Abstract
Plant diseases caused by pathogens result in a marked decrease in crop yield and quality annually, greatly threatening food production and security worldwide. The creation and cultivation of disease-resistant cultivars is one of the most effective strategies to control plant diseases. Broad-spectrum resistance (BSR) is highly preferred by breeders because it confers plant resistance to diverse pathogen species or to multiple races or strains of one species. Recently, accumulating evidence has revealed the roles of 2-oxoglutarate (2OG)-dependent oxygenases (2OGDs) as essential regulators of plant disease resistance. Indeed, 2OGDs catalyze a large number of oxidative reactions, participating in the plant-specialized metabolism or biosynthesis of the major phytohormones and various secondary metabolites. Moreover, several 2OGD genes are characterized as negative regulators of plant defense responses, and the disruption of these genes via genome editing tools leads to enhanced BSR against pathogens in crops. Here, the recent advances in the isolation and identification of defense-related 2OGD genes in plants and their exploitation in crop improvement are comprehensively reviewed. Also, the strategies for the utilization of 2OGD genes as targets for engineering BSR crops are discussed.
1. Introduction
Plant diseases caused by phytopathogens lead to substantial yield losses and a notable reduction in crop quality annually, becoming one of the important factors limiting agricultural production worldwide. The global crop losses due to pathogens and pests were estimated to be approximately 30.0% (24.6–40.9%) in rice, 22.5% (19.5–41.1%) in maize, 21.5% (10.1–28.1%) in wheat, 21.4% (11.0–32.4%) in soybean, and 17.2% (8.1–21.0%) in potato, respectively [1]. As a countermeasure, plants possess a two-layered immune surveillance system against a wide variety of invading microbes, including pattern-triggered immunity (PTI), activated through the recognition of highly conserved pathogen-associated molecular patterns (PAMPs) by cell surface-localized pattern-recognition receptors (PRRs), and effector-triggered immunity (ETI), initiated by intracellular resistance (R) proteins that directly or indirectly sense pathogen-encoded virulence effectors [2,3,4,5,6]. Although the activation mechanisms and the early signaling cascades of PTI and ETI are different, these two types of plant immunity converge on overlapping downstream cellular responses, including the induction of defense genes, the activation of mitogen-activated protein kinases (MAPKs), the accumulation of reactive oxygen species (ROS), and callose deposition [3,7].
Broad-spectrum resistance (BSR) is defined as plant disease resistance against at least two types of pathogen species or multiple members of one species [8]. Many genes involved in BSR have been identified and characterized in plants, such as those encoding membrane-associated PRRs, intracellular resistance proteins, defense-regulator proteins, pathogenesis-related (PR) proteins, susceptibility proteins, and nonhost resistance proteins [9]. For instance, FLAGELLIN SENSING 2 (FLS2), RLK-PRR EF-TU RECEPTOR (EFR), Xa21, lysin motif-containing proteins 4 (LYP4), LYP6, and elicitin response (ELR) function as PRRs to activate plant defense responses via sensing the matched PAMPs, and the ectopic expression of these genes in other plant species induces BSR to multiple pathogens [10,11,12,13,14,15,16,17,18,19,20,21,22,23,24]. Overexpressing the antimicrobial protein genes CaAMP1 or LjAMP2 in transgenic lines led to enhanced resistance against multiple fungal pathogens [25,26,27]. TaPsIPK1, encoding a receptor-like cytoplasmic protein kinase employed by the rust effector PsSpg1, has been found to be an important susceptibility gene in wheat, and the knockout of this gene confers BSR against Puccinia striiformis f. sp. tritici (Pst) races CYR32, CYR33, and CYR34 [28]. NONHOST 1 (NHO1), PENETRATION1 (PEN1), PEN2, and PEN3 are four reported nonhost resistance genes that contribute to BSR against several host-nonspecific pathogens [29,30,31,32,33,34,35]. Additionally, quantitative trait loci (QTLs) conferring BSR have been cloned in different plants. Two QTLs, Lr67/Yr46 and Lr34/Yr18/Pm38, were found to positively regulate BSR against various fungal pathogens that cause powdery mildew, stripe rust, and leaf rust in wheat [36,37,38,39,40]. The cucumber-resistant QTL dm1/psl/cla, a loss-of-susceptibility mutation of the STAYGREEN (CsSGR) gene, confers BSR to the obligate biotrophic oomycete Pseudoperonospora cubensis, the hemibiotrophic fungal Colletotrichum orbicular, and the bacterial Pseudomonas syringae pv. Lachrymans [41].
Moreover, 2-oxoglutarate (2OG)-dependent oxygenases (2OGDs) are soluble non-heme proteins distributed ubiquitously throughout the kingdoms of life, and they catalyze a variety of oxidative reactions, including hydroxylation, epimerization, demethylation, sequential oxidation, ring formation, ring expansion, cyclization, halogenation, epoxidation, and desaturation [42,43,44,45,46,47]. Commonly, 2OGDs utilize ferrous iron Fe(II) as a cofactor and employ 2OG and molecular oxygen (O2) as cosubstrates to catalyze the oxidation of a substrate, generating the desired product, carbon dioxide (CO2), and succinate. The 2OGD family can be categorized into three functionally distinct groups (designated as DOXA, DOXB, and DOXC) based on phylogenetic analyses [43]. The DOXA group includes the homologs of Escherichia coli AlkB, a DNA repair enzyme, and demethylates the alkylated DNA, RNA, and histones [48,49,50,51]. The DOXB group containing a prolyl 4-hydroxylase (P4Hc) domain functions as the key enzymes in the biosynthesis of collagen [52] and also catalyzes the post-translational modifications of the proline hydroxylation of plant cell wall proteins [53]. The DOXC group, the most functionally diverse and largest subfamily of plant 2OGDs, participates in the specialized biosynthesis or metabolism of various phytohormones and secondary metabolites, such as gibberellins, auxin, ethylene, salicylic acid (SA), jasmonic acid (JA), flavonoids, tropane alkaloids, benzylisoquinoline alkaloids, coumarins, and glucosinolates, etc. [42,43,47].
In this review, the landmark discoveries of the roles of 2OGD genes in plant immunity are revisited, and the recent advances regarding 2OGD-mediated BSR are also summarized. Additionally, the molecular mechanisms of how 2OGDs confer BSR to pathogens and the application of 2OGDs in crop improvement for disease resistance are discussed.
2. The Discoveries of 2OGD-Mediated Broad-Spectrum Disease Resistance
In 2005, eight downy mildew-resistant (dmr) mutants corresponding to six different loci were found to show reduced susceptibility to the downy mildew pathogen Hyaloperonospora parasitica via screening the ethyl methane sulfonate (EMS) mutants in the highly susceptible genetic background of the Arabidopsis line Ler eds1-2 [54]. Among them, DMR6 was then isolated via map-based cloning and characterized as At5g24530, which encodes an oxidoreductase belonging to the 2OG-Fe(II)-dependent oxygenase superfamily [55]. Intriguingly, the loss of function mutant downy mildew-resistant 6 (dmr6) also exhibited elevated resistance against the oomycete Phytophthora capsici (P. capsici) and the bacterium Pseudomonas syringae (P. syringae), and overexpressing DMR6 in Arabidopsis led to enhanced susceptibility to Hyaloperonospora arabidopsidis (H. arabidopsidis), P. capsici, and P. syringae [56]. Additionally, the overexpression of DMR6-LIKE OXYGENASE 1 (DLO1) and DMR6-LIKE OXYGENASE 2 (DLO2), two closely related homologues of DMR6, restored the susceptibility of the dmr6 mutant to downy mildew H. arabidopsidis, similar to DMR6 [56]. These important findings confirm that the DMR6 and DMR6 homologues serve as negative regulators of BSR in Arabidopsis, and represent the first example of 2OGD functioning as a key component in BSR, which has attracted many colleagues to focus on advancing our understanding of this new field.
The knockdown of StDMR6 via RNAi silencing and the knockout of StDMR6-1 in potatoes resulted in enhanced resistance against the necrotrophic pathogen Botrytis cinerea (B. cinerea) and the hemibiotrophic pathogen Phytophthora infestans (P. infestans), respectively [57,58]. Sldmr6-1 mutants exhibited increased disease resistance against three evolutionarily different classes of pathogens, including a fungus (Pseudoidium neolycopersici), an oomycete (P. capsici), and bacteria (P. syringae pv. tomato, Xanthomonas gardneri, and Xanthomonas perforans) [59]. The mutagenesis of ObDMR6 confers resistance to the downy mildew pathogen Peronospora belbahrii in sweet basil, which is supported by the decreased pathogen biomass and sporangia production on the infected leaves [60]. The ectopic overexpression of AhS5H1 or AhS5H2, the first susceptible gene characterized in peanut, blocked the induction of defense-related genes (AtPR1 and AtPR2) upon chitin treatment and enhanced the susceptibility to P. syringae pv. tomato DC3000 in Arabidopsis [61]. The introduction of Hv2OGO into dmr6 mutants fully restored the susceptibility of Arabidopsis to Fusarium graminearum [62]. The inactivation of MusaDMR6 promoted the banana resistance against Xanthomonas campestris pv. musacearum, the causal agent of a devastating bacterial disease called Banana Xanthomonas wilt (BXW) [63]. In addition, OsSAH2 and OsSAH3 were recently reported to negatively regulate the rice resistance to Magnaporthe oryzae (M. oryzae), Xanthomonas oryzae pv. Oryzae, Bipolaris oryzae, and Rhizoctonia solani [64,65,66].
Apart from the DMR6 and DMR6 orthologs, other members of the DOXC subfamily have also been found to play essential roles in plant immunity. The transcripts of various 1-aminocyclopropane-1-carboxylic acid oxidase (ACO) genes were induced by different pathogens in plants [67,68,69,70], suggesting the involvement of ACOs in the plant defense against pathogen infections. Silencing of NbACO1 enhanced the host susceptibility of Nicotiana benthamiana to the hemibiotrophic fungi Colletotrichum orbiculare, with an accelerated switch from biotrophy to necrotrophy during the course of infection [67]. TuACO3 encodes the 1-aminocyclopropane-1-carboxylic acid (ACC) oxidase and has been confirmed to positively regulate the wheat defense against the powdery mildew pathogen Blumeria graminis f. sp. tritici [70]. Four JA-induced 2OGDs named JASMONIC ACID OXIDASES (JAO1-4 or JOX1-4) act as important suppressors of Arabidopsis defenses against the fungal pathogen B. cinerea and the larvae of Mamestra brassicae [71,72]. JAO2-deficient lines display the constitutive expression of marker genes for the JA defense cascade and increased accumulation of multiple classes of antimicrobial compounds [72,73]. Furthermore, the resistance phenotype of jao2 mutants can be recovered by the ectopic overexpression of JAO2, JAO3, or JAO4 [72,73].
The information regarding the 2OGD genes described above is summarized in Table 1.

Table 1.
2OGD genes involved in plant disease resistance.
3. The Roles of Phytohormones in Broad-Spectrum Disease Resistance
SA acts as an important player in the activation of plant innate immunity, including PTI, ETI, and systemic acquired resistance (SAR), which restricts the growth of invading pathogens [74,75]. The endogenous SA levels are demonstrated to be increased in plants upon stimulation with distinct pathogens, and SA-induction-insensitive mutants display compromised pathogen resistance [76,77,78,79,80,81,82,83,84,85]. The disruption of endogenous SA accumulation via ectopically expressing the bacterial salicylate hydroxylase gene, NahG, results in decreased resistance to the biotrophic and semi-biotrophic pathogens in transgenic tobacco and Arabidopsis plants [86,87]. Furthermore, the application of exogenous SA or SA analogs enhances the defense responses in plants against infectious pathogens [61,88,89,90,91]. Plants employ the primary metabolite chorismate as a precursor to synthesize SA via two different pathways with multiple steps, the isochorismate (IC) pathway and the phenylalanine ammonia-lyase (PAL) pathway [75,92]. Both the IC pathway and the PAL pathway contribute to the pathogen-induced SA accumulation and its functions in plant resistance because the inactivation mutants in the key components of these two pathways display lowered SA levels and enhanced susceptibility to many pathogens [78,79,84,85]. The alterations in the endogenous SA levels are sensed by the SA receptors (non-expressers of PR genes, NPRs), and the response signals are then transduced to activate the expression of the downstream PR genes, which encode anti-microbial proteins. In Arabidopsis, NPR1 is well-known to positively regulate the expression of downstream defense-related genes; however, NPR3 and NPR4 act as negative players in SA-mediated plant defense [93,94,95]. SA can be reversibly or irreversibly converted to bioactive derivatives via glycosylation, sulfonation, AA conjugation, methylation, and hydroxylation [75], thus helping plants to fine-tune SA homeostasis.
JA, the lipid-derived phytohormone, plays an essential role in the plant immunity against necrotrophic pathogens. The infection of these pathogens triggers the rapid accumulation of JA and jasmonoyl-L-isoleucine (JA-Ile), the major bioactive derivative of JA, which activate the downstream signaling to upregulate the production of the defense-related secondary metabolites and proteins in plants, such as alkaloids, phenylpropanoids, terpenoids, and several types of PR proteins [96,97]. JA is formed from 12-oxophytodienoic acid (OPDA) via an enzymatic reaction of 2-oxophytodienoic acid reductases (OPRs), and OPDA is synthesized from chloroplast membrane lipids by sequential catalytic reactions of lipoxygenases, allene oxide synthase, and allene oxide cyclase in different intracellular compartments [98,99]. JA can be further conjugated with isoleucine to synthesize JA-Ile via JA amido synthetase 1 (JAR1) in the cytosol [98,99]. The binding of JA-Ile with its co-receptor, the CORONATINE INSENSITIVE1 (COI1)/JASMONATE ZIM DOMAIN (JAZ) complex, leads to the degradation of JAZ through the 26S proteasome pathway, thereby releasing the suppression of the JAZ-interacting transcription factor MYC2 that activates various JA-responsive genes [98,99,100]. The JA biosynthetic and insensitive mutant plants displayed a remarkable reduction in resistance to necrotrophic pathogens. Arabidopsis fad3/fad7/fad8 triple mutants were highly susceptible to the fungus Pythium mastophorum, and the exogenous application of methyl jasmonate could nearly restore the resistance against P. mastophorum [101]. In tomato, jasmonate-deficient mutants def1 showed enhanced susceptibility to five pathogens, including two bacteria (Xanthomonas campestris and P. syringae), two fungi (Fusarium oxysporum f. sp. lycopersici and Verticillium dahliae), and an oomycete (P. infestans) [102]. Maize opr7/opr8 double mutants lost the defensive ability of damping-off disease caused by the oomycete Pythium aristosporum [103]. The coi1 mutants failed to activate the pathogen-induced expression of PR-3, PR-4, and PDF1.2 in Arabidopsis, and exhibited elevated susceptibility to the fungal pathogens B. cinerea and Alternaria brassicicola but not Peronospora parasitica [104]. Although JA signaling was reported to function negatively regarding the defense against some hemibiotrophic bacteria in dicotyledonous plants [105,106], it also acts as a positive player in resistance against the hemibiotrophic pathogen M. oryzae in rice and viruses in both Arabidopsis and rice [107,108,109,110,111,112,113]. For instance, JA accumulation was triggered by the coat protein (CP) of Rice stripe virus (RSV), and then JAMYB, a transcription factor downstream of the JA signaling, upregulated the expression of Argonaute 18 (AGO18), thereby activating the RNA silencing pathway for the rice antiviral defense [113].
The synthesis of ethylene starts from methionine (Met), and the amino acid is catalyzed to S-adenosyl-methionine (AdoMet) via the S-AdoMet synthase [114,115,116]. AdoMet is then catalyzed to 1-aminocyclopropane-1-carboxylic acid (ACC) through the 1-aminocyclopropane-1-carboxylate synthase (ACS), followed by the formation of ethylene from ACC via the ACC oxidase (ACO) [114,115,116]. Ethylene biosynthesis was induced by M. oryzae, and the increased ethylene production enhanced the rice blast disease resistance [68,117,118]. Furthermore, the inhibition of ethylene biosynthesis by aminoethoxyvinylglycine (AVG) decreased the rice resistance to blast [68,117,119]. Intriguingly, transgenic rice lines expressing OsACS2 by a strong pathogen-induced promoter showed significant accumulation of ethylene upon M. oryzae infection and exhibited elevated resistance against rice blast and sheath blight [120]. A recent report found that PTI- and ETI-inducible OsMETS1, a methionine synthase deubiquitinated and stabilized by OsPICI1, promoted blast resistance via increasing the ethylene levels in rice [119]. Moreover, ethylene was confirmed to positively regulate the disease resistance against the hemibiotrophic and biotrophic pathogens in other plants, including soybean, tobacco, A. thaliana, and wheat [70,121,122,123]. Apart from ethylene accumulation, the components of ethylene signaling also contribute to plants’ defense against pathogens. For instance, overexpression of ERF, the ethylene responsive factor gene, elevated the host resistance to invading pathogens in many plants [124,125,126,127,128,129,130,131,132]. However, there are some cases showing that ethylene and ethylene signaling play negative regulatory roles in plant defense responses [133,134,135,136,137]. It seems that ethylene functions as a negative or positive regulator of disease resistance, relying on the pathogen type, plant species, and environmental conditions, which may be explained via the intricate mediation of the JA signaling and SA signaling through ethylene interaction [138,139].
4. Regulating Plant BSR with 2OGDs via Altering the Levels of Phytohormones
The dmr6 mutants did not exhibit the constitutive expression of PR-1, and the pathogen growth was inhibited in the dmr6 mutants without the accumulation of reactive oxygen intermediates (ROI) or obvious cell death [54], suggesting the important role of DMR6 in the cellular processes during pathogen infection. Subsequently, dmr6-mediated resistance was deduced from the increased expression of some defense-related genes or the altered levels of the DMR6 substrate and product [55]. It is worth mentioning that the SA levels in wild-type plants are approximately 10 times lower than in the dmr6 mutant and more than 200 times lower than in the dmr6-3_dlo1 double mutant [56]. Notably, DLO1 was identified and characterized as an SA 3-hydroxylase (S3H) that regulates leaf senescence via modulating the SA catabolism in Arabidopsis (Figure 1a) [140]. DLO1/S3H catalyzes SA to both 2,5-dihydroxybenzoic acid (2,5-DHBA) and 2,3-DHBA in vitro but only 2,3-DHBA in planta (Figure 1a) [140]. Given that DMR6 functions redundantly with DLO1 and the SA accumulation in the dmr6 mutant is higher than that in the dlo1 mutant, it is possible that DMR6 also catabolizes SA in a similar way to DLO1. As expected, DMR6 was discovered and functionally characterized as an SA 5-hydroxylase (S5H) via an enzymatic assay procedure (Figure 1a) [141]. DMR6/S5H catalyzes the specific hydroxylation of SA to 2,5-DHBA in vitro and in planta, with higher catalytic efficiency than DLO1/S3H [141].
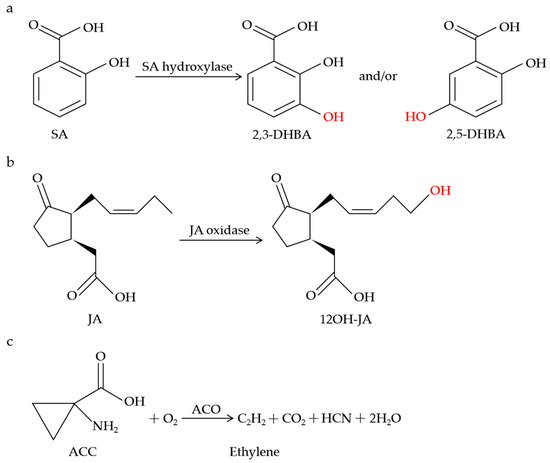
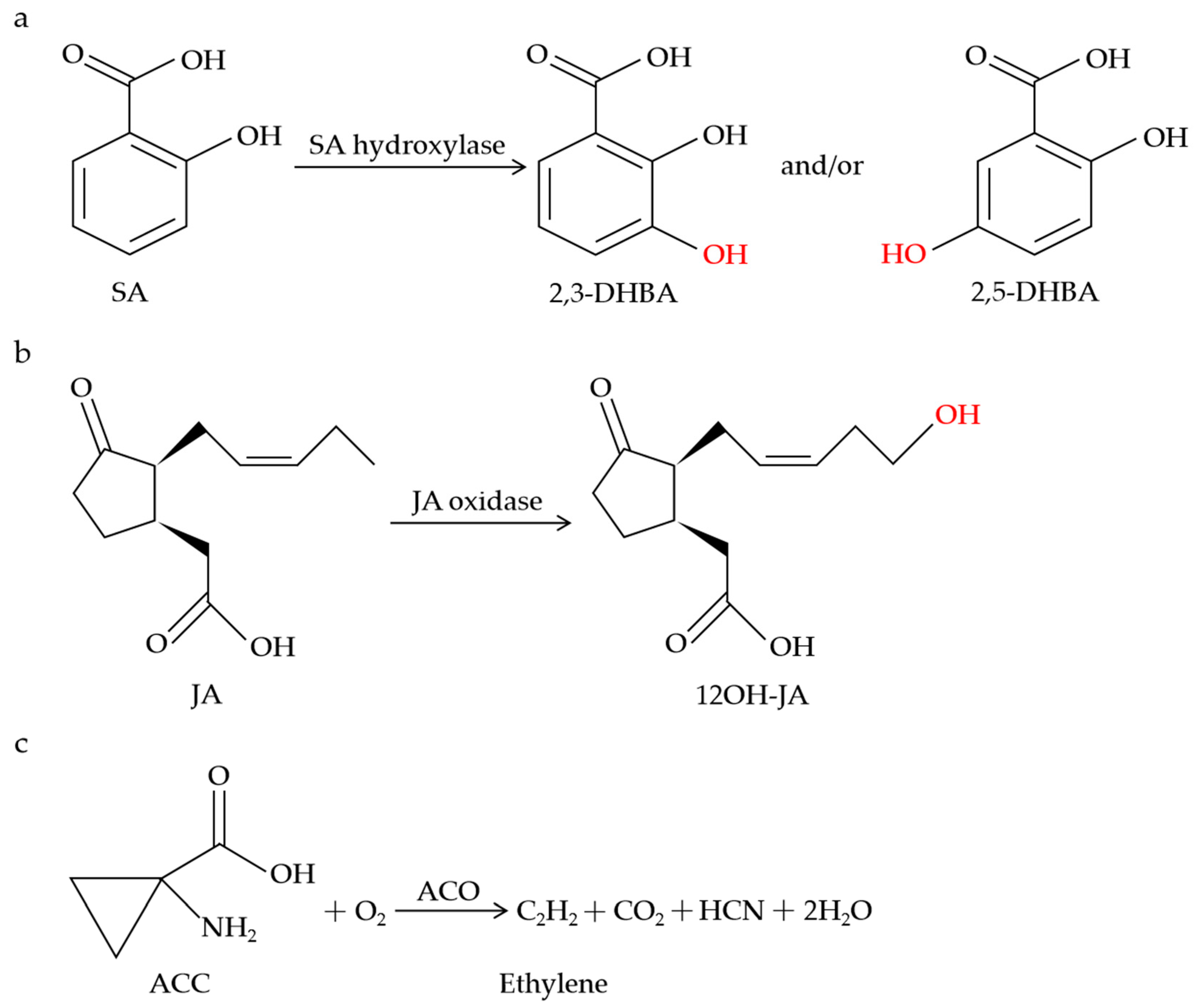
Figure 1.
Schematic of metabolic conversions of SA, JA, and ethylene by 2OGDs. SA hydroxylase catalyzes SA to 2,3-DHBA and/or 2,5-DHBA (a). JA oxidase catalyzes the specific hydroxylation of JA to 12OH-JA (b). ACO catalyzes the biosynthesis of ethylene (c).
JAO1-4 clustered between DLO1 (S3H) and amino-cyclopropane oxidase (ACCox), belonging to a four-member subclade named DOX 46 [43,72]. Based on this phylogenetic proximity, it was hypothesized that JAO1-4 could function in the oxygenation or hydroxylation reactions using hormone compounds as substrates. In support of this hypothesis, JAO2-4 specifically oxidized JA to 12OH-JA in vitro (Figure 1b), with no catalytic activity towards indole-acetic acid (IAA), SA, 12-oxophytodienoic acid (OPDA), or JA-Ile [72]. In B. cinerea-infected leaves, the accumulation of 12OH-JA was strongly decreased in the jao2, jao3, and jao4 mutants relative to the WT, indicating that JAO2, JAO3, and JAO4 contribute to the production of 12OH-JA upon B. cinerea stimulation [72]. Furthermore, molecular genetic evidence showed that the constitutive expression of ORA59, PDF1.2, and PR4 in the jao2 mutants was strongly reduced or abolished by the inactivation of JAR1 or COI1, clearly demonstrating that the jao2 defense phenotype relies on the generation and recognition of JA-Ile [72].
ACO catalyzes the final step of the ethylene biosynthetic pathway in which the ethylene precursor ACC is converted to ethylene and hydrogen cyanide (Figure 1c) [114,115,116]. It has been found that ACO-mediated resistance is associated with the formation of ethylene. Upon blast fungus (Magnaporthe grisea) infection, both the OsACO7 mRNA transcripts and the ACO activity were highly increased, accompanied by elevated ethylene emissions, contributing to the resistance of young rice plants [68]. The rapid activation of NtACO1 and NtACO2 expression coincided with a dramatic increase in the ethylene levels and the activation of the NtMEK2-SIPK/WIPK cascade-mediated defense responses in tobacco plants infected by Tobacco mosaic virus (TMV) [142]. Also, a marked increase in the ACO activity and ACO gene expression was observed in tomato plants inoculated with Xanthomonas campestris pv. vesicatoria (XCV), similar to ethylene biosynthesis [143]. TuACO3 was reported to upregulate the production of ethylene to enhance the defense against powdery mildew in einkorn wheat [70]. TuACO3-silenced plants displayed reduced ethylene production and increased susceptibility to Bgt, whereas TuACO3-overexpressing transgenic lines exhibited elevated ethylene levels and enhanced resistance to Bgt in wheat [70]. Intriguingly, the expression of TuACO3 was suppressed by a transcription factor, TuMYB46L, and the downregulation of TuMYB46L in wheat following Bgt infection led to increased expression of TuACO3, thus accumulating more ethylene to promote the einkorn wheat defense [70].
5. Conclusions and Perspectives
Plant diseases cause devastating yield losses in various crops annually and become a major threat to global food production. Different management practices have been adopted to control the diseases, among which the generation and cultivation of disease-resistant varieties is usually thought to be one of the most effective and promising strategies. In addition, durable and broad-spectrum disease resistance can be obtained via the inactivation of plant susceptibility (S) genes that are exploited by pathogens to facilitate successful infection and proliferation [144,145]. Recently, the rapid development of genome editing technologies has made it possible to precisely and efficiently mutate plant endogenous genes [146], which provides a foundation for creating disease-resistant plants via editing target genes. Taking advantage of this technology, the disease resistance was enhanced in many plants through the dysfunction of the S gene. For instance, Mildew resistance locus O (Mlo) is a well-known S gene to powdery mildew and is ubiquitously present across almost all higher monocots and dicots [147,148,149]. Mlo encodes a membrane-anchored protein with seven conserved transmembrane domains and functions in the processes of PM fungal penetration [150,151]. So far, researchers have successfully created PM-resistant tomato, grapevine, wheat, and soybean via the targeted mutagenesis of Mlo utilizing the CRISPR/Cas9 system [152,153,154,155,156].
As described in this paper, some 2OGDs are S genes that negatively regulate the BSR in plants, including JAO1-4 (JOX1-4), DMR6, and DMR6 orthologs. This implies that these genes may be candidate targets for engineering crops with enhanced resistance to pathogens (Figure 2a). Until now, the CRISPR/Cas9-mediated mutation of a DMR6 ortholog has conferred BSR in tomato and rice [59,64,65,66]. Intriguingly, the disruption of the DMR6 ortholog does not influence the yield per plant of tomato and rice [59,65]. Moreover, elevated resistance to a single pathogen has been reported in banana, sweet basil, and potato via the inactivation of a DMR6 ortholog by the CRISPR/Cas9 system [58,60,63]. It still needs to be evaluated whether these mutants exhibit enhanced resistance to other pathogens. Considering that the mutation of the DMR6 ortholog may have potential negative effects on growth and yield, the CRISPR/Cas9-mediated knockdown of this gene represents another effective path for creating BSR crops without fitness costs. Recently, three amino acids in the substrate-binding sites were found to affect the enzyme activity of the DMR6 ortholog in rice, and substitutions of these amino acids led to a decrease in the oxidation of SA to 2,5-DHBA in vitro [64]. This suggests that base substitutions of the codes for key amino acids via base editors provide a possible strategy for enhancing the resistance of crops.
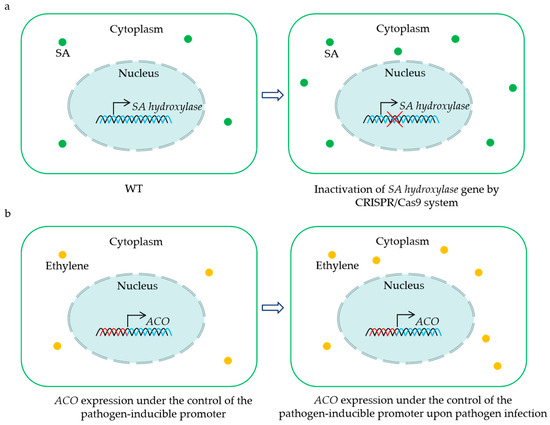
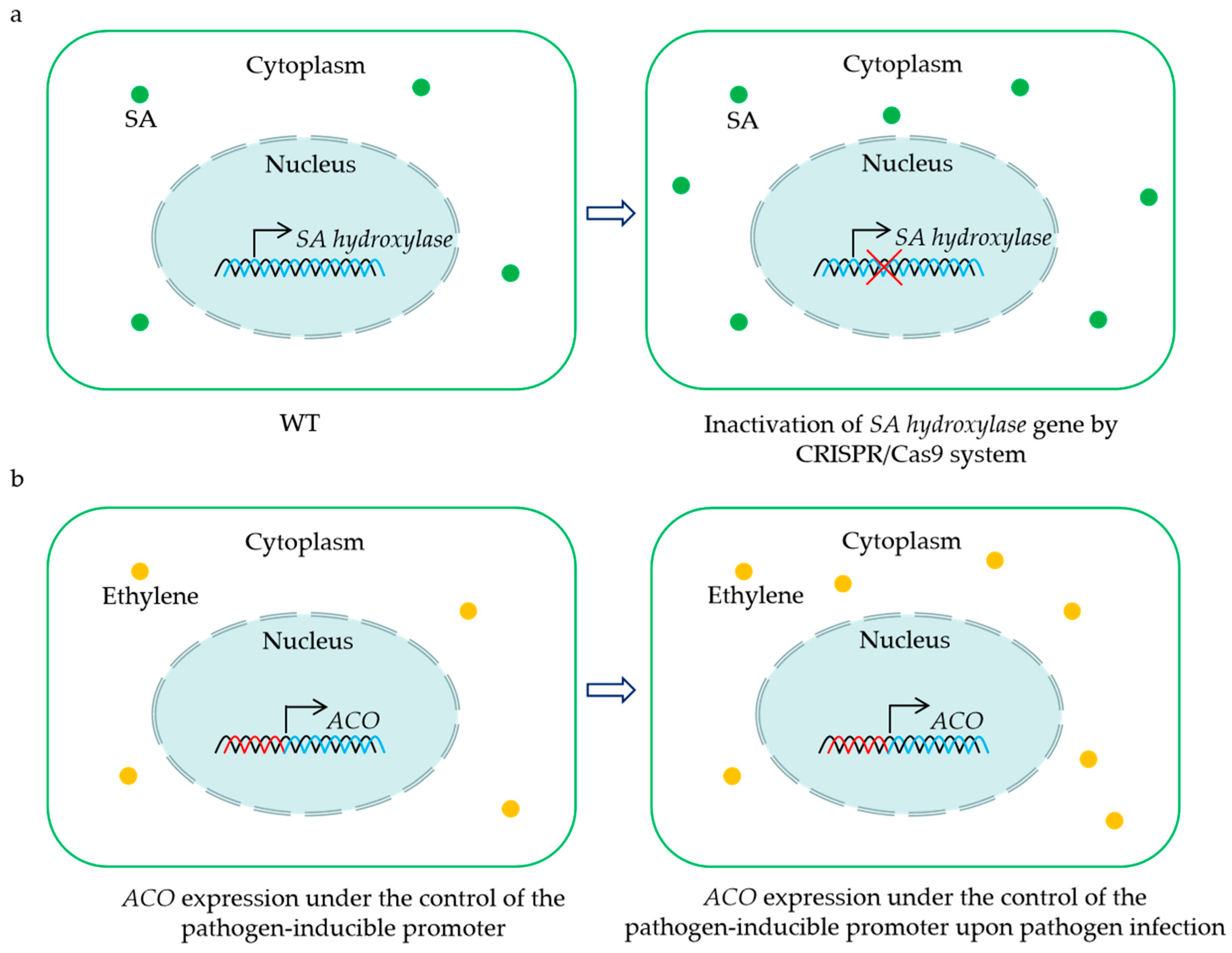
Figure 2.
Representative strategies for the utilization of 2OGD genes for creating crops with BSR. CRISPR/Cas9-mediated mutation of the SA hydroxylase gene results in sufficient SA accumulation to enhance disease resistance (a). Placing ACO gene under the control of the pathogen-inducible promoter leads to an induction of ethylene biosynthesis upon pathogen infection, and the increased levels of ethylene contribute to plant BSR (b). The green dots indicate salicylic acid (SA), and the yellow dots show ethylene.
The overexpression of the key positive regulator of immunity in plants is also recognized as one of the strategies to engineer BSR crops. There are successful cases where AtNPR1 overexpression has led to significantly enhanced BSR in diverse crop species without obvious alterations in their growth phenotype [157,158,159,160,161,162]. However, penalties in terms of plant development, yield, and virus resistance were also found in the transgenic strawberry or rice overexpressing AtNPR1, respectively [163,164,165]. Of note, expressing AtNPR1, which is controlled by the pathogen-induced promoter and the 5′ leader sequence, confers rice resistance to the bacterial leaf streak, blast, and blight in the field without fitness costs [166]. These findings establish an upgrade strategy for enhancing BSR in crops via the rapid and transient activation of plant defenses upon pathogen attacks. Based on this concept, we propose a possible method for engineering BSR crops by placing the ACO gene under the control of the pathogen-inducible promoter (Figure 2b). Although the positive role of ACO in BSR has been confirmed, this solution still needs to be further tested for controlling diseases and agricultural application under field conditions.
The 2OGD superfamily comprises a large number of members and ranks second in the plant enzyme family [43]. Generally, 2OGDs function in plants’ primary or secondary metabolism, which plays an essential role in plant defenses. However, there are only a few 2OGD genes that are demonstrated to act as important players in plant immunity. To enrich our knowledge regarding the roles of 2OGD genes in plant disease resistance, more research on the isolation and characterization of these genes is needed in the future. Moreover, understanding the detailed molecular mechanism of how 2OGD genes contribute to the plant immune response may provide novel insights for engineering BSR crops. Additionally, the rapid development and application of CRISPR/Cas9-mediated genome editing technology in different crops will accelerate the use of 2OGD genes to create BSR crops.
Author Contributions
W.F. and Q.C. conceptualized the project. H.W. and W.F. drafted the manuscript. All authors contributed to the editing of the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Hainan Provincial Natural Science Foundation of China (grant no. 321CXTD437 to Q.C.), the Scientific Research Foundation of Hainan University (grant no. KYQD(ZR)-22090 to W.F.), and the National Key R&D Program of China (grant no. 2024YFD1400100 to W.F.).
Data Availability Statement
Acknowledgments
We apologize for incomplete citations because of space limitations.
Conflicts of Interest
The authors declare no competing interests.
References
- Savary, S.; Willocquet, L.; Pethybridge, S.J.; Esker, P.; McRoberts, N.; Nelson, A. The global burden of pathogens and pests on major food crops. Nat. Ecol. Evol. 2019, 3, 430–439. [Google Scholar] [CrossRef] [PubMed]
- Jones, J.D.; Dangl, J.L. The plant immune system. Nature 2006, 444, 323–329. [Google Scholar] [CrossRef] [PubMed]
- Tsuda, K.; Katagiri, F. Comparing signaling mechanisms engaged in pattern-triggered and effector-triggered immunity. Curr. Opin. Plant Biol. 2010, 13, 459–465. [Google Scholar] [CrossRef] [PubMed]
- Cui, H.T.; Tsuda, K.; Parker, J.E. Effector-triggered immunity: From pathogen perception to robust defense. Annu. Rev. Plant Biol. 2015, 66, 487–511. [Google Scholar] [CrossRef] [PubMed]
- Couto, D.; Zipfel, C. Regulation of pattern recognition receptor signalling in plants. Nat. Rev. Immunol. 2016, 16, 537–552. [Google Scholar] [CrossRef]
- Yu, X.; Feng, B.M.; He, P.; Shan, L.B. From chaos to harmony: Responses and signaling upon microbial pattern recognition. Annu. Rev. Phytopathol. 2017, 55, 109–137. [Google Scholar] [CrossRef]
- Peng, Y.J.; van Wersch, R.; Zhang, Y.L. Convergent and divergent signaling in PAMP-triggered immunity and effector-triggered immunity. Mol. Plant Microbe Interact. 2018, 31, 403–409. [Google Scholar] [CrossRef]
- Kou, Y.J.; Wang, S.P. Broad-spectrum and durability: Understanding of quantitative disease resistance. Curr. Opin. Plant Biol. 2010, 13, 181–185. [Google Scholar] [CrossRef]
- Li, W.; Deng, Y.W.; Ning, Y.S.; He, Z.H.; Wang, G.L. Exploiting broad-spectrum disease resistance in crops: From molecular dissection to breeding. Annu. Rev. Plant Biol. 2020, 71, 575–603. [Google Scholar] [CrossRef] [PubMed]
- Forsyth, A.; Mansfield, J.W.; Grabov, N.; de Torres, M.; Sinapidou, E.; Grant, M.R. Genetic dissection of basal resistance to Pseudomonas syringae pv. phaseolicola in accessions of Arabidopsis. Mol. Plant Microbe Interact. 2010, 23, 1545–1552. [Google Scholar]
- Robatzek, S.; Chinchilla, D.; Boller, T. Ligand-induced endocytosis of the pattern recognition receptor FLS2 in Arabidopsis. Genes Dev. 2006, 20, 537–542. [Google Scholar] [CrossRef]
- Robatzek, S.; Bittel, P.; Chinchilla, D.; Köchner, P.; Felix, G.; Shiu, S.H.; Boller, T. Molecular identification and characterization of the tomato flagellin receptor LeFLS2, an orthologue of Arabidopsis FLS2 exhibiting characteristically different perception specificities. Plant Mol. Biol. 2007, 64, 539–547. [Google Scholar] [CrossRef]
- Kunze, G.; Zipfel, C.; Robatzek, S.; Niehaus, K.; Boller, T.; Felix, G. The N terminus of bacterial elongation factor Tu elicits innate immunity in Arabidopsis plants. Plant Cell 2004, 16, 3496–3507. [Google Scholar] [CrossRef]
- Lacombe, S.; Rougon-Cardoso, A.; Sherwood, E.; Peeters, N.; Dahlbeck, D.; van Esse, H.P.; Smoker, M.; Rallapalli, G.; Thomma, B.P.; Staskawicz, B.; et al. Interfamily transfer of a plant pattern-recognition receptor confers broad-spectrum bacterial resistance. Nat. Biotechnol. 2010, 28, 365–369. [Google Scholar] [CrossRef]
- Boschi, F.; Schvartzman, C.; Murchio, S.; Ferreira, V.; Siri, M.I.; Galván, G.A.; Smoker, M.; Stransfeld, L.; Zipfel, C.; Vilaró, F.L.; et al. Enhanced bacterial wilt resistance in potato through expression of Arabidopsis EFR and introgression of quantitative resistance from Solanum commersonii. Front. Plant Sci. 2017, 8, 1642. [Google Scholar] [CrossRef]
- Pfeilmeier, S.; George, J.; Morel, A.; Roy, S.; Smoker, M.; Stransfeld, L.; Downie, J.A.; Peeters, N.; Malone, J.G.; Zipfel, C. Expression of the Arabidopsis thaliana immune receptor EFR in Medicago truncatula reduces infection by a root pathogenic bacterium, but not nitrogen-fixing rhizobial symbiosis. Plant Biotechnol. J. 2019, 17, 569–579. [Google Scholar] [CrossRef]
- Schwessinger, B.; Bahar, O.; Thomas, N.; Holton, N.; Nekrasov, V.; Ruan, D.L.; Canlas, P.E.; Daudi, A.; Petzold, C.J.; Singan, V.R.; et al. Transgenic expression of the dicotyledonous pattern recognition receptor EFR in rice leads to ligand-dependent activation of defense responses. PLoS Pathog. 2015, 11, e1004809. [Google Scholar] [CrossRef]
- Song, W.Y.; Wang, G.L.; Chen, L.L.; Kim, H.S.; Pi, L.Y.; Holsten, T.; Gardner, J.; Wang, B.; Zhai, W.X.; Zhu, L.H.; et al. A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 1995, 270, 1804–1806. [Google Scholar] [CrossRef]
- Wang, G.L.; Song, W.Y.; Ruan, D.L.; Sideris, S.; Ronald, P.C. The cloned gene, Xa21, confers resistance to multiple Xanthomonas oryzae pv. oryzae isolates in transgenic plants. Mol. Plant Microbe Interact. 1996, 9, 850–855. [Google Scholar] [CrossRef]
- Holton, N.; Nekrasov, V.; Ronald, P.C.; Zipfel, C. The phylogenetically-related pattern recognition receptors EFR and XA21 recruit similar immune signaling components in monocots and dicots. PLoS Pathog. 2015, 11, e1004602. [Google Scholar] [CrossRef]
- Omar, A.A.; Murata, M.M.; El-Shamy, H.A.; Graham, J.H.; Grosser, J.W. Enhanced resistance to citrus canker in transgenic mandarin expressing Xa21 from rice. Transgenic Res. 2018, 27, 179–191. [Google Scholar] [CrossRef]
- Tripathi, J.N.; Lorenzen, J.; Bahar, O.; Ronald, P.; Tripathi, L. Transgenic expression of the rice Xa21 pattern-recognition receptor in banana (Musa sp.) confers resistance to Xanthomonas campestris pv. musacearum. Plant Biotechnol. J. 2014, 12, 663–673. [Google Scholar] [CrossRef]
- Liu, B.; Li, J.F.; Ao, Y.; Qu, J.W.; Li, Z.Q.; Su, J.B.; Zhang, Y.; Liu, J.; Feng, D.R.; Qi, K.B.; et al. Lysin motif-containing proteins LYP4 and LYP6 play dual roles in peptidoglycan and chitin perception in rice innate immunity. Plant Cell 2012, 24, 3406–3419. [Google Scholar] [CrossRef]
- Du, J.; Verzaux, E.; Chaparro-Garcia, A.; Bijsterbosch, G.; Keizer, L.C.; Zhou, J.; Liebrand, T.W.; Xie, C.; Govers, F.; Robatzek, S. Elicitin recognition confers enhanced resistance to Phytophthora infestans in potato. Nat. Plants 2015, 1, 15034. [Google Scholar] [CrossRef]
- Lee, S.C.; Hwang, I.S.; Choi, H.W.; Hwang, B.K. Involvement of the pepper antimicrobial protein CaAMP1 gene in broad spectrum disease resistance. Plant Physiol. 2008, 148, 1004–1020. [Google Scholar] [CrossRef]
- Jia, Z.C.; Gou, J.Q.; Sun, Y.M.; Yuan, L.; Tang, Q.; Yang, X.Y.; Pei, Y.; Luo, K.M. Enhanced resistance to fungal pathogens in transgenic Populus tomentosa Carr. by overexpression of an nsLTP-like antimicrobial protein gene from motherwort (Leonurus japonicus). Tree Physiol. 2010, 30, 1599–1605. [Google Scholar] [CrossRef]
- Yang, X.Y.; Li, J.; Li, X.B.; She, R.; Pei, Y. Isolation and characterization of a novel thermostable non-specific lipid transfer protein-like antimicrobial protein from motherwort (Leonurus japonicus Houtt) seeds. Peptides 2006, 27, 3122–3128. [Google Scholar] [CrossRef]
- Wang, N.; Tang, C.L.; Fan, X.; He, M.Y.; Gan, P.F.; Zhang, S.; Hu, Z.Y.; Wang, X.D.; Yan, T.; Shu, W.X.; et al. Inactivation of a wheat protein kinase gene confers broad-spectrum resistance to rust fungi. Cell 2022, 185, 2961–2974. [Google Scholar] [CrossRef]
- Kang, L.; Li, J.X.; Zhao, T.H.; Xiao, F.M.; Tang, X.Y.; Thilmony, R.; He, S.Y.; Zhou, J.M. Interplay of the Arabidopsis nonhost resistance gene NHO1 with bacterial virulence. Proc. Natl. Acad. Sci. USA 2003, 100, 3519–3524. [Google Scholar] [CrossRef]
- Lu, M.; Tang, X.Y.; Zhou, J.M. Arabidopsis NHO1 is required for general resistance against Pseudomonas bacteria. Plant Cell 2001, 13, 437–447. [Google Scholar] [CrossRef][Green Version]
- Assaad, F.F.; Qiu, J.L.; Youngs, H.; Ehrhardt, D.; Zimmerli, L.; Kalde, M.; Wanner, G.; Peck, S.C.; Edwards, H.; Ramonell, K.; et al. The PEN1 syntaxin defines a novel cellular compartment upon fungal attack and is required for the timely assembly of papillae. Mol. Biol. Cell 2004, 15, 5118–5129. [Google Scholar] [CrossRef] [PubMed]
- Collins, N.C.; Thordal-Christensen, H.; Lipka, V.; Bau, S.; Kombrink, E.; Qiu, J.L.; Hückelhoven, R.; Stein, M.; Freialdenhoven, A.; Somerville, S.C.; et al. SNARE-protein-mediated disease resistance at the plant cell wall. Nature 2003, 425, 973–977. [Google Scholar] [CrossRef] [PubMed]
- Lipka, V.; Dittgen, J.; Bednarek, P.; Bhat, R.; Wiermer, M.; Stein, M.; Landtag, J.; Brandt, W.; Rosahl, S.; Scheel, D.; et al. Pre-and postinvasion defenses both contribute to nonhost resistance in Arabidopsis. Science 2005, 310, 1180–1183. [Google Scholar] [PubMed]
- Stein, M.; Dittgen, J.; Sánchez-Rodríguez, C.; Hou, B.H.; Molina, A.; Schulze-Lefert, P.; Lipka, V.; Somerville, S. Arabidopsis PEN3/PDR8, an ATP binding cassette transporter, contributes to nonhost resistance to inappropriate pathogens that enter by direct penetration. Plant Cell 2006, 18, 731–746. [Google Scholar] [CrossRef] [PubMed]
- Loehrer, M.; Langenbach, C.; Goellner, K.; Conrath, U.; Schaffrath, U. Characterization of nonhost resistance of Arabidopsis to the Asian soybean rust. Mol. Plant Microbe Interact. 2008, 21, 1421–1430. [Google Scholar] [CrossRef] [PubMed]
- Krattinger, S.G.; Lagudah, E.S.; Spielmeyer, W.; Singh, R.P.; Huerta-Espino, J.; McFadden, H.; Bossolini, E.; Selter, L.L.; Keller, B. A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 2009, 323, 1360–1363. [Google Scholar] [PubMed]
- Lagudah, E.S.; Krattinger, S.G.; Herrera-Foessel, S.A.; Singh, R.P.; Huerta-Espino, J.; Spielmeyer, W.; Brown-Guedira, G.; Selter, L.L.; Keller, B. Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor. Appl. Genet. 2009, 119, 889–898. [Google Scholar] [CrossRef] [PubMed]
- Spielmeyer, W.; Mago, R.; Wellings, C.; Ayliffe, M. Lr67 and Lr34 rust resistance genes have much in common-they confer broad spectrum resistance to multiple pathogens in wheat. BMC Plant Biol. 2013, 13, 96. [Google Scholar] [PubMed]
- Herrera-Foessel, S.A.; Singh, R.P.; Lillemo, M.; Huerta-Espino, J.; Bhavani, S.; Singh, S.; Lan, C.X.; Calvo-Salazar, V.; Lagudah, E.S. Lr67/Yr46 confers adult plant resistance to stem rust and powdery mildew in wheat. Theor. Appl. Genet. 2014, 127, 781–789. [Google Scholar] [CrossRef] [PubMed]
- Moore, J.W.; Herrera-Foessel, S.; Lan, C.X.; Schnippenkoetter, W.; Ayliffe, M.; Huerta-Espino, J.; Lillemo, M.; Viccars, L.; Milne, R.; Periyannan, S.; et al. A recently evolved hexose transporter variant confers resistance to multiple pathogens in wheat. Nat. Genet. 2015, 47, 1494–1498. [Google Scholar] [CrossRef]
- Wang, Y.H.; Tan, J.Y.; Wu, Z.M.; VandenLangenberg, K.; Wehner, T.C.; Wen, C.L.; Zheng, X.Y.; Owens, K.; Thornton, A.; Bang, H.H.; et al. STAYGREEN, STAY HEALTHY: A loss-of-susceptibility mutation in the STAYGREEN gene provides durable, broadspectrum disease resistances for over 50 years of US cucumber production. New Phytol. 2019, 221, 415–430. [Google Scholar] [CrossRef] [PubMed]
- Farrow, S.C.; Facchini, P.J. Functional diversity of 2-oxoglutarate/Fe(II)-dependent dioxygenases in plant metabolism. Front. Plant Sci. 2014, 5, 524. [Google Scholar] [CrossRef] [PubMed]
- Kawai, Y.; Ono, E.; Mizutani, M. Evolution and diversity of the 2-oxoglutarate-dependent dioxygenase superfamily in plants. Plant J. 2014, 78, 328–343. [Google Scholar] [CrossRef]
- Martinez, S.; Hausinger, R.P. Catalytic mechanisms of Fe(II)- and 2-oxoglutarate-dependent oxygenases. J. Biol. Chem. 2015, 290, 20702–20711. [Google Scholar] [CrossRef] [PubMed]
- Markolovic, S.; Wilkins, S.E.; Schofield, C.J. Protein hydroxylation catalyzed by 2-oxoglutarate-dependent oxygenases. J. Biol. Chem. 2015, 290, 20712–20722. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.S.; Leissing, T.M.; Chowdhury, R.; Hopkinson, R.J.; Schofield, C.J. 2-oxoglutarate-dependent oxygenases. Annu. Rev. Biochem. 2018, 87, 585–620. [Google Scholar] [CrossRef] [PubMed]
- Nadi, R.; Mateo-Bonmati, E.; Juan-Vicente, L.; Micol, J.L. The 2OGD superfamily: Emerging functions in plant epigenetics and hormone metabolism. Mol. Plant 2018, 11, 1222–1224. [Google Scholar] [CrossRef]
- Falnes, P.Ø.; Johansen, R.F.; Seeberg, E. AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature 2002, 419, 178–182. [Google Scholar] [CrossRef] [PubMed]
- Trewick, S.C.; Henshaw, T.F.; Hausinger, R.P.; Lindahl, T.; Sedgwick, B. Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature 2002, 419, 174–178. [Google Scholar] [CrossRef]
- Martínez-Pérez, M.; Aparicio, F.; López-Gresa, M.P.; Bellés, J.M.; Sánchez-Navarro, J.A.; Pallás, V. Arabidopsis m6A demethylase activity modulates viral infection of a plant virus and the m6A abundance in its genomic RNAs. Proc. Natl. Acad. Sci. USA 2017, 114, 10755–10760. [Google Scholar] [CrossRef]
- Duan, H.C.; Wei, L.H.; Zhang, C.; Wang, Y.; Chen, L.; Lu, Z.; Chen, P.R.; He, C.; Jia, G. ALKBH10B is an RNA N6-methyladenosine demethylase affecting Arabidopsis floral transition. Plant Cell 2017, 29, 2995–3011. [Google Scholar] [CrossRef]
- Ozer, A.; Bruick, R.K. Non-heme dioxygenases: Cellular sensors and regulators jelly rolled into one? Nat. Chem. Biol. 2007, 3, 144–153. [Google Scholar] [CrossRef]
- Keskiaho, K.; Hieta, R.; Sormunen, R.; Myllyharju, J. Chlamydomonas reinhardtii has multiple prolyl 4-hydroxylases, one of which is essential for proper cell wall assembly. Plant Cell 2007, 19, 256–269. [Google Scholar] [CrossRef] [PubMed]
- Van Damme, M.; Andel, A.; Huibers, R.P.; Panstruga, R.; Weisbeek, P.J.; Van den Ackerveken, G. Identification of Arabidopsis loci required for susceptibility to the downy mildew pathogen Hyaloperonospora parasitica. Mol. Plant Microbe Interact. 2005, 6, 583–592. [Google Scholar] [CrossRef]
- Van Damme, M.; Huibers, R.P.; Elberse, J.; Van den Ackerveken, G. Arabidopsis DMR6 encodes a putative 2OG-Fe(II) oxygenase that is defense-associated but required for susceptibility to downy mildew. Plant J. 2008, 54, 785–793. [Google Scholar] [CrossRef]
- Zeilmaker, T.; Ludwig, N.R.; Elberse, J.; Seidl, M.F.; Berke, L.; Van Doorn, A.; Schuurink, R.C.; Snel, B.; Van den Ackerveken, G. DOWNY MILDEW RESISTANT 6 and DMR6-LIKE OXYGENASE 1 are partially redundant but distinct suppressors of immunity in Arabidopsis. Plant J. 2015, 81, 210–222. [Google Scholar] [CrossRef]
- Sun, K.L.; van Tuinen, A.; van Kan, J.A.L.; Wolters, A.A.; Jacobsen, E.; Visser, R.G.F.; Bai, Y.L. Silencing of DND1 in potato and tomato impedes conidial germination, attachment and hyphal growth of Botrytis cinerea. BMC Plant Biol. 2017, 17, 235. [Google Scholar] [CrossRef]
- Kieu, N.P.; Lenman, M.; Wang, E.S.; Petersen, B.L.; Andreasson, E. Mutations introduced in susceptibility genes through CRISPR/Cas9 genome editing confer increased late blight resistance in potatoes. Sci. Rep. 2021, 11, 4487. [Google Scholar] [CrossRef]
- Thomazella, D.P.T.; Seong, K.; Mackelprang, R.; Dahlbeck, D.; Geng, Y.; Gill, U.S.; Qi, T.C.; Pham, J.; Giuseppe, P.; Lee, C.Y.; et al. Loss of function of a DMR6 ortholog in tomato confers broad-spectrum disease resistance. Proc. Natl. Acad. Sci. USA 2021, 118, e2026152118. [Google Scholar] [CrossRef] [PubMed]
- Hasley, J.A.R.; Navet, N.; Tian, M.Y. CRISPR/Cas9-mediated mutagenesis of sweet basil candidate susceptibility gene ObDMR6 enhances downy mildew resistance. PLoS ONE 2021, 16, e0253245. [Google Scholar] [CrossRef]
- Liang, B.B.; Bai, Y.J.; Zang, C.Q.; Pei, X.; Xie, J.H.; Lin, Y.; Liu, X.Z.; Ahsan, T.; Liang, C.H. Overexpression of the first peanut-susceptible gene, AhS5H1 or AhS5H2, enhanced susceptibility to Pst DC3000 in Arabidopsis. Int. J. Mol. Sci. 2023, 24, 14210. [Google Scholar] [CrossRef]
- Low, Y.C.; Lawton, M.A.; Di, R. Validation of barley 2OGO gene as a functional orthologue of Arabidopsis DMR6 gene in Fusarium head blight susceptibility. Sci. Rep. 2020, 10, 9935. [Google Scholar] [CrossRef]
- Tripathi, J.N.; Ntui, V.O.; Shah, T.; Tripathi, L. CRISPR/Cas9-mediated editing of DMR6 orthologue in banana (Musa spp.) confers enhanced resistance to bacterial disease. Plant Biotechnol. J. 2021, 19, 1291–1293. [Google Scholar] [CrossRef] [PubMed]
- Liang, B.B.; Wang, H.; Yang, C.; Wang, L.Y.; Qi, L.L.; Guo, Z.J.; Chen, X.J. Salicylic acid is required for broad-spectrum disease resistance in rice. Int. J. Mol. Sci. 2022, 23, 1354. [Google Scholar] [CrossRef]
- Zhang, Y.J.; Yu, Q.L.; Gao, S.L.; Yu, N.N.; Zhao, L.; Wang, J.B.; Zhao, J.Z.; Huang, P.; Yao, L.B.; Wang, M.; et al. Disruption of the primary salicylic acid hydroxylases in rice enhances broad-spectrum resistance against pathogens. Plant Cell Environ. 2022, 45, 2211–2225. [Google Scholar] [CrossRef]
- Liu, X.; Yu, Y.; Yao, W.; Yin, Z.L.; Wang, Y.B.; Huang, Z.J.; Zhou, J.Q.; Liu, J.L.; Lu, X.D.; Wang, F.; et al. CRISPR/Cas9-mediated simultaneous mutation of three salicylic acid 5-hydroxylase (OsS5H) genes confers broad-spectrum disease resistance in rice. Plant Biotechnol. J. 2023, 21, 1873–1886. [Google Scholar] [CrossRef]
- Shan, X.C.; Goodwin, P.H. Silencing an ACC oxidase gene affects the susceptible host response of Nicotiana benthamiana to infection by Colletotrichum orbiculare. Plant Cell Rep. 2006, 25, 241–247. [Google Scholar] [CrossRef]
- Iwai, T.; Miyasaka, A.; Seo, S.; Ohashi, Y. Contribution of ethylene biosynthesis for resistance to blast fungus infection in young rice plants. Plant Physiol. 2006, 142, 1202–1215. [Google Scholar] [CrossRef]
- Dziurka, M.; Janeczko, A.; Juhász, C.; Gullner, G.; Oklestková, J.; Novák, O.; Saja, D.; Skoczowski, A.; Tóbiás, I.; Barna, B. Local and systemic hormonal responses in pepper leaves during compatible and incompatible pepper-tobamovirus interactions. Plant Physiol. Biochem. 2016, 109, 355–364. [Google Scholar] [CrossRef]
- Zheng, H.Y.; Dong, L.L.; Han, X.Y.; Jin, H.B.; Yin, C.C.; Han, Y.L.; Li, B.; Qin, H.J.; Zhang, J.S.; Shen, Q.H.; et al. The TuMYB46L-TuACO3 module regulates ethylene biosynthesis in einkorn wheat defense to powdery mildew. New Phytol. 2020, 225, 2526–2541. [Google Scholar] [CrossRef]
- Caarls, L.; Elberse, J.; Awwanah, M.; Ludwig, N.R.; De Vries, M.; Zeilmaker, T.; Van Wees, S.C.; Schuurink, R.C.; Ackerveken, G.V.D. Arabidopsis JASMONATE-INDUCED OXYGENASES down-regulate plant immunity by hydroxylation and inactivation of the hormone jasmonic acid. Proc. Natl. Acad. Sci. USA 2017, 114, 6388–6393. [Google Scholar] [CrossRef]
- Smirnova, E.; Marquis, V.; Poirier, L.; Aubert, Y.; Zumsteg, J.; Ménard, R.; Miesch, L.; Heitz, T. Jasmonic Acid Oxidase 2 hydroxylates jasmonic acid and represses basal defense and resistance responses against Botrytis cinerea infection. Mol. Plant 2017, 10, 1159–1173. [Google Scholar] [CrossRef]
- Marquis, V.; Smirnova, E.; Graindorge, S.; Delcros, P.; Villette, C.; Zumsteg, J.; Heintz, D.; Heitz, T. Broad-spectrum stress tolerance conferred by suppressing jasmonate signaling attenuation in Arabidopsis JASMONIC ACID OXIDASE mutants. Plant J. 2022, 109, 856–872. [Google Scholar] [CrossRef]
- Zhang, Y.L.; Li, X. Salicylic acid: Biosynthesis, perception, and contributions to plant immunity. Curr. Opin. Plant Biol. 2019, 50, 29–36. [Google Scholar] [CrossRef]
- Ding, P.T.; Ding, Y.L. Stories of salicylic acid: A plant defense hormone. Trends Plant Sci. 2020, 25, 549–565. [Google Scholar] [CrossRef]
- Malamy, J.; Carr, J.P.; Klessig, D.F.; Raskin, I. Salicylic acid: A likely endogenous signal in the resistance response of tobacco to viral infection. Science 1990, 250, 1002–1004. [Google Scholar] [CrossRef]
- Métraux, J.P.; Signer, H.; Ryals, J.; Ward, E.; Wyss-Benz, M.; Gaudin, J.; Raschdorf, K.; Schmid, E.; Blum, W.; Inverardi, B. Increase in salicylic acid at the onset of systemic acquired resistance in cucumber. Science 1990, 250, 1004–1006. [Google Scholar] [CrossRef]
- Kim, D.S.; Hwang, B.K. An important role of the pepper phenylalanine ammonia-lyase gene (PAL1) in salicylic acid-dependent signalling of the defence response to microbial pathogens. J. Exp. Bot. 2014, 65, 2295–2306. [Google Scholar] [CrossRef]
- Hao, Q.Q.; Wang, W.Q.; Han, X.L.; Wu, J.Z.; Lyu, B.; Chen, F.J.; Caplan, A.; Li, C.X.; Wu, J.J.; Wang, W.; et al. Isochorismate-based salicylic acid biosynthesis confers basal resistance to Fusarium graminearum in barley. Mol. Plant Pathol. 2018, 19, 1995–2010. [Google Scholar] [CrossRef]
- Li, Y.X.; Zhang, W.; Dong, H.X.; Liu, Z.Y.; Ma, J.; Zhang, X.Y. Salicylic acid in Populus tomentosa is a remote signalling molecule induced by Botryosphaeria dothidea infection. Sci. Rep. 2018, 8, 14059. [Google Scholar] [CrossRef]
- Ullah, C.; Tsai, C.J.; Unsicker, S.B.; Xue, L.; Reichelt, M.; Gershenzon, J.; Hammerbacher, A. Salicylic acid activates poplar defense against the biotrophic rust fungus Melampsora larici-populina via increased biosynthesis of catechin and proanthocyanidins. New Phytol. 2019, 221, 960–975. [Google Scholar] [CrossRef]
- Ullah, C.; Unsicker, S.B.; Reichelt, M.; Gershenzon, J.; Hammerbacher, A. Accumulation of catechin and proanthocyanidins in black poplar stems after infection by plectosphaerella populi: Hormonal regulation, biosynthesis and antifungal activity. Front. Plant Sci. 2019, 10, 1441. [Google Scholar] [CrossRef]
- Wang, H.; Gong, W.F.; Wang, Y.; Ma, Q. Contribution of a WRKY transcription factor, ShWRKY81, to powdery mildew resistance in wild tomato. Int. J. Mol. Sci. 2023, 24, 2583. [Google Scholar] [CrossRef]
- Nawrath, C.; Métraux, J.P. Salicylic acid induction-deficient mutants of Arabidopsis express PR-2 and PR-5 and accumulate high levels of camalexin after pathogen inoculation. Plant Cell 1999, 11, 1393–1404. [Google Scholar]
- Wildermuth, M.C.; Dewdney, J.; Wu, G.; Ausubel, F.M. Isochorismate synthase is required to synthesize salicylic acid for plant defence. Nature 2001, 414, 562–565. [Google Scholar] [CrossRef]
- Gaffney, T.; Friedrich, L.; Vernooij, B.; Negrotto, D.; Nye, G.; Uknes, S.; Ward, E.; Kessmann, H.; Ryals, J. Requirement of salicylic acid for the induction of systemic acquired resistance. Science 1993, 261, 754–756. [Google Scholar] [CrossRef]
- Delaney, T.P.; Uknes, S.; Vernooij, B.; Friedrich, L.; Weymann, K.; Negrotto, D.; Gaffney, T.; Gut-Rella, M.; Kessmann, H.; Ward, E.; et al. A central role of salicylic acid in plant disease resistance. Science 1994, 266, 1247–1250. [Google Scholar] [CrossRef]
- Conrath, U.; Chen, Z.; Ricigliano, J.R.; Klessig, D.F. Two inducers of plant defense responses, 2,6-dichloroisonicotinec acid and salicylic acid, inhibit catalase activity in tobacco. Proc. Natl. Acad. Sci. USA 1995, 92, 7143–7147. [Google Scholar] [CrossRef]
- Knoth, C.; Salus, M.S.; Girke, T.; Eulgem, T. The synthetic elicitor 3,5-dichloroanthranilic acid induces NPR1-dependent and NPR1-independent mechanisms of disease resistance in Arabidopsis. Plant Physiol. 2009, 150, 333–347. [Google Scholar] [CrossRef]
- Cui, Z.N.; Ito, J.; Dohi, H.; Amemiya, Y.; Nishida, Y. Molecular design and synthesis of novel salicyl glycoconjugates as elicitors against plant diseases. PLoS ONE 2014, 9, e108338. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, J.H. Exogenous treatment with salicylic acid attenuates occurrence of citrus canker in susceptible navel orange (Citrus sinensis Osbeck). J. Plant Physiol. 2012, 169, 1143–1149. [Google Scholar] [CrossRef]
- Lefevere, H.; Bauters, L.; Gheysen, G. Salicylic acid biosynthesis in plants. Front. Plant Sci. 2020, 11, 338. [Google Scholar] [CrossRef]
- Cao, H.; Bowling, S.A.; Gordon, A.S.; Dong, X.N. Characterization of an Arabidopsis mutant that is nonresponsive to inducers of systemic acquired resistance. Plant Cell 1994, 6, 1583–1592. [Google Scholar] [CrossRef]
- Castelló, M.J.; Medina-Puche, L.; Lamilla, J.; Tornero, P. NPR1 paralogs of Arabidopsis and their role in salicylic acid perception. PLoS ONE 2018, 13, e0209835. [Google Scholar] [CrossRef]
- Ding, Y.L.; Sun, T.J.; Ao, K.; Peng, Y.J.; Zhang, Y.X.; Li, X.; Zhang, Y.L. Opposite roles of salicylic acid receptors NPR1 and NPR3/NPR4 in transcriptional regulation of plant immunity. Cell 2018, 173, 1454–1467. [Google Scholar] [CrossRef]
- van Loon, L.C.; Rep, M.; Pieterse, C.M. Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef]
- De Geyter, N.; Gholami, A.; Goormachtig, S.; Goossens, A. Transcriptional machineries in jasmonate elicited plant secondary metabolism. Trends Plant Sci. 2012, 17, 349–359. [Google Scholar] [CrossRef]
- Wasternack, C.; Hause, B. Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 2013, 111, 1021–1058. [Google Scholar] [CrossRef]
- Li, M.Y.; Yu, G.H.; Cao, C.L.; Liu, P. Metabolism, signaling, and transport of jasmonates. Plant Commun. 2021, 2, 100231. [Google Scholar] [CrossRef]
- Howe, G.A.; Major, I.T.; Koo, A.J. Modularity in jasmonate signaling for multistress resilience. Annu. Rev. Plant Biol. 2018, 69, 387–415. [Google Scholar] [CrossRef]
- Vijayan, P.; Shockey, J.; Lévesque, C.A.; Cook, R.J.; Browse, J. A role for jasmonate in pathogen defense of Arabidopsis. Proc. Natl. Acad. Sci. USA 1998, 95, 7209–7214. [Google Scholar] [CrossRef]
- Thaler, J.S.; Owen, B.; Higgins, V.J. The role of the jasmonate response in plant susceptibility to diverse pathogens with a range of lifestyles. Plant Physiol. 2004, 135, 530–538. [Google Scholar] [CrossRef]
- Yan, Y.X.; Christensen, S.; Isakeit, T.; Engelberth, J.; Meeley, R.; Hayward, A.; Emery, R.J.; Kolomiets, M.V. Disruption of OPR7 and OPR8 reveals the versatile functions of jasmonic acid in maize development and defense. Plant Cell 2012, 24, 1420–1436. [Google Scholar] [CrossRef] [PubMed]
- Thomma, B.P.; Eggermont, K.; Penninckx, I.A.; Mauch-Mani, B.; Vogelsang, R.; Cammue, B.P.; Broekaert, W.F. Separate jasmonate-dependent and salicylate-dependent defense-response pathways in Arabidopsis are essential for resistance to distinct microbial pathogens. Proc. Natl. Acad. Sci. USA 1998, 95, 15107–15111. [Google Scholar] [CrossRef] [PubMed]
- Kloek, A.P.; Verbsky, M.L.; Sharma, S.B.; Schoelz, J.E.; Vogel, J.; Klessig, D.F.; Kunkel, B.N. Resistance to Pseudomonas syringae conferred by an Arabidopsis thaliana coronatine-insensitive (coi1) mutation occurs through two distinct mechanisms. Plant J. 2001, 26, 509–522. [Google Scholar] [CrossRef]
- Major, I.T.; Yoshida, Y.; Campos, M.L.; Kapali, G.; Xin, X.F.; Sugimoto, K.; Ferreira, D.d.O.; He, S.Y.; Howe, G.A. Regulation of growth-defense balance by the JASMONATE ZIM-DOMAIN (JAZ)-MYC transcriptional module. New Phytol. 2017, 215, 1533–1547. [Google Scholar] [CrossRef]
- Qiu, J.H.; Xie, J.H.; Chen, Y.; Shen, Z.N.; Shi, H.B.; Naqvi, N.I.; Qian, Q.; Liang, Y.; Kou, Y.J. Warm temperature compromises JA-regulated basal resistance to enhance Magnaporthe oryzae infection in rice. Mol. Plant 2022, 15, 723–739. [Google Scholar] [CrossRef]
- Yang, J.Y.; Iwasaki, M.; Machida, C.; Machida, Y.; Zhou, X.P.; Chua, N.H. βC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid responses. Genes Dev. 2008, 22, 2564–2577. [Google Scholar] [CrossRef]
- Lozano-Durán, R.; Rosas-Díaz, T.; Gusmaroli, G.; Luna, A.P.; Taconnat, L.; Deng, X.W.; Bejarano, E.R. Geminiviruses subvert ubiquitination by altering CSN-mediated derubylation of SCF E3 ligase complexes and inhibit jasmonate signaling in Arabidopsis thaliana. Plant Cell 2011, 23, 1014–1032. [Google Scholar] [CrossRef]
- Wu, D.W.; Qi, T.C.; Li, W.X.; Tian, H.X.; Gao, H.; Wang, J.J.; Ge, J.; Yao, R.F.; Ren, C.M.; Wang, X.B.; et al. Viral effector protein manipulates host hormone signaling to attract insect vectors. Cell Res. 2017, 27, 402–415. [Google Scholar] [CrossRef]
- Zhang, C.; Ding, Z.M.; Wu, K.C.; Yang, L.; Li, Y.; Yang, Z.; Shi, S.; Liu, X.J.; Zhao, S.S.; Yang, Z.R.; et al. Suppression of jasmonic acid-mediated defense by viral-inducible microRNA319 facilitates virus infection in rice. Mol. Plant 2016, 9, 1302–1314. [Google Scholar] [CrossRef] [PubMed]
- He, Y.Q.; Zhang, H.H.; Sun, Z.T.; Li, J.M.; Hong, G.J.; Zhu, Q.S.; Zhou, X.B.; MacFarlane, S.; Yan, F.; Chen, J.P. Jasmonic acid-mediated defense suppresses brassinosteroid-mediated susceptibility to Rice black streaked dwarf virus infection in rice. New Phytol. 2017, 214, 388–399. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.R.; Huang, Y.; Yang, J.L.; Yao, S.Z.; Zhao, K.; Wang, D.H.; Qin, Q.Q.; Bian, Z.; Li, Y.; Lan, Y.; et al. Jasmonate signaling enhances RNA silencing and antiviral defense in rice. Cell Host Microbe 2020, 28, 89–103. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.F.; Hoffman, N.E. Ethylene biosynthesis and its regulation in higher plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1984, 35, 155–189. [Google Scholar] [CrossRef]
- Kende, H. Ethylene biosynthesis. Annu. Rev. Plant Physiol. Plant Mol. Biol. 1993, 44, 283–307. [Google Scholar] [CrossRef]
- Zarembinski, T.I.; Theologis, A. Ethylene biosynthesis and action: A case of conservation. Plant Mol. Biol. 1994, 26, 1579–1597. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.P.; Lee, F.N.; Counce, P.A.; Gibbons, J.H. Mediation of partial resistance to rice blast through anaerobic induction of ethylene. Phytopathology 2004, 94, 819–825. [Google Scholar] [CrossRef]
- Yang, C.; Li, W.; Cao, J.D.; Meng, F.W.; Yu, Y.Q.; Huang, J.K.; Jiang, L.; Liu, M.X.; Zhang, Z.G.; Chen, X.W.; et al. Activation of ethylene signaling pathways enhances disease resistance by regulating ROS and phytoalexin production in rice. Plant J. 2017, 89, 338–353. [Google Scholar] [CrossRef]
- Zhai, K.R.; Liang, D.; Li, H.L.; Jiao, F.Y.; Yan, B.X.; Liu, J.; Lei, Z.Y.; Huang, L.; Gong, X.Y.; Wang, X. NLRs guard metabolism to coordinate pattern- and effector-triggered immunity. Nature 2022, 601, 245–251. [Google Scholar] [CrossRef]
- Helliwell, E.E.; Wang, Q.; Yang, Y.N. Transgenic rice with inducible ethylene production exhibits broad-spectrum disease resistance to the fungal pathogens Magnaporthe oryzae and Rhizoctonia solani. Plant Biotechnol. J. 2013, 11, 33–42. [Google Scholar] [CrossRef]
- Hoffman, T.; Schmidt, J.S.; Zheng, X.Y.; Bent, A.F. Isolation of ethylene-insensitive soybean mutants that are altered in pathogen susceptibility and gene-for-gene disease resistance. Plant Physiol. 1999, 119, 935–950. [Google Scholar] [CrossRef] [PubMed]
- Shibata, Y.; Kawakita, K.; Takemoto, D. Age-related resistance of Nicotiana benthamiana against hemibiotrophic pathogen Phytophthora infestans requires both ethylene- and salicylic acid-mediated signaling pathways. Mol. Plant Microbe Interact. 2010, 23, 1130–1142. [Google Scholar] [CrossRef] [PubMed]
- Tintor, N.; Ross, A.; Kanehara, K.; Yamada, K.; Fan, L.; Kemmerling, B.; Nürnberger, T.; Tsuda, K.; Saijo, Y. Layered pattern receptor signaling via ethylene and endogenous elicitor peptides during Arabidopsis immunity to bacterial infection. Proc. Natl. Acad. Sci. USA 2013, 110, 6211–6216. [Google Scholar] [CrossRef]
- He, P.; Warren, R.F.; Zhao, T.H.; Shan, L.B.; Zhu, L.H.; Tang, X.Y.; Zhou, J.M. Overexpression of Pti5 in tomato potentiates pathogen-induced defense gene expression and enhances disease resistance to Pseudomonas syringae pv. tomato. Mol. Plant Microbe Interact. 2001, 14, 1453–1457. [Google Scholar] [CrossRef]
- Gu, Y.Q.; Wildermuth, M.C.; Chakravarthy, S.; Loh, Y.T.; Yang, C.M.; He, X.H.; Han, Y.; Martin, G.B. Tomato transcription factors Pti4, Pti5, and Pti6 activate defense responses when expressed in Arabidopsis. Plant Cell 2002, 14, 817–831. [Google Scholar] [CrossRef] [PubMed]
- Berrocal-Lobo, M.; Molina, A.; Solano, R. Constitutive expression of ETHYLENERESPONSE-FACTOR1 in Arabidopsis confers resistance to several necrotrophic fungi. Plant J. 2002, 29, 23–32. [Google Scholar] [CrossRef]
- Berrocal-Lobo, M.; Molina, A. Ethylene response factor 1 mediates Arabidopsis resistance to the soilborne fungus Fusarium oxysporum. Mol. Plant Microbe Interact. 2004, 17, 763–770. [Google Scholar] [CrossRef]
- Fischer, U.; Dröge-Laser, W. Overexpression of NtERF5, a new member of the tobacco ethylene response transcription factor family enhances resistance to tobacco mosaic virus. Mol. Plant Microbe Interact. 2004, 17, 1162–1171. [Google Scholar] [CrossRef]
- McGrath, K.C.; Dombrecht, B.; Manners, J.M.; Schenk, P.M.; Edgar, C.I.; Maclean, D.J.; Scheible, W.R.; Udvardi, M.K.; Kazan, K. Repressor- and activator-type ethylene response factors functioning in jasmonate signaling and disease resistance identified via a genome-wide screen of Arabidopsis transcription factor gene expression. Plant Physiol. 2005, 139, 949–959. [Google Scholar] [CrossRef]
- Zhu, X.L.; Qi, L.; Liu, X.; Cai, S.B.; Xu, H.J.; Huang, R.F.; Li, J.R.; Wei, X.N.; Zhang, Z.Y. The wheat ethylene response factor transcription factor pathogen-induced ERF1 mediates host responses to both the necrotrophic pathogen Rhizoctonia cerealis and freezing stresses. Plant Physiol. 2014, 164, 1499–1514. [Google Scholar] [CrossRef]
- Dong, L.; Cheng, Y.; Wu, J.; Cheng, Q.; Li, W.; Fan, S.; Jiang, L.; Xu, Z.; Kong, F.; Zhang, D.; et al. Overexpression of GmERF5, a new member of the soybean EAR motif-containing ERF transcription factor, enhances resistance to Phytophthora sojae in soybean. J. Exp. Bot. 2015, 66, 2635–2647. [Google Scholar] [CrossRef] [PubMed]
- Xing, L.P.; Di, Z.C.; Yang, W.W.; Liu, J.Q.; Li, M.N.; Wang, X.J.; Cui, C.F.; Wang, X.Y.; Wang, X.E.; Zhang, R.Q.; et al. Overexpression of ERF1-V from Haynaldia villosa can enhance the resistance of wheat to powdery mildew and increase the tolerance to salt and drought stresses. Front. Plant Sci. 2017, 8, 1948. [Google Scholar] [CrossRef] [PubMed]
- Lund, S.T.; Stall, R.E.; Klee, H.J. Ethylene regulates the susceptible response to pathogen infection in tomato. Plant Cell 1998, 10, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.M.; Xue, L.; Chintamanani, S.; Germain, H.; Lin, H.Q.; Cui, H.T.; Cai, R.; Zuo, J.R.; Tang, X.Y.; Li, X.; et al. ETHYLENE INSENSITIVE3 and ETHYLENE INSENSITIVE3-LIKE1 repress SALICYLIC ACID INDUCTION DEFICIENT2 expression to negatively regulate plant innate immunity in Arabidopsis. Plant Cell 2009, 21, 2527–2540. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Steed, A.; Travella, S.; Keller, B.; Nicholson, P. Fusarium graminearum exploits ethylene signalling to colonize dicotyledonous and monocotyledonous plants. New Phytol. 2009, 182, 975–983. [Google Scholar] [CrossRef] [PubMed]
- Shen, X.L.; Liu, H.B.; Yuan, B.; Li, X.H.; Xu, C.G.; Wang, S.P. OsEDR1 negatively regulates rice bacterial resistance via activation of ethylene biosynthesis. Plant Cell Environ. 2011, 34, 179–191. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, T.R.; Jia, J.Z.; Sun, J.Q. The wheat mediator subunit TaMED25 interacts with the transcription factor TaEIL1 to negatively regulate disease resistance against powdery mildew. Plant Physiol. 2016, 170, 1799–1816. [Google Scholar] [CrossRef] [PubMed]
- Pieterse, C.M.; Leon-Reyes, A.; Van der Ent, S.; Van Wees, S.C. Networking by small-molecule hormones in plant immunity. Nat. Chem. Biol. 2009, 5, 308–316. [Google Scholar] [CrossRef] [PubMed]
- Berens, M.L.; Berry, H.M.; Mine, A.; Argueso, C.T.; Tsuda, K. Evolution of hormone signaling networks in plant defense. Annu. Rev. Phytopathol. 2017, 55, 401–425. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.W.; Halitschke, R.; Yin, C.X.; Liu, C.J.; Gan, S.S. Salicylic acid 3-hydroxylase regulates Arabidopsis leaf longevity by mediating salicylic acid catabolism. Proc. Natl. Acad. Sci. USA 2013, 110, 14807–14812. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhao, L.; Zhao, J.; Li, Y.; Wang, J.; Guo, R.; Gan, S.; Liu, C.-J.; Zhang, K. S5H/DMR6 encodes a salicylic acid 5-hydroxylase that fine-tunes salicylic acid homeostasis. Plant Physiol. 2017, 175, 1082–1093. [Google Scholar] [CrossRef] [PubMed]
- Kim, C.Y.; Liu, Y.D.; Thorne, E.T.; Yang, H.P.; Fukushige, H.; Gassmann, W.; Hildebrand, D.; Sharp, R.E.; Zhang, S.Q. Activation of a stress-responsive mitogen-activated protein kinase cascade induces the biosynthesis of ethylene in plants. Plant Cell 2003, 15, 2707–2718. [Google Scholar] [CrossRef] [PubMed]
- Yim, W.J.; Kim, K.Y.; Lee, Y.W.; Sundaram, S.P.; Lee, Y.; Sa, T.M. Real time expression of ACC oxidase and PR-protein genes mediated by Methylobacterium spp. in tomato plants challenged with Xanthomonas campestris pv. vesicatoria. J. Plant Physiol. 2014, 171, 1064–1075. [Google Scholar] [CrossRef] [PubMed]
- van Schie, C.C.; Takken, F.L. Susceptibility genes 101: How to be a good host. Annu. Rev. Phytopathol. 2014, 52, 551–581. [Google Scholar] [CrossRef] [PubMed]
- Zaidi, S.S.; Mukhtar, M.S.; Mansoor, S. Genome editing: Targeting susceptibility genes for plant disease resistance. Trends Biotechnol. 2018, 36, 898–906. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.X. Genome engineering for crop improvement and future agriculture. Cell 2021, 184, 1621–1635. [Google Scholar] [CrossRef] [PubMed]
- Jørgensen, J.H. Discovery, characterization and exploitation of Mlo powdery mildew resistance in barley. Euphytica 1992, 63, 141–152. [Google Scholar] [CrossRef]
- Büschges, R.; Hollricher, K.; Panstruga, R.; Simons, G.; Wolter, M.; Frijters, A.; van Daelen, R.; van der Lee, T.; Diergaarde, P.; Groenendijk, J. The barley Mlo gene: A novel control element of plant pathogen resistance. Cell 1997, 88, 695–705. [Google Scholar] [CrossRef]
- Acevedo-Garcia, J.; Kusch, S.; Panstruga, R. Magical mystery tour: MLO proteins in plant immunity and beyond. New Phytol. 2014, 204, 273–281. [Google Scholar] [CrossRef]
- Kim, M.C.; Panstruga, R.; Elliott, C.; Müller, J.; Devoto, A.; Yoon, H.W.; Park, H.C.; Cho, M.J.; Schulze-Lefert, P. Calmodulin interacts with MLO protein to regulate defence against mildew in barley. Nature 2002, 416, 447–451. [Google Scholar] [CrossRef]
- Aist, J.R.; Gold, R.E.; Bayles, C.J. Evidence for the involvement of molecular components of papillae in ml-o resistance to barley powdery mildew. Phytopathology 1987, 77, 17–32. [Google Scholar]
- Nekrasov, V.; Wang, C.M.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci. Rep. 2017, 7, 482. [Google Scholar] [CrossRef]
- Wan, D.Y.; Guo, Y.; Cheng, Y.; Hu, Y.; Xiao, S.Y.; Wang, Y.J.; Wen, Y.Q. CRISPR/Cas9-mediated mutagenesis of VvMLO3 results in enhanced resistance to powdery mildew in grapevine (Vitis vinifera). Hortic. Res. 2020, 7, 116. [Google Scholar] [CrossRef]
- Pramanik, D.; Shelake, R.M.; Park, J.; Kim, M.J.; Hwang, I.; Park, Y.; Kim, J.Y. CRISPR/Cas9-mediated generation of pathogen-resistant tomato against tomato yellow leaf curl virus and powdery mildew. Int. J. Mol. Sci. 2021, 22, 1878. [Google Scholar] [CrossRef] [PubMed]
- Li, S.N.; Lin, D.X.; Zhang, Y.W.; Deng, M.; Chen, Y.X.; Lv, B.; Li, B.S.; Lei, Y.; Wang, Y.P.; Zhao, L.; et al. Genome-edited powdery mildew resistance in wheat without growth penalties. Nature 2022, 602, 455–460. [Google Scholar] [CrossRef]
- Bui, T.P.; Le, H.; Ta, D.T.; Nguyen, C.X.; Le, N.T.; Tran, T.T.; Van Nguyen, P.; Stacey, G.; Stacey, M.G.; Pham, N.B.; et al. Enhancing powdery mildew resistance in soybean by targeted mutation of MLO genes using the CRISPR/Cas9 system. BMC Plant Biol. 2023, 23, 533. [Google Scholar] [CrossRef]
- Lin, W.C.; Lu, C.F.; Wu, J.W.; Cheng, M.L.; Lin, Y.M.; Yang, N.S.; Black, L.; Green, S.K.; Wang, J.F.; Cheng, C.P. Transgenic tomato plants expressing the Arabidopsis NPR1 gene display enhanced resistance to a spectrum of fungal and bacterial diseases. Transgenic Res. 2004, 13, 567–581. [Google Scholar] [CrossRef]
- Wally, O.; Jayaraj, J.; Punja, Z.K. Broad-spectrum disease resistance to necrotrophic and biotrophic pathogens in transgenic carrots (Daucus carota L.) expressing an Arabidopsis NPR1 gene. Planta 2009, 231, 131–141. [Google Scholar] [CrossRef] [PubMed]
- Parkhi, V.; Kumar, V.; Campbell, L.M.; Bell, A.A.; Shah, J.; Rathore, K.S. Resistance against various fungal pathogens and reniform nematode in transgenic cotton plants expressing Arabidopsis NPR1. Transgenic Res. 2010, 19, 959–975. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Joshi, S.G.; Bell, A.A.; Rathore, K.S. Enhanced resistance against Thielaviopsis basicola in transgenic cotton plants expressing Arabidopsis NPR1 gene. Transgenic Res. 2013, 22, 359–368. [Google Scholar] [CrossRef]
- Narváez, I.; Pliego Prieto, C.; Palomo-Ríos, E.; Fresta, L.; Jiménez-Díaz, R.M.; Trapero-Casas, J.L.; Lopez-Herrera, C.; Arjona-Lopez, J.M.; Mercado, J.A.; Pliego-Alfaro, F. Heterologous expression of the AtNPR1 gene in olive and its effects on fungal tolerance. Front. Plant Sci. 2020, 11, 308. [Google Scholar] [CrossRef] [PubMed]
- Qiu, W.M.; Soares, J.; Pang, Z.Q.; Huang, Y.X.; Sun, Z.H.; Wang, N.; Grosser, J.; Dutt, M. Potential mechanisms of AtNPR1 mediated resistance against Huanglongbing (HLB) in citrus. Int. J. Mol. Sci. 2020, 21, 2009. [Google Scholar] [CrossRef] [PubMed]
- Silva, K.J.; Brunings, A.; Peres, N.A.; Mou, Z.L.; Folta, K.M. The Arabidopsis NPR1 gene confers broad-spectrum disease resistance in strawberry. Transgenic Res. 2015, 24, 693–704. [Google Scholar] [CrossRef] [PubMed]
- Fitzgerald, H.A.; Chern, M.S.; Navarre, R.; Ronald, P.C. Overexpression of (At)NPR1 in rice leads to a BTH- and environment-induced lesion-mimic/cell death phenotype. Mol. Plant Microbe Interact. 2004, 17, 140–151. [Google Scholar] [CrossRef] [PubMed]
- Quilis, J.; Peñas, G.; Messeguer, J.; Brugidou, C.; San Segundo, B. The Arabidopsis AtNPR1 inversely modulates defense responses against fungal, bacterial, or viral pathogens while conferring hypersensitivity to abiotic stresses in transgenic rice. Mol. Plant Microbe Interact. 2008, 21, 1215–1231. [Google Scholar] [CrossRef]
- Xu, G.Y.; Yuan, M.; Ai, C.R.; Liu, L.J.; Zhuang, E.; Karapetyan, S.; Wang, S.P.; Dong, X.N. uORF-mediated translation allows engineered plant disease resistance without fitness costs. Nature 2017, 545, 491–494. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).