Single Cell Center of Mass for the Analysis of BMP Receptor Heterodimers Distributions
Abstract
:1. Introduction
2. Results
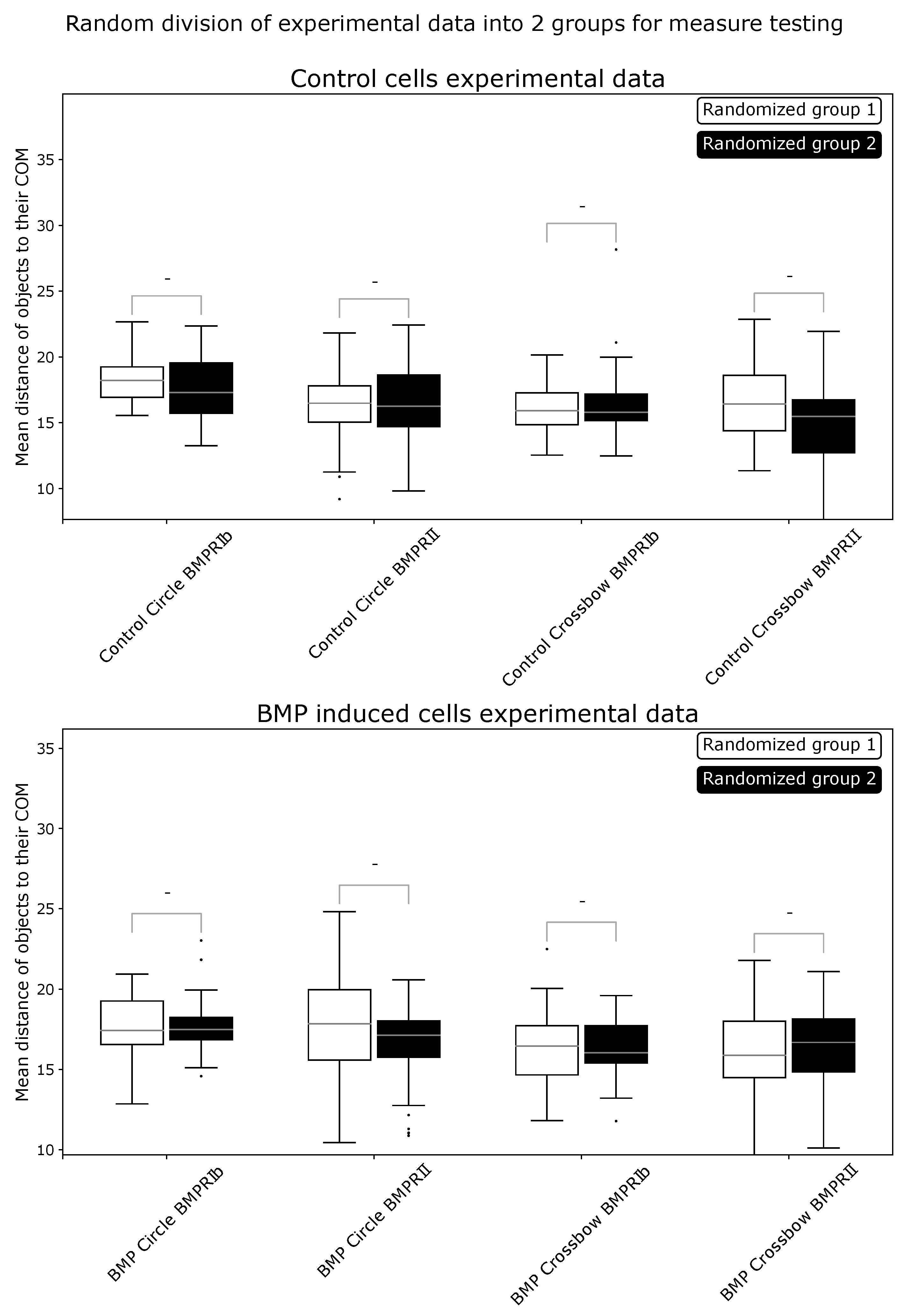
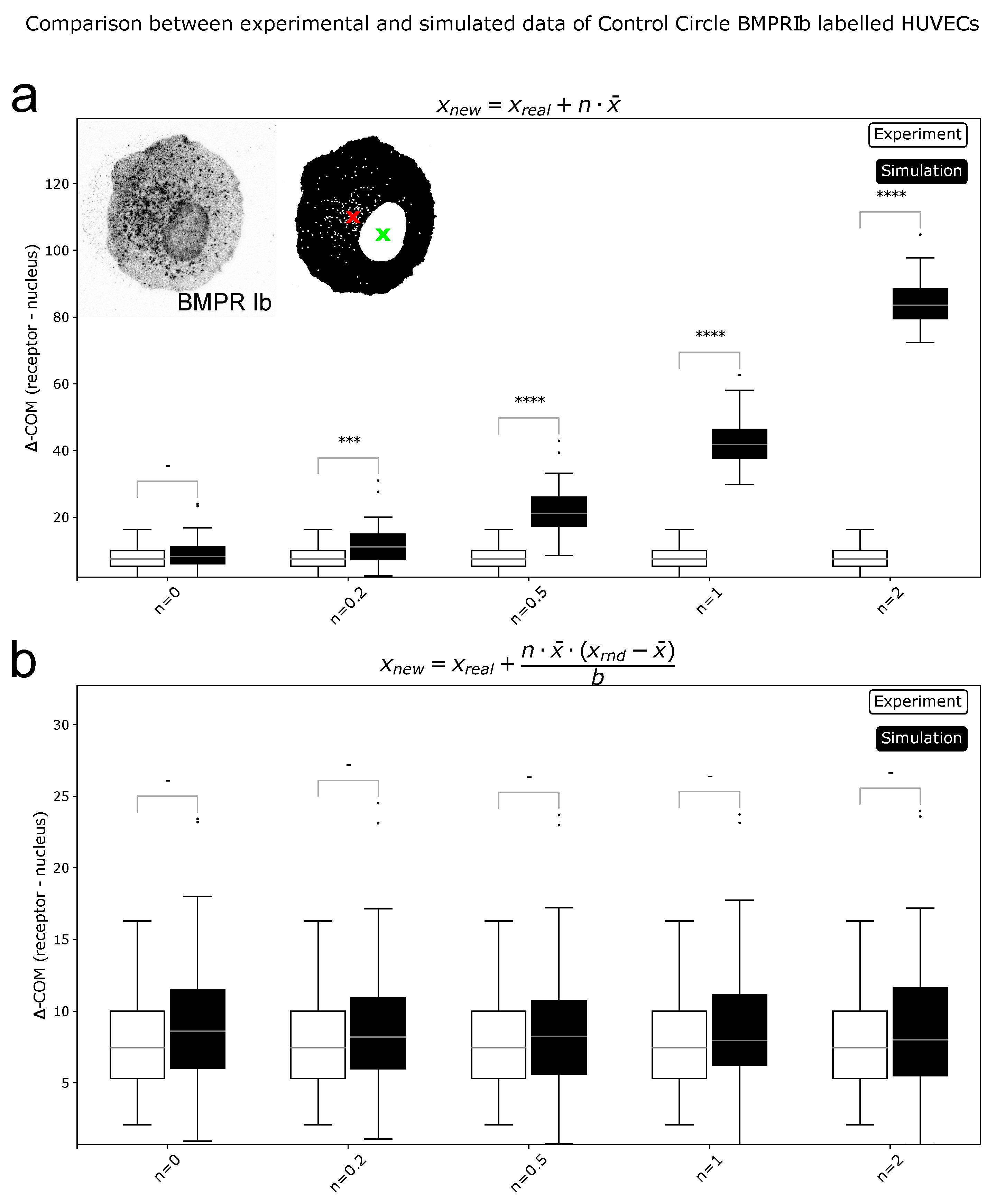
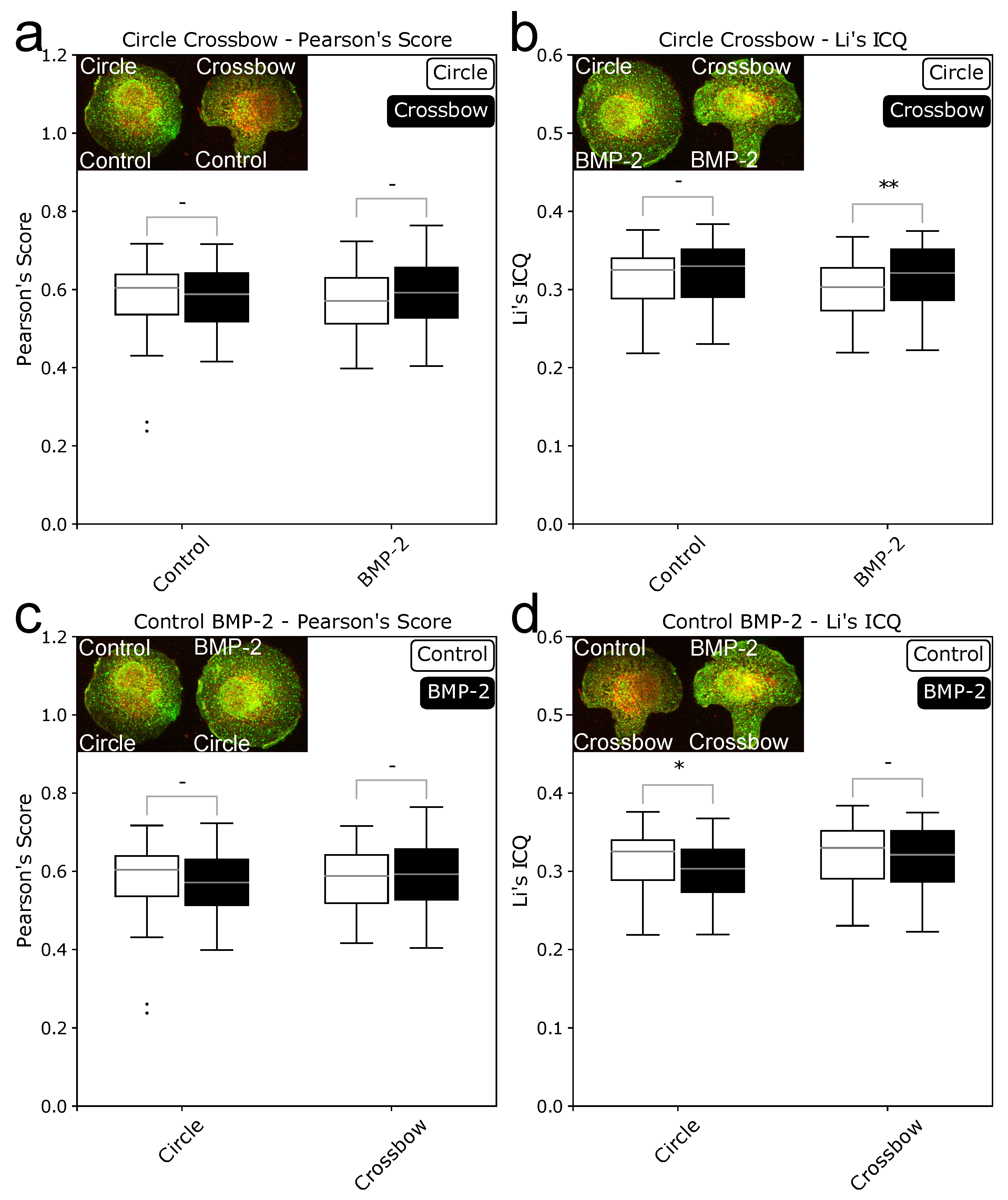
2.1. ComRed-Based Analysis Detects Changes in Experimental Data
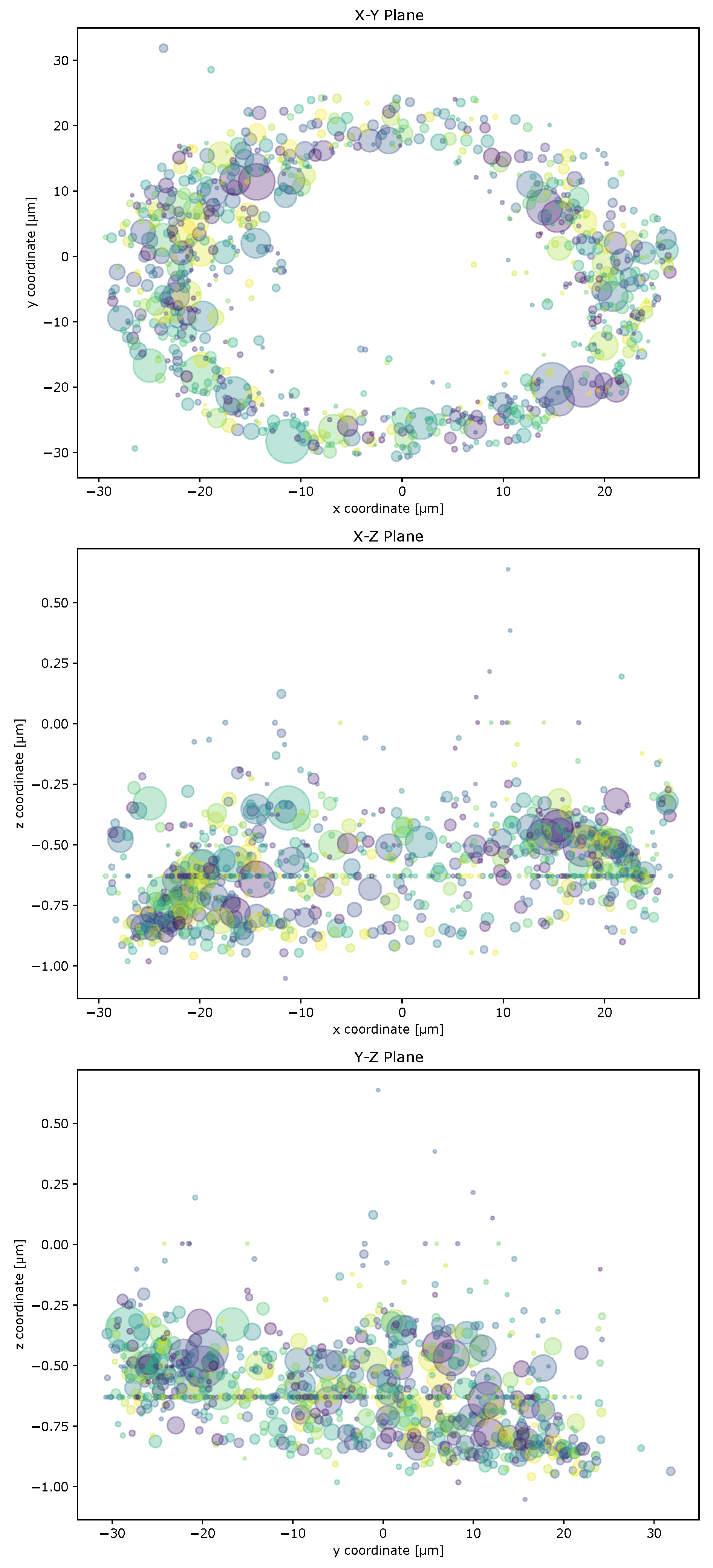
2.2. Cell Geometry Strongly Influences BMPR Subunit Distribution
3. Discussion
4. Materials and Methods
4.1. Photo-Micropatterning
4.2. Cell Culture
4.3. Immunofluorescence Staining
4.4. Microscopy
4.5. Seeding Human Umbilical Vein Endothelial Cells on Micropatterns and Induction with Bone Morphogenetic Protein 2 (BMP-2)
4.6. Image Processing and Analysis
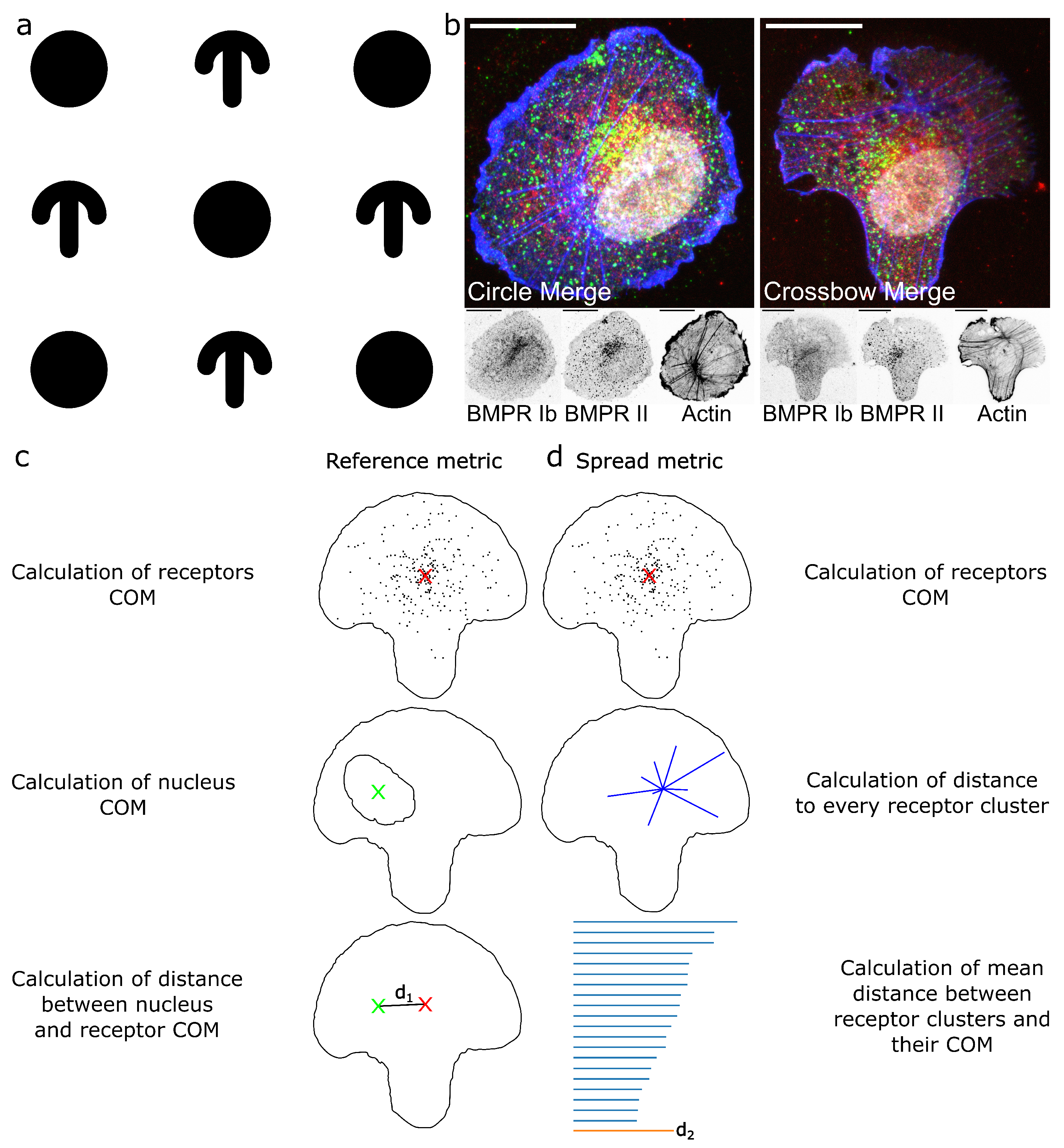
4.7. Center of Mass Calculations
4.8. Statistics
4.9. Data Simulation
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| BISC | BMP-induced signaling complex |
| BMP | Bone morphogenetic protein |
| BMPR | Bone morphogenetic protein receptor |
| ComRed | Center Of Mass REeptor Distribution |
| DAPI | 4,6-Diamidine-2-phenylindole-dihydrochloride |
| HUVEC | Human umbilical vein endothelial cell |
| PFC | Preformed complex |
Appendix A


References
- Kang, Q.; Song, W.X.; Luo, Q.; Tang, N.; Luo, J.; Luo, X.; Chen, J.; Bi, Y.; He, B.C.; Park, J.K.; et al. A comprehensive analysis of the dual roles of BMPs in regulating adipogenic and osteogenic differentiation of mesenchymal progenitor cells. Stem Cells Dev. 2009, 18, 545–559. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cunningham, N.S.; Paralkar, V.; Reddi, A.H. Osteogenin and recombinant bone morphogenetic protein 2B are chemotactic for human monocytes and stimulate transforming growth factor 81 mRNA expression. Proc. Natl. Acad. Sci. USA 1992, 89, 11740–11744. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gamell, C.; Osses, N.; Bartrons, R.; Rückle, T.; Camps, M.; Rosa, J.L.; Ventura, F. BMP2 induction of actin cytoskeleton reorganization and cell migration requires PI3-kinase and Cdc42 activity. J. Cell Sci. 2008, 23, 3960–3970. [Google Scholar] [CrossRef] [Green Version]
- Nohe, A.; Hassel, S.; Ehrlich, M.; Neubauer, F.; Sebald, W.; Henis, Y.I.; Knaus, P. The mode of bone morphogenetic protein (BMP) receptor oligomerization determines different BMP-2 signaling pathways. J. Biol. Chem. 2002, 277, 5330–5338. [Google Scholar] [CrossRef] [Green Version]
- Hartung, A.; Bitton-Worms, K.; Rechtman, M.M.; Wenzel, V.; Boergermann, J.H.; Hassel, S.; Henis, Y.I.; Knaus, P. Different routes of bone morphogenic protein (BMP) receptor endocytosis influence BMP signaling. Mol. Cell. Biol. 2006, 26, 7791–7805. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pohl, T.L.M.; Boergermann, J.H.; Schwaerzer, G.K.; Knaus, P.; Cavalcanti-Adam, E.A. Surface immobilization of bone morphogenetic protein 2 via a self-assembled monolayer formation induces cell differentiation. Acta Biomater. 2012, 8, 772–780. [Google Scholar] [CrossRef] [PubMed]
- Migliorini, E.; Valat, A.; Picart, C.; Cavalcanti-Adam, E.A. Tuning cellular responses to BMP-2 with material surfaces. Cytokine Growth Factor Rev. 2016, 27, 43–54. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bragdon, B.; Thinakaran, S.; Bonor, J.; Underhill, T.M.; Petersen, N.O.; Nohe, A. FRET reveals novel protein-receptor interaction of bone morphogenetic proteins receptors and adaptor protein 2 at the cell surface. Biophys. J. 2009, 97, 1428–1435. [Google Scholar] [CrossRef] [Green Version]
- Sieber, C.; Kopf, J.; Hiepen, C.; Knaus, P. Recent advances in BMP receptor signaling. Cytokine Growth Factor Rev. 2009, 20, 343–355. [Google Scholar] [CrossRef]
- Paralkar, V.M.; Hammonds, R.G.; Reddi, A.H. Identification and characterization of cellular binding proteins (receptors) for recombinant human bone morphogenetic protein 2B, an initiator of bone differentiation cascade. Proc. Natl. Acad. Sci. USA 1991, 88, 3397–3401. [Google Scholar] [CrossRef] [Green Version]
- Hardwick, J.C.H.; van den Brink, G.R.; Bleuming, S.A.; Ballester, I.; van den Brande, J.M.H.; Keller, J.J.; Offerhaus, G.J.A.; van Deventer, S.J.H.; Peppelenbosch, M.P. Bone morphogenetic protein 2 is expressed by, and acts upon, mature epithelial cells in the colon. Gastroenterology 2004, 126, 111–121. [Google Scholar] [CrossRef]
- Benn, A.; Hiepen, C.; Osterland, M.; Schütte, C.; Zwijsen, A.; Knaus, P. Role of bone morphogenetic proteins in sprouting angiogenesis: Differential BMP receptor-dependent signaling pathways balance stalk vs. tip cell competence. FASEB J. 2017, 31, 4720–4733. [Google Scholar] [CrossRef] [Green Version]
- Zhou, J.; Lee, P.L.; Lee, C.I.; Wei, S.Y.; Lim, S.H.; Lin, T.E.; Chien, S.; Chiu, J.J. BMP receptor-integrin interaction mediates responses of vascular endothelial Smad1/5 and proliferation to disturbed flow. J. Thromb. Haemost. JTH 2013, 11, 741–755. [Google Scholar] [CrossRef] [PubMed]
- Benn, A.; Bredow, C.; Casanova, I.; Vukičević, S.; Knaus, P. VE-cadherin facilitates BMP-induced endothelial cell permeability and signaling. J. Cell Sci. 2016, 129, 206–218. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zuo, W.H.; Zeng, P.; Chen, X.; Lu, Y.J.; Li, A.; Wu, J.B. Promotive effects of bone morphogenetic protein 2 on angiogenesis in hepatocarcinoma via multiple signal pathways. Sci. Rep. 2016, 6, 37499. [Google Scholar] [CrossRef]
- Finkenzeller, G.; Hager, S.; Stark, G.B. Effects of bone morphogenetic protein 2 on human umbilical vein endothelial cells. Microvasc. Res. 2012, 84, 81–85. [Google Scholar] [CrossRef]
- Fernandez, S.M.; Berlin, R.D. Cell surface distribution of lectin receptors determined by resonance energy transfer. Nature 1976, 264, 411–415. [Google Scholar] [CrossRef] [PubMed]
- Algar, W.R.; Hildebrandt, N.; Vogel, S.S.; Medintz, I.L. FRET as a biomolecular research tool-understanding its potential while avoiding pitfalls. Nat. Methods 2019, 16, 815–829. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Lau, A.; Morris, T.J.; Guo, L.; Fordyce, C.B.; Stanley, E.F. A syntaxin 1, Galpha(o), and N-type calcium channel complex at a presynaptic nerve terminal: Analysis by quantitative immunocolocalization. J. Neurosci. Off. J. Soc. Neurosci. 2004, 24, 4070–4081. [Google Scholar] [CrossRef] [Green Version]
- Bolte, S.; Cordelières, F.P. A guided tour into subcellular colocalization analysis in light microscopy. J. Microsc. 2006, 224, 213–232. [Google Scholar] [CrossRef]
- Medda, R.; Giske, A.; Cavalcanti-Adam, E.A. Challenges in imaging cell surface receptor clusters. Opt. Lasers Eng. 2016, 76, 3–8. [Google Scholar] [CrossRef]
- Bacia, K.; Kim, S.A.; Schwille, P. Fluorescence cross-correlation spectroscopy in living cells. Nat. Methods 2006, 3, 83–89. [Google Scholar] [CrossRef]
- Lippincott-Schwartz, J.; Snapp, E.L.; Phair, R.D. The Development and Enhancement of FRAP as a Key Tool for Investigating Protein Dynamics. Biophys. J. 2018, 115, 1146–1155. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nohe, A.; Keating, E.; Underhill, T.M.; Knaus, P.; Petersen, N.O. Effect of the distribution and clustering of the type I A BMP receptor (ALK3) with the type II BMP receptor on the activation of signalling pathways. J. Cell Sci. 2003, 116, 3277–3284. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Théry, M. Micropatterning as a tool to decipher cell morphogenesis and functions. J. Cell Sci. 2010, 123, 4201–4213. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Sommer, C.; Straehle, C.; Köthe, U.; Hamprecht, F.A. ilastik: Interactive Learning and Segmentation Toolkit. In Proceedings of the Eighth IEEE International Symposium on Biomedical Imaging (ISBI), Chicago, IL, USA, 30 March–2 April 2011; pp. 230–233. [Google Scholar]
- Berg, S.; Kutra, D.; Kroeger, T.; Straehle, C.N.; Kausler, B.X.; Haubold, C.; Schiegg, M.; Ales, J.; Beier, T.; Rudy, M.; et al. ilastik: Interactive machine learning for (bio)image analysis. Nat. Methods 2019, 16, 1226–1232. [Google Scholar] [CrossRef] [PubMed]
- Théry, M.; Racine, V.; Piel, M.; Pépin, A.; Dimitrov, A.; Chen, Y.; Sibarita, J.B.; Bornens, M. Anisotropy of cell adhesive microenvironment governs cell internal organization and orientation of polarity. Proc. Natl. Acad. Sci. USA 2006, 103, 19771–19776. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jortikka, L.; Laitinen, M.; Lindholm, T.; Marttinen, A. Internalization and Intracellular Processing of Bone Morphogenetic Protein (BMP) in Rat Skeletal Muscle Myoblasts (L6). Cell. Signal. 1997, 9, 47–51. [Google Scholar] [CrossRef]
- O’Sullivan, M.J.; Lindsay, A.J. The Endosomal Recycling Pathway-At the Crossroads of the Cell. Int. J. Mol. Sci. 2020, 21, 6074. [Google Scholar] [CrossRef]
- Mitchell, H.; Choudhury, A.; Pagano, R.E.; Leof, E.B. Ligand-dependent and -independent transforming growth factor-beta receptor recycling regulated by clathrin-mediated endocytosis and Rab11. Mol. Biol. Cell 2004, 15, 4166–4178. [Google Scholar] [CrossRef]
- Lin, H.Y.; Wang, X.F.; Ng-Eaton, E.; Weinberg, R.A.; Lodish, H.F. Expression and cloning of the TGF-β Type II Receptor, a Functional Transmembrane Serine/Threonine Kinase. Cell 1992, 68, 775–785. [Google Scholar] [CrossRef]
- Franzén, P.; ten Dijke, P.; Ichijo, H.; Yamashita, H.; Schulz, P.; Heldin, C.H.; Miyazono, K. Cloning of a TGFβ Type I Receptor That Forms a Heteromeric Complex with the TGFβ Type II Receptor. Cell 1993, 75, 681–692. [Google Scholar] [CrossRef]
- Di Guglielmo, G.M.; Le Roy, C.; Goodfellow, A.F.; Wrana, J.L. Distinct endocytic pathways regulate TGF-beta receptor signalling and turnover. Nat. Cell Biol. 2003, 5, 410–421. [Google Scholar] [CrossRef] [PubMed]
- Gleason, R.J.; Akintobi, A.M.; Grant, B.D.; Padgett, R.W. BMP signaling requires retromer-dependent recycling of the type I receptor. Proc. Natl. Acad. Sci. USA 2014, 111, 2578–2583. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [Green Version]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Boog, H.; Medda, R.; Cavalcanti-Adam, E.A. Single Cell Center of Mass for the Analysis of BMP Receptor Heterodimers Distributions. J. Imaging 2021, 7, 219. https://doi.org/10.3390/jimaging7110219
Boog H, Medda R, Cavalcanti-Adam EA. Single Cell Center of Mass for the Analysis of BMP Receptor Heterodimers Distributions. Journal of Imaging. 2021; 7(11):219. https://doi.org/10.3390/jimaging7110219
Chicago/Turabian StyleBoog, Hendrik, Rebecca Medda, and Elisabetta Ada Cavalcanti-Adam. 2021. "Single Cell Center of Mass for the Analysis of BMP Receptor Heterodimers Distributions" Journal of Imaging 7, no. 11: 219. https://doi.org/10.3390/jimaging7110219
APA StyleBoog, H., Medda, R., & Cavalcanti-Adam, E. A. (2021). Single Cell Center of Mass for the Analysis of BMP Receptor Heterodimers Distributions. Journal of Imaging, 7(11), 219. https://doi.org/10.3390/jimaging7110219





