Integrated Transcriptomic and Biochemical Analyses Reveal the Root Development-Promoting Mechanism of Piriformospora indica on Blueberry Under Tap Water Irrigation
Abstract
1. Introduction
2. Results
2.1. P. indica Colonization Improved the Root Biomass of Blueberry Plants Under Long-Term Tap Water Irrigation
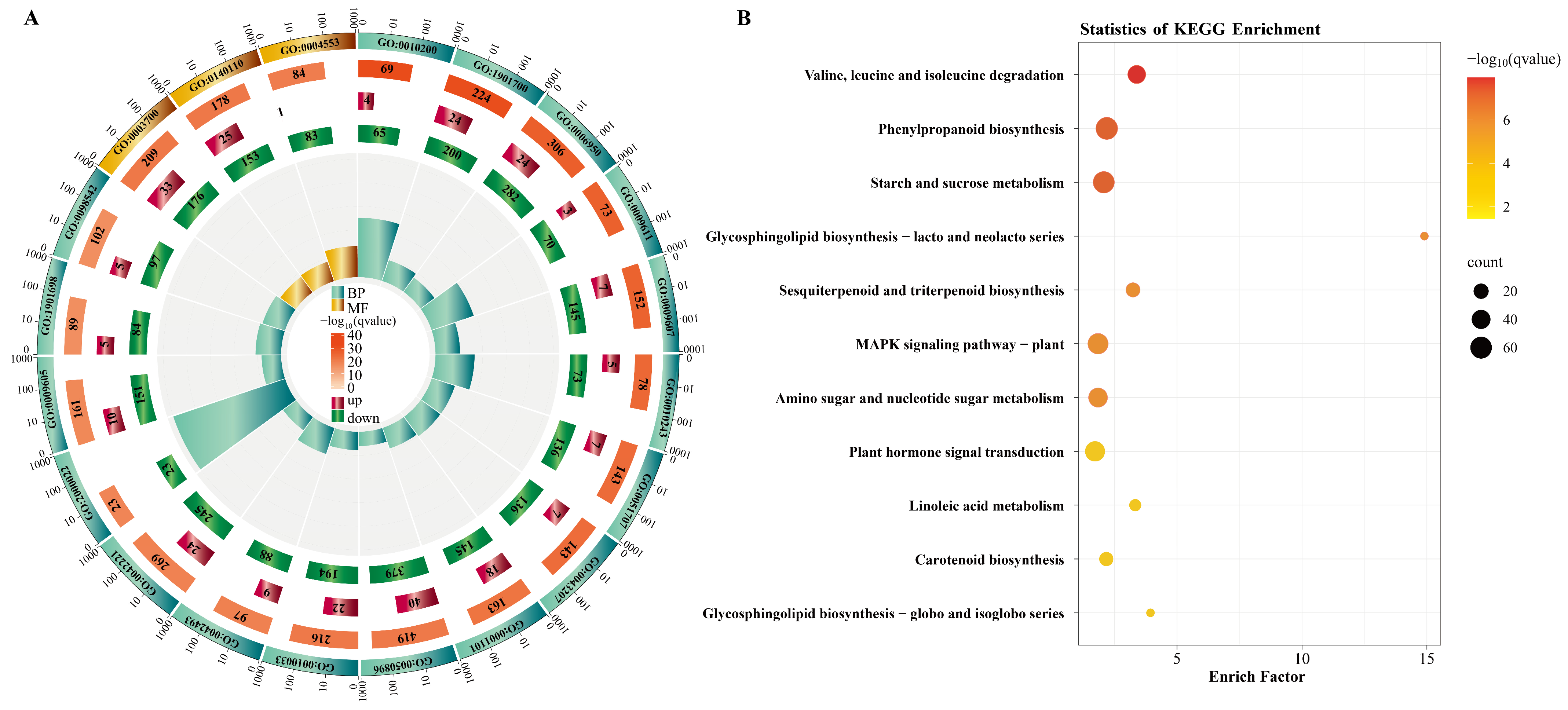
2.2. GO and KEGG Enrichment Analysis of DEGs
2.3. MapMan Annotation and PageMan Enrichment Analysis of DEGs
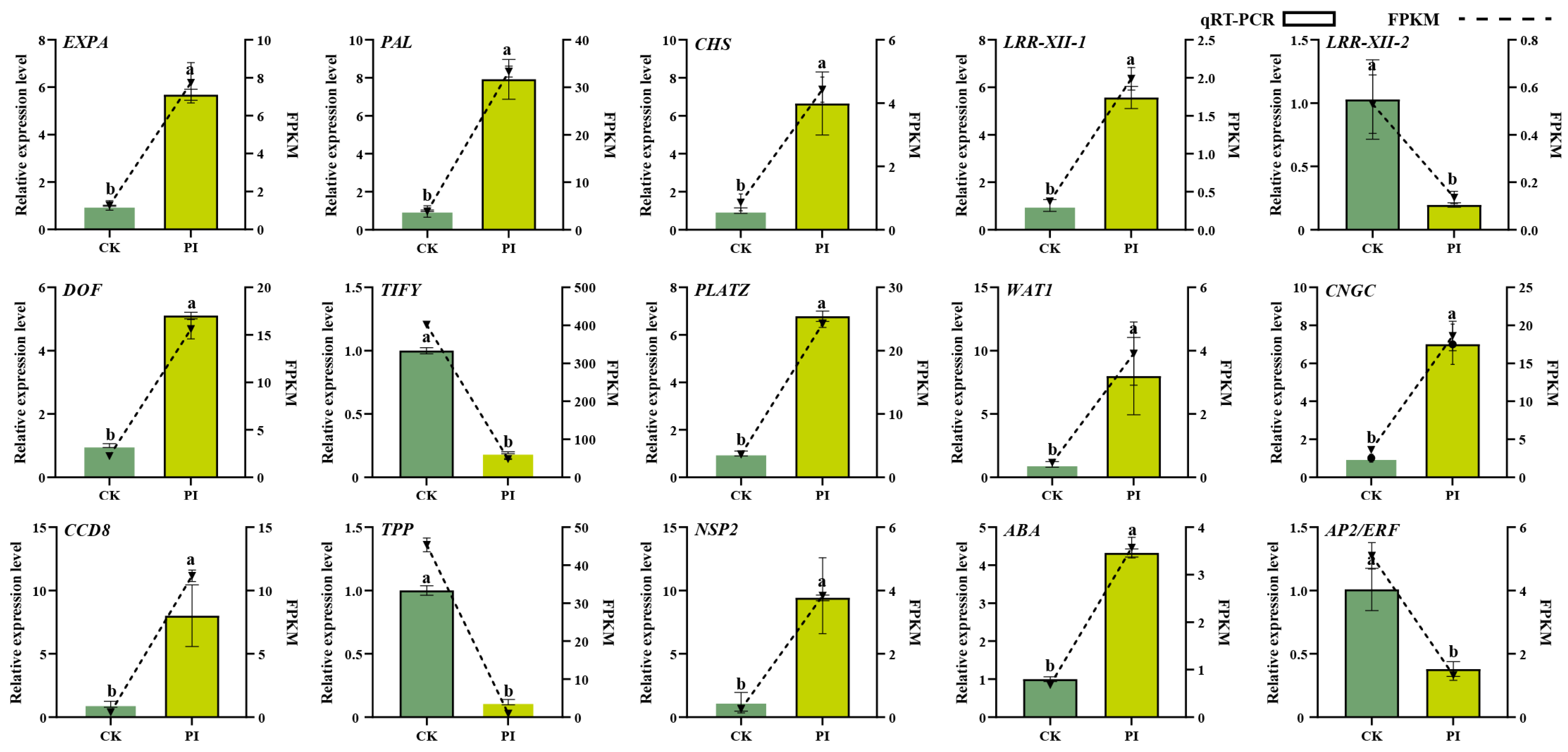
2.4. Gene Expression Validation Results of Selected DEGs
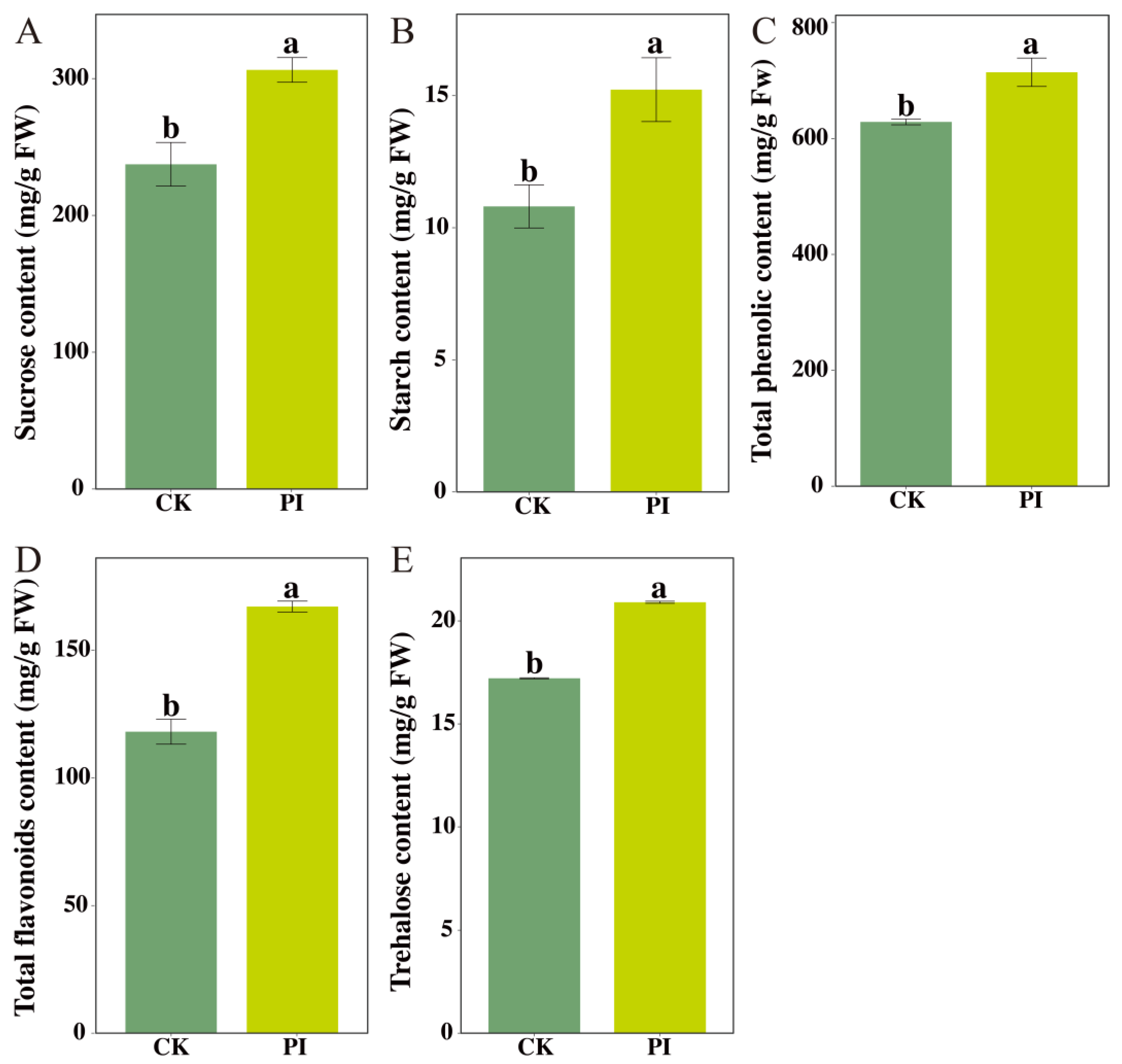
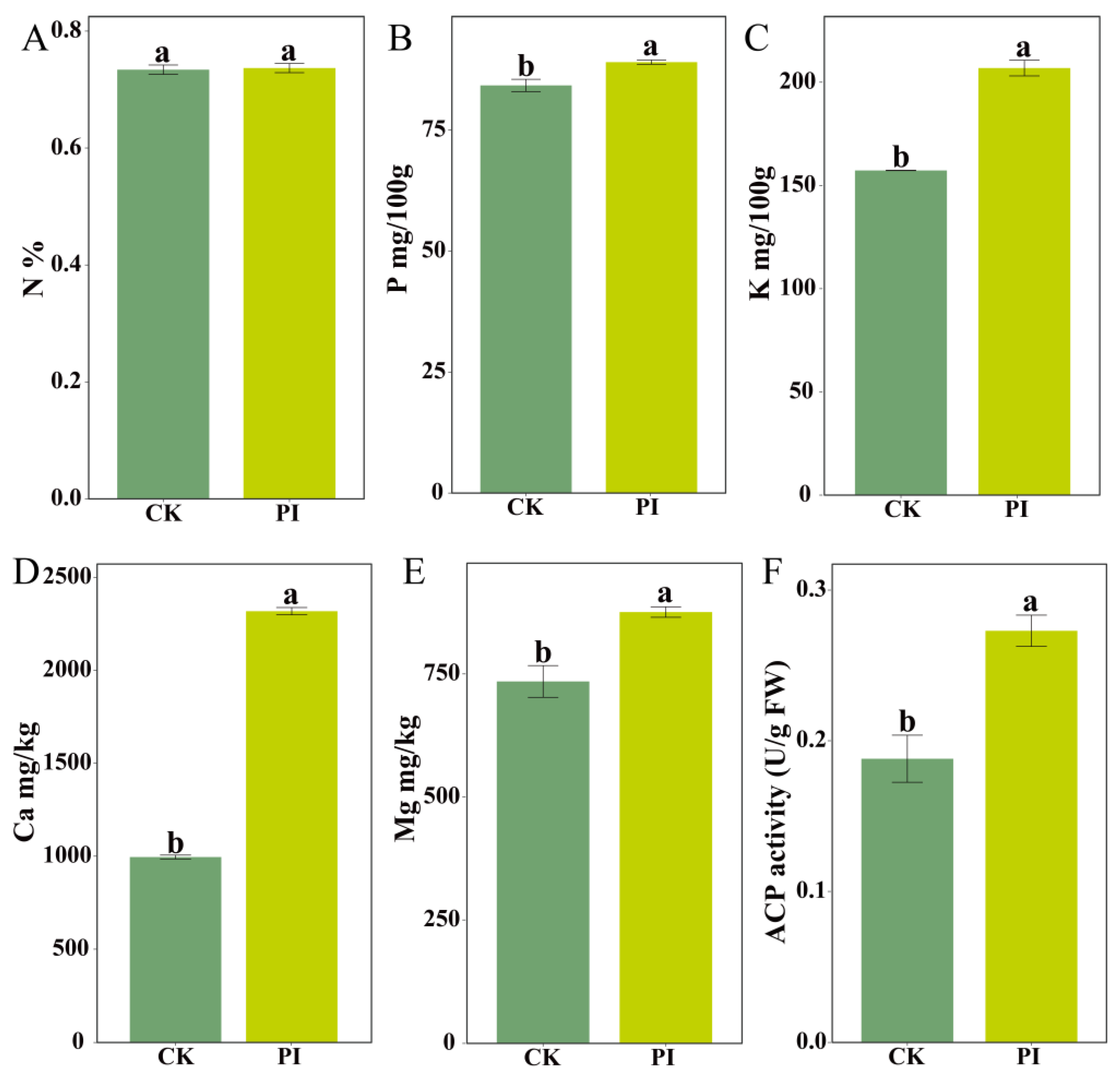
2.5. Validation of Key Biochemical Parameters Associated with Pathways Significantly Enriched by DEGs
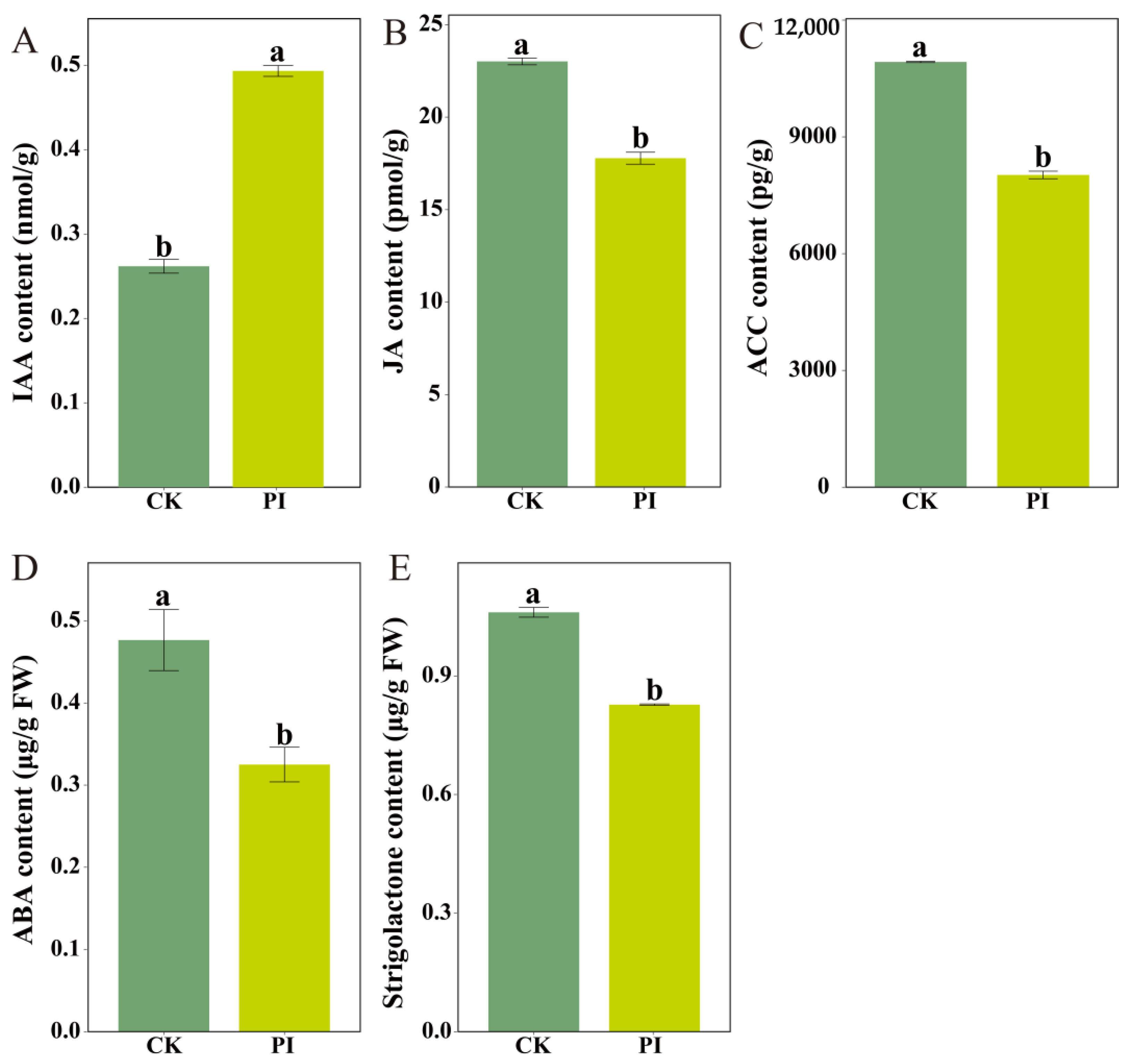
2.6. Determination of Endogenous Phytohormone Contents in CK and PI Blueberry Roots
3. Discussion
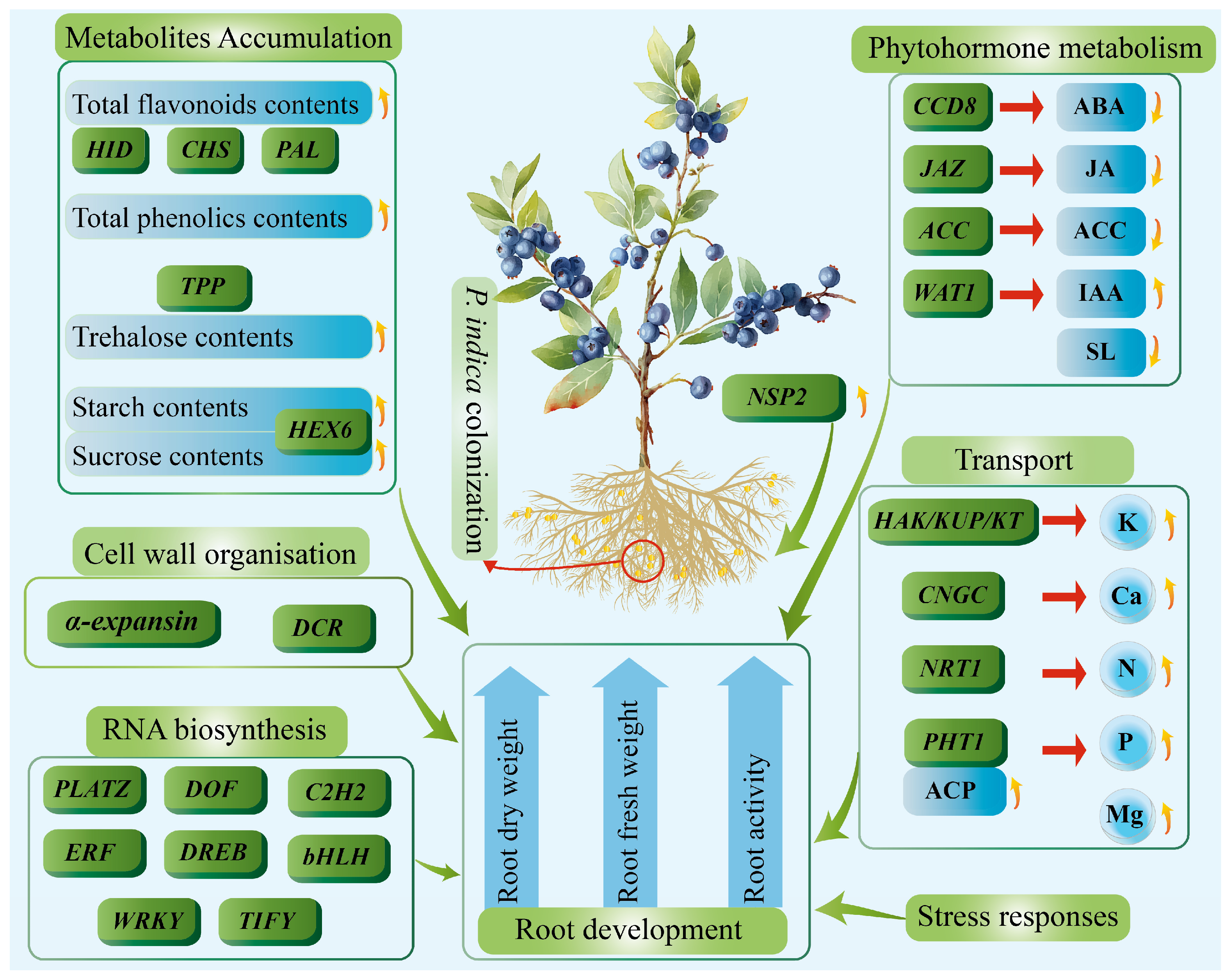
3.1. P. indica Colonization Modulates the Symbiotic Signaling Pathway and Cell Wall Metabolism in Blueberry Roots
3.2. P. indica Colonization Promotes Blueberry Root Development Through Modulating Carotenoid and Phytohormone Metabolism, Improving Secondary Metabolism, and Strengthening Stress Responses
3.3. P. indica Promotes Blueberry Root Development by Enhancing Mineral Element Absorption and Sugar Transport Metabolism
4. Materials and Methods
4.1. Plant Materials and P. indica Inoculation
4.2. Determination of Root Fresh and Dry Weight and Root Activity
4.3. Transcriptome Sequencing
4.4. Identification and Enrichment Analysis of DEGs
4.5. Gene Expression Validation by Using Quantitative Real-Time PCR (qRT-PCR)
4.6. Determination of ACP Activity and Mineral Element Contents in Blueberry Roots
4.7. Measurement of Sucrose, Starch, Total Phenolic, Total Flavonoids, and Phytohormones Contents in Blueberry Roots
4.8. Root Carotenoids Profiling and Analysis
4.9. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| ABA | Abscisic acid |
| ACC | 1-aminocyclopropane-1-carboxylic acid |
| ACP | Acid phosphatase |
| AMF | Arbuscular mycorrhizal fungus |
| bHLH | Basic helix–loop–helix |
| BP | Biological processes |
| bp | Base pair |
| Ca | Calcium |
| CC | Cellular components |
| CCD8 | Carotenoid cleavage dioxygenase 8 |
| cDNA | Complementary DNA |
| C2H2 | Cys2-His2 zinc finger |
| CHS | Chalcone synthase |
| CNGC | Cyclic nucleotide-gated channel |
| DEGs | Differentially expressed genes |
| DOF | DNA-binding with one finger |
| DREB | Dehydration-responsive element binding protein |
| EC | Electrical conductivity |
| ERF | Ethylene response factor |
| EXPA | Alpha-class expansin |
| Gb | Gigabase |
| GO | Gene ontology |
| HEX6 | Hexose carrier 6 |
| HID | 2-hydroxyisoflavone dehydratase |
| IAA | Indole-3-acetic acid |
| JA | Jasmonic acid |
| K | Potassium |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| LRR-XII | Leucine-rich repeat receptor kinases |
| MF | Molecular function |
| Mg | Magnesium |
| N | Nitrogen |
| NSP2 | Nodulation signaling pathway 2 |
| P | Phosphorus |
| PAL | Phenylalanine ammonia-lyase |
| PI | Piriformospora indica-colonized |
| PLATZ | Plant AT-rich sequence and zinc-binding protein |
| qRT-PCR | Quantitative real-time PCR |
| SL | Strigolactone |
| TF | Transcription factor |
| TPP | Trehalose-phosphate phosphatase |
| TTC | Triphenyltetrazolium chloride |
| WAT1 | Wall are thin 1 |
References
- Varma, A.; Savita, V.; Sudha; Sahay, N.; Butehorn, B.; Franken, P. Piriformospora indica, a cultivable plant-growth-promoting root endophyte. Appl. Environ. Microbiol. 1999, 65, 2741–2744. [Google Scholar] [CrossRef]
- Yang, L.; Zou, Y.-N.; Tian, Z.-H.; Wu, Q.-S.; Kuča, K. Effects of beneficial endophytic fungal inoculants on plant growth and nutrient absorption of trifoliate orange seedlings. Sci. Hortic. 2021, 277, 109815. [Google Scholar] [CrossRef]
- Hu, J.R.; Li, J.M.; Wang, H.Y.; Sun, M.L.; Huang, C.Y.; Wang, H.C. Analysis of growth dynamics in five different media and metabolic phenotypic characteristics of Piriformospora indica. Front. Microbiol. 2023, 14, 1301743. [Google Scholar] [CrossRef] [PubMed]
- Mensah, R.A.; Li, D.; Liu, F.; Tian, N.; Sun, X.; Hao, X.; Lai, Z.; Cheng, C. Versatile Piriformospora indica and its potential applications in horticultural crops. Hortic. Plant J. 2020, 6, 111–121. [Google Scholar] [CrossRef]
- Druege, U.; Baltruschat, H.; Franken, P. Piriformospora indica promotes adventitious root formation in cuttings. Sci. Hortic. 2007, 112, 422–426. [Google Scholar] [CrossRef]
- Li, D.; Mensah, R.A.; Liu, F.; Tian, N.; Qi, Q.; Yeh, K.; Xuhan, X.; Cheng, C.; Lai, Z. Effects of Piriformospora indica on rooting and growth of tissue-cultured banana (Musa acuminata cv. Tianbaojiao) seedlings. Sci. Hortic. 2019, 257, 108649. [Google Scholar] [CrossRef]
- Wu, H.; Wang, B.; Hao, X.; Zhang, Y.; Wang, T.; Lu, Z.; Lai, Z.; Cheng, C. Piriformospora indica promotes the growth and enhances the root rot disease resistance of gerbera. Sci. Hortic. 2022, 297, 110946. [Google Scholar] [CrossRef]
- Rong, Z.-Y.; Zhang, Z.-Z.; Alqahtani, M.D.; Wu, Q.-S.; Gao, X.-B. Serendipita indica is a biostimulant that improves tea growth at low P levels by modulating P acquisition and hormone levels. Rhizosphere 2023, 28, 100796. [Google Scholar] [CrossRef]
- Yin, L.; Qu, P.; Wang, D.; Yan, S.; Gong, Q.; Yang, R.; Hu, Y.; Liu, N.; Cheng, C.; Wang, P.; et al. The influence of Piriformospora indica colonization on the root development and growth of Cerasus humilis cuttings. Plants 2024, 13, 1482. [Google Scholar] [CrossRef]
- Qu, P.; Zhang, Z.; Li, R.; Liu, R.; Zhang, Y.; Cheng, C. Insights into the rooting and growth-promoting effects of endophytic fungus Serendipita indica in blueberry (Vaccinium corymbosum). J. Plant Growth Regul. 2025, 44, 2235–2246. [Google Scholar] [CrossRef]
- Boorboori, M.R.; Zhang, H.Y. The role of Serendipita indica (Piriformospora indica) in improving plant resistance to drought and salinity stresses. Biology 2022, 11, 952. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Wang, X.; Zhang, Z.; Li, C.; Xing, Y.; Luo, Y.; Li, D.; Ma, Z.; Cai, H. Symbiotic system establishment between Piriformospora indica and Glycine max and its effects on the antioxidant activity and ion-transporter-related gene expression in soybean under salt stress. Int. J. Mol. Sci. 2022, 23, 14961. [Google Scholar] [CrossRef] [PubMed]
- Sabeem, M.; Abdul Aziz, M.; Mullath, S.K.; Brini, F.; Rouached, H.; Masmoudi, K. Enhancing growth and salinity stress tolerance of date palm using Piriformospora indica. Front. Plant Sci. 2022, 13, 1037273. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Abdul Aziz, M.; Sabeem, M.; Kutty, M.S.; Sivasankaran, S.K.; Brini, F.; Xiao, T.T.; Blilou, I.; Masmoudi, K. Date palm transcriptome analysis provides new insights on changes in response to high salt stress of colonized roots with the endophytic fungus Piriformospora indica. Front. Plant Sci. 2024, 15, 1400215. [Google Scholar] [CrossRef]
- Xu, Z.; Pehlivan, N.; Ghorbani, A.; Wu, C. Effects of Azorhizobium caulinodans and Piriformospora indica co-inoculation on growth and fruit quality of tomato (Solanum lycopersicum L.) under salt stress. Horticulturae 2022, 8, 302. [Google Scholar] [CrossRef]
- Abdelaziz, M.E.; Kim, D.; Ali, S.; Fedoroff, N.V.; Al-Babili, S. The endophytic fungus Piriformospora indica enhances Arabidopsis thaliana growth and modulates Na(+)/K(+) homeostasis under salt stress conditions. Plant Sci. 2017, 263, 107–115. [Google Scholar] [CrossRef]
- Balbontín, C.; Flores, S.; Reyes, M.; Urrutia, V.; Parra-Palma, C.; Morales-Quintana, L.; Ramos, P. Antarctic fungal inoculation enhances drought tolerance and modulates fruit physiology in blueberry plants. Curr. Plant Biol. 2025, 42, 100462. [Google Scholar] [CrossRef]
- Su, H.; Guo, Y.; Gu, L.; Shi, X.; Zhou, Y.; Wu, F.; Wang, L. A novel growth-promoting dark septate endophytic fungus improved drought tolerance in blueberries by modulating phytohormones and non-structural carbohydrates. Tree Physiol. 2024, 44, tpae105. [Google Scholar] [CrossRef]
- Qu, D.; Wu, F.; Zhao, X.; Zhu, D.; Gu, L.; Yang, L.; Zhao, W.; Sun, Y.; Yang, J.; Tian, W.; et al. A bZIP transcription factor VabZIP12 from blueberry induced by dark septate endocyte improving the salt tolerance of transgenic Arabidopsis. Plant Sci. 2022, 315, 111135. [Google Scholar] [CrossRef]
- Trzewik, A.; Marasek-Ciolakowska, A.; Orlikowska, T. Protection of highbush blueberry plants against Phytophthora cinnamomi using Serendipita indica. Agronomy. 2020, 10, 1598. [Google Scholar] [CrossRef]
- Zhang, Y.; Qu, P.; Zhang, J.; Li, R.; Liu, R.; Cheng, C. Insights into the underlying mechanism of the Piriformospora indica-enhanced drought tolerance in blueberry. Horticulturae 2025, 11, 605. [Google Scholar] [CrossRef]
- Ma, W.; Wang, Y.; Li, K.; Lu, C.; Hou, D.; Li, Y.; Liu, H.; Wu, L.; Li, J. Physiological responses and assessment of salt tolerance of different blueberry cultivars under chloride stress. Agronomy 2025, 15, 494. [Google Scholar] [CrossRef]
- Song, H.; Cao, Y.; Zhao, X.; Zhang, L. Na+-preferential ion transporter HKT1;1 mediates salt tolerance in blueberry. Plant Physiol. 2023, 194, 511–529. [Google Scholar] [CrossRef] [PubMed]
- Machado, R.M.A.; Bryla, D.R.; Vargas, O. Effects of salinity induced by ammonium sulfate fertilizer on root and shoot growth of highbush blueberry. In Acta Horiculturae; ISHS: Leuven, Belgium, 2014; pp. 407–414. [Google Scholar]
- Kalo, P.; Gleason, C.; Edwards, A.; Marsh, J.; Mitra, R.M.; Hirsch, S.; Jakab, J.; Sims, S.; Long, S.R.; Rogers, J.; et al. Nodulation signaling in legumes requires NSP2, a member of the GRAS family of transcriptional regulators. Science 2005, 308, 1786–1789. [Google Scholar] [CrossRef] [PubMed]
- Fonouni-Farde, C.; Tan, S.; Baudin, M.; Brault, M.; Wen, J.; Mysore, K.S.; Niebel, A.; Frugier, F.; Diet, A. DELLA-mediated gibberellin signalling regulates Nod factor signalling and rhizobial infection. Nat. Commun. 2016, 7, 12636. [Google Scholar] [CrossRef]
- Ariel, F.; Brault-Hernandez, M.; Laffont, C.; Huault, E.; Brault, M.; Plet, J.; Moison, M.; Blanchet, S.; Ichante, J.L.; Chabaud, M.; et al. Two direct targets of cytokinin signaling regulate symbiotic nodulation in Medicago truncatula. Plant Cell 2012, 24, 3838–3852. [Google Scholar] [CrossRef]
- Kovacs, S.; Fodor, L.; Domonkos, A.; Ayaydin, F.; Laczi, K.; Rakhely, G.; Kalo, P. Amino acid polymorphisms in the VHIID conserved motif of nodulation signaling pathways 2 distinctly modulate symbiotic signaling and nodule morphogenesis in Medicago truncatula. Front. Plant Sci. 2021, 12, 709857. [Google Scholar] [CrossRef]
- Liu, W.; Kohlen, W.; Lillo, A.; Op den Camp, R.; Ivanov, S.; Hartog, M.; Limpens, E.; Jamil, M.; Smaczniak, C.; Kaufmann, K.; et al. Strigolactone biosynthesis in Medicago truncatula and rice requires the symbiotic GRAS-type transcription factors NSP1 and NSP2. Plant Cell 2011, 23, 3853–3865. [Google Scholar] [CrossRef]
- Marowa, P.; Ding, A.; Kong, Y. Expansins: Roles in plant growth and potential applications in crop improvement. Plant Cell Rep. 2016, 35, 949–965. [Google Scholar] [CrossRef]
- McQueen-Mason, S.; Durachko, D.M.; Cosgrove, D.J. Two endogenous proteins that induce cell wall extension in plants. Plant Cell 1992, 4, 1425–1433. [Google Scholar]
- Cosgrove, D.J.; Bedinger, P.; Durachko, D.M. Group I allergens of grass pollen as cell wall-loosening agents. Proc. Natl. Acad. Sci. USA. 1997, 94, 6559–6564. [Google Scholar] [CrossRef]
- Sampedro, J.; Cosgrove, D.J. The expansin superfamily. Genome Biol. 2005, 6, 242. [Google Scholar] [CrossRef][Green Version]
- Bashline, L.; Lei, L.; Li, S.; Gu, Y. Cell wall, cytoskeleton, and cell expansion in higher plants. Mol. Plant 2014, 7, 586–600. [Google Scholar] [CrossRef]
- Xu, H.; Lantzouni, O.; Bruggink, T.; Benjamins, R.; Lanfermeijer, F.; Denby, K.; Schwechheimer, C.; Bassel, G.W. A molecular signal integration network underpinning Arabidopsis seed germination. Curr. Biol. 2020, 30, 3703–3712.e4. [Google Scholar] [CrossRef] [PubMed]
- Cho, H.T.; Cosgrove, D.J. Regulation of root hair initiation and expansin gene expression in Arabidopsis. Plant Cell 2002, 14, 3237–3253. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.W.; Kim, J. EXPANSINA17 up-regulated by LBD18/ASL20 promotes lateral root formation during the auxin response. Plant Cell Physiol. 2013, 54, 1600–1611. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wang, Y.; Liu, M.; Yan, P.; Niu, F.; Ma, F.; Hu, J.; He, S.; Cui, J.; Yuan, X.; et al. OsEXPA7 encoding an expansin affects grain size and quality traits in rice (Oryza sativa L.). Rice 2024, 17, 36. [Google Scholar] [CrossRef]
- Jadamba, C.; Kang, K.; Paek, N.C.; Lee, S.I.; Yoo, S.C. Overexpression of rice expansin7 (osexpa7) confers enhanced tolerance to salt stress in rice. Int. J. Mol. Sci. 2020, 21, 454. [Google Scholar] [CrossRef]
- ZhiMing, Y.; Bo, K.; XiaoWei, H.; ShaoLei, L.; YouHuang, B.; WoNa, D.; Ming, C.; Hyung-Taeg, C.; Ping, W. Root hair-specific expansins modulate root hair elongation in rice. Plant J. 2011, 66, 725–734. [Google Scholar] [CrossRef]
- Li, Y.; Chen, Y.; Fu, Y.; Shao, J.; Liu, Y.; Xuan, W.; Xu, G.; Zhang, R. Signal communication during microbial modulation of root system architecture. J. Exp. Bot. 2024, 75, 526–537. [Google Scholar] [CrossRef]
- Wei, Z.; Li, J. Brassinosteroids regulate root growth, development, and symbiosis. Mol. Plant. 2016, 9, 86–100. [Google Scholar] [CrossRef] [PubMed]
- Ranocha, P.; Denance, N.; Vanholme, R.; Freydier, A.; Martinez, Y.; Hoffmann, L.; Kohler, L.; Pouzet, C.; Renou, J.P.; Sundberg, B.; et al. Walls are thin 1 (WAT1), an Arabidopsis homolog of Medicago truncatula NODULIN21, is a tonoplast-localized protein required for secondary wall formation in fibers. Plant J. 2010, 63, 469–483. [Google Scholar] [CrossRef] [PubMed]
- Ranocha, P.; Dima, O.; Nagy, R.; Felten, J.; Corratge-Faillie, C.; Novak, O.; Morreel, K.; Lacombe, B.; Martinez, Y.; Pfrunder, S.; et al. Arabidopsis WAT1 is a vacuolar auxin transport facilitator required for auxin homoeostasis. Nat. Commun. 2013, 4, 2625. [Google Scholar] [CrossRef]
- Ladwig, F.; Stahl, M.; Ludewig, U.; Hirner, A.A.; Hammes, U.Z.; Stadler, R.; Harter, K.; Koch, W. Siliques are Red1 from Arabidopsis acts as a bidirectional amino acid transporter that is crucial for the amino acid homeostasis of siliques. Plant Physiol. 2012, 158, 1643–1655. [Google Scholar] [CrossRef]
- Fiorilli, V.; Wang, J.Y.; Bonfante, P.; Lanfranco, L.; Al-Babili, S. Apocarotenoids: Old and new mediators of the arbuscular mycorrhizal symbiosis. Front. Plant Sci. 2019, 10, 1186. [Google Scholar] [CrossRef]
- Singh, S.; Sakshi; Annapurna, K.; Shrivastava, N.; Varma, A. Symbiotic interplay of Piriformospora indica and Azotobacter chroococcum augments crop productivity and biofortification of Zinc and Iron. Microbiol. Res. 2022, 262, 127075. [Google Scholar] [CrossRef]
- Ke, D.; Guo, J.; Li, K.; Wang, Y.; Han, X.; Fu, W.; Miao, Y.; Jia, K.P. Carotenoid-derived bioactive metabolites shape plant root architecture to adapt to the rhizospheric environments. Front. Plant Sci. 2022, 13, 986414. [Google Scholar] [CrossRef]
- Yin, C.C.; Ma, B.; Collinge, D.P.; Pogson, B.J.; He, S.J.; Xiong, Q.; Duan, K.X.; Chen, H.; Yang, C.; Lu, X.; et al. Ethylene responses in rice roots and coleoptiles are differentially regulated by a carotenoid isomerase-mediated abscisic acid pathway. Plant Cell 2015, 27, 1061–1081. [Google Scholar] [CrossRef]
- Mashiguchi, K.; Seto, Y.; Yamaguchi, S. Strigolactone biosynthesis, transport and perception. Plant J. 2021, 105, 335–350. [Google Scholar] [CrossRef]
- Akiyama, K.; Matsuzaki, K.; Hayashi, H. Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 2005, 435, 824–827. [Google Scholar] [CrossRef] [PubMed]
- Nawaz, Z.; Kakar, K.U.; Ullah, R.; Yu, S.; Zhang, J.; Shu, Q.Y.; Ren, X.L. Genome-wide identification, evolution and expression analysis of cyclic nucleotide-gated channels in tobacco (Nicotiana tabacum L.). Genomics 2019, 111, 142–158. [Google Scholar] [CrossRef]
- Saand, M.A.; Xu, Y.P.; Li, W.; Wang, J.P.; Cai, X.Z. Cyclic nucleotide gated channel gene family in tomato: Genome-wide identification and functional analyses in disease resistance. Front. Plant Sci. 2015, 6, 303. [Google Scholar] [CrossRef]
- Tian, W.; Hou, C.; Ren, Z.; Wang, C.; Zhao, F.; Dahlbeck, D.; Hu, S.; Zhang, L.; Niu, Q.; Li, L.; et al. A calmodulin-gated calcium channel links pathogen patterns to plant immunity. Nature 2019, 572, 131–135. [Google Scholar] [CrossRef]
- Meena, M.K.; Prajapati, R.; Krishna, D.; Divakaran, K.; Pandey, Y.; Reichelt, M.; Mathew, M.K.; Boland, W.; Mithofer, A.; Vadassery, J. The Ca(2+) Channel CNGC19 regulates Arabidopsis defense against Spodoptera herbivory. Plant Cell 2019, 31, 1539–1562. [Google Scholar] [CrossRef] [PubMed]
- Charpentier, M.; Sun, J.; Vaz Martins, T.; Radhakrishnan, G.V.; Findlay, K.; Soumpourou, E.; Thouin, J.; Very, A.A.; Sanders, D.; Morris, R.J.; et al. Nuclear-localized cyclic nucleotide-gated channels mediate symbiotic calcium oscillations. Science 2016, 352, 1102–1105. [Google Scholar] [CrossRef] [PubMed]
- Brost, C.; Studtrucker, T.; Reimann, R.; Denninger, P.; Czekalla, J.; Krebs, M.; Fabry, B.; Schumacher, K.; Grossmann, G.; Dietrich, P. Multiple cyclic nucleotide-gated channels coordinate calcium oscillations and polar growth of root hairs. Plant J. 2019, 99, 910–923. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.Q.; Yang, Y.; Zhang, A.; Fei, C.F.; Gu, L.L.; Sun, S.J.; Xu, W.; Wang, L.; Liu, H.; Wang, Y.F. Three CNGC family members, CNGC5, CNGC6, and CNGC9, are required for constitutive growth of Arabidopsis root hairs as Ca(2+)-permeable channels. Plant Commun. 2020, 1, 100001. [Google Scholar] [CrossRef]
- Zhang, S.; Pan, Y.; Tian, W.; Dong, M.; Zhu, H.; Luan, S.; Li, L. Arabidopsis CNGC14 mediates calcium influx required for tip growth in root hairs. Mol. Plant 2017, 10, 1004–1006. [Google Scholar] [CrossRef]
- Zeb, Q.; Wang, X.; Hou, C.; Zhang, X.; Dong, M.; Zhang, S.; Zhang, Q.; Ren, Z.; Tian, W.; Zhu, H.; et al. The interaction of CaM7 and CNGC14 regulates root hair growth in Arabidopsis. J. Integr. Plant Biol. 2020, 62, 887–896. [Google Scholar] [CrossRef]
- Wu, M.; Wei, Q.; Xu, L.; Li, H.; Oelmüller, R.; Zhang, W. Piriformospora indica enhances phosphorus absorption by stimulating acid phosphatase activities and organic acid accumulation in Brassica napus. Plant Soil. 2018, 432, 333–344. [Google Scholar] [CrossRef]
- Su, Z.Z.; Wang, T.; Shrivastava, N.; Chen, Y.Y.; Liu, X.; Sun, C.; Yin, Y.; Gao, Q.K.; Lou, B.G. Piriformospora indica promotes growth, seed yield and quality of Brassica napus L. Microbiol. Res. 2017, 199, 29–39. [Google Scholar]
- Johnson, D.A.; Thomas, M.A. The monosaccharide transporter gene family in Arabidopsis and rice: A history of duplications, adaptive evolution, and functional divergence. Mol. Biol. Evol. 2007, 24, 2412–2423. [Google Scholar] [CrossRef]
- Ahmad, S.; Aftab, M.N.; Aslam, M.S.; Rehman, A.U.; Haq, I.U.; Ali, S.; Usman, M. Trehalose–abscisic acid pathway feedback loops in wheat—Piriformospora indica symbiosis: Mechanisms and drought resilience. Microbe 2025, 9, 100561. [Google Scholar] [CrossRef]
- Cheng, C.; Li, D.; Wang, B.; Liao, B.; Qu, P.; Liu, W.; Zhang, Y.; Lü, P. Piriformospora indica colonization promotes the root growth of Dimocarpus longan seedlings. Sci. Hortic. 2022, 301, 111137. [Google Scholar] [CrossRef]
- Mylavarapu, R.; Hochmuth, G.; Mackowiak, C.; Wright, A.; Silveira, M. Lowering soil pH to optimize nutrient management and crop production. EDIS 2016, 2016, 4. [Google Scholar] [CrossRef]
- Bütehorn, B.; Rhody, D.; Franken, P. Isolation and characterisation of Pitef1 encoding the translation elongation factor EF-1α of the root endophyte Piriformospora indica. Plant Biol. 2008, 2, 687–692. [Google Scholar] [CrossRef]
- Tanaka, J.; Kiyoshi, K.; Kadokura, T.; Suzuki, K.-I.; Nakayama, S. Elucidation of the enzyme involved in 2,3,5-triphenyl tetrazolium chloride (TTC) staining activity and the relationship between TTC staining activity and fermentation profiles in Saccharomyces cerevisiae. J. Biosci. Bioeng. 2021, 131, 396–404. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Jin, X.; Lin, H.; He, J.; Chen, Y. Comparative transcriptome sequencing and endogenous phytohormone content of annual grafted branches of Zelkova schneideriana and its dwarf variety Hentiangao. Int. J. Mol. Sci. 2023, 24, 16902. [Google Scholar] [CrossRef]
- Zhang, Z.; Qu, P.; Hao, S.; Li, R.; Zhang, Y.; Zhao, Q.; Wen, P.; Cheng, C. Characterization and functional analysis of chalcone synthase genes in highbush blueberry (Vaccinium corymbosum). Int. J. Mol. Sci. 2023, 24, 13882. [Google Scholar] [CrossRef]
- Jaakola, L.; Maatta, K.; Pirttila, A.M.; Torronen, R.; Karenlampi, S.; Hohtola, A. Expression of genes involved in anthocyanin biosynthesis in relation to anthocyanin, proanthocyanidin, and flavonol levels during bilberry fruit development. Plant Physiol. 2002, 130, 729–739. [Google Scholar] [CrossRef]
- Kirk, P.L. Kjeldahl Method for Total Nitrogen. Anal. Chem. 1950, 22, 354–358. [Google Scholar] [CrossRef]
- Brookes, P.C.; Powlson, D.S. Preventing phosphorus losses during perchloric acid digestion of sodium bicarbonate soil extracts. J. Sci. Food Agr. 2006, 32, 671–674. [Google Scholar] [CrossRef]
- Roy, S.J.; Huang, W.; Wang, X.J.; Evrard, A.; Schmockel, S.M.; Zafar, Z.U.; Tester, M. A novel protein kinase involved in Na(+) exclusion revealed from positional cloning. Plant Cell Environ. 2013, 36, 553–568. [Google Scholar] [CrossRef]
- Oliveira, S.R.; Gomes Neto, J.A.; Nóbrega, J.A.; Jones, B.T. Determination of macro- and micronutrients in plant leaves by high-resolution continuum source flame atomic absorption spectrometry combining instrumental and sample preparation strategies. Spectrochim. Acta B. 2010, 65, 316–320. [Google Scholar] [CrossRef]
- Perez, M.; Dominguez-Lopez, I.; Lamuela-Raventos, R.M. The chemistry behind the Folin-Ciocalteu method for the estimation of (Poly)phenol content in food: Total phenolic intake in a mediterranean dietary pattern. J. Agric. Food Chem. 2023, 71, 17543–17553. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.C.; Yang, M.H.; Wen, H.M.; Chern, J.C. Estimation of total flavonoid content in propolis by two complementary colometric methods. J. Biosci. Bioeng. 2002, 10, 3. [Google Scholar] [CrossRef]
| Root Growth Parameters | CK Group | PI Group |
|---|---|---|
| Root fresh weight (g) | 72.33 ± 7.76 b | 100.66 ± 5.85 a |
| Root dry weight (g) | 26.33 ± 3.05 a | 32.66 ± 4.04 a |
| Root activity (μg·g−1·h−1) | 4.15 ± 0.55 a | 5.85 ± 1.01 a |
| BIN ID | BIN Name | All DEGs | Up | Down | p-Value |
|---|---|---|---|---|---|
| 15.5.7 | RNA biosynthesis; transcriptional regulation; and AP2/ERF transcription factor superfamily. | 73 | 6 | 67 | 1.19 × 10−4 |
| 18.4.1.12 | Protein modification; phosphorylation.; TKL protein kinase superfamily; and protein kinase (LRR-XII). | 18 | 15 | 3 | 1.19 × 10−4 |
| 24.1.2 | Solute transport; primary active transport; and P-type ATPase superfamily. | 20 | 0 | 20 | 1.19 × 10−4 |
| 24.1.2.2 | Solute transport; primary active transport; P-type ATPase superfamily; and P2 family. | 20 | 0 | 20 | 1.19 × 10−4 |
| 24.1.2.2.2 | Solute transport; primary active transport; P-type ATPase superfamily; P2 family; and P2B-type calcium cation-transporting ATPase (ACA). | 20 | 0 | 20 | 1.19 × 10−4 |
| 15.5.1.5 | RNA biosynthesis; transcriptional regulation; C2C2 transcription factor superfamily; and transcription factor (DOF). | 9 | 9 | 0 | 8.34 × 10−4 |
| 21.4.2.1 | Cell wall organization; cell wall proteins; expansin activities; and alpha-class expansin | 10 | 10 | 0 | 8.48 × 10−4 |
| 15.5.7.2 | RNA biosynthesis; transcriptional regulation; AP2/ERF transcription factor superfamily; and transcription factor (DREB). | 43 | 3 | 40 | 1.98 × 10−3 |
| 11.9 | Phytohormone action and strigolactone. | 8 | 8 | 0 | 2.50 × 10−3 |
| 24.1 | Solute transport and primary active transport. | 44 | 1 | 43 | 1.01 × 10−3 |
| 24.2.1.1 | Solute transport; carrier-mediated transport; DMT superfamily; and NST-TPT group. | 13 | 0 | 13 | 1.01 × 10−2 |
| 24.2.1.5 | Solute transport; carrier-mediated transport; DMT superfamily; and solute transporter (UmamiT). | 7 | 7 | 0 | 1.01 × 10−2 |
| 11.7.3.3 | Phytohormone action; jasmonic acid; conjugation and degradation; and jasmonoyl-amino acid carboxylase (CYP94C). | 5 | 0 | 5 | 1.01 × 10−2 |
| 15.5.42 | RNA biosynthesis; transcriptional regulation; and transcription factor (TIFY). | 15 | 0 | 15 | 1.01 × 10−2 |
| 7.9.6 | Coenzyme metabolism; NAD/NADP biosynthesis; and NAD homeostasis. | 7 | 0 | 7 | 1.19 × 10−2 |
| 7.9.6.2 | Coenzyme metabolism; NAD/NADP biosynthesis; NAD homeostasis; and pyrophosphohydrolase (NUDX). | 7 | 0 | 7 | 1.19 × 10−2 |
| 3.3 | Carbohydrate metabolism and trehalose metabolism. | 7 | 0 | 7 | 1.49 × 10−2 |
| 3.3.2 | Carbohydrate metabolism; trehalose metabolism; and trehalose-6-phosphate phosphatase. | 7 | 0 | 7 | 1.49 × 10−2 |
| 15.5.40 | RNA biosynthesis; transcriptional regulation; and transcription factor (PLATZ). | 6 | 6 | 0 | 1.66 × 10−2 |
| 15.5.1 | RNA biosynthesis; transcriptional regulation; and C2C2 transcription factor superfamily. | 16 | 10 | 6 | 1.92 × 10−2 |
| 27.1.4 | Multi-process regulation; circadian clock system; and time-of-day-dependent expressed repressor (PRR). | 7 | 7 | 0 | 2.04 × 10−2 |
| 24.2.1.1.11 | Solute transport; carrier-mediated transport; DMT superfamily; NST-TPT group; and nucleotide sugar transporter (UUAT). | 9 | 0 | 9 | 2.46 × 10−2 |
| 21.4.2 | Cell wall organization; cell wall proteins; and expansin activities. | 19 | 10 | 9 | 2.46 × 10−2 |
| 19.2.2.1.4.2 | Protein homeostasis; ubiquitin–proteasome system; and ubiquitin-fold protein conjugation; ubiquitin conjugation (ubiquitylation); ubiquitin-ligase E3 activities; and U-Box E3 ligase activities. | 49 | 0 | 49 | 3.28 × 10−2 |
| 26.10.1.6 | External stimuli response; symbiont; common symbiotic signaling pathway (CSSP); and NSP1-NSP2 nodulation initiation complex. | 4 | 4 | 0 | 3.28 × 10−2 |
| 26.10.1.6.2 | External stimuli response; symbiont; common symbiotic signaling pathway (CSSP); NSP1-NSP2 nodulation initiation complex; and component NSP2. | 4 | 4 | 0 | 3.28 × 10−2 |
| 27.1 | Multi-process regulation and circadian clock system. | 9 | 7 | 2 | 3.51 × 10−2 |
| 26.4 | External stimuli response and temperature. | 17 | 0 | 17 | 3.81 × 10−2 |
| 26.4.3 | External stimuli response; temperature; and cold response. | 17 | 0 | 17 | 3.81 × 10−2 |
| 15.5.22 | RNA biosynthesis; transcriptional regulation; and WRKY transcription factor activity. | 41 | 0 | 41 | 3.82 × 10−2 |
| 15.5.22.1 | RNA biosynthesis; transcriptional regulation; WRKY transcription factor activity; and transcription factor (WRKY). | 41 | 0 | 41 | 3.82 × 10−2 |
| 9.1.4 | Secondary metabolism; terpenoids; and terpene biosynthesis. | 14 | 0 | 14 | 3.92 × 10−2 |
| 26.4.3.4 | External stimuli response; temperature; cold response; and ICE-CBF cold acclimation transcriptional cascade. | 14 | 0 | 14 | 4.09 × 10−2 |
| 26.4.3.4.2 | External stimuli response; temperature; cold response; ICE-CBF cold acclimation transcriptional cascade; and transcription factor (CBF/DREB1). | 14 | 0 | 14 | 4.09 × 10−2 |
| 15.5.30 | RNA biosynthesis; transcriptional regulation; and transcription factor (bHLH). | 21 | 1 | 20 | 4.82 × 10−2 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Guo, S.; Qu, P.; Du, S.; Liu, R.; Zhang, Y.; Cheng, C. Integrated Transcriptomic and Biochemical Analyses Reveal the Root Development-Promoting Mechanism of Piriformospora indica on Blueberry Under Tap Water Irrigation. Plants 2025, 14, 3646. https://doi.org/10.3390/plants14233646
Guo S, Qu P, Du S, Liu R, Zhang Y, Cheng C. Integrated Transcriptomic and Biochemical Analyses Reveal the Root Development-Promoting Mechanism of Piriformospora indica on Blueberry Under Tap Water Irrigation. Plants. 2025; 14(23):3646. https://doi.org/10.3390/plants14233646
Chicago/Turabian StyleGuo, Sijian, Pengyan Qu, Shitao Du, Rui Liu, Yongyan Zhang, and Chunzhen Cheng. 2025. "Integrated Transcriptomic and Biochemical Analyses Reveal the Root Development-Promoting Mechanism of Piriformospora indica on Blueberry Under Tap Water Irrigation" Plants 14, no. 23: 3646. https://doi.org/10.3390/plants14233646
APA StyleGuo, S., Qu, P., Du, S., Liu, R., Zhang, Y., & Cheng, C. (2025). Integrated Transcriptomic and Biochemical Analyses Reveal the Root Development-Promoting Mechanism of Piriformospora indica on Blueberry Under Tap Water Irrigation. Plants, 14(23), 3646. https://doi.org/10.3390/plants14233646






