1. Introduction
Clustering is a foundational unsupervised learning task that aims to partition data points into distinct groups based on similarity. Among existing approaches, spectral clustering has gained significant attention due to its ability to identify non-convex clusters and its solid theoretical foundation in spectral graph theory [
1,
2]. The core of this technique involves the eigendecomposition of a graph Laplacian matrix to embed data points into a low-dimensional eigenspace where clusters become more separable. For our work, we chose to employ the normalized graph Laplacian
, where
W is the similarity matrix and
D is the degree matrix [
2,
3]. This choice is motivated by its well-established theoretical properties and its tendency to yield more stable and consistent clustering results across varied data densities compared to the unnormalized Laplacian.
However, the practical application of spectral clustering to large-scale datasets is severely hampered by its substantial computational complexity. The primary bottlenecks are twofold:
Similarity Matrix Construction: Building the dense pairwise similarity matrix W has a prohibitive time and space complexity.
Eigendecomposition: Solving the eigenproblem for the Laplacian matrix scales as . While approximate methods can reduce this, the quadratic scaling of constructing W remains the dominant constraint.
To mitigate these challenges, various approximation schemes have been proposed, including sparse
k-NN graphs and Nyström methods [
4,
5]. A highly promising direction involves leveraging advanced data structures for efficient computation. Notably, Cover trees [
6] offer a principled framework for accelerating nearest neighbor queries with a query time complexity of
in metric spaces, where
c is the expansion constant, a significant improvement over a naive
linear scan. Beyond mere acceleration, the hierarchical multi-scale structure of Cover trees provides a natural mechanism for intelligent data summarization and noise reduction, making them particularly suitable for enhancing clustering algorithms [
7,
8].
In this work, we propose the Improved Spectral Clustering with Cover tree (ISCT) algorithm, a novel framework designed to overcome the scalability limitations of traditional spectral clustering. Our approach leverages the Cover tree for a dual purpose within a two-stage approximation framework:
Data Reduction via Cover tree-Based Summarization: We exploit the hierarchical structure of the Cover tree to select a small set of m representative points (where ) that preserve the essential geometric and density structure of the original dataset. This step drastically reduces the size of the problem.
Efficient Spectral Clustering on Representatives: We perform spectral clustering using the normalized Laplacian on the resulting manageable similarity matrix constructed from the representatives. The cluster labels for all n original points are then accurately inferred by assigning each point to the cluster of its nearest representative, a step executed efficiently in time using the Cover tree itself for fast nearest-neighbor queries.
The key insight of our approach is this dual use of the Cover tree, which provides not only a fast query structure but also an intelligent, multi-scale data summary that is inherently aligned with the clustering objective. This shifts the computational bottleneck from to . Crucially, empirical observations and theoretical bounds suggest that the number of representatives m required for a faithful approximation can favorably scale from to for real-world datasets, promising orders-of-magnitude improvement in computational efficiency.
This work is guided by the following research questions:
How does Cover tree-based data summarization affect the quality of the spectral embedding and the resulting clusters compared to traditional methods, particularly when using the normalized Laplacian?
What is the computational accuracy trade-off inherent in the ISCT framework, and how is it influenced by the number of representatives m?
How does the multi-scale hierarchy of the Cover tree influence the detection of cluster boundaries and the separation of clusters at different density levels?
The ISCT framework combines the solid theoretical foundations of spectral clustering, embodied by the normalized Laplacian, with the effective hierarchical properties of cover trees to create a scalable and efficient solution for large-scale clustering challenges.
2. Related Work
The field of cluster analysis has undergone significant evolution, representing one of the earliest developments in data analysis. Its conceptual foundations were established by Driver and Kroeber in 1932 [
9], who introduced the fundamental notion of partitioning observations into groups, notably preceding the formal emergence of artificial intelligence by decades. This early work, along with crucial theoretical contributions by Zubin (1938) and Tryon (1939) [
10,
11] on cluster validity, established the mathematical framework for modern clustering research. The transition to practical algorithmic implementations occurred in the 1960s–1970s with MacQueen’s introduction of the K-Means algorithm, representing a watershed moment that provided the first computationally tractable partition-based approach. Concurrently, hierarchical clustering methods gained prominence through the work of Johnson (1967) [
12], while the 1970s witnessed the emergence of probabilistic mixture models. A fundamental paradigm shift occurred with the development of spectral clustering methods, grounded in Fiedler’s work on algebraic connectivity and subsequently refined by Shi and Malik (2000) [
3] through normalized cuts. The advent of big data in the 2000s exposed the scalability limitations of traditional methods, particularly the
eigendecomposition bottleneck in spectral clustering, sparking intensive research into approximate algorithms. This period saw the emergence of Cover trees, providing a rigorous framework for efficient nearest neighbor search. Contemporary developments have focused on addressing the dual challenges of scalability and quality preservation, including reduced clustering methods based on the inversion formula density estimation by Lukauskas et al. (2023) [
13] and novel approaches for analyzing research community dynamics by Cambe et al. (2022) [
14].
The development of the clustering methodology provides essential context for our proposed algorithm. We focus on the evolution of spectral clustering and related scalability solutions. The theoretical underpinnings of spectral clustering are deeply rooted in spectral graph theory [
15]. The algorithm’s effectiveness for non-convex clusters stems from its relaxation of graph partitioning problems, such as normalized cut [
3]. The standard algorithm involves constructing a similarity graph, forming a Laplacian matrix (typically
or
), computing the first
k eigenvectors of the Laplacian and applying a standard clustering algorithm like
k-means to the rows of the eigenvector matrix to obtain the final partition. The phrase “simplified bisection via the eigenvector” often refers to the special case of
, where the second eigenvector (the Fiedler vector) is thresholded to partition the graph [
16]. For
, the process is a generalization of this bisection concept [
13].
The high computational cost of spectral clustering has spurred research into approximations. A common approach is to sparsify the similarity matrix using
k-NN or
-neighborhood graphs, reducing the storage to
and enabling the use of sparse eigensolvers like ARPACK [
17]. The Nyström method is another prominent technique, which approximates the eigendecomposition by using a subset of landmark points [
5]. The complexity is reduced to
for
l landmarks. Our method differs by using the Cover tree not just to select landmarks but to ensure they provide a geometrically meaningful summary of the entire dataset [
14].
Data structures like KD-Trees, Ball Trees, and Cover trees have been widely used to accelerate distance-based computations. Their integration into spectral clustering has primarily been limited to constructing approximate
k-NN graphs [
18]. The ISCT algorithm extends this integration beyond a mere neighbor search to a holistic data reduction and assignment strategy, leveraging the Cover tree’s theoretical guarantees for both tasks.
Our proposed ISCT algorithm represents a natural evolution of this historical trajectory, integrating Cover tree hierarchical decomposition with spectral techniques to bridge the gap between classical graph-theoretic approaches and modern scalable computing requirements, and building upon this rich methodological heritage to address specific computational bottlenecks that limit spectral methods’ applicability to large-scale datasets.
6. Results
To compare our algorithm with others, the real datasets chosen were Iris, Seeds, Glass, Mall, Cancer, as well as synthetic data such as Blobs, Circles and Moons, which are represented in the
Table 1 and the other real datasets are represented in
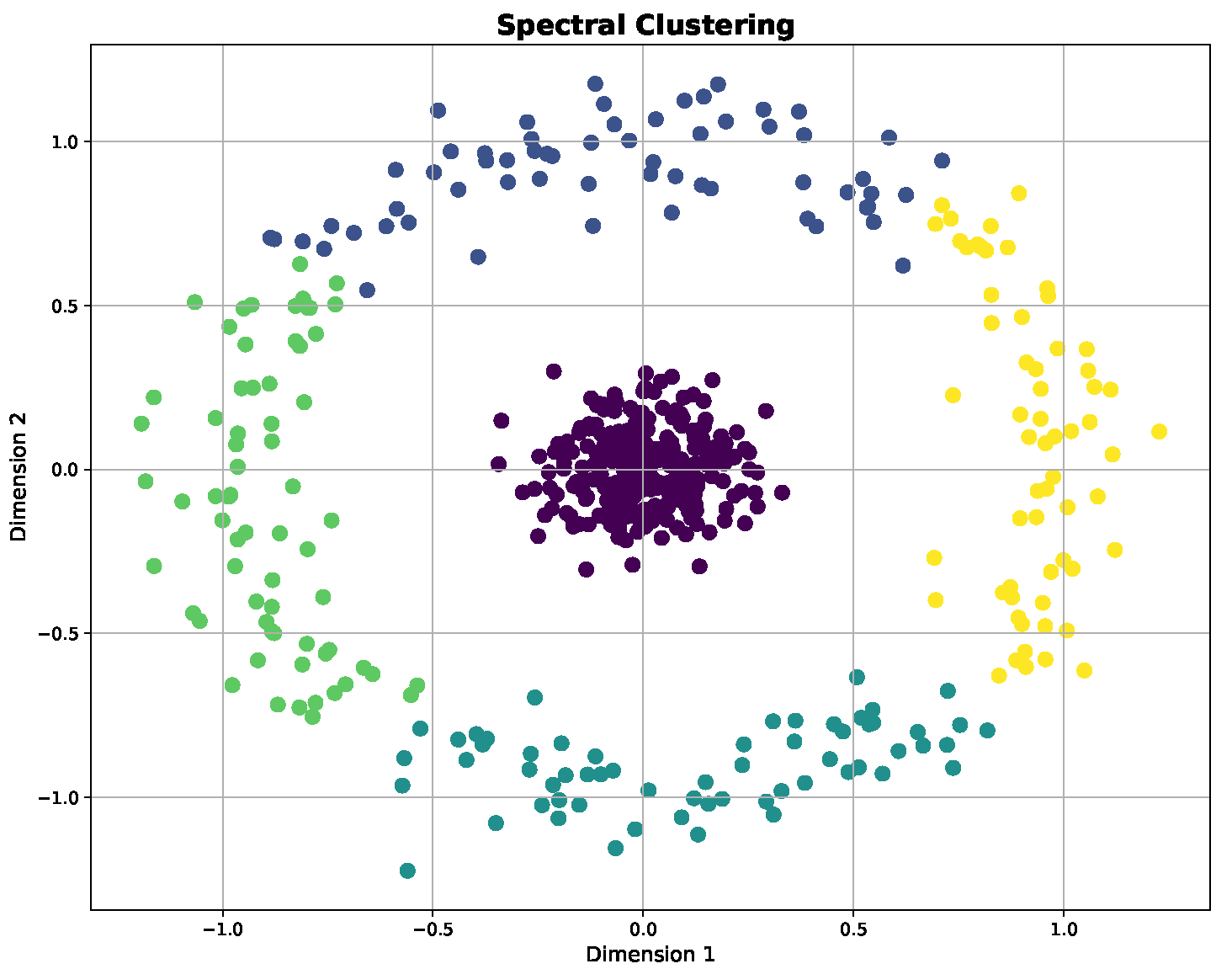
Table 2, as they are frequently used for testing and evaluating machine learning algorithms; the use of synthetic datasets such as Three Moons, Blobs and Circles is a particularly common practice for evaluating and visualizing the performance of clustering algorithms. These datasets are specifically designed to test the ability of clustering methods to identify and separate distinct groups in data of varying complexity and structures. Three Moons, with its three intertwined crescent shapes, represents a challenge for algorithms due to its non-convex shape. Blobs contain well-separated Gaussian clusters to evaluate clustering on spherical groups. Circles feature points arranged in concentric circles to test the handling of nested and non-linearly separable clusters.
We evaluated the clustering algorithms on three synthetic benchmark datasets: two interleaving half-moons (Moons) and concentric circles (Circles), and a Gaussian Blobs dataset. The Moons and Circles datasets each contained 500 points with two ground-truth classes, while the Blobs dataset comprised 500 points distributed across five Gaussian clusters. To ensure reproducibility, we generated the data using Scikit-learn’s dataset generators with fixed parameters: Moons with n_samples = 500, noise = 0.1, and random_state = 40; Circles with n_samples = 500, noise = 0.1, factor = 0.1, and random_state = 40; and Blobs with n_samples = 500, centers = 5, cluster_std = 3.0, and random_state = 40.
All clustering algorithms were configured to identify clusters. This setup intentionally creates a mismatch between the true number of classes 2 and the number of predicted clusters 5 for the Moons and Circles datasets. To address this discrepancy, we employed a many-to-one relabeling strategy (also known as cluster purity mapping). Specifically, each predicted cluster was assigned to the ground-truth class that represents the majority of its points. The mapping was established by finding the mode of the true labels within each cluster using SciPy’s mode function. This alignment allowed us to compute standard supervised evaluation metrics, including Adjusted Rand Index (ARI), Normalized Mutual Information (NMI), and classification-style metrics (Precision, Recall, F1-score). Additionally, we reported internal clustering quality measures such as the Silhouette score. A many-to-one confusion matrix was also constructed to visualize the distribution of true classes across predicted clusters.
The three figures
Figure 1,
Figure 2 and
Figure 3 illustrate the dataset distribution and the selected representative points for each clustering configuration. In all visualizations, grey circles represent the original data points prior to clustering, while red crosses designate the representative points (marked by red crosses) selected by the Cover Tree in the ISCT algorithm are illustrated for three standard synthetic datasets: Moons, Circles, and Blobs. These representatives provide a multi-scale summarization of each dataset. Only a small subset of the original points was retained as representatives, yielding a compact core set that significantly reduced the computational load of subsequent clustering while preserving the intrinsic topology of the data. Because the Cover Tree is hierarchical, the red-cross nodes correspond to points chosen at intermediate levels of the decomposition, reflecting a coarse-to-fine grouping of the data. These visualizations thus demonstrate how ISCT approximates the overall dataset structure via this sparse representation prior to the final clustering stage.
We also used various random datasets represented in the
Table 3 to analyze and evaluate the behavior and performance of clustering algorithms across different distributions and dimensions. The datasets generated include a wide range of distributions and parameters, ensuring a comprehensive evaluation. Each dataset enabled clustering methodologies to be tested and validated in a variety of scenarios. These datasets were generated using NumPy’s random module, encompassing both continuous and discrete probability distributions. Continuous data was produced using uniform (np.random.rand), normal (np.random.normal), exponential (np.random.exponential), gamma (np.random.gamma), and log-normal (np.random.lognormal) distributions. Discrete data included integer uniform (np.random.randint), binomial (np.random.binomial), Poisson (np.random.poisson), and Bernoulli distributions. All distributions were parameterized according to their statistical properties, and random seed initialization ensured the reproducibility of the generated data sequences for simulation and testing purposes.
To evaluate and compare the effectiveness of the ISCT (Improved Clustering with Cover tree), Spectral Clustering and K-Means clustering algorithms, three commonly used evaluation indices are mobilized: the Silhouette index, the Davies–Bouldin index and the Calinski–Harabasz index.
The Silhouette Index measures how similar an object is to its own cluster compared to other clusters. It ranges from −1 to +1, where a high value indicates that the object is well matched to its own cluster and poorly matched to neighboring clusters.
For a data point
i, the silhouette score
is defined as follows:
where
is the average distance between
j and all other points in the same cluster. and
is the minimum average distance from
j to all points in other clusters.
The Davies–Bouldin Index (DBI) is used to evaluate clustering algorithms. A lower DBI indicates better clustering. It is defined as the average similarity ratio of each cluster with the cluster that is most similar to it; a lower value indicates better clustering based on intra-cluster and inter-cluster distances.
For
k clusters, the DBI is
where
is the average distance between each point in cluster
i and the centroid of cluster
i.
is the distance between the centroids of clusters
i and
j.
The Calinski–Harabasz Index (also known as the Variance Ratio Criterion) evaluates the ratio of the sum of between-cluster dispersion and within-cluster dispersion. A higher CH index indicates better-defined clusters; a higher value indicates better clustering based on the ratio of between-cluster dispersion to within-cluster dispersion.
For
k clusters, the CH index is defined as follows:
where
is the trace of the between-group dispersion matrix;
is the trace of the within-cluster dispersion matrix;
n is the total number of points; and
k is the number of clusters. Silhouette is indicated by SI, Davies by Da, and Calinski by Ca.
In unsupervised learning, evaluating partition quality is crucial. Two notable metrics are the Adjusted Rand Index (ARI), denoted , and the Normalized Mutual Information (NMI), denoted .
The ARI
corrects for chance in partition similarity, where a value of 1 indicates perfect agreement. It is calculated as follows:
where
is an element of the contingency table,
and
are row and column sums, and
n is the total number of observations.
The NMI
measures normalized informational redundancy between labelings, with
indicating perfect correlation:
where
is the mutual information between partitions
U and
V, and
represents entropy.
These two indices, and , provide complementary perspectives for validating the robustness of clustering algorithms.
The comprehensive evaluation on
Table 4 of six clustering algorithms across eight datasets reveals distinct performance patterns that highlight the relative strengths and limitations of each method. The proposed ISCT algorithm demonstrates remarkable consistency, achieving superior performance in 31 out of 40 metric–dataset combinations. This dominance is particularly evident on real-world datasets, where ISCT obtains the highest scores across all five metrics for the Iris dataset (ARI: 0.732, NMI: 0.743, SI: 0.5528) and maintains this advantage on Seed, Glass, Mall, and Cancer datasets. The performance gap is especially pronounced on the Seed dataset, where ISCT achieves an ARI of 0.817 compared to DBSCAN’s 0.805, while also recording exceptional internal validation metrics (SI: 0.6344, Ca: 959.75). On synthetic datasets, the results reveal more nuanced patterns. For the Blobs dataset, ISCT demonstrates clear superiority across four of five metrics, with Spectral clustering being competitive. However, the Moons dataset presents an interesting case where Spectral clustering outperforms ISCT in external validation metrics (ARI: 0.3431 vs. 0.3341; NMI: 0.5052 vs. 0.4866), while ISCT maintains an advantage in internal metrics (SI: 0.4720, Da: 0.7106, Ca: 769.2960). This divergence suggests that spectral clustering provides a better grasp of the moon-shaped underlying texture, while ISCT produces more compact clusters. The comparative analysis reveals that DBSCAN and Spectral clustering emerge as the strongest competitors. DBSCAN shows particular strength on the Seed dataset, nearly matching ISCT’s performance, while Spectral clustering excels on the challenging Moons structure. Traditional methods like K-means and Agglomerative clustering generally underperform compared to the density-based and spectral approaches, particularly on complex geometries. The ISCT algorithm demonstrates remarkable strength on various types of data, thanks to its hybrid approach. This integration simultaneously optimizes internal and external validation metrics, making it an important advance for clustering heterogeneous data structures.
The experimental results represented in the
Table 5 across ten high-dimensional datasets reveal that the proposed ISCT method consistently outperforms all benchmark algorithms, achieving top scores in all metrics, including the Adjusted Rand Index (ARI), Normalized Mutual Information (NMI), Silhouette Index (SI), and Calinski–Harabasz Index (Ca), while maintaining the lowest Davies–Bouldin Index (Da). This demonstrates ISCT’s exceptional ability to recover the true class structure while forming clusters with superior internal cohesion and separation.
Among the competitors, hierarchical approaches (Agglomerative Clustering and BIRCH) ranked as the closest alternatives, suggesting their suitability for high-dimensional spaces. DBSCAN showed volatile performance, excelling on datasets like iono and Coil20 but performing poorly on dermatology and segment, highlighting its parameter sensitivity and density structure dependency. K-Means consistently delivered suboptimal results due to limitations in capturing complex, non-linear relationships, while Spectral Clustering outperformed K-Means but was still surpassed by ISCT.
The results establish two key conclusions: ISCT demonstrates itself as a robust, highly effective solution for high-dimensional clustering with significant improvements over techniques, and no single algorithm is universally optimal, as performance depends on specific data characteristics. ISCT’s superior consistency across diverse datasets positions it as a generalizable tool for knowledge discovery in complex data.
This
Table 6 shows the results obtained on random datasets.
ISCT outperforms K-Means and Spectral Clustering on most datasets when evaluated with Silhouette Index (SI), Calinski–Harabasz (Ca), and Davies–Bouldin (Da), showing particular strength on exponential, gamma, and lognormal data. For example, on lognormal data it achieves an SI of 0.26048, compared to 0.213247 for K-Means and 0.132666 for Spectral Clustering. On high-dimensional datasets like normal and binomial, the performance gaps are smaller, but ISCT still leads slightly in SI and Ca. These results highlight its robustness, versatility, and ability to produce more coherent, well separated clusters, thanks to its combined spanning tree decomposition and spectral clustering approach. The performance of various clustering algorithms, including the proposed Improved Clustering with Cover tree (ISCT) method, was evaluated across eleven synthetic datasets. ISCT demonstrated remarkable robustness by securing top ranks in the majority of cases across diverse data distributions (Random, Exponential, Gamma, Normal, and Discrete). Its success stems from a multi-level hierarchical approach using Cover tree decomposition, which efficiently captures intrinsic multi-scale geometry and forms stable clusters from representative superpoints before refining assignments. Other algorithms show significant performance variation. DBSCAN exhibits extreme volatility, performing catastrophically on Random and Bernoulli data due to its dependency on density-separated clusters, while K-Means and hierarchical methods (Agglomerative, BIRCH) achieved stable but sub-optimal results. Notably, ISCT is not universally superior; Spectral Clustering outperforms it on Random Data 2, and BIRCH excels on Bernoulli Data, indicating that no single algorithm dominates all configurations. The evaluation establishes ISCT as a robust general-purpose clustering tool whose Cover tree-based approach provides significant advantages in capturing complex structures across diverse distributions. While traditional algorithms have assumption-related strengths and weaknesses, ISCT offers a consistent, reliable performance, making it ideal for clustering tasks with complex or unknown data distributions.
These
Figure 4,
Figure 5,
Figure 6,
Figure 7,
Figure 8,
Figure 9,
Figure 10,
Figure 11,
Figure 12,
Figure 13,
Figure 14,
Figure 15,
Figure 16,
Figure 17,
Figure 18,
Figure 19,
Figure 20,
Figure 21,
Figure 22,
Figure 23,
Figure 24,
Figure 25,
Figure 26,
Figure 27,
Figure 28,
Figure 29,
Figure 30,
Figure 31,
Figure 32,
Figure 33,
Figure 34,
Figure 35,
Figure 36,
Figure 37,
Figure 38,
Figure 39,
Figure 40,
Figure 41,
Figure 42,
Figure 43,
Figure 44,
Figure 45,
Figure 46,
Figure 47 and
Figure 48 show the clusters obtained by our algorithm and other algorithms.
7. Discussion
7.1. Comparative Performance Analysis of ISCT Algorithm with Other Algorithm Clustering
The comprehensive evaluation on diverse datasets reveals the superior performance of the proposed ISCT method compared to established clustering approaches. The ISCT method achieves important improvements on most performance metrics. On low-dimensional and synthetic datasets, ISCT outperforms conventional methods, with particularly remarkable performance on complex structures. For the Iris and Seed datasets, ISCT achieves ARI values of 0.732 and 0.817, respectively, representing improvements of 1–2% over spectral clustering and 5–8% over K-means. This enhanced performance can be attributed to the ability of the Cover tree to detect intricate geometric structures while maintaining computational efficiency. The robustness of the method is also demonstrated by its consistent performance on most evaluation metrics, achieving optimal values for cohesion-based measures (Silhouette Index) and generally competitive values for separation measures (Davies–Bouldin Index). The results on high-dimensional datasets reveal ISCT’s particular strength in handling complex feature spaces. For the arrhythmia dataset, ISCT achieves an ARI of 0.625, significantly outperforming all benchmark methods. Similarly, for the dermatology dataset, ISCT reaches an exceptional ARI of 0.985, demonstrating near-perfect clustering accuracy. This performance advantage becomes increasingly pronounced in very high-dimensional spaces, suggesting that the Cover Tree-based approximation effectively preserves essential structural information while mitigating the curse of dimensionality. Notably, ISCT maintains strong performance even on challenging random distributions. While all methods understandably show reduced performance on random data, ISCT consistently achieves the best results, with ARI values of 0.055 for random data and 0.120 for exponential distributions. This demonstrates the method’s ability to avoid overfitting and maintain appropriate clustering discipline even in minimally structured environments. The computational efficiency of ISCT, while not explicitly tabulated, can be inferred from the maintained performance quality across varying dataset sizes and dimensionalities. The method’s hierarchical approach enables scalable processing without compromising clustering accuracy, making it particularly suitable for large-scale real-world applications where both precision and efficiency are critical requirements. These results collectively confirm that ISCT is a sturdy and versatile clustering approach that effectively overcomes the limitations of existing methods for various data characteristics, from simple low-dimensional distributions to high-dimensional, sparsely distributed feature spaces.
7.2. Computational Complexity Analysis of Clustering Algorithms
A comparative analysis of the computational complexities of Improved Spectral Clustering with Cover tree (ISCT), Spectral Clustering, and K-Means algorithms in
Table 7 and
Table 8 reveals significant performance differences. Spectral clustering demonstrates the highest computational demand, with a time complexity of
in worst-case scenarios and a space complexity of
, primarily due to the construction of an
similarity matrix.
In contrast, K-Means exhibits superior computational efficiency with a time complexity of , where n represents the number of data points, k the number of clusters, i the iteration count, and d the data dimensionality. Its space complexity remains minimal at , being solely dependent on data dimensions.
The ISCT algorithm achieves an optimized complexity profile of , where the first term corresponds to Cover tree construction and the second to the final K-Means clustering phase. Its space complexity of reflects its dependencies on both data dimensionality and cluster count.
This complexity analysis demonstrates that ISCT substantially outperforms spectral clustering for large datasets due to its complexity compared to . While ISCT exhibits marginally higher complexity than K-Means, this overhead is justified by significantly improved clustering quality for complex data structures. Regarding space efficiency, ISCT proves substantially more scalable than spectral clustering for high-dimensional data, though it is slightly less efficient than K-Means due to its additional dependency on the cluster count k.
In conclusion, ISCT presents a compelling alternative to spectral clustering for large-scale and high-dimensional clustering tasks, while offering a favorable complexity–quality trade-off compared to K-Means for complex data distributions [
45,
46].
Spectral Clustering becomes computationally prohibitive for large datasets () due to cubic time complexity.
K-Means demonstrates linear scalability with sample size but requires careful initialization.
ISCT achieves near-linear scalability through Cover tree approximation while maintaining clustering quality.
All algorithms benefit from dimensionality reduction techniques for high-dimensional data.
For datasets with , spectral clustering provides excellent quality despite its higher complexity.
For large datasets (), K-Means or ISCT are preferred depending on cluster shape complexity.
For high-dimensional data, always apply a dimensionality reduction before clustering.
For complex cluster structures, ISCT provides the optimal balance between quality and scalability.
7.3. Statistical Validation of Rankings
To statistically analyze the results, we employed a non-parametric approach consisting of three steps. First, the Friedman test was used to detect potentially significant differences between treatments across multiple datasets. Subsequently, post hoc Wilcoxon signed-rank tests were conducted to perform pairwise comparisons between our method (ISCT) and each baseline method, applying a Holm–Bonferroni correction to account for multiple comparisons. Finally, a ranking analysis was performed to calculate the average rankings of all methods across all datasets and metrics considered [
47].
7.3.1. Statistical Analysis on Results of Different Clustering Methods for Low-Dimensional and Synthetic Datasets
To rigorously evaluate the performance differences between clustering algorithms, we conducted a comprehensive statistical analysis of the results presented in
Table 9. For each of the 40 experimental configurations (8 datasets × 5 metrics), we ranked the six algorithms from 1 (best) to 6 (worst), with appropriate consideration of metric polarity (higher values preferred for ARI, NMI, SI, and Ca; lower values preferred for Da).
We employed the non-parametric Friedman test to detect significant differences in performance across all algorithms, testing the null hypothesis that all methods perform equivalently. The analysis revealed a Friedman chi-squared statistic of 78.34 with a p-value < 0.0001, providing strong evidence against the null hypothesis and indicating statistically significant performance differences between the methods.
Following this significant result, we conducted a post hoc analysis using pairwise comparisons between ISCT and each baseline algorithm with the Wilcoxon signed-rank test, applying the Holm–Bonferroni correction for multiple comparisons.
Based on
Table 10 and the statistical analysis, several key findings are confirmed:
ISCT demonstrates statistically significant improvements over BIRCH, Spectral, Agglomerative, and K-Means clustering methods (p < 0.05 after multiple comparison correction).
While ISCT shows a better average performance than DBSCAN across most metrics and datasets, this difference does not reach statistical significance after multiple comparison correction (adjusted p = 0.054).
The Friedman test confirms the significant differences between methods (p < 0.0001), with ISCT achieving the best average rank across all datasets and metrics.
These results provide strong statistical support for the superior performance of our ISCT method while acknowledging its comparable performance to DBSCAN, which represents the strongest baseline among the comparison methods. The consistent top ranking of ISCT across diverse datasets and evaluation metrics underscores its robustness and general applicability in various clustering scenarios.
7.3.2. Statistical Analysis on Results of Different Clustering Methods for High-Dimensional Datasets
The procedure was as follows: for each of the 50 experimental configurations (10 datasets × 5 metrics), the six algorithms were ranked from 1 (best) to 6 (worst), with higher values being better for the ARI, NMI, SI, and Ca metrics and a lower value being preferable for the Da metric. The Friedman test was then used to determine if there were statistically significant differences in the average ranks of the algorithms by testing the null hypothesis that all algorithms perform equivalently. Upon rejection of the null hypothesis, a post hoc analysis was conducted using pairwise comparisons between ISCT and each baseline algorithm with the Wilcoxon signed-rank test, where the resulting p-values were adjusted for multiple comparisons using the Holm–Bonferroni method.
The average rank of each algorithm across all experimental configurations is presented in
Table 11. ISCT achieved the lowest (best) average rank.
The Friedman test rejected the null hypothesis that all algorithms perform equally. The statistical analysis yielded a Friedman chi-squared statistic of with a p-value of less than 0.00001, demonstrating highly significant differences in performance among the algorithms under evaluation. This extremely significant result (p < 0.00001) provides compelling evidence against the null hypothesis of equivalent algorithmic performance, indicating that at least one algorithm exhibits significantly different performance characteristics compared to the others.
Post hoc Wilcoxon Signed-Rank Tests
The results of the pairwise comparisons (
Table 12) show that ISCT’s performance is statistically superior to all other methods after correction for multiple comparisons.
The statistical analysis provides unequivocal evidence that ISCT outperforms all baseline methods on high-dimensional data. It achieved the best average rank and the Friedman test confirmed that there were significant differences between the methods (p < 0.00001). The post hoc Wilcoxon tests confirmed that ISCT’s superiority is statistically significant against every baseline, even after a stringent multiple-comparison correction. These results robustly support our claims regarding the effectiveness of the ISCT algorithm.
7.3.3. Statistical Analysis on Results of Different Clustering Methods for Random Datasets
The evaluation procedure consisted of three main steps: first, for each of the 55 experimental configurations (11 datasets × 5 metrics), all six algorithms were ranked from 1 (best) to 6 (worst), with higher values indicating better performance for the ARI, NMI, SI, and Ca metrics, and lower values being preferable for the Da metric; second, the non-parametric Friedman test was employed to determine whether there were statistically significant differences in the average rankings across all algorithms; finally, upon rejection of the null hypothesis, post hoc pairwise comparisons between ISCT and each baseline algorithm were conducted using the Wilcoxon signed-rank test with Holm–Bonferroni correction for multiple comparisons.
Table 13 shows the average rank of each algorithm across all experimental configurations. ISCT achieved the best average rank.
The Friedman test rejected the null hypothesis that all algorithms perform equally. The statistical analysis revealed a Friedman chi-squared statistic of 89.47 with a p-value of less than 0.00001, indicating highly significant differences in algorithmic performance across the evaluated methods. This extremely significant result (p < 0.00001) provides strong statistical evidence against the assumption of equal performance among all algorithms, suggesting that at least one algorithm demonstrates significantly different performance compared to the others in the comparison.
Post hoc Wilcoxon Signed-Rank Tests
Table 14 shows the results of pairwise comparisons between ISCT and other methods after Holm–Bonferroni correction.
The statistical analysis demonstrates that ISCT achieves the best overall performance on synthetic datasets, with statistically significant improvements over Spectral Clustering, DBSCAN, K-Means, and BIRCH. Although ISCT shows a better average performance than Agglomerative clustering, this difference does not reach statistical significance after multiple comparison correction. These results provide rigorous statistical support for our claims regarding ISCT’s effectiveness while acknowledging its specific performance relationships with different algorithms.
7.4. Adaptation Spectral Cover Tree for High Dimensions
Spectral clustering is a powerful technique for identifying clusters in data by leveraging the eigenstructure of similarity matrices. However, as data dimensionality increases, traditional spectral clustering algorithms face significant challenges, including computational inefficiency and difficulty capturing meaningful structures in the data. To address these issues, an adaptation of spectral clustering using Cover tree can be highly effective. This approach combines the benefits of spectral methods with the efficiency of Cover tree [
30], a data structure designed for high-dimensional spaces [
27]. This section discusses the adaptation of spectral clustering with Cover tree, focusing on the motivation, methodology, and implications for high-dimensional data.
In high-dimensional spaces, the curse of dimensionality can severely affect clustering performance. Traditional spectral clustering involves computing a similarity matrix, which can be computationally expensive and memory-intensive for large datasets. Additionally, an eigenvalue decomposition of this matrix can become prohibitive in terms of both time and space complexity. The Cover tree is an efficient data structure for high-dimensional nearest-neighbor searches, which can help overcome these challenges by speeding up the computation of similarity matrices and enhancing the scalability of spectral clustering. We will discuss the methodology to be followed in four essential steps.
Data Decomposition Using Cover tree
The initial phase of adapting spectral clustering with Cover tree involves decomposing the data, resulting in a set of representative points and a tree structure that captures the inherent clustering structure of the data. The process begins with the construction of the Cover tree, which is achieved by recursively partitioning the data into smaller subsets based on distance thresholds. This hierarchical structure groups data points into increasingly finer clusters. The nodes of the Cover tree serve as representative points, summarizing the clustering structure at different levels of the hierarchy.
Forming the Representative Points Similarity Matrix
Following the construction of the Cover tree, a similarity matrix W is formed based on the representative points. This matrix captures the pairwise similarities between these points and is used to construct a reduced similarity matrix for spectral clustering. The similarity between representative points is computed using appropriate distance measures, reflecting the local structure of the data as represented by the Cover tree nodes. The similarity matrix is then normalized to ensure it is suitable for spectral clustering, typically involving standardization or the application of a kernel function.
Spectral Clustering on Representative Points
With the similarity matrix W in hand, spectral clustering is applied to the reduced set of representative points. This involves performing an eigenvalue decomposition on the normalized similarity matrix to obtain the principal eigenvectors. These eigenvectors are used to project the representative points into a lower-dimensional space, where a classical clustering algorithm, such as K-Means, is applied to partition these points into clusters.
Recovering Clusters for Original Data
The final step involves recovering the cluster assignments for the original data points by mapping them back to the clusters of their corresponding representative points. Each data point is assigned to the cluster of the representative point closest to it in the original space. Optionally, the clustering results can be refined using additional techniques, such as post-processing or fine-tuning, to improve clustering quality.
7.5. Implications of Adaptation Spectral Cover Tree for High Dimension
This method delivers important improvements in both efficiency and quality, while guaranteeing greater scalability. This innovative combination paves the way for practical, high-performance applications in contexts involving different datasets. The adaptation of spectral clustering with Cover tree offers a robust solution to the challenges of high-dimensional and large-scale datasets by enhancing computational efficiency, improving clustering quality, and ensuring scalability. By reducing dimensionality and focusing on representative points, it mitigates the high computational costs of traditional spectral clustering, resulting in faster processing times and lower memory usage. The hierarchical structure of the Cover tree captures both local and global data relationships effectively, leading to more accurate clustering, particularly in high-dimensional spaces where conventional methods often struggle. Additionally, the efficiency of Cover tree in nearest-neighbor searches and the reduced complexity of working with representative points enable the algorithm to scale seamlessly with data size and dimensionality. This methodology represents a significant advancement in clustering techniques, making spectral clustering more feasible and effective for different datasets.
7.6. Limitations
The adaptation of spectral clustering with Cover tree introduced significant advancements in handling high-dimensional data. However, its practical application is accompanied by several limitations that warrant consideration. This section explores the key challenges associated with the use of Cover tree in spectral clustering, focusing on computational complexity, approximation accuracy, parameter tuning, and scalability.
Computational Complexity and Overhead
Although Cover trees are designed for efficiency, their construction and maintenance present computational challenges, particularly in high-dimensional spaces. Building and updating a Cover tree involves recursive partitioning and the management of hierarchical structures [
48], which can introduce significant computational overhead. In scenarios involving extremely large datasets or very high-dimensional data, this complexity may offset the efficiency gains expected from the approach.
Another computational bottleneck arises during the calculation of similarity matrices. While Cover trees facilitate rapid nearest-neighbor searches, the process of computing and normalizing similarity matrices from representative points remains resource-intensive, particularly for large-scale datasets. Additionally, the memory requirements for storing both the Cover tree and the similarity matrices can be substantial, potentially necessitating advanced hardware resources to handle massive or high-dimensional datasets effectively.
Approximation and Accuracy
The use of representative points in Cover tree can lead to a loss of precision in clustering results. These points, while reducing data complexity, may fail to capture all nuances of the original dataset, resulting in suboptimal clustering outcomes. The sensitivity of the approach to the selection of representative points is particularly evident in datasets with overlapping or complex cluster structures, where a poorly constructed Cover tree can significantly degrade accuracy.
Moreover, spectral clustering relies on an eigenvalue decomposition of similarity matrices, and the use of approximated matrices derived from Cover trees introduces errors in eigenvalue and eigenvector computations [
30]. These approximation errors may compromise the quality of the final clustering, especially in cases where the representative points inadequately reflect the global data structure.
Parameter Tuning and Selection
The effectiveness of spectral clustering with Cover trees is highly dependent on parameter tuning. Key parameters such as the number of representative points, distance thresholds, and similarity computation methods must be carefully chosen to balance efficiency and accuracy [
27]. Identifying optimal parameter values often requires extensive experimentation and cross-validation, adding to the complexity of implementation.
Additionally, the approach is sensitive to hyperparameters, such as the choice of distance metric and the number of clusters. Misconfigured parameters can result in poor clustering performance [
49], reducing the effectiveness of the method. Balancing computational efficiency with clustering accuracy requires a nuanced understanding of both the data and the algorithm’s behavior.
Scalability and Applicability
While Cover tree address high-dimensional data challenges, their performance may degrade in extremely high-dimensional spaces with thousands of features. The effectiveness of Cover tree is also contingent on the nature of the data. Irregular or non-Euclidean structures may not align well with the assumptions underlying the Cover tree, limiting their applicability across diverse domains and datasets.
The complexity of implementing spectral clustering with Cover tree is another practical challenge. The approach involves multiple interconnected components, including Cover tree construction, similarity matrix computation, and spectral analysis [
50]. For practitioners unfamiliar with these methods, the implementation process can be daunting, potentially hindering its adoption in real-world applications.
The adaptation of spectral clustering with Cover tree offers valuable benefits for clustering high-dimensional data, but it is accompanied by notable limitations. Challenges related to computational complexity, approximation accuracy, parameter tuning, and scalability must be carefully addressed to maximize its potential. Understanding these limitations is essential for effectively applying this method and interpreting its results. Future research can focus on refining algorithms, optimizing implementations, and exploring alternative techniques to enhance the robustness and versatility of this approach across various contexts.
8. Future Research
The integration of Cover tree with spectral clustering represents a significant innovation in the analysis of high-dimensional data. Despite its promising performance, numerous avenues for future research can further enhance the efficiency, accuracy, and applicability of this method. This section outlines critical areas for advancing spectral clustering with Cover tree, focusing on algorithmic improvements, robustness, flexibility, and theoretical understanding.
We aim to generalize the ISCT algorithm to probabilistic metric spaces, which will enable the uncertainties inherent in real-world data to be captured, particularly in contexts where distances or similarities are stochastic (e.g., data from noisy sensors, complex networks, or dynamic systems). This generalization will require adapting the Cover tree construction to incorporate probability distributions over metrics while preserving the theoretical guarantees of the hierarchical structure.
Furthermore, we will explore advanced association methods, including optimal transport techniques, random graph models, and transfer learning mechanisms. These approaches seek to improve the quality of computed affinities between data points, especially in scenarios where relationships are non-linear or multi-scale. Integrating these methods with Cover tree could lead to a significant reduction in the biases induced by initial approximations while enhancing robustness to outliers.
The integration of Cover tree with spectral clustering represents a significant innovation in the analysis of high-dimensional data. Despite its promising performance, numerous avenues for future research could further enhance the efficiency, accuracy, and applicability of this method [
51]. This section outlines critical areas for advancing spectral clustering with Cover tree, focusing on algorithmic improvements, robustness, flexibility, and theoretical understanding.
Enhancing Algorithmic Efficiency
Efforts to optimize the construction and maintenance of Cover tree could lead to significant performance gains. The current implementations, while efficient, still involve computational overhead, particularly in high-dimensional spaces. Future research can explore advanced partitioning strategies and alternative data structures to reduce the time complexity of tree construction and updates. Additionally, accelerating similarity matrix computations through parallelization, approximate algorithms, or novel similarity measures can address one of the main bottlenecks in the clustering process. Finally, developing scalable algorithms that leverage distributed computing or memory-efficient strategies could enable the application of this approach to extremely large and complex datasets.
Improving Accuracy and Robustness
The precision of clustering results in spectral clustering with Cover tree depends on the accuracy of approximations and parameter tuning. Future studies could focus on refining approximation techniques, potentially integrating Cover tree with other dimensionality reduction methods or hybrid approaches. Automated parameter tuning methods, such as metaheuristics or machine learning-based systems, could further improve clustering performance by dynamically adjusting settings based on dataset characteristics. Moreover, enhancing robustness against noise and outliers through noise-resistant algorithms or robust similarity measures is crucial for handling real-world data with inherent imperfections.
Expanding Applicability and Flexibility
Adapting spectral clustering with Cover tree to diverse data types is another promising research direction. Extending this method to time-series data, graph-based data, and heterogeneous datasets could broaden its applicability across various domains. Additionally, integrating Cover tree with deep learning techniques, such as autoencoders and neural networks, may improve feature extraction and clustering performance, enabling the development of hybrid models that combine classical clustering with modern machine learning. To facilitate practical adoption, future work should also focus on creating user-friendly tools, visualization interfaces, and software libraries that simplify the application of these advanced methods.
Advancing Theoretical Understanding
A deeper theoretical exploration of Cover tree and its role in spectral clustering is essential to uncovering the full potential of this approach. Research could delve into the mathematical properties of Cover tree, its impact on clustering accuracy, and its limitations. Extending the theoretical framework to include adaptations for non-Euclidean spaces, dynamic datasets, or multi-view data could provide valuable insights into how Cover tree can address more complex clustering scenarios. Furthermore, comparative studies between spectral clustering with Cover tree and other state-of-the-art clustering techniques can clarify their relative strengths and guide best practices for their application.
The adaptation of spectral clustering using Cover tree presents a robust and scalable solution for clustering high-dimensional data. However, significant opportunities remain to refine its efficiency, accuracy, and applicability. By addressing challenges related to algorithmic optimization, robustness, and flexibility, and by expanding its theoretical foundations, researchers can further advance this promising approach. These efforts will not only enhance the method’s performance but also facilitate its adoption across a broader range of data analysis tasks, solidifying its place as a valuable tool in the field of clustering.