PeTaL (Periodic Table of Life) and Physiomimetics
Abstract
1. Introduction
2. Misconceptions, Cautions and Opportunities
- Life supports life or life supports conditions for life. There are many ways to interpret this statement. It is possible that an organism may inflict harm on another organisms, but that this is in fact a benefit to the larger ecosystem. An example of this is predation. Predation is beneficial to the predator because it provides an energy source. It keeps the population of prey in check and thus ensures ecosystem stability. If the rate of predation is too large this stability is no longer viable.
- There is no waste in living systems and a corollary, all elements of a living system must have a purpose. The obvious exceptions are spandrels [14]. Spandrels are remnants from adaptations that may not have an obvious function. These may eventually turn out to be exaptations given a change in the external ecosystem. An example of this is the development of vectored ejection of noxious fluids by bombardier beetles [15] that could have resulted from the chemicals used in producing its exoskeleton.
- Trade-offs are always the best way to identify solutions inspired by nature. Trade-offs are inevitable in any optimized design given a fixed set of resources [16,17]. An increase in one performance metric must result in a reduction of another assuming that the performance has been optimized globally. To apply trade-offs to natural systems we must assume that they are optimized in some manner. If not, trade-offs, the way engineers use them in a biological context:
- Do not necessarily comport with the laws of logic, namely, given that there exists a claim p, then
- Either p or “not p” must be true
- p and “not p” cannot both be true
- there are no other options other than p and not p.
The statement p would take the form “denticles reduce drag on shark skin”. This does not imply that “denticles increase mass on shark skin” or “denticles reduce shark life”. These may very well be true, but there is no evidence that this is a general law. Energy and material may trade among numerous functions and one may not in fact see trade-offs between two variables that trade a particular quantity, because there are more than two functions trading that quantity. For human systems, by design, we have introduced trade-offs given the limitations of available energy, cost and resources to build and therefore, these trade-offs are an appropriate method of assessing systems or proposing solutions. In nature, trade-offs are more dynamic and comparing trade-offs amongst multiple species requires a way to scale from one species to another. This does not mean that trade-offs are not useful as an innovation tool. To go beyond suggesting solutions, we must define the quantity being traded (e.g., energy), the resulting functions that trade the quantity or quality (e.g., swim speed, turn radius, target tracking precision, body mass) and ensure that these are the only functions (or the major functions) involved in trading the quantity. - Assume linearity in trades—given that we have the complete set of contradictions over which quantities such as energy or matter are traded, we assume that an increase in one, necessarily correlates with a decrease in the other. They are thus only a reflection of our understanding of biology in an engineering context and may not help us develop multifunctional systems without additional data.
- Can be used for a relatively (to the number of species available) limited (but large in number) class of problems. There are some organisms and living systems that do in fact show clear trades between two functions [18,19] such as speed and accuracy. This does not mean other functions are not varying, but rather that these are the primary functions involved in trading the particular quantity.
- Multivariate trade-offs, by virtue of being a necessary feature of optimized systems, are a great way of characterizing multifunctional ideas such as structural heat exchangers or social strategies such as leadership. In nature, leadership takes many forms. The alpha male in a group of chimpanzees may show more empathy toward others, favor egalitarianisms and objectivity in disputes. At the same time, populations may give the leader or dominant individual(s) a majority of food or other resources in exchange for protection. The two scenarios seem contradictory. However, if viewed through the lens of context they make sense. As the population (team) becomes more stable with fewer threats from the organization or environment, leadership is more about getting the most out of the team. If the team is focused mostly on self-preservation, then it makes sense for the team to invest in defensive capability (by feeding the dominant individuals for example). This is an example of a trade-off between progressive and conservative values that depend on the perceived or real stability in the environment.
- Biological systems are tending toward some optimal or best design. In reality, there are numerous examples of features that are clearly not optimal, but are not fatal and thus persist. Some examples of these are:
- Adaptations can be reversed. The Inaccessible Island Rail (Atlantisia rogersi) is the smallest extant flightless bird [20]. Atlantis elpenor, Aphanocrex podarces and Atlantisia rogersi are believed to have shared a common ancestor that was blown to isolated islands where all three species evolved into flightless birds. The Island rail lacks mammalian predators and thus flight is not selected for. In fact, reducing flight muscle size results in energy savings. In addition, the bird has low basal metabolic rate. Evidence thus points to an adaptation that was once useful but is no longer necessary and might actually be a waste of energy.
- Context is key. Adaptations are based on the history of the system and thus, even successful adaptations are not optimal. An example is the human optical system [21].
- For the greater good: Some adaptations are not particularly beneficial to the individuals or the species itself, but do result in benefits to the super system. Squirrels burying nuts results in wasted energy. However, the nuts that are not found result in new trees. Ants forming an ant bridge do not all survive, but this emergent behavior is beneficial to the ant colony.
- Adaptations favor short term gains: While successive selection pressures may eventually lead to a robust set of adaptations, no particular adaptation is suited to long term benefit [22]. Clune et al. [22] used digital simulations to show that natural selection preferred mutation rates that were suboptimal in the long run.
- There are other areas of philosophical conflict such as the differences and similarities between bio-utilization and biomimicry. In one sense, bio-utilization, or the use of biology directly in technology can be thought of as the most sustainable form of biomimicry. For example, recreating the earth’s ecosystem on another planet to minimize the use of terrestrial technology would save on energy and mass of raw materials that need to be sent off-world. In another sense, bio-utilization such as the use of leather or other animal products may require the use of large quantities of water energy for processing and are associated with ethical questions. In this case bio-utilization is not sustainable. Thus, the question is not so much about terminology but about sustainability.
3. Expanding Our Domain of Inquiry
3.1. Physioteleology
- Longevity of species, L (years): The average lifespan of a species is between 1 and 10 million years [23]. For mammals, the average is 1 million years. If a species survives longer than this time it is reasonable to expect that it has survived minor climatic variations, ecosystem pressures or competition from other species adapted to its environment. Examples of this are living fossils such as goblin sharks (origin in fossil records 125 Mya) or horseshoe crabs (origin in fossil records 450 Mya) that have remained virtually unchanged over millions of years [24,25]. This criteria alone is not sufficient to evaluate the bio-fitness of an organism. Photosynthetic cyanobacteria for example, existed for billions of years but caused an oxygen crisis due to there not being any organisms to breathe oxygen. Thus, we may want to give the lifespan more weight if the total number of species alive at the time is large.
- Longevity of adaptation, La (years): This metric would count the time a particular adaptation has existed, such as the eye, in a stable form. Adaptations that exist together within a given species may then be thought of as highly compatible and successful. An example is denticles on sharks.
- Variation in genus, Vg (number of species in the genus): This is an indicator of the success of the general body plan, set of adaptations or morphology. A related criteria may be convergence.
- Convergence, C (number of times a given body plan or feature has evolved independently): Computer vision may be used to create clusters of body plans. In the short term, it may be sufficient to identify genera and compare similar genera and note the number of species per genera as a metric.
- Survival of extinction events, Next (number of mass extinction events survived in same geographic location): This may be an indicator of adaptability to large variations in climate, atmospheric composition, temperature, pressure or seismic events. The adaptations may be behavioral, such as being able to burrow or exist on the surface, or morphological, such as extremophiles that survive cosmic radiation.
- Types of events and reasons for surviving
- Adaptations that allow the species to survive drastic changes.
- Sufficient geographic extent: Some members of species were wiped out but those in a different geographical location persisted. This would mean that survival was largely due to chance but having wide geographic extent is in itself a merit.
- Evolutionary fitness, f: This is a measure of reproductive success of an organism and is characterized by absolute or relative fitness.
- Ecosystem diversity and networking, Ed (number of species connected to a given species through food web or other interactions given Es, the total number of species in the ecosystem): This is an indication of the size of ecosystem supported and the number of connections within the ecosystem. For practitioners of biomimetics, choosing an appropriate model organism is useful to minimize contradictions between the system being designed and the inspirational model. The connections may be gleaned from an ontology that identifies connections between species through NLP (natural language processing) using sources such as Animal Diversity Web [26] or Encyclopedia of Life [27].
- Ecosystem stability (temporal), Et (turbulence of the ecosystem or rms (root mean square) deviation form basal extinction rate): This answers the question of how the ecosystem reacts to change. Typically this is measured by global parameters such as biodiversity, but we could also study the individual components of an ecosystem to see how long the system has remained relatively unchanged (extinction rate of species that are part of the ecosystem). Eavg is the average life of an ecosystem.
- Number of ecosystems supported, En: For the geographical range of a species, how many ecosystems does it support? Etot is the total number of ecosystems with overlap.
- Scalability, S (Variance of spatial scales with 0 indicating high specialization and Smax being the maximum variance ever found in nature): For a species or pattern, this metric would indicate the morphological spatial scalability. This may be measured by relating the patterns that comprise the overall morphology to functions. The patterns in turn have mathematical rules that can be scaled. For example, the venation patterns on dragonfly wings may be characterized by angles, cell widths and thicknesses of veins. Given the knowledge of performance at different scales, one can determine what kind of design is required for thermal, aerodynamic and structural performance at a new scale. Given sufficient information, we may use nondimensional scaling parameters such as Reynolds number, Froude number and Biot number to display natural system information in a manner that is directly applicable to engineering application.
- Suitability of a model may then be calculated by applying the formula:
3.2. Anthropomimesis
3.3. Paleomimesis
4. PeTaL (Periodic Table of Life)
4.1. Intent of PeTaL
- (a)
- Problem definition
- Identify a set of words or phrases (topics) relevant to the field (not exhaustive): distribution, generation, management of heat, energy, stress, convection, conduction, cooling, heating, evaporation, condensation. These terms form the basis for searching articles in the literature, patents, blogs or books. Ideally, these words themselves are connected in a ‘bag of words’ to related terminology to broaden the search and reduce the burden on the user.
- Define the problem as a set of goals (minimum heat flux, maximum stress and minimum weight), boundary conditions (temperature, pressure, humidity, volume) and constraints (gravity, material must be continuous and supported, properties must exist in the natural world).
- (b)
- Identify possible models based on historical work
- Human generated literature
- i.
- Patents, literature: Use NLP (natural language processing) to identify articles related to the problem at hand through search terms related to the problem. The objective is to uncover solutions that have direct implementation, data that may be used to train models, or literature that identifies human technology (heat pipes, pulsating heat pipes, pins, heat exchangers, fluidic devices, cooling schemes for turbines, anti-icing technology) that may be further investigated thought the generation of synthetic data.
- Natural databases: The objective is to identify strategies in nature that enable solutions to the problem at hand. In the case of thermal management, one would search for articles that discuss distribution, generation, management of heat, energy, stress, convection, conduction, cooling, heating, evaporation, condensation. These terms form a word cloud. If articles from the literature are already classified or clustered into word clouds, then the search is greatly simplified. This may be done through the use of a search tool that translates user search terms to biology or through algorithms that find occurrences of noun, verb, noun combinations that represent function, object, and environment. Sources may include:
- ii.
- natural data sets such as Encyclopedia of Life, iDigBio, Wikipedia provide general information on organisms.
- iii.
- research data and information that may be gleaned from publications such as the Journal of Experimental Biology.
- (c)
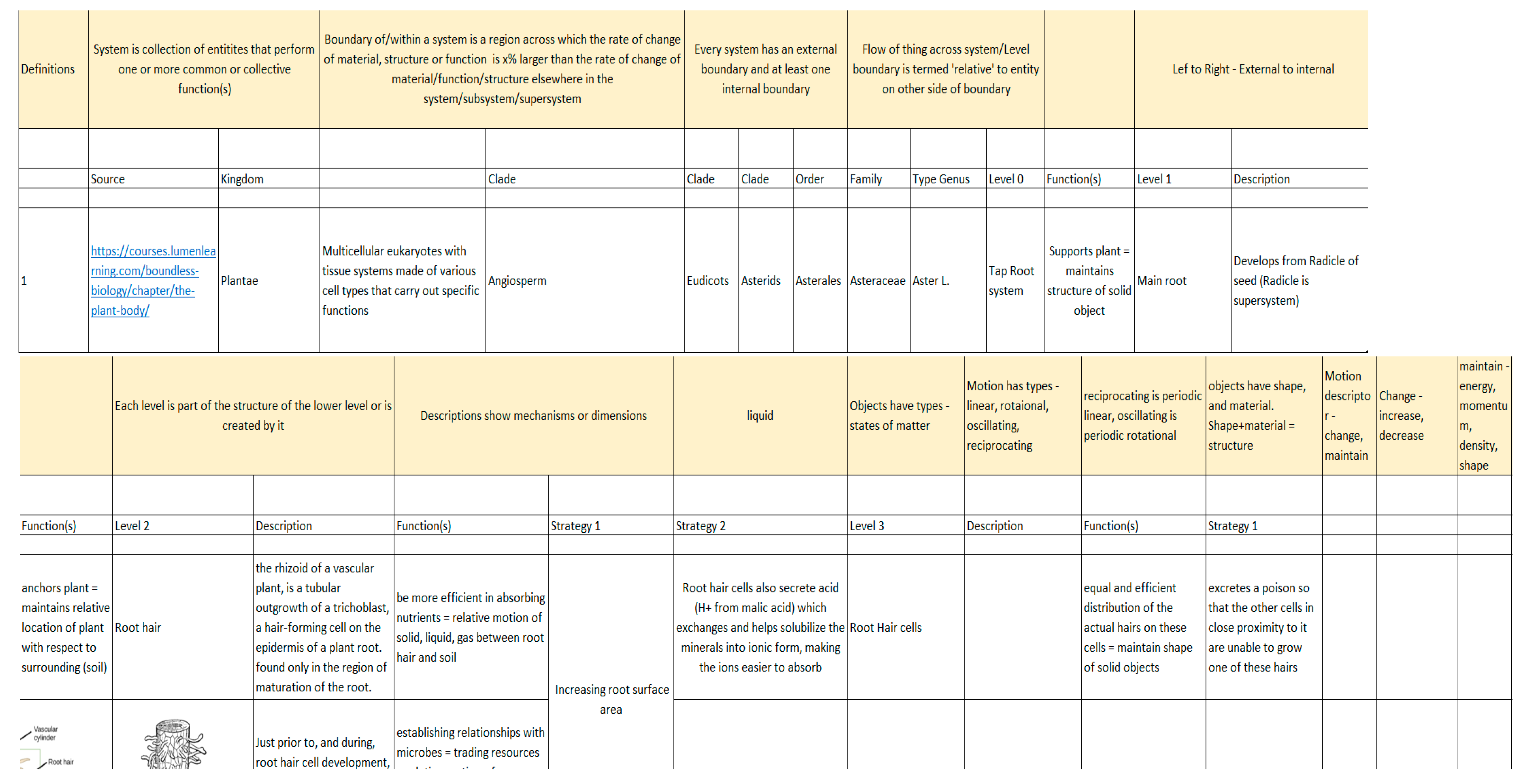
- Based on the models identified, identify the structure, pattern or mechanism associated with the specific function(s) being performed by the model (whether natural or human-made). For example, given tuna, penguins, toucan bills, falcon wings as models, we would identify that a vascular system or loop-like system should be considered with a hierarchy of diameters, branches and lengths for a set of structural, weight and heat exchange requirements.
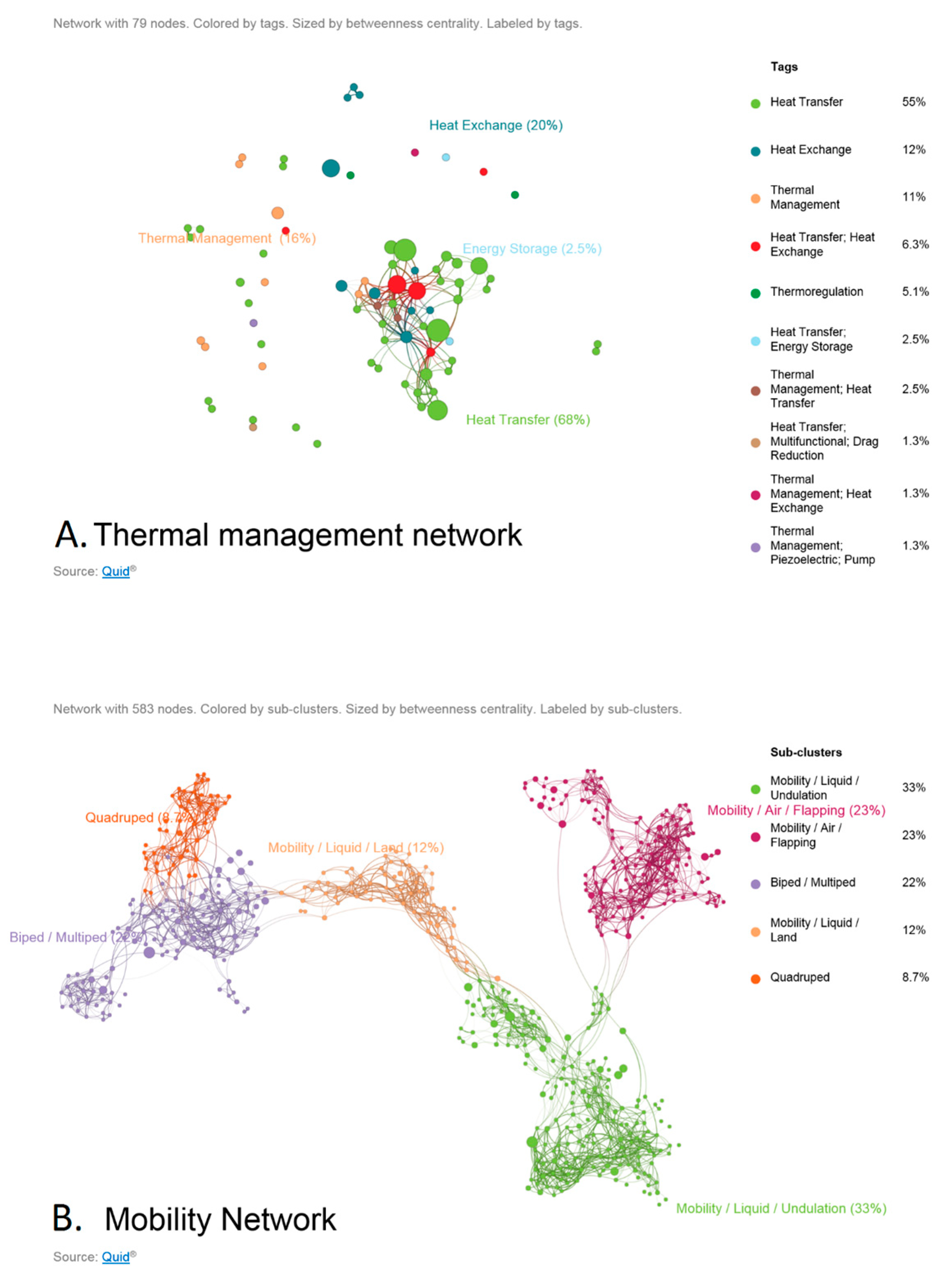
- Unsupervised learning (clustering or topic modeling for example) may be used to determine the parameter set associated with the environment we are designing for. Due to the sparse nature of most data available today, we may only be able to glean qualitative trends such as ‘more branches on the heat sink’ or more layers leads to increased heat flux. This may lead us to design a counter current heat exchanger system with parameters derived from a model that is most suitable. We may also see that certain types of thermal management solutions co-occur with a certain type of structure.
- Synthetic data sets may be generated based on the patterns found in nature to train machine learning algorithms that can provide insights into the relationship between form and multiple functions.
- Optimization may be conducted to customize the structure to meet a specific distribution of objectives.
- (d)
- Down-select by comparing metrics (weight, heat flux, structural loads) to reference architecture or goals.
- To enable broad access to the platform and development of code by all interested entities, the platform must be open and use an open source language. Python was chosen for this purpose although early development and proof of concept was carried out in R and RStudio. The platform must be accessible and a way of doing this is through a virtual environment or website. V.I.N.E. (Virtual Interchange for Nature-inspired Exploration) [42] is intended to be that virtual location and is described in subsequent sections.
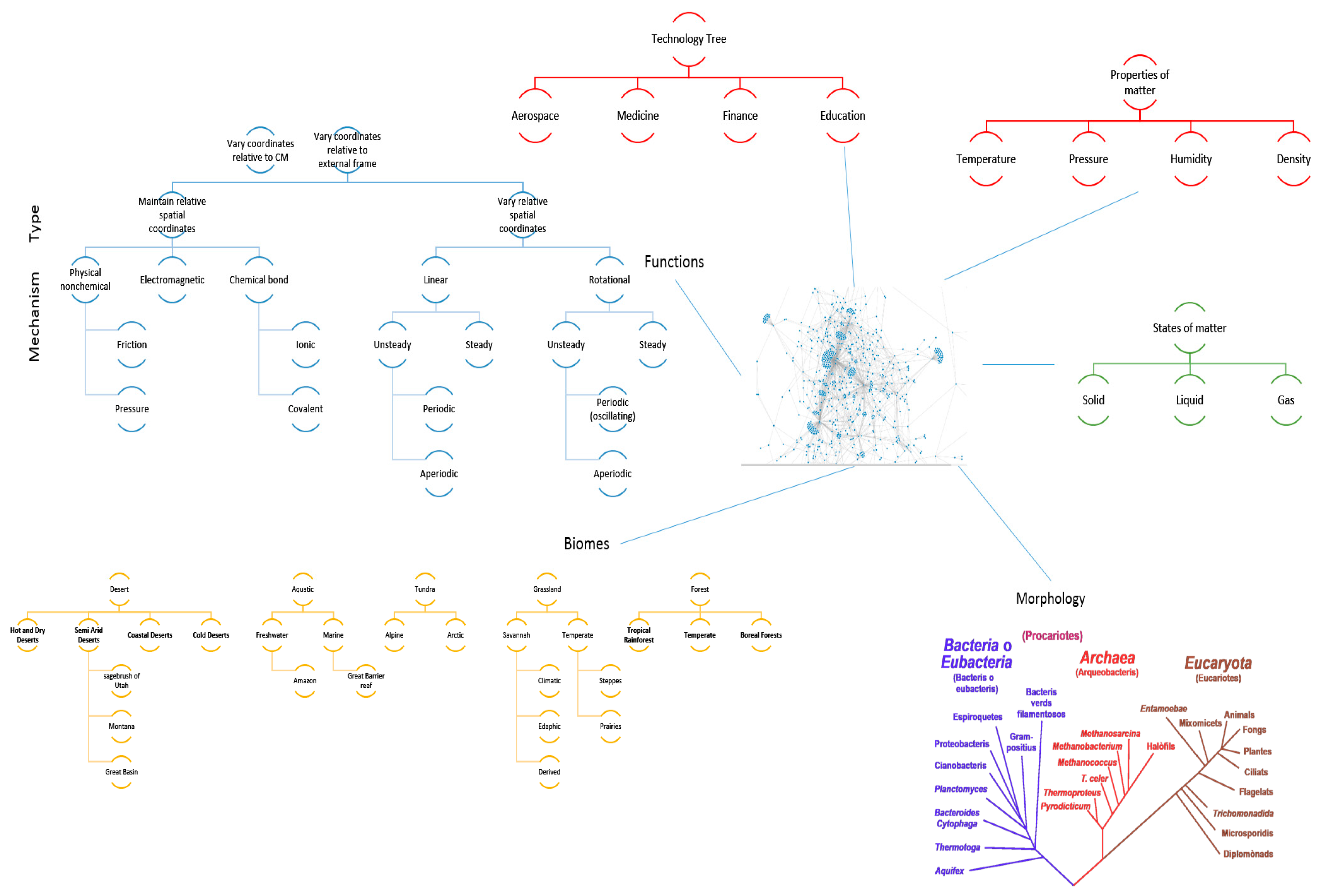
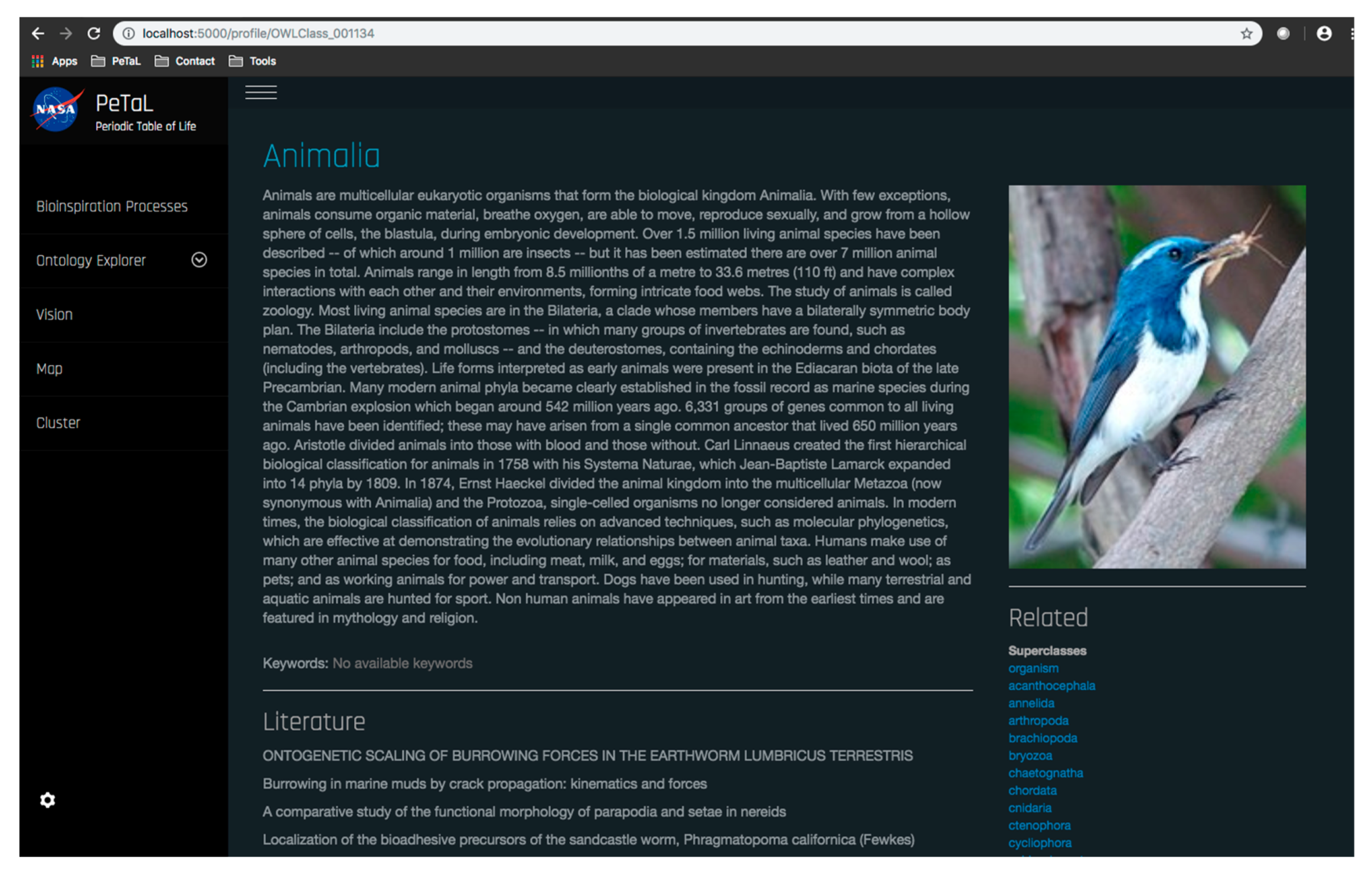
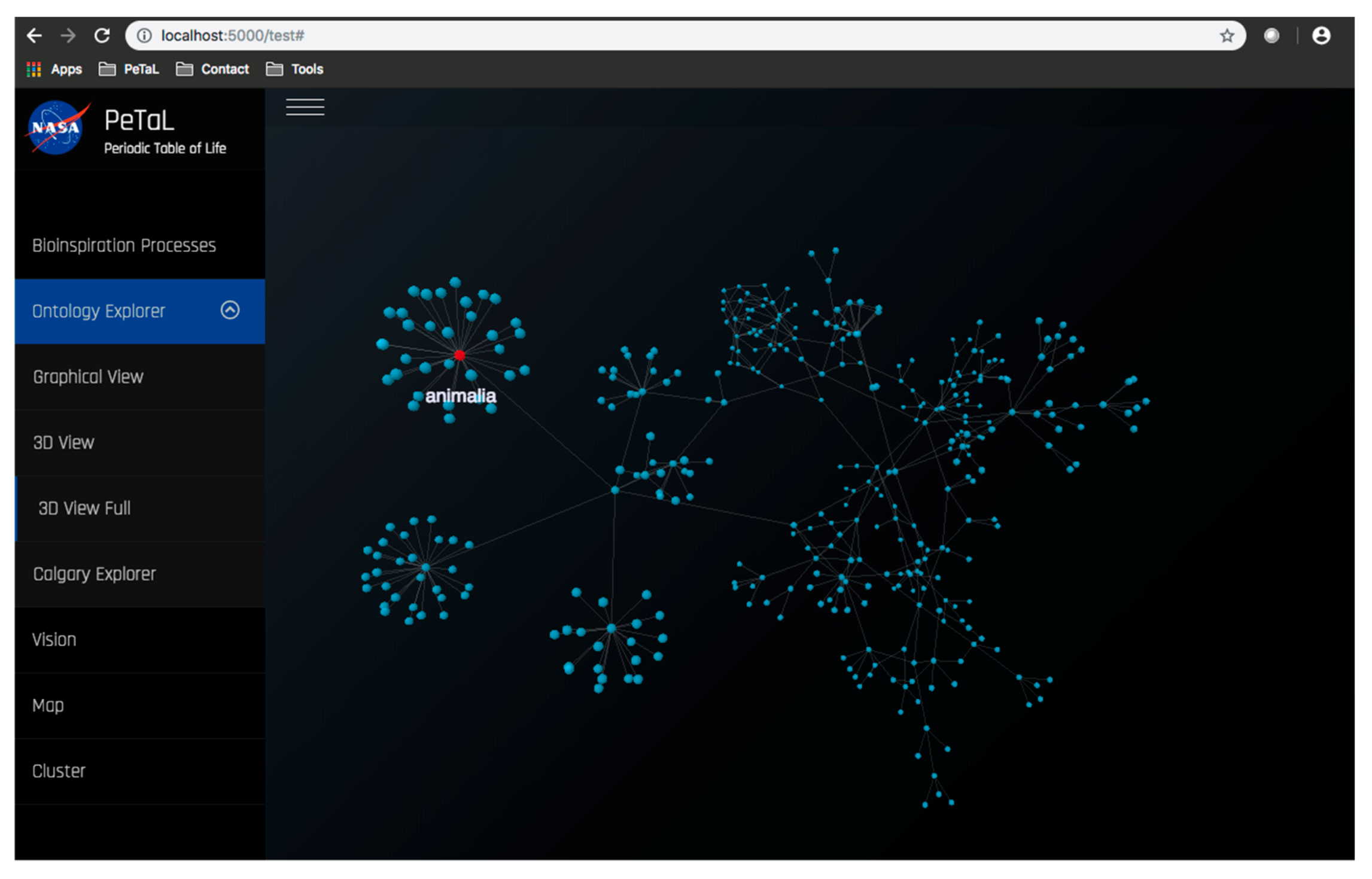
- The platform must enable new apps and tools to communicate through some common language and have some common reference of the universe. To this end an ontology is proposed that uses existing ontologies for morphology (phylogenetic tree), physics, ecosystems, and states of matter and properties of matter. The ontology must relate biological organisms to their ecosystem or habitat that includes descriptions of environment. The environment must include descriptions of states of matter such as solid, liquid or gas and these must in turn connect to properties such as temperature, pressure, density, specific heat capacity etc. The phylogenetic tree must include evolutionary history of organisms and their inner composition, down to the cellular level. This gives the ability to zoom in on any part of the organism and to consider it as a model system for analysis. Thus, the entire ontology has the ability to define ecosystems. Given a network diagram and having performed a clustering operation on this network, such ecosystems may emerge visually. Alternatively, clustering algorithms may produce similar groups in evolutionary time or in function. Figure 12 shows an example of such an ontology. The entities shown must be connected by the laws of physics (force is mass times acceleration, pressure is force per unit area etc.) A distinguishing feature from existing tools is the presence of a ‘human’ ontology. Given biological solutions such as ‘a snake, shark jaw and bat ultrasound’ inspired sensor, one must be able to identify materials that are flexible and poses the strength, finish and other qualities dictated by the biological model to realize a technological analog. This requires classification of human technology and a method of translation across domains.
- The platform must provide a common database where data related to the elements of the ontology may reside. Data may be in the form of videos, images, text, or numerical values. An unstructured database would be a suitable choice, for example, MongoDB.
- Tools must be able to communicate with the base entities of the platform (ontology, database) regardless of domain specificity or level of user expertise. Thus a natural language processing method is essential. One solution may be a thesaurus that takes queries or variables from users or user-defined tools and translates them to a form amenable to the PeTaL platform.
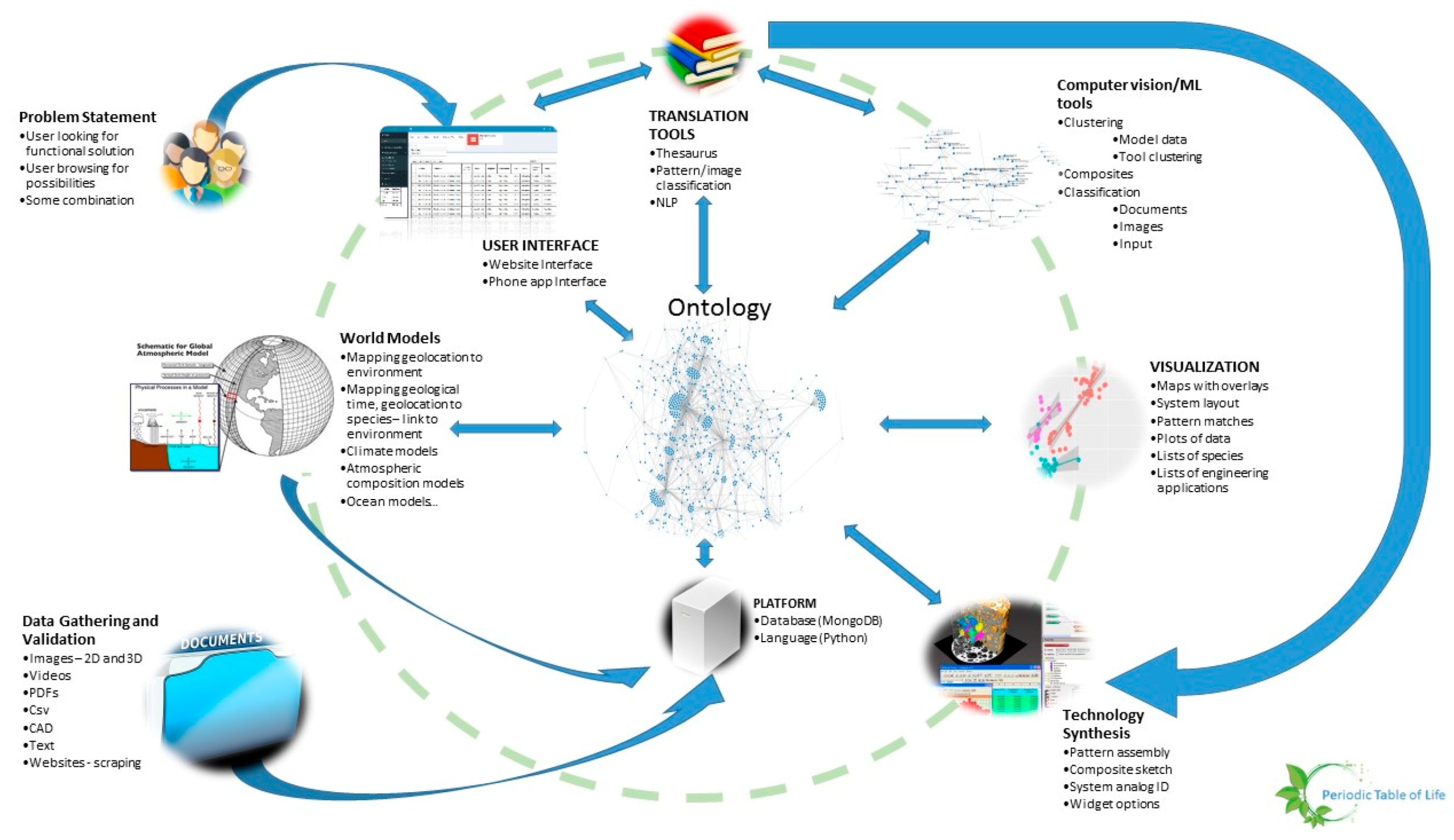
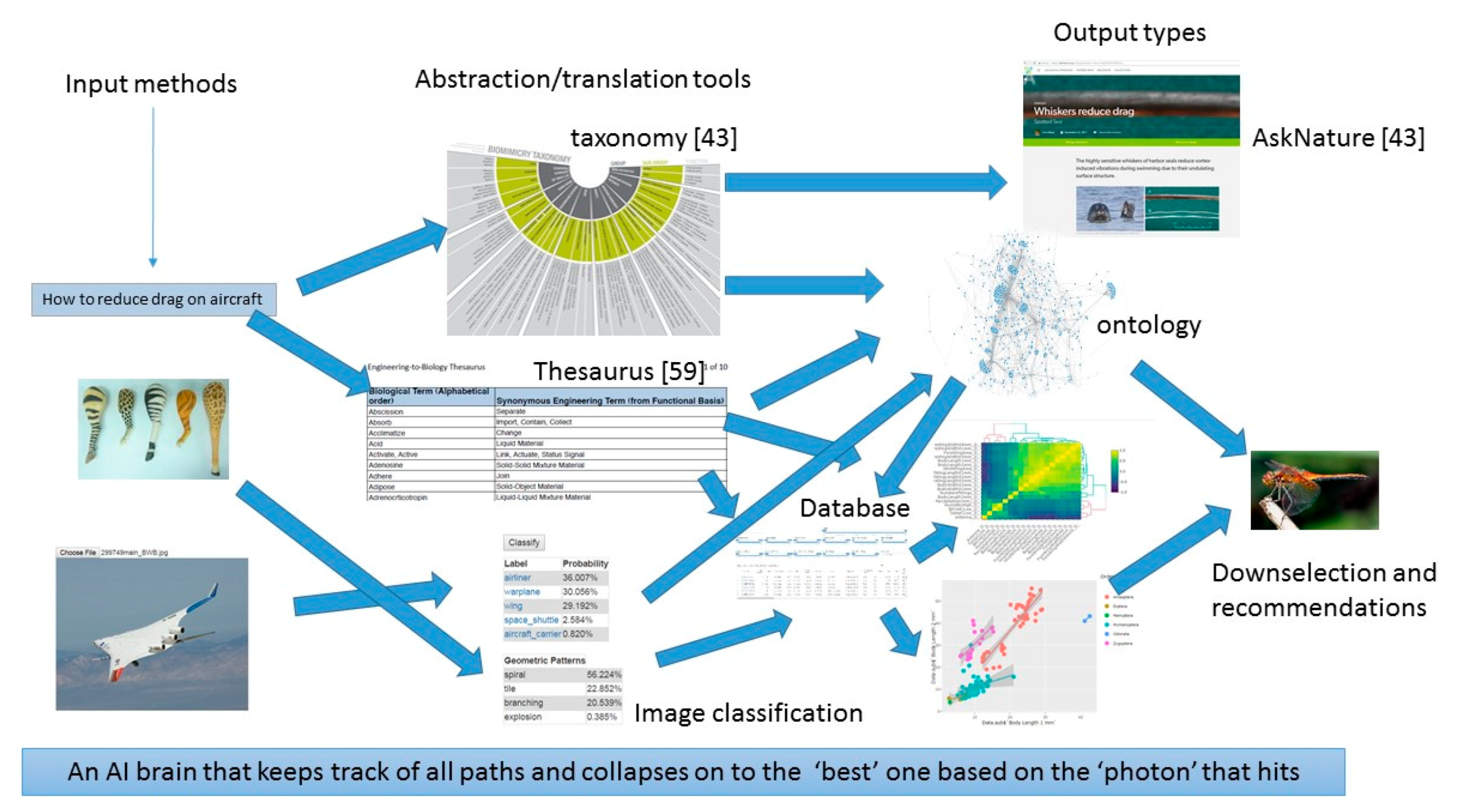
- Given a platform and a set of tools such as classification algorithms, image analysis, design or modeling tools, there should be a machine learning wrapper that navigates these tools in a manner that is likely to produce better outcomes given a particular set of user objectives, design philosophy and constraints. An example of such an algorithm is random forest, where several decision trees may be navigated depending on the set of input parameters, quality and quantity of data, time available to solve the problem, computational resources. The idea here is to mimic the evolutionary process where several mutations are permitted and the ones that are not flawed, propagate. Thus, the platform is the earth consisting of an ontology, database and language. These are themselves subject to change over time. The organisms are the tools that rely on the platform. The user problems and constraints are the forcing functions that ensure that problem solving pathways are constantly updated. Some deviations from evolution may be beneficial such as lack of extinction (storing old tools that may be shown to have a place in the toolset at a later time). Figure 13 shows the architecture of PeTaL. The primary components include:
- User Interface
- Ontology and database
- Translation Tools such as thesauri, mapping tools, abstraction tools
- Data science/machine learning/computer vision tools
- Physical models such as relationship between geolocation (including under water bodies) and temperature, pressure, density, humidity.
- Data management (validation, entry, modification)
- Technology synthesis tools (given solutions and known constraints and resources, how to construct a functioning system?)
4.2. Taxonomy
- Type of question or problem being posed
- Abstract problem: Ways to move through a fluid. Identify articles that have been classified with terms related to fluid flow and efficiency using topic modeler. A bag of words will be used for the classification based on unsupervised clustering or online dictionaries. Unsupervised clustering would be performed on articles, a subset of which contain references to fluid flow or motion.
- Well-posed problem—design a widget or change something: Reduce drag on aircraft wing. We may use existing natural language processing tools to identify the noun, verb and adverb triad (wing, drag, reduce). We associate these triads with their respective bags of words. Then, we search classified articles for biological models that match the triad bags of words.
- Well-posed problem at system level: Break into subsystems and follow process for step 1.b.
- Type of solution sought
- Background resources regarding a topic: Provide articles, websites or images based on classification results.
- Inspiration for design without a specific end goal: Use the visual graph network to enable the user to click and explore data.
- Proposed solutions given a problem: Once the problem has been abstracted and a set of possible biological models has been identified, show the data on a plot to allow the user to assess feasibility.
- Completed system solution for a high level challenge: This requires completion of a tool to create a human technology ontology and mapping from biology to technology. Supporting tools such as the ontology graph and clustering tool would be common between biological entities and human technology entities. This would require the abstraction of biological systems to enable their comparison to human systems perhaps using the taxonomy in Table 5 in addition to knowledge of degrees of freedom.
- Format of question
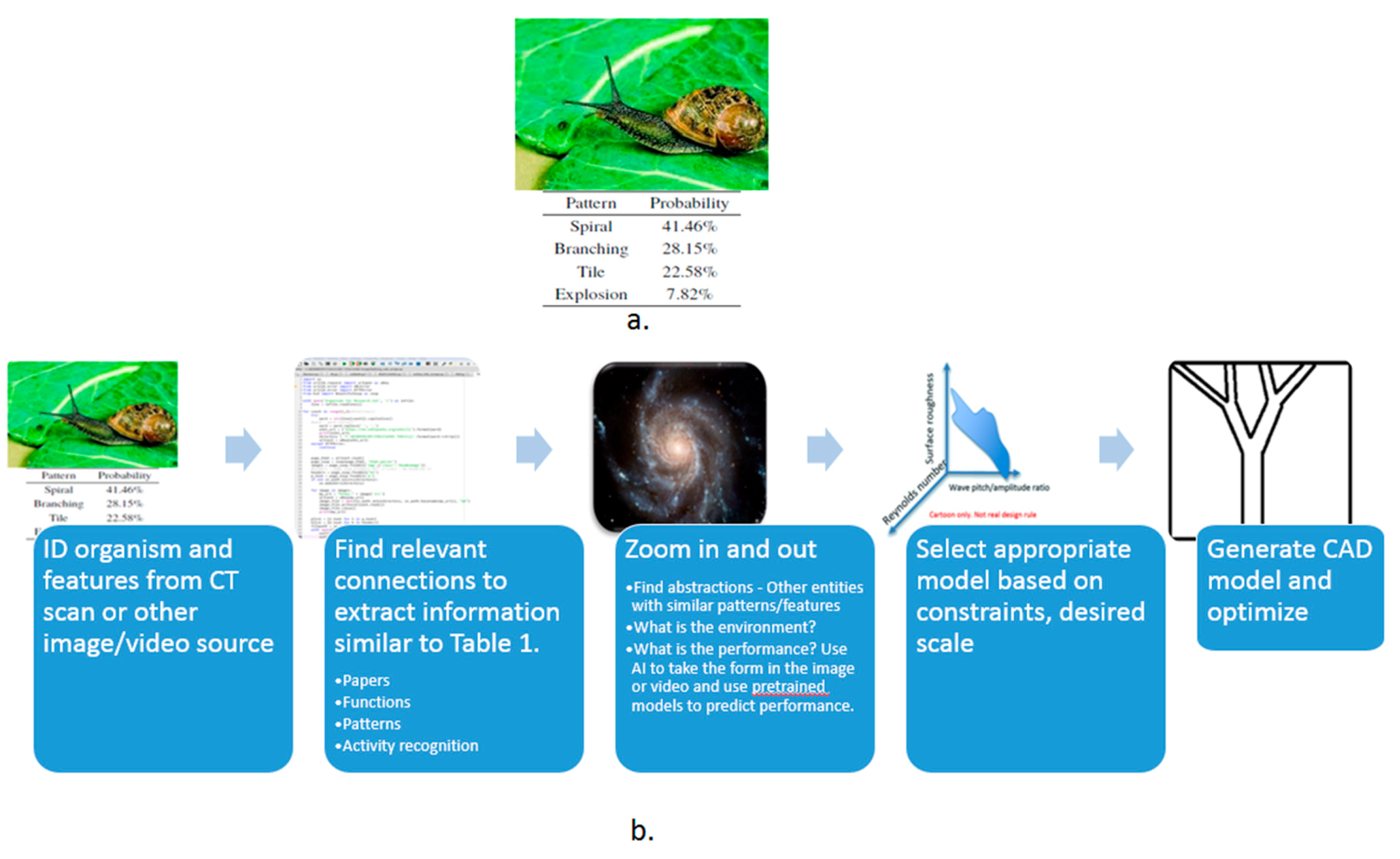
- Image: The user may submit a picture of an organism or a man-made entity. Computer vision would be used to identify the primary pattern in the image, identify the entity or organism and the environment of the entity. Feature recognition would be valuable. Once identified, the data in PeTaL that is tagged with the features, patterns or organism would be available. Similar patterns may be suggested for inspiration or functions associated with the pattern may be presented as a suggestion.
- Video: Use of activity recognition would enable explanation of mechanisms and their relationship to functions and morphology.
- Text, key phrases: This relates to 1.b. and the manner of data classification and tagging.
- Amount of data available
- For Insights on patterns, features and applications beyond a single use case, a large quantity of data is required that contains sufficient connections to other entities.
- For scaling:
- Qualitative
- Clusters are sufficient and sparse data sets may be able to provide this information. For example, we know that thick, cambered airfoils generate high lift and drag compared to thin airfoils without camber.
- Quantitative
- Sufficient data required to generate statistics across scales.
- For inspiration:
- Number of articles, models, case studies available
- Soundness of NLP to identify linkage between query and model.
- Quality of data available
- Is data from respected source like journal or curated database? How do we measure this: number of citations, impact factor?
- Is data resolution sufficient?
4.3. Status of PeTaL
- a.
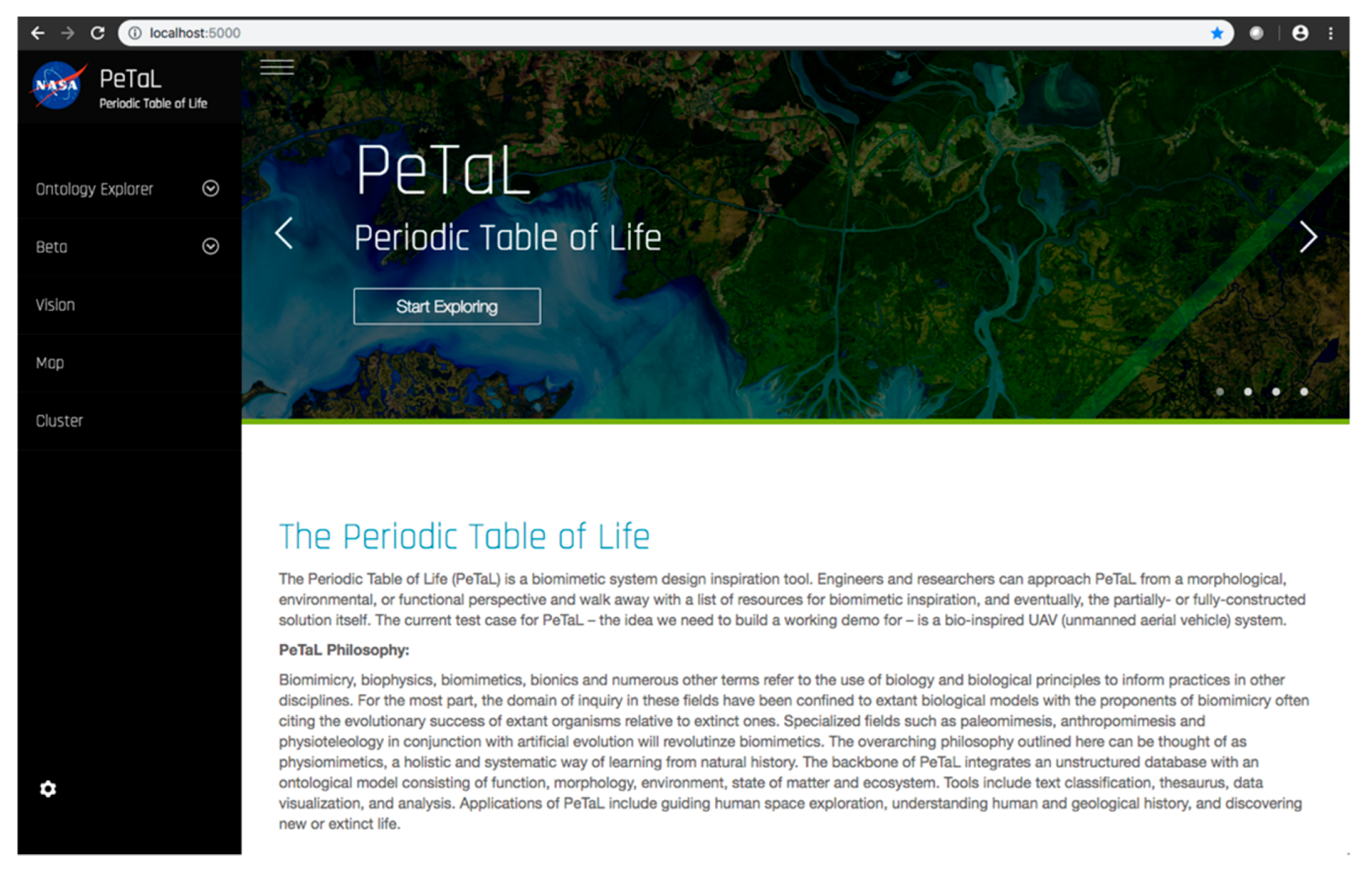
- User Interface: HTML, CSS, and JavaScript were used to create a barebones PeTaL user interface with a home, profile, map, vision, and graph page, implemented using the Bootstrap framework. The python-based home page for PeTaL is shown in Figure 24. SCSS, a superset of CSS, was introduced that allowed the writing of maintainable CSS in an object-oriented manner and enabled speeding up the process of styling elements and introducing new features and sections for the site. CSS was moved to more structured SCSS files. Doing so facilitated creating consistency and enhancing the platform with interactions, such as subtle SCSS driven scroll-triggered animations. Proper use of version control Git was key to develop new features and debugging. To reduce loading time of the application the number of HTTP requests was minimized.
- b.
- Ontology and database: The diversity and size associated with “Big Data” endeavors have enforced the adoption of non-traditional data storage systems. Thus, we have witnessed the dawn of NoSQL systems that allows for drastic increases in size over time, a concept known as scalability, as well as flexibility in formatting [60]. From this order of database systems there exists the genus of “Document-Oriented Databases” that utilize key-value systems to store data encoded in standardized language encoded documents. One particular species of note from this genus is known as MongoDB and has been widely lauded by industry and academia for its combined strengths of speed and schema-less formatting. PeTaL uses this database to store data. This database uses the Binary Javascript Object Notation (BSON) format, which is extremely similar to the popular Javascript Object Notation (JSON) format [61]. Figure 25 depicts the PeTaL database being represented in the NoSQL Booster [62]. The database has given each entry its own ObjectId. The intent is to store data gathered from researchers, citizen scientists and other databases in this format.
- c.
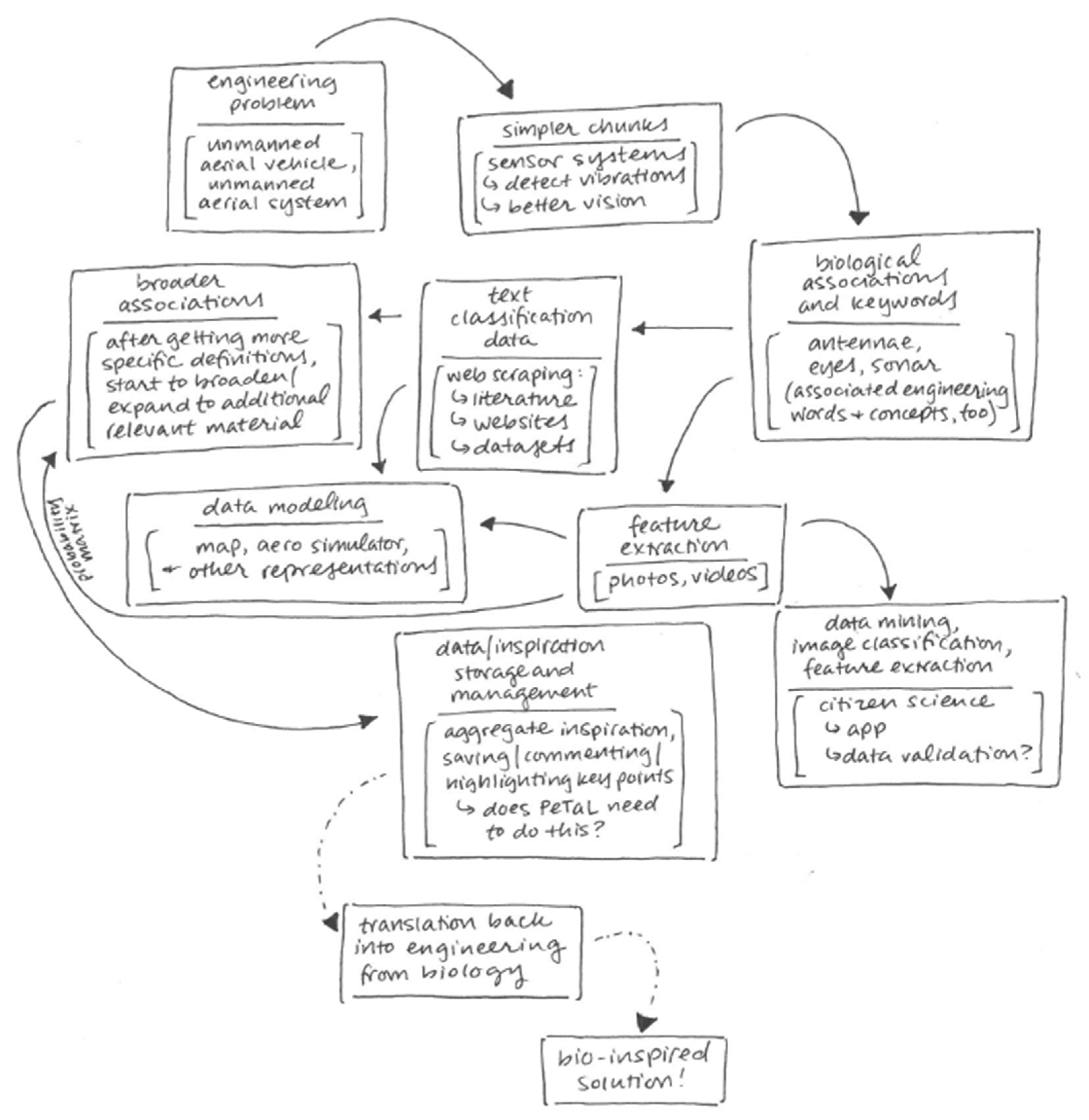
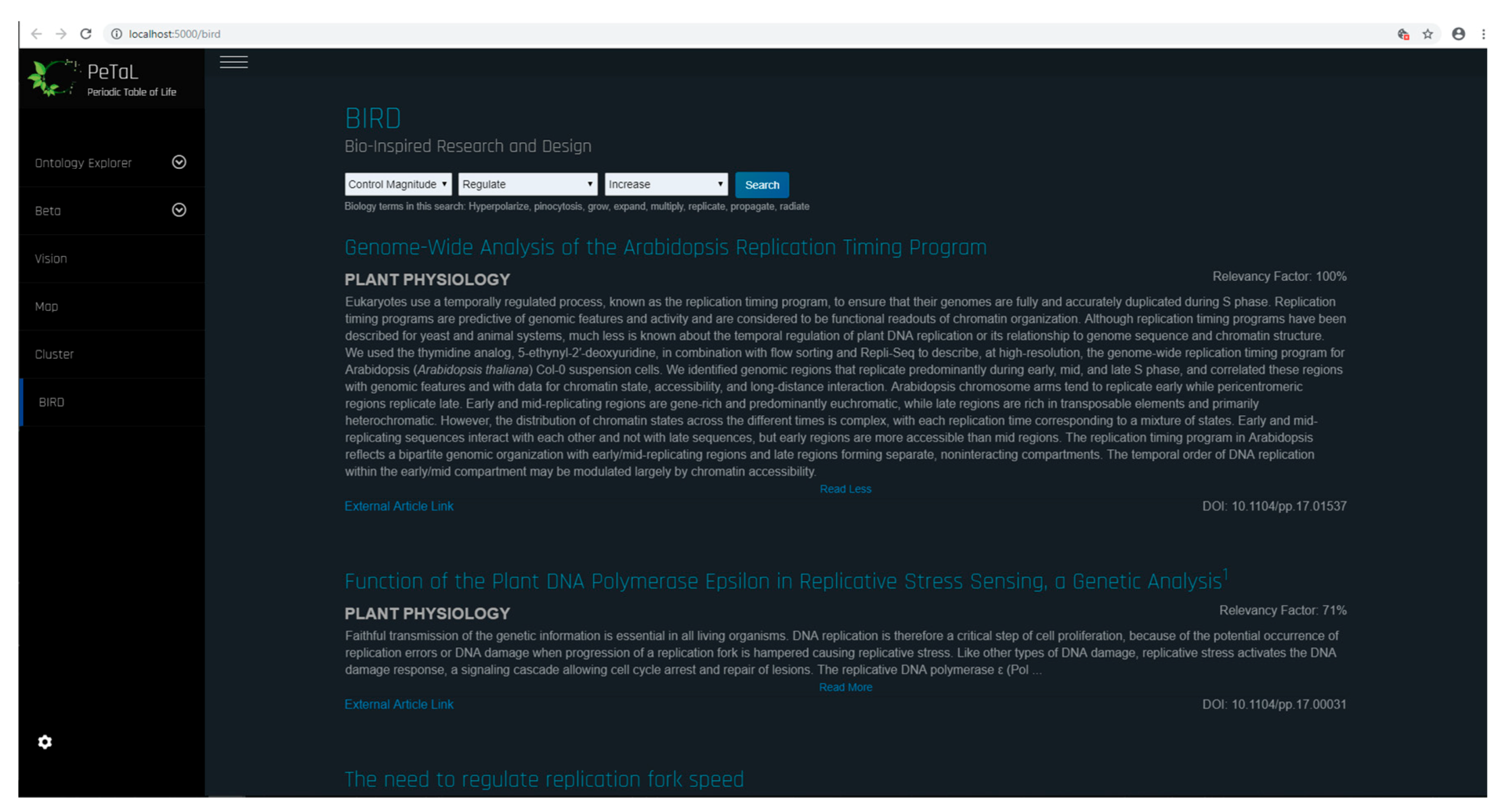
- Translation Tools such as thesauri, mapping tools, abstraction tools: Figure 28 shows the bio-inspired research and design (BIRD) tool for the PeTaL interface. This tool provides easy to use biology data engineering services for engineers. Because the translation and discover phases are often the most difficult to overcome when beginning a biomimicry project, BIRD provides specific tools to overcome these challenges. At the heart of BIRD is a JSON file that translates common engineering functions to their biology counterparts in the appropriate language. The program takes biology primary research articles and scrapes them against this list of relevant biology terms, ranks the top 50 results, and displays them back to the user. The output of this literature search can be either in the format to be human readable as shown in Figure 28, or in a format compatible for use with PeTaL’s clustering tools. Figure 29 shows the software architecture of BIRD’s back end. One of the largest benefits of using BIRD is the high degree to which a user can customize their search. Figure 29 shows the overall structure of BIRD as it is presented to future users upon download. The working directory has a folder labeled “highwire” that contains all of the tar.gz files taken from the HighWire [63] repository. Additionally, there are 3 other empty folders labeled cluster, data, and article checkpoint. The text file “journal_inputs.txt” contains a list of journal abbreviations that the user can update to add or remove journals from their search. The “e2b.json” file contains the modified engineering to biology thesaurus. Both the e2b and the journal_inputs files were designed to be highly manipulated by the user to enable a custom experience. Throughout the process, at designated checkpoints the dataclass instances are exported as pickle files into the folder “article_checkpoint” which is originally presented empty. In combine.py, the user is able to specify whether they want the output of the ranking and sorting tool to be in human and PeTaL compatible format, or in a format compatible with a clustering tool also housed in PeTaL. If the user specifies human readable output, the final text files representing the top 50 results for each engineering term are written to the “data” folder. If the user specifies the cluster compatible output, the text files are written to the “cluster” folder. Overall, combine.py will run highwire_scrapter.py and hand off the result, a list of Article dataclass instances, to article_rank.py which produces the final output. Figure 30 shows a contributed tool that uses AskNature [43] by Dr. Marjan Eggermont [64]. The intent is to continue adding tools from contributing developers. Several independent projects have already begun to combine tools so as to enable easier integration into PeTaL.
- d.
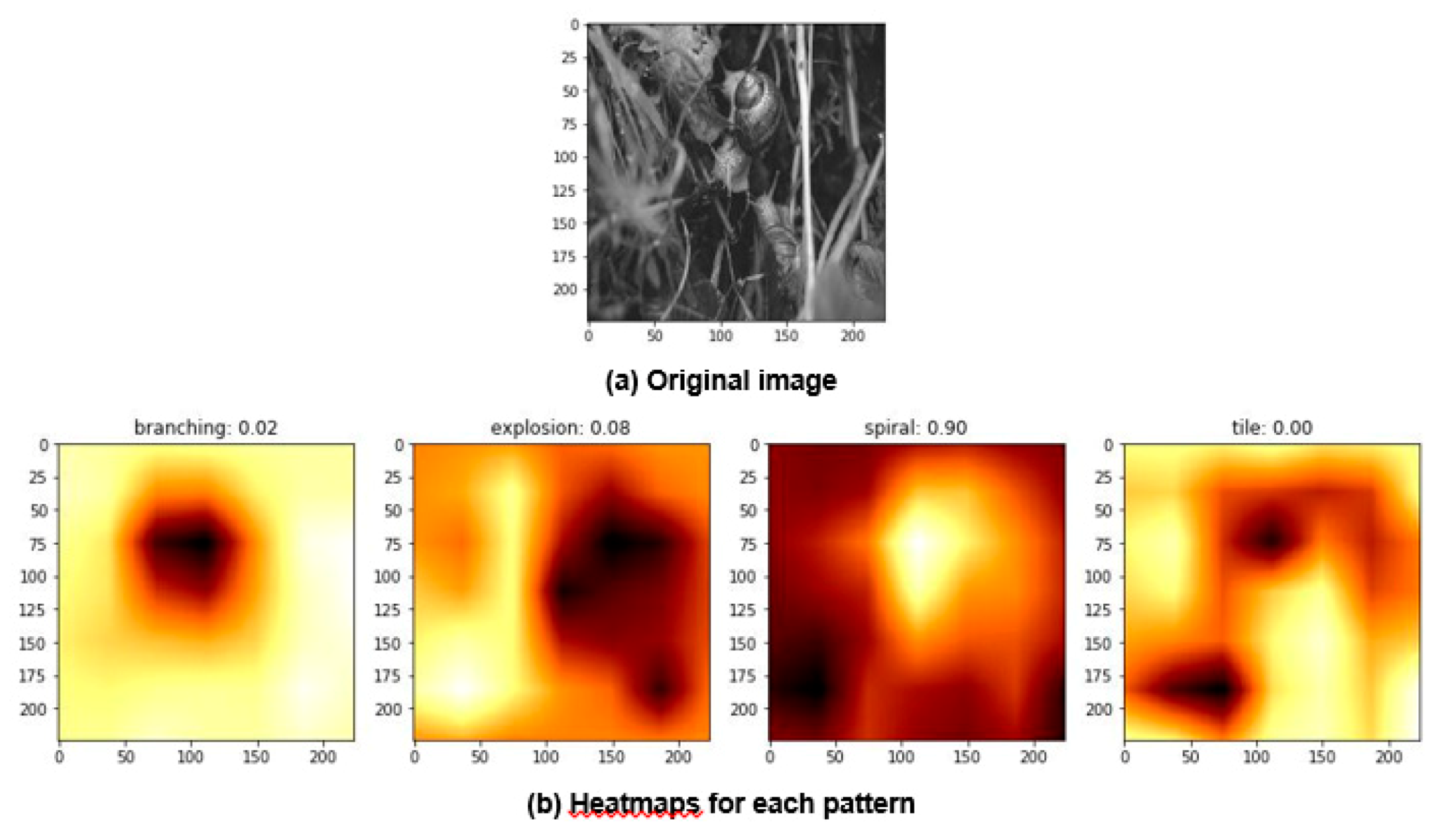
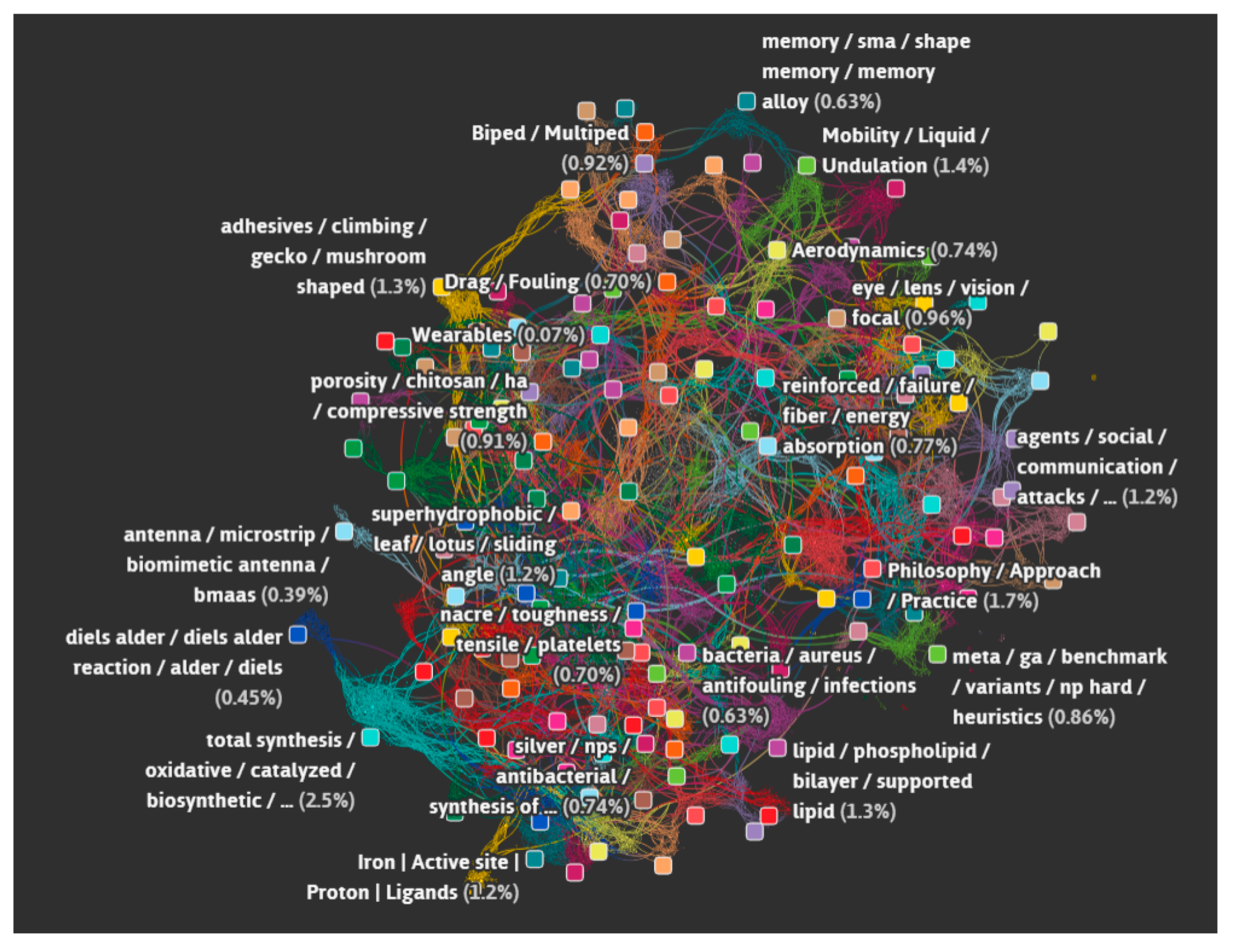
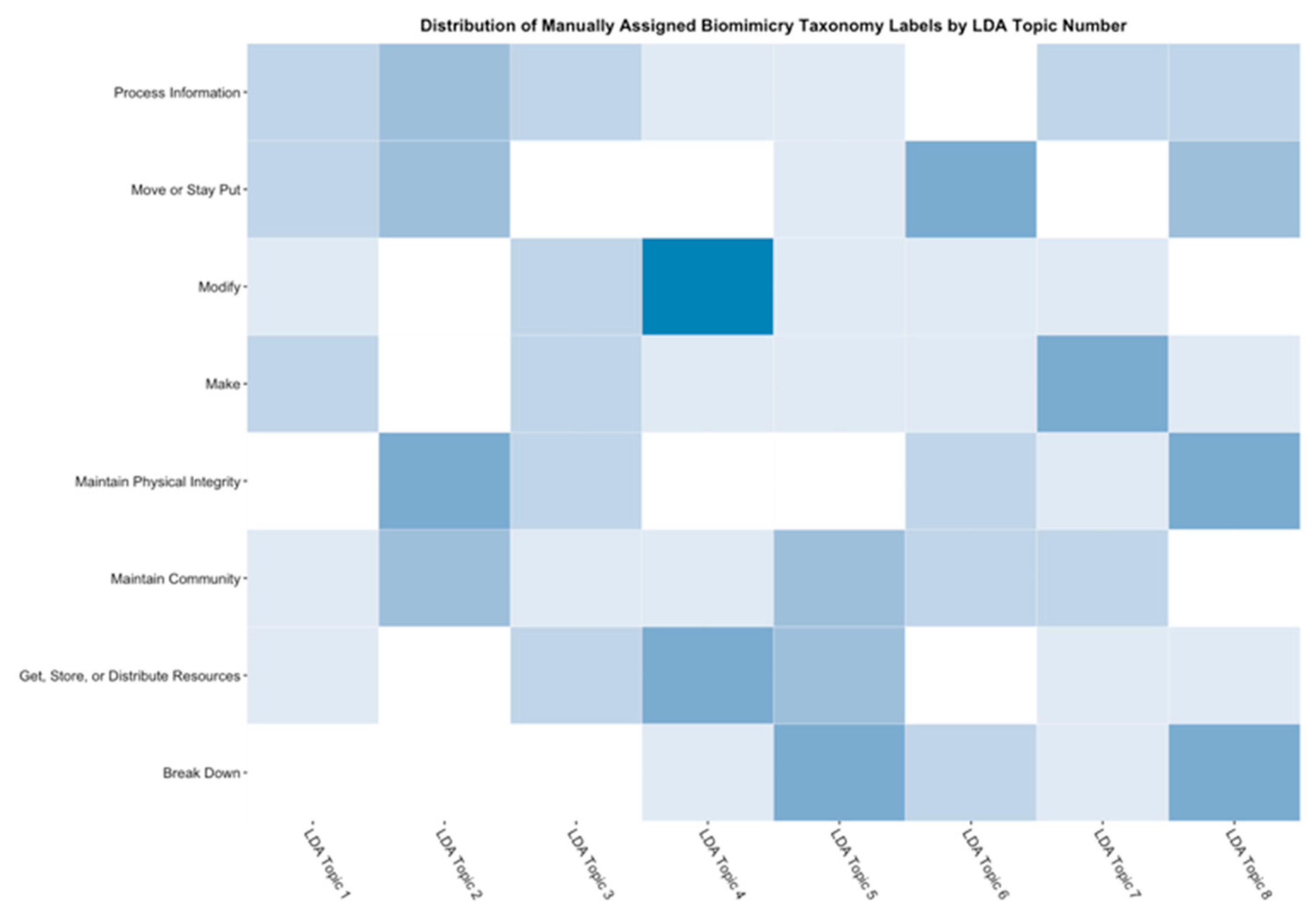
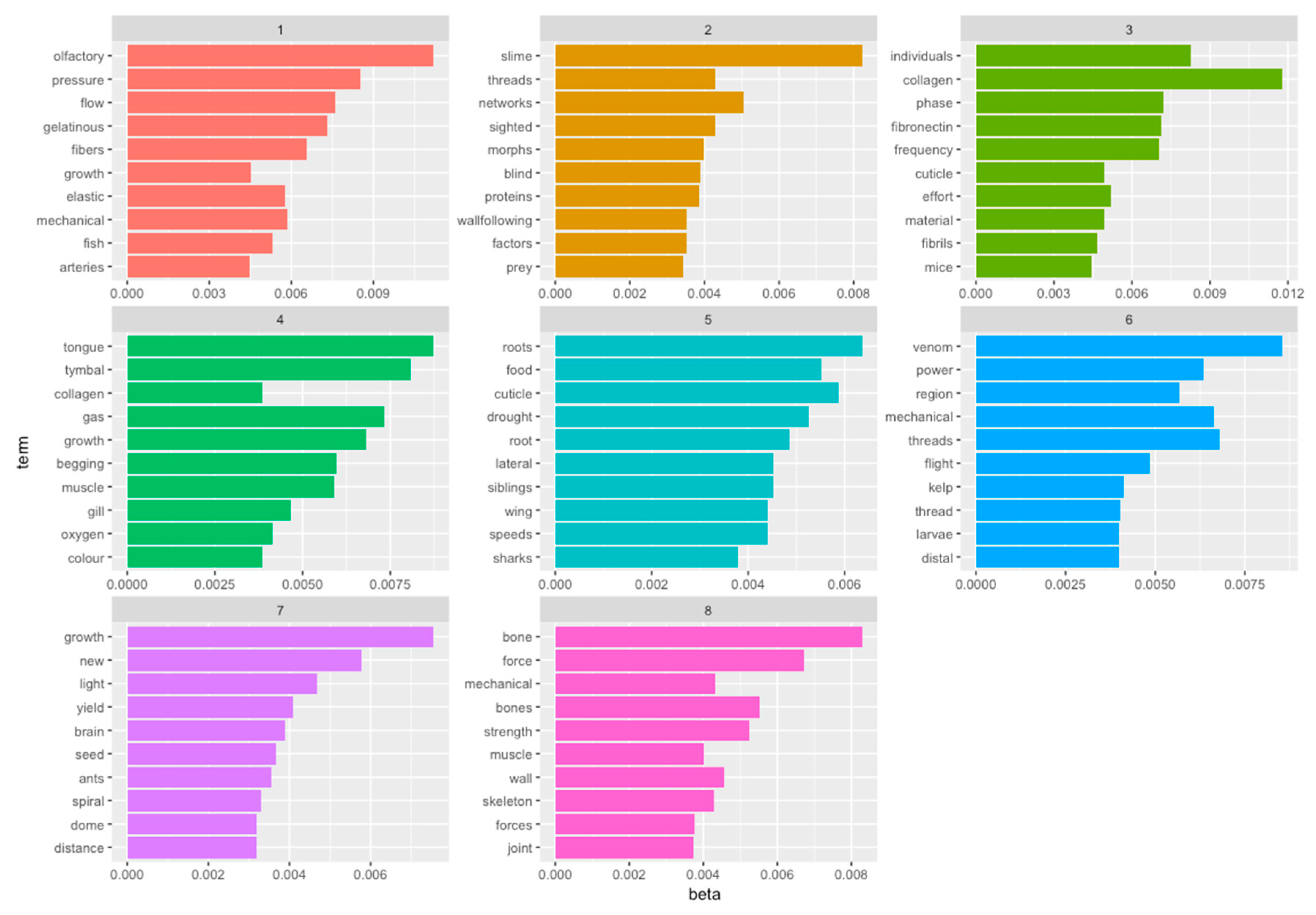
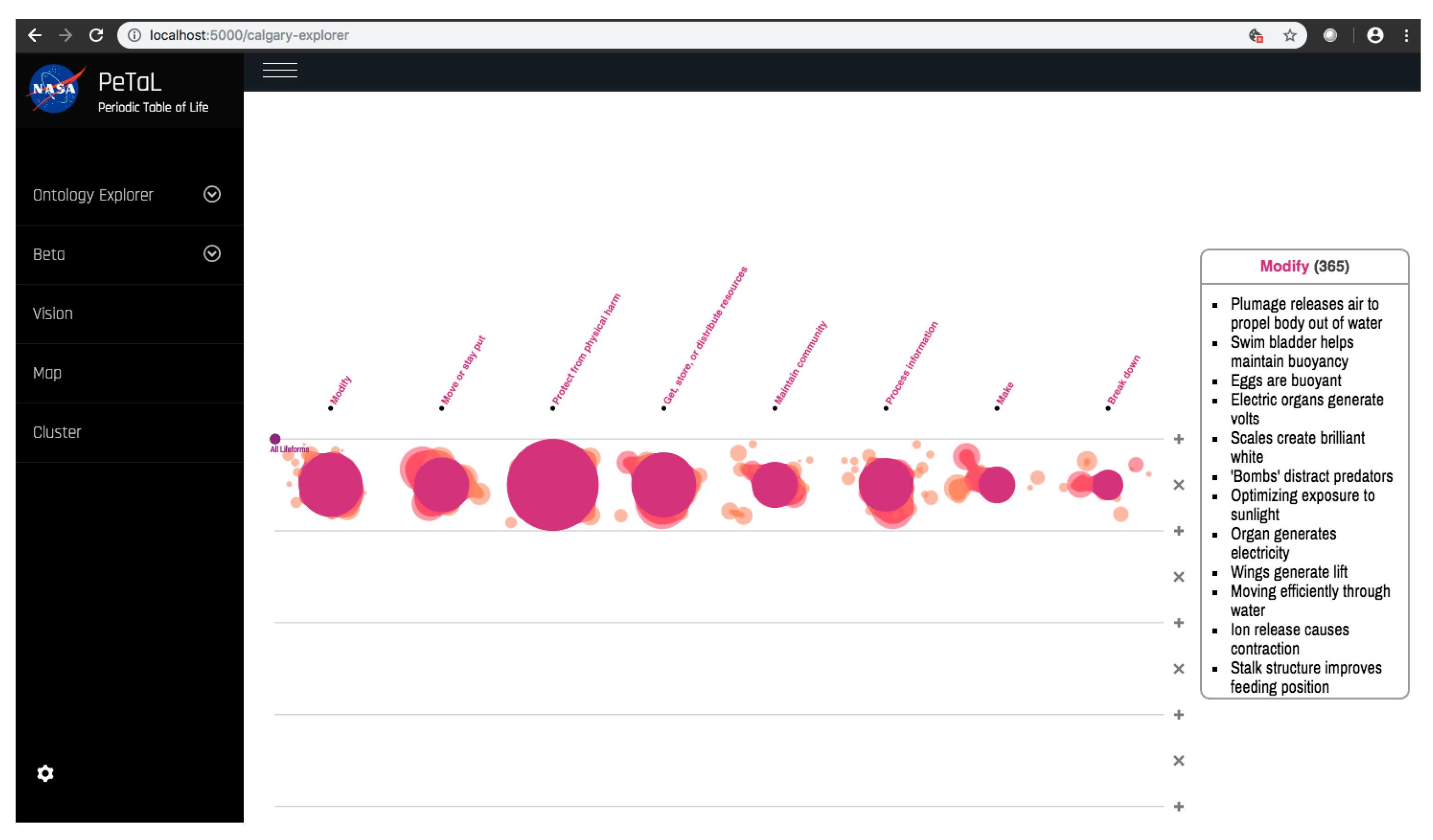
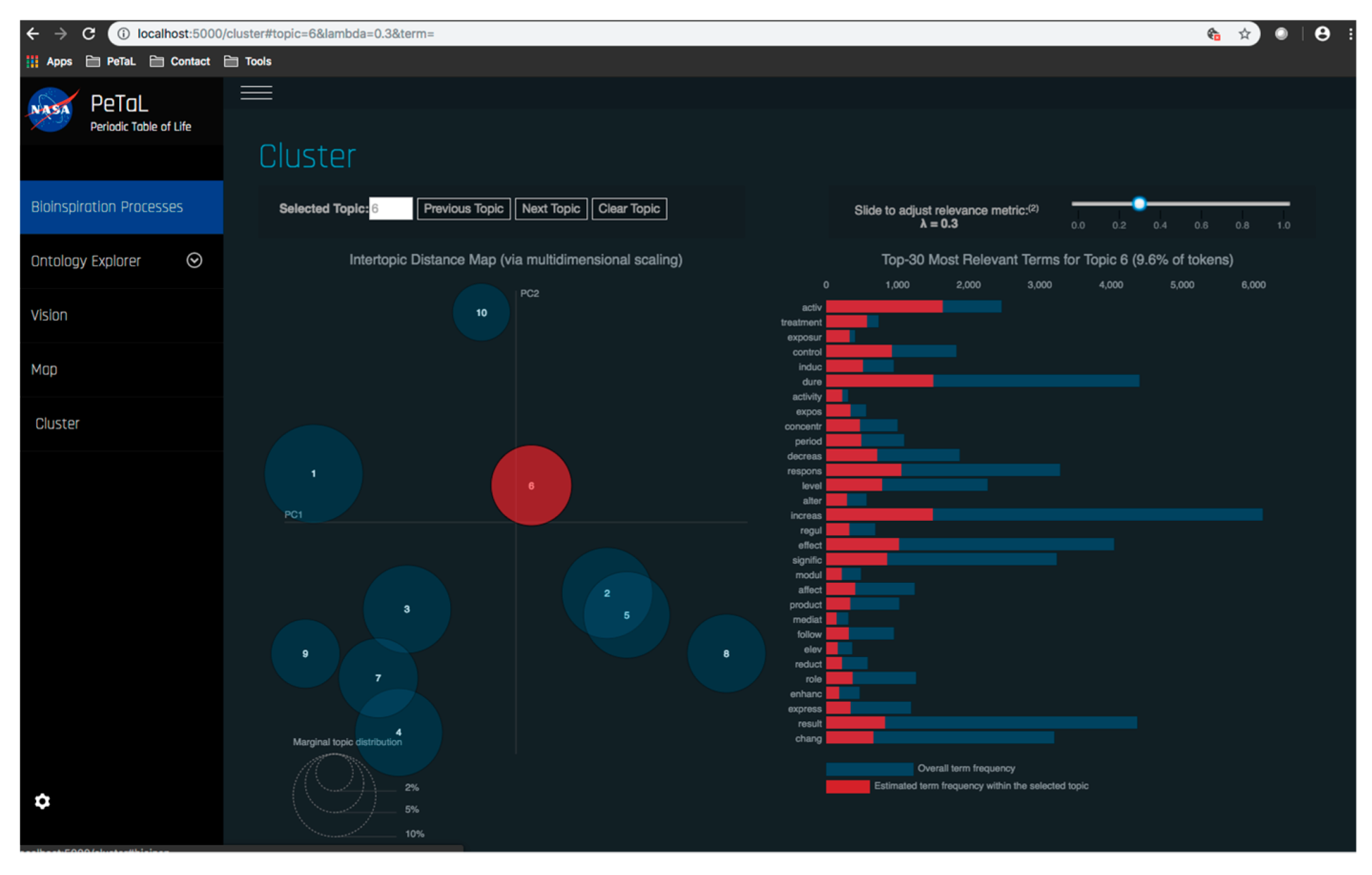
- Data science/machine learning/computer vision tools: Figure 31 shows a topic modeling tool that helps researchers identify models for inspiration in the short term and provides a method to generate automated taxonomies in the long term based on the methods that are presented in Section 4.2. Given a set of input documents (a corpus), Latent Dirichlet Allocation (LDA) and nonnegative matrix factorization (NMF) with Kullback–Leibler (KL) divergence are used to provide topic clusters to the user. Topic clusters are the latent underlying themes of a body of text. For the text modeling problem, NMF-KL is the equivalent of probabilistic latent semantic analysis. Both LDA and NMF-KL are preferred modeling tools within the topic modeling community [65,66,67,68,69]. These tools are used to identify biological specimens relevant to the user. The user can search for a cluster with relevant topics, find articles and identify the biological models and mechanisms associated with the function of interest. It is also possible to sub-cluster a given topic to learn of underlying topics. Another tool developed for PeTaL is an image classifier that determines the organism(s) and patterns present in an image. Convolutional neural networks (CNN) have demonstrated an impressive ability to learn abstract features from raw pixel information in order to classify images [70,71]. ResNet [72], pretrained on 1000 ImageNet classes, was used for organism classification. This was implemented with the Keras Applications package and no training or validation was required. The relevant output classes that pertain to organisms in PeTaL are mapped to their respective phylum page for relevant information and literature within the PeTaL platform. For geometric pattern classification, a standard convolutional neural network was trained with four convolution layers, each followed by batch normalization, ReLU non-linear activation, and max pooling. We will simply refer to this model as the CNN. A more complex CNN architecture, called MobileNet [73], was also trained. MobileNet is an image classification network designed to perform well with limited computing resources. MobileNet was chosen because it is a good compromise between classification performance and required time to train. The classifier was trained on naturally occurring patterns [74]. Examples include trees and roots for branching, dandelions for explosion, spiral galaxies and staircases for spiral, and tile roofing and giraffe spots for tile. Images were sourced from ImageNet. Images were automatically gray-scaled to avoid biasing patterns to certain colors. All images were down-sampled to 224 × 224 resolution. Figure 32 shows the image classifier workflow in PeTaL. Ongoing work includes identifying pattern locations on the images using heatmaps (Figure 33) to allow for verification of the classification tool and to also enable extraction of multiple patterns and organisms from an image to reveal connections of systems to their environment. Eventually, computed tomography (CT) scans might be available for large databases of specimen and an autonomous approach to identifying the relationship between function, morphology and pattern might be useful.
4.4. V.I.N.E. (Virtual Interchange for Nature-Inspired Exploration)
4.4.1. Goals
- Establish clusters of researchers, scientists, engineers, and subject matter experts.
- Select areas for focused biomimetic exploration and/or application.
- Establish workspaces (both traditional and virtual) where network members can connect and collaborate.
- Convene at least one annual Biomimicry Summit, at a NASA Center or other agreeable venue.
4.4.2. Objectives of each V.I.N.E. Technology Cluster
- Convene and leverage a diverse network of multi-disciplinary practitioners and experts, with the purpose of utilizing bio-inspired philosophies, tools, and research to benefit NASA, the nation, and the planet.
- Contribute standardized data to PeTaL.
- Assess current policies, plans, programs, funds, research, and technology.
- Recommend steps for future research.
- Recommend steps to ensure a viable posture for future funding.
5. Recommendations and Future Work
- A collaboration between academia and industry is vital for the success of this endeavor. Industry can supply design databases to train algorithms at the system level. These data points do not need to be state of the art.
- Blockchain and privacy preserving technology may be vital in allowing open source design tools such as PeTaL to democratize design and yet protect intellectual property. It is possible that blockchain could solve the issue of sharing proprietary data by using weights as the quantities that are bookkept and updated. Contributions would be tracked and any resulting intellectual property could be traced to contributing sources. This would allow for zoos, museums and other institutions to provide consulting services with proceeds going to research and conservation.
- Funding organizations such as the National Science Foundation (NSF), Department of Defense (DoD) and National Aeronautics and Space administration (NASA), must include the collection and curation of datasets and databases as a requirement for proposed work. We must move beyond the existing Darwin Core standards to include data such as shown in Table 1 as well as functional information described earlier in this paper.
- Work will continue on expanding PeTaL’s database, tools and applications. We project public and private contributions once the tool becomes available as an open source platform. Workshops for tool development, optimization, and integration with others and within the PeTaL framework will be held [42], along with the development of an interface through which researchers and subject matter experts may contribute data in a standardized format. Future work would include a more robust neural network architecture that is more tailored to the task of geometric pattern recognition than MobileNet. One architecture that seems promising is Microsoft’s PatternNet. It is a more generalized network that looks for repetitions in images and identifies them as patterns. After the pattern recognition, it makes use of a deConvolutional Neural Network to localize this pattern in the original image. Furthermore, it would be valuable to use a multi-headed neural network with binary classification for each individual pattern since multiple patterns can be detected in one image.
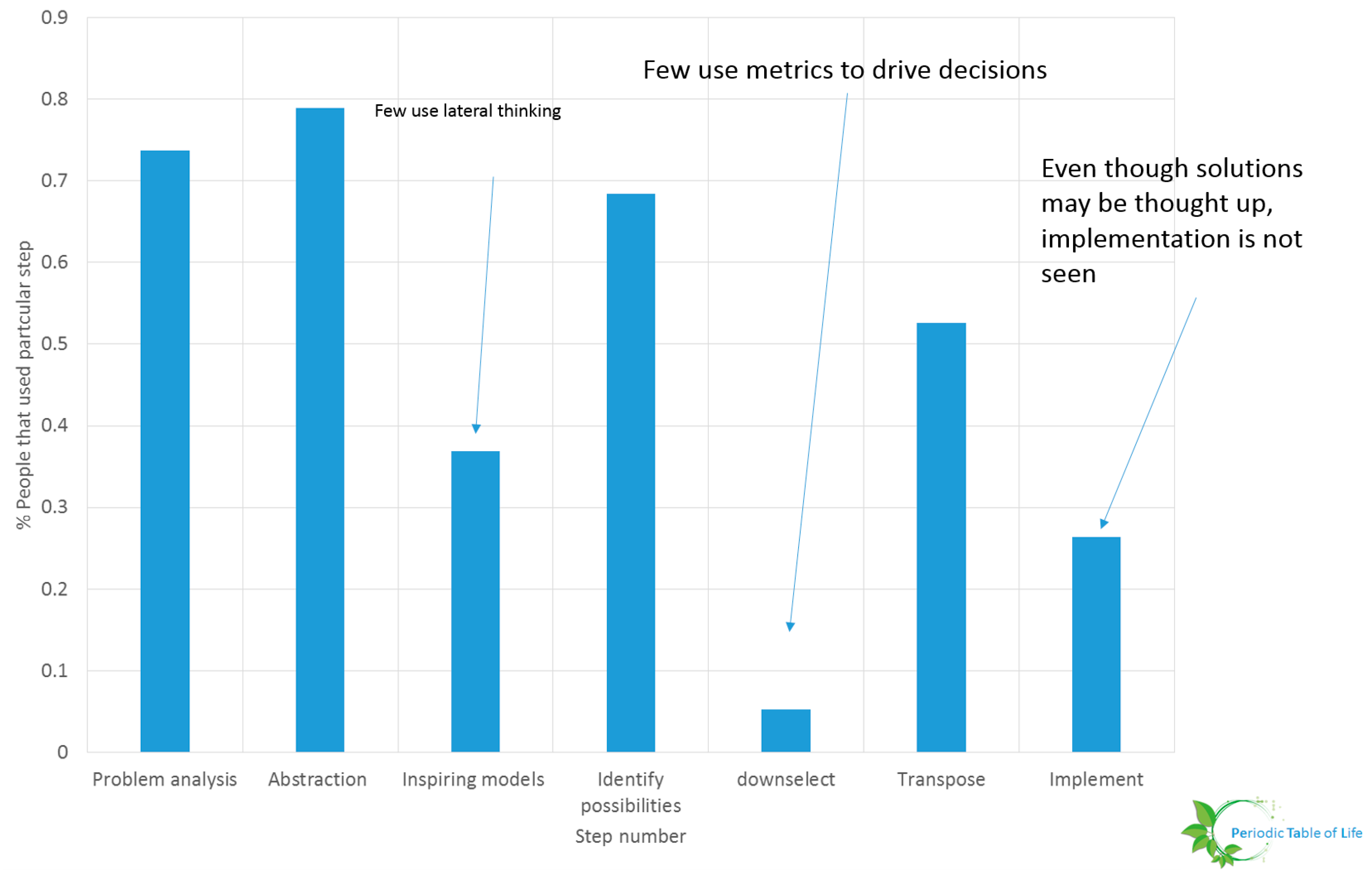
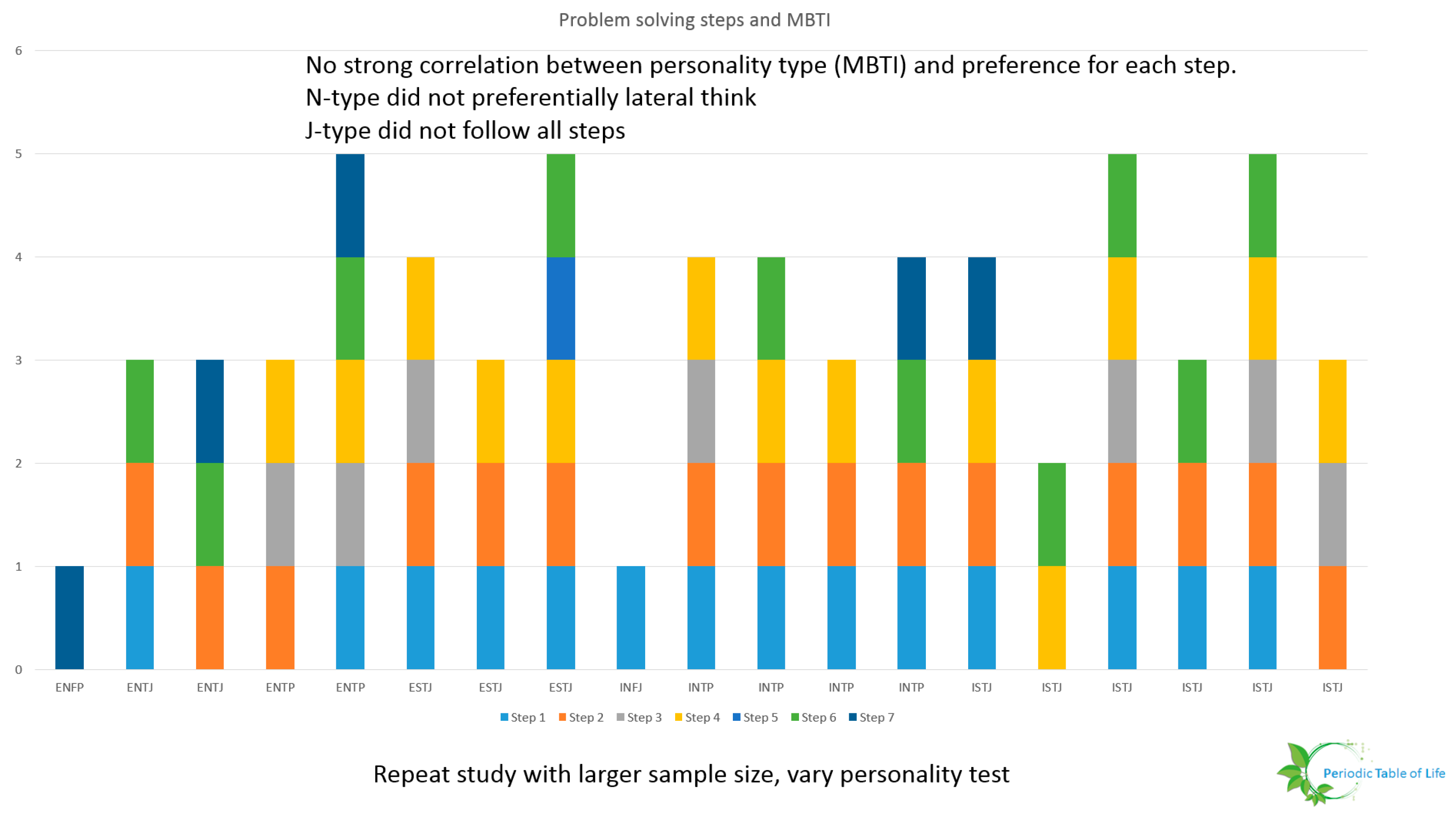
- Researchers must determine metrics and ways to evaluate design choices. A preliminary study conducted at NASA Glenn Research Center including 19 individuals form varying backgrounds (age, expertise, gender, race) showed that few individuals down select ideas. This is shown in Figure 36 and Figure 37. The participants were asked to improve the process used to procure food for their households. It is possible that the problem was too generic or that the sample size is too small. This study should be repeated. The problem solving steps of Wanieck et al. [40] were used to study the link between personality type and steps pursued. No obvious pattern emerged. One possibility is that there is no correlation between the way people solve certain problems and the way they identify their personalities. The Meyers Briggs personality test (MBTI®) was used for this study. Both the sample size and the personality test should be varied and the test repeated for a range of problems to extract meaningful insight.
- Creating a useful interactive design tool requires iteratively integrating feedback from use cases. Two preliminary demos of discovery tools on the PeTaL interface were shared with engineers at NASA Glenn Research Center to demonstrate current features and access points. Work will continue offering demos and use case opportunities to researchers in academia and industry, and other potential users.
- Several applications of this tool are either underway or planned through NASA-funded contracts as well as NASA-internal work to add data, explore probabilistic design (stochastic design) and to use machine learning algorithms to quantify the performance of systems based on images and videos.
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bai, H.; Guo, X.; Narisu, N.; Lan, T.; Wu, Q.; Xing, Y.; Zhang, Y.; Bond, S.R.; Pei, Z.; Zhang, Y.; et al. Whole-genome sequencing of 175 Mongolians uncovers population-specific genetic architecture and gene flow throughout North and East Asia. Nat. Genet. 2018, 50, 1696–1704. [Google Scholar] [CrossRef]
- Koren, S.; Rhie, A.; Walenz, B.P.; Dilthey, A.T.; Bickhart, D.M.; Kingan, S.B.; Hiendleder, S.; Williams, J.L.; Smith, T.P.L.; Phillippy, A.M. De novo assembly of haplotype-resolved genomes with trio binning. Nat. Biotechnol. 2018, 36, 1174–1182. [Google Scholar] [CrossRef]
- Anon. The future of human genome editing. Nat. Genet. 2017, 49, 653. [Google Scholar] [CrossRef]
- Luisi, P.L. The Emergence of Life: From Chemical Origins to Synthetic Biology, 2nd ed.; Cambridge University Press: Cambridge, UK, 2016. [Google Scholar]
- Goodall, J. The Chimpanzees of Gombe: Patterns of Behavior; Harvard University Press: Cambridge, MA, USA, 1986. [Google Scholar]
- Bar-Cohen, Y. Biomimetics: Mimicking and inspired-by biology. In Proceedings of the Smart Structures and Materials 2005: Electroactive Polymer Actuators and Devices (EAPAD), San Diego, CA, USA, 7–10 March 2005; Volume 5759, p. 1. [Google Scholar] [CrossRef]
- Taylor, B. Encyclopedia of Religion and Nature; Bloomsbury: London, UK, 2008; Volume 1. [Google Scholar]
- Green, T.A. Martial Arts of the World: Encyclopedia; ABC-CLIO: Santa Barbara, CA, USA, 2001; Volume 1: A-Q. [Google Scholar]
- Needham, J. Science and Civilization in China: Physics and Physical Technology, Part 1, Physics; Cambridge University Press: Cambridge, UK, 1962; Volume 4. [Google Scholar]
- White, L.T., Jr. Eilmer of Malmesbury, an Eleventh Century Aviator: A Case Study of Technological Innovation, Its Context and Tradition. Technol. Cult. 1961, 2, 97–111. [Google Scholar] [CrossRef]
- Haslam, M.; Luncz, L.V.; Staff, R.A.; Bradshaw, F.; Ottoni, E.B.; Falótico, T. Pre-Columbian monkey tools. Curr. Biol. 2016, 26, R515–R522. [Google Scholar] [CrossRef]
- Haslam, M.; Hernandez-Aguilar, A.; Ling, V.; Carvalho, S.; de la Torre, I.; DeStefano, A.; Du, A.; Hardy, B.L.; Harris, J.; Marchant, L.; et al. Primate archaeology. Nature 2009, 460, 339–344. [Google Scholar] [CrossRef]
- Eggermont, M. Biotechnik and the Bauhaus. Personal Communication, 2016. [Google Scholar]
- Gould, S.J. The exaptive excellence of spandrels as a term and prototype. Proc. Natl. Acad. Sci. USA 2015, 94, 10750–10755. [Google Scholar] [CrossRef]
- Arndt, E.M.; Moore, W.; Lee, W.-K.; Ortiz, C. Mechanistic origins of bombardier beetle (Brachinini) explosion-induced defensive spray pulsation. Science 2015, 348, 563–567. [Google Scholar] [CrossRef]
- Bourg, S.; Jacob, L.; Menu, F.; Rajon, E. How evolution draws trade-offs. bioRxiv 2017, 1–12. [Google Scholar] [CrossRef]
- Roff, D.A.; Mostowy, S.; Fairbairn, D.J. The evolution of trade-offs: Testing predictions on response to selection and environmental variation. Evolution 2002, 56, 84–95. [Google Scholar] [CrossRef]
- Weinstein, B.S. Evolutionary Trade-Offs: Emergent Constraints and Their Adaptive Consequences. Ph.D. Thesis, University of Michigan, Ann Arbor, MI, USA, 2009. [Google Scholar]
- Vincent, J.F.V. The trade-off: A central concept for biomimetics. Bioinspired Biomim. Nanobiomater. 2017, 6, 67–76. [Google Scholar] [CrossRef]
- Olson, S.L. Evolution of the rails of the South Atlantic islands (Aves: Rallidae). Smithson. Contrib. Zool. 1973, 152, 1–53. [Google Scholar] [CrossRef]
- Fishman, R.S. Evolution and the eye: The Darwin bicentennial and the sesquicentennial of the origin of species. Arch. Ophthalmol. 2008, 126, 1586–1592. [Google Scholar] [CrossRef][Green Version]
- Clune, J.; Misevic, D.; Ofria, C.; Lenski, R.E.; Elena, S.F.; Sanjuán, R. Natural Selection Fails to Optimize Mutation Rates for Long-Term Adaptation on Rugged Fitness Landscapes. PLoS Comput. Boil. 2008, 4, e1000187. [Google Scholar] [CrossRef]
- Newman, M. A Model of Mass Extinction. J. Theor. Boil. 1997, 189, 235–252. [Google Scholar] [CrossRef]
- Stein, R.W.; Mull, C.G.; Kuhn, T.S.; Aschliman, N.C.; Davidson, L.N.K.; Joy, J.B.; Smith, G.J.; Dulvy, N.K.; Mooers, A.O. Global priorities for conserving the evolutionary history of sharks, rays and chimaeras. Nat. Ecol. Evol. 2018, 2, 288–298. [Google Scholar] [CrossRef]
- Avise, J.C.; Nelson, W.S.; Sugita, H. A speciational history of “living fossils”: Molecular evolutionary patterns in horseshoe crabs. Evolution 1994, 48, 1986–2001. [Google Scholar] [CrossRef]
- ADW: Home. (n.d.). Available online: https://animaldiversity.org/ (accessed on 31 May 2019).
- Encyclopedia of Life. (n.d.). Available online: https://eol.org/ (accessed on 31 May 2019).
- “Polar Bear”, Wikipedia. (n.d.). Available online: https://en.wikipedia.org/wiki/Polar_bear (accessed on 31 May 2019).
- Simonis, P.; Rattal, M.; Oualim, E.M.; Mouhse, A.; Vigneron, J.-P. Radiative contribution to thermal conductance in animal furs and other woolly insulators. Opt. Express 2014, 22, 1940–1951. [Google Scholar] [CrossRef]
- Cui, Y.; Gong, H.; Wang, Y.; Li, D.; Bai, H. A Thermally Insulating Textile Inspired by Polar Bear Hair. Adv. Mater. 2018, 30, 1706807. [Google Scholar] [CrossRef]
- Porter, W.P.; Kearney, M. Size, shape, and the thermal niche of endotherms. Proc. Natl. Acad. Sci. USA 2009, 106, 19666–19672. [Google Scholar] [CrossRef]
- Dawson, C.; Vincent, J.F.; Jeronimidis, G.; Rice, G.; Forshaw, P. Heat Transfer through Penguin Feathers. J. Theor. Biol. 1999, 199, 291–295. [Google Scholar]
- Clarke, J.A.; Ksepka, D.T.; Salas-Gismondi, R.; Altamirano, A.J.; Shawkey, M.; D’Alba, L.; Vinther, J.; Devries, T.J.; Baby, P.; D’Alba, L. Fossil Evidence for Evolution of the Shape and Color of Penguin Feathers. Science 2010, 330, 954–957. [Google Scholar] [CrossRef]
- Norberg, U.M.L.; Norberg, R.A. Scaling of wingbeat frequency with body mass in bats and limits to maximum bat size. J. Exp. Biol. 2012, 215, 711–722. [Google Scholar] [CrossRef]
- Rian, I.M.; Sassone, M. Tree-inspired dendriforms and fractal-like branching structures in architecture: A brief historical overview. Front. Arch. Res. 2014, 3, 298–323. [Google Scholar] [CrossRef]
- Maleki, B.A. Traditional Sustainable Solutions in Iranian Desert Architecture to Solve the Energy Problem. Int. J. Tech. Phys. Probl. Eng. 2011, 3, 84–91. [Google Scholar] [CrossRef]
- Meganeura. Wikipedia. Available online: https://en.wikipedia.org/wiki/Meganeura (accessed on 13 September 2018).
- Shyam, V.; Ameri, A.; Poinsatte, P.; Thurman, D.; Wroblewski, A.; Snyder, C. Application of Pinniped Vibrissae to Aeropropulsion. In Turbo Expo: Power for Land, Sea, and Air, Volume 2A: Turbomachinery, V02AT38A023; ASME: New York, NY, USA, 2015. [Google Scholar] [CrossRef]
- Goel, A.K.; McAdams, D.A.; Stone, R.B. Biologically Inspired Design; Springer: London, UK, 2015. [Google Scholar]
- Wanieck, K.; Fayemi, P.-E.; Maranzana, N.; Zollfrank, C.; Jacobs, S. Biomimetics and its tools. Bioinspired Biomim. Nanobiomater. 2017, 6, 53–66. [Google Scholar] [CrossRef]
- Scopus Preview—Scopus—Welcome to Scopus. (n.d.). Available online: https://www.scopus.com/home.uri (accessed on 31 May 2019).
- V.I.N.E. (n.d.). Available online: https://www.grc.nasa.gov/vine/ (accessed on 31 May 2019).
- Biomimicry Institute 2016. Available online: http://www.asknature.org/ (accessed on 16 October 2017).
- Natekin, A.; Knoll, A. Gradient boosting machines, a tutorial. Front. Neurorobot. 2013, 7, 21. [Google Scholar]
- Hadi, A.S.; Kaufman, L.; Rousseeuw, P.J. Finding Groups in Data: An Introduction to Cluster Analysis. Technometrics 1992, 34, 111. [Google Scholar]
- Blei, D.M.; Ng, A.Y.; Jordan, M.I. Latent Dirichlet Allocation. J. Mach. Learn. Res. 2003, 3, 993–1022. [Google Scholar]
- Schimpf, N.G.; Matthews, P.G.D.; Wilson, R.S.; White, C.R. Cockroaches breathe discontinuously to reduce respiratory water loss. J. Exp. Biol. 2009, 212, 2773–2780. [Google Scholar] [CrossRef]
- Fudge, D.S. Composition, morphology and mechanics of hagfish slime. J. Exp. Biol. 2005, 208, 4613–4625. [Google Scholar]
- Cronin, A.L. Consensus decision making in the ant Myrmecina nipponica: house-hunters combine pheromone trails with quorum responses. Anim. Behav. 2012, 84, 1243–1251. [Google Scholar]
- Malcom, J.W. Evolution of Competitive Ability: An Adaptation Speed vs. Accuracy Tradeoff Rooted in Gene Network Size. PLoS ONE 2011, 6, e14799. [Google Scholar]
- Faury, G. Function-structure relationship of elastic arteries in evolution: from microfibrils to elastin and elastic fibres. Pathol. Biol. 2001, 49, 310–325. [Google Scholar]
- Penick, C.A.; Prager, S.S.; Liebig, J. Juvenile hormone induces queen development in late-stage larvae of the ant Harpegnathos saltator. J. Insect Physiol. 2012, 58, 1643–1649. [Google Scholar]
- Sharma, S.; Coombs, S.; Patton, P.; Perera, T.B.D. The function of wall-following behaviors in the Mexican blind cavefish and a sighted relative, the Mexican tetra (Astyanax). J. Comp. Physiol. A 2008, 195, 225–240. [Google Scholar]
- Alonso-Alvarez, C.; Pérez-Rodríguez, L.; Mateo, R.; Chastel, O.; Viñuela, J. The oxidation handicap hypothesis and the carotenoid allocation trade-off. J. Evolut. Biol. 2008, 21, 1789–1797. [Google Scholar] [CrossRef]
- Jasmin, J.-N.; Dillon, M.M.; Zeyl, C. The yield of experimental yeast populations declines during selection. Proc. R. Soc. B: Biol. Sci. 2012, 279, 4382–4388. [Google Scholar]
- Gage, J.D. Why are there so many species in deep-sea sediments? J. Exp. Mar. Biol. Ecol. 1996, 200, 257–286. [Google Scholar]
- Bunge, M. System Boundary. Int. J. Gen. Syst. 1992, 20, 215–219. [Google Scholar] [CrossRef]
- List of Systems of the Human Body–Wikipedia. (n.d.). Available online: https://en.wikipedia.org/wiki/List_of_systems_of_the_human_body (accessed on 31 May 2019).
- Nagel, J.K.S. A Thesaurus for Bioinspired Engineering Design. In Biologically Inspired Design; Goel, A., Stone, R.B., McAdams, D., Eds.; Springer: London, UK, 2014. [Google Scholar]
- Bhatia, J. NoSQL: A Panorama for Scalable Databases in Web. Int. J. Mod. Trends Eng. Res. 2017, 4, 142–147. [Google Scholar]
- Kumar, K.B.S.; Srividya; Mohanavalli, S. A performance comparison of document oriented NoSQL databases. In Proceedings of the 2017 International Conference on Computer, Communication and Signal Processing (ICCCSP), Chennai, India, 10–11 January 2017. [Google Scholar]
- NoSQLBooster—The Smartest GUI Admin Tool for MongoDB. (n.d.). Available online: https://nosqlbooster.com/ (accessed on 31 May 2019).
- HighWire Current. HighWire Press, Inc. Available online: http://highwire.stanford.edu/librarians/hwcurrent.dtl (accessed on 31 May 2019).
- Eggermont, M.; Knudsen, S.; Carpendale, S.; Pusch, R. Personal Communication, University of Calgary: Calgary, AB, Canada, 2018.
- Lee, D.; Seung, H. Algorithms for non-negative matrix factorization. In Advances in Neural Information Processing Systems; MIT Press: Cambridge, MA, USA, 2001; pp. 556–562. [Google Scholar]
- Sievert, C.; Shirley, K. LDAvis: A method for visualizing and interpreting topics. In Proceedings of the Workshop on Interactive Language Learning, Visualization, and Interfaces, Baltimore, MD, USA, 27 June 2014; pp. 63–70. [Google Scholar]
- Aggarwal, C. Machine Learning for Text, 2nd ed.; Chaps. 1, 3, 4; Springer International Publishing AG: Cham, Switzerland, 2018. [Google Scholar]
- Aggarwal, C.; Reddy, C. Data Clustering: Algorithms and Applications; Chaps. 3, 4; CRC Press: New York, NY, USA, 2014. [Google Scholar]
- Mabey, B. pyLDAvis. GitHub Repository. 2015. Available online: https://github.com/bmabey/pyLDAvis (accessed on 31 May 2019).
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolutional neural networks. In Advances in Neural Information Processing Systems; MIT Press: Cambridge, MA, USA, 2012; pp. 1097–1105. [Google Scholar]
- LeCun, Y.; Huang, F.J.; Bottou, L. Learning methods for generic object recognition with invariance to pose and lighting. In Proceedings of the 2004 IEEE Computer Society Conference on Computer Vision and Pattern Recognition, 2004. CVPR 2004, Washington, DC, USA, 27 June–2 July 2004; Volume 2, pp. 97–104. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv 2017, arXiv:1704.04861. [Google Scholar]
- na2ure. (n.d.). Available online: https://www.na2ure.com/ (accessed on 31 May 2019).
- Hymenoptera Online (HOL) > Hymenoptera. (n.d.). Available online: https://hol.osu.edu/ (accessed on 31 July 2019).













| Metric | Polar Bear Fur | Penguin Feathers | Method of Data Retrieval |
|---|---|---|---|
| Function | Insulation [28] | Containment of air [32] | NLP: Journal of experimental Biology |
| Mechanisms | Backscattering (radiation), low thermal conductivity [29] | reduce convection, radiation [32] | NLP: Journal of experimental Biology |
| Supersystem | Polar Bear | Gentoo Penguin | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Subsystem | underhair, guard hair, oil [28] | afterfeather (barbules, cilia) | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Paralel adaptations | Fat (buoyancy) [28] | Fat, trapped air (insulation, drag reduction) | NLP |
| Mass | 350–700 kg [28] | 4.5 kg–8.5 kg [26] | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Size | 182 cm to 274 cm [28] | 70 cm to 95 cm [26] | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Ecosystem Connections | Prey: Seals. Weak connections: walrus, whale carcasses, birds’ eggs, muskox, reindeer, rodents, crabs, other crustaceans and other polar bears, plants, berries, roots, and kelp. Dependencies: Foxes and gulls scavenge polar bear kills. Dependent on: abundance of ringed seals. Predators: Orcas, brown bears wolves may kill polar bear (cubs) | Prey: crustaceans, such as krill, with fish making up only about 15% of the diet. Weak interactions: fish, squat lobsters, squid. Predators: leopard seals, sea lions, and killer whales. Dependencies: Skuas and giant petrels, kelp gulls, snowy sheathbill, parasites [26,27] | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Ecosystems supported | Arctic Ring of Life, polynya, Newfoundland to James Bay [28] | Falkland Islands, South Georgia, and Kerguelen Islands; smaller populations are found on Macquarie Island, Heard Islands, South Shetland Islands, and the Antarctic Peninsula [32] | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Longevity | <500,000 years [28] | 38 Mya [32] | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Longevity of adaptation | (divergence of mammals) 220 Mya | >36 Mya, feathers 75 to 80 Mya [33] | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| Variation in genus | 4 (black, brown, polar, asian black) | 3 (brush-tailed penguins) | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| convergence | 1 | 1 | NLP |
| extinction events survived | Quaternary extinction event (74,000, 13,000 years ago). Climate related event | K-T extinction | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| ecosystem stability | Uncertain | Least concern (conservation status) [26] | Web Scraping: Animal Diversity Web, Wikipedia, Encyclopedia of Life |
| scalability | Table 1 in [31] | [32] | Data mining from technical articles |
| exaptation potential | sensory, non-hair epidermal structure, hydrophobic | Flight, mobility through air, water | Multi-label classficiation of technical articles, computer vision |
| Temperature range | −34 ℃ to −69 ℃ [28] | −30 ℃ on land, −2.2 ℃ in water [26] | Mixed |
| Speed | low (due to overheating) | 36 km/h in water [32] | NLP |
| Function related parameters | Sink temperature: −34 to −69 ℃. Source: 37 ℃. Thermal conductivity: 0.0272 W/mK, Wind speed: Low [31] | Table 1 in [32], 655ft dive depth | Data mining from technical articles.Technical databases with equations for nondimensional and scaling laws such as Graschoff number, Peclet number, Strouhal number, Knudsen number etc. |
| Function related pattern | Dendritic (branching) | networked, branched, chaotic | Computer vision |
| Hypertuned XGBoost 5-Fold Cross Validation Results | ||||
|---|---|---|---|---|
| Round No. | Training Mlogloss Mean | Training Mlogloss Std. | Testing Mlogloss Mean | Testing Mlogloss Std. |
| 35 | 0.3416 | 0.0057 | 1.846 | 0.1046 |
| Hold-Out Testing XGBoost Results | |||||||
|---|---|---|---|---|---|---|---|
| Break Down | Get, Store, or Distribute Resources | Maintain Community | Maintain Physical Integrity | Make | Modify | Move or Stay Put | Process Information |
| Cockroaches breathe discontinuously to reduce respiratory water loss [47] | |||||||
| 0.1662 | 0.1335 | 0.1153 | 0.1298 | 0.0978 | 0.1509 | 0.0984 | 0.1078 |
| Composition morphology and mechanics of hagfish slime [48] | |||||||
| 0.1006 | 0.1312 | 0.1513 | 0.1286 | 0.1003 | 0.1656 | 0.1098 | 0.1122 |
| Consensus decision making in the ant Myrmecina nipponica house hunters combine pheromone trails with quorum responses [49] | |||||||
| 0.2130 | 0.0890 | 0.1290 | 0.1235 | 0.1132 | 0.1607 | 0.0837 | 0.0874 |
| Evolution of Competitive Ability An Adaptation Speed vs Accuracy Trade off Rooted in Gene Network Size [50] | |||||||
| 0.0975 | 0.1273 | 0.1596 | 0.1307 | 0.1164 | 0.1606 | 0.0988 | 0.1087 |
| Function structure relationship of elastic arteries in evolution from microfibrils to elastin and elastic fibres [51] | |||||||
| 0.0888 | 0.0927 | 0.1031 | 0.0948 | 0.1563 | 0.2749 | 0.0902 | 0.0988 |
| Juvenile hormone induces queen development in late stage larvae of the ant Harpegnathos saltatory [52] | |||||||
| 0.0972 | 0.1356 | 0.1517 | 0.1232 | 0.1269 | 0.1600 | 0.1072 | 0.0979 |
| The function of wall following behaviors in the Mexican blind cavefish and a sighted relative the Mexican tetra Astyanax [53] | |||||||
| 0.0786 | 0.1189 | 0.1299 | 0.0956 | 0.0737 | 0.1342 | 0.0863 | 0.2824 |
| The oxidation handicap hypothesis and the carotenoid allocation trade off [54] | |||||||
| 0.1069 | 0.1272 | 0.1359 | 0.1458 | 0.1089 | 0.1779 | 0.0988 | 0.0982 |
| The yield of experimental yeast populations declines during selection [55] | |||||||
| 0.0984 | 0.1373 | 0.1353 | 0.1169 | 0.1067 | 0.1838 | 0.0997 | 0.1215 |
| Why are there so many species in deep sea sediments [56] | |||||||
| 0.1463 | 0.0936 | 0.1546 | 0.1249 | 0.1399 | 0.1465 | 0.0845 | 0.1092 |
| Abstracted Action | Action Mirror/ Disambiguation if Exists | Tertiary Terms (from E2B) | Biological Terms | Context | Mechanism of Action | Search Terms (Specific Combinations of Action, Object, Mechanism) | Asknature |
|---|---|---|---|---|---|---|---|
| Change quantity in self | Generate | increase | produce, create, spawn, form, originate, synthesize, meiosis, mitosis | increase within model | chemical, physical, electromagnetic | Generate resources chemically in a solid, generate resources chemically in a liquid, generate resources chemically in a gas, generate resources physically in a solid... | Make/Break Down |
| Destroy | divide | Division, prophase, metaphase, anaphase, cleave, cytokinesis, differentiate, branch | decrease within model | chemical, physical, electromagnetic | ... | ||
| Change boundary condition/ Interface | Attach | couple, join, link | bond, hook, interlock, join, adhere, adsorb | bring into direct association with external model | chemical, physical, electromagnetic | attach resources to solid through chemicals, attach resources to solid through physical, attach resources to solid through electromagnetic... | Stay Put |
| Detach | separate | Bleach, meiosis, abscission, mitosis, segment, electrophoresis, dialysis, denature, free, detach, release, divide, differentiate, disarticulate, branch, isolated | end direct association with external model | chemical, physical, electromagnetic | ... | Move/Break Down | |
| Change geometry of environment | Transport | move, push, pull, rotate, oscillate, remove, guide | move, push, pull, rotate, oscillate, remove, guide | Change in spatial architecture of environment relative to model | chemical, physical, electromagnetic | transport solid resources through chemicals, transport solid resources through physical, transport solid resources through electromagnetic... | Move |
| Transfer | copy, transmit | Communicate, transduce, signal, relay, reproduce, deliver | Interaction of external environment(s) through model | chemical, physical, electromagnetic | transfer resources to gas through chemicals, transfer resources to gas through physical, transfer resources to liquid through electromagnetic... | Distribute | |
| Change composition | Exchange | import, export | trade, osmosis, reflect, react, communicate, remove, symport | Interaction between model and environment resulting in composition (resource, energy, information) change | chemical, physical, electromagnetic | ... | N/A |
| Maintain configuration | Regulate | regulate | Homeostasis, cling, hold, bind, connect, stabilize, balance, adjust | Maintain internal state | chemical, physical, electromagnetic | ... | Protect, Maintain |
| Change external quantity | Promote | actuate, change | catalyze, foster, enhance, increase | Change external state | chemical, physical, electromagnetic | ... | Modify |
| Inhibit | inhibit | Camouflage, reduce, decrease | Change external state | chemical, physical, electromagnetic | ... | Modify |
| Action | Entity | Nature | Resource | Phase | Mechanism |
|---|---|---|---|---|---|
| Modify/Change | Self | Composition | Energy | Solid | Physical |
| Maintain/Keep | Other | Quantity | Mass | Liquid | Electromagnetic |
| Geometry | Gas | Chemical | |||
| Interface | Plasma |
| Technology Cluster | Catalyst(s) |
|---|---|
| Big Data, Artificial Intelligence, Machine Learing, Sensors, Robotics | Herb Schilling |
| Synthetic Biology, Artificial Evolution and Human Persistence in Space (SHIPS) | Anita Alexander Marjan Eggermont |
| Multi-functional Materials, Structures, Processing | Chris Maurer Tiffany Williams |
| In-Situ Resource Utilization (ISRU)/Hybrid and Alternative Manufacturing, Design, & Architecture (HAMDA) | Andrew Trunek Emil Reyes |
| Information, Communication, Education (ICE) | Marjan Eggermont |
| Systemology/Systems Engineering | Jacquelyn Nagel Curt McNamara |
| Photonics | Tim Peshek |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shyam, V.; Friend, L.; Whiteaker, B.; Bense, N.; Dowdall, J.; Boktor, B.; Johny, M.; Reyes, I.; Naser, A.; Sakhamuri, N.; et al. PeTaL (Periodic Table of Life) and Physiomimetics. Designs 2019, 3, 43. https://doi.org/10.3390/designs3030043
Shyam V, Friend L, Whiteaker B, Bense N, Dowdall J, Boktor B, Johny M, Reyes I, Naser A, Sakhamuri N, et al. PeTaL (Periodic Table of Life) and Physiomimetics. Designs. 2019; 3(3):43. https://doi.org/10.3390/designs3030043
Chicago/Turabian StyleShyam, Vikram, Lauren Friend, Brian Whiteaker, Nicholas Bense, Jonathan Dowdall, Bishoy Boktor, Manju Johny, Isaias Reyes, Angeera Naser, Nikhitha Sakhamuri, and et al. 2019. "PeTaL (Periodic Table of Life) and Physiomimetics" Designs 3, no. 3: 43. https://doi.org/10.3390/designs3030043
APA StyleShyam, V., Friend, L., Whiteaker, B., Bense, N., Dowdall, J., Boktor, B., Johny, M., Reyes, I., Naser, A., Sakhamuri, N., Kravets, V., Calvin, A., Gabus, K., Goodman, D., Schilling, H., Robinson, C., Reid II, R. O., & Unsworth, C. (2019). PeTaL (Periodic Table of Life) and Physiomimetics. Designs, 3(3), 43. https://doi.org/10.3390/designs3030043





