Abstract
Rapid eye movement sleep (REM) is a critical sleep stage associated with several sleep disorders, including sleep apnea and rapid eye movement sleep behavior disorder. Polysomnography (PSG) is the gold standard for identifying REM periods and diagnosing sleep disorders. However, PSG is typically conducted in sleep medicine centers using specialized equipment, where sleep experts assess sleep conditions through measurements such as brain activity, respiration, heart activity, and eye movements. An overnight stay in a sleep laboratory can adversely affect a patient’s natural sleep quality, introducing the risk of iatrogenic sleep disturbances. Recent studies have explored sleep stage detection using lightweight wearable devices, such as smartwatches, which offer lower cost but rely on a limited set of psychological signals. In this study, we propose a machine learning approach for REM sleep staging based solely on breathing rate (BR) and heart rate (HR), without relying on PSG recordings. Experimental evaluations conducted on the Dreamt dataset demonstrate the feasibility of the proposed approach and its potential to provide meaningful information for sleep staging. Future work will focus on developing a fully non-contact REM detection framework by integrating video-based estimation of HR and BR.
1. Introduction
Sleep staging provides essential information related to sleep quality and has a significant impact on individual health. Traditionally, accurate identification of sleep stages is performed by physicians or trained technicians based on polysomnography (PSG) signals, which makes the analysis process labor-intensive and costly. With the rapid advancement of data-driven and machine learning techniques, numerous automated and advanced sleep staging methods have been proposed [1,2]. However, sleep analysis based on overnight PSG recordings remains both expensive and time-consuming. To address this limitation, lightweight solutions leveraging wearable devices have been developed to enable basic sleep assessment in a more accessible and convenient manner [3,4]. In this work, we propose a REM detection method that relies solely on heart rate and breathing rate signals at a limited sample rate.
2. Method
To provide a low-cost and convenient approach for identifying REM periods, we aim to develop a data-driven solution that relies solely on two relatively accessible physiological signals: heart rate and breathing rate.
2.1. Dreamt Dataset
The Dreamt dataset [5] was used to develop and evaluate the proposed method. The dataset was released by the Duke Sleep Disorders Center and comprises signals sampled at two different rates, 64 Hz and 100 Hz. The 64 Hz data are based on measurements collected using the Empatica E4 smartwatch, while the 100 Hz data originate from polysomnography (PSG) recordings. The Empatica E4 was used to collect four physiological signals, including blood volume pulse, accelerometry, skin temperature, and electrodermal activity. In contrast, the PSG data include electroencephalography (EEG), electrocardiography (ECG), electrooculography (EOG), and respiratory channels. Both data sources are accompanied by sleep stage labels annotated by professional technicians, with detailed annotation procedures described in the original dataset publication. Because breathing rate is not directly provided in the dataset, it was estimated from the thoracic (Thorax) respiratory channel of the PSG signals. Breathing rate was derived by identifying the dominant frequency corresponding to the maximum magnitude in the Fourier transform spectrum within each 30 s segment.
2.2. Feature Engineering
According to previous studies [4,5,6], heart rate variability (HRV) is an important indicator for REM sleep identification. However, to obtain accurate HRV values, physicians must first measure RR intervals—defined as the time intervals between consecutive R peaks in electrocardiography (ECG) signals—and then compute the variability among these RR intervals. One commonly used HRV metric is the standard deviation of NN intervals (SDNN), which is calculated as follows:
where denotes the n-th RR interval among N successive heartbeats, is the mean RR interval, and HRV represents the heart rate variability within the analyzed period.
Instead of utilizing SDNN as described above, we adopt an alternative measure that relies solely on coarse-grained heart rate values. Following the original concept of HRV, which characterizes variations between consecutive heartbeats within a specified time window, we propose an indicator that captures similar information in a simplified manner. Specifically, we define the sequence of heart rate difference (SHRD) as the difference between heart rate values at two successive time steps.
where and denote the heart rate and the corresponding heart rate difference at time step n, respectively. Using the raw heart rate and the estimated breathing rate, we construct a three-dimensional feature vector at each time step. A sequence of length m is then used as the input to the proposed machine learning model.
2.3. Deep Learning Model Architecture
A gated recurrent unit (GRU) [7] was used as the architecture of the proposed prediction model to process sequential input data. Given an input sequence of length m, the resulting m × 3 feature matrix is fed into a unidirectional two-layer GRU. The last component of the output sequence is then processed by two consecutive fully connected layers to perform the REM sleep classification task. The overall model architecture is summarized in Table 1.

Table 1.
Summary of proposed model architecture.
3. Results
Multiple REM experiments were conducted under different settings to evaluate the effectiveness of the proposed method. We compared model performance using three different input lengths: 30, 60, and 90 s.
3.1. Experiments
The original Dreamt dataset consists of 100 recordings from individual patients. To reduce noise and ensure sufficient representation of REM events, we excluded records with total REM duration of less than 20 min throughout the night. After filtering, 74 recordings remained, which were divided into training and testing sets using an 80:20 split. In addition, 25% of the training set was reserved for validation during the training process. Because the annotated REM periods vary in length, we segmented them into uniform time intervals. After preprocessing, approximately 14,000 segments were obtained for training, while both the validation and testing sets contained approximately 5000 segments each. Across the training, validation, and testing sets, the ratio of REM to NREM samples remained approximately 30:70.
Sequential HR and BR features were used as model inputs, and recurrent neural network (RNN) architectures were adopted for classification. The model was optimized using the AdamW optimizer with a learning rate of 0.001, and training was conducted for 100 epochs.
3.2. Numerical Results
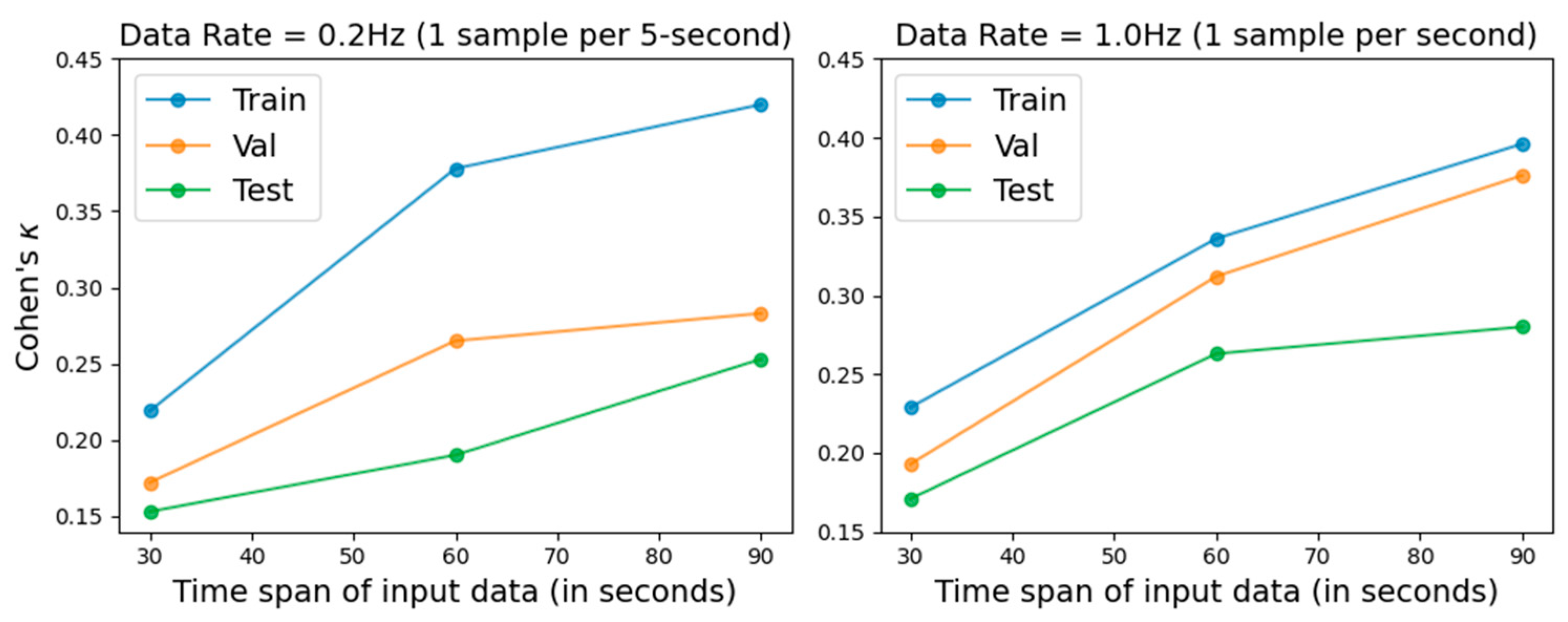
We evaluated model performance using input time spans of 30, 60, and 90 s, with Cohen’s kappa coefficient employed as the primary performance metric. In addition, two sampling rates, 0.2 Hz and 1.0 Hz, were evaluated. The results are illustrated in Figure 1, and the numerical results are presented in Table 2 and Table 3. Longer input spans improved prediction performance across all datasets. In addition, the 1.0 Hz sampling rate produced more robust results than 0.2 Hz, which aligns with the intuitive expectation that higher temporal resolution can capture more informative physiological variations for the machine learning model. Previous PSG-based studies [7] reported Cohen’s kappa values ranging from 0.47 to –0.52.
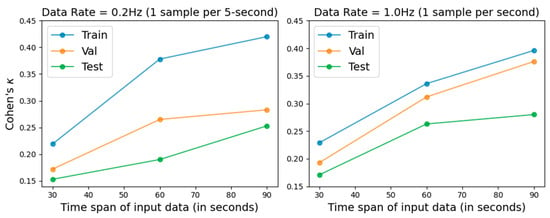
Figure 1.
REM detection performance under different sampling rates and input time spans, evaluated using Cohen’s kappa on the training, validation, and testing sets.

Table 2.
REM detection performance with a sampling interval of 5 s per sample (0.2 Hz).

Table 3.
REM detection performance with a sampling interval of 1 s per sample (1.0 Hz).
4. Conclusions
In this study, a data-driven REM recognition approach based on heart rate and breathing rate was presented. We proposed a simple REM indicator based on the difference between consecutive heart rate values as an alternative to traditional HRV measures. Leveraging modern deep learning techniques, an RNN-based REM prediction model was developed. A series of experiments were conducted to evaluate the effectiveness of the proposed approach. The experimental results indicated that, although a performance gap remains compared with PSG-based approaches, the proposed method provides a promising basis for further improvement. As the developed method relies on HR and BR signals, future work will focus on integrating video-based HR and BR estimation to enable a fully non-contact REM detection framework.
Author Contributions
Conceptualization, C.-Y.J. and T.-I.T.; methodology, C.-Y.J.; software, C.-Y.J.; validation, T.-I.T., C.-Y.J. and S.-H.H.; writing—original draft preparation, C.-Y.J.; project administration, S.-H.H.; funding acquisition, S.-H.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Institutes of Applied Research, Taiwan, R.O.C.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
The data that support the findings of this study are openly available in https://doi.org/10.13026/7r9r-7r24 (accessed on 28 June 2025).
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Lee, H.; Choi, Y.R.; Lee, H.K.; Jeong, J.; Hong, J.; Shin, H.W.; Kim, H.S. Explainable vision transformer for automatic visual sleep staging on multimodal PSG signals. NPJ Digit. Med. 2025, 8, 55. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.K.; Yang, J.; Hershkovich, L.; Jeong, H.; Chen, B.; Singh, K.; Roghanizad, A.R.; Shandhi, M.M.H.; Spector, A.R.; Dunn, J. Addressing Wearable Sleep Tracking Inequity: A New Dataset and Novel Methods for a Population with Sleep Disorders. In Proceedings of the Fifth Conference on Health, Inference, and Learning, New York, NY, USA, 27–28 June 2024. [Google Scholar]
- Yaso, M.; Nuruki, A.; Tsujimura, S.; Yunokuchi, K. Detection of REM Sleep by Heart Rate. In Proceedings of the First International Workshop on Kansei, Fukuoka, Japan, 2–3 February 2006. [Google Scholar]
- Klein, A.; Velicu, O.R.; Madrid, N.M.; Seepold, R. Sleep Stages Classification Using Vital Signals Recordings. In Proceedings of the 12th International Workshop on Intelligent Solutions in Embedded Systems (WISES), Ancona, Italy, 29–30 October 2015. [Google Scholar]
- Wang, K.; Yang, J.; Shetty, A.; Dunn, J. DREAMT: Dataset for real-time sleep stage estimation using multisensor wearable technology (version 2.1.0). PhysioNet 2025, RRID:SCR_007345. [Google Scholar] [CrossRef]
- Chung, J.; Gülçehre, Ç.; Cho, K.; Bengio, Y. Empirical Evaluation of Gated Recurrent Neural Networks on Sequence Modeling. In Proceedings of the NIPS 2014 Deep Learning and Representation Learning Workshop, Montreal, QC, Canada, 8–13 December 2014. [Google Scholar]
- Radha, M.; Fonseca, P.; Moreau, A.; Ross, M.; Cerny, A.; Anderer, P.; Long, X.; Aarts, R.M. Sleep stage classification from heart-rate variability using long short-term memory neural networks. Sci. Rep. 2019, 9, 14149. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.