- Article
Genetic Characterization of the Arabic-Speaking Population from the Casablanca-Settat Region Using Autosomal STR Markers: Understanding the Interplay of Geography and Language in Moroccan Population History
- Othmane Essoubaiy,
- Adnane Hakem and
- Brahim El Houate
- + 4 authors
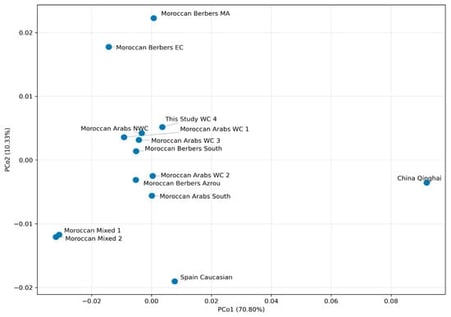
Background/Objectives: The Casablanca-Settat region of Morocco, located at the interface between Arab and Amazigh cultural zones, has only recently been investigated using autosomal short tandem repeat (STR) markers. The objective of this study was to characterize the genetic diversity and forensic efficiency of 15 autosomal STR loci in the Casablanca-Settat population and to evaluate its genetic relationships with other Moroccan populations. Methods: Fifteen autosomal STR loci were genotyped in 138 unrelated Arabic-speaking individuals from the Casablanca-Settat region. Allele frequencies, Hardy–Weinberg equilibrium, and standard forensic parameters were calculated. The genetic structure of the population was further examined through comparative analyses with 12 previously published Moroccan reference populations using multivariate and phylogenetic approaches. Results: A total of 146 distinct alleles were identified across the 15 loci. D18S51 was the most polymorphic marker (Ho = 0.9203), whereas D3S1358, TPOX, D5S818, and D16S539 exhibited lower allelic diversity. No statistically significant deviation from Hardy–Weinberg equilibrium was detected after correction for multiple testing. The combined power of discrimination exceeded 0.99, and the combined power of exclusion reached 0.99999965, demonstrating the high forensic efficiency of the STR panel. Population structure analyses positioned the Casablanca-Settat population within an intermediate genetic cluster, closely related to central Moroccan populations, consistent with historical gene flow and admixture. Conclusions: This study provides robust autosomal STR reference data for the Casablanca-Settat population, confirming the suitability of these markers for forensic identification in Morocco and offering valuable insights into regional population structure and genetic diversity.
21 February 2026