Effects of Advective-Diffusive Transport of Multiple Chemoattractants on Motility of Engineered Chemosensory Particles in Fluidic Environments
Abstract
1. Introduction
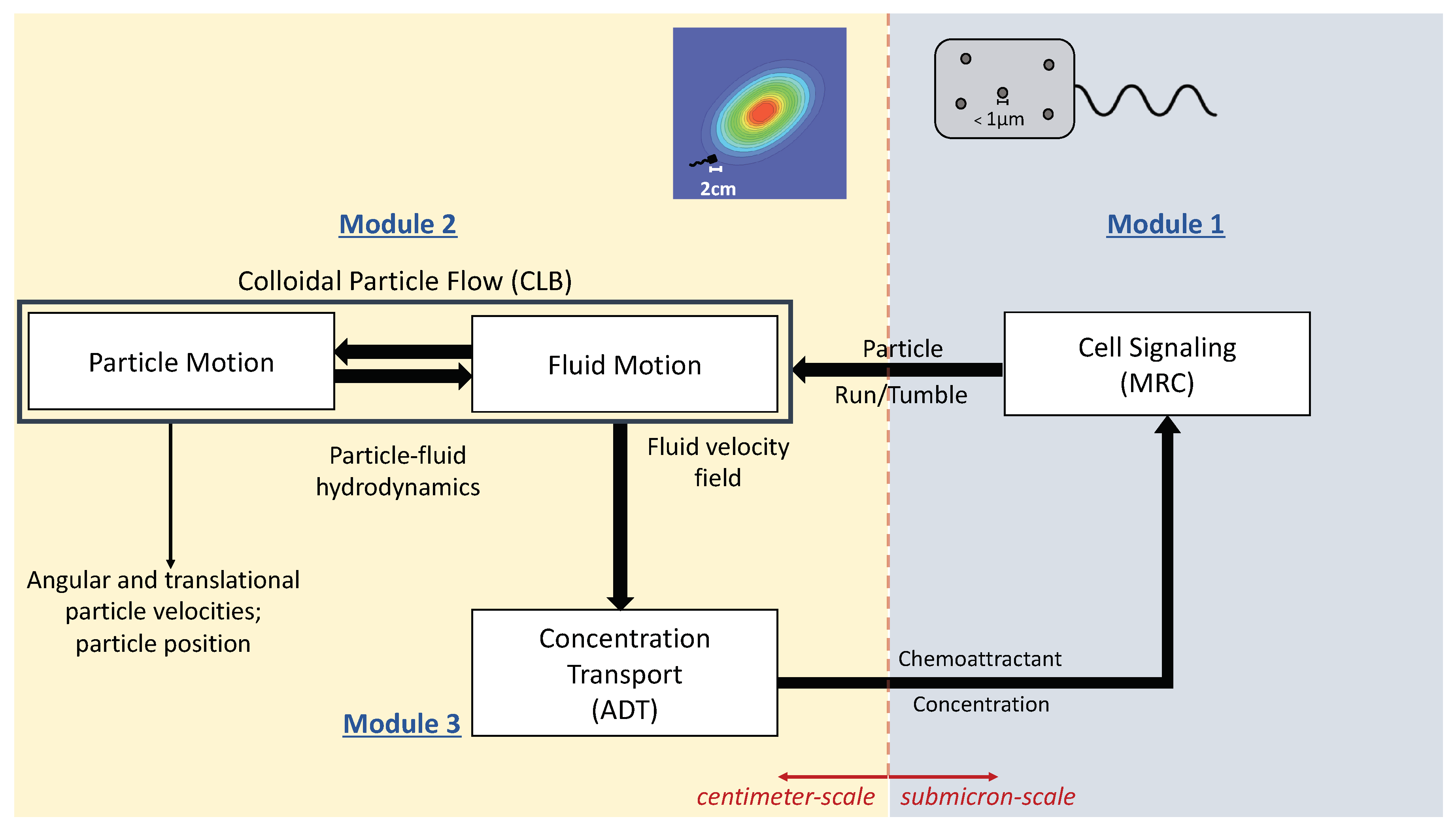
2. Mathematical Framework of the MRC-CLB-ADT Model
2.1. Module 1. Modified RapidCell (MRC) Model for Particle Chemosensing in Two Chemoattractant Fields
2.1.1. Static (Time-Invariant) Concentration Fields
2.1.2. Dynamic (Time-Variant) Concentration Fields
2.2. Module 2. Colloidal Lattice Boltzmann (CLB) Model for Particle-Fluid Interactions
2.2.1. Fluid Flow Submodule (FFS)
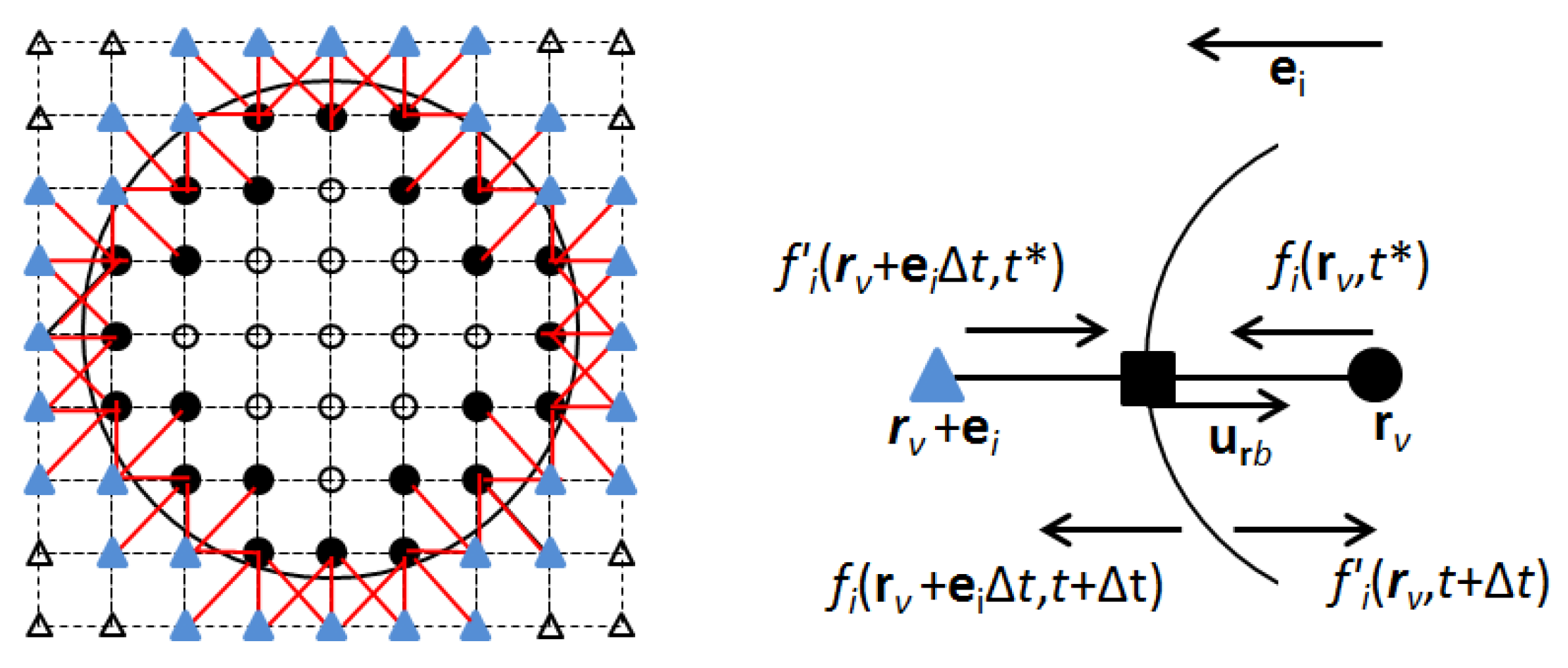
2.2.2. Particle Flow Submodule
2.3. Module 3. Advective-Diffusive Transport (ADT) Model for Chemoattractant Distributions
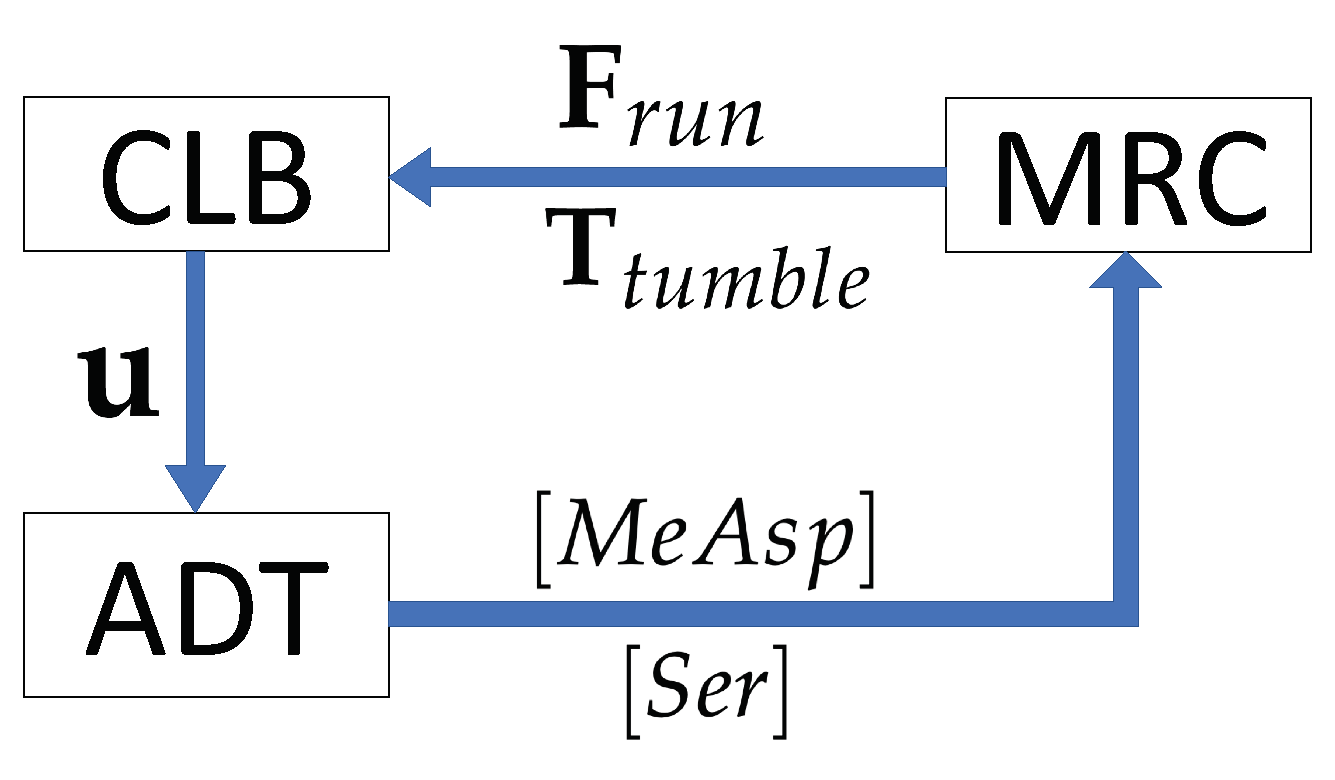
2.4. Coupling of the Modules, MRC-CLB-ADT Model
2.5. Simulation Parameters
3. Results
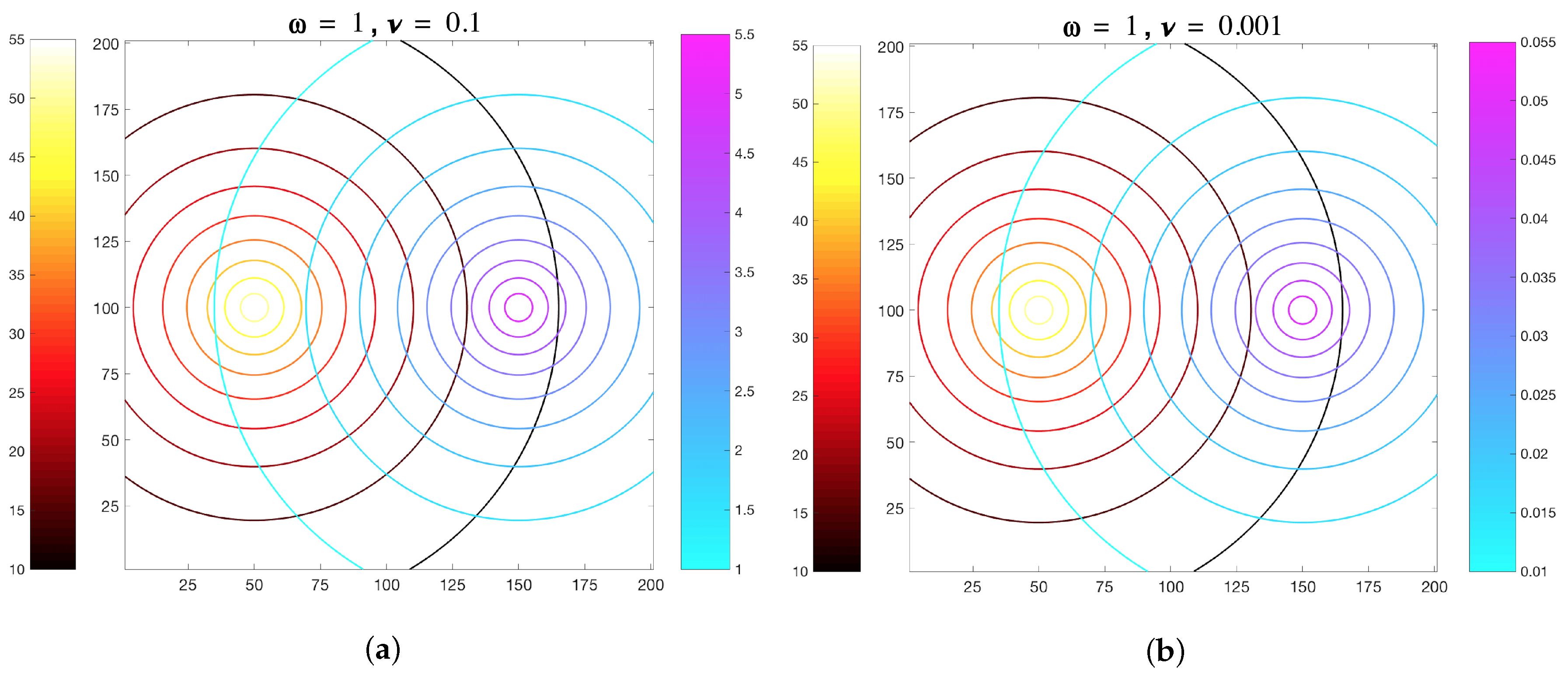
3.1. Simulations with Imposed Temporally-Invariant, Spatially-Variant Chemoattractant Concentrations
3.2. Simulations with Spatiotemporal Variations in Chemoattractant Concentrations Computed via ADT Model
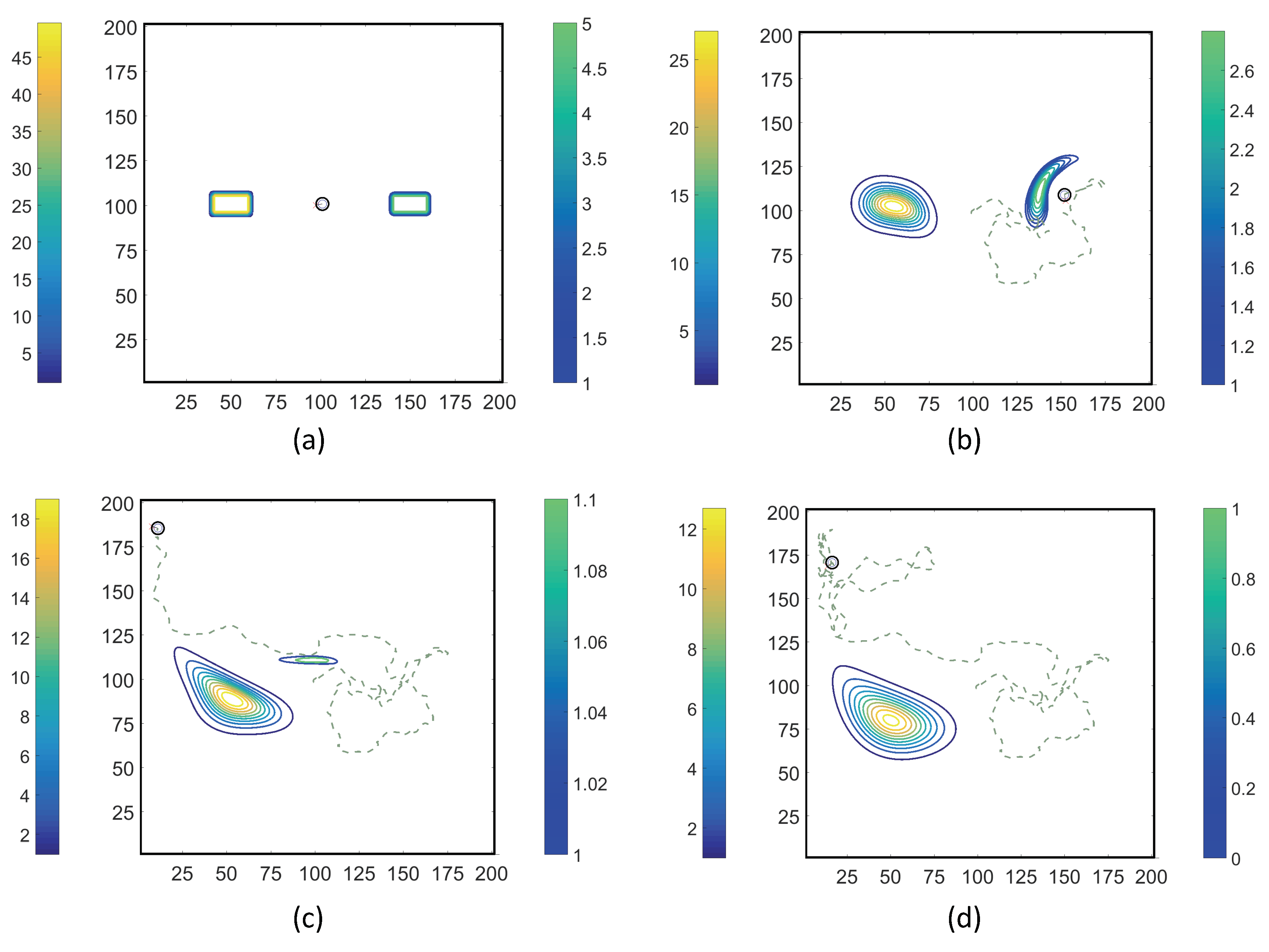
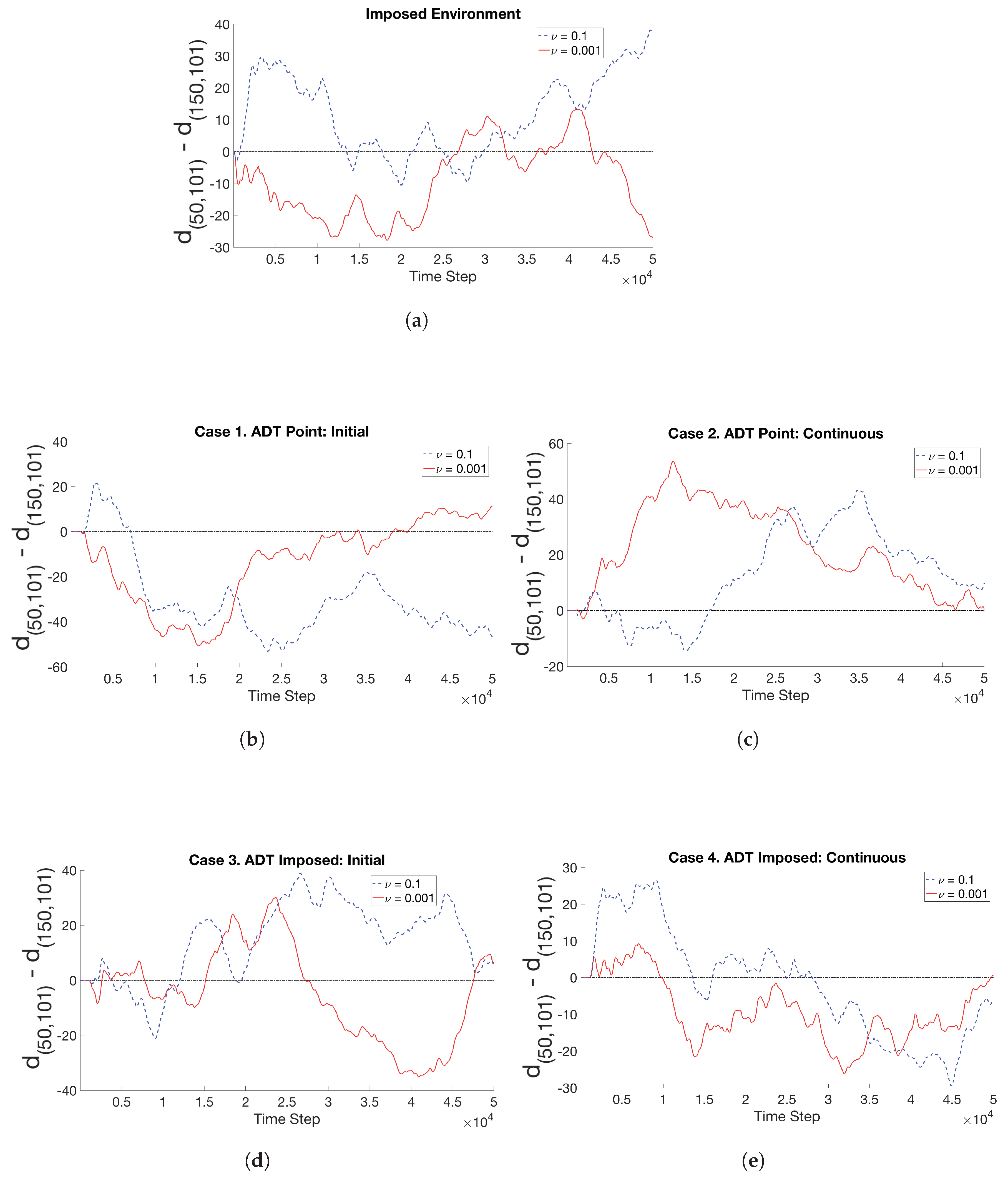
- Case 1. “ADT Point: Initial”: At , MeAsp and Ser were released into the fluid from point sources at and , respectively. No additional chemoattractant releases occurred for . Snapshots from this simulation are shown in Figure 8.
- Case 2. “ADT Point: Continuous”: After the initial condition was set up as in Case 1, of each chemoattractant was released into the fluid in each time-step for from their respective point source locations, at which their maximum concentrations were maintained throughout the simulation. Snapshots from this simulation are shown in Figure 9.
- Case 4. “ADT Imposed: Continuous”: After the initial concentration fields of chemoattractants were established as in Case 3, of each chemoattractant was released into the fluid in each time-step for from their respective point source locations. Snapshots from this simulation are shown in Figure 11.
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
Appendix A.1. Numerical Validation of the ADT Model




Appendix A.2. ADT Model Simulations of Advective-Diffusive Substrate Transport in a Flow Channel

Appendix A.3. With Simulated ADT-Concentrations Moving around Obstacles



Appendix A.4. Tables of Parameters and Variables for All the Modules
Appendix A.4.1. Prescribed Parameters for the MRC Module
| Parameter | Description [Source] | Value |
|---|---|---|
| N | Number of chemoreceptors in receptor cluster [28] | 18 |
| Ratio of Tar to Tsr receptors [28] | 1:1.4 | |
| Dissociation constant in the on state of Tar receptors [17] | 0.012 M | |
| Dissociation constant in the off state of Tar receptors [17] | 0.0017 M | |
| Dissociation constant in the on state of Tsr receptors [17] | 10 M | |
| Dissociation constant in the off state of Tsr receptors [17] | 100 M | |
| Total concentration [17] | 0.16 M | |
| Total concentration [17] | 0.28 M | |
| Total concentration [17] | * | |
| Basal motor bias [17] | ||
| H | Motor Hill coefficient [17] | |
| a | Scaling factor for methylation [17] | |
| b | Scaling factor for demethylation [17] | |
| Rate constant [17] | M | |
| Rate constant [17] | 100 M | |
| Scaling coefficient [17] | M | |
| Rate constant [17] | 0.1 s | |
| Minimum chemoattractant concentration for MeAsp | 0.1 M | |
| Minimum chemoattractant concentration for Ser | 0.1 M | |
| Scaling parameter for MeAsp gradient | 1 | |
| Scaling parameter for Ser gradient | 0.1 or 0.001 | |
| Location of the maximum MeAsp concentration initially | (14.3 cm, 28.9 cm) | |
| Location of the maximum Ser concentration initially | (42.9 cm, 28.9 cm) | |
| Domain length | 57 cm | |
| r | Scaling parameter for domain size | 14.3 cm |
| Variable | Description [Source] |
|---|---|
| F | Total free energy differences between ‘on’ or ‘off’ state [28] |
| Offset energy given by [28] | |
| Chemoattractant MeAsp concentration [28] | |
| Chemoattractant Ser concentration [28] | |
| Probability of the cluster activity [17] | |
| [CheY-P] | Concentration of phosphorylated CheY [17] |
| m | Receptor methylation [17] |
| Motor bias [17] | |
| Horizontal and vertical coordinates |
Appendix A.4.2. System Parameters and Variables for the CLB ad ADT Modules
| Notation | Type | Description | Used by |
|---|---|---|---|
| parameter | speed of sound | FFS, ADT | |
| parameter | unit velocity vectors | FFS, PFS , ADT | |
| variable | population densities associated with fluid flow | FFS, PFS | |
| variable | equilibrium distribution associated fluid flow | FFS | |
| parameter | particle force strength | PFS | |
| parameter | acceleration due to external forces | FFS, PFS | |
| variable | population densities associated with substrate transport | ADT | |
| variable | equilibrium distribution associated substrate transport | ADT | |
| i | variable | index | FFS, PFS, ADT |
| parameter | particle mass | PFS | |
| variable | position vector | FFS, PFS, ADT | |
| variable | position of boundary nodes of ECP | PFS | |
| variable | covered lattice nodes by ECP motion | PFS | |
| variable | uncovered lattice nodes by ECP motion | PFS | |
| variable | position of the ECP’s centroid | PFS | |
| variable | location of the cluster receptor | PFS | |
| variable | distance vector | PFS | |
| parameter | repulsive threshold distance | PFS | |
| parameter | particle radius | PFS | |
| variable | surface to surface distance between the wall and ECP | PFS | |
| variable | position of intra-particle lattice node | PFS | |
| t | variable | time | FFS, PFS, ADT |
| variable | post-collision time | PFS | |
| variable | fluid velocity | FFS, PFS, ADT | |
| C | variable | chemoattractant concentration | ADT |
| D | parameter | diffusion coefficient of chemoatractant | ADT |
| variable | forces associated with running motion of ECP | PFS | |
| variable | hydrodynamic forces | PFS | |
| variable | forces associated with (un)covered lattice nodes | PFS | |
| variable | steric interaction forces between ECP and wall | PFS | |
| variable | total force imposed on ECP | PFS | |
| parameter | moment of inertia of ECP | PFS | |
| parameter | domain length | FFS | |
| M | parameter | Mach number | FFS |
| variable | torque associated with tumble motion of ECP | PFS | |
| variable | translation velocity of of ECP | PFS | |
| variable | rotation angle of the receptor cluster | PFS | |
| parameter | fluid kinematic viscosity | FFS | |
| variable | fluid density | FFS, PFS | |
| parameter | relaxation parameter associated fluid flow | FFS | |
| parameter | relaxation parameter associated substrate transport | PFS | |
| parameter | stiffness parameter associated with steric interaction forces | PFS | |
| parameter | weights associated with the D2Q9 lattice geometry | FFS, PFS, ADT | |
| variable | uniform deviate | PFS | |
| parameter | pressure differential | FFS | |
| parameter | temporal increment | FFS, PFS, ADT | |
| parameter | lattice spacing | FFS, PFS, ADT | |
| variable | angular rotation of ECP | FFS | |
| variable | angular velocity of ECP due to its tumbling motion only | PFS | |
| variable | angular velocity of of ECP | PFS | |
| parameter | time-scale factor associated with ECP’s angular rotation | PFS |
References
- Stanton, M.M.; Sanchez, S. Pushing bacterial biohybrids to in vivo applications. Trends Biotechnol. 2017, 35, 910–913. [Google Scholar] [CrossRef] [PubMed]
- Chien, T.; Doshi, A.; Danino, T. Advances in bacterial cancer therapies using synthetic biology. Curr. Opin. Syst. Biol. 2017, 5, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Felicetti, L.; Femminella, M.; Reali, G.; Lio, P. Applications of molecular communications to medicine: A survey. Nano Commun. Netw. 2016, 7, 27–45. [Google Scholar] [CrossRef]
- Sylvain, M. Bacterial microsystems and microrobots. Biomed. Microdevices 2012, 14, 1033–1045. [Google Scholar]
- Ceylan, H.; Giltinan, J.; Kozielski, K.; Sitti, M. Mobile microrobots for bioengineering applications. Lab Chip 2017, 17, 1705–1724. [Google Scholar] [CrossRef]
- Carlsen, R.W.; Sitti, M. Bio-hybrid cell-based actuators for microsystems. Small 2014, 10, 3831–3851. [Google Scholar]
- Purcell, E.M. Life at low Reynolds number. Am. J. Phys. 1977, 45, 3–11. [Google Scholar] [CrossRef]
- Zhang, L.; Abbott, J.J.; Dong, L.; Kratochvil, B.E.; Bell, D.; Nelson, B.J. Artificial bacterial flagella: Fabrication and magnetic control. Appl. Phys. Lett. 2009, 94, 064107. [Google Scholar] [CrossRef]
- Patino, T.; Mestre, R.; Mestre, S. Miniaturized soft bio-hybrid robotics: A step forward into healthcare applications. Lab Chip 2016, 16, 3626–3630. [Google Scholar] [CrossRef] [PubMed]
- Park, S.J.; Park, S.H.; Cho, S.; Kim, D.M.; Lee, Y.; Ko, S.Y.; Hong, Y.; Choy, H.E.; Min, J.J.; Park, J.O.; et al. New paradigm for tumor theranostic methodology using bacteria-based microrobot. Sci. Rep. 2013, 3, 1–8. [Google Scholar] [CrossRef]
- Shao, J.; Xuan, M.; Zhang, H.; Lin, X.; Wu, Z.; He, Q. Chemotaxis-Guided Hybrid Neutrophil Micromotor for Actively Targeted Drug Transport. Angew. Chem. Int. Ed. 2017, 56, 12935–12939. [Google Scholar] [CrossRef] [PubMed]
- Nelson, B.J.; Kaliakatsos, I.K.; Abbott, J.J. Microrobots for minimally invasive medicine. Annu. Rev. Biomed. Eng. 2010, 12, 55–85. [Google Scholar] [CrossRef]
- Anderson, J.C.; Clarke, E.J.; Arkin, A.P.; Voigt, C.A. Environmentally controlled invasion of cancer cells by engineered bacteria. J. Mol. Biol. 2006, 355, 619–627. [Google Scholar] [CrossRef]
- Zoaby, N.; Shainsky-Roitman, J.P.; Badarneh, S.; Abumanhal, H.; Leshansky, A.; Yaron, S.; Schroeder, A. Autonomous bacterial nanoswimmers target cancer. J. Control. Release 2017, 257, 68–75. [Google Scholar] [CrossRef]
- Felfoul, O.; Mohammadi, M.; Gaboury, L.; Martel, S. Tumor targeting by computer controlled guidance of magnetotactic bacteria acting like autonomous microrobots. In Proceedings of the 2011 IEEE/RSJ International Conference on Intelligent Robots and Systems (IROS), San Francisco, CA, USA, 25–30 September 2011; pp. 1304–1308. [Google Scholar]
- Colin, R.; Sourjik, V. Emergent properties of bacterial chemotaxis pathway. Curr. Opin. Microbiol. 2017, 39, 24–33. [Google Scholar] [CrossRef] [PubMed]
- Vladimirov, N.; Løvdok, L.; Lebiedz, D.; Sourjik, V. Dependence of Bacterial Chemotaxis on Gradient Shape and Adaptation Rate. PLOS Comput. Biol. 2008, 4, e1000242. [Google Scholar] [CrossRef]
- Cortez, R. The method of regularized Stokeslets. SIAM J Sci. Comput. 2001, 23, 1204–1225. [Google Scholar] [CrossRef]
- Başağaoğlu, H.; Allwein, S.; Succi, S.; Dixon, H.; Carrola, J.T., Jr.; Stothoff, S. Two- and three-dimensional lattice-Boltzmann simulations of particle migration in microchannels. Microfluid Nanofluid 2013, 15, 785–796. [Google Scholar] [CrossRef]
- Başağaoğlu, H.; Carrola, J.T., Jr.; Freitas, C.J.; Başağaoğlu, B.; Succi, S. Lattice Boltzmann simulations of vortex entrapment of particles in a microchannel with curved and flat edges. Microfluid Nanofluid 2015, 18, 1165–1175. [Google Scholar] [CrossRef]
- Nguyen, H.; Başağaoğlu, H.; McKay, C.; Carpenter, A.; Succi, S.; Healy, F. Coupled RapidCell and lattice-Boltzmann models to simulate hydrodynamics of bacterial transport in response to chemoattractant gradients in confined domains. Microfluid Nanofluid 2016, 20, 1–14. [Google Scholar] [CrossRef]
- Xu, F.; Bierman, R.; Healy, F.; Nguyen, H. A multiscale model of Escherichia coli chemotaxis from intracellular signaling pathway to motility and nutrient uptake in nutrient gradient and isotropic fluid environments. Comput. Math. Appl. 2016, 71, 2466–2478. [Google Scholar] [CrossRef]
- Lai, R.Z.; Gosink, K.K.; Parkinson, J.S. Signaling Consequences of Structural Lesions that Alter the Stability of Chemoreceptor Trimers of Dimers. J. Mol. Biol. 2017, 429, 2823–2835. [Google Scholar] [CrossRef]
- Pan, W.; Dahlquist, F.W.; Hazelbauer, G.L. Signaling complexes control the chemotaxis kinase by altering its apparent rate constant of autophosphorylation. Protein Sci. 2017, 26, 1535–1546. [Google Scholar] [CrossRef] [PubMed]
- Ud-Din, A.I.M.S.; Roujeinikova, A. Flagellin glycosylation with pseudaminic acid in Campylobacter and Helicobacter: Prospects for development of novel therapeutics. Cell. Mol. Life Sci. 2017, 2018, 1163–1178. [Google Scholar]
- Ma, Q.; Sowa, Y.; Baker, M.A.; Bai, F. Bacterial Flagellar Motor Switch in Response to CheY-P Regulation and Motor Structural Alterations. Biophys. J. 2016, 110, 1411–1420. [Google Scholar] [CrossRef][Green Version]
- Krembel, A.; Colin, R.; Sourjikr, V. Importance of multiple methylation sites in Escherichia coli chemotaxis. PloS ONE 2015, 10, e0145582. [Google Scholar] [CrossRef]
- Edgington, M.; Tindall, M. Understanding the link between single cell and population scale responses of Escherichia coli in differing ligand gradients. Comput. Struct. Biotechnol. J. 2015, 13, 528–538. [Google Scholar] [CrossRef]
- Succi, S. The lattice-Boltzmann Equation for Fluid Dynamics and Beyond; Oxford University Press: New York, NY, USA, 2001. [Google Scholar]
- Bhatnagar, P.L.; Gross, E.P.; Krook, M. A model for collision process in gases. I. Small amplitude processes in charged and neutral one-component systems. Phys. Rev. 1954, 94, 511–525. [Google Scholar] [CrossRef]
- Qian, Y.H.; D’Humieres, D.; Lallemand, P. Lattice BGK models for Navier-Stokes equation. Europhys. Lett. 1992, 17, 479–484. [Google Scholar] [CrossRef]
- Buick, J.M.; Greated, C.A. Gravity in a lattice Boltzmann model. Phys. Rev. E. 2000, 61, 5307–5320. [Google Scholar] [CrossRef]
- Hanasoge, S.M.; Succi, S.; Orszag, S. Lattice Boltzmann method for electromagnetic wave propagation. Europhys. Lett. 2011, 96, 14002. [Google Scholar] [CrossRef]
- Ladd, A.J.C. Numerical simulations of particulate suspensions via a discretized Boltzmann equation. Part 1. Theoretical foundation. J. Fluid Mech. 1994, 271, 285–309. [Google Scholar] [CrossRef]
- Ding, E.J.; Aidun, C. Extension of the Lattice-Boltzmann method for direct simulation of suspended particles near contact. J. Stat. Phys. 2003, 112, 685–708. [Google Scholar] [CrossRef]
- Başağaoğlu, H.; Succi, S.; Wyrick, D.; Blount, J. Particle shape influences settling and sorting behavior in microfluidic domains. Sci. Rep. 2018, 8, 8583. [Google Scholar] [CrossRef]
- Aidun, C.K.; Lu, Y.; Ding, E.J. Direct analysis of particulate suspensions with inertia using the discrete Boltzmann equation. J. Fluid Mech. 1998, 373, 287–311. [Google Scholar] [CrossRef]
- Başağaoğlu, H.; Succi, S. Lattice-Boltzmann simulations of repulsive particle-particle and particle-wall interactions: Coughing and choking. J. Chem. Phys. 2010, 132, 134111. [Google Scholar] [CrossRef] [PubMed]
- Başağaoğlu, H.; Meakin, P.; Succi, S.; Redden, G.R.; Ginn, T.R. Two-dimensional lattice-Boltzmann simulation of colloid migration in rough-walled narrow flow channels. Phys. Rev. E 2008, 77, 031405. [Google Scholar] [CrossRef]
- Feng, J.; Hu, H.H.; Joseph, D.D. Direct simulation of initial value problems for the motion of solid bodies in a Newtonian fluid Part 1. Sedimentation. J. Fluid. Mech. 1994, 261, 95–134. [Google Scholar] [CrossRef]
- Gibbs, R.J.; Matthews, M.D.; Link, D.A. The relationship between sphere size and settling velocity. J. Sedimentary Petrol. 1971, 41, 7–18. [Google Scholar]
- Hilpert, M. Lattice-Boltzmann model for bacterial chemotaxis. J. Math. Biol. 2005, 51, 302–332. [Google Scholar] [CrossRef] [PubMed]
- Kang, Q.; Zhang, D.; Chen, S.; He, X. Lattice Boltzmann simulation of chemical dissolution in porous media. Phys. Rev. E 2002, 85, 036318. [Google Scholar] [CrossRef]
- Landsberg, P.T. Grad v or grad(Dv)? J. Appl. Phys. 1984, 56, 1119. [Google Scholar] [CrossRef]
- Schnitzer, M.J. Theory of continuum random walks and application to chemotaxis. Phys. Rev. E 1993, 48, 2553–2568. [Google Scholar] [CrossRef]
- Boon, J.P.; Lutsko, J.F. Temporal Diffusion: From Microscopic Dynamics to Generalised Fokker–Planck and Fractional Equations. J. Stat. Phys. 2017, 166, 1441–1454. [Google Scholar] [CrossRef]
- Andreucci, D.; Cirillo, E.N.; Colangeli, M.; Gabrielli, D. Fick and Fokker-Planck diffusion law in inhomogeneous media. J. Stat. Phys. 2019, 174, 469–493. [Google Scholar] [CrossRef]
- Wu, F.; Shi, W.; Liu, F. A lattice Boltzmann model for the Fokker–Planck equation. Commun. Nonlinear Sci. Numer. Simul. 2012, 17, 2776–2790. [Google Scholar] [CrossRef]
- Ma, Y.; Zhu, C.; Ma, P.; Yu, K. Studies on the Diffusion Coefficients of Amino Acids in Aqueous Solutions. J. Chem. Eng. Data 2005, 50, 1192–1196. [Google Scholar] [CrossRef]
- Frankel, N.W.; Pontius, W.; Dufour, Y.S.; Long, J.; Hernandez-Nunez, L.; Emonet, T. Adaptability of non-genetic diversity in bacterial chemotaxis. eLife 2014, 3, e03526. [Google Scholar] [CrossRef]
- Jasuja, R.; Lin, Y.; Trentham, D.R.; Khan, S. Response tuning in bacterial chemotaxis. Proc. Natl. Acad. Sci. USA 1999, 96, 11346–11351. [Google Scholar] [CrossRef]
- Fetter, G.W. Contaminant Hydrogeology; Prentice-Hall Inc.: Upper Saddle River, NJ, USA, 1993. [Google Scholar]
- Matlab R2017a. Available online: https://www.mathworks.com/ (accessed on 4 May 2019).
- Mello, B.A.; Tu, Y. Effects of adaptation in maintaining high sensitivity over a wide range of backgrounds for Escherichia coli chemotaxis. Biophys. J. 2007, 92, 2329–2337. [Google Scholar] [CrossRef] [PubMed]
- Frank, V.; Piñas, G.E.; Cohen, H.; Parkinson, J.S.; Vaknin, A. Networked Chemoreceptors Benefit Bacterial Chemotaxis Performance. mBIO 2016, 6, e01824-16. [Google Scholar] [CrossRef] [PubMed]
- Bolster, D.; Dentz, M.; Le Borgn, T. Hypermixing in linear shear flow. Water Resour. Res. 2011, 47, W09602. [Google Scholar] [CrossRef]






© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
King, D.; Başağaoğlu, H.; Nguyen, H.; Healy, F.; Whitman, M.; Succi, S. Effects of Advective-Diffusive Transport of Multiple Chemoattractants on Motility of Engineered Chemosensory Particles in Fluidic Environments. Entropy 2019, 21, 465. https://doi.org/10.3390/e21050465
King D, Başağaoğlu H, Nguyen H, Healy F, Whitman M, Succi S. Effects of Advective-Diffusive Transport of Multiple Chemoattractants on Motility of Engineered Chemosensory Particles in Fluidic Environments. Entropy. 2019; 21(5):465. https://doi.org/10.3390/e21050465
Chicago/Turabian StyleKing, Danielle, Hakan Başağaoğlu, Hoa Nguyen, Frank Healy, Melissa Whitman, and Sauro Succi. 2019. "Effects of Advective-Diffusive Transport of Multiple Chemoattractants on Motility of Engineered Chemosensory Particles in Fluidic Environments" Entropy 21, no. 5: 465. https://doi.org/10.3390/e21050465
APA StyleKing, D., Başağaoğlu, H., Nguyen, H., Healy, F., Whitman, M., & Succi, S. (2019). Effects of Advective-Diffusive Transport of Multiple Chemoattractants on Motility of Engineered Chemosensory Particles in Fluidic Environments. Entropy, 21(5), 465. https://doi.org/10.3390/e21050465






