Dissection of Resistance Genes to Pseudomonas syringae pv. phaseolicola in UI3 Common Bean Cultivar
Abstract
1. Introduction
2. Results
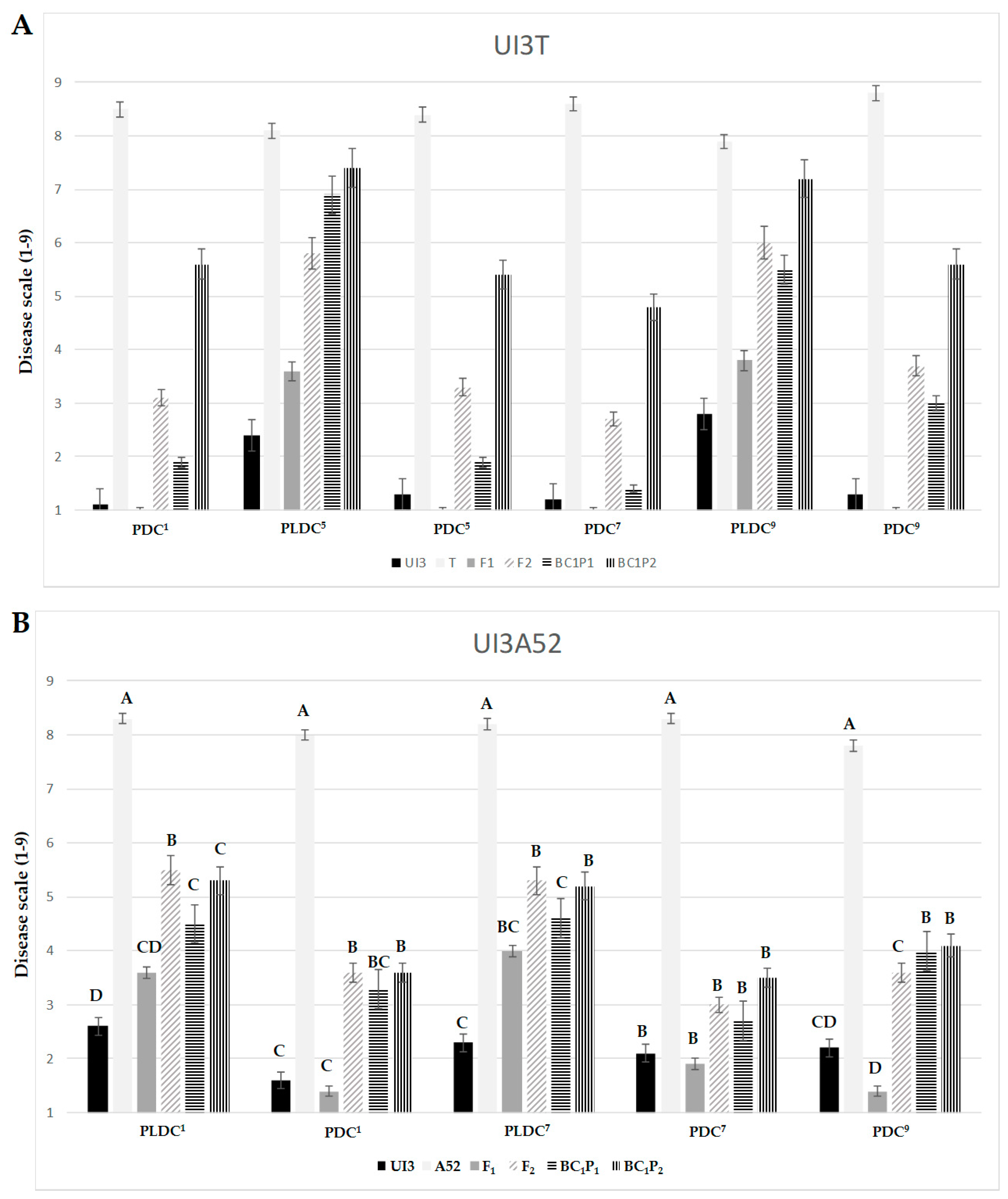
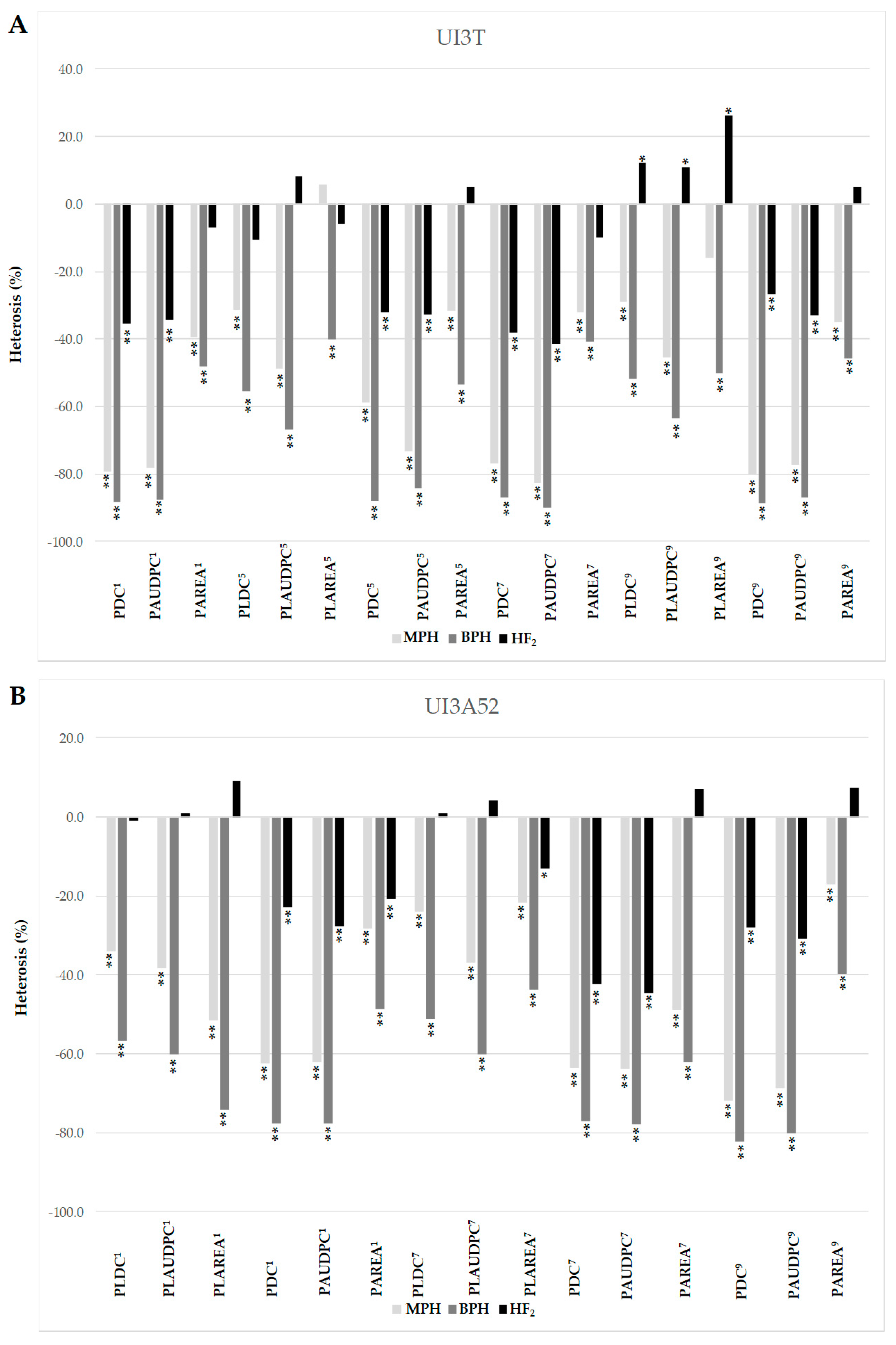
2.1. Potential Genetic Mechanisms of Common Bean Resistance to Races 1, 5, 7 and 9 of P. syringae pv. phaseolicola
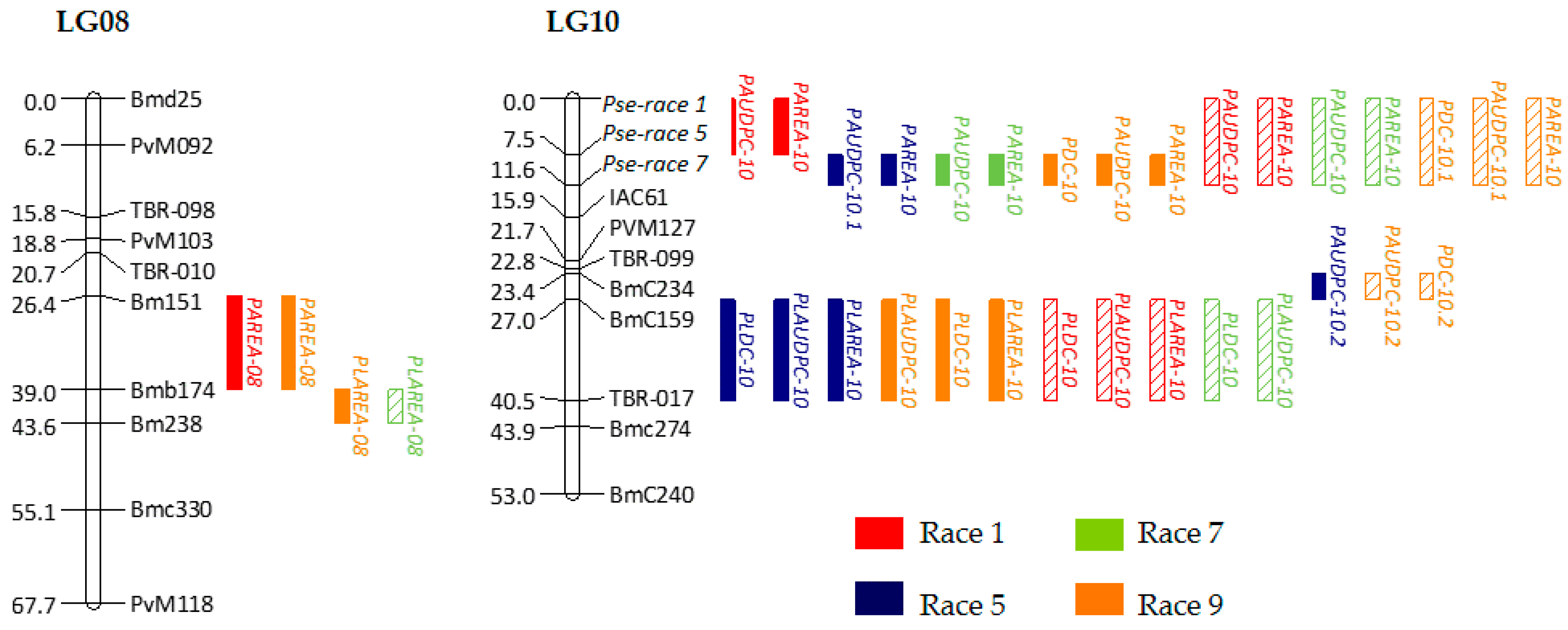
2.2. Genetic Linkage Analysis and Quantitative Trait Loci Mapping
2.3. Genes from the Whole Genome Located within Potential Quantitative Trait Loci Regions
3. Discussion
3.1. Different Modes of Inheritance Reveal Organ and Race Specific Resistance to Psp
3.2. Putative Candidate Genes Underlying Psp Resistance
4. Material and Methods
4.1. Plant Material and Phenotypic Assay
4.2. DNA Isolation and Molecular Marker Analysis
4.3. Quantitative Data Analysis
4.4. Linkage Map Construction and QTL Mapping
4.5. Database Searches of QTLs in Common Bean Genome
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| Psp | Pseudomonas syringae pv. phaseolicola |
| R | Resistance |
| LGs | Linkage groups |
| BCMNV | Bean common mosaic necrotic virus |
| MAMP or PAMP | Microbial- or pathogen-associated molecular patterns |
| PTI | PAMP-triggered immunity |
| ETI | Effector triggered immunity |
| NB-LRRs | Nucleotide-binding site and leucine-rich repeats |
| TIR | Toll-interleukin 1 receptor |
| CC | Coiled-coil |
| RIN4 | RPM1-interacting protein 4 |
| RI | Recombinant Inbred |
| QTL | Quantitative trait loci |
| MPH | Mid-parent heterosis |
| BPH | Better-parent heterosis |
| PFC | Primary flower color |
| SSR | Simple sequence repeat |
| Chr | Chromosome |
| RING finger | C3HC4-type zinc finger |
| TF | Transcription factor |
| NRAMP | Natural resistance associated macrophage protein |
| PL | Primary leaves |
| VC | Vegetative cotyledonary |
| P | Pods |
| DAI | Days after inoculation |
| DC | Disease score |
| AUDPC | Area under disease progress curve |
| AREA | Lesion area |
References
- Prosen, D.; Hatziloukas, E.; Schaad, N.W.; Panopoulos, N.J. Specific detection of Pseudomonas syringae pv. phaseolicola DNA in bean seed by polymerase chain reaction-based amplification of a phaseolotoxin gene region. Phytopathology 1993, 83, 965–970. [Google Scholar] [CrossRef]
- Duncan, R.W.; Gilbertson, R.L.; Lema, M.; Singh, S.P. Inheritance of resistance to the widely distributed race 6 of Pseudomonas syringae pv. phaseolicola in common bean pinto US14HBR6. Can. J. Plant Sci. 2014, 94, 923–928. [Google Scholar] [CrossRef]
- Schwartz, H.F. Halo blight. In Bean Production Problems in the Tropics, 2nd ed.; Schwartz, H.F., Pastor Corrales, M.A., Eds.; CIAT: Cali, Colombia, 1989; pp. 285–301. [Google Scholar]
- Taylor, J.D.; Teverson, D.M.; Allen, D.J.; Pastor-Corrales, M.A. Identification and origin of races of Pseudomonas syringae pv. phaseolicola from Africa and other bean growing areas. Plant Pathol. 1996, 45, 469–478. [Google Scholar] [CrossRef]
- Miklas, P.N.; Fourie, D.; Wagner, J.; Larsen, R.C.; Mienie, C. Tagging and mapping Pse-1 gene for resistance to halo blight in common bean differential cultivar UI-3. Crop Sci. 2009, 49, 41. [Google Scholar] [CrossRef]
- Miklas, P.N.; Fourie, D.; Trapp, J.; Larsen, R.C.; Chavarro, C.; Blair, M.W.; Gepts, P. Genetic characterization and molecular mapping Pse-2 gene for resistance to halo blight in common bean. Crop Sci. 2011, 51, 2439. [Google Scholar] [CrossRef]
- Walker, J.; Patel, P. Inheritance of resistance to halo blight in bean. Phytopathology 1964, 54, 952–954. [Google Scholar]
- Teverson, D.M. Genetics of Pathogenicity and Resistance in Halo-Blight Disease of Beans in Africa. Master’s Thesis, University of Birmingham, Birmingham, UK, 1991. [Google Scholar]
- Taylor, J.D.; Teverson, D.M.; Davis, J.H.C. Sources of resistance to Pseudomonas syringae pv. phaseolicola races in Phaseolus vulgaris. Plant Pathol. 1996, 45, 479–485. [Google Scholar] [CrossRef]
- Miklas, P.N.; Fourie, D.; Trapp, J.; Davis, J.; Myers, J.R. New loci including Pse-6 conferring resistance to halo bacterial blight on chromosome Pv04 in common bean. Crop Sci. 2014, 54, 2099. [Google Scholar] [CrossRef]
- Chisholm, S.T.; Coaker, G.; Day, B.; Staskawicz, B.J. Host-microbe interactions: Shaping the evolution of the plant immune response. Cell 2006, 124, 803–814. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Boller, T.; Felix, G. A renaissance of elicitors: Perception of microbe-associated molecular patterns and danger signals by pattern-recognition receptors. Annu. Rev. Plant Biol. 2009, 60, 379–406. [Google Scholar] [CrossRef] [PubMed]
- Innes, R.W. Guarding the goods. New insights into the central alarm system of plants. Plant Physiol. 2004, 135, 695–701. [Google Scholar] [CrossRef] [PubMed]
- Dangl, J.L.; McDowell, J.M. Two modes of pathogen recognition by plants. Proc. Natl. Acad. Sci. USA 2006, 103, 8575–8576. [Google Scholar] [CrossRef] [PubMed]
- Meyers, B.C.; Kaushik, S.; Nandety, R.S. Evolving disease resistance genes. Curr. Opin. Plant Biol. 2005, 8, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Collier, S.M.; Moffett, P. NB-LRRs work a “bait and switch” on pathogens. Trends Plant Sci. 2009, 14, 521–529. [Google Scholar] [CrossRef] [PubMed]
- Rairdan, G.J.; Collier, S.M.; Sacco, M.A.; Baldwin, T.T.; Boettrich, T.; Moffett, P. The coiled-coil and nucleotide binding domains of the potato Rx disease resistance protein function in pathogen recognition and signaling. Plant Cell 2008, 20, 739–751. [Google Scholar] [CrossRef] [PubMed]
- Meyers, B.C.; Kozik, A.; Griego, A.; Kuang, H.; Michelmore, R.W. Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 2003, 15, 809–834. [Google Scholar] [CrossRef] [PubMed]
- Ameline-Torregrosa, C.; Wang, B.B.; O’Bleness, M.S. Identification and characterization of nucleotide-binding site-leucine-rich repeat genes in the model plant Medicago truncatula. Plant Physiol. 2008, 146, 5–21. [Google Scholar] [CrossRef] [PubMed]
- Kohler, A.; Rinaldi, C.; Duplessis, S.; Baucher, M.; Geelen, D.; Duchaussoy, F.; Meyers, B.C.; Boerjan, W.; Martin, F. Genome-wide identification of NBS resistance genes in Populus trichocarpa. Plant Mol. Biol. 2008, 66, 619–636. [Google Scholar] [CrossRef] [PubMed]
- Grant, M.R.; Godiard, L.; Straube, E.; Ashfield, T.; Lewald, J.; Sattler, A.; Innes, R.W.; Dangl, J.L. Structure of the Arabidopsis RPM1 gene enabling dual specificity disease resistance. Science 1995, 269, 843–846. [Google Scholar] [CrossRef] [PubMed]
- Leister, D. Tandem and segmental gene duplication and recombination in the evolution of plant disease resistance gene. Trends Genet. 2004, 20, 116–122. [Google Scholar] [CrossRef] [PubMed]
- McDowell, J.M.; Simon, S.A. Recent insights into R gene evolution. Mol. Plant Pathol. 2006, 7, 437–448. [Google Scholar] [CrossRef] [PubMed]
- McDowell, J.M.; Simon, S.A. Molecular diversity at the plant-pathogen interface. Dev. Comp. Immunol. 2008, 32, 736–744. [Google Scholar] [CrossRef] [PubMed]
- Schmutz, J.; McClean, P.E.; Mamidi, S.; Wu, G.A.; Cannon, S.B.; Grimwood, J.; Jenkins, J.; Shu, S.; Song, Q.; Chavarro, C.; et al. A reference genome for common bean and genome-wide analysis of dual domestications. Nat. Genet. 2014, 46, 7–13. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.R.; Shepherd, K.W.; Mayo, G.M. Recombination among genes at the L group in flax conferring resistance to rust. Theor. Appl. Genet. 1989, 77, 540–546. [Google Scholar] [CrossRef] [PubMed]
- Hulbert, S.H.; Bennetzen, J.L. Recombination at the Rp1 locus of maize. Mol. Gen. Genet. 1991, 226, 377–382. [Google Scholar] [CrossRef] [PubMed]
- Witsenboer, H.; Kesseli, R.V.; Fortin, M.G.; Stanghellini, M.; Michelmore, R.W. Sources and genetic structure of a cluster of genes for resistance to three pathogens in lettuce. Theor. Appl. Genet. 1995, 91, 178–188. [Google Scholar] [CrossRef] [PubMed]
- Belkhadir, Y.; Nimchuk, Z.; Hubert, D.A.; Mackey, D.; Dangl, J.L. Arabidopsis RIN4 negatively regulates disease resistance mediated by RPS2 and RPM1 downstream or independent of the NDR1 signal modulator and is not required for the virulence functions of bacterial type III effectors AvrRpt2 or AvrRpm1. Plant Cell 2004, 16, 2822–2835. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.J.; Lin, N.C.; Martin, G.B. Two distinct Pseudomonas effector proteins interact with the Pto kinase and activate plant immunity. Cell 2002, 109, 589–598. [Google Scholar] [CrossRef]
- Mackey, D.; Belkhadir, Y.; Alonso, J.M.; Ecker, J.R.; Dangl, J.L. Arabidopsis RIN4 is a target of the Type III virulence effector AvrRpt2 and modulates RPS2-mediated resistance. Cell 2003, 112, 379–389. [Google Scholar] [CrossRef]
- Ashfield, T.; Bocian, A.; Held, D.; Henk, A.D.; Marek, L.F.; Danesh, D.; Peñuela, S.; Meksem, K.; Lightfoot, D.A.; Young, N.D.; et al. Genetic and physical localization of the soybean Rpg1-b disease resistance gene reveals a complex locus containing several tightly linked families of NBS-LRR genes. Mol. Plant Microbe Interact. 2003, 16, 817–826. [Google Scholar] [CrossRef] [PubMed]
- Ashfield, T.; Ong, L.E.; Nobuta, K.; Schneider, C.M.; Innes, R.W. Convergent evolution of disease resistance gene specificity in two flowering plant families. Plant Cell 2004, 16, 309–318. [Google Scholar] [CrossRef] [PubMed]
- McDowell, J.M. Convergent evolution of disease resistance genes. Trends Plant Sci. 2004, 9, 315–317. [Google Scholar] [CrossRef] [PubMed]
- Ashfield, T.; Keen, N.T.; Buzzell, R.I.; Innes, R.W. Soybean resistance genes specific for different Pseudomonas syringae avirulence genes are allelic, or closely linked, at the RPG1 locus. Genetics 1995, 141, 1597–1604. [Google Scholar] [PubMed]
- Chen, N.W.; Sevignac, M.; Thareau, V.; Magdelenat, G.; David, P.; Ashfield, T.; Innes, R.W.; Geffroy, V. Specific resistances against Pseudomonas syringae effectors AvrB and AvrRpm1 have evolved differently in common bean (Phaseolus vulgaris), soybean (Glycine max), and Arabidopsis thaliana. New Phytol. 2010, 187, 941–956. [Google Scholar] [CrossRef] [PubMed]
- Tao, Y.; Xie, Z.; Chen, W.; Glazebrook, J.; Chang, H.S.; Han, B.; Zhu, T.; Zou, G.; Katagiri, F. Quantitative nature of Arabidopsis responses during compatible and incompatible interactions with the bacterial pathogen Pseudomonas syringae. Plant Cell 2003, 15, 317–330. [Google Scholar] [CrossRef] [PubMed]
- Rant, J.C.; Arraiano, L.S.; Chabannes, M.; Brown, J.K.M. Quantitative trait loci for partial resistance to Pseudomonas syringae pv. maculicola in Arabidopsis thaliana. Mol. Plant Pathol. 2013, 14, 828–837. [Google Scholar] [CrossRef] [PubMed]
- Stuthman, D.D.; Leonard, K.J.; Miller-Garvin, J. Breeding crops for durable resistance to disease. Adv. Agron. 2007, 95, 319–367. [Google Scholar] [CrossRef]
- St. Clair, D.A. Quantitative disease resistance and quantitative resistance loci in breeding. Annu. Rev. Phytopathol. 2010, 48, 247–268. [Google Scholar] [CrossRef] [PubMed]
- Poland, J.A.; Balint-Kurti, P.J.; Wisser, R.J.; Pratt, R.C.; Nelson, R.J. Shades of gray: The world of quantitative disease resistance. Trends Plant Sci. 2009, 14, 21–29. [Google Scholar] [CrossRef] [PubMed]
- Zaiter, Z.H.; Coyne, D.P. Testing inoculation methods and sources of resistance to the halo blight bacterium (Pseudomonas syringae pv. phaseolicola) in Phaseolus vulgaris. Euphytica 1984, 33, 133–141. [Google Scholar] [CrossRef]
- Ariyaratne, H.M.; Coyne, D.P.; Jung, G.; Skroch, P.V.; Vidaver, A.K.; Steadman, J.R.; Miklas, P.N.; Bassett, M.J. Molecular mapping of disease resistance genes for halo blight, common bacterial blight and bean common mosaic virus in a segregating population of common bean. J. Am. Soc. Hortic. Sci. 1999, 124, 654–662. [Google Scholar]
- Trabanco, N.; Asensio-Manzanera, M.C.; Pérez-Vega, E.; Ibeas, A.; Campa, A.; Ferreira, J.J. Identification of quantitative trait loci involved in the response of common bean to Pseudomonas syringae pv. phaseolicola. Mol. Breed. 2014, 33, 577–588. [Google Scholar] [CrossRef]
- González, A.M.; Yuste-Lisbona, F.J.; Godoy, L.; Fernández-Lozano, A.; Rodiño, A.P.; De Ron, A.M.; Lozano, R.; Santalla, M. Exploring the quantitative resistance to Pseudomonas syringae pv. phaseolicola in common bean (Phaseolus vulgaris L.). Mol. Breed. 2016, 36, 166. [Google Scholar] [CrossRef]
- Tock, A.J.; Fourie, D.; Walley, P.G.; Holub, E.B.; Soler, A.; Cichy, K.A.; Pastor-Corrales, M.A.; Song, Q.; Porch, T.G.; Hart, J.P.; et al. Genome-Wide Linkage and Association Mapping of Halo Blight Resistance in Common Bean to Race 6 of the Globally Important Bacterial Pathogen. Front. Plant Sci. 2017, 8, 1170. [Google Scholar] [CrossRef] [PubMed]
- Taylor, J.D.; Innes, N.L.; Dudley, C.L.; Griffiths, W.A. Sources and inheritance of resistance to halo-blight of Phaseolus beans. Annu. Appl. Biol. 1978, 90, 101–110. [Google Scholar] [CrossRef]
- Fourie, D.; Miklas, P.; Ariyaranthe, H. Genes conditioning halo blight resistance to races 1, 7 and 9 occur in a tight cluster. Annu. Rep. Bean Improv. Coop. 2004, 9, 103–104. [Google Scholar]
- David, P.; Chen, N.W.G.; Pedrosa-Harand, A.; Thareau, V.; Sévignac, M.; Cannon, S.B.; Debouck, D.; Langin, T.; Geffroy, V. A nomadic subtelomeric disease resistance gene cluster in common bean. Plant Physiol. 2009, 151, 1048–1065. [Google Scholar] [CrossRef] [PubMed]
- Campa, A.; Giraldez, R.; Ferreira, J.J. Genetic dissection of the resistance to nine anthracnose races in the common bean differential cultivars MDRK and TU. Theor. Appl. Genet. 2009, 119, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Kelly, J.D.; Vallejo, V.A. A comprehensive review of the major genes conditioning resistance to anthracnose in common bean. HortScience 2004, 39, 1196–1207. [Google Scholar]
- Vallejo, V.A.; Kelly, J.D. Molecular tagging and characterization of alleles at the Co-1 anthracnose resistance locus in common. ICFAI Univ. J. Genet. Evol. 2008, 1, 7–20. [Google Scholar]
- Campa, A.; Giraldez, R.; Ferreira, J.J. Genetic analysis of the resistance to eight anthracnose races in the common bean differential cultivar Kaboon. Phytopathology 2011, 101, 757–764. [Google Scholar] [CrossRef] [PubMed]
- Campa, A.; Rodríguez-Suárez, C.; Giraldez, R.; Ferreira, J.J. Genetic analysis of the response to eleven Colletotrichum lindemuthianum races in a RIL population of common bean (Phaseolus vulgaris L.). BMC Plant Biol. 2014, 14, 115. [Google Scholar] [CrossRef] [PubMed]
- Mather, K.; Jinks, J.L. Components of means: Additive and dominance effects. In Biometrical Genetics: The Study of Continuous Variation, 3rd ed.; Chapman & Hall: London, UK, 1982; pp. 65–81. ISBN 978-0-412-22890-2. [Google Scholar]
- Lynch, M.; Walsh, B. Genetics and Analysis of Quantitative Traits; Sinauer Associates: Sunderland, MA, USA, 1998. [Google Scholar]
- Arnaud-Santana, E.; Coyne, D.P.; Steadman, J.R.; Eskridge, K.M.; Beaver, J.S. Heritabilities of seed transmission, leaf and pod reactions to common blight, leaf reaction to web blight and plant architecture and their associations in dry beans. Annu. Rep. Bean Improv. Coop. 1994, 37, 46–47. [Google Scholar]
- Santos, A.; Bressan, R.; Pereira, M.; Rodrigues, R.; Ferreira, C. Genetic linkage map of Phaseolus vulgaris and identification of QTLs responsible for resistance to Xanthomonas axonopodis pv. phaseoli. Fitopatología Brasileira 2003, 28, 5–10. [Google Scholar] [CrossRef]
- Borel, J.C.; Ramalho, M.A.P.; Abreu, A.F.B.; Maia, L.G.S. Genetic control of angular leaf spot reaction in common bean leaves and pods. Sci. Agric. 2011, 68, 661–664. [Google Scholar] [CrossRef]
- Genchev, D.; Christova, P.; Kiryakov, I.; Beleva, M.; Batchvarova, R. Breeding of common bean for resistance to the physiological races of anthracnose identified in Bulgaria. Biotechnol. Biotechnol. Equip. 2010, 24, 1814–1823. [Google Scholar] [CrossRef]
- González, A.M.; Yuste-Lisbona, F.J.; Rodiño, A.P.; De Ron, A.M.; Capel, C.; García-Alcázar, M.; Lozano, R.; Santalla, M. Uncovering the genetic architecture of Colletotrichum lindemuthianum resistance through QTL mapping and epistatic interaction analysis in common bean. Front. Plant Sci. 2015, 6, 141. [Google Scholar] [CrossRef] [PubMed]
- Shaner, G.; Finney, R.E. The effect of nitrogen fertilization on the expression of slow-mildew in resistance in Knox wheat. Phytopathology 1977, 67, 1051–1056. [Google Scholar] [CrossRef]
- Pedrosa-Harand, A.; Porch, T.; Gepts, P. Standard nomenclature for common bean chromosomes and linkage groups. Annu. Rep. Bean Improv. Coop. 2008, 51, 106–107. [Google Scholar]
- Koinange, E.; Singh, S.; Gepts, P. Genetic Control of the domestication syndrome in common bean. Crop Sci. 1996, 36, 1037–1045. [Google Scholar] [CrossRef]
- McClean, P.E.; Lee, R.K.; Otto, C.; Gepts, P.; Bassett, M.J. Molecular and phenotypic mapping of genes controlling seed coat pattern and color in common bean (Phaseolus vulgaris L.). J. Hered. 2002, 93, 148–152. [Google Scholar] [CrossRef] [PubMed]
- Stuber, C.W.; Edwards, M.D.; Wendel, J.F. Molecular-marker facilitated investigations of quantitative trait loci in maize. II. Factors influencing yield and its component traits. Crop Sci. 1987, 27, 639–648. [Google Scholar] [CrossRef]
- Garcia, A.A.F.; Wang, S.; Melchinger, A.E.; Zeng, Z.B. Quantitative trait loci mapping and the genetic basis of heterosis in maize and rice. Genetics 2008, 180, 1707–1724. [Google Scholar] [CrossRef] [PubMed]
- Shah, J.; Chaturvedi, R. Lipid signals in plant–pathogen interactions. Annu. Plant Rev. 2009, 34, 292–333. [Google Scholar] [CrossRef]
- Day, B.; Dahlbeck, D.; Staskawicz, B.J. NDR1 interaction with RIN4 mediates the differential activation of multiple disease resistance pathways in Arabidopsis. Plant Cell 2006, 18, 2782–2791. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Elmore, J.M.; Lin, Z.J.; Coaker, G. A receptor-like cytoplasmic kinase phosphorylates the host target RIN4, leading to the activation of a plant innate immune receptor. Cell Host Microbe 2011, 9, 137–146. [Google Scholar] [CrossRef] [PubMed]
- Zipfel, C.; Robatzek, S.; Navarro, L.; Oakeley, E.J.; Jones, J.D.; Felix, G.; Boller, T. Bacterial disease resistance in Arabidopsis through flagellin perception. Nature 2004, 428, 764–767. [Google Scholar] [CrossRef] [PubMed]
- Riehs-Kearnan, N.; Gloggnitzer, J.; Dekrout, B.; Jonak, C.; Riha, K. Aberrant growth and lethality of Arabidopsis deficient in nonsense-mediated RNA decay factors is caused by autoimmune-like response. Nucleic Acids Res. 2012, 40, 5615–5624. [Google Scholar] [CrossRef] [PubMed]
- Segond, D.; Dellagi, A.; Lanquar, V.; Rigault, M.; Patrit, O.; Thomine, S.; Expert, D. NRAMP genes function in Arabidopsis thaliana resistance to Erwinia chrysanthemi infection. Plant J. 2009, 58, 195–207. [Google Scholar] [CrossRef] [PubMed]
- Bent, A.F.; Kunkel, B.N.; Dahlbeck, D.; Brown, K.L.; Schmidt, R.; Giraudat, J.; Leung, J.; Staskawicz, B.J. RPS2 of Arabidopsis thaliana: A leucine-rich repeat class of plant disease resistance genes. Science 1994, 265, 1856–1860. [Google Scholar] [CrossRef] [PubMed]
- Katiyar-Agarwal, S.; Gao, S.; Vivian-Smith, A.; Jin, H. A novel class of bacteria-induced small RNAs in Arabidopsis. Genes Dev. 2007, 21, 3123–3134. [Google Scholar] [CrossRef] [PubMed]
- Yaish, M.W.F.; Sosa, D.; Vences, F.J.; Vaquero, F. Genetic mapping of quantitative resistance to race 5 of Pseudomonas syringae pv. phaseolicola in common bean. Euphytica 2006, 152, 397–404. [Google Scholar] [CrossRef]
- Velich, I.; Szarka, J.; Neda, P. Biotic and abiotic stresses in the bean: I. Genetic background of complex resistance to bacterial diseases. Hort. Sci. 1994, 26, 49–53. [Google Scholar]
- Young, N.D. QTL mapping and quantitative disease resistance in plants. Annu. Rev. Phytopathol. 1996, 34, 479–501. [Google Scholar] [CrossRef] [PubMed]
- Marcel, T.C.; Gorguet, B.; Ta, M.T.; Kohutova, Z.; Vels, A.; Niks, R.E. Isolate specificity of quantitative trait loci for partial resistance of barley to Puccinia hordei confirmed in mapping populations and near-isogenic lines. New Phytol. 2008, 177, 743–755. [Google Scholar] [CrossRef] [PubMed]
- Kou, Y.; Wang, S. Broad-spectrum and durability: Understanding of quantitative disease resistance. Curr. Opin. Plant Biol. 2010, 13, 181–185. [Google Scholar] [CrossRef] [PubMed]
- Asensio, C.; Martin, E.; Montoya, J.L. Inheritance of resistance to race 1 of Pseudomonas syringae pv. phaseolicola in some varieties of beans. Investigación Agraria Producción y Protección Vegetal 1993, 8, 445–456. [Google Scholar]
- Cardenas, F.; Adams, M.W.; Andersen, A. The genetic system for reaction of field beans (Phaseolus vulgaris L.) to infection by three physiologic races of Colletotrichum lindemuthianum. Euphytica 1964, 13, 178–186. [Google Scholar]
- Blazey, D.; Smith, P.; Gentile, A.; Miyagawa, S. Nematode resistance in common beans. J. Hered. 1964, 55, 20–22. [Google Scholar] [CrossRef]
- Finke, M.L.; Coyne, D.P.; Steadman, J.R. The inheritance and association of resistance to rust, common bacterial blight, plant habit and foliar abnormalities in Phaseolus vulgaris L. Euphytica 1986, 35, 969–982. [Google Scholar] [CrossRef]
- Chataika, B.Y.E.; Bokosi, J.M.; Kwapata, M.B.; Chirwa, R.M.; Mwale, V.M.; Mnyenyembe, P.; Myers, J.R. Performance of parental genotypes and inheritance of Angular Leaf Spot (Phaeosariopsis griseola) resistance in the common bean (Phaseolus vulgaris). Afr. J. Biotechnol. 2011, 9, 4398–4406. [Google Scholar]
- Miklas, P.N.; Smith, J.R.; Riley, R.; Grafton, K.F.; Singh, S.P.; Jung, G.; Coyne, D.P. Marker-assisted breeding for pyramided resistance to common bacterial blight in common bean. Annu. Rep. Bean Improv. Coop. USA 2000, 43, 39–40. [Google Scholar]
- Mutlu, N.; Miklas, P.N.; Coyne, D.P. Resistance gene analog polymorphism (RGAP) Markers co-localize with disease resistance genes and QTL in common bean. Mol. Breed. 2006, 17, 127–135. [Google Scholar] [CrossRef]
- Fall, A.L.; Byrne, P.F.; Jung, J.; Coyne, D.P.; Brick, M.A.; Schwartz, H.F. Detection and mapping of a major locus for fusarium wilt resistance in common bean. Crop Sci. 2001, 41, 1494–1498. [Google Scholar] [CrossRef]
- Duncan, R.W.; Singh, S.P.; Gilbertson, R.L. Interaction of common bacterial blight bacteria with disease resistance quantitative trait loci in common bean. Phytopathology 2011, 101, 425–435. [Google Scholar] [CrossRef] [PubMed]
- Thompson, D.L.; Bergquist, R.R. Inheritance of mature plant resistance to Helminthosporium maydis race 0 in maize. Crop Sci. 1984, 24, 807–811. [Google Scholar] [CrossRef]
- Robertson, D.S. A possible technique for isolating genic DNA for quantitative traits in plants. J. Theor. Biol. 1985, 117, 1–10. [Google Scholar] [CrossRef]
- Maleck, K.; Levine, A.; Eulgem, T.; Morgan, A.; Schmid, J.; Lawton, K.A.; Dangl, J.L.; Dietrich, R.A. The transcriptome of Arabidopsis thaliana during systemic acquired resistance. Nat. Genet. 2000, 26, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Jeuken, M.J.W.; Zhang, N.W.; McHale, L.K.; Pelgrom, K.; den Boer, E.; Lindhout, P.; Michelmore, R.W.; Visser, R.G.F.; Niks, R.E. Rin4 causes hybrid necrosis and race-specific resistance in an interspecific lettuce hybrid. Plant Cell 2009, 21, 3368–3378. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Caldwell, K.S.; Wroblewski, T.; Wright, M.E.; Michelmore, R.W. Proteolysis of a negative regulator of innate immunity is dependent on resistance genes in tomato and Nicotiana benthamiana and induced by multiple bacterial effectors. Plant Cell 2009, 21, 2458–2472. [Google Scholar] [CrossRef] [PubMed]
- Selote, D.; Kachroo, A. RIN4-like proteins mediate resistance protein-derived soybean defense against Pseudomonas syringae. Plant Signal. Behav. 2010, 5, 1453–1456. [Google Scholar] [CrossRef] [PubMed]
- Ghosh, A.K.; Neha, C.; Rajakumari, S.; Daum, G.; Rajasekharan, R. AT4G24160, a soluble acyl-coenzyme A-dependent lysophosphatidic acid acyltransferase. Plant Physiol. 2009, 151, 869–881. [Google Scholar] [CrossRef] [PubMed]
- Garcia, R.A.V.; Rangel, P.N.; Brondani, C.; Martins, W.S.; Melo, L.C.; Carneiro, M.S.; Borba, T.C.O.; Brondani, R.P.V. The characterization of a new set of EST-derived simple sequence repeat (SSR) markers as a resource for the genetic analysis of Phaseolus vulgaris. BMC Genet. 2011, 12, 41. [Google Scholar] [CrossRef] [PubMed]
- King, E.D.; Ward, M.K.; Raney, D.E. Two simple media for the demonstration of pyocyanin and fluorescin. J. Lab. Clin. Med. 1954, 44, 301–307. [Google Scholar] [PubMed]
- Schwartz, H.F.; Brick, M.; Harveson, R.; Franc, G. Disease Management. In Dry Bean Production and Pest Management, 2nd ed.; Schwartz, H.F., Brick, M., Harveson, R., Franc, G., Eds.; APS Press: Minneapolis, MN, USA, 2004; pp. 109–143. [Google Scholar]
- Mills, L.J.; Silbernagel, M.J. A rapid screening technique to combine resistance to halo blight and bean common mosaic virus in Phaseolus vulgaris L. Euphytica 1992, 58, 201–208. [Google Scholar] [CrossRef]
- Chen, D.H.; Ronald, P.C. A rapid DNA minipreparation method suitable for AFLP and other PCR applications. Plant Mol. Biol. Rep. 1999, 17, 53–57. [Google Scholar] [CrossRef]
- Foolad, M.R.; Jones, R.A. Mapping salt-tolerance genes in tomato (Lycopersicon esculentum) using trait-based marker analysis. Theor. Appl. Genet. 1993, 87, 184–192. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.P.; Lin, G.Y.; Niño-Liu, D.; Foolad, M.R. Mapping QTLs conferring early blight (Alternaria solani) resistance in a Lycopersicon esculentum X L. hirsutum cross by selective genotyping. Mol. Breed. 2003, 12, 3–19. [Google Scholar] [CrossRef]
- Coque, M.; Gallais, A. Genomic regions involved in response to grain yield selection at high and low nitrogen fertilization in maize. Theor. Appl. Genet. 2006, 112, 1205–1220. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Wang, J.; Crouch, J.H.; Xu, Y. Efficiency of selective genotyping for genetic analysis of complex traits and potential applications in crop improvement. Mol. Breed. 2010, 26, 493–511. [Google Scholar] [CrossRef]
- Benchimol, L.L.; de Campos, T.; Carbonell, S.A.M.; Colombo, C.A.; Chioratto, A.F.; Formighieri, E.F.; Gouvêa, L.R.L.; de Souza, A.P. Structure of genetic diversity among common bean (Phaseolus vulgaris L.) varieties of Mesoamerican and Andean origins using new developed microsatellite markers. Genet. Resour. Crop Evol. 2007, 54, 1747–1762. [Google Scholar] [CrossRef]
- Cardoso, J.M.K.; Oblessuc, P.R.; Campos, T.; Sforça, D.A.; Carbonell, S.A.M.; Chioratto, A.F.; Formighieri, E.F.; de Souza, A.P.; Benchimol, L.L. New microsatellite markers developed from an enriched microsatellite common bean library. Pesquisa Agropecuária Brasileira 2008, 43, 929–936. [Google Scholar] [CrossRef]
- Campos, T.; Oblessuc, P.R.; Sforça, D.A.; Cardoso, J.M.K.; Baroni, R.M.; Sousa, A.B.; Carbonell, S.A.M.; Chioratto, A.F.; Garcia, A.A.F.; Rubiano, L.B.; et al. Inheritance of growth habit detected by genetic linkage analysis using microsatellites in the common bean (Phaseolus vulgaris L.). Mol. Breed. 2011, 27, 549–560. [Google Scholar] [CrossRef]
- Gaitán-Solís, E.; Duque, M.C.; Edwards, K.J.; Tohme, J. Microsatellite repeats in common bean (Phaseolus vulgaris) isolation, characterization, and cross species amplification in Phaseolus ssp. Crop Sci. 2002, 42, 2128–2136. [Google Scholar] [CrossRef]
- Blair, M.W.; Torres, M.M.; Pedraza, F.; Giraldo, M.C.; Buendía, H.F.; Hurtado, N. Development of microsatellite markers for common bean (Phaseolus vulgaris L.) based on screening of non-enriched, small-insert genomic libraries. Genome 2009, 52, 772–782. [Google Scholar] [CrossRef] [PubMed]
- Córdoba, J.M.; Chavarro, C.; Schlueter, J.A.; Jackson, S.A.; Blair, M.W. Integration of physical and genetic maps of common bean through BAC-derived microsatellite markers. BMC Genom. 2010, 11, 436. [Google Scholar] [CrossRef] [PubMed]
- Blair, M.W.; Muñoz-Torres, M.; Giraldo, M.C.; Pedraza, F. Development and diversity assessment of Andean-derived, gene-based microsatellites for common bean (Phaseolus vulgaris L.). BMC Plant Biol. 2009, 9, 100. [Google Scholar] [CrossRef] [PubMed]
- Blair, M.W.; Hurtado, N.; Chavarro, C.M.; Muñoz-Torres, M.; Giraldo, M.C.; Pedraza, F.; Tomkins, J.; Wing, R. Gene-based SSR markers for common bean (Phaseolus vulgaris L.) derived from root and leaf tissue ESTs: An integration of the BMc series. BMC Plant Biol. 2011, 11, 50. [Google Scholar] [CrossRef] [PubMed]
- Blair, M.W.; Pedraza, F.; Buendía, H.F.; Gaitán-Solís, E.; Beebe, S.E.; Gepts, P.; Tohme, J. Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L.). Theor. Appl. Genet. 2003, 107, 1362–1374. [Google Scholar] [CrossRef] [PubMed]
- Buso, G.S.C.; Amaral, Z.P.S.; Brondani, R.P.V.; Ferreira, M.E. Microsatellite markers for the common bean Phaseolus vulgaris. Mol. Ecol. Notes 2006, 6, 252–254. [Google Scholar] [CrossRef]
- Grisi, M.C.M.; Blair, M.W.; Gepts, P.; Brondani, C.; Pereira, P.A.A.; Brondani, R.P.V. Genetic mapping of a new set of microsatellite markers in a reference common bean (Phaseolus vulgaris) population BAT93 × Jalo EEP558. Genet. Mol. Res. 2007, 6, 691–706. [Google Scholar] [PubMed]
- Hanai, L.R.; de Campos, T.; Camargo, L.E.; Benchimol, L.L.; Souza, A.P.; Melotto, M.; Carbonell, S.A.; Chioratto, A.F.; Consoli, L.; Formighieri, E.F.; et al. Development, characterization, and comparative analysis of polymorphism at common bean SSR loci isolated from genic and genomic sources. Genome 2007, 50, 266–277. [Google Scholar] [CrossRef] [PubMed]
- SAS. SAS/STAT 9.4 User’s Guide; SAS Institute Inc.: Cary, NC, USA, 2013. [Google Scholar]
- Davis, D.D. Hybrid cotton: Specific problems and potentials. Adv. Agron. 1978, 43, 514–516. [Google Scholar]
- Falconer, D.S.; Mackay, T.F.C. Introduction to Quantitative Genetics, 4th ed.; Pearson: London, UK, 1996. [Google Scholar]
- Wynne, J.C.; Emery, D.A.; Rice, P.W. Combining ability estimates in Arachis hypogaea L. II. Field performance of F1 hybrids. Crop Sci. 1970, 10, 713. [Google Scholar] [CrossRef]
- Gusmini, G.; Wehner, T.C.; Donaghy, S.B. SASQuant: A SAS software program to estimate genetic effects and heritabilities of quantitative traits in populations consisting of 6 related generations. J. Hered. 2007, 98, 345–350. [Google Scholar] [CrossRef] [PubMed]
- Van Ooijen, J. JoinMap 4: Software for the Calculation of Genetic Linkage Maps in Experimental Populations. Kyazma BV: Wageningen, The Netherlands, 2006. Available online: https://www.kyazma.nl/index.php/JoinMap/ (accessed on 12 December 2016).
- Kosambi, D.D. The estimation of map distances from recombination values. Ann. Eugen. 1943, 12, 172–175. [Google Scholar] [CrossRef]
- Stam, P. Construction of integrated genetic linkage maps by means of a new computer package: JoinMap. Plant J. 1993, 3, 739–744. [Google Scholar] [CrossRef]
- Yang, J.; Hu, C.; Hu, H.; Yu, R.; Xia, Z.; Ye, X.; Zhu, J. QTLNetwork: Mapping and visualizing genetic architecture of complex traits in experimental populations. Bioinformatics 2008, 24, 721–723. [Google Scholar] [CrossRef] [PubMed]
- Voorrips, R.E. MapChart: Software for the graphical presentation of linkage maps and QTLs. J. Hered. 2002, 93, 77–78. [Google Scholar] [CrossRef] [PubMed]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef] [PubMed]
- Swarbreck, D.; Wilks, C.; Lamesch, P.; Berardini, T.Z.; Garcia-Hernandez, M.; Foerster, H.; Li, D.; Meyer, T.; Muller, R.; Ploetz, L.; et al. The Arabidopsis Information Resource (TAIR): Gene structure and function annotation. Nucleic Acids Res. 2008, 36, 1009–1014. [Google Scholar] [CrossRef] [PubMed]

| Trait a | Parents | F2 | ||||||
|---|---|---|---|---|---|---|---|---|
| UI3 | Tendergreen | PPAR b | N c | Mean | Range | PF2 b | h2 | |
| Race 1 | ||||||||
| PDC | 1.1 ± 0.10 | 8.5 ± 0.22 | ** | 272 | 3.1 ± 0.17 | 1.0–9.0 | ** | 0.93 |
| PAUDPC | 60.9 ±5.29 | 427.4 ± 20.27 | ** | 272 | 160.4 ± 9.37 | 55.6–500.0 | ** | 0.90 |
| PAREA | 1.8 ± 0.09 | 2.5 ± 0.22 | ** | 267 | 2.0 ± 0.04 | 0.9–5.0 | ** | 0.82 |
| Race 5 | ||||||||
| PLDC | 2.4 ± 0.14 | 8.1 ± 0.14 | ** | 459 | 5.8 ± 0.11 | 1.0–9.0 | ** | 0.84 |
| PLAUDPC | 357.8 ± 22.11 | 1175.5 ± 20.62 | ** | 459 | 830.4 ± 16.44 | 155.6–1400.0 | ** | 0.93 |
| PLAREA | 0.2 ± 0.11 | 1.5 ± 0.14 | ** | 164 | 0.8 ± 0.02 | 0.2–2.5 | ** | 0.70 |
| PDC | 1.3 ± 0.16 | 8.4 ± 0.21 | ** | 271 | 3.3 ± 0.18 | 1.0–9.0 | ** | 0.96 |
| PAUDPC | 70.1 ± 7.58 | 420.9 ± 17.18 | ** | 272 | 165.3 ± 9.65 | 27.8–500.0 | ** | 0.94 |
| PAREA | 1.0 ± 0.12 | 2.8 ± 0.11 | ** | 266 | 2.0 ± 0.04 | 0.9–4.5 | ** | 0.78 |
| Race 7 | ||||||||
| PDC | 1.2 ± 0.13 | 8.6 ± 0.18 | ** | 272 | 2.7 ± 0.17 | 1.0–9.0 | ** | 0.95 |
| PAUDPC | 66.1 ± 7.29 | 463.7 ± 14.20 | ** | 272 | 136.3 ± 8.59 | 55.6–500.0 | ** | 0.94 |
| PAREA | 1.9 ± 0.14 | 2.9 ± 0.22 | ** | 267 | 2.3 ± 0.04 | 1.2–4.7 | ** | 0.73 |
| Race 9 | ||||||||
| PLDC | 2.8 ± 0.08 | 7.9 ± 0.14 | ** | 471 | 6.0 ± 0.11 | 1.0–9.0 | ** | 0.97 |
| PLAUDPC | 393.6 ± 16.24 | 1172.0 ± 24.04 | ** | 471 | 868.3 ± 16.42 | 155.6–1400.0 | ** | 0.95 |
| PLAREA | 0.3 ± 0.05 | 1.6 ± 0.03 | ** | 172 | 1.2 ± 0.06 | 0.6–3.2 | ** | 0.95 |
| PDC | 1.3 ± 0.10 | 8.8 ± 0.17 | ** | 272 | 3.7 ± 0.18 | 1.0–9.0 | ** | 0.96 |
| PAUDPC | 70.1 ± 5.29 | 482.9 ± 9.71 | ** | 272 | 185.1 ± 9.70 | 55.6–500.0 | ** | 0.97 |
| PAREA | 1.6 ± 0.11 | 2.4 ± 0.20 | ** | 266 | 2.2 ± 0.04 | 1.0–4.9 | ** | 0.78 |
| Trait a | Parents | F2 | ||||||
|---|---|---|---|---|---|---|---|---|
| UI3 | A52 | PPAR b | N c | Mean | Range | PF2 b | h2 | |
| Race 1 | ||||||||
| PLDC | 2.6 ± 0.11 | 8.3 ± 0.18 | ** | 402 | 5.5 ± 0.13 | 1.0–9.0 | ** | 0.88 |
| PLAUDPC | 346.1 ± 17.40 | 1182.2 ± 33.40 | ** | 402 | 771.7 ± 18.95 | 38.9–1400.0 | ** | 0.89 |
| PLAREA | 0.2 ± 0.11 | 3.1 ± 0.22 | ** | 161 | 1.4 ± 0.06 | 0.2–4.1 | ** | 0.93 |
| PDC | 1.6 ± 0.13 | 8.0 ± 0.26 | ** | 214 | 3.6 ± 0.21 | 1.0–9.0 | ** | 0.92 |
| PAUDPC | 77.4 ± 5.19 | 429.0 ± 19.61 | ** | 214 | 183.3 ± 10.57 | 27.8–500.0 | ** | 0.88 |
| PAREA | 1.6 ± 0.15 | 3.7 ± 0.07 | ** | 213 | 2.1 ± 0.05 | 0.9–4.5 | ** | 0.81 |
| Race 7 | ||||||||
| PLDC | 2.3 ± 0.12 | 8.2 ± 0.17 | ** | 402 | 5.3 ± 0.13 | 1.0–9.0 | ** | 0.86 |
| PLAUDPC | 307.2 ± 16.67 | 1170.6 ± 27.63 | ** | 402 | 769.5 ± 18.78 | 155.6–1400.0 | ** | 0.84 |
| PLAREA | 0.4 ± 0.08 | 3.2 ± 0.17 | ** | 174 | 2.1 ± 0.04 | 0.4–4.0 | ** | 0.86 |
| PDC | 2.1 ± 0.23 | 8.3 ± 0.18 | ** | 213 | 3.0 ± 0.18 | 1.0–9.0 | ** | 0.88 |
| PAUDPC | 103.2 ± 9.40 | 446.0 ± 8.90 | ** | 214 | 147.8 ± 9.21 | 55.6–500.0 | ** | 0.89 |
| PAREA | 1.4 ± 0.07 | 2.9 ± 0.12 | ** | 214 | 2.3 ± 0.06 | 0.8–5.8 | ** | 0.83 |
| Race 9 | ||||||||
| PDC | 2.2 ± 0.24 | 7.8 ± 0.22 | ** | 214 | 3.6 ± 0.18 | 1.0–9.0 | ** | 0.92 |
| PAUDPC | 107.1 ± 10.44 | 393.5 ± 19.22 | ** | 214 | 173.3 ± 9.39 | 55.6–500.0 | ** | 0.91 |
| PAREA | 1.3 ± 0.07 | 2.8 ± 0.15 | ** | 213 | 2.2 ± 0.05 | 0.7–4.3 | ** | 0.80 |
| Organ | UI3T | UI3A52 | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Ratio | R a | S | χ2 | P b | Ratio | R | S | χ2 | P b | |
| Race 1 | ||||||||||
| Primary leaf | NR | 7:9 | 155 | 246 | 4.2 | 0.04 | ||||
| Pod | 3:1 | 191 | 81 | 3.3 | 0.07 | 3:1 | 150 | 64 | 2.8 | 0.10 |
| Race 5 | ||||||||||
| Primary leaf | 1:3 | 147 | 312 | 12.1 | 0.00 | NR | ||||
| Pod | 3:1 | 190 | 81 | 3.5 | 0.06 | NR | ||||
| Race 7 | ||||||||||
| Primary leaf | NR | 7:9 | 173 | 230 | 0.1 | 0.07 | ||||
| Pod | 3:1 | 204 | 68 | 0.0 | 1.00 | 3:1 | 150 | 64 | 2.8 | 0.10 |
| Race 9 | ||||||||||
| Primary leaf | 1:3 | 151 | 320 | 12.5 | 0.00 | NR | ||||
| Pod | 3:1 | 166 | 106 | 28.3 | 0.00 | 3:1 | 125 | 60 | 32.6 | 0.00 |
| LG | Map Length (cM) | No. Markers | Marker Density (cM/Marker) | Marker Types | |||||
|---|---|---|---|---|---|---|---|---|---|
| SSR a | Fin | PFC b | Pse-race 1 | Pse-race 5 | Pse-race 7 | ||||
| 1 | 46.52 | 7 | 6.6 | 6 | 1 | – | |||
| 2 | 112.69 | 15 | 7.5 | 15 | |||||
| 3 | 58.95 | 13 | 4.5 | 13 | |||||
| 4 | 59.62 | 9 | 6.6 | 9 | |||||
| 5 | 45.96 | 5 | 9.2 | 5 | |||||
| 6 | 88.23 | 12 | 7.4 | 11 | 1 | ||||
| 7 | 34.83 | 3 | 11.6 | 3 | |||||
| 8 | 67.67 | 10 | 6.8 | 10 | |||||
| 9 | 102.30 | 14 | 7.3 | 14 | |||||
| 10 | 53.00 | 11 | 4.8 | 8 | 1 | 1 | 1 | ||
| 11 | 64.23 | 7 | 9.2 | 7 | |||||
| Total | 733.98 | 106 | 81.5 | 101 | 1 | 1 | 1 | 1 | 1 |
| QTL | Marker Interval | LG (Position) a | F Value b | A c | D | h2 | Gene Action d |
|---|---|---|---|---|---|---|---|
| Race 1 | |||||||
| Threshold F value: 7.3 (PAUDPC 1), 6.3 (PAREA1) | |||||||
| PAUDPC1-10 | Pse-race 1-Pse-race 5 | 10 (0.0–7.6) | 18.8 | −133.0 *** | −272.08 *** | 50.09 | OD |
| PAREA1-8 | BM151-BMB174 | 08 (53.9–60.3) | 8.8 | −0.26 *** | −0.84 *** | 35.19 | OD |
| PAREA1-10 | Pse-race1-Pse-race5 | 10 (0.0–7.6) | 30.7 | −0.16 *** | 0.003 | 42.47 | A |
| Race 5 | |||||||
| Threshold F value: 6.0 (PLDC5), 6.0 (PLAUDPC5), 5.5 (PLAREA5), 11.2 (PAUDPC5), 9.0 (PAREA5) | |||||||
| PLDC5-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 6.3 | −1.17 *** | −1.14 *** | 14.01 | D |
| PLAUDPC5-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 7.4 | −151.5 *** | −147.2 *** | 13.09 | D |
| PLAREA5-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 6.4 | −0.39 *** | 0.09 *** | 19.38 | A |
| PAUDPC5-10.1 | Pse-race5-Pse-race7 | 10 (7.6–10.9) | 52.5 | −65.46 *** | −90.14 *** | 83.08 | OD |
| PAUDPC5-10.2 | BMC234-BMC159 | 10 (16.6–19.3) | 17.2 | −133.6 *** | −262.1 *** | 12.35 | OD |
| PAREA5-10 | Pse-race5-Pse-race7 | 10 (7.6–10.9) | 29.6 | −0.18 *** | −0.24 ** | 41.07 | OD |
| Race 7 | |||||||
| Threshold F value: 11.0 (PAUDPC7), 6.3 (PAREA7) | |||||||
| PAUDPC7-10 | Pse-race5-Pse-race7 | 10 (7.6–10.9) | 350.9 | −2.87 *** | −5.84 *** | 89.62 | OD |
| PAREA7-10 | Pse-race5-Pse-race7 | 10 (7.6–10.9) | 6.4 | −0.10 *** | −0.33 *** | 14.27 | OD |
| Race 9 | |||||||
| Threshold F value: 6.0 (PLDC9), 6.1 (PLAUDPC9), 6.0 (PLAREA9), 7.2 (PDC9), 7.5 (PAUDPC9), 6.4 (PAREA9) | |||||||
| PLDC9-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 7.1 | −1.79 *** | −1.71 *** | 6.56 | D |
| PLAUDPC9-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 6.4 | −162.42 *** | −153.13 *** | 9.09 | D |
| PLAREA9-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 6.3 | −0.48 *** | 0.10 ** | 6.42 | A |
| PLAREA9-8 | BMB174-BM238 | 08 (43.0–53.7) | 6.1 | −0.49 *** | −0.11 ** | 5.28 | A |
| PDC9-10 | Pse-race5-Pse-race7 | 10 (7.6–10.9) | 85.1 | −13.2 *** | −2.50 | 69.38 | A |
| PAUDPC9-10 | Pse-race5-Pse-race7 | 10 (7.6–10.9) | 85.3 | −28.5 *** | −22.1 *** | 68.26 | PD |
| PAREA9-8 | BM151-BMB174 | 08 (53.9–60.3) | 8.9 | −0.26 *** | −0.84 *** | 11.34 | OD |
| PAREA9-10 | Pse-race5-Pse-race7 | 10 (7.6–10.9) | 30.7 | −0.16 *** | 0 | 42.54 | A |
| QTL | Marker Interval | LG (Position) a | F Value b | A c | D | h2 | Gene Action d |
|---|---|---|---|---|---|---|---|
| Race 1 | |||||||
| Threshold F value: 6.0 (PLDC1), 6.0 (PLAUDPC1), 6.4 (PLAREA1), 6.3 (PAUDPC1), 6.8 (PAREA1) | |||||||
| PLDC1-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 6.4 | −1.54 *** | −1.42 *** | 8.94 | D |
| PLAUDPC1-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 7.8 | −220.4 *** | −218.0 *** | 10.55 | D |
| PLAREA1-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 7.0 | −0.75 *** | 0 | 25.81 | A |
| PAUDPC1-10 | Pse-race1-Pse-race7 | 10 (0.0–13.3) | 70.9 | −176.7 *** | −699.8 *** | 75.29 | OD |
| PAREA1-10 | Pse-race1-Pse-race7 | 10 (0.0–13.3) | 15.3 | −0.17 *** | −0.65 *** | 32.06 | OD |
| Race 7 | |||||||
| Threshold F value: 6.2 (PLDC7), 6.1 (PLAUDPC7), 8.8 (PLAREA7), 6.5 (PAUDPC7), 9.0 (PAREA5) | |||||||
| PLDC7-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 6.3 | −2.67 *** | −2.51 *** | 9.45 | D |
| PLAUDPC7-10 | BMC159-TBR-017 | 10 (19.3–28.1) | 6.3 | −253.1 *** | −201.5 *** | 9.25 | PD |
| PLAREA7-8 | BMB174-BM238 | 08 (43.0–53.7) | 23.4 | −0.54 *** | −0.46 *** | 33.87 | D |
| PAUDPC7-10 | Pse-race1-Pse-race7 | 10 (0.0–13.3) | 64.9 | −122.7 *** | −179.2 *** | 73.14 | OD |
| PAREA7-10 | Pse-race1-Pse-race7 | 10 (7.6–10.9) | 29.6 | −0.18 *** | −0.24 ** | 41.07 | OD |
| Race 9 | |||||||
| Threshold F value: 6.5 (PDC9), 6.4 (PAUDPC9), 6.2 (PAREA9) | |||||||
| PDC9-10.1 | Pse-race1-Pse-race7 | 10 (0.0–13.3) | 27.5 | −1.88 *** | −1.47*** | 54.51 | PD |
| PDC9-10.2 | BMC234-BMC159 | 10 (16.6–19.3) | 8.9 | −1.47 *** | −2.33*** | 21.56 | OD |
| PAUDPC9-10.1 | Pse-race1-Pse-race7 | 10 (0.0–13.3) | 27.1 | −93.4 *** | −5.9 | 53.58 | A |
| PAUDPC9-10.2 | BmC234-BMC159 | 10 (16.6–19.3) | 9.2 | −75.6 *** | −113.0 *** | 21.89 | OD |
| PAREA9-10 | Pse-race1-Pse-race7 | 10 (0.0–13.3) | 12.9 | −0.23 *** | −0.16 *** | 18.33 | PD |
| Organ | QTL (Races of Psp) | Potential Candidate Gen | Physical Position | Putative Gene Function | Arabidopsis Homolog |
|---|---|---|---|---|---|
| Pod | PAREA (races 1 and 9) | Phvul.008G130600 | Chr08: 21193569…21195702 | RPM1-interacting protein 4 (RIN4) | AT3G25070 |
| Primary Leaf | PLAREA (races 7 and 9) | Phvul.008G139200 | Chr08: 29375892…29378417 | Serine/Threonine protein kinase | AT2G32800 |
| Pod | PDC, PAUDPC, PAREA (races 1, 5, 7 and 9) | Phvul.010G021200 | Chr10: 3035922…3042052 | RPM1-interacting protein 4 (RIN4) | AT3G25070 |
| Pod | PDC, PAUDPC (races 5 and 9) | Phvul.010G088200 | Chr10: 33784293…33794256 | Lysophospholipid acyltransferase 2 | AT2G45670 |
| Primary Leaf | PLDC, PLAUDPC, PLAREA (races 1, 5, 7 and 9) | Phvul.010G110500 | Chr10: 38414889…38418585 | Resistance Associated Macrophage protein (NRAMP) | AT1G47240 |
| Phvul.010G104300 | Chr10: 37472766…37482584 | NL-like protein | AT4G27190 | ||
| Phvul.010G112400 | Chr10: 38745119…38748532 | Pentatricopeptide repeat family (PPR) | AT4G01030 | ||
| Phvul.010G114100 | Chr10: 39045743…39062551 | Pentatricopeptide repeat family (PPR) | AT1G01320 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
González, A.M.; Godoy, L.; Santalla, M. Dissection of Resistance Genes to Pseudomonas syringae pv. phaseolicola in UI3 Common Bean Cultivar. Int. J. Mol. Sci. 2017, 18, 2503. https://doi.org/10.3390/ijms18122503
González AM, Godoy L, Santalla M. Dissection of Resistance Genes to Pseudomonas syringae pv. phaseolicola in UI3 Common Bean Cultivar. International Journal of Molecular Sciences. 2017; 18(12):2503. https://doi.org/10.3390/ijms18122503
Chicago/Turabian StyleGonzález, Ana M., Luís Godoy, and Marta Santalla. 2017. "Dissection of Resistance Genes to Pseudomonas syringae pv. phaseolicola in UI3 Common Bean Cultivar" International Journal of Molecular Sciences 18, no. 12: 2503. https://doi.org/10.3390/ijms18122503
APA StyleGonzález, A. M., Godoy, L., & Santalla, M. (2017). Dissection of Resistance Genes to Pseudomonas syringae pv. phaseolicola in UI3 Common Bean Cultivar. International Journal of Molecular Sciences, 18(12), 2503. https://doi.org/10.3390/ijms18122503






