Steroid Hydroxylation by Mutant Cytochrome P450 BM3-LG23 Using Two Expression Chassis
Abstract
1. Introduction
2. Results
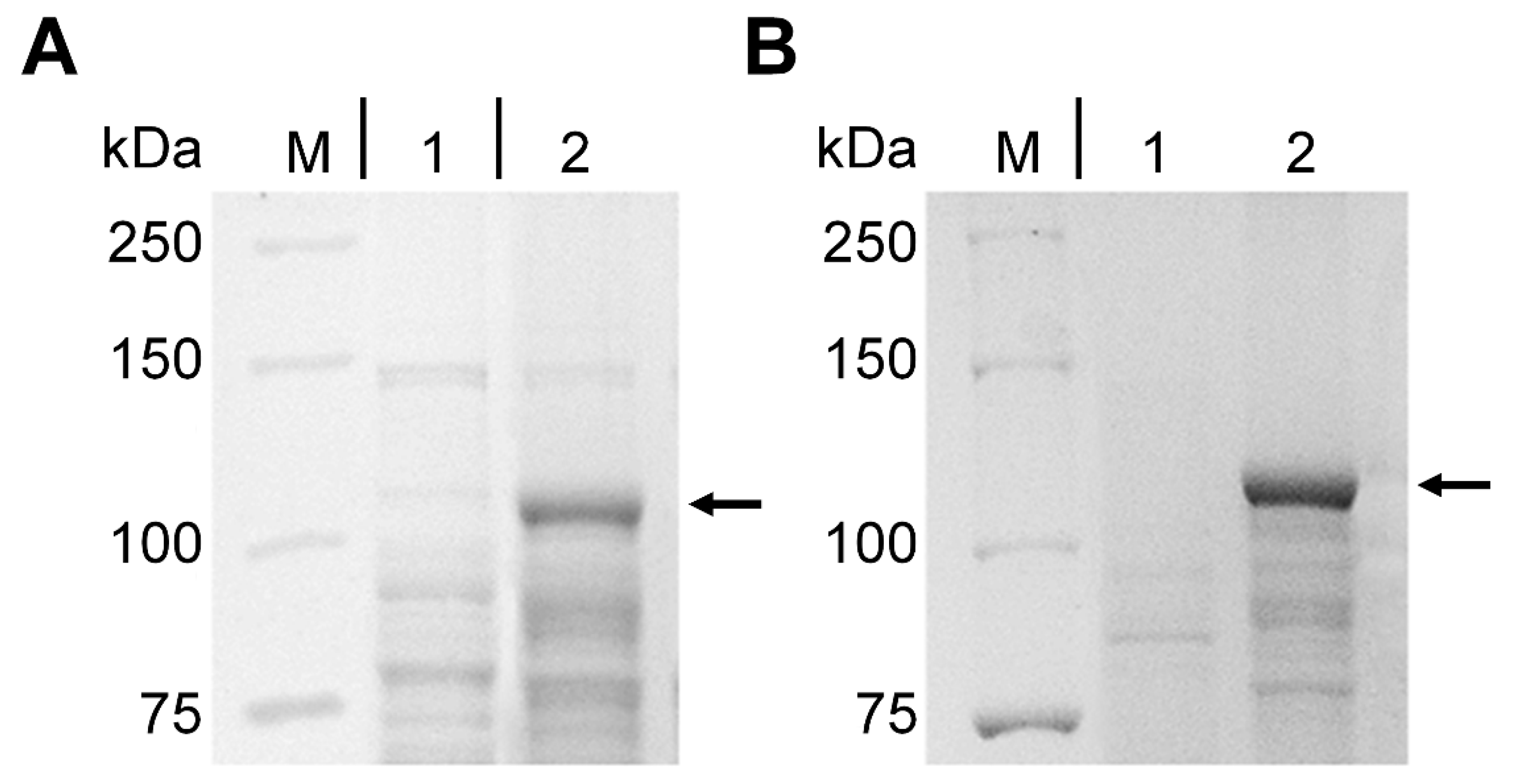
2.1. Expression of cyp102A1-LG23
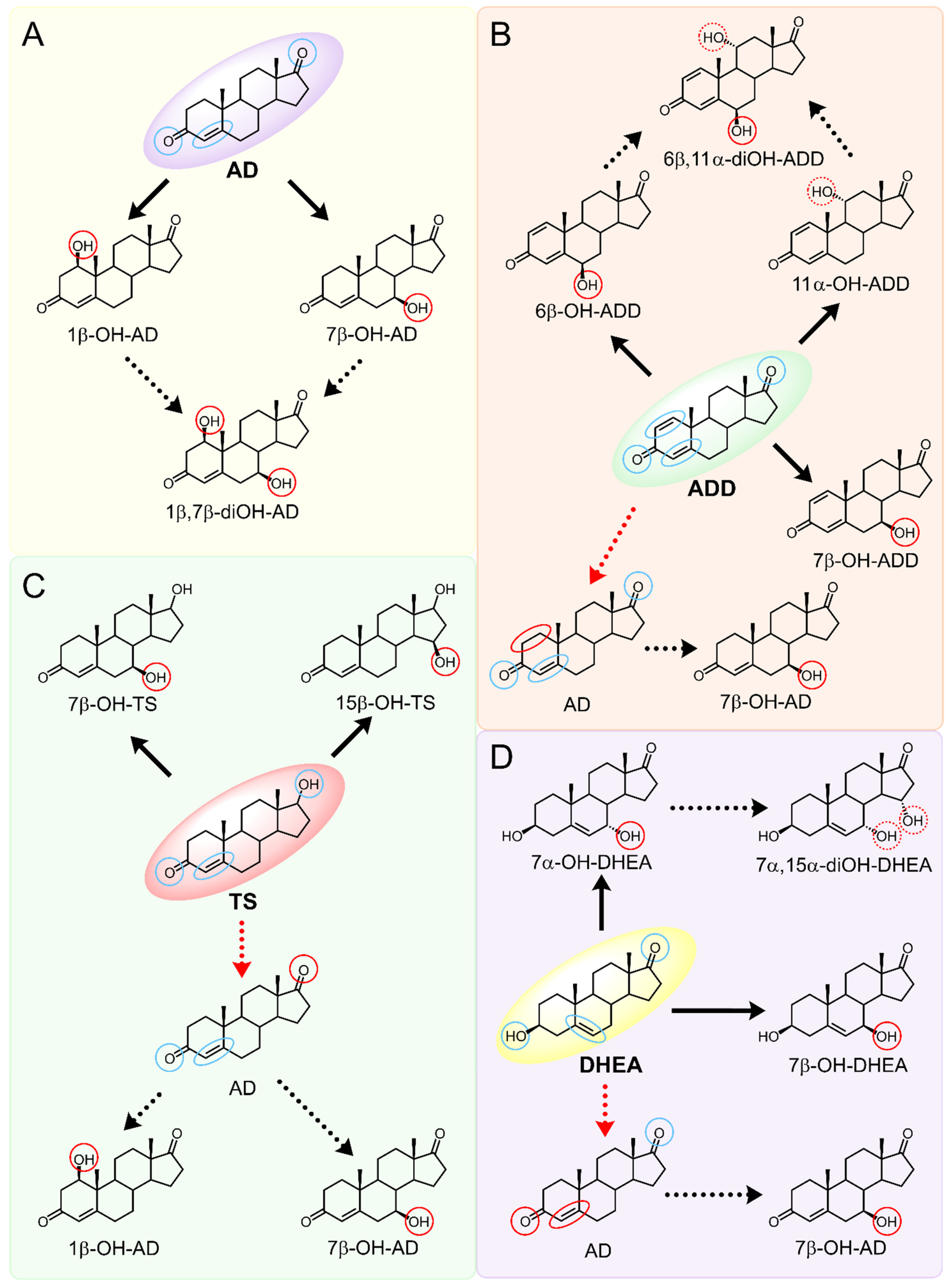
2.2. Bioconversion of AD
2.3. Bioconversion of ADD
2.4. Bioconversion of Testosterone
2.5. Bioconversion of DHEA
3. Discussion
4. Materials and Methods
4.1. Chemicals
4.2. Strains and Plasmids
4.3. Heterologous Expression
4.4. Steroid Bioconversion
4.5. Thin Layer Chromatography (TLC)
4.6. High Performance Liquid Chromatography (HPLC)
4.7. Steroid Isolation
4.8. Mass-Spectrometry (MS) and 1H NMR Spectroscopy
4.9. Statistical Data Processing
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pellissier, H.; Santelli, M. Chemical and biochemical hydroxylations of steroids, a review. Org. Prep. Proced. Int. 2001, 33, 1–58. [Google Scholar] [CrossRef]
- Berrie, J.R.; Williams, R.A.D.; Smith, K.E. Microbial transformations of steroids-XI. Progesterone transformation by Streptomyces roseochromogenes—Purification and characterisation of the 16α-hydroxylase system. J. Steroid Biochem. Mol. Biol. 1999, 71, 153–165. [Google Scholar] [CrossRef]
- Prosser, D.E.; Jones, G. Enzymes involved in the activation and inactivation of vitamin D. Trends Biochem. Sci. 2004, 29, 664–673. [Google Scholar] [CrossRef]
- Shah, A.A.; Ultanultan, S.S.; Adnan, H.S. A whole-cell biocatalysis application of steroidal drugs. Orient. J. Chem. 2013, 29, 389–403. [Google Scholar] [CrossRef]
- Marcello, A.; Civra, A.; Bonotto, R.M.; Alves, L.N.; Rajasekharan, S.; Giacobone, C.; Caccia, C.; Cavalli, R.; Adami, M.; Brambilla, P.; et al. The cholesterol metabolite 27-hydroxycholesterol inhibits SARS-CoV-2 and is markedly decreased in COVID-19 patients. Redox Biol. 2020, 36, 101682. [Google Scholar] [CrossRef] [PubMed]
- Traish, A.M.; Kang, H.P.; Saad, F.; Guay, A.T. Dehydroepiandrosterone (DHEA)—A precursor steroid or an active hormone in human physiology. J. Sex. Med. 2011, 8, 2960–2982. [Google Scholar] [CrossRef] [PubMed]
- Kozłowska, E.; Urbaniak, M.; Kancelista, A.; Dymarska, M.; Kostrzewa-Susłow, E.; Łukasz, S.; Janeczko, T. Biotransformation of dehydroepiandrosterone (DHEA) by environ-mental strains of filamentous fungi. RSC Adv. 2017, 7, 31493–31501. [Google Scholar] [CrossRef]
- Wulfert, E.; Pringle, A.K.; Sundstrom, L.E. Neuroprotective 7-Beta-Hydroxysteroids. Patent WO2002000224A1, 3 January 2002. [Google Scholar]
- Tonin, F.; Arends, I.W.C.E. Latest development in the synthesis of ursodeoxycholic acid (UDCA): A critical review. Beilstein J. Org. Chem. 2018, 14, 470–483. [Google Scholar] [CrossRef]
- Eggert, T.; Bakonyi, D.; Hummel, W. Enzymatic routes for the synthesis of ursodeoxycholic acid. J. Biotechnol. 2014, 191, 11–21. [Google Scholar] [CrossRef]
- Haal, S.; Guman, M.S.S.; Boerlage, T.C.C.; Acherman, Y.I.Z.; Brauw, D.L.M.; Bruin, S.; de Castro, S.M.M.; van Hooft, J.E.; van de Laar, A.W.J.M.; Moes, D.E.; et al. Ursodeoxycholic acid for the prevention of symptomatic gallstone disease after bariatric surgery (UPGRADE): A multicentre, double-blind, randomised, placebo-controlled superiority trial. Lancet Gastroenterol. Hepatol. 2021, 6, 993–1001. [Google Scholar] [CrossRef]
- Byelyayeva, N. Effectiveness of ursodeoxycholic acid (UDCA) in patients with chronic pancreatitis (CP) and biliary sludge. Pancreatology 2015, 15, 69. [Google Scholar] [CrossRef]
- Colombo, C.; Alicandro, G.; Oliver, M.; Lewindon, P.J.; Ramm, G.A.; Ooi, C.Y.; Alghisi, F.; Kashirskaya, N.; Kondratyeva, E.; Corti, F.; et al. Ursodeoxycholic acid and liver disease associated with cystic fibrosis: A multicenter cohort study. J. Cyst. Fibros. 2021, 21, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Abdelkader, N.F. Ursodeoxycholic and tauroursodeoxycholic acids as antiapoptotic agents: Modulation of Parkinson’s disease. Diagn. Manag. Park. Dis. 2020, 1, 653–664. [Google Scholar] [CrossRef]
- Huang, F. Ursodeoxycholic acid as a potential alternative therapeutic approach for neurodegenerative disorders: Effects on cell apoptosis, oxidative stress and inflammation in the brain. Brain Behav. Immun. Health 2021, 18, 100348. [Google Scholar] [CrossRef]
- Goossens, J.F.; Bailly, C. Ursodeoxycholic acid and cancer: From chemoprevention to chemotherapy. Pharmacol. Ther. 2019, 203, 107396. [Google Scholar] [CrossRef] [PubMed]
- Lugini, A.; Verna, S.; Buzzacchino, F.; Minelli, M.; Cristofani, L. 1134P Prevention of hepatic toxicities associated with anaplastic lymphoma kinase inhibitors in the treatment of non-small cell lung cancer by administration of ursodeoxycholic acid: Analysis from the monoinstitutional analysis. Ann. Oncol. 2022, 33, 1069. [Google Scholar] [CrossRef]
- Loewenthal, H. Selective reactions and modification of functional groups in steroid chemistry. Tetrahedron 1959, 6, 269–303. [Google Scholar] [CrossRef]
- Nassiri-Koopaei, N.; Faramarzi, M.A. Recent developments in the fungal transformation of steroids. Biocatal. Biotransform. 2015, 33, 1–28. [Google Scholar] [CrossRef]
- Munro, A.W.; Daff, S.; Coggins, J.R.; Lindsay, J.G.; Chapman, S.K. Probing electron transfer in flavocytochrome P-450 BM3 and its component domains. Eur. J. Biochem. 1996, 239, 403–409. [Google Scholar] [CrossRef]
- Munro, A.W.; Leys, D.G.; McLean, K.J.; Marshall, K.R.; Ost, T.W.B.; Daff, S.; Miles, C.S.; Chapman, S.K.; Lysek, D.A.; Moser, C.C.; et al. P450 BM3: The very model of a modern flavocytochrome. Trends Biochem. Sci. 2002, 27, 250–257. [Google Scholar] [CrossRef]
- Cirino, P.C.; Arnold, F.H. A self-sufficient peroxide-driven hydroxylation biocatalyst. Angew. Chem. Int. Ed. 2003, 42, 3299–3301. [Google Scholar] [CrossRef] [PubMed]
- Girvan, H.M.; Munro, A.W. Applications of microbial cytochrome P450 enzymes in biotechnology and synthetic biology. Curr. Opin. Chem. Biol. 2016, 31, 136–145. [Google Scholar] [CrossRef] [PubMed]
- Urlacher, V.B.; Girhard, M. Cytochrome P450 monooxygenases: An update on perspectives for synthetic application. Trends Biotechnol. 2012, 30, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Kille, S.; Zilly, F.E.; Acevedo, J.P.; Reetz, M.T. Regio- and stereoselectivity of P450-catalysed hydroxylation of steroids controlled by laboratory evolution. Nat. Chem. 2011, 3, 738–743. [Google Scholar] [CrossRef]
- Li, A.; Acevedo-Rocha, C.G.; D’Amore, L.; Chen, J.; Peng, Y.; Garcia-Borràs, M.; Gao, C.; Zhu, J.; Rickerby, H.; Osuna, S.; et al. Regio- and stereoselective steroid hydroxylation at C7 by cytochrome P450 monooxygenase mutants. Angew. Chem. Int. Ed. 2020, 59, 12499–12505. [Google Scholar] [CrossRef]
- Poshekhontseva, V.Y.; Strizhov, N.I.; Karpov, M.V.; Nikolaeva, V.M.; Kazantsev, A.V.; Sazonova, O.I.; Shutov, A.A.; Donova, M.V. Expression of Synthetic cyp102A1-LG23 Gene and Functional Analysis of Recombinant Cytochrome P450 BM3-LG23 in the Actinobacterium Mycolicibacterium smegmatis. Biochemistry 2023, 88, 1347–1355. [Google Scholar] [CrossRef]
- Miura, Y.; Fulco, A.J. (ω–2) Hydroxylation of Fatty Acids by a Soluble System from Bacillus megaterium. J. Biol. Chem. 1974, 249, 1880–1888. [Google Scholar] [CrossRef]
- Cha, G.S.; Ryu, S.H.; Ahn, T.; Yun, C.-H. Regioselective hydroxylation of 17β-estradiol by mutants of CYP102A1 from Bacillus megaterium. Biotechnol. Lett. 2014, 36, 2501–2506. [Google Scholar] [CrossRef]
- Thistlethwaite, S.; Jeffreys, L.N.; Girvan, H.M.; McLean, K.J.; Munro, A.W. A promiscuous bacterial P450: The unparalleled diversity of BM3 in pharmaceutical metabolism. Int. J. Mol. Sci. 2021, 22, 11380. [Google Scholar] [CrossRef]
- Venkataraman, H.; de Beer, S.B.A.; van Bergen, L.A.H.; van Essen, N.; Geerke, D.P.; Vermeulen, N.P.E.; Commandeur, J.N.M. A single active site mutation inverts stereoselectivity of 16-hydroxylation of testosterone catalyzed by engineered cytochrome P450 BM3. ChemBioChem 2012, 13, 520–523. [Google Scholar] [CrossRef]
- Dodson, R.M.; Kraychy, S.; Nicholson, R.T.; Mizuba, S. Microbiological transformations. IX. The 1β-hydroxylation of androstenedione. J. Org. Chem. 1962, 27, 3159–3164. [Google Scholar] [CrossRef]
- Li, H.; Dai, W.; Qin, S.; Li, S.; Yu, Y.; Zhang, L. Regio- and stereo-selective 1β-hydroxylation of lithocholic acid by cytochrome P450 BM3 mutants. Biotechnol. Bioeng. 2023, 120, 2230–2241. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wang, Z.-B.; Wang, Y.-N.; Kong, J.-Q. Probing steroidal substrate specificity of cytochrome P450 BM3 variants. Molecules 2016, 21, 760. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Kong, J.-Q. Steroids hydroxylation catalyzed by the monooxygenase mutant 139-3 from Bacillus megaterium BM3. Acta Pharm. Sin. B 2017, 7, 510–516. [Google Scholar] [CrossRef] [PubMed]
- Lobastova, T.G.; Gulevskaya, S.A.; Sukhodolskaya, G.V.; Donova, M.V. Dihydroxylation of dehydroepiandrosterone in positions 7α and 15α by mycelial fungi. Appl. Biochem. Microbiol. 2009, 45, 617–622. [Google Scholar] [CrossRef]
- Wu, Y.; Li, H.; Zhang, X.-M.; Gong, J.-S.; Li, H.; Rao, Z.-M.; Shi, Z.-H.; Xu, J.-S. Improvement of NADPH-dependent P450-mediated biotransformation of 7a,15a-diOH-DHEA from DHEA by a dual cosubstrate-coupled system. Steroids 2015, 101, 15–20. [Google Scholar] [CrossRef]
- Zhao, Y.-Q.; Liu, Y.-J.; Ji, W.-T.; Liu, K.; Gao, B.; Tao, X.-Y.; Zhao, M.; Wang, F.-Q.; Wei, D.-Z. One-pot biosynthesis of 7β-hydroxyandrost-4-ene-3,17-dione from phytosterols by cofactor regeneration system in engineered Mycolicibacterium neoaurum. Microb. Cell Fact. 2022, 21, 59. [Google Scholar] [CrossRef]
- Mohn, W.W.; van der Geize, R.; Stewart, G.R.; Okamoto, S.; Liu, J. The actinobacterial mce4 locus encodes a steroid transporter. J. Biol. Chem. 2008, 283, 35368–35374. [Google Scholar] [CrossRef]
- Klepp, L.I.; Forrellad, M.A.; Osella, A.V.; Blanco, F.C.; Stella, E.J.; Bianco, M.V.; de la Paz Santangelo, M.; Sassetti, C.; Jackson, M.; Cataldi, A.A.; et al. Impact of the deletion of the six mce operons in Mycobacterium smegmatis. Microbes Infect. 2012, 14, 590–599. [Google Scholar] [CrossRef]
- Bansal-Mutalik, R.; Nikaido, H. Quantitative lipid composition of cell envelopes of Corynebacterium glutamicum elucidated through reverse micelle extraction. Proc. Natl. Acad. Sci. USA 2011, 108, 15360–15365. [Google Scholar] [CrossRef]
- Korycka-Machala, M.; Ziolkowski, A.; Rumijowska-Galewicz, A.; Lisowska, K.; Sedlaczek, L. Polycations increase the permeability of Mycobacterium vaccae cell envelopes to hydrophobic compounds. Microbiol. 2001, 147, 2769–2781. [Google Scholar] [CrossRef][Green Version]
- Kim, S.J.; Kweon, O.; Cerniglia, C.E. Degradation of Polycyclic Aromatic Hydrocarbons by Mycobacterium Strain. In Handbook of Hydrocarbon and Lipid Microbiology; Timmis, K.N., Ed.; Springer: Berlin/Heidelberg, Germany, 2010; pp. 1865–1880. [Google Scholar]
- Tekucheva, D.N.; Nikolayeva, V.M.; Karpov, M.V.; Timakova, T.A.; Shutov, A.V.; Donova, M.V. Bioproduction of testosterone from phytosterol by Mycolicibacterium neoaurum strains: “one-pot”, two modes. Bioresour. Bioprocess. 2022, 9, 116. [Google Scholar] [CrossRef]
- Egorova, O.; Nikolayeva, V.; Sukhodolskaya, G.; Donova, M. Transformation of C19-steroids and testosterone production by sterol-transforming strains of Mycobacterium spp. J. Mol. Catal. B Enzym. 2009, 57, 198–203. [Google Scholar] [CrossRef]
- Deo, A.K.; Bandiera, S.M. Biotransformation of lithocholic acid by rat hepatic microsomes: Metabolite analysis by liquid chromatography/mass spectrometry. Drug Metab. Dispos. 2008, 36, 442–451. [Google Scholar] [CrossRef] [PubMed]
- Karpov, M.V.; Nikolaeva, V.M.; Fokina, V.V.; Shutov, A.A.; Kazantsev, A.V.; Strizhov, N.I.; Donova, M.V. Creation and functional analysis of Mycolicibacterium smegmatis recombinant strains carrying the bacillary cytochromes CYP106A1 and CYP106A2 genes. Appl. Biochem. Microbiol. 2022, 58, 947–957. [Google Scholar] [CrossRef]
- Daugelat, S.; Kowall, J.; Mattow, J.; Bumann, D.; Winter, R.; Hurwitz, R.; Kaufmann, S.H.E. The RD1 proteins of Mycobacterium tuberculosis: Expression in Mycobacterium smegmatis and biochemical characterization. Microbes Infect. 2003, 5, 1082–1095. [Google Scholar] [CrossRef]
- Tartoff, K.D.; Hobbs, C.A. Improved media for growing plasmid and cosmid clones. Bethesda Res. Lab. Focus 1987, 9, 12–16. [Google Scholar]
- Lobastova, T.G.; Gulevskaya, S.A.; Sukhodolskaya, G.V.; Turchin, K.V.; Donova, M.V. Screening of mycelial fungi for 7α- and 7β-hydroxylase activity towards dehydroepiandrosterone. Biocatal. Biotransform. 2007, 25, 434–442. [Google Scholar] [CrossRef]

| Steroid Substrate | Transformation Product (TP) | Concentration (% mol.) | TP Ratio Ms/Ec | |
|---|---|---|---|---|
| E. coli (Ec) | M. smegmatis (Ms) | |||
| Androstenedione (AD) | 1β-OH-AD | 0.62 | 6.05 | 9.8 |
| 7β-OH-AD | 11.78 | 26.55 | 2.3 | |
| 1β,7β-diOH-AD | n.d. | 1.15 | – | |
| Androstadienedione (ADD) | 6β-OH-ADD | 1.76 | 11.54 | 6.6 |
| 7β-OH-ADD | 2.39 | 19.79 | 8.3 | |
| 11α-OH-ADD | 1.94 | 8.96 | 4.6 | |
| 6β,11α-diOH-ADD | n.d. | 7.25 | – | |
| 7β-OH-AD | trace | 3.30 | – | |
| Testosterone (TS) | 7β-OH-TS | 0.65 | 2.07 | 3.2 |
| 15β-OH-TS | 1.10 | 1.79 | 1.6 | |
| 1β-OH-AD | n.d. | 3.52 | – | |
| 7β-OH-AD | n.d. | 9.62 | – | |
| Dehydroepiandrosterone (DHEA) | 7α-OH-DHEA | 0.29 | 1.93 | 6.7 |
| 7β-OH-DHEA | 0.49 | 2.33 | 4.8 | |
| 7α,15α-diOH-DHEA | 0.32 | 4.97 | 15.5 | |
| 7β-OH-AD | n.d. | trace | – | |
| Compound | RT (min) | m/z | Structure |
|---|---|---|---|
| Substrates | |||
| Androst-4-ene-3,17-dione (AD) | 26.36 (I) | n.d. | 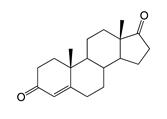 |
| Androsta-1,4-diene-3,17-dione (ADD) | 23.69 (I) | n.d. | 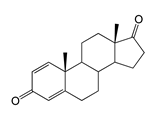 |
| Testosterone (TS) | 26.13 (I) | n.d. | 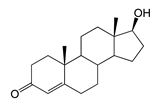 |
| Dehydroepiandrosterone (DHEA) | 8.49 (II) | n.d. | 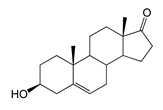 |
| Bioconversion products | |||
| 1β-Hydroxyandrost-4-ene-3,17-dione (1β-OH-AD) | 18.78 (I) | 303 |  |
| 7β-Hydroxyandrost-4-ene-3,17-dione (7β-OH-AD) | 12.34 (I) | 303.1 | 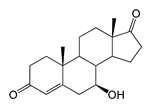 |
| 1β,7β-Dihydroxyandrost-4-ene-3,17-dione (1β,7β-diOH-AD) | 4.96 (I) | 318.9 | 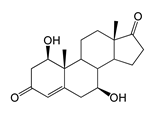 |
| 6β-Hydroxyandrosta-1,4-diene-3,17-dione (6β-OH-ADD) | 12.83 (I) | 300.8 | 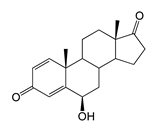 |
| 7β-Hydroxyandrosta-1,4-diene-3,17-dione (7β-OH-ADD) | 10.71 (I) | 300.9 | 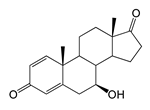 |
| 11α-Hydroxyandrosta-1,4-diene-3,17-dione (11α-OH-ADD) | 9.28 (I) | 300.9 | 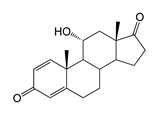 |
| 6β,11α-Dihydroxyandrosta-1,4-diene-3,17-dione (6β,11α-diOH-ADD) | 4.79 (I) | 316.8 |  |
| 7β-Hydroxytestosterone (7β-OH-TS) | 9.49 (I) | 305.1 |  |
| 15β-Hydroxytestosterone (15β-OH-TS) | 10.64 (I) | 305 | 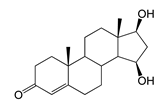 |
| 7α-Hydroxydehydroepiandrosterone (7α-OH-DHEA) | 3.58 (II) | n.d. |  |
| 7β-Hydroxydehydroepiandrosterone (7β-OH-DHEA) | 3.36 (II) | n.d. | 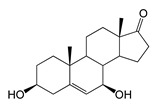 |
| 7α,15α-Dihydroxydehydroepiandrosterone (7α,15α-diOH-DHEA) | 2.99 (II) | n.d. | 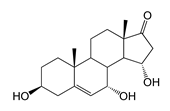 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Poshekhontseva, V.; Nikolaeva, V.; Shutov, A.; Kazantsev, A.; Sazonova, O.; Strizhov, N.; Donova, M. Steroid Hydroxylation by Mutant Cytochrome P450 BM3-LG23 Using Two Expression Chassis. Int. J. Mol. Sci. 2025, 26, 10728. https://doi.org/10.3390/ijms262110728
Poshekhontseva V, Nikolaeva V, Shutov A, Kazantsev A, Sazonova O, Strizhov N, Donova M. Steroid Hydroxylation by Mutant Cytochrome P450 BM3-LG23 Using Two Expression Chassis. International Journal of Molecular Sciences. 2025; 26(21):10728. https://doi.org/10.3390/ijms262110728
Chicago/Turabian StylePoshekhontseva, Veronika, Vera Nikolaeva, Andrey Shutov, Alexey Kazantsev, Olesya Sazonova, Nicolai Strizhov, and Marina Donova. 2025. "Steroid Hydroxylation by Mutant Cytochrome P450 BM3-LG23 Using Two Expression Chassis" International Journal of Molecular Sciences 26, no. 21: 10728. https://doi.org/10.3390/ijms262110728
APA StylePoshekhontseva, V., Nikolaeva, V., Shutov, A., Kazantsev, A., Sazonova, O., Strizhov, N., & Donova, M. (2025). Steroid Hydroxylation by Mutant Cytochrome P450 BM3-LG23 Using Two Expression Chassis. International Journal of Molecular Sciences, 26(21), 10728. https://doi.org/10.3390/ijms262110728







