.; Zhang, J.-Y.; Liu, Y.; Wang, X.; Wang, Y.; Chen, J.; Yang, Y.-H.; Li, H.-Y.; Zhang, S.-Y.; et al. Trastuzumab Resistance in HER2-Positive Breast Cancer: Mechanisms, Emerging Biomarkers and Targeting Agents. Front. Oncol. 2022, 12, 1006429. [Google Scholar] [CrossRef]
- Lin, N.U.; Murthy, R.K.; Abramson, V.; Anders, C.; Bachelot, T.; Bedard, P.L.; Borges, V.; Cameron, D.; Carey, L.A.; Chien, A.J.; et al. Tucatinib versus Placebo, Both in Combination With Trastuzumab and Capecitabine, for Previously Treated ERBB2 (HER2)-Positive Metastatic Breast Cancer in Patients With Brain Metastases: Updated Exploratory Analysis of the HER2CLIMB Randomized Clinical Trial. JAMA Oncol. 2023, 9, 197–205. [Google Scholar] [CrossRef]
- Saleh, K.; Khoury, R.; Khalife, N.; Chahine, C.; Ibrahim, R.; Tikriti, Z.; Le Cesne, A. Mechanisms of action and resistance to anti-HER2 antibody-drug conjugates in breast cancer. Cancer Drug Resist. 2024, 7, 22. [Google Scholar] [CrossRef] [PubMed]
- Poskus, M.D.; McDonald, J.; Laird, M.; Li, R.; Norcoss, K.; Zervantonakis, I.K. Rational Design of HER2-Targeted Combination Therapies to Reverse Drug Resistance in Fibroblast-Protected HER2+ Breast Cancer Cells. Cell. Mol. Bioeng. 2024, 17, 491–506. [Google Scholar] [CrossRef]
- Dent, R.; Oliveira, M.; Isakoff, S.J.; Im, S.-A.; Espié, M.; Blau, S.; Tan, A.R.; Saura, C.; Wongchenko, M.J.; Xu, N.; et al. Final results of the double-blind placebo-controlled randomized phase 2 LOTUS trial of first-line ipatasertib plus paclitaxel for inoperable locally advanced/metastatic triple-negative breast cancer. Breast Cancer Res. Treat. 2021, 189, 377–386. [Google Scholar] [CrossRef] [PubMed]
- Harding, J.J.; Fan, J.; Oh, D.Y.; Choi, H.J.; Kim, J.W.; Chang, H.M.; Bao, L.; Sun, H.C.; Macarulla, T.; Xie, F.; et al. Zanidatamab for HER2-amplified, unresectable, locally advanced or metastatic biliary tract cancer (HERIZON-BTC-01): A multicentre, single-arm, phase 2b study. Lancet Oncol. 2023, 24, 772–782. [Google Scholar] [CrossRef] [PubMed]
- Dent, R.A.; Kim, S.-B.; Oliveira, M.; Barrios, C.; O’Shaughnessy, J.; Isakoff, S.J.; Saji, S.; Freitas-Junior, R.; Philco, M.; Bondarenko, I.; et al. Ipatasertib plus Paclitaxel for Patients with PIK3CA/AKT1/PTEN-Altered Locally Advanced Unresectable or Metastatic Triple-Negative Breast Cancer in the IPATunity130 Phase III Trial. Clin. Cancer Res. 2024, 30, 4329–4338. [Google Scholar] [CrossRef]
- Danishuddin; Jamal, M.S.; Song, K.S.; Lee, K.W.; Kim, J.J.; Park, Y.M. Revolutionizing drug targeting strategies: Integrating artificial intelligence and structure-based methods in PROTAC development. Pharmaceuticals 2023, 16, 1649. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.K.; Matulonis, U.A. PARP Inhibitor Resistance Mechanisms and Implications for Post-Progression Combination Therapies. Cancers 2020, 12, 2054. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, K.; Hayashi, H.; Kawano, R.; Ishikawa, M.; Aimono, E.; Mizuno, T.; Kuroda, H.; Kojima, Y.; Niikura, N.; Kawanishi, A.; et al. BRCA1/2Reversion Mutations in a Pan-Cancer Cohort. Cancer Sci. 2024, 115, 635–647. [Google Scholar] [CrossRef]
- Wang, C.; Qu, L.; Li, S.; Yin, F.; Ji, L.; Peng, W.; Luo, H.; Lu, D.; Liu, X.; Chen, X.; et al. Discovery of First-in-Class Dual PARP and EZH2 Inhibitors for Triple-Negative Breast Cancer with Wild-Type BRCA. J. Med. Chem. 2021, 64, 12630–12650. [Google Scholar] [CrossRef]
- Rossi, T.; Iorio, E.; Chirico, M.; Pisanu, M.E.; Amodio, N.; Cantafio, M.E.G.; Perrotta, I.; Colciaghi, F.; Fiorillo, M.; Gianferrari, A.; et al. BET Inhibitors (BETi) Influence Oxidative Phosphorylation Metabolism by Affecting Mitochondrial Dynamics Leading to Alterations in Apoptotic Pathways in Triple-Negative Breast Cancer (TNBC) Cells. Cell Prolif. 2024, 57, e13730. [Google Scholar] [CrossRef]
- Karimpour, M.; Totonchi, M.; Behmanesh, M.; Montazeri, H. Pathway-Driven Analysis of Synthetic Lethal Interactions in Cancer Using Perturbation Screens. Life Sci. Alliance 2024, 7, e202302268. [Google Scholar] [CrossRef]
- Wang, X.Y.; Luo, H.Y. The Roles of Histone Acetylation Key Enzymes HAT, HDAC and BET Proteins in Neuropathic Pain: Selection of Drug Targets. Pharmacol. Res. 2025, 217, 107813. [Google Scholar] [CrossRef]
- Luque, M.; Sanz-Álvarez, M.; Santamaría, A.; Zazo, S.; Cristóbal, I.; de la Fuente, L.; Mínguez, P.; Eroles, P.; Rovira, A.; Albanell, J.; et al. Targeted therapy modulates the secretome of cancer-associated fibroblasts to induce resistance in HER2-positive breast cancer. Int. J. Mol. Sci. 2021, 22, 13297. [Google Scholar] [CrossRef]
- Guo, Y.; Zou, Y.; Chen, Y.; Deng, D.; Zhang, Z.; Liu, K.; Tang, M.; Yang, T.; Fu, S.; Zhang, C. Design, Synthesis and Biological Evaluation of Purine-Based Derivatives as Novel JAK2/BRD4(BD2) Dual Target Inhibitors. Bioorg. Chem. 2023, 132, 106386. [Google Scholar] [CrossRef] [PubMed]
- Liu, K.; Zhou, Z.; Gao, H.; Yang, F.; Qian, Y.; Jin, H.; Guo, Y.; Liu, Y.; Li, H.; Zhang, C.; et al. JQ1, a BET-Bromodomain Inhibitor, Inhibits Human Cancer Growth and Suppresses PD-L1 Expression. Cell Biol. Int. 2019, 43, 642–650. [Google Scholar] [CrossRef]
- Zhang, Y.; Zheng, L.; Ma, L.; Yin, F.; Luo, Z.; Li, S.; Jiang, Y.; Kong, L.; Wang, X. Discovery of Dual CDK6/BRD4 Inhibitor Inducing Apoptosis and Increasing the Sensitivity of Ferroptosis in Triple-Negative Breast Cancer. J. Med. Chem. 2024, 67, 21186–21207. [Google Scholar] [CrossRef]
- Andrieu, G.; Denis, G.V. Bromodomain and Extraterminal Proteins Regulate PD-L1/PD-1 Signaling in Breast Cancer. Cancer Immunol. Res. 2018, 6, A17. [Google Scholar] [CrossRef]
- Zheng, B.; Gold, S.; Iwanaszko, M.; Howard, B.C.; Wang, L.; Shilatifard, A. Distinct Layers of BRD4-PTEFb Reveal Bromodomain-Independent Function in Transcriptional Regulation. Mol. Cell 2023, 83, 2917–2931.e6. [Google Scholar] [CrossRef]
- Berlin, M.; Cantley, J.; Bookbinder, M.; Bortolon, E.; Broccatelli, F.; Cadelina, G.; Chan, E.W.; Chen, H.F.; Chen, X.; Cheng, Y.; et al. PROTACs Targeting BRM (SMARCA2) Afford SelectiveIn VivoDegradation over BRG1 (SMARCA4) and Are Active in BRG1 Mutant Xenograft Tumor Models. J. Med. Chem. 2024, 67, 1262–1313. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Yan, L.; Zhang, L.; Ma, H.; Wang, H.; Bu, P.; Xi, Y.; Lian, J. Genomic Characterization Reveals Distinct Mutational Landscapes and Therapeutic Implications Between Different Molecular Subtypes of Triple-Negative Breast Cancer. Sci. Rep. 2024, 14, 12386. [Google Scholar] [CrossRef]
- Masuda, H.; Harano, K.; Miura, S.; Wang, Y.; Hirota, Y.; Harada, O.; Jolly, M.K.; Matsunaga, Y.; Lim, B.; Wood, A.L.; et al. Changes in Triple-Negative Breast Cancer Molecular Subtypes in Patients Without Pathologic Complete Response After Neoadjuvant Systemic Chemotherapy. JCO Precis. Oncol. 2022, 6, e2000368. [Google Scholar] [CrossRef]
- Marqués, M.; Sorolla, M.A.; Urdanibia, I.; Parisi, E.; Hidalgo, I.; Morales, S.; Salud, A.; Sorolla, A. Are transcription factors plausible oncotargets for triple negative breast cancers? Cancers 2022, 14, 1101. [Google Scholar] [CrossRef]
- Chen, X.; Zhao, J.; Herjan, T.; Hong, L.; Liao, Y.; Liu, C.; Vasu, K.; Wang, H.; Thompson, A.; Fox, P.L.; et al. IL-17-Induced HIF1α Drives Resistance to Anti-PD-L1 via Fibroblast-Mediated Immune Exclusion. J. Exp. Med. 2022, 219, e20210693. [Google Scholar] [CrossRef]
- Sun, S.; Zhou, J.; Bai, Y.; Gao, W.; Lin, L.; Jiang, T.; You, C.; Gu, Y. Role of Oedema and Shrinkage Patterns for Prediction of Response to Neoadjuvant Chemotherapy and Survival Outcomes in Luminal Breast Cancer. Clin. Radiol. 2024, 79, e1010–e1020. [Google Scholar] [CrossRef]
- Zhang, Y.; You, C.; Pei, Y.; Yang, F.; Li, D.; Jiang, Y.; Shao, Z. Integration of Radiogenomic Features for Early Prediction of Pathological Complete Response in Patients with Triple-Negative Breast Cancer and Identification of Potential Therapeutic Targets. J. Transl. Med. 2022, 20, 256. [Google Scholar] [CrossRef]
- Caballo, M.; Sanderink, W.B.G.; Han, L.; Gao, Y.; Athanasiou, A.; Mann, R.M. Four-Dimensional Machine Learning Radiomics for the Pretreatment Assessment of Breast Cancer Pathologic Complete Response to Neoadjuvant Chemotherapy in Dynamic Contrast-Enhanced MRI. J. Magn. Reson. Imaging 2023, 57, 97–110. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, I.; Rahman, R. Radiogenomics as an Integrated Approach to Glioblastoma Precision Medicine. Curr. Oncol. Rep. 2024, 26, 1213–1222. [Google Scholar] [CrossRef] [PubMed]
- Bhattarai, S.; Saini, G.; Li, H.; Seth, G.; Fisher, T.B.; Janssen, E.A.M.; Kiraz, U.; Kong, J.; Aneja, R. Predicting Neoadjuvant Treatment Response in Triple-Negative Breast Cancer Using Machine Learning. Diagnostics 2024, 14, 74. [Google Scholar] [CrossRef]
- Li, W.; Zhao, X.; Ren, C.; Gao, S.; Han, Q.; Lu, M.; Li, X. The Therapeutic Role of γδT Cells in TNBC. Front. Immunol. 2024, 15, 1420107. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.; Lee, Y. Targeting the undruggable: Recent progress in PROTAC-induced transcription factor degradation. Cancers 2025, 17, 1871. [Google Scholar] [CrossRef]
- Kurokawa, K.; Mitsuishi, Y.; Shimada, N.; Kawakami, Y.; Miura, K.; Miyawaki, T.; Asao, T.; Ko, R.; Shukuya, T.; Shibayama, R.; et al. Association Between the Efficacy and Immune-Related Adverse Events of Pembrolizumab and Chemotherapy in Non-Small Cell Lung Cancer Patients: A Retrospective Study. BMC Cancer 2022, 22, 1047. [Google Scholar] [CrossRef]
- Im, S.A.; Cortes, J.; Cescon, D.W.; Yusof, M.M.; Iwata, H.; Masuda, N.; Takano, T.; Huang, C.S.; Chung, C.F.; Tsugawa, K.; et al. Results from the Randomized KEYNOTE-355 Study of Pembrolizumab Plus Chemotherapy for Asian Patients with Advanced TNBC. NPJ Breast Cancer 2024, 10, 79. [Google Scholar] [CrossRef] [PubMed]
- Pusztai, L.; Denkert, C.; O’Shaughnessy, J.; Cortes, J.; Dent, R.; McArthur, H.; Kümmel, S.; Bergh, J.; Park, Y.H.; Hui, R.; et al. Event-Free Survival by Residual Cancer Burden with Pembrolizumab in Early-Stage TNBC: Exploratory Analysis from KEYNOTE-522. Ann. Oncol. 2024, 35, 429–436. [Google Scholar] [CrossRef] [PubMed]
- Wood, S.J.; Gao, Y.; Lee, J.H.; Chen, J.S.C.; Wang, Q.; Meisel, J.L.; Li, X.X. High Tumor Infiltrating Lymphocytes Are Significantly Associated with Pathological Complete Response in Triple Negative Breast Cancer Treated with Neoadjuvant KEYNOTE-522 Chemoimmunotherapy. Breast Cancer Res. Treat. 2024, 205, 193–199. [Google Scholar] [CrossRef]
- McGregor, B.A.; Sonpayde, G.P.; Bellmunt, J. The Double Antibody Drug Conjugate (DAD) Phase I Trial: Sacituzumab Govitecan Plus Enfortumab Vedotin for Metastatic Urothelial Carcinoma. Ann. Oncol. 2024, 35, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Parmar, A.; Lu, B.D.; Luo, J.; Chan, K.K. Real-World Comparative Effectiveness and Safety of Pembrolizumab for PD-L1 ≥ 50% Metastatic Non-Small-Cell Lung Cancer. Future Oncol. 2024, 20, 2879–2888. [Google Scholar] [CrossRef]
- Ramalingam, S.S.; Thara, E.; Awad, M.M.; Dowlati, A.; Haque, B.; Stinchcombe, T.E.; Dy, G.K.; Spigel, D.R.; Lu, S.; Singh, N.I.; et al. JASPER: Phase 2 Trial of First-Line Niraparib Plus Pembrolizumab in Patients with Advanced Non-Small Cell Lung Cancer. Cancer 2022, 128, 65–74. [Google Scholar] [CrossRef]
- Donisi, C.; Pretta, A.; Pusceddu, V.; Ziranu, P.; Lai, E.; Puzzoni, M.; Mariani, S.; Massa, E.; Madeddu, C.; Scartozzi, M. Immunotherapy and cancer: The multi-omics perspective. Int. J. Mol. Sci. 2024, 25, 3563. [Google Scholar] [CrossRef]
- Krebs, M.G.; Delord, J.P.; Evans, T.R.J.; De Jonge, M.; Kim, S.W.; Meurer, M.; Postel-Vinay, S.; Lee, J.S.; Angell, H.K.; Rocher-Ros, V.; et al. Olaparib and Durvalumab in Patients with Relapsed Small Cell Lung Cancer (MEDIOLA): An Open-Label, Multicenter, Phase 1/2, Basket Study. Lung Cancer 2023, 180, 107216. [Google Scholar] [CrossRef]
- Wilton, J.; Abdulmenan, J.; Janjua, N.Z. Cohort Profile: The British Columbia COVID-19 Cohort (BCC19C)-A Dynamic, Linked Population-Based Cohort. Front. Public Health 2024, 12, 1248905. [Google Scholar] [CrossRef]
- Smoots, S.G.; Schreiber, A.R.; Pitts, T.M. Overcoming Doxorubicin Resistance in Triple-Negative Breast Cancer Using the Class I-Targeting HDAC Inhibitor Bocodepsin/OKI-179 to Promote Apoptosis. Breast Cancer Res. 2024, 26, 35. [Google Scholar] [CrossRef]
- da Rocha, M.N.; de Sousa, D.S.; Marinho, E.S. Ligand and Structure-Based Virtual Screening Approaches in Drug Discovery: Minireview. Mol. Divers. 2025, 29, 2799–2809. [Google Scholar] [CrossRef] [PubMed]
- Matricon, P.; Nguyen, A.T.N.; Vo, D.D.; Baltos, J.A.; Jaiteh, M.; Luttens, A.; Kampen, S.; Christopoulos, A.; Kihlberg, J.; May, L.T.; et al. Structure-Based Virtual Screening Discovers Potent and Selective Adenosine A1 Receptor Antagonists. Eur. J. Med. Chem. 2023, 257, 115419. [Google Scholar] [CrossRef]
- Berlin, K.; O’Leary, D.P.; Fushman, D. Fast Approximations of the Rotational Diffusion Tensor and Their Application to Structural Assembly of Molecular Complexes. Proteins 2011, 79, 2268–2281. [Google Scholar] [CrossRef]
- Isett, C.; Atz, K.; Schneider, G. Exploring Protein-Ligand Binding Affinity Prediction with Electron Density-Based Geometric Deep Learning. RSC Adv. 2024, 14, 4492–4502. [Google Scholar] [CrossRef]
- Ahmed, M.; Maldonado, A.M.; Durrant, J.D. From Byte to Bench to Bedside: Molecular Dynamics Simulations and Drug Discovery. BMC Biol. 2023, 21, 299. [Google Scholar] [CrossRef]
- Song, L.F.; Merz, K.M., Jr. Evolution of Alchemical Free Energy Methods in Drug Discovery. J. Chem. Inf. Model. 2020, 60, 5308–5318. [Google Scholar] [CrossRef]
- Cai, H.; Zhang, H.; Zhao, D.; Wu, J.; Wang, L. FP-GNN: A Versatile Deep Learning Architecture for Enhanced Molecular Property Prediction. Brief. Bioinform. 2022, 23, bbac408. [Google Scholar] [CrossRef]
- Fralish, Z.; Chen, A.; Skaluba, P.; Reker, D. DeepDelta: Predicting ADMET Improvements of Molecular Derivatives with Deep Learning. J. Cheminform. 2023, 15, 69. [Google Scholar] [CrossRef]
- Yang, W.; Duan, L.; Zhao, X.; Niu, L.; Wang, C.; Fan, D.; Hong, L. Integration of Machine Learning in Biomarker Discovery for Esophageal Squamous Cell Carcinoma: Applications and Future Directions. Pathol. Res. Pract. 2025, 272, 156083. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wang, S.; Li, Y.; Guo, J.; Wei, Y.; Mu, Y.; Zheng, L.; Li, W. A New Paradigm for Applying Deep Learning to Protein-Ligand Interaction Prediction. Brief. Bioinform. 2024, 25, bbae145. [Google Scholar] [CrossRef] [PubMed]
- Dhakal, A.; McKay, C.; Tanner, J.J.; Cheng, J. Artificial Intelligence in the Prediction of Protein-Ligand Interactions: Recent Advances and Future Directions. Brief. Bioinform. 2022, 23, bbab476. [Google Scholar] [CrossRef]
- Fluetsch, A.; Di Lascio, E.; Gerebtzoff, G.; Rodriguez-Pérez, R. Adapting Deep Learning QSPR Models to Specific Drug Discovery Projects. Mol. Pharm. 2024, 21, 1817–1826. [Google Scholar] [CrossRef] [PubMed]
- Shihoya, W.; Iwama, A.; Sano, F.K.; Nureki, O. Cryo-EM Advances in GPCR Structure Determination. J. Biochem. 2024, 176, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Bryant, P.; Noé, F. Improved Protein Complex Prediction with AlphaFold-Multimer by Denoising the MSA Profile. PLoS Comput. Biol. 2024, 20, e1012253. [Google Scholar] [CrossRef] [PubMed]
- Nam, K.H. Evaluation of AlphaFold 3 for the Fatty Acids Docking to Human Fatty Acid-Binding Proteins. J. Mol. Graph. Model. 2024, 131, 108872. [Google Scholar] [CrossRef]
- Zhang, R.; Wen, H.; Lin, Z. Artificial Intelligence-Driven Drug Toxicity Prediction: Advances, Challenges, and Future Directions. Toxics 2025, 13, 525. [Google Scholar] [CrossRef]
- York, D.M. Modern alchemical free energy methods for drug discovery explained. ACS Phys. Chem. Au. 2023, 3, 478–491. [Google Scholar] [CrossRef]
- Renaud, N.; Jung, Y.; Honavar, V.; Geng, C.; Bonvin, A.M.J.J.; Xue, L.C. iScore: An MPI Supported Software for Ranking Protein-Protein Docking Models Based on a Random Walk Graph Kernel and Support Vector Machines. SoftwareX 2020, 11, 100462. [Google Scholar] [CrossRef]
- Federico, A.; Fratello, M.; Scala, G.; Möbus, L.; Pavel, A.; Del Giudice, G.; Ceccarelli, M.; Costa, V.; Ciccodicola, A.; Fortino, V.; et al. Integrated Network Pharmacology Approach for Drug Combination Discovery: A Multi-Cancer Case Study. Cancers 2022, 14, 2043. [Google Scholar] [CrossRef]
- Vittorio, S.; Lunghini, F.; Morerio, P.; Gadioli, D.; Orlandini, S.; Silva, P.; Martinovic, J.; Pedretti, A.; Bonanni, D.; Del Bue, A.; et al. Addressing Docking Pose Selection with Structure-Based Deep Learning: Recent Advances, Challenges and Opportunities. Comput. Struct. Biotechnol. J. 2024, 23, 2141–2151. [Google Scholar] [CrossRef]
- Weisser, N.E.; Sanches, M.; Escobar-Cabrera, E.; O’Toole, J.; Whalen, E.; Chan, P.W.Y.; Wickman, G.; Abraham, L.; Choi, K.; Harbourne, B.; et al. An Anti-HER2 Biparatopic Antibody That Induces Unique HER2 Clustering and Complement-Dependent Cytotoxicity. Nat. Commun. 2023, 14, 1394. [Google Scholar] [CrossRef]
- Hu, Y.; Li, Z.; Zhao, W.; Wang, H.; Zhou, J.; Li, X.; Zhang, L.; Zhao, Y.; Chen, Y. Machine Learning-Developed LKB1-AMPK Signaling-Related Signature for Prognosis and Drug Sensitivity in Hepatocellular Carcinoma. Sci. Rep. 2025, 15, 20738. [Google Scholar] [CrossRef]
- Popovic, L.S.; Matovina-Brko, G.; Popovic, M.; Punie, K.; Cvetanovic, A.; Lambertini, M. Targeting triple-negative breast cancer: A clinical perspective. Oncol. Res. 2023, 31, 221–238. [Google Scholar] [CrossRef]
- Challita-Eid, P.M.; Satpayev, D.; Yang, P.; An, Z.; Morrison, K.; Shostak, Y.; Raitano, A.; Nadell, R.; Liu, W.; Lortie, D.R.; et al. Enfortumab Vedotin Antibody-Drug Conjugate Targeting Nectin-4 Is a Highly Potent Therapeutic Agent in Multiple Preclinical Cancer Models. Cancer Res. 2016, 76, 3003–3013. [Google Scholar] [CrossRef]
- Gentile, F.; Yaacoub, J.-C.; Gleave, J.; Fernandez, M.; Ton, A.-T.; Ban, F.; Stern, A.; Cherkasov, A. Artificial intelligence-enabled virtual screening of ultra-large chemical libraries with deep docking. Nat. Protoc. 2022, 17, 672–697. [Google Scholar] [CrossRef]
- Yang, K.; Swanson, K.; Jin, W.; Coley, C.; Eiden, P.; Gao, H.; Guzman-Perez, A.; Hopper, T.; Kelley, B.; Mathea, M.; et al. Analyzing Learned Molecular Representations for Property Prediction. J. Chem. Inf. Model. 2019, 59, 3370–3388. [Google Scholar] [CrossRef]
- Chen, Q.; Liang, X.; Wu, T.; Jiang, J.; Jiang, Y.; Zhang, S.; Ruan, Y.; Zhang, H.; Zhang, C.; Chen, P.; et al. Integrative Analysis of Metabolomics and Proteomics Reveals Amino Acid Metabolism Disorder in Sepsis. J. Transl. Med. 2022, 20, 123. [Google Scholar] [CrossRef] [PubMed]
- Askin, S.; Burkhalter, D.; Calado, G.; El Dakrouni, S. Artificial Intelligence Applied to Clinical Trials: Opportunities and Challenges. Health Technol. 2023, 13, 203–211. [Google Scholar] [CrossRef]
- Zhang, Y.; Li, X.; Wang, Z.; Chen, H. A Comprehensive Review of Deep Learning Applications with Multi-Omics. Genes 2025, 16, 648. [Google Scholar] [CrossRef] [PubMed]
- Tobias, A.V.; Wahab, A. Autonomous ‘Self-Driving’ Laboratories: A Review of Technology and Policy Implications. R. Soc. Open Sci. 2025, 12, 250646. [Google Scholar] [CrossRef]
- Butakova, M.A.; Chernov, A.V.; Kartashov, O.O.; Soldatov, A.V. Data-Centric Architecture for Self-Driving Laboratories with Autonomous Discovery of New Nanomaterials. Nanomaterials 2022, 12, 12. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Li, J.; Xu, Y.; Wang, J.; Zhang, H.; Chen, Y.; Liu, M.; Zhao, X.; Sun, L.; Zhou, Q.; et al. The role of artificial intelligence in drug screening, drug design, and clinical trials. Front. Pharmacol. 2024, 15, 1459954. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Wang, Y.; Zhang, Y.; Zhao, H.; Xu, J.; Chen, L.; Liu, Q.; Yang, F. Advances in spatial transcriptomics and related data analysis strategies. J. Transl. Med. 2023, 21, 276. [Google Scholar] [CrossRef] [PubMed]

| Subtype and Features | Design Focus | CADD Methods | Tools/Software/Servers | Representative Agents |
|---|---|---|---|---|
| Luminal-ER+/PR+; typically HER2−; frequent ESR1 and PIK3CA mutations; endocrine resistance/brain exposure considerations | ER Degradation | Docking | AutoDock Vina; Glide; GOLD; GNINA | Elacestrant [48,49,50,51,52]; GDC-0810 [53]; Camizestrant [54]; Imlunestrant [55]; AZD9496 [56,57,58,59,60,61,62,63,64] |
| VS | ZINC; Enamine REAL | Elacestrant [48,49,50,51,52]; Imlunestrant [55] | ||
| MD simulations | GROMACS; AMBER; OpenMM | Elacestrant [48,49,50,51,52]; Imlunestrant [55]; ESR1-Y537S/D538G mutants [58,59] | ||
| Structural Insights & Rational Design | Cryo-EM | RELION; cryoSPARC; EMDB | GDC-0810 structural insights [57] | |
| X-ray Crystallography | Phenix; Coot; PDB | GDC-0810 [53]; ESR1 mutant scaffolds [58,59] | ||
| NMR | TopSpin; NMRPipe | Allosteric SERD scaffolds [52] | ||
| AlphaFold Predictions | AlphaFold/ColabFold; MODELLER; SWISS-MODEL; Rosetta | ESR1-Y537S/D538G mutant studies [58,59] | ||
| Free-Energy Calculations | FEP+; PyAutoFEP (OpenMM) | GDC-0810 [53]; conformational refinements [57,58] | ||
| Pharmacokinetics/ADME Optimization | QSAR Modeling; ADME PredictionML Models | scikit-learn; Chemprop; DeepChem; pkCSM; ADMETlab; ADMET-AI | Elacestrant [48,49,50,51,52]; AZD9496 [60,61,62,63,64] | |
| PBPK simulations | GastroPlus; Simcyp | Elacestrant [48,49,50,51,52] | ||
| HER2+-HER2 overexpression/amplification (IHC 3+ or ISH+); frequent PIK3CA co-alterations; spatial heterogeneity; higher CNS risk+ | Antibodies, ADCs, and TKIs | Docking | AutoDock Vina; Glide; GNINA | Tucatinib [72]; Lapatinib [72]; Neratinib [72] |
| Pharmacophore Modeling | Rosetta; MODELLER; SWISS-MODE | Trastuzumab [72]; Pertuzumab [72] | ||
| MD | GROMACS; OpenMM | T-DM1 [72]; T-DXd [78] | ||
| AI-Assisted Antibody Design | RosettaAntibody; DeepAb; IgFold | Zanidatamab [132] | ||
| Targeted HER2 Degradation (PROTACs) | Linker Optimization | OpenMM; Rosetta | Tucatinib-based PROTAC [69] | |
| E3 Ligase Selection | PRosettaC; Rosetta | HER2-directed PROTACs [75,76] | ||
| Ternary Complex Docking & MD | Rosetta; OpenMM | HER2 PROTAC prototypes [75,76] | ||
| Multi-Omics Integration | Multi-Omics ML | scikit-learn (survival/classifiers) | PI3K-mutant subgroup predictions [77] | |
| AI-Driven Pathology | QuPath; HALO AI; MONAI; CLAM | Trastuzumab deruxtecan-sensitive subtypes [78] | ||
| Digital Pathology (HER2 IHC) | QuPath; MONAI | Automated HER2 IHC scoring [65] | ||
| TNBC-ER−/PR−/HER2−; often basal-like; BRCA1/2 or high HRD; genomic instability; TROP-2 expression | Pathway Targeting (PI3K/AKT, DDR, Epigenetic) | Docking | AutoDock Vina; Glide | AKT inhibitors [73,75] |
| MD | OpenMM; GROMACS | PARP inhibitors [85,86]; PARP/HDAC hybrids [79,83] | ||
| ADME/T Profiling | pkCSM; ADMETlab; ADMET-AI | BET inhibitors [93,94,96] | ||
| Dual-Warhead Modeling | Rosetta; OpenMM | EGFR/BRD4 dual inhibitors [84,86]; JAK2/BRD4 dual inhibitors [95] | ||
| PROTACs for Transcriptional Regulators | Linker Design | OpenMM; Rosetta | BRD4 PROTACs (CRBN recruiting) [75,76] | |
| Ternary Complex MD | OpenMM; Rosetta | BRD4 PROTACs suppressing KLF5 [75,76] | ||
| Solvent Mapping | FTMap; MixMD | BRD4 PROTACs in TNBC models [75,76] | ||
| Multi-Omics Integration | Subtype-Guided Docking | AutoDock Vina; Glide | BRD4 inhibitors in basal-like TNBC [95] | |
| ML Predictors | Chemprop; scikit-learn | Subtype-specific stratification [95] | ||
| Radiomics/Radiogenomics | PyRadiomics; scikit-image; MONAI | Imaging-guided therapeutic ranking [95] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tian, W.; Hu, Y.; Gao, X.; Yang, J.; Jiang, W. Computer-Aided Drug Design Across Breast Cancer Subtypes: Methods, Applications and Translational Outlook. Int. J. Mol. Sci. 2025, 26, 10744. https://doi.org/10.3390/ijms262110744
Tian W, Hu Y, Gao X, Yang J, Jiang W. Computer-Aided Drug Design Across Breast Cancer Subtypes: Methods, Applications and Translational Outlook. International Journal of Molecular Sciences. 2025; 26(21):10744. https://doi.org/10.3390/ijms262110744
Chicago/Turabian StyleTian, Wei, Ying Hu, Xinyu Gao, Jinghui Yang, and Wei Jiang. 2025. "Computer-Aided Drug Design Across Breast Cancer Subtypes: Methods, Applications and Translational Outlook" International Journal of Molecular Sciences 26, no. 21: 10744. https://doi.org/10.3390/ijms262110744
APA StyleTian, W., Hu, Y., Gao, X., Yang, J., & Jiang, W. (2025). Computer-Aided Drug Design Across Breast Cancer Subtypes: Methods, Applications and Translational Outlook. International Journal of Molecular Sciences, 26(21), 10744. https://doi.org/10.3390/ijms262110744







