Abstract
Background: BRAF is a core component of the RAS–MAPK signaling pathway and an established oncogenic driver in several solid tumors and selected hematologic malignancies. In myeloid neoplasms, BRAF mutations are rare, and their prevalence, molecular context, and clinical significance remain incompletely defined. Available evidence is scattered across heterogeneous reports involving acute myeloid leukemia, myelodysplastic syndromes, myeloproliferative neoplasms, and overlap myelodysplastic/myeloproliferative neoplasms, with variable descriptions of mutation subtypes, co-mutational profiles, cytogenetic associations, therapeutic approaches, and clinical outcomes. To address these gaps, this review synthesizes data from the published literature up to 2025, summarizing the distribution, genetic landscape, and clinical impact of molecularly confirmed BRAF mutations across the spectrum of myeloid neoplasms. Results: Across published cohorts, BRAF mutations occurred in less than 1% of unselected myeloid neoplasms, with enrichment in chronic myelomonocytic leukemia and therapy-related or secondary acute myeloid leukemia. Both V600E and non-V600E variants were observed, typically within a complex genomic background involving ASXL1, TET2, DNMT3A, SRSF2, and RAS-pathway mutations. Acute myeloid leukemia cases showed poor prognosis, with median overall survival measured in months, whereas myelodysplastic syndromes and chronic myelomonocytic leukemia demonstrated relatively longer survival. Targeted MAPK inhibition produced hematologic responses in selected cases but rarely resulted in durable molecular clearance. Conclusions: BRAF mutations in myeloid neoplasms are rare, heterogeneous, and usually represent secondary events in clonal evolution. Although mutation clearance appears prognostically relevant, current targeted approaches provide limited durability, underscoring the need for prospective studies in this setting.
1. Introduction
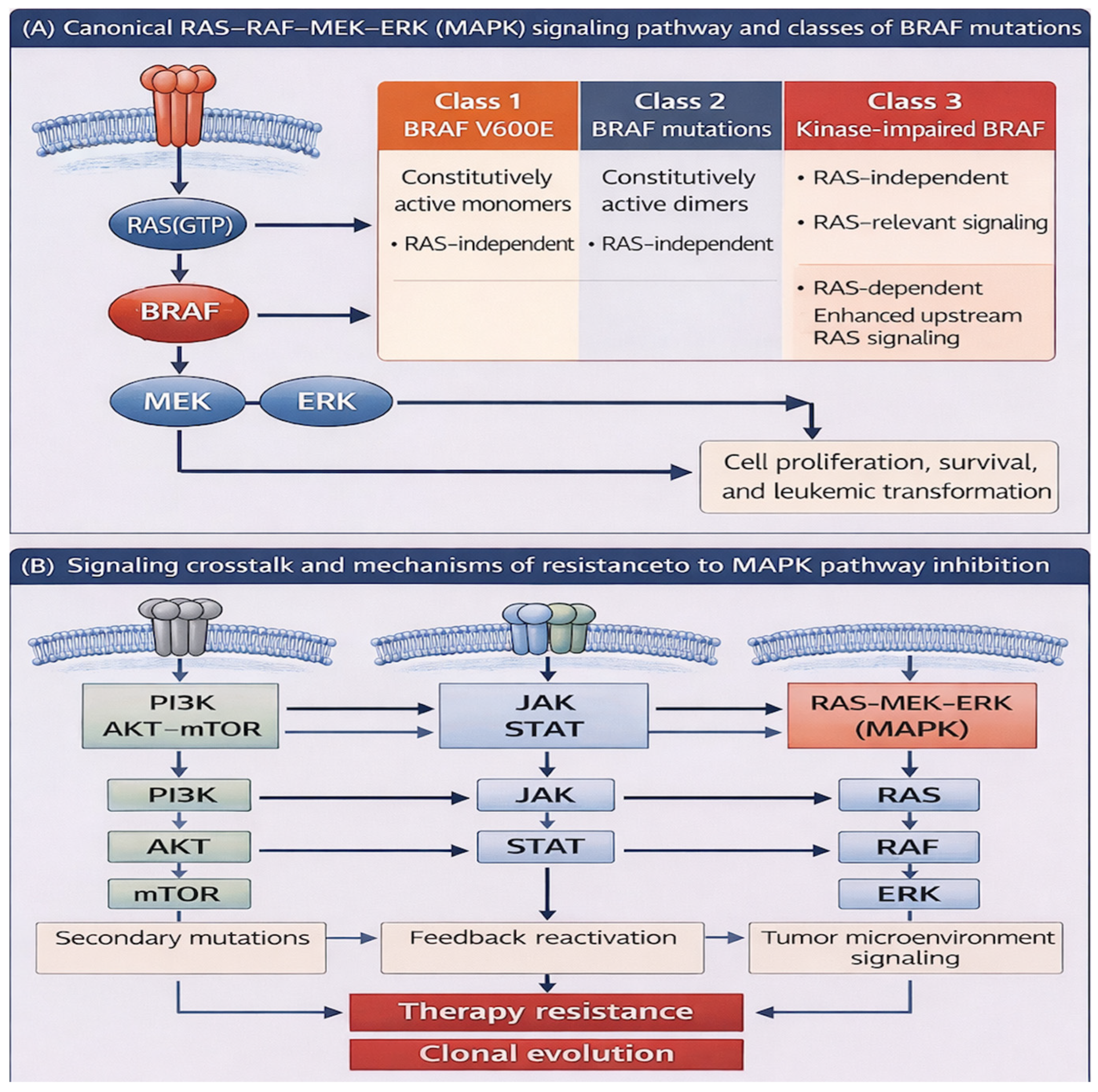
The B-Raf proto-oncogene, serine/threonine kinase (BRAF) is a key component of the RAS–MAPK/ERK signaling pathway, which regulates cellular proliferation, differentiation, and survival [1]. Located on chromosome 7q34, BRAF encodes a 766–amino acid protein with three conserved regions: CR1, which contains the RAS-binding domain (RBD); CR2, a serine-rich hinge region; and CR3, the kinase domain responsible for MEK phosphorylation [2]. Under normal conditions, BRAF remains inactive through intramolecular interactions, with activation occurring when GTP-bound RAS engages the RBD. This conformational shift halts autoinhibition, promotes membrane localization and dimerization, and enables phosphorylation of downstream MEK–ERK targets [1,3,4]. The canonical RAS–RAF–MEK–ERK signaling cascade and its interaction with other major signaling pathways implicated in leukemogenesis are illustrated in Figure 1. Oncogenic mutations disrupt this regulation, driving constitutive pathway activation and uncontrolled proliferation [2].
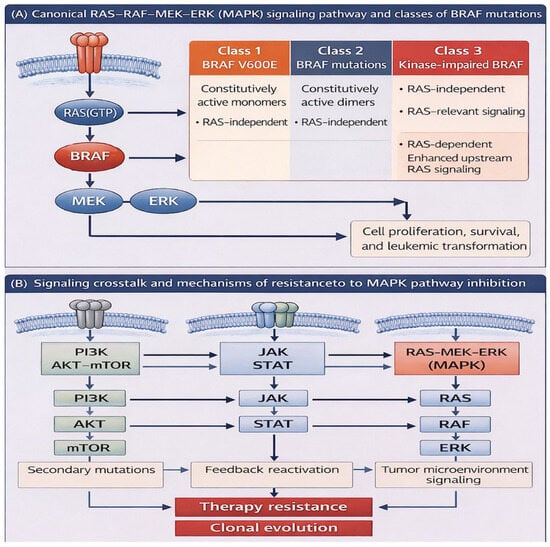
Figure 1.
Overview of MAPK signaling and BRAF mutation classes in myeloid neoplasms. (A) The canonical RAS-RAF-MEK-ERK signaling cascade activated by receptor tyrosine kinases (RTKs). Activating BRAF mutations are classified into three functional classes: class 1 mutations (e.g., V600E), which signal as constitutively active monomers independent of RAS; class 2 mutations, which signal as constitutive dimers and are also RAS-independent; and class 3 mutations, which are kinase-impaired and enhance upstream RAS-dependent signaling. (B) Crosstalk between MAPK signaling and other major intracellular pathways relevant to leukemogenesis, including PI3K–AKT–mTOR and JAK–STAT signaling pathways.
BRAF mutations are established oncogenic drivers in multiple solid tumors, including melanoma (approximately 60%), papillary thyroid carcinoma (40–60%), colorectal cancer (10–15%), and non-small-cell lung cancer (1–4%), as well as subsets of glioblastoma and lymphoma [2,5,6]. More than 200 variants have been described, with nearly 30 functionally characterized. These mutations fall into three classes: class 1 (e.g., V600E), which act as constitutively active monomers; class 2, which signal as active dimers independent of RAS; and class 3, which are kinase-impaired but amplify upstream RAS activity [1,7]. This classification has therapeutic relevance, as class 1 and 2 tumors often respond to direct BRAF inhibition, whereas class 3 mutations are typically dependent on upstream RAS or receptor tyrosine kinase signaling [7].
In hematologic malignancies, BRAF mutations are rare. They are clinically significant in hairy cell leukemia, where V600E serves as both a diagnostic marker and therapeutic target [8,9], but have been reported in fewer than 1–2% of myeloid neoplasms. Notably, the spectrum of BRAF mutations in myeloid disease appears to include a higher proportion of non-V600E variants, whose functional and clinical significance remain unclear [5]. Given the scarcity of cases, available data is limited to small series and case reports with heterogeneous methods and outcomes. This review synthesizes published evidence up to 2025 on molecularly confirmed BRAF mutations across the major categories of myeloid neoplasms, including acute myeloid leukemia, myelodysplastic syndromes, myeloproliferative neoplasms, and myelodysplastic/myeloproliferative overlap disorders. We integrate available data on mutation prevalence, variant subtype, variant allele frequency, co-occurring genetic alterations, cytogenetic features, treatment strategies, and clinical outcomes, with particular attention to patterns of mutation acquisition, therapeutic response, and prognostic implications. By consolidating these dispersed observations, this review aims to clarify the emerging biological and clinical landscape of BRAF-mutated myeloid neoplasms and to highlight areas requiring further investigation.
2. Results
2.1. Study Characteristics
Across 12 published studies, research designs included retrospective and prospective cohorts, cross-sectional analyses, and case series. Sample sizes ranged from 2 patients to more than 6000. Mutation detection methods also varied over time.
Early studies used PCR-based assays, such as restriction fragment length polymorphism (RFLP) analysis [10], single-stranded conformational polymorphism (SSCP) [11,12], and allele-specific PCR for BRAF V600E [13].
In contrast, more recent studies relied on next-generation sequencing (NGS). These included targeted myeloid panels of different sizes, such as 28-gene, 53-gene, or 81-gene assays [14], as well as broader clinical panels like the HopeSeq Heme panel [15]. Other approaches included capture-based and amplicon-based sequencing [16,17,18,19,20].
2.2. Prevalence, Clinical Features, and Genetic Landscape
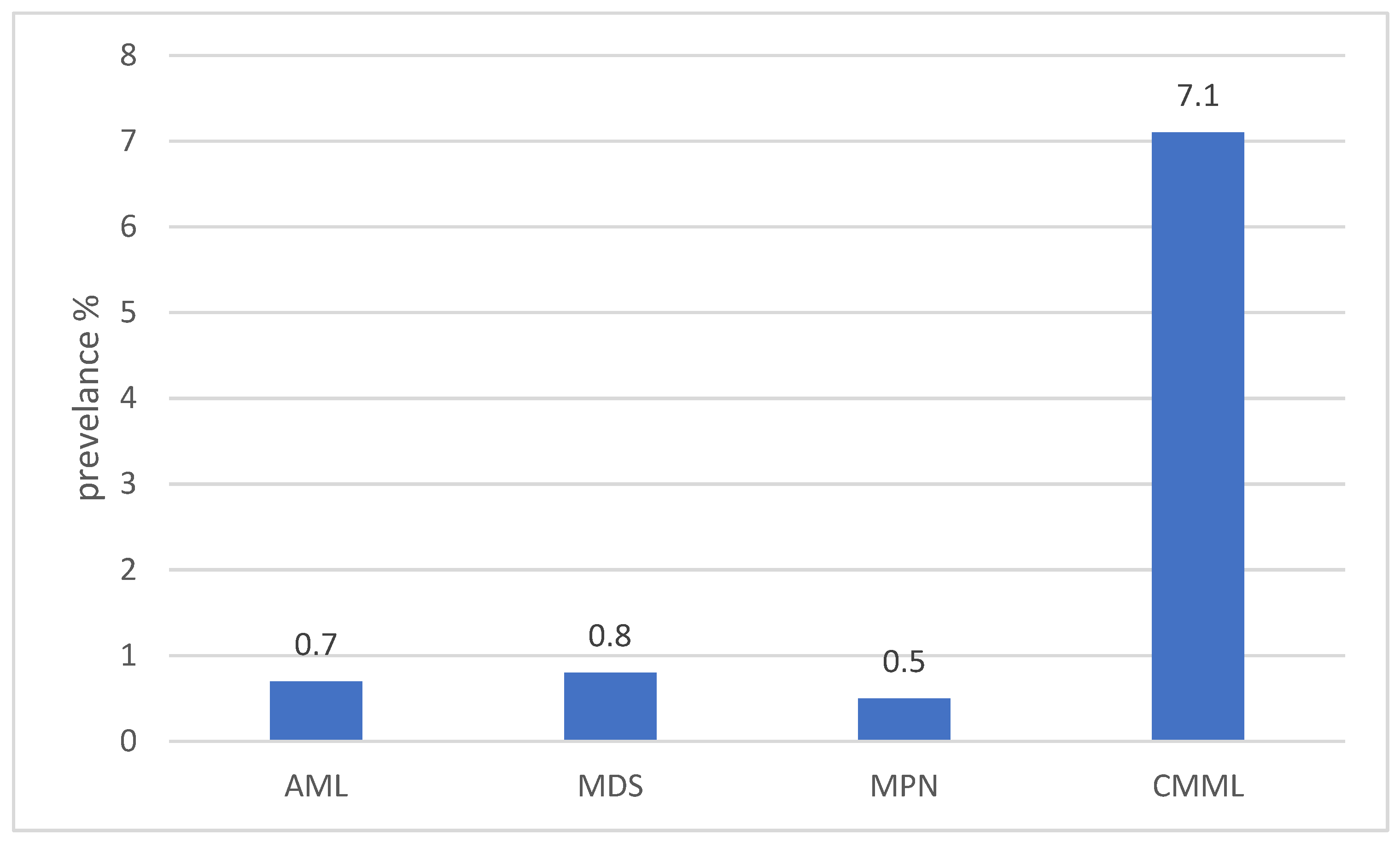
The overall prevalence of BRAF mutations in myeloid neoplasms was consistently low, generally below 1%. Reported frequencies were 0.5–1% in acute myeloid leukemia (AML), myelodysplastic syndromes (MDS), and chronic myelomonocytic leukemia (CMML), though CMML-specific cohorts demonstrated higher rates, reaching up to 7.1% (Figure 2). Sporadic cases were also described in acute promyelocytic leukemia (APL), myeloid sarcoma, and biphenotypic acute leukemias (Table 1).


Table 1.
Summary of published studies on BRAF mutations in myeloid neoplasms.
Table 1.
Summary of published studies on BRAF mutations in myeloid neoplasms.
| ID | Study | DOI | Study Design | Total Sample Size | BRAF Prevalence | Age | Gender (M/F) |
|---|---|---|---|---|---|---|---|
| 1 | Abuasab T et al. (2024) [14] | 10.1080/10428194.2024.2347539 | Retrospective cohort study | 6667 | 48/6667 (0.7%) - AML: 18/2438 (0.7%) - MDS/CMML: 22/2538 (0.8%) - MPN: 5/939 (0.5%) | Median: 70 yo (24–89) | 28/20 |
| 2 | Zhang et al. (2014) [16] | 10.1002/ajh.23652 | Retrospective cohort study | 70 | 5/70 (7.1%) | Median: 67.8 yo (28–86) | 37/33 |
| 3 | Christiansen et al. (2005) [10] | 10.1038/sj.leu.2404009 | Retrospective cohort study | 140 | t-AML: 3/51 (5.8%) t-MDS: 0 | 47, 65, 69 yo | 1/2 |
| 4 | Fei et al. (2024) [15] | 10.3390/ijms25105183 | Retrospective cohort study | 2632 | 14/2632 (0.53%) | Mean: 63.9 yo (23–89) | 10/4 |
| 5 | George et al. (2024) [17] | 10.3390/genes15111383 | Case series | 2 | 16/1600 (1%) 2 cases detailed | 2, 75 yo | 2/0 |
| 6 | Abu-Shihab et al. (2023) [18] | 10.1093/heqpro/daad094 | Cross-sectional study | 42 | NA | Median: 67 yo (19–84) | 24/18 |
| 7 | Kandarpa et al. (2017) [19] | 10.1002/ajh.24728 | Retrospective cohort study | 8 | 2/7 | 65, 83 yo | 1/1 |
| 8 | Santos et al. (2014) [21] | 10.1182/blood.V124.2.1.3172.3172 | Case series | 87 | 1/87 (1.2%) | NA | NA |
| 9 | Papaemmanuil et al. (2016) [20] | 10.1056/NEJMoa1516192 | Prospective cohort study | 1540 | 9/1540 (0.6%) | 18–84 yo | 823/719 |
| 10 | Xu et al. (2017) [22] | 10.1080/10428194.2016.1213830 | Cross-sectional study | 339 | 4/339 (1%) | 0.5, 49, 59, 60 yo | 2/2 |
| 11 | Lee et al. (2004) [11] | 10.1038/sj.leu.2403201 | Retrospective cohort study | 90 | 2/90 (2.2%) | 20–80 yo | NA |
| 12 | Lee et al. (2025) [23] | 10.1101/2025.10.14.682328 | Retrospective cohort study | 5779 | 50/5779 (1%) | Median: 67 yo (19–84) | 27/23 |
Abbreviations: yo = years old; M = male; F = female; NA = not available; AML = acute myeloid leukemia; MDS = myelodysplastic syndrome; MPN = myeloproliferative neoplasm; CMML = chronic myelomonocytic leukemia; t-AML = therapy-related AML. Note: For studies reporting multiple myeloid neoplasm (MN) subtypes, the number of cases in each subtype is indicated within the corresponding table cell. Across larger cohorts, BRAF-mutated patients were typically older adults, with median ages in the sixth to seventh decade; sex distribution was generally balanced when reported [10,11,14,15,16,17,18,19,20,21,22]. Mutations occurred across a range of myeloid neoplasms (Table 2). However, they were relatively enriched in chronic myelomonocytic leukemia and in therapy-related or secondary AML, where frequencies exceeded those seen in de novo AML [11,14,18].
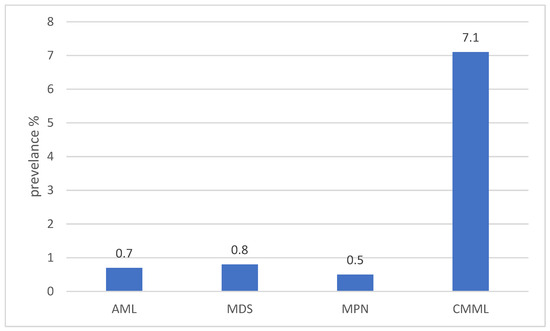
Figure 2.
Distribution of BRAF mutations across myeloid neoplasms. Prevalence values reflect representative estimates reported in published cohorts ([14,15,16,20,21]). Data illustrate the relative enrichment of BRAF mutations in CMML compared to AML, MDS, and MPN, rather than precise pooled estimates.
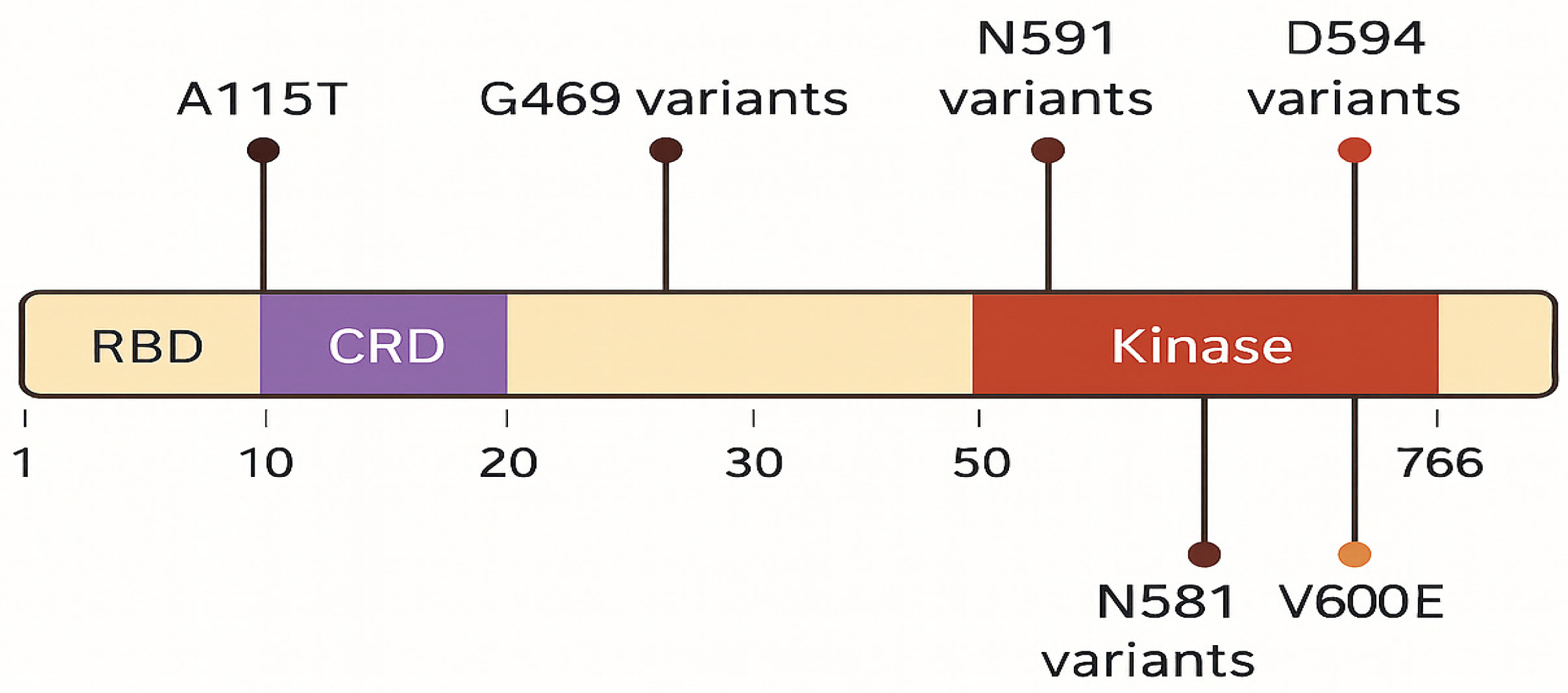
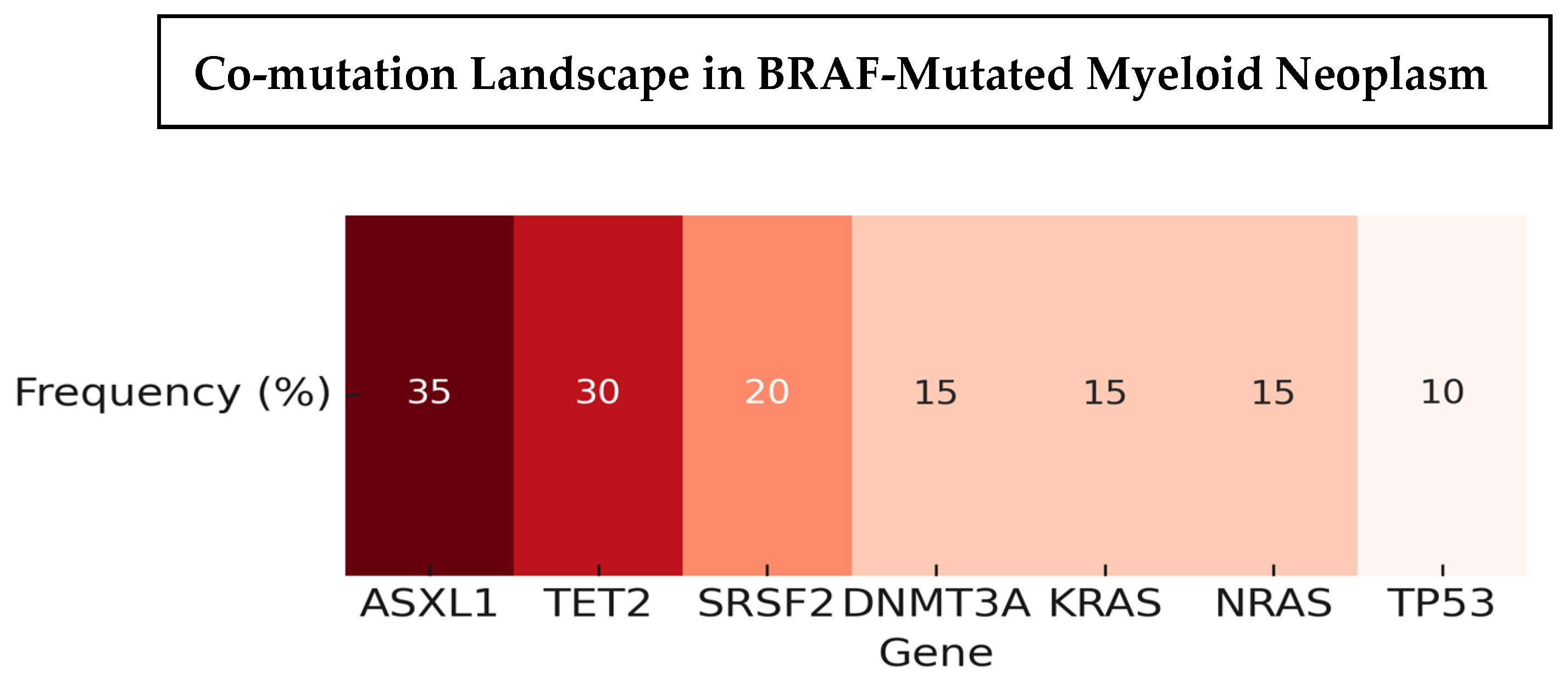
The spectrum of BRAF variants was heterogeneous, including V600E, G469A/V, D594E/G, L597Q/R, N581S/I/K, and K601E, with variant allele frequencies ranging from 1% to over 80% (Figure 3). Co-mutations were frequent and recurrent across studies. The most common partners included ASXL1, TET2, DNMT3A, and splicing factor mutations such as SRSF2, along with recurrent RAS-pathway lesions (KRAS/NRAS) (Figure 4). Other reported partners included IDH1, EZH2, FLT3-ITD, RUNX1, JAK2, TP53, and NPM1. Only a minority of cases carried BRAF mutations as solitary events, suggesting that BRAF usually occurs within a multi-hit genomic context. Cytogenetic profiles ranged from normal karyotype to high-risk abnormalities, such as monosomy 7, complex karyotypes, and KMT2A rearrangements (Table 3).
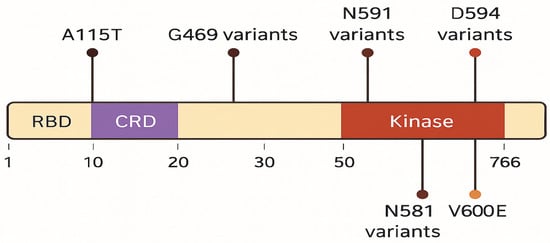
Figure 3.
Distribution of BRAF mutations across functional protein domains.
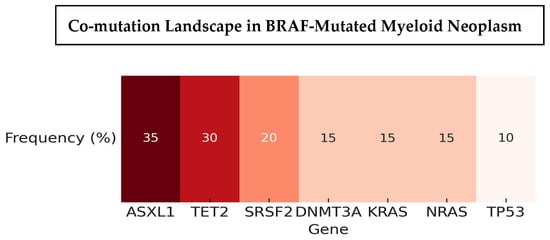
Figure 4.
Heatmap showing recurrent co-mutations in BRAF-mutated myeloid neoplasms. Frequencies reflect aggregated estimates from the largest available studies ([14,15,18,20]). Values are illustrative to highlight recurrent mutation partners; exact frequencies vary by cohort and sequencing method.

Table 2.
Underlying disease and classification of BRAF-mutated cases in published studies.
Table 2.
Underlying disease and classification of BRAF-mutated cases in published studies.
| ID | Study | Total BRAF-Mutated Cases | Underlying Disease Category (n) | Reported Subtype/Clinical Classification (n) |
|---|---|---|---|---|
| 1 | Abuasab et al., 2024 [14] | 48 | AML (18), CMML (12), MDS (10), MF (5), MDS/MPN (1), APL (1), Myeloid sarcoma (1) | s-AML (12), t-MDS (4) |
| 2 | Zhang et al., 2014 [16] | 5 | CMML | CMML-1 (2), CMML-2 (3) |
| 3 | Christiansen et al. (2005) [10] | 3 | Therapy-related AML | AML-M5 |
| 4 | Fei et al. (2024) [15] | 14 | AML (7), MPN (3), MDS (2), MDS/MPN (1), unclear (1) | Acute monocytic AML; AML with monocytic differentiation; ET |
| 5 | George et al. (2024) [17] | 2 | s-AML | AML with monocytic features |
| 6 | Abu-Shihab et al. (2023) [18] | 32 | AML | De novo AML (19), relapsed/refractory AML (8), secondary AML (15) |
| 7 | Kandarpa et al. (2017) [19] | 2 | MPN | Post-ET myelofibrosis |
| 8 | Santos et al. (2014) [21] | 1 | MDS/MPN overlap | Ph-negative MDS/MPN-U |
| 9 | Papaemmanuil et al. (2016) [20] | 9 | AML | De novo AML; therapy-related AML; secondary AML |
| 10 | Xu et al. (2017) [22] | 4 | AML | Monoblastic AML |
| 11 | Lee et al. (2004) [11] | 2 | AML | Biphenotypic AML (1); AML with maturation (1) |
| 12 | Lee et al. (2025) [23] | 50 | AML | De novo AML (21), secondary AML (20), relapsed/refractory AML (9), AML-MR (34) |
Abbreviations: AML = acute myeloid leukemia; MDS = myelodysplastic syndrome; MPN = myeloproliferative neoplasm; CMML = chronic myelomonocytic leukemia; MF = myelofibrosis; APL = acute promyelocytic leukemia; ET = essential thrombocythemia; s-AML = secondary AML; MDS/MPN-U = myelodysplastic/myeloproliferative neoplasm, unclassifiable; AML-MR = Acute Myeloid Leukemia with Myelodysplasia-Related changes. Note: For studies reporting multiple myeloid neoplasm (MN) subtypes, the number of cases in each subtype is indicated within the corresponding table cell.

Table 3.
Genetic characteristics of BRAF-mutated myeloid neoplasm cases.
Table 3.
Genetic characteristics of BRAF-mutated myeloid neoplasm cases.
| ID | Study | BRAF Mutation/VAF | Co-Mutations | Karyotype/Cytogenetics |
|---|---|---|---|---|
| 1 | Abuasab et al., 2024 [14] | G469A, V600E, others; median VAF 8.6% (1.3–86.6) | KRAS, NRAS, ASXL1, TET2, SRSF2, TP53, CBL, DNMT3A | 44% diploid; 19% high-risk; poor risk in AML |
| 2 | Zhang et al., 2014 [16] | D594E, N581S, L597Q, G466E (exon 11/15) | RAS WT | Low risk |
| 3 | Christiansen et al. (2005) [10] | V600E | AML1, CBFb, MLL, RARa, KRAS | Recurrent balanced translocations; +8; MLL-rearrangements |
| 4 | Fei et al. (2024) [15] | V600E, D594G, N581S, others (exons 6, 11, 15, 17) | NRAS, KRAS, DNMT3A, TET2, ASXL1, IDH1, JAK2, TP53, etc. | Normal: 6; Abnormal: 7 (del5q, +8, del9q, del20q, etc.); KMT2A fusions |
| 5 | George et al. (2024) [17] | V600E, N581S | TET2, KRAS, ZRSR2, EZH2, RUNX1T1, RAF1 | Complex karyotype; KMT2A rearrangements |
| 6 | Abu-Shihab et al. (2023) [18] | G469, D594, others; VAF 1–83% | TET2, ASXL1, NRAS, KRAS, RUNX1, DNMT3A, FLT3, NPM1, SRSF2 | NA |
| 7 | Kandarpa et al. (2017) [19] | D594E, V600E, G469V | JAK2, ASXL1, ASXL2, TP53, NF1, PIK3R3, KMT2C | NA |
| 8 | Santos et al. (2014) [21] | D594G | JAK2 | del(5q) |
| 9 | Papaemmanuil et al. (2016) [20] | V600E, D594N, L597Q, A115T | ASXL1, DNMT3A, EZH2, FLT3, IDH1/2, NRAS, RUNX1, TP53, etc. | Favorable: 205; Intermediate: 960; Adverse: 253 |
| 10 | Xu et al. (2017) [22] | V600E, D594G, K601E (exons 11,15) | NPM1, ASXL | Normal: 2; Abnormal: +8, der (1;12), t(10;11) |
| 11 | Lee et al. (2004) [11] | Exon 11 | NA | NA |
| 12 | Lee et al. (2025) [23] | V600, G469, D594, others Median VAF 15% (1–83%) | TET2, NPM1, NRAS, KRAS, BRAF, TP53, and SRSF2. | MECOM-rearrangements, RUNX1:RUNX1T1 fusion |
Abbreviations: VAF = variant allele frequency; NA = not available; WT = wild type. Cytogenetic notation follows ISCN standards.
2.3. Survival Outcomes
Survival outcomes varied by disease subtype and genetic background. In AML, prognosis was generally poor, with median overall survival (OS) ranging from 126 days to 7 months [14,15,17,18]. Clearance of the BRAF mutation, when documented, correlated with improved outcomes (OS 34.8 vs. 10.4 months, p = 0.047) [11,14]. In MDS and CMML, survival was somewhat longer, typically 16–22 months [11,14,17], particularly in patients with diploid cytogenetics and fewer co-mutations (Supplementary Table S2) In many cases, BRAF mutations occurred within complex genomic backgrounds that included established adverse-risk mutations (e.g., ASXL1, TP53, DNMT3A, and RAS-pathway genes), making it difficult to attribute clinical outcomes specifically to BRAF alterations.
2.4. Treatment Approaches and Outcomes
Differences in treatment strategies and clinical outcomes across disease subtypes are summarized in Table 4. In AML, most patients received intensive induction chemotherapy, with generally poor outcomes; occasional prolonged survival was observed in those who achieved BRAF clearance. MAPK inhibitors (dabrafenib, trametinib, vemurafenib) were used mainly in relapsed or refractory settings, producing transient hematologic or morphologic responses without durable molecular remissions [24,25,26,27,28,29]. In MDS and CMML, hypomethylating agents were the most common therapy, with modest clinical improvements but rare molecular clearance; hydroxyurea was also frequently used in CMML. In MPN, patients generally received conventional agents such as hydroxyurea or ruxolitinib, with no consistent evidence for BRAF-targeted therapy.

Table 4.
Clinicogenetic features, treatment patterns, and outcomes of BRAF-mutated myeloid neoplasms by disease subtype.
Outcomes differed substantially by disease subtype. AML harboring BRAF mutations carried the poorest prognosis, with median OS typically ranging from 4 to 7 months [14,17,19]. Clearance of the BRAF mutation, when achieved, was associated with longer survival (34.8 vs. 10.4 months) and may be enhanced by HSCT [11]. Secondary and therapy-related AML were enriched for high-risk cytogenetic abnormalities, including KMT2A rearrangements and monosomy 7, and were associated with inferior outcomes, often measured in weeks [10,15,21]. In contrast, de novo AML cases could achieve initial remissions with intensive chemotherapy, although relapse was common and durable molecular clearance of BRAF was rare [17,18]. Patients with MDS and CMML demonstrated relatively more favorable outcomes, with median OS of approximately 16–22 months in larger cohorts [11,14,17]. Notably, Zhang et al. (2014) reported improved survival among CMML-1 patients with low-risk cytogenetics, a finding that was not consistently reproduced in subsequent datasets [16]. Data on MPN were limited; however, co-mutation with RAS-pathway genes was associated with aggressive disease biology and progression to AML despite therapy [20,22].
2.5. Published Case Reports of BRAF-Altered Myeloid Neoplasms
Six detailed case reports published between 2015 and 2025 were also identified, originating from Australia, the United States, France, and India. Patient ages ranged from early childhood to 78 years, with equal sex distribution (Table S1). Underlying diagnoses included chronic myeloid leukemia (CML), therapy-related AML, CMML, therapy-related MDS, and post–acute lymphoblastic leukemia AML. There was also one case of CMML with Langerhans cell histiocytoma, the mutation was harbored in both myeloid neoplasms. BRAF p.V600E was present in five of six cases, while the remaining report described a BRAF-wild-type patient with MAPK pathway activation. Co-mutations included KRAS, TET2, and SRSF2, often accompanied by high-risk cytogenetics such as t(9;11) KMT2A–MLLT3, t(9;22) BCR–ABL1, +8, del(7q), and +5. Treatments ranged from conventional chemotherapy and hypomethylating agents to MAPK inhibitors, with responses varying from transient cytoreduction to complete remission. Durable responses were rare; relapses often coincided with KRAS-mutated subclones. Death occurred in four cases, typically within a short time frame due either to disease progression or treatment-related complications. One patient was reported to be in preparation for haploidentical bone marrow transplantation at last follow-up.
3. Discussion
BRAF mutations are rare in myeloid neoplasms, occurring in <1% of unselected cohorts and more commonly in CMML and therapy-related AML [10,11,14,15,16,17,18,19,20,21,22]. Most reported BRAF mutations were clustered within the kinase domain, specifically involving the activation segment (exon 15, including V600). Despite the overall poor outcomes, achieving clearance of BRAF mutations was associated with longer survival, and this clearance may be augmented by HSCT. These findings highlight the clinical relevance of identifying BRAF mutations despite their low prevalence.
When compared with other hematologic malignancies, the role of BRAF in myeloid disease is far less definitive. In hairy cell leukemia, BRAF V600E is disease-defining and therapeutically targetable [8,9]. In Langerhans cell histiocytosis and Erdheim–Chester disease, BRAF or MAPK pathway mutations occur in 50–70% of cases and underpin the dramatic success of BRAF/MEK inhibitors, now approved in these settings [30,31,32,33,34]. In multiple myeloma, BRAF V600E is rare but associated with more aggressive disease and inferior outcomes [35,36]. By contrast, in myeloid neoplasms, BRAF mutations arise infrequently and within a heterogeneous genomic background, limiting their utility as stand-alone biomarkers or therapeutic targets [10,11,14,15,16,17,18,19,20,21,22]. Indeed, some studies such as Shin et al. (2016) have reported no BRAF mutations at all, underscoring the variability across cohorts [37].
The biological implications of these findings are important. Canonical class 1 mutations, such as V600E, act as constitutively active monomers, while class 2 and class 3 variants rely on dimerization or upstream RAS activation. Their frequent co-occurrence with epigenetic regulators (ASXL1, TET2, DNMT3A), splicing factors (SRSF2), and RAS pathway mutations (KRAS/NRAS) suggest that BRAF is rarely an initiating lesion in myeloid disease but rather contributes to clonal evolution and progression. These co-occurrences likely reflect shared clonal architecture and disease biology rather than a direct functional interaction between BRAF and specific co-mutated genes. Given the reported adverse prognostic impact of many of these co-mutations, the available data do not allow reliable assessment of an independent biological or prognostic contribution attributable specifically to BRAF mutations. However, we also acknowledge that some cases describe BRAF as the sole detectable abnormality (within the limits of the assays used), raising the possibility that BRAF could act as a more central driver in select patients. Because the included studies span different testing eras and platforms (PCR vs broader NGS), improvements in sequencing breadth and sensitivity may increasingly clarify whether there exists a subset with more “isolated” BRAF lesions or more interpretable clonal architecture.
Reports of clonal dynamics support this view: in some patients, BRAF persists or expands at relapse, while in others it is replaced by alternative drivers, underscoring its instability as a therapeutic target [11,14].
From a clinical standpoint, BRAF testing may be most relevant in contexts where it is enriched, such as CMML and therapy-related AML [11,14,19]. The relative enrichment of BRAF mutations in myeloid neoplasms with prominent monocytic differentiation, including CMML and acute monocytic leukemia, suggests a potential lineage-specific biological context. Although MAPK signaling is ubiquitous across cell types, monocytic lineage cells exhibit heightened dependence on tightly regulated MAPK activity during differentiation and inflammatory activation [38]. Parallels can be drawn with histiocytic disorders such as Langerhans cell histiocytosis and Erdheim–Chester disease, which arise from monocyte-derived cells and are characterized by frequent MAPK pathway mutations. These observations raise the possibility that monocytic-lineage cells may be particularly permissive to MAPK-driven clonal expansion; however, this association remains speculative and requires dedicated mechanistic and functional studies for validation. Standard therapies remain ineffective, with short survival in AML and modest benefit in MDS and CMML. Responses to BRAF/MEK inhibitors have been observed but are typically transient, without durable molecular clearance [11,14,30,31,32,33,34]. Achieving molecular mutation clearance has been reported to be prognostically meaningful, and HSCT may offer a survival benefit when clearance is attained. However, this observation is currently supported primarily by a single retrospective study (Tariq et al.) and therefore remains hypothesis-generating and in need of validation in larger, independent cohorts and prospective studies [11,14]. These observations suggest that BRAF clearance could be explored as a potential biomarker of treatment response.
Several limitations affect interpretation. Most data come from retrospective cohorts or case reports with a small number of BRAF-positive cases. Testing strategies varied; while earlier studies used PCR-based hotspot assays, more recent cohorts used NGS, complicating prevalence estimates [10,11,12,13,14,15,16,17,18,19,20,21,22,39]. PCR-based approaches are inherently limited in their ability to detect non-hot-spot substitutions, indels, and low–allele frequency variants, potentially leading to underestimation of both the true prevalence and the full mutational spectrum of BRAF alterations in earlier cohorts. Treatment approaches were heterogeneous and often anecdotal, and publication bias may favor reports of positive targeted therapy outcomes. Although publicly available genomic databases may provide additional insights into mutation frequency and co-mutation patterns, the rarity of BRAF alterations in myeloid neoplasms, coupled with variable diagnostic annotation and clinical granularity, limits their utility for definitive genotype–phenotype correlations in this setting.
Future work should prioritize prospective multicenter studies with standardized sequencing to define prevalence and outcomes more precisely. Functional studies are needed to clarify the significance of non-V600E variants, particularly class 2 and 3 mutations. Clinical trials of BRAF and MEK inhibitors, ideally in rational combinations with chemotherapy, HMAs, or HSCT, will be required to establish whether targeting MAPK signaling can provide a durable benefit. Finally, systematic evaluation of molecular clearance as a biomarker should be incorporated into trial design.
4. Conclusions
BRAF mutations in myeloid neoplasms are rare but clinically relevant. They are enriched in CMML and therapy-related AML, usually occur in a complex genomic background, and portend poor outcomes. Unlike HCL or histiocytic disorders, where BRAF is a defining driver, in myeloid neoplasms, these mutations appear secondary and unstable. While targeted therapy responses have been modest and short-lived, clearance of BRAF mutations and subsequent HSCT may provide a path to improved outcomes. Collaborative efforts will be essential to clarify the prognostic and therapeutic role of this rare subset.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/biomedicines14030672/s1. Table S1: Clinical and Molecular Characteristics of BRAF-Mutated Myeloid Neoplasm Case Reports. Table S2: Clinical outcomes of BRAF-mutated myeloid neoplasm cases.
Author Contributions
Conceptualization: S.F.M. and G.B.; methodology: S.F.M. and T.A.; data curation: A.M., M.F.M., A.E.A.A., A.F.A.-M. and M.A.; writing—original draft preparation: A.M., M.F.M., A.E.A.A., T.A., H.C. and G.B.; writing—review and editing: S.F.M. and R.M.; supervision: G.B. and H.C.; project administration: S.F.M. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
The study did not require ethical approval.
Informed Consent Statement
Not applicable.
Data Availability Statement
This review is based on previously published studies. No new datasets were generated or analyzed during the current study.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| AML | Acute Myeloid Leukemia |
| APL | Acute Promyelocytic Leukemia |
| BRAF | B-Raf Proto-Oncogene, Serine/Threonine Kinase |
| CMML | Chronic Myelomonocytic Leukemia |
| HCL | Hairy Cell Leukemia |
| HSCT | Hematopoietic Stem Cell Transplantation |
| MAPK | Mitogen-Activated Protein Kinase |
| MDS | Myelodysplastic Syndromes |
| MPN | Myeloproliferative Neoplasms |
References
- Wellbrock, C.; Karasarides, M.; Marais, R. The RAF proteins take centre stage. Nat. Rev. Mol. Cell Biol. 2004, 5, 875–885. [Google Scholar] [CrossRef] [PubMed]
- Davies, H.; Bignell, G.R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Teague, J.; Woffendin, H.; Garnett, M.J.; Bottomley, W.; et al. Mutations of the BRAF gene in human cancer. Nature 2002, 417, 949–954. [Google Scholar] [CrossRef] [PubMed]
- Chambard, J.-C.; Lefloch, R.; Pouyssegur, J.; Lenormand, P. ERK implication in cell cycle regulation. Biochim. Biophys. Acta 2007, 1773, 1299–1310. [Google Scholar] [CrossRef]
- Avruch, J.; Khokhlatchev, A.; Kyriakis, J.M.; Luo, Z.; Tzivion, G.; Vavvas, D.; Zhang, X.-F. Ras activation of the Raf kinase: Tyrosine kinase recruitment of the MAP kinase cascade. Recent Prog. Horm. Res. 2001, 56, 127–155. [Google Scholar] [CrossRef]
- Naoki, K.; Chen, T.-H.; Richards, W.G.; Sugarbaker, D.J.; Meyerson, M. Missense mutations of the BRAF gene in human lung adenocarcinoma. Cancer Res. 2002, 62, 7001–7003. [Google Scholar]
- Caunt, C.J.; Keyse, S.M. Dual-specificity MAP kinase phosphatases (MKPs): Shaping the outcome of MAP kinase signalling. FEBS J. 2013, 280, 489–504. [Google Scholar] [CrossRef]
- Jones, D.T.; Kocialkowski, S.; Liu, L.; Pearson, D.M.; Backlund, L.M.; Ichimura, K.; Collins, V.P. Tandem duplication producing a novel oncogenic BRAF fusion gene defines the majority of pilocytic astrocytomas. Cancer Res. 2008, 68, 8673–8677. [Google Scholar] [CrossRef] [PubMed]
- Tiacci, E.; Trifonov, V.; Schiavoni, G.; Holmes, A.; Kern, W.; Martelli, M.P.; Pucciarini, A.; Bigerna, B.; Pacini, R.; Wells, V.A.; et al. BRAF mutations in hairy-cell leukemia. N. Engl. J. Med. 2011, 364, 2305–2315. [Google Scholar] [CrossRef]
- Tiacci, E.; Park, J.H.; De Carolis, L.; Chung, S.S.; Broccoli, A.; Scott, S.; Zaja, F.; Devlin, S.; Pulsoni, A.; Chung, Y.R.; et al. Targeting mutant BRAF in relapsed or refractory hairy-cell leukemia. N. Engl. J. Med. 2015, 373, 1733–1747. [Google Scholar] [CrossRef] [PubMed]
- Christiansen, D.H.; Andersen, M.K.; Desta, F.; Pedersen-Bjergaard, J. Mutations of genes in the receptor tyrosine kinase/RAS-BRAF signal transduction pathway in therapy-related myelodysplasia and acute myeloid leukemia. Leukemia 2005, 19, 2232–2240. [Google Scholar] [CrossRef]
- Lee, J.W.; Soung, Y.H.; Park, W.S.; Kim, S.Y.; Nam, S.W.; Min, W.S.; Lee, J.Y.; Yoo, N.J.; Lee, S.H. BRAF mutations in acute leukemias. Leukemia 2004, 18, 170–172. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.L.; Snaddon, J.; Neat, M.; Cambal-Parrales, M.; Arch, R.; Lister, T.A.; Fitzgibbon, J. Mutation of BRAF is uncommon in AML FAB type M1 and M2. Leukemia 2003, 17, 274–275. [Google Scholar] [CrossRef][Green Version]
- Trifa, A.P.; Popp, R.A.; Cucuianu, A.; Coadă, C.A.; Urian, L.G.; Militaru, M.S.; Bănescu, C.; Dima, D.; Farcaş, M.F.; Crişan, T.O.; et al. Absence of BRAF V600E mutation in a cohort of 402 patients with various chronic and acute myeloid neoplasms. Leuk. Lymphoma 2012, 53, 2496–2497. [Google Scholar] [CrossRef]
- Abuasab, T.; Mohamed, S.; Pemmaraju, N.; Kadia, T.M.; Daver, N.; DiNardo, C.D.; Ravandi, F.; Qiao, W.; Montalban-Bravo, G.; Borthakur, G. BRAF mutation in myeloid neoplasm: Incidences and clinical outcomes. Leuk. Lymphoma 2024, 65, 1344–1349. [Google Scholar] [CrossRef]
- Fei, F.; Caporale, C.; Chang, L.; Fortini, B.K.; Ali, H.; Bell, D.; Stein, A.; Marcucci, G.; Telatar, M.; Afkhami, M. BRAF mutations in patients with myeloid neoplasms: A cancer center multigene next-generation sequencing analysis experience. Int. J. Mol. Sci. 2024, 25, 5183. [Google Scholar] [CrossRef]
- Zhang, L.; Singh, R.R.; Patel, K.P.; Stingo, F.; Routbort, M.; You, M.J.; Miranda, R.N.; Garcia-Manero, G.; Kantarjian, H.M.; Medeiros, L.J.; et al. BRAF kinase domain mutations are present in a subset of chronic myelomonocytic leukemia with wild-type RAS. Am. J. Hematol. 2014, 89, 499–504. [Google Scholar] [CrossRef]
- George, G.V.; Evans, A.G.; Jajosky, A.N. BRAF V600E-mutant acute myeloid leukemia: A case series and literature review of a rare entity. Genes 2024, 15, 1383. [Google Scholar] [CrossRef]
- Abu-Shihab, Y.; Nicolet, D.; Mrózek, K.; Routbort, M.; Patel, K.P.; Walker, C.J.; Buss, J.; Stiff, A.; Laganson, A.; DiNardo, C.D.; et al. BRAF-mutated acute myeloid leukemia represents a distinct, prognostically poor subgroup enriched in myelodysplasia-related AML. Blood 2023, 142, 1575. [Google Scholar] [CrossRef]
- Kandarpa, M.; Wu, Y.-M.; Robinson, D.; Burke, P.W.; Chinnaiyan, A.M.; Talpaz, M. Clinical characteristics and whole exome/transcriptome sequencing of coexisting chronic myeloid leukemia and myelofibrosis. Am. J. Hematol. 2017, 92, 555–561. [Google Scholar] [CrossRef]
- Papaemmanuil, E.; Gerstung, M.; Bullinger, L.; Gaidzik, V.I.; Paschka, P.; Roberts, N.D.; Potter, N.E.; Heuser, M.; Thol, F.; Bolli, N.; et al. Genomic classification and prognosis in acute myeloid leukemia. N. Engl. J. Med. 2016, 374, 2209–2221. [Google Scholar] [CrossRef] [PubMed]
- Santos, F.P.; Lisboa, B.; Datoguia, T.S.; Helman, R.; Pereira, W.O.; Puga, R.D.; Miyagi, M.; da Mata, E.H.; Santucci, R.; Perini, G.F.; et al. Incidence and impact of activating mutations of the RAS–RAF–MEK–ERK pathway in patients with Philadelphia-negative myeloproliferative neoplasms and myelodysplastic/myeloproliferative disorders-unclassified. Blood 2014, 124, 3172. [Google Scholar] [CrossRef]
- Xu, Y.; Wertheim, G.; Morrissette, J.J.D.; Bagg, A. BRAF kinase domain mutations in de novo acute myeloid leukemia with monocytic differentiation. Leuk. Lymphoma 2017, 58, 743–745. [Google Scholar] [CrossRef]
- Lee, D.; Abu-Shihab, Y.; Plas, K.; Nicolet, D.; Mrózek, K.; Routbort, M.J.; Patel, K.P.; Walker, C.J.; Buss, J.; Stiff, A.R.; et al. Prognostic and therapeutic implications of BRAF mutations in acute myeloid leukemia. bioRxiv 2025. [Google Scholar] [CrossRef]
- Andrews, M.C.; Turner, N.; Boyd, J.; Roberts, A.W.; Grigg, A.P.; Behren, A.; Cebon, J. Cellular mechanisms underlying complete hematological response of chronic myeloid leukemia to BRAF and MEK1/2 inhibition in a patient with concomitant metastatic melanoma. Clin. Cancer Res. 2015, 21, 5222–5234. [Google Scholar] [CrossRef] [PubMed]
- McNew, B.R.; Darbro, B.W.; Ma, D.; Gordon, D.J. Development of secondary acute myeloid leukemia in a pediatric patient concurrently receiving primary therapy for Ewing sarcoma. J. Pediatr. Hematol. Oncol. 2017, 39, e370–e372. [Google Scholar] [CrossRef]
- Konstantinou, M.P.; Lucas, P.; Uthurriague, C.; Severino-Freire, M.; Spenatto, N.; Gaudin, C.; Lamant, L.; Tournier, E.; Bulai-Livideanu, C.; Meyer, N.; et al. Langerhans cell histiocytosis associated with chronic myelomonocytic leukaemia both harbouring the same BRAF V600E mutation: Efficacy of vemurafenib. J. Eur. Acad. Dermatol. Venereol. 2021, 35, e120–e121. [Google Scholar] [CrossRef]
- Tedjaseputra, A.; Vilcassim, F.S.; Grigoriadis, G. Ocular infiltration as initial presentation of acute monocytic leukaemia transformed from chronic myelomonocytic leukaemia associated with BRAF V600E mutation. BMJ Case Rep. 2019, 12, e228519. [Google Scholar] [CrossRef] [PubMed]
- Mamidala, S.; Tripathi, P.; Agarwal, S.; Khera, S. Acute myeloid leukaemia post-cytotoxic therapy with BRAF V600E mutation in a child. BMJ Case Rep. 2025, 18, e263083. [Google Scholar] [CrossRef] [PubMed]
- Wander, S.A.; Hasserjian, R.P.; Oduro, K.; Glomski, K.; Nardi, V.; Cote, G.M.; Graubert, T.A.; Brunner, A.M.; Chen, Y.A.; Fathi, A.T. Combined targeted therapy for BRAF-mutant, treatment-related acute myeloid leukemia. JCO Precis. Oncol. 2017, 1, 1–7. [Google Scholar] [CrossRef]
- Badalian-Very, G.; Vergilio, J.-A.; Degar, B.A.; Rodriguez-Galindo, C.; Rollins, B.J. Recurrent BRAF mutations in Langerhans cell histiocytosis. Blood 2010, 116, 1919–1923. [Google Scholar] [CrossRef]
- Haroche, J.; Charlotte, F.; Arnaud, L.; von Deimling, A.; Hélias-Rodzewicz, Z.; Hervier, B.; Cohen-Aubart, F.; Launay, D.; Lesot, A.; Mokhtari, K.; et al. High prevalence of BRAF V600E mutations in Erdheim–Chester disease but not in other non-Langerhans cell histiocytoses. Blood 2012, 120, 2700–2703. [Google Scholar] [CrossRef]
- Diamond, E.L.; Durham, B.H.; Haroche, J.; Yao, Z.; Ma, J.; Parikh, S.A.; Wang, Z.; Choi, J.; Kim, E.; Cohen-Aubart, F.; et al. Diverse and targetable kinase alterations drive histiocytic neoplasms. Cancer Discov. 2016, 6, 154–165. [Google Scholar] [CrossRef] [PubMed]
- Go, H.; Jeon, Y.K.; Huh, J.; Choi, S.J.; Choi, Y.D.; Cha, H.J.; Kim, J.E.; Lee, H.; Kim, C.W. Frequent detection of BRAF(V600E) mutations in histiocytic and dendritic cell neoplasms. Histopathology 2014, 65, 261–272. [Google Scholar] [CrossRef] [PubMed]
- Bautista, F.; Paci, A.; Minard-Colin, V.; Dufour, C.; Grill, J.; Lacroix, L.; Valteau-Couanet, D.; Geoerger, B. Vemurafenib in pediatric patients with BRAF V600E-mutated high-grade gliomas and secondary histiocytic sarcoma. Pediatr. Blood Cancer 2014, 61, 1101–1103. [Google Scholar] [CrossRef]
- Andrulis, M.; Lehners, N.; Capper, D.; Penzel, R.; Heining, C.; Huellein, J.; Zenz, T.; von Deimling, A.; Schirmacher, P.; Ho, A.D.; et al. Targeting the BRAF V600E mutation in multiple myeloma. Cancer Discov. 2013, 3, 862–869. [Google Scholar] [CrossRef]
- Chapman, M.A.; Lawrence, M.S.; Keats, J.J.; Cibulskis, K.; Sougnez, C.; Schinzel, A.C.; Harview, C.L.; Brunet, J.-P.; Ahmann, G.J.; Adli, M.; et al. Initial genome sequencing and analysis of multiple myeloma. Nature 2011, 471, 467–472. [Google Scholar] [CrossRef]
- Shin, S.Y.; Lee, S.T.; Kim, H.J.; Cho, E.H.; Kim, J.W.; Park, S.; Jung, C.W.; Kim, S.H. Mutation profiling of 19 candidate genes in acute myeloid leukemia suggests significance of DNMT3A mutations. Oncotarget 2016, 7, 54825–54837. [Google Scholar] [CrossRef]
- Arthur, J.S.; Ley, S.C. Mitogen-activated protein kinases in innate immunity. Nat. Rev. Immunol. 2013, 13, 679–692. [Google Scholar] [CrossRef]
- de Vries, A.C.H.; Stam, R.W.; Kratz, C.P.; Zenker, M.; Niemeyer, C.M.; van den Heuvel-Eibrink, M.M. Mutation analysis of the BRAF oncogene in juvenile myelomonocytic leukemia. Haematologica 2007, 92, 1574–1575. [Google Scholar] [CrossRef] [PubMed][Green Version]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.