Application of Antisense Oligonucleotides as an Alternative Approach for Gene Expression Control and Functional Studies
Abstract
1. Introduction
2. Gene Expression Control by ASOs Utilizing Naturally Occurring Mechanisms
2.1. RNA-Level Control—Regulation of Transcription and Translation
2.2. Mechanisms of Action of Synthetic Oligonucleotides
- Steric blocking—ASOs physically obstruct mRNA or hnRNA molecules, preventing further maturation (e.g., capping, poly(A) tail synthesis) or transcript function [28].
- Splicing modulation—ASOs can bind to splice sites within pre-mRNA, influencing exon inclusion and alternative splicing, ultimately altering the repertoire of protein isoforms produced [29]. For instance, ASO binding to splicing enhancers may block the recruitment of splice-promoting factors, resulting in exon skipping, whereas binding to splicing silencers can prevent repressor binding, thereby relieving inhibition and promoting exon inclusion [30].
- Poly(A) tail degradation—ASOs may shorten the poly(A) tail, thereby reducing mRNA stability and accelerating transcript degradation [31].
- Nonsense-mediated mRNA decay (NMD) pathway—ASO binding to pre-mRNA can induce the formation of transcripts containing premature termination codons (PTCs), resulting in aberrant mRNAs that are subsequently degraded via the NMD pathway [32].
- RNA interference (RNAi) pathway—by mimicking siRNAs or miRNAs, ASOs can be incorporated into the RNA-induced silencing complex (RISC), leading to selective mRNA degradation or translational repression [2].
- RNase H pathway—specific to DNA-based oligonucleotides, in which RNase H recognizes DNA-RNA hybrids and degrades the RNA strand. It occurs primarily in the nucleus and constitutes one of the principal applications of antisense oligonucleotides [33].
- RNA activation (RNAa) pathway—some ASOs can activate gene expression through interactions with promoters, enhancers, translation start sites, or transcription factors. RNAa typically involves ~21-nt double-stranded RNAs resembling siRNAs, which, instead of repressing expression, enhance transcriptional activity of the target gene [34].
- Anti-miRNA activity (antagomirs)—ASOs can bind endogenous miRNAs, preventing their interaction with target mRNAs. This results in derepression of genes previously inhibited by the miRNA [35].
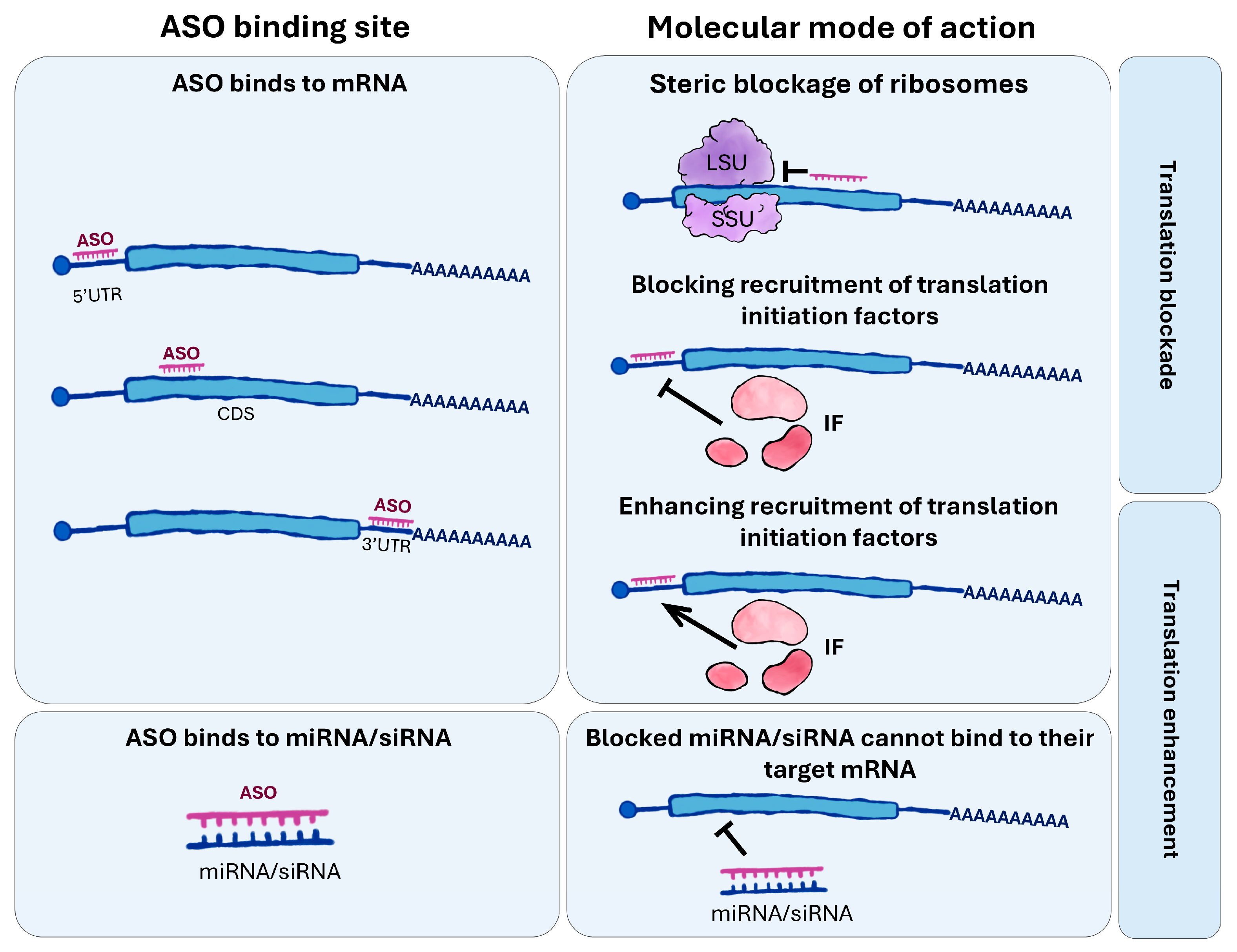
2.3. Regulation of Translation by Antisense Oligonucleotides
2.4. DNA-Level Control—Epigenetic Imprinting
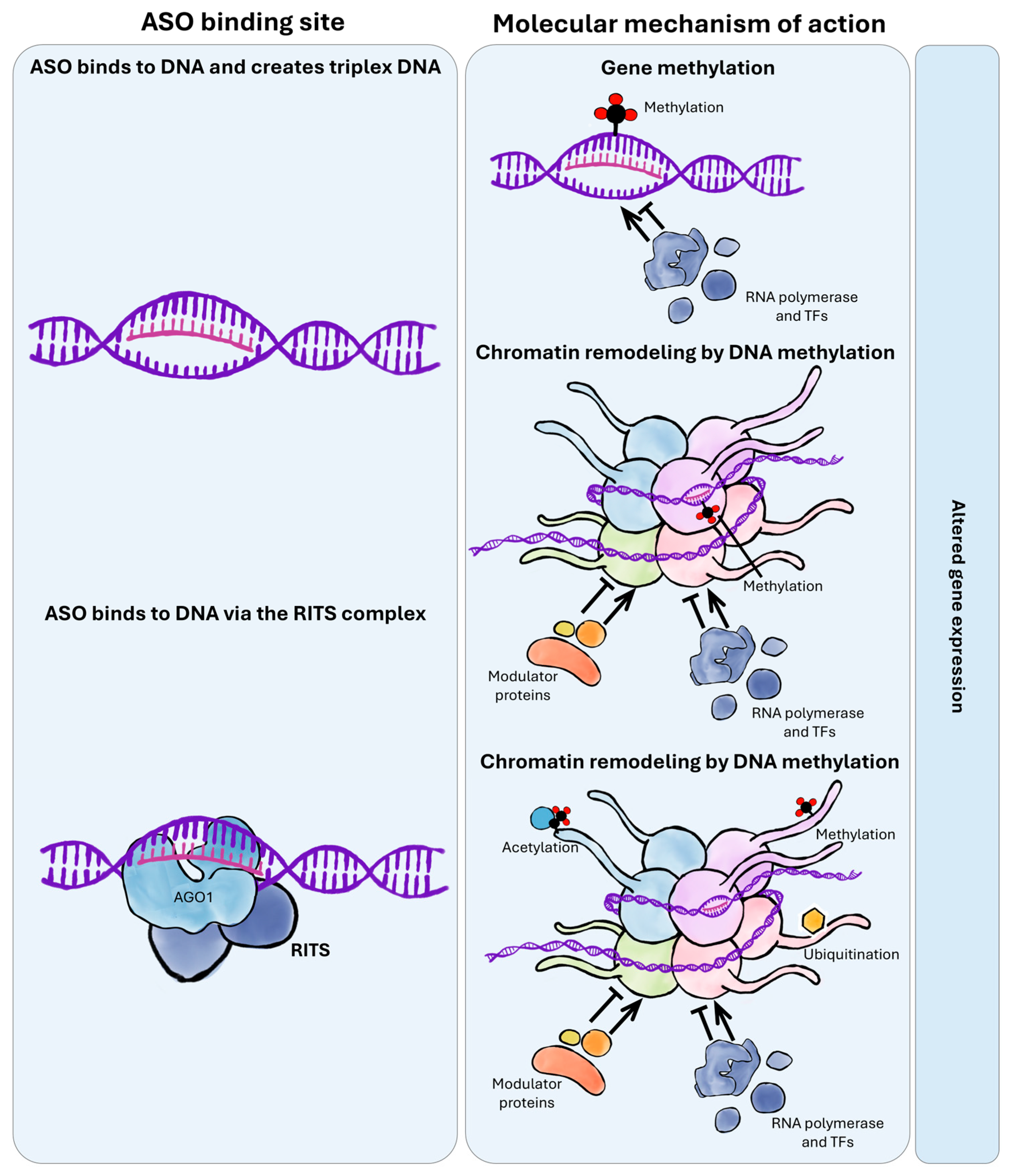
2.5. Epigenetic Modulation by Antisense Oligonucleotides
3. Design and Chemical Modifications of ASOs for Optimized Biological Effect
3.1. ASO Design
3.2. Chemical Modifications
3.2.1. First Generation—Phosphorothioate Backbone Modifications
- Phosphorothioates (PS)—this modification involves replacing an oxygen atom in the phosphate backbone with a sulfur atom [64]. PS modifications are most commonly applied at the ends of ASOs in both in vitro and in vivo studies [57]. They increase stability but reduce the melting temperature of the ASO–mRNA heteroduplex by approximately 0.5 °C per modified nucleotide.
- Methylphosphonates—these modifications replace one of the oxygen atoms in the phosphate group with a methyl group. This results in the ASO molecule losing its negative charge, which enhances stability in biological environments but simultaneously decreases solubility and membrane permeability. Nevertheless, methylphosphonates can be internalized via endocytosis. A significant limitation of their use is the inability to activate RNase H, which precludes their application in strategies requiring target RNA degradation [65].
3.2.2. Second Generation—Sugar (Ribose) Modifications
- 2′-O-methyl (2′-OMe) and 2′-O-methoxyethyl (2′-MOE)—these are the most commonly used sugar modifications, often combined with phosphorothioate (PS) modifications in gapmer designs [56]. They enhance ASO stability; however, due to the absence of the 2′-OH group, they do not activate the RNase H pathway [66].
- Hexitol nucleic acids (HNA)—these are synthetic nucleic acid analogs in which the conventional ribose sugar is replaced by a six-membered hexitol ring. This structural alteration of the sugar backbone increases resistance to enzymatic degradation by nucleases and improves the thermal stability of HNA–DNA and HNA–RNA hybrids. HNAs do not induce RNase H cleavage but can effectively block splicing and translation processes [67,68].
3.2.3. Third Generation—Base Analogs and Modified Furanose Rings
- Peptide Nucleic Acids (PNA)—synthetic DNA analogs in which the sugar-phosphate backbone is replaced by a pseudopeptide chain composed of repeating N-(2-aminoethyl)-glycine units. Nitrogenous bases are attached to this backbone in a manner analogous to natural nucleic acids, enabling specific hybridization with complementary DNA or RNA. PNAs are electrically neutral and exhibit enhanced resistance to nucleases and proteases. Their mode of action involves inhibition of translation or splicing, as well as modulation of transcription [67,69].
- Locked Nucleic Acids (LNA)—synthetic nucleotide analogs in which the ribose ring is chemically “locked” through the introduction of an additional methylene bridge (-CH2-) connecting the 2′-O and 4′-C atoms of the furanose ring. This modification rigidifies the ribose conformation, increasing duplex stability and hybridization affinity. LNAs markedly enhance oligonucleotide resistance to enzymatic degradation both in vitro and in vivo. They are used to suppress mRNA expression by interfering with splicing or obstructing ribosomal translation. When incorporated into gapmer constructs, LNAs can indirectly activate target mRNA degradation through RNase H [67,69].
- Phosphorodiamidate Morpholino Oligomers (PMO)—synthetic oligonucleotide analogs in which the conventional phosphodiester backbone is replaced by an uncharged phosphorodiamidate backbone and the ribose is substituted with a morpholine ring [70]. PMOs exhibit high stability in vitro and in vivo due to strong resistance to nucleases and proteases. They do not activate RNase H but function by blocking translation initiation and disrupting pre-mRNA splicing [67]. Being electrically neutral, their cellular uptake is highly limited; thus, delivery is enhanced by conjugation with cell-penetrating peptides (CPPs), such as arginine-rich peptides (ARPs), which significantly improve cellular internalization and functional efficacy [71].
3.3. Gapmers and Mixmers—Chimeric Oligonucleotides
- Gapmers consist of a central “gap” of approximately 10 phosphorothioate (PS) nucleotides, flanked at the 5′ and 3′ ends by roughly five modified nucleotides (second- or third-generation modifications) [72,73]. This configuration allows RNase H to access the cleavage site, recognize the target and degrade the mRNA, while simultaneously providing protection against exonucleases [56]. Gapmers enable the degradation of target nucleic acids using modifications that would otherwise not activate RNase H due to the absence of the 2′-OH group in the ribose moiety.
- Mixmers contain alternating modified and natural nucleotides (e.g., LNA/DNA). They are designed primarily to block translation, typically targeting the 5′ untranslated region (5′ UTR) of mRNAs [74].
4. Methods of ASO Delivery
4.1. Delivery of ASOs into Animal Cells
4.2. Delivery of ASOs into Plants
5. Applications of ASO Technology
6. Summary and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ASO | Antisense oligonucleotides |
| ODNs | Oligodeoxynucleotides |
| miRNAs | microRNAs |
| siRNAs | small interfering RNAs |
| RITS | RNA-induced transcriptional silencing |
| RISC | RNA-induced silencing complex |
References
- Crooke, S.T. Progress in Antisense Technology. Annu. Rev. Med. 2004, 55, 61–95. [Google Scholar] [CrossRef]
- Bennett, C.F.; Swayze, E.E. RNA Targeting Therapeutics: Molecular Mechanisms of Antisense Oligonucleotides as a Therapeutic Platform. Annu. Rev. Pharmacol. Toxicol. 2010, 50, 259–293. [Google Scholar] [CrossRef]
- Bennett, C.F. Therapeutic Antisense Oligonucleotides Are Coming of Age. Annu. Rev. Med. 2019, 70, 307–321. [Google Scholar] [CrossRef]
- Zhao, M.; Zhang, N.; Gao, T.; Jin, J.; Jing, T.; Wang, J.; Wu, Y.; Wan, X.; Schwab, W.; Song, C. Sesquiterpene Glucosylation Mediated by Glucosyltransferase UGT91Q2 Is Involved in the Modulation of Cold Stress Tolerance in Tea Plants. New Phytol. 2020, 226, 362–372. [Google Scholar] [CrossRef]
- Liang, X.; Shen, W.; Sun, H.; Migawa, M.T.; Vickers, T.A.; Crooke, S.T. Translation Efficiency of mRNAs Is Increased by Antisense Oligonucleotides Targeting Upstream Open Reading Frames. Nat. Biotechnol. 2016, 34, 875–880. [Google Scholar] [CrossRef]
- Stephenson, M.L.; Zamecnik, P.C. Inhibition of Rous Sarcoma Viral RNA Translation by a Specific Oligodeoxyribonucleotide. Proc. Natl. Acad. Sci. USA 1978, 75, 285–288. [Google Scholar] [CrossRef]
- Crooke, S.T.; Witztum, J.L.; Bennett, C.F.; Baker, B.F. RNA-Targeted Therapeutics. Cell Metab. 2018, 27, 714–739. [Google Scholar] [CrossRef] [PubMed]
- Sang, A.; Zhuo, S.; Bochanis, A.; Manautou, J.E.; Bahal, R.; Zhong, X.; Rasmussen, T.P. Mechanisms of Action of the US Food and Drug Administration-Approved Antisense Oligonucleotide Drugs. BioDrugs 2024, 38, 511–526. [Google Scholar] [CrossRef] [PubMed]
- Charleston, J.S.; Schnell, F.J.; Dworzak, J.; Donoghue, C.; Lewis, S.; Chen, L.; Young, G.D.; Milici, A.J.; Voss, J.; DeAlwis, U.; et al. Eteplirsen Treatment for Duchenne Muscular Dystrophy. Neurology 2018, 90, e2146–e2154. [Google Scholar] [CrossRef] [PubMed]
- Finkel, R.S.; Mercuri, E.; Darras, B.T.; Connolly, A.M.; Kuntz, N.L.; Kirschner, J.; Chiriboga, C.A.; Saito, K.; Servais, L.; Tizzano, E.; et al. Nusinersen versus Sham Control in Infantile-Onset Spinal Muscular Atrophy. N. Engl. J. Med. 2017, 377, 1723–1732. [Google Scholar] [CrossRef]
- Tabrizi, S.J.; Leavitt, B.R.; Landwehrmeyer, G.B.; Wild, E.J.; Saft, C.; Barker, R.A.; Blair, N.F.; Craufurd, D.; Priller, J.; Rickards, H.; et al. Targeting Huntingtin Expression in Patients with Huntington’s Disease. N. Engl. J. Med. 2019, 380, 2307–2316. [Google Scholar] [CrossRef]
- Kuijper, E.C.; Bergsma, A.J.; Pijnappel, W.W.M.P.; Aartsma-Rus, A. Opportunities and Challenges for Antisense Oligonucleotide Therapies. J. Inherit. Metab. Dis. 2021, 44, 72–87. [Google Scholar] [CrossRef]
- Buthelezi, L.A.; Pillay, S.; Ntuli, N.N.; Gcanga, L.; Guler, R. Antisense Therapy for Infectious Diseases. Cells 2023, 12, 2119. [Google Scholar] [CrossRef]
- Takakusa, H.; Iwazaki, N.; Nishikawa, M.; Yoshida, T.; Obika, S.; Inoue, T. Drug Metabolism and Pharmacokinetics of Antisense Oligonucleotide Therapeutics: Typical Profiles, Evaluation Approaches, and Points to Consider Compared with Small Molecule Drugs. Nucleic Acid Ther. 2023, 33, 83–94. [Google Scholar] [CrossRef]
- Angrish, N.; Khare, G. Antisense Oligonucleotide Based Therapeutics and Its Applications against Bacterial Infections. Med. Drug Discov. 2023, 20, 100166. [Google Scholar] [CrossRef]
- Dzialo, M.; Szopa, J.; Hnitecka, A.; Zuk, M. Transgenerational Perpetuation of CHS Gene Expression and DNA Methylation Status Induced by Short Oligodeoxynucleotides in Flax (Linum usitatissimum). Int. J. Mol. Sci. 2019, 20, 3983. [Google Scholar] [CrossRef] [PubMed]
- Becker, A.; Lange, M. VIGS—Genomics Goes Functional. Trends Plant Sci. 2010, 15, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Pandey, P.; Senthil-Kumar, M.; Mysore, K.S. Advances in Plant Gene Silencing Methods. In Plant Gene Silencing; Springer: Berlin/Heidelberg, Germany, 2015; pp. 3–23. [Google Scholar]
- Bartel, D.P. MicroRNAs. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef] [PubMed]
- Matzke, M.A.; Mosher, R.A. RNA-Directed DNA Methylation: An Epigenetic Pathway of Increasing Complexity. Nat. Rev. Genet. 2014, 15, 394–408. [Google Scholar] [CrossRef]
- Qian, Y.; Liu, C.; Zeng, X.; Li, L.-C. RNAa: Mechanisms, Therapeutic Potential, and Clinical Progress. Mol. Ther. Nucleic Acids 2025, 36, 102494. [Google Scholar] [CrossRef]
- Paunovska, K.; Loughrey, D.; Dahlman, J.E. Drug Delivery Systems for RNA Therapeutics. Nat. Rev. Genet. 2022, 23, 265–280. [Google Scholar] [CrossRef]
- Papp, I.; Mette, M.F.; Aufsatz, W.; Daxinger, L.; Schauer, S.E.; Ray, A.; van der Winden, J.; Matzke, M.; Matzke, A.J.M. Evidence for Nuclear Processing of Plant Micro RNA and Short Interfering RNA Precursors. Plant Physiol. 2003, 132, 1382–1390. [Google Scholar] [CrossRef]
- Sunkar, R.; Zhu, J.-K. Novel and Stress-Regulated MicroRNAs and Other Small RNAs from Arabidopsis. Plant Cell 2004, 16, 2001–2019. [Google Scholar] [CrossRef] [PubMed]
- Jones-Rhoades, M.W.; Bartel, D.P. Computational Identification of Plant MicroRNAs and Their Targets, Including a Stress-Induced MiRNA. Mol. Cell 2004, 14, 787–799. [Google Scholar] [CrossRef]
- Kolovos, P.; Knoch, T.A.; Grosveld, F.G.; Cook, P.R.; Papantonis, A. Enhancers and Silencers: An Integrated and Simple Model for Their Function. Epigenetics Chromatin 2012, 5, 1. [Google Scholar] [CrossRef] [PubMed]
- Anand, P.; Zhang, Y.; Patil, S.; Kaur, K. Metabolic Stability and Targeted Delivery of Oligonucleotides: Advancing RNA Therapeutics Beyond the Liver. J. Med. Chem. 2025, 68, 6870–6896. [Google Scholar] [CrossRef] [PubMed]
- Desterro, J.; Bak-Gordon, P.; Carmo-Fonseca, M. Targeting MRNA Processing as an Anticancer Strategy. Nat. Rev. Drug Discov. 2020, 19, 112–129. [Google Scholar] [CrossRef]
- Singh, R.K.; Cooper, T.A. Pre-MRNA Splicing in Disease and Therapeutics. Trends Mol. Med. 2012, 18, 472–482. [Google Scholar] [CrossRef]
- Chen, S.; Heendeniya, S.N.; Le, B.T.; Rahimizadeh, K.; Rabiee, N.; Zahra, Q.U.A.; Veedu, R.N. Splice-Modulating Antisense Oligonucleotides as Therapeutics for Inherited Metabolic Diseases. BioDrugs 2024, 38, 177–203. [Google Scholar] [CrossRef]
- Licatalosi, D.D.; Darnell, R.B. RNA Processing and Its Regulation: Global Insights into Biological Networks. Nat. Rev. Genet. 2010, 11, 75–87. [Google Scholar] [CrossRef]
- Ward, A.J.; Norrbom, M.; Chun, S.; Bennett, C.F.; Rigo, F. Nonsense-Mediated Decay as a Terminating Mechanism for Antisense Oligonucleotides. Nucleic Acids Res. 2014, 42, 5871–5879. [Google Scholar] [CrossRef]
- Mata-Ventosa, A.; Vila-Planas, A.; Solsona-Pujol, A.; de la Dueña, J.; Torrents, M.; Izquierdo-García, E.; Pastor-Anglada, M.; Pérez-Torras, S.; Terrazas, M. RNase H-Sensitive Multifunctional ASO-Based Constructs as Promising Tools for the Treatment of Multifactorial Complex Pathologies. Bioorg. Chem. 2024, 150, 107595. [Google Scholar] [CrossRef]
- Tan, C.P.; Sinigaglia, L.; Gomez, V.; Nicholls, J.; Habib, N.A. RNA Activation—A Novel Approach to Therapeutically Upregulate Gene Transcription. Molecules 2021, 26, 6530. [Google Scholar] [CrossRef]
- Krützfeldt, J.; Rajewsky, N.; Braich, R.; Rajeev, K.G.; Tuschl, T.; Manoharan, M.; Stoffel, M. Silencing of MicroRNAs in a with ‘Antagomirs’. Nature 2005, 438, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Falese, J.P.; Donlic, A.; Hargrove, A.E. Targeting RNA with Small Molecules: From Fundamental Principles towards the Clinic. Chem. Soc. Rev. 2021, 50, 2224–2243. [Google Scholar] [CrossRef] [PubMed]
- Liang, X.; Sun, H.; Shen, W.; Wang, S.; Yao, J.; Migawa, M.T.; Bui, H.-H.; Damle, S.S.; Riney, S.; Graham, M.J.; et al. Antisense Oligonucleotides Targeting Translation Inhibitory Elements in 5′ UTRs Can Selectively Increase Protein Levels. Nucleic Acids Res. 2017, 45, 9528–9546. [Google Scholar] [CrossRef]
- Khvorova, A. Modulation of DNA Transcription: The Future of ASO Therapeutics? Cell 2022, 185, 2011–2013. [Google Scholar] [CrossRef] [PubMed]
- Jaenisch, R.; Bird, A. Epigenetic Regulation of Gene Expression: How the Genome Integrates Intrinsic and Environmental Signals. Nat. Genet. 2003, 33, 245–254. [Google Scholar] [CrossRef]
- Liu, X.S.; Wu, H.; Ji, X.; Stelzer, Y.; Wu, X.; Czauderna, S.; Shu, J.; Dadon, D.; Young, R.A.; Jaenisch, R. Editing DNA Methylation in the Mammalian Genome. Cell 2016, 167, 233–247.e17. [Google Scholar] [CrossRef] [PubMed]
- Kouzarides, T. Chromatin Modifications and Their Function. Cell 2007, 128, 693–705. [Google Scholar] [CrossRef]
- Feng, S.; Cokus, S.J.; Zhang, X.; Chen, P.-Y.; Bostick, M.; Goll, M.G.; Hetzel, J.; Jain, J.; Strauss, S.H.; Halpern, M.E.; et al. Conservation and Divergence of Methylation Patterning in Plants and Animals. Proc. Natl. Acad. Sci. USA 2010, 107, 8689–8694. [Google Scholar] [CrossRef]
- Zhang, H.; Lang, Z.; Zhu, J.-K. Dynamics and Function of DNA Methylation in Plants. Nat. Rev. Mol. Cell Biol. 2018, 19, 489–506. [Google Scholar] [CrossRef] [PubMed]
- Verdel, A.; Jia, S.; Gerber, S.; Sugiyama, T.; Gygi, S.; Grewal, S.I.S.; Moazed, D. RNAi-Mediated Targeting of Heterochromatin by the RITS Complex. Science 2004, 303, 672–676. [Google Scholar] [CrossRef] [PubMed]
- Hisey, J.A.; Masnovo, C.; Mirkin, S.M. Triplex H-DNA Structure: The Long and Winding Road from the Discovery to Its Role in Human Disease. NAR Mol. Med. 2024, 1, ugae024. [Google Scholar] [CrossRef] [PubMed]
- Li, H.-L.; Wang, Y.; Guo, D.; Zhu, J.-H.; Wang, Y.; Dai, H.-F.; Peng, S.-Q. Reprogramming of DNA Methylation and Changes of Gene Expression in Grafted Hevea brasiliensis. Front. Plant Sci. 2024, 15, 1407700. [Google Scholar] [CrossRef]
- Bonev, B.; Cavalli, G. Organization and Function of the 3D Genome. Nat. Rev. Genet. 2016, 17, 661–678. [Google Scholar] [CrossRef]
- Banushi, B.; Collova, J.; Milroy, H. Epigenetic Echoes: Bridging Nature, Nurture, and Healing Across Generations. Int. J. Mol. Sci. 2025, 26, 3075. [Google Scholar] [CrossRef]
- Szyf, M.; Pakneshan, P.; Rabbani, S.A. DNA Methylation and Breast Cancer. Biochem. Pharmacol. 2004, 68, 1187–1197. [Google Scholar] [CrossRef]
- Baylin, S.B. DNA Methylation and Gene Silencing in Cancer. Nat. Clin. Pract. Oncol. 2005, 2, S4–S11. [Google Scholar] [CrossRef]
- Litman, A.; Sauerwald, N.; Green Snyder, L.; Foss-Feig, J.; Park, C.Y.; Hao, Y.; Dinstein, I.; Theesfeld, C.L.; Troyanskaya, O.G. Decomposition of Phenotypic Heterogeneity in Autism Reveals Underlying Genetic Programs. Nat. Genet. 2025, 57, 1611–1619. [Google Scholar] [CrossRef]
- Yu, Q.; Liu, S.; Yu, L.; Xiao, Y.; Zhang, S.; Wang, X.; Xu, Y.; Yu, H.; Li, Y.; Yang, J.; et al. RNA Demethylation Increases the Yield and Biomass of Rice and Potato Plants in Field Trials. Nat. Biotechnol. 2021, 39, 1581–1588. [Google Scholar] [CrossRef] [PubMed]
- Goyal, N.; Narayanaswami, P. Making Sense of Antisense Oligonucleotides: A Narrative Review. Muscle Nerve 2018, 57, 356–370. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.; Lee, D.; Hwang, G.; Lee, K.; Kang, M. ASOptimizer: Optimizing Chemical Diversity of Antisense Oligonucleotides through Deep Learning. Nucleic Acids Res. 2025, 53, W39–W44. [Google Scholar] [CrossRef]
- Matveeva, O.V. Identification of Sequence Motifs in Oligonucleotides Whose Presence Is Correlated with Antisense Activity. Nucleic Acids Res. 2000, 28, 2862–2865. [Google Scholar] [CrossRef][Green Version]
- Chan, J.H.; Lim, S.; Wong, W.F. ANTISENSE OLIGONUCLEOTIDES: FROM DESIGN TO THERAPEUTIC APPLICATION. Clin. Exp. Pharmacol. Physiol. 2006, 33, 533–540. [Google Scholar] [CrossRef]
- Wdowikowska, A.; Janicka, M. Antisense Oligonucleotide Technology as a Research Tool in Plant Biology. Funct. Plant Biol. 2021, 49, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Krasnodębski, C.; Sawuła, A.; Kaźmierczak, U.; Żuk, M. Oligo—Not Only for Silencing: Overlooked Potential for Multidirectional Action in Plants. Int. J. Mol. Sci. 2023, 24, 4466. [Google Scholar] [CrossRef]
- Dhuri, K.; Bechtold, C.; Quijano, E.; Pham, H.; Gupta, A.; Vikram, A.; Bahal, R. Antisense Oligonucleotides: An Emerging Area in Drug Discovery and Development. J. Clin. Med. 2020, 9, 2004. [Google Scholar] [CrossRef]
- Hofman, C.R.; Corey, D.R. Targeting RNA with Synthetic Oligonucleotides: Clinical Success Invites New Challenges. Cell Chem. Biol. 2024, 31, 125–138. [Google Scholar] [CrossRef]
- Dzialo, M.; Szopa, J.; Czuj, T.; Zuk, M. Oligodeoxynucleotides Can Transiently Up- and Downregulate CHS Gene Expression in Flax by Changing DNA Methylation in a Sequence-Specific Manner. Front. Plant Sci. 2017, 8, 755. [Google Scholar] [CrossRef]
- Wojtasik, W.; Kulma, A.; Boba, A.; Szopa, J. Oligonucleotide Treatment Causes Flax β-Glucanase up-Regulation via Changes in Gene-Body Methylation. BMC Plant Biol. 2014, 14, 261. [Google Scholar] [CrossRef]
- Crooke, S.T. Molecular Mechanisms of Antisense Oligonucleotides. Nucleic Acid Ther. 2017, 27, 70–77. [Google Scholar] [CrossRef]
- Eckstein, F. Phosphorothioate Oligodeoxynucleotides: What Is Their Origin and What Is Unique About Them? Antisense Nucleic Acid Drug Dev. 2000, 10, 117–121. [Google Scholar] [CrossRef] [PubMed]
- Migawa, M.T.; Shen, W.; Wan, W.B.; Vasquez, G.; Oestergaard, M.E.; Low, A.; De Hoyos, C.L.; Gupta, R.; Murray, S.; Tanowitz, M.; et al. Site-Specific Replacement of Phosphorothioate with Alkyl Phosphonate Linkages Enhances the Therapeutic Profile of Gapmer ASOs by Modulating Interactions with Cellular Proteins. Nucleic Acids Res. 2019, 47, 5465–5479. [Google Scholar] [CrossRef]
- Kurreck, J. Antisense Technologies. Eur. J. Biochem. 2003, 270, 1628–1644. [Google Scholar] [CrossRef]
- Juliano, R.; Alam, M.R.; Dixit, V.; Kang, H. Mechanisms and Strategies for Effective Delivery of Antisense and siRNA Oligonucleotides. Nucleic Acids Res. 2008, 36, 4158–4171. [Google Scholar] [CrossRef]
- De, S.; Jabgunde, A.M.; Patil, R.S.; De Jonghe, S.; Beigelman, L.; Herdewijn, P. Synthesis of Protected Amino Hexitol Nucleosides as Building Blocks for Oligonucleotide Synthesis. J. Org. Chem. 2018, 83, 15155–15169. [Google Scholar] [CrossRef]
- Scoles, D.R.; Minikel, E.V.; Pulst, S.M. Antisense Oligonucleotides. Neurol. Genet. 2019, 5, e323. [Google Scholar] [CrossRef]
- Amantana, A.; Iversen, P. Pharmacokinetics and Biodistribution of Phosphorodiamidate Morpholino Antisense Oligomers. Curr. Opin. Pharmacol. 2005, 5, 550–555. [Google Scholar] [CrossRef] [PubMed]
- Nelson, M.H.; Stein, D.A.; Kroeker, A.D.; Hatlevig, S.A.; Iversen, P.L.; Moulton, H.M. Arginine-Rich Peptide Conjugation to Morpholino Oligomers: Effects on Antisense Activity and Specificity. Bioconjugate Chem. 2005, 16, 959–966. [Google Scholar] [CrossRef] [PubMed]
- McKay, R.A.; Miraglia, L.J.; Cummins, L.L.; Owens, S.R.; Sasmor, H.; Dean, N.M. Characterization of a Potent and Specific Class of Antisense Oligonucleotide Inhibitor of Human Protein Kinase C-α Expression. J. Biol. Chem. 1999, 274, 1715–1722. [Google Scholar] [CrossRef]
- Kurreck, J. Design of Antisense Oligonucleotides Stabilized by Locked Nucleic Acids. Nucleic Acids Res. 2002, 30, 1911–1918. [Google Scholar] [CrossRef]
- Le, B.T.; Raguraman, P.; Kosbar, T.R.; Fletcher, S.; Wilton, S.D.; Veedu, R.N. Antisense Oligonucleotides Targeting Angiogenic Factors as Potential Cancer Therapeutics. Mol. Ther. Nucleic Acids 2019, 14, 142–157. [Google Scholar] [CrossRef]
- Gagliardi, M.; Ashizawa, A.T. The Challenges and Strategies of Antisense Oligonucleotide Drug Delivery. Biomedicines 2021, 9, 433. [Google Scholar] [CrossRef] [PubMed]
- Rustgi, S.; Naveed, S.; Windham, J.; Zhang, H.; Demirer, G.S. Plant Biomacromolecule Delivery Methods in the 21st Century. Front. Genome Ed. 2022, 4, 1011934. [Google Scholar] [CrossRef] [PubMed]
- Höfte, H.; Voxeur, A. Plant Cell Walls. Curr. Biol. 2017, 27, R865–R870. [Google Scholar] [CrossRef]
- Sun, C.; Ridderstråle, K.; Höglund, A.; Larsson, L.; Jansson, C. Sweet Delivery—Sugar Translocators as Ports of Entry for Antisense Oligodeoxynucleotides in Plant Cells. Plant J. 2007, 52, 1192–1198. [Google Scholar] [CrossRef]
- Yagi, Y.; Hosokawa, T.; Yamada, T.; Nii, T.; Mori, T.; Katayama, Y. Intravenous Administration of Antisense Oligonucleotide Incorporated into PLGA Nanoparticles Alters the Pattern of Organ Distribution and Gene Knockdown Effects. Mol. Pharm. 2025, 22, 4794–4803. [Google Scholar] [CrossRef] [PubMed]
- Ämmälä, C.; Drury, W.J.; Knerr, L.; Ahlstedt, I.; Stillemark-Billton, P.; Wennberg-Huldt, C.; Andersson, E.-M.; Valeur, E.; Jansson-Löfmark, R.; Janzén, D.; et al. Targeted Delivery of Antisense Oligonucleotides to Pancreatic β-Cells. Sci. Adv. 2018, 4, eaat3386. [Google Scholar] [CrossRef]
- Cao, Y.; Matsubara, T.; Zhao, C.; Gao, W.; Peng, L.; Shan, J.; Liu, Z.; Yuan, F.; Tang, L.; Li, P.; et al. Antisense Oligonucleotide and Thyroid Hormone Conjugates for Obesity Treatment. Sci. Rep. 2017, 7, 9307. [Google Scholar] [CrossRef]
- Mignani, S.; Shi, X.; Zablocka, M.; Majoral, J.-P. Dendrimer-Enabled Therapeutic Antisense Delivery Systems as Innovation in Medicine. Bioconjugate Chem. 2019, 30, 1938–1950. [Google Scholar] [CrossRef]
- Haque, U.S.; Yokota, T. Enhancing Antisense Oligonucleotide-Based Therapeutic Delivery with DG9, a Versatile Cell-Penetrating Peptide. Cells 2023, 12, 2395. [Google Scholar] [CrossRef]
- Qiao, Y.; Wotring, J.W.; Zhang, C.J.; Jiang, X.; Xiao, L.; Watt, A.; Gattis, D.; Scandalis, E.; Freier, S.; Zheng, Y.; et al. Antisense Oligonucleotides to Therapeutically Target SARS-CoV-2 Infection. PLoS ONE 2023, 18, e0281281. [Google Scholar] [CrossRef]
- Zhu, C.; Lee, J.Y.; Woo, J.Z.; Xu, L.; Wrynla, X.H.; Yamashiro, L.H.; Ji, F.; Biering, S.B.; Van Dis, E.; Gonzalez, F.; et al. An Intranasal ASO Therapeutic Targeting SARS-CoV-2. Nat. Commun. 2022, 13, 4503. [Google Scholar] [CrossRef] [PubMed]
- Bennett, M.; Deikman, J.; Hendrix, B.; Iandolino, A. Barriers to Efficient Foliar Uptake of dsRNA and Molecular Barriers to dsRNA Activity in Plant Cells. Front. Plant Sci. 2020, 11, 816. [Google Scholar] [CrossRef]
- Sun, C.; Ghebramedhin, H.; Höglund, A.-S.; Jansson, C. Antisense Oligodeoxynucleotide Inhibition as a Potent Diagnostic Tool for Gene Function in Plant Biology. Plant Signal. Behav. 2008, 3, 328–330. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Sundström, J.F.; Jin, Y.; Liu, C.; Jansson, C.; Sun, C. A Selection Strategy in Plant Transformation Based on Antisense Oligodeoxynucleotide Inhibition. Plant J. 2014, 77, 954–961. [Google Scholar] [CrossRef] [PubMed]
- Dinç, E.; Tóth, S.Z.; Schansker, G.; Ayaydin, F.; Kovács, L.; Dudits, D.; Garab, G.; Bottka, S. Synthetic Antisense Oligodeoxynucleotides to Transiently Suppress Different Nucleus- and Chloroplast-Encoded Proteins of Higher Plant Chloroplasts. Plant Physiol. 2011, 157, 1628–1641. [Google Scholar] [CrossRef]
- Zhang, H.; Demirer, G.S.; Zhang, H.; Ye, T.; Goh, N.S.; Aditham, A.J.; Cunningham, F.J.; Fan, C.; Landry, M.P. DNA Nanostructures Coordinate Gene Silencing in Mature Plants. Proc. Natl. Acad. Sci. USA 2019, 116, 7543–7548. [Google Scholar] [CrossRef]
- Dias, N.; Stein, C.A. Antisense Oligonucleotides: Basic Concepts and Mechanisms. Mol. Cancer Ther. 2002, 1, 347–355. [Google Scholar]
- Moutinho, A.; Hussey, P.J.; Trewavas, A.J.; Malhó, R. CAMP Acts as a Second Messenger in Pollen Tube Growth and Reorientation. Proc. Natl. Acad. Sci. USA 2001, 98, 10481–10486. [Google Scholar] [CrossRef]
- Zamecnik, P.C.; Stephenson, M.L. Inhibition of Rous Sarcoma Virus Replication and Cell Transformation by a Specific Oligodeoxynucleotide. Proc. Natl. Acad. Sci. USA 1978, 75, 280–284. [Google Scholar] [CrossRef]
- Sun, C.; Höglund, A.; Olsson, H.; Mangelsen, E.; Jansson, C. Antisense Oligodeoxynucleotide Inhibition as a Potent Strategy in Plant Biology: Identification of SUSIBA2 as a Transcriptional Activator in Plant Sugar Signalling. Plant J. 2005, 44, 128–138. [Google Scholar] [CrossRef] [PubMed]
- Roldán, E.L.; Stelinski, L.L.; Pelz-Stelinski, K.S. Evaluation of Tree-injected Oxytetracycline and Antisense Oligonucleotides Targeting Candidatus Liberibacter Asiaticus in Citrus. Pest Manag. Sci. 2025, 81, 1487–1500. [Google Scholar] [CrossRef]
- de Lima, L.F.F.; Carvalho, I.G.B.; de Souza-Neto, R.R.; dos Santos, L.d.S.; Nascimento, C.A.; Takita, M.A.; Távora, F.T.P.K.; Mehta, A.; de Souza, A.A. Antisense Oligonucleotide as a New Technology Application for CsLOB1 Gene Silencing Aiming at Citrus Canker Resistance. Phytopathology 2024, 114, 1802–1809. [Google Scholar] [CrossRef] [PubMed]
- Kochevenko, A.; Willmitzer, L. Chimeric RNA/DNA Oligonucleotide-Based Site-Specific Modification of the Tobacco Acetolactate Syntase Gene. Plant Physiol. 2003, 132, 174–184. [Google Scholar] [CrossRef] [PubMed]
- Liao, F.; Wang, L.; Yang, L.-B.; Zhang, L.; Peng, X.; Sun, M. Antisense Oligodeoxynucleotide Inhibition as an Alternative and Convenient Method for Gene Function Analysis in Pollen Tubes. PLoS ONE 2013, 8, e59112. [Google Scholar] [CrossRef]
- Faltus, T. The Applicability of the European GMO Legislation to Epigenetically Modified Organisms. Front. Bioeng. Biotechnol. 2023, 11, 1124131. [Google Scholar] [CrossRef]
- Crooke, S.T.; Liang, X.-H.; Baker, B.F.; Crooke, R.M. Antisense Technology: A Review. J. Biol. Chem. 2021, 296, 100416. [Google Scholar] [CrossRef]

| Generation | Type of Modification | Chemically Modified Component | Silencing Mechanism | Target Binding | Stability | Adverse Effects |
|---|---|---|---|---|---|---|
| Reference | DNA | No modification | RNase H activity | + | – | Low resistance to nuclease activity: t1/2 ≈ 20 min; possible nonspecific interactions; toxic degradation products (dNMP) |
| First | Phosphorothioate (PS) | Replacement of a non-bridging oxygen in the phosphate backbone with sulfur | RNase H activity | – | ++ Increased resistance to nuclease degradation; t1/2 up to 35 h | Interactions with cell surface and intracellular proteins, potentially affecting cell physiology; possible toxicity |
| Second | 2′-O-Methyl (2′-OMe) | Methyl substitution at the 2′ position of ribose | Steric blockade of translational machinery | ++ | + | Reduced solubility and cellular delivery (restricted to endocytosis) |
| 2′-O-Methoxyethyl (2′-MOE) | Methoxyethyl substitution at the 2′ position of ribose | Steric blockade of translational machinery | ++ | + | Reduced solubility and cellular delivery (restricted to endocytosis) | |
| Hexitol Nucleic Acids (HNA) | Insertion of a methylene group between O4′ and C1′ of the sugar to form a hexose, shifting the nucleobase to the 2′ position | Steric blockade of translational and splicing machinery | ++ | ++ Increased resistance to nuclease and thermal degradation | ||
| Third | Peptide Nucleic Acids (PNA) | Pseudopeptidic backbone (N-(2-aminoethyl)glycine) instead of phosphate; nucleobases attached via methylene-carbonyl linkage | Steric blockade of translational machinery; inhibition of transcriptional elongation, transcription factor binding, and splicing | +++ | +++ Increased resistance to nuclease and peptidase degradation; higher stability due to absence of a negatively charged phosphate backbone | Reduced solubility and cellular delivery |
| Locked Nucleic Acids (LNA) | 2′-O,4′-C-methylene bridge in β-D-furanosyl ribose | Steric blockade of translational and splicing machinery | +++ | +++ Significantly enhanced hybridization affinity and thermodynamic stability of duplexes; resistance to nuclease degradation | Potential toxic effects | |
| Phosphorodiamidate Morpholino Oligomers (PMO) | Six-membered morpholine ring instead of ribose and phosphorodiamidate linkage instead of phosphate backbone | Steric blockade of translational and splicing machinery | +++ | +++ Resistance to nuclease and protease degradation | Possible interactions with nucleic acid–binding proteins due to lack of charge |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Szukowska, A.; Żuk, M.; Sztompke, J.; Bednarz, B.; Kaźmierczak, U. Application of Antisense Oligonucleotides as an Alternative Approach for Gene Expression Control and Functional Studies. Int. J. Mol. Sci. 2025, 26, 10524. https://doi.org/10.3390/ijms262110524
Szukowska A, Żuk M, Sztompke J, Bednarz B, Kaźmierczak U. Application of Antisense Oligonucleotides as an Alternative Approach for Gene Expression Control and Functional Studies. International Journal of Molecular Sciences. 2025; 26(21):10524. https://doi.org/10.3390/ijms262110524
Chicago/Turabian StyleSzukowska, Amelia, Magdalena Żuk, Julia Sztompke, Bartosz Bednarz, and Urszula Kaźmierczak. 2025. "Application of Antisense Oligonucleotides as an Alternative Approach for Gene Expression Control and Functional Studies" International Journal of Molecular Sciences 26, no. 21: 10524. https://doi.org/10.3390/ijms262110524
APA StyleSzukowska, A., Żuk, M., Sztompke, J., Bednarz, B., & Kaźmierczak, U. (2025). Application of Antisense Oligonucleotides as an Alternative Approach for Gene Expression Control and Functional Studies. International Journal of Molecular Sciences, 26(21), 10524. https://doi.org/10.3390/ijms262110524







