Towards Explainable Machine Learning from Remote Sensing to Medical Images—Merging Medical and Environmental Data into Public Health Knowledge Maps
Abstract
1. Introduction
2. Materials and Methods
2.1. Image Processing Workflow
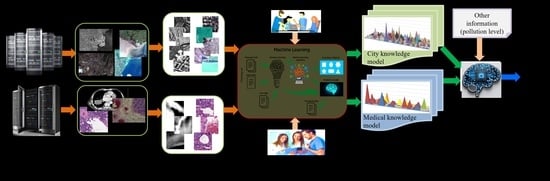
- Step 1: During a preparatory phase, the experts select the dates and target areas acquired by different satellites and download them to typical archives such as the one from the ESA Copernicus hub [17] or the DLR archives [18]. Each EO acquisition from the archive has two parts: the image and the metadata. Later, these two parts are used during operations, as shown in Figure 1. Like EO images, the medical images are acquired by various devices/sensors operated by experts and stored on servers as images and metadata associated with each medical image.
- Step 2: Tile each EO image into patches (with no overlapping) and with a pre-selected size, depending on the actual pixel ground sampling distance to cover the objects on the ground (see Table 2). The tiling is applied in the same way for the medical images, with no overlapping between the patches and with a patch size adapted to the content of the medical image (see Table 3). The size of the patch should be adapted to the image resolution and its content, so that the patch includes (as much as possible) a single object [11,12].
- Step 3: Extract the primitive features that describe the content of each original patch. For the EO images, one applies Gabor filters with 5 scales and 6 orientations [13,14,15], while for the medical images, one applies Weber local descriptors [15] with 8 orientations and 18 excitation levels or multispectral histograms 64 bins. In the case of Gabor filtering, we extract the Gabor coefficients from each image patch and compute the means and standard deviations of each set of coefficients (in total, 5 × 6 × 2 = 60 coefficients). In the case of Weber local descriptors, the features are extracted from each patch in a set of 144 (i.e., 8 × 18) coefficients. A detailed study in the use of various primitive feature extraction methods and different values of their parameters can be found in [5,19].
- Step 4: The classification of the primitive features of each original patch is made automatically, and the patch features are grouped into clusters (i.e., “mathematical groupings”) using a Support Vector Machine (SVM) [16] with relevant feedback. The aim is to obtain a feature-based image patch classification by assigning a single semantic label to each patch using a user-oriented terminology of real-world classes. For the SVM, a chi-squared kernel is selected, and a one-against-all approach is used. The activities of the expert users are called “active learning”, referring to the interactive selection of randomly selected positive and negative examples of target classes based on a proper visualisation of the individual patches, a visual comparison of the selected patches (using for comparison the Google Earth maps in the case of EO images and reference health datasets in the case of medical case), and human expert judgements about the actual patch content.
- Step 5: Generate a set of patches that are semantically correctly labelled. This step is finished once all the given patches have been labelled. However, some patches may remain unlabelled. If the missing labelling represents problem cases, an expert user must identify the most probable class, or the data can be assigned to an unclassified class. There is already a hierarchical semantic labelling scheme in the case of EO images (see [16]). However, regarding the medical images, the experience of expert physicians is significant for defining the semantic labels allocated to the classes.
- Step 6: Interpretation of the produced results. The first data product is the semantic classification results/maps of each image—or, in the case of image time series—the corresponding change maps. The second product is domain ontology representations [5], which help users extract the information and knowledge from the images. Finally, knowledge graphs can be created to explain the entire chain of relations (starting from the data, information extraction up to the semantic classes and their relations).
2.2. Dataset Description
2.2.1. Earth Observation Datasets
2.2.2. Medical Datasets
- (1)
- The first dataset of images is from 8 patients with colorectal adenocarcinoma; the total collected images at different magnifications (5×, 10×, 20×) are 180. They were collected using an optical microscopic device equipped with an RGB camera, being selected by a medical expert based on their expertise to include typical normal structures of the colon wall and pathological structures commonly found in colon adenocarcinoma. Usually, the prototypic colorectal cancer is a well-to-moderately differentiated adenocarcinoma consisting of tubular, anastomosing, and branching glands in a desmoplastic stroma. The surface component may be ulcerated or show papillary or villous architecture. In addition, residual adenoma is often present at the edge of the tumour [30,31].
- (2)
- The second dataset of images is from the patients with lung tumours; there are 11,210 CT images and 25 pathology slices collected from 6 patients. From these, we selected 10 images from 2 patients with lung adenocarcinoma. Usually, lung adenocarcinomas show an admixture of many architectural patterns such as acinar, papillary, micropapillary, lepidic, and solid growth patterns [32,33].
- (3)
- The third dataset of images is extracted from a collection of 52,072 images from 422 patients with non-small cell lung cancer (NSCLC) [34]. For these patients, pre-treatment CT scans lung tumours; manual delineation by a radiation oncologist of the 3D volume of the gross tumour volume and clinical outcome data are available in [31] for the Lung1 dataset. Typically, lung cancer pathology can identify two groups of cancer cells: small cell lung cancer (SCLC) and non-small cell lung cancer (NSCLC). Then, the last ones, the NSCLC, are divided again into squamous cell cancer (SCC), large cell cancer, and lung adenocarcinoma. Finally, in situ (ISA) and invasive are the two types of lung adenocarcinoma.
3. Results
3.1. Semantic Classification Based on the Extracted Information
3.1.1. Earth Observation Images
3.1.2. Medical Images
3.2. Knowledge Representation
4. Discussion
4.1. Machine Learning Systems and Semantic Labelling
4.2. Knowledge Graphs and Interpretability
4.3. From AI for EO to AI for Health: The Case of Lung and Colorectal Cancers
4.4. Limitations and Perspectives of AI Applications
4.5. Limitations of This Study
- Fixation using chemicals such as formalin to stabilise proteins and cellular structures to prevent autolysis and degradation. This process can induce artefacts by coagulating proteins and altering the appearance of blood vessels, causing contractions or stiffening.
- Dehydration of tissue samples with increasing concentrations of alcohol, which can lead to a reduction in the volume of blood vessels and small airways and their collapse, thus altering their microscopic appearance.
- Clearing, in which tissues are passed through clearing solutions (usually xylene or toluene), making them transparent and ready for paraffin infiltration. This can lead to additional alterations, such as vessel collapse and changes in the spatial relationships between structures, including lung airways.
- Paraffin infiltration leading to mechanical artefacts, such as distortion or displacement of tissue structures, including blood vessels. This process can make vessels and airways appear more rigid and collapsed than in their natural state.
- Microtomy or fine cutting of thin sections for microscopy, which can induce mechanical artefacts, such as cracking or distorting blood vessels. Sections may show compressed or deformed vessels and airways due to the pressure of the microtome blade.
- Staining involving various chemicals (e.g., haematoxylin and eosin), which can accentuate or blur certain structural details of blood vessels. Sometimes, dyes can cause precipitates or other colouring artefacts, which can mask or alter the natural appearance of the dishes.
- Fitting the tissue ultrathin sections onto the slides and covering them with another slide can induce mechanical pressure, which could compress or deform the blood vessels and airways.
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AI | artificial intelligence |
| AI4EO | artificial intelligence for Earth observation |
| AI4Health | artificial intelligence for Health applications |
| CRC | colorectal cancer |
| CRCLM | colorectal cancer liver metastasis |
| CT | computed tomography |
| DL | deep learning |
| DLR | Deutsches Zentrum für Luft- und Raumfahrt (German Aerospace Centre) |
| DP | digital pathology |
| EO | Earth observation |
| ESA | European Space Agency |
| HE | haematoxylin–eosin |
| LARC | locally advanced rectal cancer |
| MRI | magnetic resonance imaging |
| SAR | synthetic aperture radar |
| SVM | Support Vector Machine |
References
- Sudmanns, M.; Tiede, D.; Lang, S.; Bergstedt, H.; Trost, G.; Augustin, H.; Baraldi, A.; Blaschke, T. Big Earth Data: Disruptive Changes in Earth Observation Data Management and Analysis? Int. J. Digit. Earth 2020, 13, 832–850. [Google Scholar] [CrossRef]
- Satellite Missions Directory—Earth Observation Missions—eoPortal. Available online: https://eoportal.org/web/eoportal/satellite-missions/ (accessed on 28 April 2022).
- DLR—Earth Observation Center—60 Petabytes for the German Satellite Data Archive D-SDA. Available online: https://www.dlr.de/eoc/en/desktopdefault.aspx/tabid-12632/22039_read-51751 (accessed on 28 April 2022).
- Copernicus Annual Reports, Open Access Hub. Available online: https://dataspace.copernicus.eu/news/2024-11-5-copernicus-data-space-ecosystem-cdse-releases-annual-report-2023 (accessed on 15 October 2025).
- Dumitru, C.O.; Schwarz, G.; Datcu, M. Semantic Labeling of Globally Distributed Urban and Nonurban Satellite Images Using High-Resolution SAR Data. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2021, 14, 6009–6068. [Google Scholar] [CrossRef]
- Mallappallil, M.; Sabu, J.; Gruessner, A.; Salifu, M. A Review of Big Data and Medical Research. SAGE Open Med. 2020, 8, 2050312120934839. [Google Scholar] [CrossRef] [PubMed]
- Sfikas, G. List of Medical Imaging Datasets. Github Repository. Available online: https://github.com/sfikas/medical-imaging-datasets (accessed on 28 April 2022).
- The Cancer Imaging Archive (TCIA). Available online: https://www.cancerimagingarchive.net/ (accessed on 28 April 2022).
- High Quality Pathology Images—WebPathology. Available online: http://webpathology.com (accessed on 28 April 2022).
- Dumitru, C.O.; Cui, S.; Faur, D.; Datcu, M. Data Analytics for Rapid Mapping: Case Study of a Flooding Event in Germany and the Tsunami in Japan Using Very High Resolution SAR Images. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2015, 8, 114–129. [Google Scholar] [CrossRef]
- Dumitru, C.O.; Schwarz, G.; Datcu, M. AL4SLEO: An Active Learning Solution for the Semantic Labelling of Earth Observation Satellite Images—Part 1. In Benchmarks and Hybrid Algorithms in Optimization and Applications; Yang, X.-S., Ed.; Springer Tracts in Nature-Inspired Computing; Springer Nature: Singapore, 2023; pp. 105–118. ISBN 978-981-99-3970-1. [Google Scholar]
- Dumitru, C.O.; Schwarz, G.; Datcu, M. AL4SLEO: An Active Learning Solution for the Semantic Labelling of Earth Observation Satellite Images—Part 2. In Benchmarks and Hybrid Algorithms in Optimization and Applications; Yang, X.-S., Ed.; Springer Tracts in Nature-Inspired Computing; Springer Nature: Singapore, 2023; pp. 119–146. ISBN 978-981-99-3970-1. [Google Scholar]
- MPEG-7 Standard. Available online: https://mpeg.chiariglione.org/standards/mpeg-7.html (accessed on 15 October 2025).
- Manjunath, B.S.; Ma, W.Y. Texture Features for Browsing and Retrieval of Image Data. IEEE Trans. Pattern Anal. Mach. Intell. 1996, 18, 837–842. [Google Scholar] [CrossRef]
- Chen, J.; Shan, S.; He, C.; Zhao, G.; Pietikäinen, M.; Chen, X.; Gao, W. WLD: A Robust Local Image Descriptor. IEEE Trans. Pattern Anal. Mach. Intell. 2010, 32, 1705–1720. [Google Scholar] [CrossRef]
- Cui, S.; Dumitru, C.O.; Datcu, M. Semantic Annotation in Earth Observation Based on Active Learning. Int. J. Image Data Fusion. 2014, 5, 152–174. [Google Scholar] [CrossRef]
- Sentinels Scientific Data Hub. Available online: https://scihub.copernicus.eu/dhus/#/home (accessed on 28 April 2022).
- EOWEB GeoPortal—TerraSAR-X Archive Portal. Available online: https://eoweb.dlr.de/egp/ (accessed on 28 April 2022).
- Dumitru, C.O.; Datcu, M. Information Content of Very High Resolution SAR Images: Study of Feature Extraction and Imaging Parameters. IEEE Trans. Geosci. Remote Sens. 2013, 51, 4591–4610. [Google Scholar] [CrossRef]
- The European Space Agency. Earth Observation Satellite Missions. Available online: https://earth.esa.int/eogateway/search?skipDetection=true&text=&category=Missions (accessed on 28 April 2022).
- Gaofen-3 Sensor Parameter Description. Available online: https://directory.eoportal.org/web/eoportal/satellite-missions/g/gaofen-3 (accessed on 28 April 2022).
- Canadian Space Agency. RADARSAT Satellites: Technical Comparison. Available online: https://www.asc-csa.gc.ca/eng/satellites/radarsat/technical-features/radarsat-comparison.asp (accessed on 28 April 2022).
- Sentinel-1—Missions. Available online: https://sentiwiki.copernicus.eu/web/s1-mission (accessed on 15 October 2025).
- Sentinel-2—Missions. Available online: https://sentinels.copernicus.eu/copernicus/sentinel-2 (accessed on 15 October 2025).
- TerraSAR-X Sensor Parameter Description and Data Access. Available online: https://sss.terrasar-x.dlr.de/ (accessed on 28 April 2022).
- WorldView Sensor Parameter Description. Available online: https://www.satimagingcorp.com/satellite-sensors/worldview-2/ (accessed on 28 April 2022).
- CANDELA (Copernicus Access Platform Intermediate Layers Small Scale Demonstrator) Project. Available online: http://www.candela-h2020.eu/ (accessed on 28 April 2022).
- Dumitru, C.O.; Schwarz, G.; Pulak-Siwiec, A.; Kulawik, B.; Albughdadi, M.; Lorenzo, J.; Datcu, M. Understanding Satellite Images: A Data Mining Module for Sentinel Images. Big Earth Data 2020, 4, 367–408. [Google Scholar] [CrossRef]
- Murphy, A. Radiological Medical Image Datasets Build for Artificial Intelligence, Radiopaedia.Org. Available online: https://radiopaedia.org/articles/imaging-data-sets-artificial-intelligence (accessed on 28 April 2022).
- WebPathology: Case Colo-Rectal Adenocarcinoma. Available online: https://www.webpathology.com/search-result?query=colorectal%20adenocarcinoma (accessed on 27 April 2022).
- Aerts, H.J.W.L.; Velazquez, E.R.; Leijenaar, R.T.H.; Parmar, C.; Grossmann, P.; Carvalho, S.; Bussink, J.; Monshouwer, R.; Haibe-Kains, B.; Rietveld, D.; et al. Decoding Tumour Phenotype by Noninvasive Imaging Using a Quantitative Radiomics Approach. Nat. Commun. 2014, 5, 4006. [Google Scholar] [CrossRef]
- WebPathology: Case Lung Adenocarcinoma. Available online: https://www.webpathology.com/case.asp?case=416 (accessed on 28 April 2022).
- Rusu, M.; Rajiah, P.; Gilkeson, R.; Yang, M.; Donatelli, C.; Thawani, R.; Jacono, F.J.; Linden, P.; Madabhushi, A. Co-Registration of Pre-Operative CT with Ex Vivo Surgically Excised Ground Glass Nodules to Define Spatial Extent of Invasive Adenocarcinoma on in Vivo Imaging: A Proof-of-Concept Study. Eur. Radiol. 2017, 27, 4209–4217. [Google Scholar] [CrossRef]
- Aerts, H.J.W.L.; Wee, L.; Rios Velazquez, E.; Leijenaar, R.T.H.; Parmar, C.; Grossmann, P.; Carvalho, S.; Bussink, J.; Monshouwer, R.; Haibe-Kains, B.; et al. Data from NSCLC-Radiomics. 2019. Available online: https://www.cancerimagingarchive.net/collection/nsclc-radiomics/ (accessed on 27 April 2022).
- Colapicchioni, A. KES: Knowledge Enabled Services for Better EO Information Use. In Proceedings of the IEEE International Geoscience and Remote Sensing Symposium (IGARSS’04), Anchorage, AK, USA, 20–24 September 2004; IEEE: Anchorage, AK, USA, 2004; Volume 1, pp. 176–179. [Google Scholar]
- Dumitru, C.O.; Schwarz, G.; Datcu, M. Machine Learning Techniques for Knowledge Extraction from Satellite Images: Application to Specific Area Types. Int. Arch. Photogramm. Remote Sens. Spat. Inf. Sci. 2021, XLIII-B3-2021, 455–462. [Google Scholar] [CrossRef]
- Earth from Space: Bucharest, Romania. 2021. Available online: https://www.esa.int/ESA_Multimedia/Videos/2021/04/Earth_from_Space_Bucharest (accessed on 27 April 2022).
- Dumitru, C.O.; Schwarz, G.; Cui, S.; Datcu, M. Improved Image Classification by Proper Patch Size Selection: TerraSAR-X vs. Sentinel-1A. In Proceedings of the 2016 International Conference on Systems, Signals and Image Processing (IWSSIP), Bratislava, Slovakia, 23–25 May 2016; IEEE: New York, NY, USA; pp. 1–4. [Google Scholar]
- TerraSAR-X and TanDEM-X—Earth Online. Available online: https://earth.esa.int/eogateway/missions/terrasar-x-and-tandem-x (accessed on 15 October 2025).
- Sentinel-1—Sentinel Online. Available online: https://sentinels.copernicus.eu/copernicus/sentinel-1 (accessed on 15 October 2025).
- ESA—Sentinel-2. Available online: https://www.esa.int/Applications/Observing_the_Earth/Copernicus/Sentinel-2 (accessed on 15 October 2025).
- Karmakar, C.; Dumitru, C.O.; Schwarz, G.; Datcu, M. Feature-Free Explainable Data Mining in SAR Images Using Latent Dirichlet Allocation. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2021, 14, 676–689. [Google Scholar] [CrossRef]
- Russell, B.C.; Torralba, A.; Murphy, K.P.; Freeman, W.T. LabelMe: A Database and Web-Based Tool for Image Annotation. Int. J. Comput. Vis. 2008, 77, 157–173. [Google Scholar] [CrossRef]
- Smeulders, A.W.M.; Worring, M.; Santini, S.; Gupta, A.; Jain, R. Content-Based Image Retrieval at the End of the Early Years. IEEE Trans. Pattern Anal. Mach. Intell. 2000, 22, 1349–1380. [Google Scholar] [CrossRef]
- Shyu, C.-R.; Klaric, M.; Scott, G.J.; Barb, A.S.; Davis, C.H.; Palaniappan, K. GeoIRIS: Geospatial Information Retrieval and Indexing System—Content Mining, Semantics Modeling, and Complex Queries. IEEE Trans. Geosci. Remote Sens. 2007, 45, 839–852. [Google Scholar] [CrossRef] [PubMed]
- Witten, I.H.; Frank, E.; Hall, M.A. Data Mining: Practical Machine Learning Tools and Techniques, 3rd ed.; Morgan Kaufmann Series in Data Management Systems; Morgan Kaufmann: Burlington, MA, USA, 2011; ISBN 978-0-12-374856-0. [Google Scholar]
- Wang, S.; Cao, J.; Yu, P.S. Deep Learning for Spatio-Temporal Data Mining: A Survey. arXiv 2019, arXiv:1906.04928. [Google Scholar] [CrossRef]
- Huang, Z.; Dumitru, C.O.; Ren, J. Physics-Aware Feature Learning of Sar Images with Deep Neural Networks: A Case Study. In Proceedings of the 2021 IEEE International Geoscience and Remote Sensing Symposium IGARSS, Brussels, Belgium, 11–16 July 2021; pp. 1264–1267. [Google Scholar]
- Stepinski, T.F.; Netzel, P.; Jasiewicz, J. LandEx—A GeoWeb Tool for Query and Retrieval of Spatial Patterns in Land Cover Datasets. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2014, 7, 257–266. [Google Scholar] [CrossRef]
- Espinoza-Molina, D.; Manilici, V.; Dumitru, C.; Reck, C.; Cui, S.; Rotzoll, H.; Hofmann, M.; Schwarz, G.; Datcu, M. The Earth Observation Image Librarian (EOLIB): The Data Mining Component of the TerraSAR-X Payload Ground Segment. In Proceedings of the Big Data from Space (BiDS’16), Santa Cruz de Tenerife, Spain, 15–17 March 2016; Soille, P., Marchetti, P.G., Eds.; European Union: Brussels, Belgium, 2016; pp. 228–231. [Google Scholar]
- Zhou, J.; Cao, R.; Kang, J.; Guo, K.; Xu, Y. An Efficient High-Quality Medical Lesion Image Data Labeling Method Based on Active Learning. IEEE Access 2020, 8, 144331–144342. [Google Scholar] [CrossRef]
- Esteva, A.; Chou, K.; Yeung, S.; Naik, N.; Madani, A.; Mottaghi, A.; Liu, Y.; Topol, E.; Dean, J.; Socher, R. Deep Learning-Enabled Medical Computer Vision. NPJ Digit. Med. 2021, 4, 5. [Google Scholar] [CrossRef]
- Baumgartner, C. Herausforderungen und Chancen von KI in der Medizinischen Bildgebung. Exzellenzcluster “Maschinelles Lernen: Neue Perspektiven für die Wissenschaft“, Universität Tübingen. Available online: https://www.mlmia-unitue.de/uploads/lndw-final.pdf (accessed on 28 April 2022).
- Dumitru, C.O.; Schwarz, G.; Datcu, M. Land Cover Semantic Annotation Derived from High-Resolution SAR Images. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2016, 9, 2215–2232. [Google Scholar] [CrossRef]
- Cook, D.J.; Holder, L.B. (Eds.) Mining Graph Data; Wiley-Interscience: Hoboken, NJ, USA, 2007; ISBN 978-0-471-73190-0. [Google Scholar]
- Samatova, N.F. (Ed.) Practical Graph Mining with R; Chapman & Hall/CRC Data Mining and Knowledge Discovery Series; Taylor & Francis: Boca Raton, FL, USA, 2014; ISBN 978-1-4398-6084-7. [Google Scholar]
- Vieira, A. Knowledge Representation in Graphs Using Convolutional Neural Networks. arXiv 2016, arXiv:1612.02255. [Google Scholar] [CrossRef]
- Sullivan, D. Google Launches Knowledge Graph to Provide Answers, Not Just Links. Available online: https://www.techmeme.com/120516/p37#a120516p37 (accessed on 28 April 2022).
- Dumitru, C.O.; Schwarz, G.; Datcu, M. Image Representation Alternatives for the Analysis of Satellite Image Time Series. In Proceedings of the 2017 9th International Workshop on the Analysis of Multitemporal Remote Sensing Images (MultiTemp), Brugge, Belgium, 27–29 June 2017; pp. 1–4. [Google Scholar]
- Karalis, N.; Mandilaras, G.; Koubarakis, M. Extending the YAGO2 Knowledge Graph with Precise Geospatial Knowledge. In The Semantic Web—ISWC 2019; Ghidini, C., Hartig, O., Maleshkova, M., Svátek, V., Cruz, I., Hogan, A., Song, J., Lefrançois, M., Gandon, F., Eds.; Lecture Notes in Computer Science; Springer International Publishing: Cham, Switzerland, 2019; Volume 11779, pp. 181–197. ISBN 978-3-030-30795-0. [Google Scholar]
- Ji, S.; Pan, S.; Cambria, E.; Marttinen, P.; Yu, P.S. A Survey on Knowledge Graphs: Representation, Acquisition and Applications. IEEE Trans. Neural Netw. Learn. Syst. 2022, 33, 494–514. [Google Scholar] [CrossRef]
- Réjichi, S.; Chaabane, F.; Tupin, F. Expert Knowledge-Based Method for Satellite Image Time Series Analysis and Interpretation. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2015, 8, 2138–2150. [Google Scholar] [CrossRef]
- Rotmensch, M.; Halpern, Y.; Tlimat, A.; Horng, S.; Sontag, D. Learning a Health Knowledge Graph from Electronic Medical Records. Sci. Rep. 2017, 7, 5994. [Google Scholar] [CrossRef] [PubMed]
- Roscher, R.; Bohn, B.; Duarte, M.F.; Garcke, J. Explainable Machine Learning for Scientific Insights and Discoveries. IEEE Access 2020, 8, 42200–42216. [Google Scholar] [CrossRef]
- Li, X.-H.; Cao, C.C.; Shi, Y.; Bai, W.; Gao, H.; Qiu, L.; Wang, C.; Gao, Y.; Zhang, S.; Xue, X.; et al. A Survey of Data-Driven and Knowledge-Aware eXplainable AI. IEEE Trans. Knowl. Data Eng. 2022, 34, 29–49. [Google Scholar] [CrossRef]
- Amann, J.; Blasimme, A.; Vayena, E.; Frey, D.; Madai, V.I. The Precise4Q consortium Explainability for Artificial Intelligence in Healthcare: A Multidisciplinary Perspective. BMC Med. Inform. Decis. Mak. 2020, 20, 310. [Google Scholar] [CrossRef]
- TELEIOS Project, Deliverable D6.4 “The Virtual Observatory for TerraSAR-X Data and Applications—Phase II”. Available online: https://cordis.europa.eu/project/id/257662 (accessed on 28 April 2022).
- Tharwat, A.; Schenck, W. A Survey on Active Learning: State-of-the-Art, Practical Challenges and Research Directions. Mathematics 2023, 11, 820. [Google Scholar] [CrossRef]
- Mosqueira-Rey, E.; Hernández-Pereira, E.; Alonso-Ríos, D.; Bobes-Bascarán, J.; Fernández-Leal, Á. Human-in-the-Loop Machine Learning: A State of the Art. Artif. Intell. Rev. 2023, 56, 3005–3054. [Google Scholar] [CrossRef]
- Lenczner, G.; Chan-Hon-Tong, A.; Le Saux, B.; Luminari, N.; Le Besnerais, G. DIAL: Deep Interactive and Active Learning for Semantic Segmentation in Remote Sensing. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2022, 15, 3376–3389. [Google Scholar] [CrossRef]
- Cong, L.; Feng, W.; Yao, Z.; Zhou, X.; Xiao, W. Deep Learning Model as a New Trend in Computer-Aided Diagnosis of Tumor Pathology for Lung Cancer. J. Cancer 2020, 11, 3615–3622. [Google Scholar] [CrossRef]
- Sakamoto, T.; Furukawa, T.; Lami, K.; Pham, H.H.N.; Uegami, W.; Kuroda, K.; Kawai, M.; Sakanashi, H.; Cooper, L.A.D.; Bychkov, A.; et al. A Narrative Review of Digital Pathology and Artificial Intelligence: Focusing on Lung Cancer. Transl. Lung Cancer Res. 2020, 9, 2255–2276. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, V.S.; Toro, P.; Corredor, G.; Mukhopadhyay, S.; Madabhushi, A. The State of the Art for Artificial Intelligence in Lung Digital Pathology. J. Pathol. 2022, 257, 413–429. [Google Scholar] [CrossRef] [PubMed]
- Cai, Y.-W.; Dong, F.-F.; Shi, Y.-H.; Lu, L.-Y.; Chen, C.; Lin, P.; Xue, Y.-S.; Chen, J.-H.; Chen, S.-Y.; Luo, X.-B. Deep Learning Driven Colorectal Lesion Detection in Gastrointestinal Endoscopic and Pathological Imaging. World J. Clin. Cases 2021, 9, 9376–9385. [Google Scholar] [CrossRef]
- Chassagnon, G.; De Margerie-Mellon, C.; Vakalopoulou, M.; Marini, R.; Hoang-Thi, T.-N.; Revel, M.-P.; Soyer, P. Artificial Intelligence in Lung Cancer: Current Applications and Perspectives. Jpn. J. Radiol. 2023, 41, 235–244. [Google Scholar] [CrossRef]
- Lee, J.; Hwang, E.; Kim, H.; Park, C. A Narrative Review of Deep Learning Applications in Lung Cancer Research: From Screening to Prognostication. Transl. Lung Cancer Res. 2022, 11, 1217–1229. [Google Scholar] [CrossRef]
- Astley, J.R.; Wild, J.M.; Tahir, B.A. Deep Learning in Structural and Functional Lung Image Analysis. Br. J. Radiol. 2022, 95, 20201107. [Google Scholar] [CrossRef]
- Sourlos, N.; Wang, J.; Nagaraj, Y.; van Ooijen, P.; Vliegenthart, R. Possible Bias in Supervised Deep Learning Algorithms for CT Lung Nodule Detection and Classification. Cancers 2022, 14, 3867. [Google Scholar] [CrossRef]
- Hou, M.; Sun, J.-H. Emerging Applications of Radiomics in Rectal Cancer: State of the Art and Future Perspectives. World J. Gastroenterol. 2021, 27, 3802–3814. [Google Scholar] [CrossRef]
- Joseph, J.; LePage, E.M.; Cheney, C.P.; Pawa, R. Artificial Intelligence in Colonoscopy. World J. Gastroenterol. 2021, 27, 4802–4817. [Google Scholar] [CrossRef]
- Stanzione, A.; Verde, F.; Romeo, V.; Boccadifuoco, F.; Mainenti, P.P.; Maurea, S. Radiomics and Machine Learning Applications in Rectal Cancer: Current Update and Future Perspectives. World J. Gastroenterol. 2021, 27, 5306–5321. [Google Scholar] [CrossRef]
- Liang, F.; Wang, S.; Zhang, K.; Liu, T.-J.; Li, J.-N. Development of Artificial Intelligence Technology in Diagnosis, Treatment, and Prognosis of Colorectal Cancer. World J. Gastrointest. Oncol. 2022, 14, 124–152. [Google Scholar] [CrossRef] [PubMed]
- Davri, A.; Birbas, E.; Kanavos, T.; Ntritsos, G.; Giannakeas, N.; Tzallas, A.T.; Batistatou, A. Deep Learning on Histopathological Images for Colorectal Cancer Diagnosis: A Systematic Review. Diagnostics 2022, 12, 837. [Google Scholar] [CrossRef]
- Thakur, N.; Yoon, H.; Chong, Y. Current Trends of Artificial Intelligence for Colorectal Cancer Pathology Image Analysis: A Systematic Review. Cancers 2020, 12, 1884. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Yu, M.; Chen, D.; Li, P.; Tang, B.; Li, J. Role of MRI-based Radiomics in Locally Advanced Rectal Cancer (Review). Oncol. Rep. 2021, 47, 34. [Google Scholar] [CrossRef]
- Alshohoumi, F.; Al-Hamdani, A.; Hedjam, R.; AlAbdulsalam, A.; Al Zaabi, A. A Review of Radiomics in Predicting Therapeutic Response in Colorectal Liver Metastases: From Traditional to Artificial Intelligence Techniques. Healthcare 2022, 10, 2075. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Deng, C.; Wu, B. Magnetic Resonance Imaging-Based Artificial Intelligence Model in Rectal Cancer. World J. Gastroenterol. 2021, 27, 2122–2130. [Google Scholar] [CrossRef]
- Bedrikovetski, S.; Dudi-Venkata, N.; Kroon, H.; Seow, W.; Vather, R.; Carneiro, G.; Moore, J.; Sammour, T. Artificial Intelligence for Pre-Operative Lymph Node Staging in Colorectal Cancer: A Systematic Review and Meta-Analysis. BMC Cancer 2021, 21. [Google Scholar] [CrossRef]
- Viganò, L.; Jayakody Arachchige, V.S.; Fiz, F. Is Precision Medicine for Colorectal Liver Metastases Still a Utopia? New Perspectives by Modern Biomarkers, Radiomics, and Artificial Intelligence. World J. Gastroenterol. 2022, 28, 608–623. [Google Scholar] [CrossRef]
























| Parameter | EO Images | Medical Images |
|---|---|---|
| Kernel type | Chi-squared kernel | Chi-squared kernel |
| Multi-class strategy | One-against-all | One-against-all |
| Regularisation parameter (C) | 1.0 (default) | 1.0 (default) |
| Gamma (kernel coefficient) | 1/(number of features) | 1/(number of features) |
| Cross-validation strategy | 5-fold cross-validation | 5-fold cross-validation |
| Active learning cycles | 3–5 iterations depending on dataset size | 3–5 iterations depending on dataset size |
| Location | Instrument | Type | Mode | No. of Sensor Bands/Selected Bands | Resolution | Polarisation | Patch Size (Pixels) | No. of Patches |
|---|---|---|---|---|---|---|---|---|
| Berlin, Germany | Gaofen-3 | C-band SAR | SpotLight (SL) | 1/1 | 1 m | HH | 256 × 256 | 2080 |
| TerraSAR-X | X-band SAR | Multi-look Ground range Detected (MGD) | 1/1 | 2.9 m | HH | 160 × 160 | 1025 | |
| Bucharest, Romania | TerraSAR-X | X-band SAR | Multi-look Ground range Detected (MGD) | 1/1 | 2.9 m | HH | 160 × 160 | 4455 |
| WorldView-2 | Multi-spectral | - | 8/3 (RGB) | 1.87 m | - | 100 × 100 | 33,930 | |
| Vancouver, Canada | RADARSAT-2 | C-band SAR | Extended high (EH) | 1/1 | 13.5 m | HH | 160 × 160 | 660 |
| TerraSAR-X | X-band SAR | Multi-look Ground range Detected (MGD) | 1/1 | 2.9 m | HH | 160 × 160 | 825 | |
| Albania and Greece | TerraSAR-X | X-band SAR | Multi-look Ground range Detected (MGD) | 1/1 | 2.9 m | HH | 160 × 160 | 1872 |
| Sentinel-1 | C-band SAR | Interferometric Wide swath (IW) | 1/1 | 20 m | VV/VH | 128 × 128 | 26,260 | |
| Sentinel-2 | Multi-spectral | - | 13/3 (RGB) | 10/20/60 m | - | 120 × 120 | 8281 |
| Data | Instrument | No. of Bands/Selected Bands | Image Dimension (Pixels) | Sub-Images (Pixels) | Patch Size (Pixels) | No. of Patches |
|---|---|---|---|---|---|---|
| Colorectal adenocarcinoma | Optical microscopy | 3/3 | 1024 × 768 | - | 4 × 4 | 49,152 |
| Lung adenocarcinoma | Optical microscopy | 3/3 | avg. 9688 × 9832 | 890 × 801 | 4 × 4 | 44,400 |
| Non-small cell lung cancer | Computer tomography (CT) scan | 1/1 | avg. 1802 × 884 | 1372 × 672 | 4 × 4 | 57,624 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bilteanu, L.; Dumitru, C.O.; Dumachi, A.; Alexandrescu, F.; Popa, R.; Buiu, O.; Serban, A.I. Towards Explainable Machine Learning from Remote Sensing to Medical Images—Merging Medical and Environmental Data into Public Health Knowledge Maps. Mach. Learn. Knowl. Extr. 2025, 7, 140. https://doi.org/10.3390/make7040140
Bilteanu L, Dumitru CO, Dumachi A, Alexandrescu F, Popa R, Buiu O, Serban AI. Towards Explainable Machine Learning from Remote Sensing to Medical Images—Merging Medical and Environmental Data into Public Health Knowledge Maps. Machine Learning and Knowledge Extraction. 2025; 7(4):140. https://doi.org/10.3390/make7040140
Chicago/Turabian StyleBilteanu, Liviu, Corneliu Octavian Dumitru, Andreea Dumachi, Florin Alexandrescu, Radu Popa, Octavian Buiu, and Andreea Iren Serban. 2025. "Towards Explainable Machine Learning from Remote Sensing to Medical Images—Merging Medical and Environmental Data into Public Health Knowledge Maps" Machine Learning and Knowledge Extraction 7, no. 4: 140. https://doi.org/10.3390/make7040140
APA StyleBilteanu, L., Dumitru, C. O., Dumachi, A., Alexandrescu, F., Popa, R., Buiu, O., & Serban, A. I. (2025). Towards Explainable Machine Learning from Remote Sensing to Medical Images—Merging Medical and Environmental Data into Public Health Knowledge Maps. Machine Learning and Knowledge Extraction, 7(4), 140. https://doi.org/10.3390/make7040140