Exploiting Genomic Relations in Big Data Repositories by Graph-Based Search Methods
Abstract
1. Introduction
2. Preliminaries
3. The Pharmacogenomics Data Repository LINCS

4. Technical Considerations
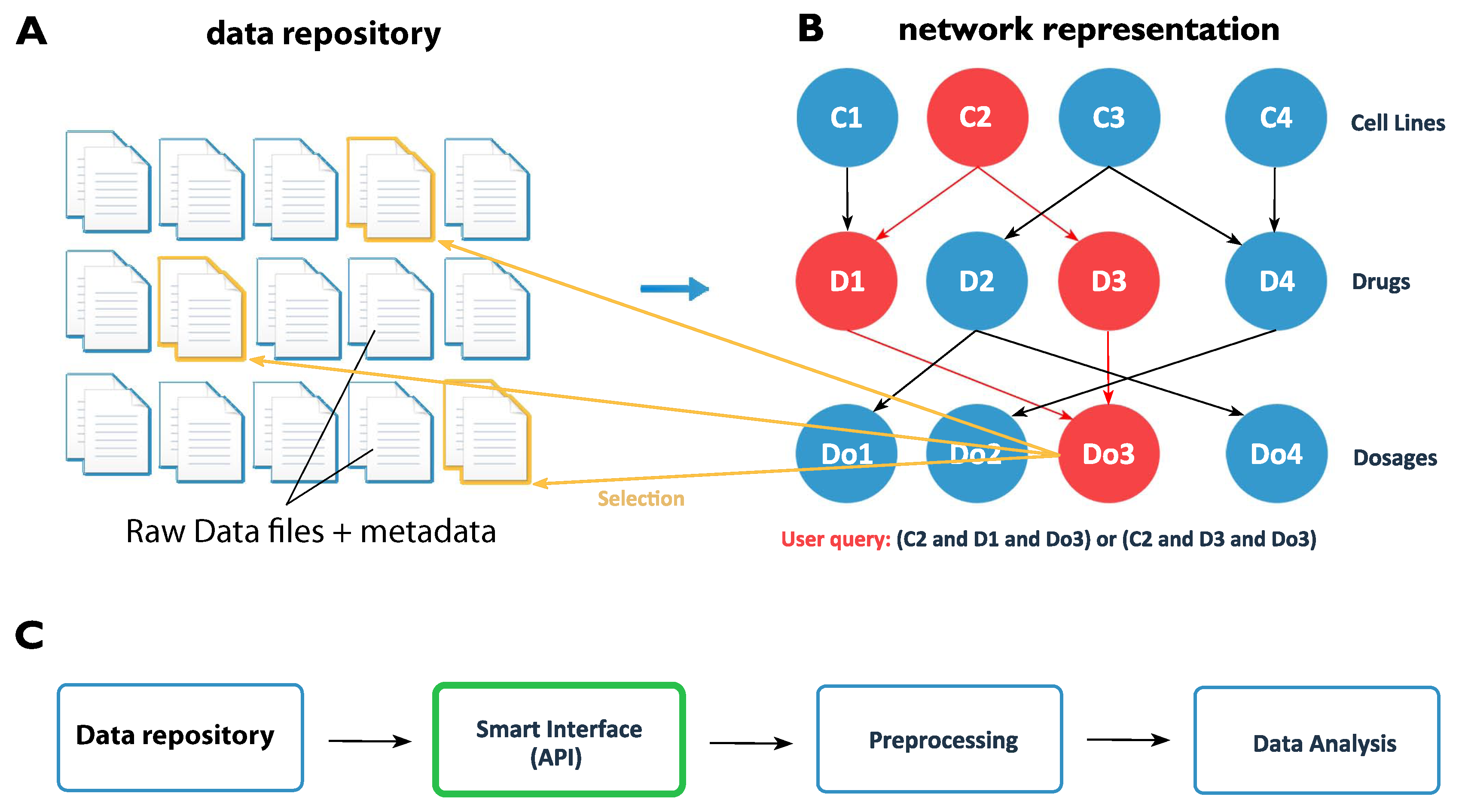
5. Conceptual Idea
6. Further Applications
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Edgar, R.; Domrachev, M.; Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002, 30, 207–210. [Google Scholar] [CrossRef] [PubMed]
- Holzinger, A.; Jurisica, I. Knowledge discovery and data mining in biomedical informatics: The future is in integrative, interactive machine learning solutions. In Interactive Knowledge Discovery and Data Mining in Biomedical Informatics; Springer: Berlin, Germany, 2014; pp. 1–18. [Google Scholar]
- Lamb, J.; Crawford, E.D.; Peck, D.; Modell, J.W.; Blat, I.C.; Wrobel, M.J.; Lerner, J.; Brunet, J.P.; Subramanian, A.; Ross, K.N.; et al. The Connectivity Map: Using gene-expression signatures to connect small molecules, genes, and disease. Science 2006, 313, 1929–1935. [Google Scholar] [CrossRef] [PubMed]
- Ma’ayan, A.; Rouillard, A.; Clark, N.; Wang, Z.; Duan, Q.; Kou, Y. Lean Big Data integration in systems biology and systems pharmacology. Trends Pharmacol. Sci. 2014, 35, 450–460. [Google Scholar] [CrossRef] [PubMed]
- Campillos, M.; Kuhn, M.; Gavin, A.C.; Jensen, L.J.; Bork, P. Drug target identification using side-effect similarity. Science 2008, 321, 263–266. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Narayan, R.; Corsello, S.M.; Peck, D.D.; Natoli, T.E.; Lu, X.; Gould, J.; Davis, J.F.; Tubelli, A.A.; Asiedu, J.K.; et al. A Next Generation Connectivity Map: L1000 Platform And The First 1,000,000 Profiles. BioRxiv 2017. [Google Scholar] [CrossRef] [PubMed]
- Musa, A.; Ghoraie, L.; Zhang, S.D.; Glazko, G.; Yli-Harja, O.; Dehmer, M.; Haibe-Kains, B.; Emmert-Streib, F. A Review of Connectivity Mapping and Computational Approaches in Pharmacogenomics. Brief. Bioinform. 2017, 19, 506–523. [Google Scholar]
- Musa, A.; Tripathi, S.; Kandhavelu, M.; Dehmer, M.; Emmert-Streib, F. Harnessing the biological complexity of Big Data from LINCS gene expression signatures. PLoS ONE 2018, 13, e0201937. [Google Scholar] [CrossRef] [PubMed]
- Vidovic, D.; A, K.; Schurer, S. Large-scale integration of small molecule-induced genome-wide transcriptional responses, Kinome-wide binding affinities and cell-growth inhibition profiles reveal global trends characterizing systems-level drug action. Front. Genet. 2014, 5, 342. [Google Scholar] [PubMed]
- Barrett, T.; Troup, D.B.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; et al. NCBI GEO: Archive for functional genomics data sets -10 years on. Nucleic Acids Res. 2011, 39, D1005–D1010. [Google Scholar] [CrossRef] [PubMed]
- Codd, E.F. A Relational Model of Data for Large Shared Data Banks. Commun. ACM 1970, 13, 377–387. [Google Scholar] [CrossRef]
- Wiese, L. Advanced Data Management: For SQL, NoSQL, Cloud and Distributed Databases; De Gruyter: Berlin, Germany, 2015. [Google Scholar]
- Angles, R.; Gutierrez, C. Survey of Graph Database Models. ACM Comput. Surv. 2008, 40, 1–39. [Google Scholar] [CrossRef]
- Zou, L.; Chen, L.; Özsu, M.T. Distance-join: Pattern match query in a large graph database. Proc. VLDB Endowment 2009, 2, 886–897. [Google Scholar] [CrossRef]
- Himmelstein, D.S.; Lizee, A.; Hessler, C.; Brueggeman, L.; Chen, S.L.; Hadley, D.; Green, A.; Khankhanian, P.; Baranzini, S.E. Systematic integration of biomedical knowledge prioritizes drugs for repurposing. eLife 2017, 6, e26726. [Google Scholar] [CrossRef] [PubMed]
- Matthews, L.; Gopinath, G.; Gillespie, M.; Caudy, M.; Croft, D.; de Bono, B.; Garapati, P.; Hemish, J.; Hermjakob, H.; Jassal, B.; et al. Reactome knowledgebase of human biological pathways and processes. Nucleic Acids Res. 2009, 37, D619–D622. [Google Scholar] [CrossRef] [PubMed]
- Swainston, N.; Batista-Navarro, R.; Carbonell, P.; Dobson, P.D.; Dunstan, M.; Jervis, A.J.; Vinaixa, M.; Williams, A.R.; Ananiadou, S.; Faulon, J.L.; et al. biochem4j: Integrated and extensible biochemical knowledge through graph databases. PLoS ONE 2017, 12, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Touré, V.; Mazein, A.; Waltemath, D.; Balaur, I.; Saqi, M.; Henkel, R.; Pellet, J.; Auffray, C. STON: Exploring biological pathways using the SBGN standard and graph databases. BMC Bioinform. 2016, 17, 494. [Google Scholar] [CrossRef] [PubMed]
- Cormen, T.; Leiserson, C.; Rivest, R.; Stein, C. Introduction to Algorithms; MIT Press: Cambridge, MA, USA, 2001. [Google Scholar]
- Lipski, W.; Marek, W. File organization, an application of graph theory. In Automata, Languages and Programming: 2nd Colloquium, University of Saarbrücken 29 July– 2 August 1974; Loeckx, J., Ed.; Springer: Berlin/Heidelberg, Germany, 1974; pp. 270–279. [Google Scholar]
- Lipski, W. Information storage and retrieval? mathematical foundations II (combinatorial problems). Theor. Comput. Sci. 1976, 3, 183–211. [Google Scholar] [CrossRef]
- Baeza-Yates, R.; Ribeiro-Neto, B. Modern Information Retrieval; ACM Press: New York, NY, USA, 1999; Volume 463. [Google Scholar]
- Chowdhury, G.G. Introduction to Modern Information Retrieval; Facet Publishing: London, UK, 2010. [Google Scholar]
- Chang, F.; Dean, J.; Ghemawat, S.; Hsieh, W.C.; Wallach, D.A.; Burrows, M.; Chandra, T.; Fikes, A.; Gruber, R.E. Bigtable: A distributed storage system for structured data. ACM Trans. Comput. Syst. 2008, 26, 4. [Google Scholar] [CrossRef]
- Shoemaker, R.H. The NCI60 human tumour cell line anticancer drug screen. Nat. Rev. Cancer 2006, 6, 813–823. [Google Scholar] [CrossRef] [PubMed]
- Brazma, A.; Parkinson, H.; Sarkans, U.; Shojatalab, M.; Vilo, J.; Abeygunawardena, N.; Holloway, E.; Kapushesky, M.; Kemmeren, P.; Lara, G.G.; et al. ArrayExpress-a public repository for microarray gene expression data at the EBI. Nucleic Acids Res. 2003, 31, 68–71. [Google Scholar] [CrossRef] [PubMed]
- Barretina, J.; Caponigro, G.; Stransky, N.; Venkatesan, K.; Margolin, A.A.; Kim, S.; Wilson, C.J.; Lehár, J.; Kryukov, G.V.; Sonkin, D.; et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012, 483, 603–607. [Google Scholar] [CrossRef] [PubMed]
- Dehmer, M.; Emmert-Streib, F. (Eds.) Analysis of Complex Networks: From Biology to Linguistics; Wiley-VCH: Weinheim, Germany, 2009. [Google Scholar]
- Emmert-Streib, F.; Moutari, S.; Dehmer, M. The process of analyzing data is the emergent feature of data science. Front. Genet. 2016, 7, 12. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Musa, A.; Dehmer, M.; Yli-Harja, O.; Emmert-Streib, F. Exploiting Genomic Relations in Big Data Repositories by Graph-Based Search Methods. Mach. Learn. Knowl. Extr. 2019, 1, 205-210. https://doi.org/10.3390/make1010012
Musa A, Dehmer M, Yli-Harja O, Emmert-Streib F. Exploiting Genomic Relations in Big Data Repositories by Graph-Based Search Methods. Machine Learning and Knowledge Extraction. 2019; 1(1):205-210. https://doi.org/10.3390/make1010012
Chicago/Turabian StyleMusa, Aliyu, Matthias Dehmer, Olli Yli-Harja, and Frank Emmert-Streib. 2019. "Exploiting Genomic Relations in Big Data Repositories by Graph-Based Search Methods" Machine Learning and Knowledge Extraction 1, no. 1: 205-210. https://doi.org/10.3390/make1010012
APA StyleMusa, A., Dehmer, M., Yli-Harja, O., & Emmert-Streib, F. (2019). Exploiting Genomic Relations in Big Data Repositories by Graph-Based Search Methods. Machine Learning and Knowledge Extraction, 1(1), 205-210. https://doi.org/10.3390/make1010012






