Genome-Wide Study of NOT2_3_5 Protein Subfamily in Cotton and Their Necessity in Resistance to Verticillium wilt
Abstract
1. Introduction
2. Results
2.1. Identification of NOT2_3_5 Domain Protein Family in Cotton
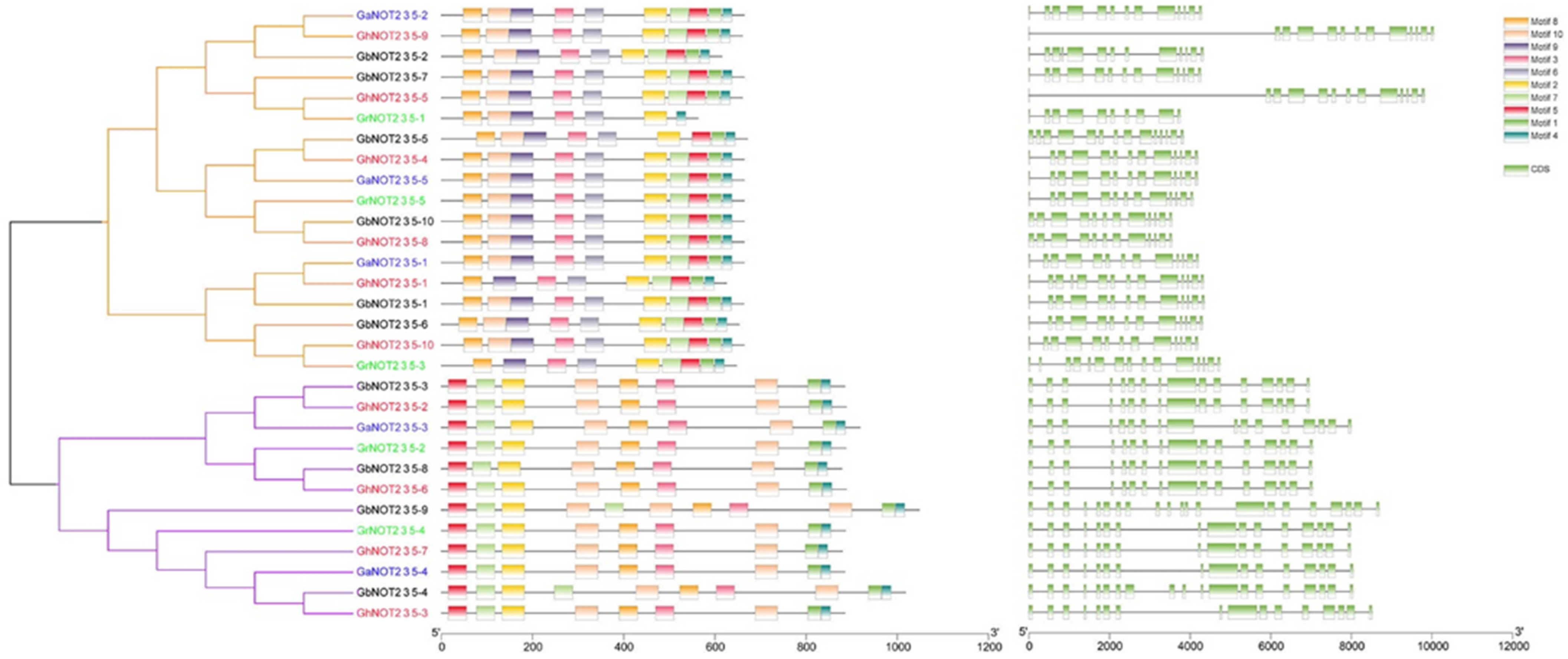
2.2. Phylogenetic and Conserved Amino Residue Analysis of NOT2_3_5 Domain Proteins
2.3. Chromosome Location and Gene Collinearity Analysis
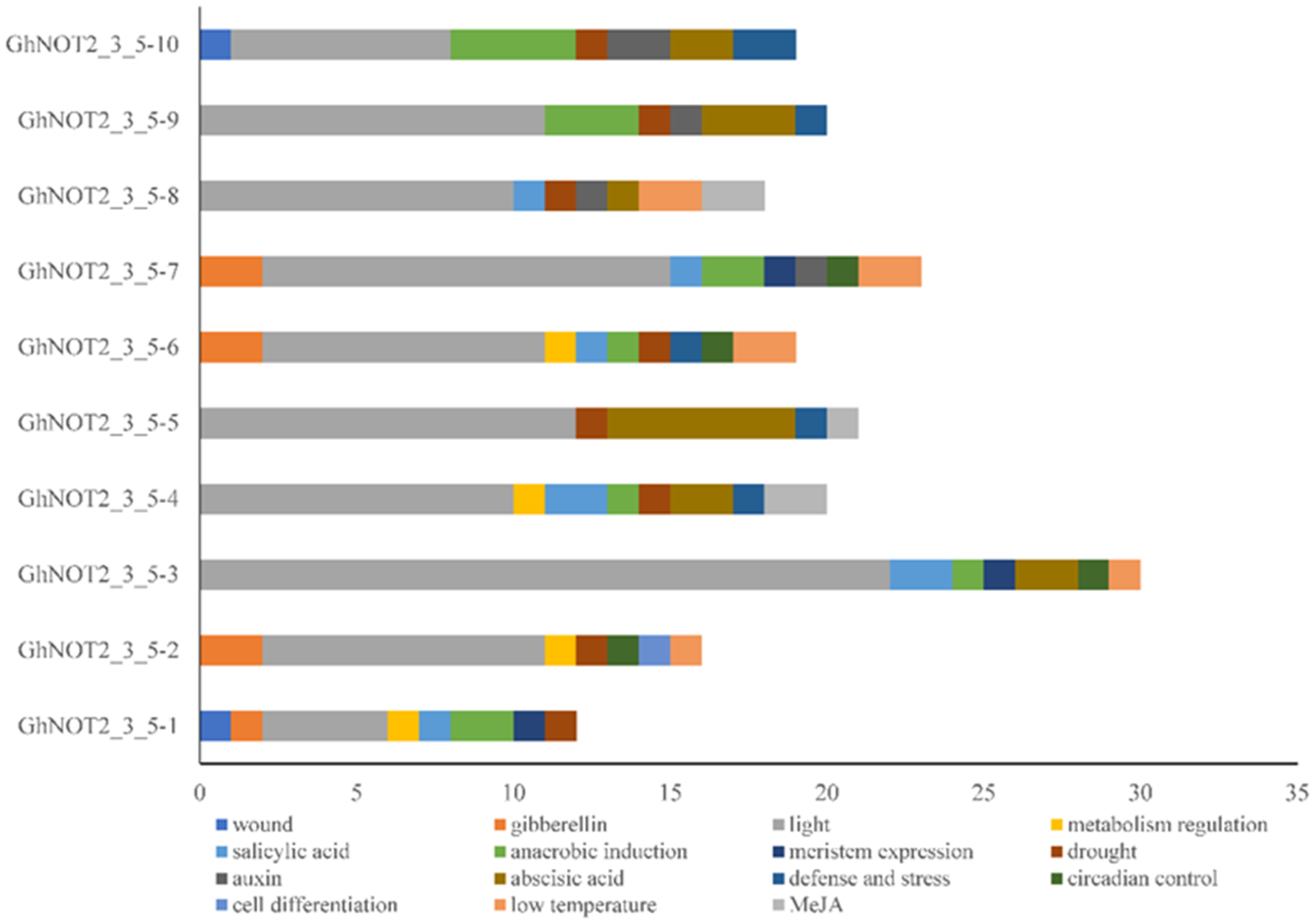
2.4. Cis-Regulatory Elements Analysis of NOT 2_3_5 Promoter Regions
2.5. Prediction of MiRNAs Targeting GhNOT2_3_5 Domain Protein Genes
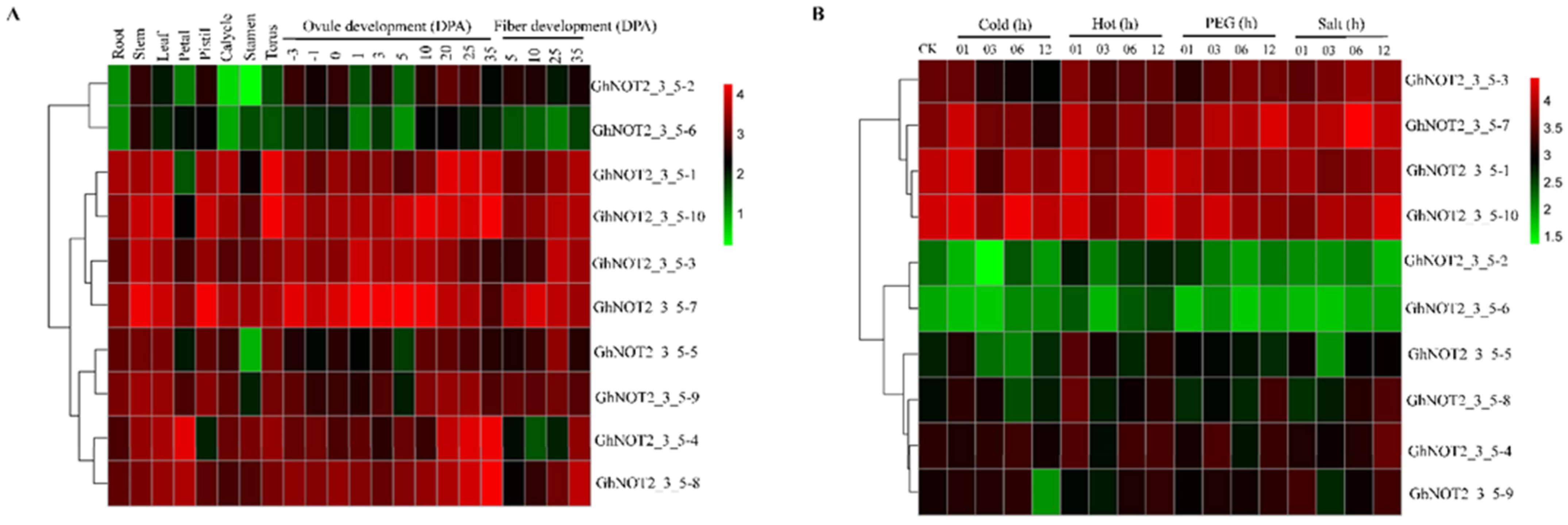
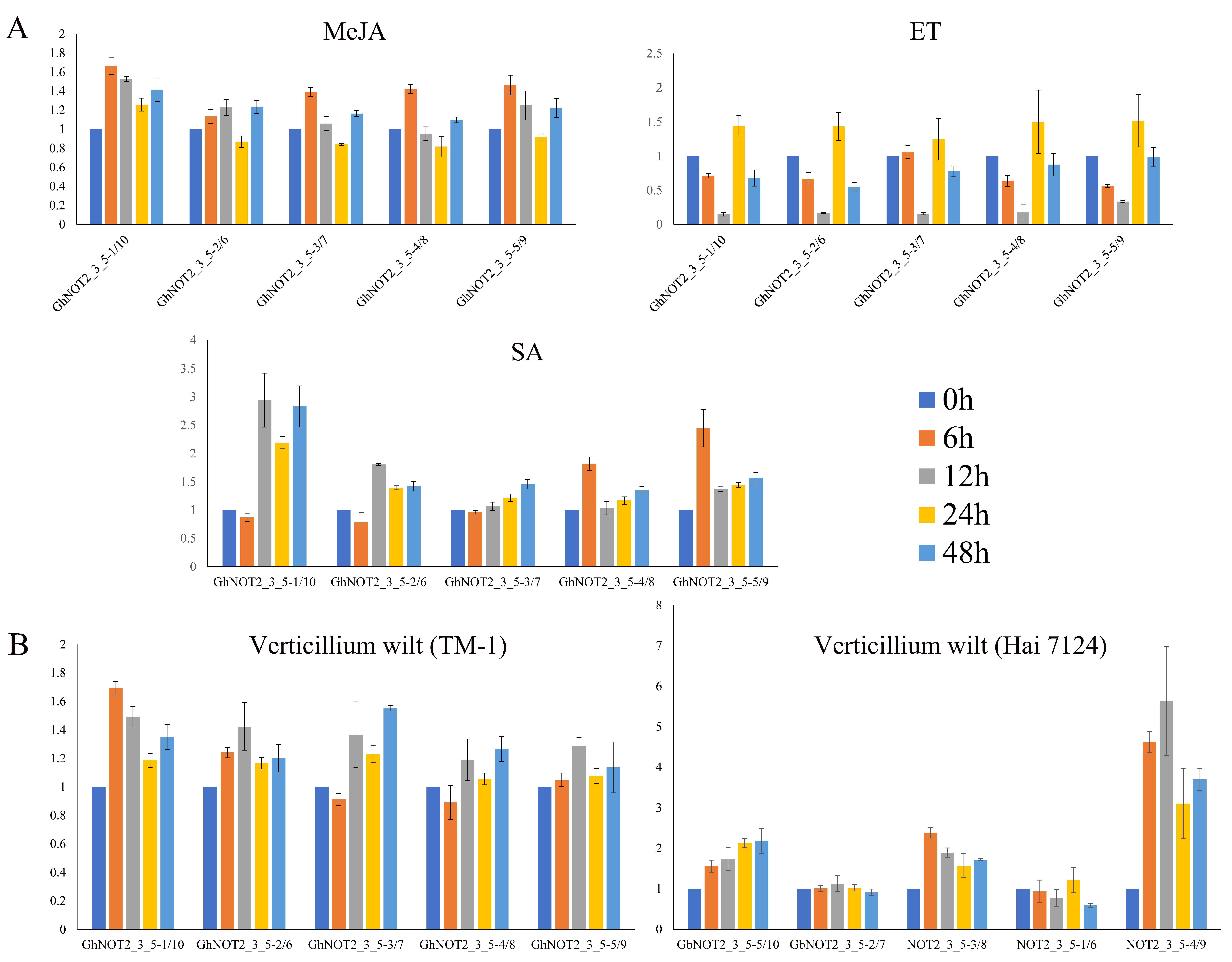
2.6. Expression Patterns of NOT2_3_5 Genes in Different Tissues and Responding to Stresses and Phytohormones
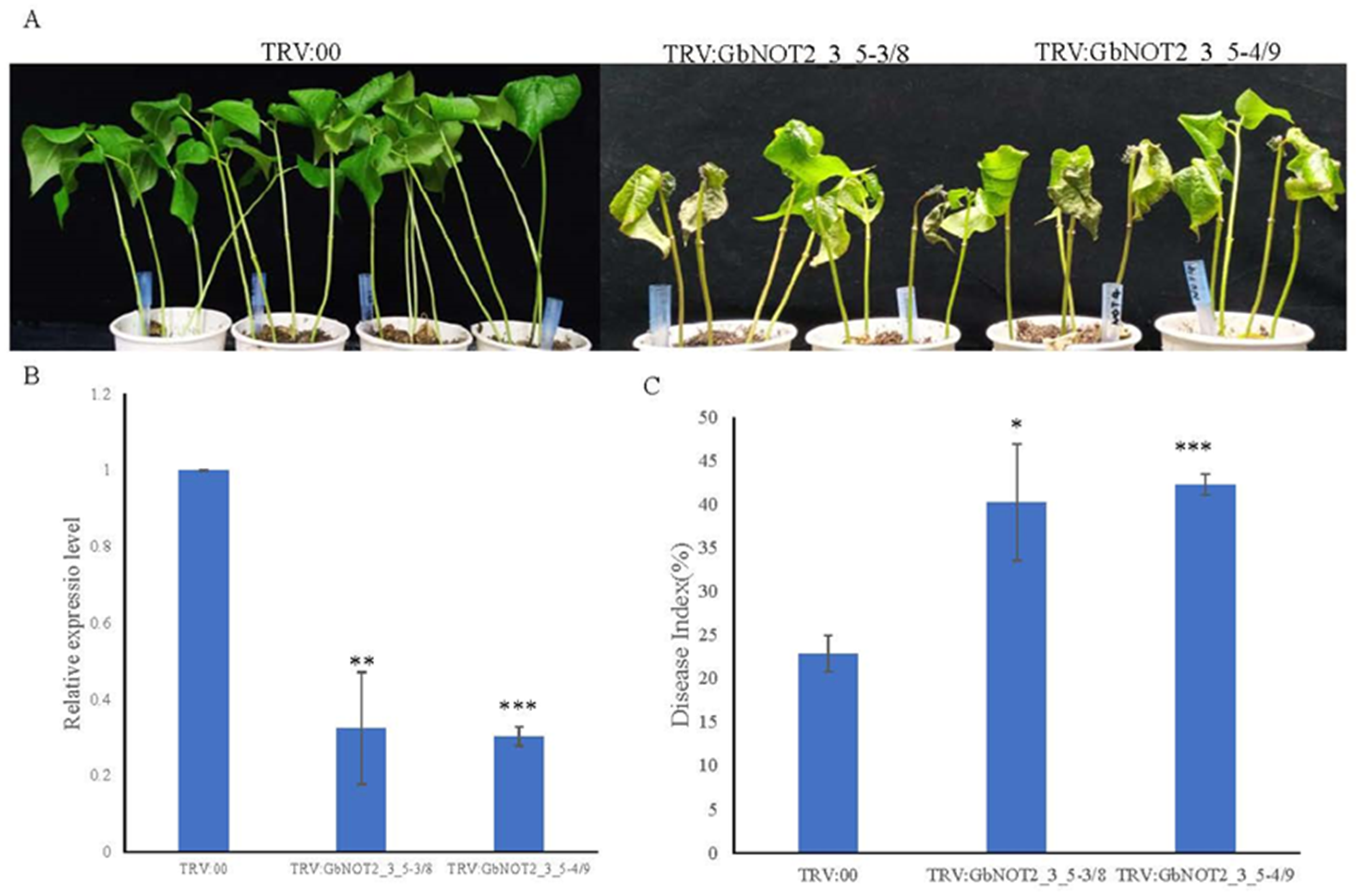
2.7. Silencing of GbNOT2_3_5 Genes Reduced Verticillium Wilt Resistance in Cotton
3. Discussion
3.1. Evolution and Duplication of NOT2_3_5 Genes
3.2. Relationship between MiRNA and NOT2_3_5 Genes
3.3. Differences between NOT2_3_5 Domain Proteins from Two Classes
3.4. Expression Patterns and Putative Disease Resistant Functions of NOT2_3_5 Domain Protein Genes
4. Materials and Methods
4.1. Identification of the NOT2_3_5 Domain Protein Family
4.2. Construction of Phylogenetic Tree and Chromosome Map
4.3. Gene Collinearity and Structural Analysis
4.4. Target Prediction between MiRNAs and GhNOT2_3_5 Members
4.5. Prediction of Upstream cis-Elements
4.6. Expression Pattern Analysis
4.7. Vector Construction and Development of Virus-Induced Gene Silencing (VIGS) in Cotton
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| NOT | Negative on TATA-less |
| CCR4-NOT | Carbon Catabolite Repression 4-NOT |
| SA | Salicylic Acid |
| ET | Ethylene |
| MeJA | Methyl Jasmonate |
| VIGS | Virus-Induced Gene Silencing |
| pI | isoelectric point |
| MW | molecular weight |
| Dpa | Days Post Anthesis |
| TRV | Tobacco Rattle Virus |
References
- Gupta, I.; Villanyi, Z.; Kassem, S.; Hughes, C.; Panasenko, O.O.; Steinmetz, L.M.; Collart, M.A. Translational capacity of a cell is determined during transcription elongation via the ccr4-not complex. Cell Rep. 2016, 15, 1782–1794. [Google Scholar] [CrossRef]
- Collart, M.A.; Struhl, K. Cdc39, an essential nuclear protein that negatively regulates transcription and differentially affects the constitutive and inducible his3 promoters. EMBO J. 1993, 12, 177–186. [Google Scholar] [CrossRef] [PubMed]
- Collart, M.A.; Struhl, K. Not1(cdc39), not2(cdc36), not3, and not4 encode a global-negative regulator of transcription that differentially affects tata-element utilization. Genes Dev. 1994, 8, 525–537. [Google Scholar] [CrossRef]
- Collart, M.A. The not, spt3, and mot1 genes functionally interact to regulate transcription at core promoters. Mol. Cell Biol. 1996, 16, 6668–6676. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Collart, M.A. The ccr4-not complex is a key regulator of eukaryotic gene expression. Wiley Interdiscip. Rev. RNA 2016, 7, 438–454. [Google Scholar] [CrossRef] [PubMed]
- Collart, M.A.; Panasenko, O.O. The ccr4-not complex: Architecture and structural insights. Subcell. Biochem. 2017, 83, 349–379. [Google Scholar] [PubMed]
- Bhaskar, V.; Roudko, V.; Basquin, J.; Sharma, K.; Urlaub, H.; Seraphin, B.; Conti, E. Structure and rna-binding properties of the not1-not2-not5 module of the yeast ccr4-not complex. Nat. Struct. Mol. Biol. 2013, 20, 1281–1288. [Google Scholar] [CrossRef]
- Boland, A.; Chen, Y.; Raisch, T.; Jonas, S.; Kuzuoglu-Ozturk, D.; Wohlbold, L.; Weichenrieder, O.; Izaurralde, E. Structure and assembly of the not module of the human ccr4-not complex. Nat. Struct. Mol. Biol. 2013, 20, 1289–1297. [Google Scholar] [CrossRef]
- Bawankar, P.; Loh, B.; Wohlbold, L.; Schmidt, S.; Izaurralde, E. Not10 and c2orf29/not11 form a conserved module of the ccr4-not complex that docks onto the not1 n-terminal domain. RNA Biol. 2013, 10, 228–244. [Google Scholar] [CrossRef]
- Mauxion, F.; Preve, B.; Seraphin, B. C2orf29/cnot11 and cnot10 form a new module of the ccr4-not complex. RNA Biol. 2013, 10, 267–276. [Google Scholar] [CrossRef]
- Bai, Y.; Salvadore, C.; Chiang, Y.C.; Collart, M.A.; Liu, H.Y.; Denis, C.L. The ccr4 and caf1 proteins of the ccr4-not complex are physically and functionally separated from not2, not4, and not5. Mol. Cell Biol. 1999, 19, 6642–6651. [Google Scholar] [CrossRef] [PubMed]
- Collart, M.A.; Panasenko, O.O.; Nikolaev, S.I. The not3/5 subunit of the ccr4-not complex: A central regulator of gene expression that integrates signals between the cytoplasm and the nucleus in eukaryotic cells. Cell Signal. 2013, 25, 743–751. [Google Scholar] [CrossRef]
- Oberholzer, U.; Collart, M.A. Characterization of not5 that encodes a new component of the not protein complex. Gene 1998, 207, 61–69. [Google Scholar] [CrossRef]
- Russell, P.; Benson, J.D.; Denis, C.L. Characterization of mutations in not2 indicates that it plays an important role in maintaining the integrity of the ccr4-not complex. J. Mol. Biol. 2002, 322, 27–39. [Google Scholar] [CrossRef]
- Kruk, J.A.; Dutta, A.; Fu, J.; Gilmour, D.S.; Reese, J.C. The multifunctional ccr4-not complex directly promotes transcription elongation. Genes Dev. 2011, 25, 581–593. [Google Scholar] [CrossRef]
- Sohn, E.J.; Jung, D.B.; Lee, J.; Yoon, S.W.; Won, G.H.; Ko, H.S.; Kim, S.H. Ccr4-not2 promotes the differentiation and lipogenesis of 3t3-l1 adipocytes via upregulation of pparxγ, cebpalpha and inhibition of p-gsk3alpha/beta and beta-catenin. Cell Physiol. Biochem. 2015, 37, 1881–1889. [Google Scholar] [CrossRef] [PubMed]
- Dai, Y.; Cao, Z.; Huang, L.; Liu, S.; Shen, Z.; Wang, Y.; Wang, H.; Zhang, H.; Li, D.; Song, F. Ccr4-not complex subunit not2 plays critical roles in vegetative growth, conidiation and virulence in watermelon fusarium wilt pathogen fusarium oxysporum f. Sp. Niveum. Front. Microbiol. 2016, 7, 1449. [Google Scholar] [CrossRef] [PubMed]
- Anand, A.; Krichevsky, A.; Schornack, S.; Lahaye, T.; Tzfira, T.; Tang, Y.; Citovsky, V.; Mysore, K.S. Arabidopsis vire2 interacting protein2 is required for agrobacterium t-DNA integration in plants. Plant Cell 2007, 19, 1695–1708. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Song, X.; Gu, L.; Li, X.; Cao, S.; Chu, C.; Cui, X.; Chen, X.; Cao, X. Not2 proteins promote polymerase ii-dependent transcription and interact with multiple microrna biogenesis factors in arabidopsis. Plant Cell 2013, 25, 715–727. [Google Scholar] [CrossRef] [PubMed]
- Zhao, P.; Wang, K.; Lin, Z.; Zhang, W.; Du, L.; Zhang, Y.; Ye, X. Cloning and characterization of tavip2 gene from triticum aestivum and functional analysis in nicotiana tabacum. Sci. Rep. 2016, 6, 37602. [Google Scholar] [CrossRef] [PubMed]
- Paterson, A.H.; Wendel, J.F.; Gundlach, H.; Guo, H.; Jenkins, J.; Jin, D.; Llewellyn, D.; Showmaker, K.C.; Shu, S.; Udall, J.; et al. Repeated polyploidization of gossypium genomes and the evolution of spinnable cotton fibres. Nature 2012, 492, 423–427. [Google Scholar] [CrossRef] [PubMed]
- Lu, P.; Magwanga, R.O.; Kirungu, J.N.; Dong, Q.; Cai, X.; Zhou, Z.; Wang, X.; Xu, Y.; Hou, Y.; Peng, R.; et al. Genome-wide analysis of the cotton g-coupled receptor proteins (gpcr) and functional analysis of gtom1, a novel cotton gpcr gene under drought and cold stress. BMC Genom. 2019, 20, 651. [Google Scholar] [CrossRef]
- Du, X.; Huang, G.; He, S.; Yang, Z.; Sun, G.; Ma, X.; Li, N.; Zhang, X.; Sun, J.; Liu, M.; et al. Resequencing of 243 diploid cotton accessions based on an updated a genome identifies the genetic basis of key agronomic traits. Nat. Genet. 2018, 50, 796–802. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Tu, L.; Yuan, D.; Zhu, D.; Shen, C.; Li, J.; Liu, F.; Pei, L.; Wang, P.; Zhao, G.; et al. Reference genome sequences of two cultivated allotetraploid cottons, gossypium hirsutum and gossypium barbadense. Nat. Genet. 2019, 51, 224–229. [Google Scholar] [CrossRef]
- Li, X.; Zhang, Y.; Ding, C.; Xu, W.; Wang, X. Temporal patterns of cotton fusarium and verticillium wilt in jiangsu coastal areas of china. Sci. Rep. 2017, 7, 12581. [Google Scholar] [CrossRef]
- Zhang, J.; Hu, H.L.; Wang, X.N.; Yang, Y.H.; Zhang, C.J.; Zhu, H.Q.; Shi, L.; Tang, C.M.; Zhao, M.W. Dynamic infection of verticillium dahliae in upland cotton. Plant Biol. 2020, 22, 90–105. [Google Scholar] [CrossRef]
- Zhu, T.; Liang, C.; Meng, Z.; Sun, G.; Meng, Z.; Guo, S.; Zhang, R. Cottonfgd: An integrated functional genomics database for cotton. BMC Plant Biol. 2017, 17, 101. [Google Scholar] [CrossRef] [PubMed]
- Rong, J.; Feltus, F.A.; Liu, L.; Lin, L.; Paterson, A.H. Gene copy number evolution during tetraploid cotton radiation. Heredity 2010, 105, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Adams, K.L.; Wendel, J.F. Polyploidy and genome evolution in plants. Curr. Opin. Plant Biol. 2005, 8, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Wendel, J.E.; Cronn, R.C. Polyploidy and the evolutionary history of cotton. Adv. Agron. 2003, 78, 139–186. [Google Scholar]
- Fang, L.; Guan, X.Y.; Zhang, T.Z. Asymmetric evolution and domestication in allotetraploid cotton (Gossypium hirsutum L.). Crop J. 2017, 5, 159–165. [Google Scholar] [CrossRef]
- Zhang, T.Z.; Hu, Y.; Jiang, W.K.; Fang, L.; Guan, X.Y.; Chen, J.D.; Zhang, J.B.; Saski, C.A.; Scheffler, B.E.; Stelly, D.M.; et al. Sequencing of allotetraploid cotton (gossypium hirsutum l. Acc. Tm-1) provides a resource for fiber improvement. Nat. Biotechnol. 2015, 33, 531–537. [Google Scholar] [CrossRef]
- Xie, Z.; Johansen, L.K.; Gustafson, A.M.; Kasschau, K.D.; Lellis, A.D.; Zilberman, D.; Jacobsen, S.E.; Carrington, J.C. Genetic and functional diversification of small rna pathways in plants. PLoS Biol. 2004, 2, E104. [Google Scholar] [CrossRef] [PubMed]
- Macovei, A.; Tuteja, N. Micrornas targeting dead-box helicases are involved in salinity stress response in rice (Oryza sativa L.). BMC Plant Biol. 2012, 12, 183. [Google Scholar] [CrossRef]
- He, F.; Xu, C.; Fu, X.; Shen, Y.; Guo, L.; Leng, M.; Luo, K. The microrna390/trans-acting short interfering rna3 module mediates lateral root growth under salt stress via the auxin pathway. Plant Physiol. 2018, 177, 775–791. [Google Scholar] [CrossRef]
- Boccara, M.; Sarazin, A.; Thiebeauld, O.; Jay, F.; Voinnet, O.; Navarro, L.; Colot, V. The arabidopsis mir472-rdr6 silencing pathway modulates pamp- and effector-triggered immunity through the post-transcriptional control of disease resistance genes. PLoS Pathog. 2014, 10, e1003883. [Google Scholar] [CrossRef]
- Feyissa, B.A.; Arshad, M.; Gruber, M.Y.; Kohalmi, S.E.; Hannoufa, A. The interplay between mir156/spl13 and dfr/wd40-1 regulate drought tolerance in alfalfa. BMC Plant Biol. 2019, 19, 434. [Google Scholar] [CrossRef]
- Chen, J.; Pan, A.; He, S.; Su, P.; Yuan, X.; Zhu, S.; Liu, Z. Different microrna families involved in regulating high temperature stress response during cotton (Gossypium hirsutum L.) anther development. Int. J. Mol. Sci. 2020, 21, 1280. [Google Scholar] [CrossRef]
- Franco-Zorrilla, J.M.; Solano, R. Identification of plant transcription factor target sequences. Biochim. Biophys. Acta Gene Regul. Mech. 2017, 1860, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.X.; Ahammed, G.J.; Wu, C.; Fan, S.Y.; Zhou, Y.H. Crosstalk among jasmonate, salicylate and ethylene signaling pathways in plant disease and immune responses. Curr. Protein. Pept. Sci. 2015, 16, 450–461. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Bateman, A.; Clements, J.; Coggill, P.; Eberhardt, R.Y.; Eddy, S.R.; Heger, A.; Hetherington, K.; Holm, L.; Mistry, J.; et al. Pfam: The protein families database. Nucleic Acids Res. 2014, 42, D222–D230. [Google Scholar] [CrossRef]
- Letunic, I.; Doerks, T.; Bork, P. Smart 7: Recent updates to the protein domain annotation resource. Nucleic Acids Res. 2012, 40, D302–D305. [Google Scholar] [CrossRef]
- Artimo, P.; Jonnalagedda, M.; Arnold, K.; Baratin, D.; Csardi, G.; de Castro, E.; Duvaud, S.; Flegel, V.; Fortier, A.; Gasteiger, E.; et al. Expasy: Sib bioinformatics resource portal. Nucleic Acids Res. 2012, 40, W597–W603. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.S.; Lin, C.J.; Hwang, J.K. Predicting subcellular localization of proteins for gram-negative bacteria by support vector machines based on n-peptide compositions. Protein. Sci. 2004, 13, 1402–1406. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal w and clustal x version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. Mega7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Voorrips, R.E. Mapchart: Software for the graphical presentation of linkage maps and qtls. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef]
- Krzywinski, M.; Schein, J.; Birol, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef]
- Dai, X.; Zhuang, Z.; Zhao, P.X. Psrnatarget: A plant small rna target analysis server (2017 release). Nucleic Acids Res. 2018, 46, W49–W54. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Lescot, M.; Dehais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouze, P.; Rombauts, S. Plantcare, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Shan, L. Functional genomic analysis of cotton genes with agrobacterium-mediated virus-induced gene silencing. Methods Mol. Biol. 2013, 975, 157–165. [Google Scholar] [PubMed]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, P.; Qin, T.; Chen, W.; Sang, X.; Zhao, Y.; Wang, H. Genome-Wide Study of NOT2_3_5 Protein Subfamily in Cotton and Their Necessity in Resistance to Verticillium wilt. Int. J. Mol. Sci. 2021, 22, 5634. https://doi.org/10.3390/ijms22115634
Zhao P, Qin T, Chen W, Sang X, Zhao Y, Wang H. Genome-Wide Study of NOT2_3_5 Protein Subfamily in Cotton and Their Necessity in Resistance to Verticillium wilt. International Journal of Molecular Sciences. 2021; 22(11):5634. https://doi.org/10.3390/ijms22115634
Chicago/Turabian StyleZhao, Pei, Tengfei Qin, Wei Chen, Xiaohui Sang, Yunlei Zhao, and Hongmei Wang. 2021. "Genome-Wide Study of NOT2_3_5 Protein Subfamily in Cotton and Their Necessity in Resistance to Verticillium wilt" International Journal of Molecular Sciences 22, no. 11: 5634. https://doi.org/10.3390/ijms22115634
APA StyleZhao, P., Qin, T., Chen, W., Sang, X., Zhao, Y., & Wang, H. (2021). Genome-Wide Study of NOT2_3_5 Protein Subfamily in Cotton and Their Necessity in Resistance to Verticillium wilt. International Journal of Molecular Sciences, 22(11), 5634. https://doi.org/10.3390/ijms22115634




