Abstract
Bats are great models for studying repetitive DNAs due to their compact genomes and extensive chromosomal rearrangements. Here, we investigated the repetitive DNA content of two phyllostomid bat species, Artibeus lituratus (2nn = 30♀/31♂) and Carollia perspicillata (2n = 20♀/21♂), both harboring a multiple XY1Y2 sex chromosome system. Satellite DNA (satDNA) libraries were isolated and characterized, revealing four and ten satDNA families in A. lituratus and C. perspicillata, respectively. These sequences, along with selected microsatellites, were in situ mapped onto chromosomes in both species and phylogenetically related taxa. SatDNAs showed strong accumulation in centromeric and subtelomeric regions, especially pericentromeric areas. Cross-species mapping with C. perspicillata-derived probes indicated terminal localization patterns in other bat species, suggesting conserved distribution. Microsatellites co-localized with 45S rDNA clusters on the neo-sex chromosomes. Additionally, genomic hybridization revealed a male-specific signal on the Y1 chromosome, pointing to potential sex-linked repetitive regions. These findings confirm that bat genomes display relatively low amounts of repetitive DNA compared to other mammals and underscore the role of these elements in genome organization and sex chromosome evolution in phyllostomid bats.
1. Introduction
Repetitive DNA elements can be broadly classified into two major categories based on their distribution: dispersed elements, such as transposable elements (TEs), and tandem repeats, which include multigene families as well as three subclasses defined by the length of their repeat units—satellite, minisatellite, and microsatellite sequences [1,2]. Except for multigene families, tandem repeats are usually non-coding sequences organized in long arrays that evolve at high rates, being considered the fastest mutating repetitive DNA class [3]. Typically, these sequences accumulate in pericentromeric and centromeric heterochromatin but can also be present in euchromatic regions [4,5,6,7], and, in certain instances, linked to chromosomal rearrangement processes [8,9,10]. Furthermore, the proportion of repetitive DNA sequences can vary significantly across the genomes of vertebrates, particularly in mammals, resulting in variations in genome size among different species [11].
Encompassing roughly 1474 living species across 21 families and 236 genera, Chiroptera represents the second most-speciose mammalian order, surpassed only by Rodentia [12]. With estimated genome sizes spanning from 1.6 to 3.54 Gb, bats displays the smallest reported genomes among mammals [11]. Their small genome sizes are frequently linked to physiological adaptations, such as flight [13,14], as the genomes of flying mammals and birds are regarded as more compact than those of their flightless counterparts [15]. One proposed explanation for this reduction is a decreased on their repetitive DNA content [16,17,18,19]. However, research on repetitive DNAs in bat genomes is scarce and has primarily focused on in situ mapping of ribosomal DNAs (rDNA), telomeric repeats, and certain transposable elements (as reviewed in [20]).
Among bat families, Phyllostomidae Gray, 1825, which includes the New World leaf-nosed bats, is the third-largest family, comprising approximately 200 species distributed across 60 genera, all endemic to the Americas [21]. This group is known for its diversity in ecological niches, morphological variation, and extensive chromosomal diversification [22,23]. Therefore, diploid numbers (2n) of this family widely vary, ranging from 14 to 46 chromosomes, with the well-established simple XY system coexisting with multiple sex chromosome systems, including species of the Carolliinae and Stenodermatinae subfamilies [20,24,25].
The genus Artibeus Leach, 1821 (Phyllostomidae: Stenodermatinae) comprises 13 species exhibiting a broad distribution throughout the Neotropics, while Carollia Gray, 1838 (Phyllostomidae: Carolliinae) includes 8 species, being the most prevalent in the Neotropical region [11,26]. In the cytogenetic context, both genera have several intriguing features, making this group an exciting model for evolutionary studies, including karyotype evolution. The great fruit-eating bat Artibeus lituratus exhibits 2n = 30♀/31♂ chromosomes, whereas Seba’s short-tailed bat C. perspicillata has 2n = 20♀/21♂, due to the presence of a multiple XY1Y2 male heterogametic system [24,27,28]. Intriguingly, other sex chromosome systems observed in Sternodermatinae evolved from an XY1Y2 system as observed in Sturnira with a compound system or Neo-XY related to Artibeus [29,30]. In Artibeus, the translocation event involved the X chromosome and a small acrocentric autosome [31], whereas in C. perspicillata, it involved a large autosome containing the 45S rDNA locus [29,32]. In situ hybridization experiments in C. perspicillata also demonstrate that eight chromosome pairs exhibit interstitial telomeric sites (ITS), representing the highest number of ITS yet documented for the family [32].
In this study, we selected two Phyllostomidae species, Artibeus lituratus and Carollia perspicillata, as models to isolate and characterize their satellitomes (i.e., the whole set of satDNA families). Firstly, to test the repetitive DNA content and genome size correlation in bats, we estimate the total content of satDNAs in the genome. Then we performed a comprehensive in situ hybridization analysis to determine the evolutionary trajectory of these satDNA sequences, and also microsatellite repeats, in those two species and also in phylogenetically related species. We select other Sternodermatinae species, including: the Gervais’s fruit-eating bat Dermanura cinerea (2n = 30 with a neo-XY system), the little-yellow-shouldered bat Sturnira lilium (2n = 30 with a neo-XY system), the tent-making bat Uroderma bilobatum (2n = 42 with a neo-XY system), and the brown tent-making bat Uroderma magnirostrum (2n = 36 with a neo-XY system), for this purpose. Comparative genomic hybridization (CGH) was performed to identify potential sex-specific regions in the multiple sex chromosomes of C. perspicillata.
2. Materials and Methods
2.1. Samples and Chromosomal Preparation
Phyllostomid bats were collected in the municipality of Abaetetuba, Pará state, Brazil (1°46′03.2″ S 48°50′15.6″ W) using mist nets. The collection sites are shown in Table 1 and Figure 1. The mitotic cells were obtained from bone marrow preparations after colchicine treatment [33], and also from fibroblast cell cultures established from skin biopsies [34]. The specimens were collected with authorization from the National System for the Management of Genetic Heritage and Associated Traditional Knowledge (SISGEN-A96FF09) and the System of Authorization and Information about Biodiversity (SISBIO) under license number 92665-1. The experiments were conducted in accordance with the Ethics Committee on Animal Experimentation of the Universidade Federal de São Carlos, Brazil (Process number 7994170423). After sample collections, animals were fixed in 10% formalin, preserved in 70% ethanol, and deposited in the Zoology Collection of the Instituto Federal do Pará, Abaetetuba campus.

Table 1.
Individuals of the Phyllostomidae family analyzed in the present work, including their respective diploid numbers (2n), sex chromosome systems (SCS), and number of sampled individuals (N). The asterisk (*) represents the target species for the satellitome analyses.
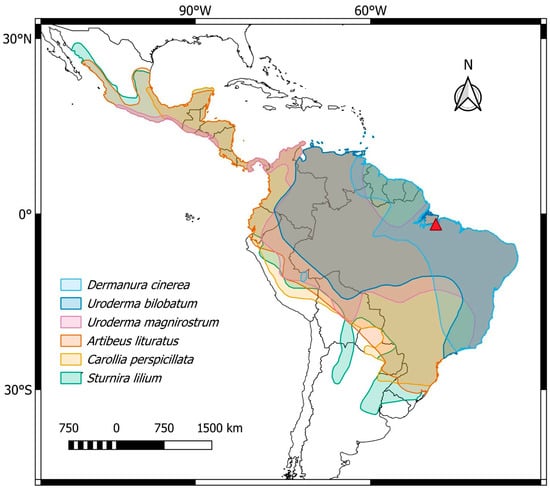
Figure 1.
Central and South America, highlighting the general distribution of the phyllostomid bat species analyzed in the present work: Dermanura cinerea (light blue), Uroderma bilobatum (blue), Uroderma magnirostrum (pink), Artibeus lituratus (orange), Carollia perspicillata (yellow), and Sturnira lilium (green). The collection site is represented by the red triangle. Map created using QGIS 3.40.7 (http://qgis.org accessed on 12 June 2025).
2.2. DNA Extraction and Genome Sequencing
Muscle samples of male individuals from A. lituratus and male and female individuals from C. perspicillata were used to obtain the genomic DNAs (gDNAs). The extraction process followed phenol-chloroform protocol [35], and the samples of one individual male of each species were then sequenced on the BGISEQ-500 platform (BGI Shenzhen Corporation, Shenzhen, China) (paired-end 2 × 150 bp), yielding 3 Gb of data for each species, representing ~1× coverage usually required for satellite assembly [36,37]. Raw reads were deposited on SRA-NCBI and are available under the accession numbers SRR32523217 for A. lituratus and SRR32523216 for C. perspicillata.
2.3. Bioinformatic Analyses and Satellitome Characterization
We isolated the satDNAs from raw reads of A. lituratus and C. perspicillata with the RepeatExplorer2 (RE2) pipeline [38]. To process the raw reads library, we used Trimmomatic (v. 033) [39] to remove low-quality reads and to crop the reads to be 100 base pairs long. After that, we randomly selected 2 × 500,000 reads from the trimmed libraries to prepare the library using the pipeline described by [36] and ran RE2 on the Galaxy server (v. 2.3.12.1) (https://repeatexplorer-elixir.cerit-sc.cz/galaxy). After the first iteration, and in each one thereafter, we filtered the previously inputted library to remove the putative satDNAs found using DeconSeq (v. 0.4.3.) [40] until no more satDNAs were found in the RE2 runs.
The catalogs of A. lituratus and C. perspicillata, (designated AliSatDNAs and CpeSatDNAs, respectively) were examined for additional tandemly repeated elements, including multigene families, utilizing the annotation function in Geneious (v. 7.3) [41], which yielded no further elements. Similarities between inter- and intra-species satellite DNAs were investigated using rm_homology_v2.py [42] and the “de novo assembly” function in Geneious 7.3 and classified according to [36], considering the percentage of similarity: around 95% the same satellite, while between 50 and 80% belonging to the same superfamily (SF).
We employed RepeatMasker (v. 4.1.9) [42] to assess the abundance and divergence of the two satellitomes, utilizing a random selection of 5,000,000 trimmed reads from each library with the “cross-match” tool. The satDNAs were arranged in descending order of abundance and renamed according to the protocol established by [36], utilizing the first letter of the genus, two letters from the specific epithet, and the size of the satDNA. The relative abundance of each satellite DNA was assessed by aligning selected reads with the complete catalog of satDNAs, where the ratio of mapped reads to the total number of nucleotides represented the relative abundance of each satDNA family. Divergence values were concurrently expressed as weighted average Kimura-2-parameter values for each sequence from the script calcDivergenceFromAlign.pl of the RepeatMasker [42]. The satDNA sequences of Artibeus lituratus were deposited in GenBank with accession numbers PV067726 to PV067729, while those for Carollia perspicillata were assigned numbers PV067174 to PV067183.
2.4. Primer Design and Conditions for Satellite Amplification
Primers were designed for five of the ten CpeSatDNAs and three of the four AliSatDNAs and then amplified by Polymerase Chain Reaction (PCR) (Table S1). The DNA template concentrations used were 0.1, 1, 10, and 100 ng. The amplification reaction consisted of the following cycles: 95 °C for 5 min for initial denaturation, followed by 34 cycles, each one consisting of denaturation at 95 °C for 45 s, primer annealing at 52–62 °C for 1 min, and extension at 72 °C for 1 min, with final extension at 72 °C for 5 min. The satDNA amplifications were checked in an electrophoretic agarose gel with 1 or 2% concentration and measured using a ThermoFisher NanoDrop spectrophotometer (ThermoFisher Scientific, Waltham, MA, USA). The remaining satDNAs with repeat unit lengths (RULs) smaller than 31 bp (CpeSat01-30, CpeSat02-24, CpeSat06-24, CpeSat08-21, CpeSat10-13, and AliSat01-19) were directly labeled with Cy3 at the 5′ end during the synthesis by ThermoFisher (ThermoFisher Scientific).
2.5. Probe Preparation and Fluorescence in Situ Hybridization (FISH)
The PCR-amplified satDNAs were labeled with Atto-550-dUTP using a nick translation protocol from Jena Bioscience (Jena, Germany). In addition, the microsatellites (A)n, (C)n, (CA)n, (GA)n, (GC)n, (TA)n, (CAA)n, (GAA)n, (TAA)n, (CGG)n, (CAC)n, (CAG)n, (CAT)n, (GAC)n, (GAG)n, and (TAC)n were also used as probes. These motifs were directly labeled with Cy3 at the 5′ end during synthesis (VBC Biotech, Vienna, Austria). Without the DNA template, telomeric sequences (TTAGGG)n were produced by PCR using the primers (TTAGGG)5 and (CCCTAA)5 according to [43]. These probes were labeled with Atto-550-dUTP using a nick translation protocol from Jena Bioscience (Jena, Germany). Following the guidelines provided by [44], high-stringency conditions were used for the fluorescence in situ hybridization (FISH) procedures. The 4,6-diamidino-2-phenylindole (DAPI) solution was used to counterstain all metaphase plates. Sequential G and C-banding was performed to identify the homologous and the sex chromosomes, according to [45,46], respectively.
2.6. Intraspecific Comparative Genomic Hybridization
Male and female gDNA of C. perspicillata were labeled using a nick translation protocol from Jena Bioscience (Jena, Germany) with Atto-550-dUTP and Atto-488-dUTP, respectively. We employed Cot-1 DNA isolated from the female gDNA of C. perspicillata to block common genomic repetitive regions. The final probe mix consisted of 10 μg of unlabeled female Cot-1 DNA and 500 ng of each male- and female-labeled gDNA, which was ethanol-precipitated, air-dried, and resuspended in 20 μL of hybridization buffer, pH 7.0. The comparative genomic hybridization (CGH) procedures were conducted using the protocol outlined in [47].
2.7. Microscopy and Image Analysis
A minimum of 30 metaphase spreads per individual were examined to validate the 2n and FISH findings. Images were acquired using an Axioplan II fluorescence microscope (Carl Zeiss, Jena GmbH, Jena, Germany) and processed using the ISIS software, version 5.5.9 (MetaSystems, Silver Spring, MD, USA).
3. Results
3.1. Satellitome Composition of A. lituratus and C. perspicillata
Using the RE2 pipeline, we identified 4 and 14 putative satDNAs for A. lituratus and C. perspicillata, respectively. From the 14 satDNAs in C. perspicillata, we revealed four putative satDNAs that exhibited around 95% similarity, which were considered the same satDNA. Considering this, the final catalog of C. perspicillata was composed of 10 CpeSatDNAs, with the repeat unit lengths ranging from 13 to 1681 base pairs and the A+T content varying between 33 and 60%, with an average of 50%. Among these ten, eight satDNA families showed more than 50% A+T content. The homology search by similarities between satellite DNAs found two superfamilies: superfamily 1 (CpeSat03-838 and CpeSat04-535) and superfamily 2 (CpeSat01-30, CpeSat02-24, CpeSat05-72, CpeSat07-1531, and CpeSat08-21) (Table 2). In A. lituratus, the satellite DNA sequences were all distinct, yielding four separate AliSatDNA families, with their repeat unit lengths ranging from 19 to 1388 base pairs, with the A+T content varying between 19.6% and 59.3% (Table 2). The repeat landscapes generated for male genomes of both species are shown in Figure S1. No significant similarity was found among the isolated satDNAs obtained from A. lituratus and C. perspicillata using the Basic local alignment search tool (BLAST)n at NCBI (https://blast.ncbi.nlm.nih.gov/Blast.cgi accessed on 7 July 2025).

Table 2.
General features of Artibeus lituratus (AliSatDNAs) and Carollia perspicillata (CpeSatDNAs) satellitomes, including the classification in superfamilies (SF).
3.2. Chromosomal Distribution of AliSatDNAs and CpeSatDNAs in A. lituratus and Carollia perspicillata
Fluorescence in situ hybridization (FISH) was employed to determine the chromosomal localization of AliSatDNAs and CpeSatDNAs. All four AliSatDNAs showed positive signals in some chromosomes of A. lituratus males (Figure 2a–d). AliSat01-19, AliSat02-51, and AliSat04-312 all exhibited hybridization signals concentrated in the terminal region. The AliSat01-19 was present in ten chromosomes, including four pairs of autosomes (5, 6, 7 and 9), and along with the X and Y2 chromosomes, AliSat02-51 was present on three pairs of autosomes (5, 6 and 7), and AliSat04-312 was dispersed on one chromosome pair (11). Meanwhile, AliSat03-1388 exhibited positive signals on the pericentromeric region of the largest metacentric chromosome pair (1) (Figure 2c).
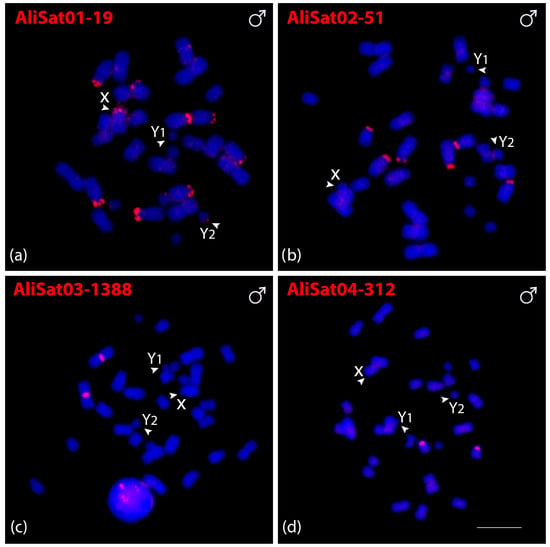
Figure 2.
Male metaphasic chromosomes of A. lituratus showing hybridization signals of AliSatDNAs as probes (a–d). The name of each satellite DNA family is detailed in red in the upper left corner (Atto-550-dUTP labeled). The arrowheads indicate the sex chromosomes X, Y1, and Y2. Scale bar: 10 µm.
Eight out of ten CpeSatDNAs showed positive hybridization signals on C. perspicillata metaphase chromosomes in both males (Figure 3) and females (Figure S2). CpeSat01-30, CpeSat02-24, CpeSat03-838, CpeSat04-535, and CpeSat05-72 showed clustering mainly on the pericentromeric region of all chromosomes, except for some autosome pairs and the sex chromosomes (Figure 3a–e). Additionally, CpeSat04-535 and CpeSat07-1531 were clustered on the centromeric and pericentromeric regions of the X and Y2 chromosomes, respectively (Figure 3d,g), while CpeSat03-838 produces signals on the centromeric region, mainly on the small chromosomes (Figure 3c). Oppositely, CpeSat06-24 shows signals only on chromosome pairs 1, 2, and 3, and CpeSat09-1681 only on the pericentromeric region of pair 1 (Figure 3f,h). All the satellite DNAs of both A. lituratus and C. perspicillata were distributed on heterochromatic regions (Figure S3).
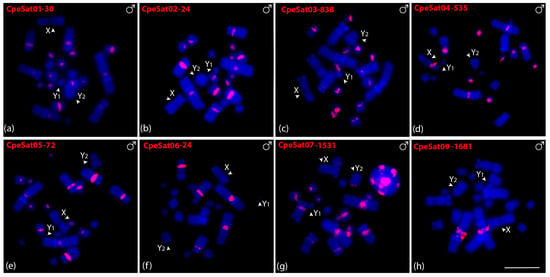
Figure 3.
Male metaphasic chromosomes of C. perspicillata showing hybridization signals of CpeSatDNAs as probes (a–h). The name of each satellite DNA family is detailed in red in the upper left corner (Atto-550-dUTP labeled). The arrowheads indicate the sex chromosomes X, Y1, and Y2. Scale bar: 10 µm.
Cross-species hybridizations with the four AliSatDNAs in C. perspicillata chromosomes did not show any positive signals. Hybridizations using the ten CpeSatDNAs in A. lituratus chromosomes revealed positive signals for only two of them (CpeSat02-24 and CpeSat05-72) distributed mainly on the terminal region of all chromosomes (Figure S4a,b).
3.3. Chromosomal Distribution of Microsatellite and Telomeric Repeats in A. lituratus and C. perspicillata
We also in situ mapped microsatellite and telomeric motifs to determine their chromosomal locations. Only three of the sixteen microsatellite repeat motifs showed positive signals on C. perspicillata chromosomes. (CA)n, (C)n, and (CGG)n motifs showed hybridization signals in female metaphase chromosomes and one signal in males, corresponding to the X chromosome (Figure 4a–c). Intriguingly, the telomeric probe (TTAGGG)5 hybridized onto the pericentromeric region instead of the telomeric regions, except for the sex chromosomes (Figure 4d). We then tested the three microsatellite motifs that showed positive signals on C. perspicillata in A. lituratus. (CA)n and (CGG)n displayed the same hybridization pattern in the terminal region of six chromosomes, while (C)n were clustered in four chromosomes (Figure 4e–g). The telomeric probe (TTAGGG)5 showed hybridization signals on the terminal region of all chromosomes. Furthermore, we identified the interstitial telomeric signal (ITS) on chromosome pair 4 in A. lituratus (Figure 4h).
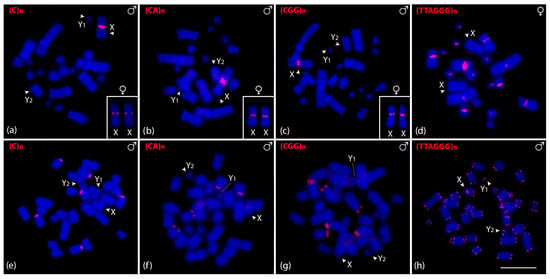
Figure 4.
Male and female metaphasic chromosomes of C. perspicillata (a–d) and male of A. lituratus (e–h) showing hybridization signals of CpeSatDNAs as probes. The name of the repeat motif is detailed in the upper left corner. The arrowheads indicate the sex chromosomes X, Y1, and Y2. Scale bar: 10 µm.
3.4. Cross-Species Mapping of satDNAs and Chromosomal Distribution of Microsatellites and Telomeric Repeats in Other Bat Species
All AliSatDNAs and CpeSatDNAs were in situ mapped in other Sternodermatinae species (D. cinerea, S. lilium, U. bilobatum, and U. magnirostrum) to check the presence of conserved satDNAs. Among all, only CpeSat02-24 and CpeSat05-72 presented positive signals (Figure S4). In D. cinerea and S. lilium, CpeSat02-24 and CpeSat05-72 showed clustering in the terminal region of all chromosomes, similar to those observed for A. lituratus. In U. bilobatum, besides CpeSat02-24, CpeSat05-72 presented strong signals on the X chromosome and on the terminal region of all chromosomes, and in U. magnistrostum, on the centromeric region of six autosome pairs (5, 6, 8, 10, 13 and 14), and on the X chromosome. Comparative hybridization using the general telomeric repeats (TTAGGG)n presented a pattern similar to that observed for the CpeSatDNAs (Figure S4c,f,i,l and Figure 4h).
The three satDNA repeats identified in C. perspicillata were additionally investigated in D. cinerea, S. lilium, Uroderma bilobatum, and U. magnirostrum (Figure 5). In D. cinerea females, (C)n and (CGG)n were mapped in one small chromosome pair and (CA)n on the short arm interstitial region of a small submetacentric pair (Figure 5a–c). The (C)n and (CA)n in the S. lilium male were clustered in the pericentromeric region of both X and Y, and (CGG)n only on the X chromosome. Additionally, we observed a centromeric signal in a large acrocentric pair using the (C)n and (CGG)n probes (Figure 5d–f). In the U. bilobatum, (C)n and (CA)n were clustered in the centromeric region of one chromosome pair and (CGG)n on two pairs, and also on the X (Figure 5g–i). The (C)n and (CA)n repeats did not present positive signals in U. magnirostrum chromosomes, and (CGG)n clustered in the terminal region of the short arms of two chromosome pairs (Figure 5j–l).
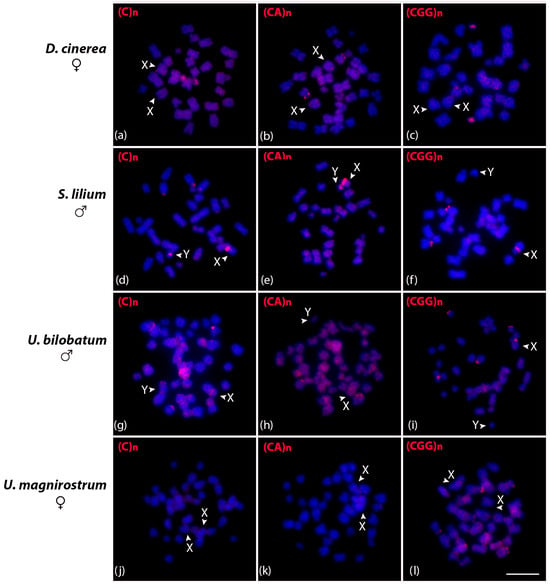
Figure 5.
Metaphasic chromosomes of D. cinerea female (a–c), S. lilium male (d–f), U. bilobatum male (g–i), and U. magnirostrum female (j–l) after in situ hybridization experiments with (C)n, (CA)n, and (GCC)n microsatellite probes indicated in red in the upper left corner. Scale bar: 10 µm.
3.5. Intraspecific Genomic Hybridization in C. perspicillata
To identify putative sex-specific regions in multiple systems, we compared male and female genomes of C. perspicillata. The results of genomic comparison detected overlapped centromeric and pericentromeric regions on all chromosomes, except for X, Y1, and Y2. An exclusive accumulation of male repeats was observed on the Y1 chromosome (Figure 6).
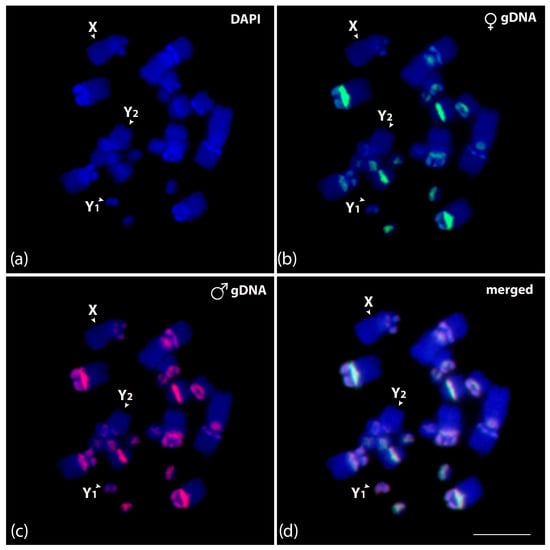
Figure 6.
Mitotic chromosome spreads of Carollia perspicillata male (a–d) after intraspecific genomic hybridization with female- and male-derived genomic probes. (a) DAPI-stained chromosomes (blue); (b) hybridization pattern for the female genomic DNA in green (Atto-488-dUTP labeled); (c) hybridization pattern for the male genomic DNA in red (Atto-550-dUTP labeled); and (d) merged images for DAPI-stained and both genomic DNA probes. According to morphology, the sex chromosomes are identified and indicated in each chromosome spread. Scale bar: 10 µm.
4. Discussion
In this study, we examined the repetitive DNA content of flying mammals at the genomic and chromosomal levels. The in silico data allowed us to identify two libraries composed of a few satellite DNA sequences, representing a low proportion of this repetitive sequence class in A. lituratus and C. perspicillata genomes; some of which present a direct association with telomeric sequences. The in situ mapping demonstrated the distribution of satDNAs in the telomeric and pericentromeric regions and microsatellite repeats in adjacent regions to the 45S rDNA cluster. The findings presented here revealed new occurrences between telomeric sequences and satellite DNA association in mammals, thereby providing insight into the involvement of these repeats in the evolutionary dynamics within these species.
4.1. General Features of A. lituratus and C. perspicillata Satellitomes
Prior characterization of satDNA in mammals revealed satellitomes consisting of a limited number of families, accounting for a minor fraction of repetitive DNA content (about 1% of the genome) [48,49]. The exception was related to rodent species of the genus Microtus, in which 17 satDNA families were identified, corresponding to 6% of the genome [50]. In bats, a significant presence of satDNAs was also observed in Perimyotis subflavus (Vespertilionidae), accounting for 6% of its genome [51]. In other bat species, however, the presence of some TE classes is higher in the genomic representation, like in A. lituratus (Phyllostomidae), in which about 15% of the genome is represented by the long interspersed element-1 (LINE-1) [51]. In our context, satDNAs constituted minor fractions of the genome, accounting for 0.014% in A. lituratus and 0.073% in C. perspicillata (Table 2), reinforcing that bats possess even smaller amounts of this repetitive DNA class than other mammals and how the satDNA content can vary within this group.
In most instances, satDNAs may be conserved among closely related organisms, including species within the same genus [50,52] or even across the same family or subfamily [50,53,54,55], as posited by the library hypothesis [56]. However, in some reports, arrays of satDNAs presented in the heterochromatic regions were considered a chromosome- or species-specific variant [57,58]. Here, we observed that the whole library was species-specific since the in-silico approach could not identify any common satellites between A. lituratus and C. perspicillata. These findings probably can be associated with the estimated divergence time of around 21.1 million years ago between these species (median time calculated by [59]) (Figure 7), along with the concept that satDNAs have been considered one of the most rapidly shifting repetitive DNAs in the genome [5]. Since we only analyzed one species per genus, comparative studies centered on satDNA analysis in other bat species within the Artibeus and Carollia genera could shed light on the conservation of these repeats.
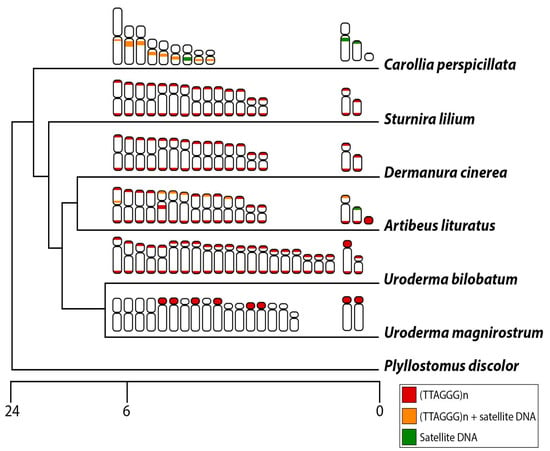
Figure 7.
Schematic representation of the phylogenetic relationship of phyllostomid species analyzed in the present work and the distribution of telomeric repeats and satellite DNA: exclusive distribution of (TTAGGG)5 (red), (TTAGGG)5 and satellite DNA (orange), and exclusive distribution of satellite DNA (green). The scale bar corresponds to million years ago (MYA).
Satellitome studies also have demonstrated that centromeric-associated repeats exhibit rapid changes and significant variability across closely related species, with mutation fixation facilitating the emergence of species-specific centromeric sequences [60,61,62]. Studies in mammals have connected satellite sequences to centromeric regions [48,53,63]. Possible CENP-B box-like motifs were found in the Neotropical deer Mazama gouazoubira and the Florida manatee subspecies Trichechus manatus latirostris [53,63]. The mostly satellite DNAs of C. perspicillata were found in the (peri)centromeric region of several chromosome pairs; however, BLAST analyses did not find any of them to be associated with putative centromeric satellite DNA or with any other repetitive sequence already described. Besides satellite DNAs, centromere-associated elements in plants also include some retrotransposons (reviewed in [64]). In this sense, investigations focused on the accurate centromere motif identification methods or even the identification of other repeats as main components of bat centromeres (e.g., LINE-1 retrotransposons, since it was largely found in the centromeric region [65]) can better elucidate the centromeric organization in bats.
4.2. Heterochromatin, Telomeric Repeats, and Satellite DNA Association
From plants to animals, the centromeres are frequently surrounded by pericentromeric regions enriched in constitutive heterochromatin (CH) and several different types of highly repetitive DNAs [66]. Among bats, incredible amounts of heterochromatin in interstitial chromosomal regions were reported [25,67,68], and the in situ mapping in phyllostomid bats demonstrates the presence of LINE elements [65] and satellite repeats, including telomeric sequences associated with these regions (present study).
According to our data, the distribution of the majority of CpeSatDNAs also coincides with the pericentromeric C-heterochromatin bands and telomeric motifs in C. perspicillata. In fact, CpeSat02-24 consists of the canonical telomeric repeat (TTAGGG)n (confirmed by in silico and in situ approaches). These repeats are also part of the CpeSat05-72 and homologies between both classified CpeSat02-24 and CpeSat05-72 as the same superfamily. The association of these satDNAs and telomeric sequences is also supported by fluorescence in situ experiments with CpeSat05-72, presenting signals associated with telomeric patterns in different bat species. Even in U. magnirostrum, with prominent centromeric signals in several chromosome pairs of CpeSat05-72, were also represented by telomeric repeats (Figure S4).
Telomeres in bats have been considered intriguing, especially regarding their unusual distribution across the chromosomes, frequently found in pericentromeric regions associated with CH [32]. In C. perspicillata, the chromosomal termini lack telomeric signals but exhibit significant accumulation in the pericentromeric region [32,69] (present work). This pattern was also identified in U. magnistrum, where telomeric repeats are highly amplified in the pericentromeric region of some chromosome pairs. Ref. [32], analyzing the distribution pattern of telomeric signals in phyllostomid bats, identified basal karyotypes possessing mainly telomeric signals. In fact, C. perspicillata and species of the Uroderma genus present the most divergent karyotypes within Phyllostomidae (reviewed in [20]). In the present study, terminal hybridization signals were observed in A. lituratus, D. cinerea, S. lilium, and U. bilobatum (Figure S4 and Figure 7). Even though U. bilobatum presents telomeric signals and U. magnirostrum centromeric accumulation of telomeric repeats, U. bilobatum species exhibit a more evolutionary derived karyotype compared to U. magnirostrum [68]. Considering the recent divergence (around 5 million years ago between U. bilobatum and U. magnirostrum), the telomeric repeats present very different distributions between Uroderma species, reinforcing how particular the evolution of telomeric sequence is in bats.
Regarding the absence of terminal signals using the canonical telomeric sequence, we must consider several possibilities, including the prospect of different sequences performing the role of chromosome stability, as seen in some insects [70,71], or even a different organization, similar to what has been reported in marsupials [72]. Furthermore, it is possible that the number of telomere repeats is below the detection limit for the method used, i.e., FISH. The observation in human molecular cytogenetics that even centromeric repeats present on a derived chromosome (detected by microdissection and reverse FISH) may not be detectable by FISH could support this possibility [73].
The telomeric repeats and satellite DNA association have been reported for some fish and plant species. In the Sparidae species Pagellus erythrinus (Acanthuriformes: Sparidae), sequencing analyses of Dra I satellite DNA indicated an association between satDNA and telomeric sequences, and in situ experiments revealed a subtelomeric distribution [74]. Similarly, a telomere-associated satellite DNA (RUSI) isolated from the Iberian sorrel Rumex induratus (Polygonales: Polygonaceae) contains different variants of degenerate telomere motifs, and in situ experiments revealed that the RUSI repeats are not only located at the subtelomeric, but also the centromeric region of some chromosomes [75]. On the other hand, in Thomas’s pine vole, Microtus thomasi (Rodentia: Cricetidae), telomeric sequences were massively amplified in the pericentromeric heterochromatin [76,77] and additionally, two variants of the canonical telomeric repeat (MthoSat4-6) were found [50]. Here we report two sequences, one corresponding to CpeSat02-24 and the other to CpeSat05-72, classified as the same superfamily (high similarity, but with some differences in the sequence content). It is reasonable that once these sequences are no longer located in the telomeric region, fulfilling their function of chromosome stability and protection, they tend to degenerate. Thus, mutational events may be the main cause for the presence of these variations of satDNA sequences.
In addition to telomeric studies, bats are also well characterized regarding the in situ mapping of multigenic rDNA families. While most bat species typically possess one autosomal pair containing the 45S rDNA cluster, episodes of sex-autosome translocation may result in this cluster being located on the sex chromosomes, similar to observations in Carollia, or distributed across multiple chromosomal sites, as noted in Artibeus [32]. The microsatellite repeats (CA)n, (C)n, and (CGG)n showed positive signals associated with the 45 rDNA cluster in C. perspicillata. This pattern was also observed in A. lituratus for (C)n and (CGG)n repeats. The association between multigene families and simple sequence repeats (SSRs) is very common and has been reported in other groups. In the armored catfish Rineloricaria latirostris (Siluriformes: Loricariidae) and in the freshwater stingray Potamotrygon schroederi (Myliobatiformes: Potamotrygonidae), SSRs were detected associated with the 5S and 45S rDNA genes, respectively [78,79]. Here, besides the microsatellites, the AliSat01-19 and AliSat02-51 satellites presented positive signals coincident with the location of the 45S rDNA sites in A. lituratus, reinforcing the view that the secondary constriction regions of bats are enriched in microsatellite and satellite DNAs. Variation in the size and composition of the intergenic spacer of rDNA is frequently associated with the presence of several repetitive DNA classes [80]. In this sense, detailed investigations, including the characterization of these regions in bats, can confirm the results obtained from in situ mapping.
4.3. Repetitive DNAs and the Evolution of Sex Chromosomes in Bats
Most Carolliinae and Sternodermatinae species are characterized by having multiple or compound sex chromosome systems with a high frequency of chromosomal rearrangements [27,29,81,82,83,84]. In multiple sex chromosomes, where a Y fission triggers two chromosomes, named Y1 and Y2 chromosomes, the true male sex chromosome of the system is Y1, while Y2 represents the autosome involved in the translocation/fission. In accordance with earlier cytogenetics reports that the Y1 is the most likely carrier of the male-specific genes [29,85], the results of the intraspecific CGH experiment in C. perspicillata highlight a male-specific region on the Y1 chromosome. A. lituratus harbors the same system as C. perspicillata; however, with a different origin, since different chromosomes are involved in the origin [81]. We hypothesized a similar pattern with the Y1-carrying male-specific sequences; however, we couldn’t assume this pattern, since no female gDNA was available for CGH experiments. Intraspecific CGH in XY systems also reveals male-specific regions in rodents and primates [47,86], marking this as the first intraspecific hybridization in a multiple sex chromosome system in mammals.
The analysis of chromosomal distribution of repetitive DNAs is sparse and limited in bats [65], especially with a focus on the sex chromosomes. In the present work, satellite distribution of repetitive DNAs in multiple sex chromosomes includes AliSat01-19 in A. lituratus, clustered in some autosome pairs but also on the X and Y2 chromosomes, being absent on the Y1. In this case, the enrichment of satellite DNA on both X and Y2 chromosomes can be related to the two autosomal segments involved in the translocation, reinforcing their common origin. In C. perspicillata, only CpeSat04-535 and CpeSat07-1531 were clustered in the X and Y2, respectively. So, specifically for CpeSat04-535, the loss of repetitive sequences after the fission event appears to be a characteristic of this multiple chromosome system, since few sequences were accumulated on the X chromosome and were absent or with few copy numbers on the Y chromosomes. Besides satDNAs, microsatellite repeats are the only types of sequences investigated in this study that are grouped on the X chromosomes of C. perspicillata, related to the rDNA 45S cluster as mentioned earlier. In this case, it is feasible that this accumulation of repetitive sequences may stabilize the chromosomal rearrangement, since they were mapped at the fusion point between the original X chromosome and the autosome (Figure 4a–c).
Compound sex chromosome systems in bats are characterized by a secondary translocation that reorganized the sex chromosomes through centric fusion, originating neo-sex chromosome systems. Between Artibeus planirostris and Sturnira lilium, the only difference observed is related to the sex chromosome system involving a reciprocal X/autosome translocation, characterizing a neo-XY system [30]. Here, in S. lilium, we observed a significant accumulation of microsatellite repeats on the sex chromosomes. While (C)n was distributed in both autosomal and sex chromosomes, (CA)n appears to be a sex-chromosome-specific marker present in both X and Y. On the other hand, the pattern present with (CGG)n indicates a loss of this sequence or diminution in copy number on the Y. In grasshoppers, other repetitive DNAs analyzed on neo-sex chromosomes highlight an intensive accumulation reinforcing the differentiation and high turnover events of neo-sex chromosomes [87,88,89,90]. Similar to those found in insects, bats also represent a group with high turnovers of neo-sex chromosomes. In this sense, studies with a focus on the characterization of other repetitive sequences can shed light on the molecular differentiation of the few explored systems in this mammal group.
Although Uroderma species do not exhibit enrichment of microsatellite repeats, similar to those repeats found in S. lilium, telomeric sequences were highly prevalent on the X chromosomes of U. bilobatum (Figure S4). In U. magnirostrum, the large accumulation of telomeric sequences in the centromeric region of several autosome pairs is also present on the X chromosome. Since we have not performed in situ hybridization on male chromosomes, it is not possible to confirm whether its accumulation was lost in the Y; however, an expected pattern similar to that of U. bilobatum is possible. Taken together, our study points to a significant contribution of different repetitive DNAs in the sex chromosome differentiation in bats. Moreover, investigations involving other microsatellite repeats and the characterization of satDNA libraries from other bat species might enhance the understanding of karyotypic evolution related to the processes that trigger chromosomal rearrangement events and sex chromosome differentiation in mammals.
5. Conclusions
This study characterized satellite DNA sequences and analyzed the distribution of repetitive repeats for the first time from multiple phyllostomid bat species, which have been poorly investigated. Our study confirmed the hypothesis that the reduced genome size in bats is possibly associated with a loss of repetitive sequences, especially satellite DNAs. Furthermore, our study also demonstrated the presence of a few satellite families that were associated with heterochromatic regions, similar to what has been reported previously for other mammals, thereby underscoring the significance of satellite DNA as a principal content of heterochromatin. We also demonstrated that in two of our species, only a few satellites accumulated on the sex chromosomes, and in C. perspicillata, some satellites were associated with telomeric sequences. The absence of terminal signals relative to canonical telomeric repeats in some bat species remains intriguing. Future investigations with whole-genome sequencing can offer previously unknown details about this organization.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/biom15091248/s1. Figure S1. Repeat landscape of (a) AliSatDNAs and (b) CpeSatDNAs in the male genome of Artibeus lituratus and Carollia perspicillata species, respectively. Figure S2. Female metaphasic chromosomes of C. perspicillata showing hybridization signals of CpeSatDNAs as probes (a–i). The name of each satellite DNA family is detailed in the upper left corner in red (Atto-550-dUTP labeled). The arrowheads indicate the sex chromosomes X, Y1, and Y2. Scale bar: 10 µm. Figure S3. Male and female metaphasic chromosomes of bat species analyzed in the present work after C-positive heterochromatin detection: (a) Artibeus lituratus, (b) Carollia perspicillata, (c) Dermanura cinerea, (d) Sturnira lilium, (e) Uroderma bilobatum, and (f) Uroderma magnirostrum. Scale bar: 10 µm. Figure S4. Male and female metaphasic chromosome plates of A. lituratus, D. cinerea, S. lilium, U. bilobatum, and U. magnirostrum for comparative hybridizations using the Telomeric (TTAGGG)n and CpeSatDNAs repeats. Scale bar: 10 µm. Table S1: Designed primers for all satellite DNAs analyzed in the present study. The satDNAs with repeat unit length smaller than 31 bp were directly labeled with Cy3 at the 5′ end during the synthesis (*).
Author Contributions
Conceptualization, M.d.B.C.; Data curation, M.d.B.C.; Formal analysis, G.A.D., A.J.B.G., P.G.S., E.H.C.d.O., F.P.-F., R.U., T.L. and M.d.B.C.; Funding acquisition, M.d.B.C.; Investigation, G.A.D., T.E., J.H.F.S., R.U. and M.d.B.C.; Methodology, G.A.D., T.E., J.H.F.S., R.Z.d.S., A.J.B.G., P.G.S., E.H.C.d.O., R.U. and M.d.B.C.; Project administration, M.d.B.C.; Resources, T.L.; Software, G.A.D., T.E., R.Z.d.S., F.P.-F., R.U. and M.d.B.C.; Supervision, M.d.B.C.; Validation, G.A.D., P.G.S., E.H.C.d.O. and T.L.; Writing—original draft, G.A.D.; Writing—review & editing, T.E., J.H.F.S., R.Z.d.S., A.J.B.G., P.G.S., E.H.C.d.O., F.P.-F., R.U., T.L. and M.d.B.C. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the São Paulo Research Foundation (FAPESP) grant 2021/14096-6 and 2023/15593-9 (G.A.D.) and 2023/00955–2 (M.B.C.), and Brazilian National Council for Scientific and Technological Development (CNPq), grant number 302928/2021–9 (M.B.C). We acknowledge support by the German Research Foundation Projekt—Nr. 512648189 (T.L.) and the Open Access Publication Fund of the Thueringer Universitaets—und Landesbibliothek Jena. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior, Brasil (CAPES), Finance Code 001.
Institutional Review Board Statement
The specimens were collected with authorization from the National System for the Management of Genetic Heritage and Associated Traditional Knowledge (SISGEN-A96FF09), the Chico Mendes Institute for Biodiversity Conservation (ICMBIO), and the System of Authorization and Information about Biodiversity (SISBIO) under license number 92665-1. The experiments were conducted in accordance with the Ethics Committee on Animal Experimentation of the Universidade Federal de São Carlos, Brazil (Process number 7994170423).
Informed Consent Statement
Not applicable.
Data Availability Statement
The datasets generated during and/or analyzed during the current study are available from the corresponding author upon reasonable request. The datasets generated and analyzed during the current study are available in the GenBank repository under the accession numbers PV067726-PV067729 and PV067174-PV067183.
Acknowledgments
The authors thank Ana Lilian Luz Rodrigues and Sara Santos da Conceição for their support in the field expeditions and chromosome preparations.
Conflicts of Interest
The authors declare no conflicts of interest.
Abbreviations
The following abbreviations are used in this manuscript:
| TE | Transposable elements |
| satDNA | Satellite DNA |
| rDNA | Ribosomal DNA |
| 2n | Diploid number |
| ITS | Interstitial Telomeric Sites |
| gDNA | Genomic DNA |
| RE2 | RepeatExplorer2 |
| PCR | Polymerase Chain Reaction |
| RUL | Repeat Unit Length |
| FISH | Fluorescence in situ Hybridization |
| CGH | Comparative Genomic Hybridization |
| CH | Constitutive Heterochromatin |
| SSRs | Simple Sequence Repeats |
References
- Biscotti, M.A.; Olmo, E.; Heslop-Harrison, J.S. Repetitive DNA in Eukaryotic Genomes. Chromosome Res. 2015, 23, 415–420. [Google Scholar] [CrossRef]
- Paço, A.; Freitas, R.; Vieira-da-Silva, A. Conversion of DNA Sequences: From a Transposable Element to a Tandem Repeat or to a Gene. Genes 2019, 10, 1014. [Google Scholar] [CrossRef]
- Padeken, J.; Zeller, P.; Gasser, S.M. Repeat DNA in Genome Organization and Stability. Curr. Opin. Genet. Dev. 2015, 31, 12–19. [Google Scholar] [CrossRef]
- Pavlek, M.; Gelfand, Y.; Plohl, M.; Meštrović, N. Genome-Wide Analysis of Tandem Repeats in Tribolium castaneum Genome Reveals Abundant and Highly Dynamic Tandem Repeat Families with Satellite DNA Features in Euchromatic Chromosomal Arms. DNA Res. 2015, 22, 387–401. [Google Scholar] [CrossRef] [PubMed]
- Garrido-Ramos, M. Satellite DNA: An Evolving Topic. Genes 2017, 8, 230. [Google Scholar] [CrossRef]
- Feliciello, I.; Pezer, Ž.; Kordiš, D.; Bruvo Mađarić, B.; Ugarković, Đ. Evolutionary History of Alpha Satellite DNA Repeats Dispersed within Human Genome Euchromatin. Genome Biol. Evol. 2020, 12, 2125–2138. [Google Scholar] [CrossRef]
- Rico-Porras, J.M.; Mora, P.; Palomeque, T.; Montiel, E.E.; Cabral-de-Mello, D.C.; Lorite, P. Heterochromatin is not the only place for satDNAs: The High Diversity of satDNAs in the Euchromatin of the Beetle Chrysolina americana (Coleoptera, Chrysomelidae). Genes 2024, 15, 395. [Google Scholar] [CrossRef]
- Hedges, D.J.; Deininger, P.L. Inviting Instability: Transposable Elements, Double-Strand Breaks, and the Maintenance of Genome Integrity. Mutat. Res./Fundam. Mol. Mech. Mutagen. 2007, 616, 46–59. [Google Scholar] [CrossRef]
- Carbone, L.; Alan Harris, R.; Gnerre, S.; Veeramah, K.R.; Lorente-Galdos, B.; Huddleston, J.; Meyer, T.J.; Herrero, J.; Roos, C.; Aken, B.; et al. Gibbon Genome and the Fast Karyotype Evolution of Small Apes. Nature 2014, 513, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Paço, A.; Adega, F.; Meštrović, N.; Plohl, M.; Chaves, R. The Puzzling Character of Repetitive DNA in Phodopus Genomes (Cricetidae, Rodentia). Chromosome Res. 2015, 23, 427–440. [Google Scholar] [CrossRef] [PubMed]
- Gregory, T.R. Animal Genome Size Database 2025; Elsevier Academic Press: San Diego, CA, USA, 2025. [Google Scholar]
- Mammal Diversity Database. 2024. Available online: https://www.mammaldiversity.org/ (accessed on 27 July 2025).
- Smith, J.D.L.; Gregory, T.R. The genome sizes of Megabats (Chiroptera: Pteropodidae) are remarkably constrained. Biol. Lett. 2009, 5, 347–351. [Google Scholar] [CrossRef]
- Smith, J.D.L.; Bickham, J.W.; Gregory, T.R. Patterns of Genome Size Diversity in Bats (Order Chiroptera). Genome 2013, 56, 457–472. [Google Scholar] [CrossRef]
- Kapusta, A.; Suh, A.; Feschotte, C. Dynamics of Genome Size Evolution in Birds and Mammals. Proc. Natl. Acad. Sci. USA 2017, 114, E1460–E1469. [Google Scholar] [CrossRef]
- de Oliveira, A.M.; Souza, G.M.; Toma, G.A.; Dos Santos, N.; Dos Santos, R.Z.; Goes, C.A.G.; Deon, G.A.; Setti, P.G.; Porto-Foresti, F.; Utsunomia, R.; et al. Satellite DNAs, Heterochromatin, and Sex Chromosomes of the Wattled Jacana (Charadriiformes; Jacanidae): A Species with Highly Rearranged Karyotype. Genome 2024, 67, 109–118. [Google Scholar] [CrossRef]
- Kretschmer, R.; Toma, G.A.; Deon, G.A.; Dos Santos, N.; Dos Santos, R.Z.; Utsunomia, R.; Porto-Foresti, F.; Gunski, R.J.; Garnero, A.D.V.; Liehr, T.; et al. Satellitome Analysis in the Southern Lapwing (Vanellus chilensis) Genome: Implications for SatDNA Evolution in Charadriiform Birds. Genes 2024, 15, 258. [Google Scholar] [CrossRef]
- Setti, P.G.; Deon, G.A.; Zeni Dos Santos, R.; Goes, C.A.G.; Garnero, A.D.V.; Gunski, R.J.; De Oliveira, E.H.C.; Porto-Foresti, F.; De Freitas, T.R.O.; Silva, F.A.O.; et al. Evolution of Bird Sex Chromosomes: A Cytogenomic Approach in Palaeognathae Species. BMC Ecol. Evol. 2024, 24, 51. [Google Scholar] [CrossRef] [PubMed]
- Souza, G.M.; Kretschmer, R.; Toma, G.A.; De Oliveira, A.M.; Deon, G.A.; Setti, P.G.; Zeni Dos Santos, R.; Goes, C.A.G.; Del Valle Garnero, A.; Gunski, R.J.; et al. Satellitome Analysis on the Pale-Breasted Thrush Turdus leucomelas (Passeriformes; Turdidae) Uncovers the Putative Co-Evolution of Sex Chromosomes and Satellite DNAs. Sci. Rep. 2024, 14, 20656. [Google Scholar] [CrossRef]
- Sotero-Caio, C.; Baker, R.; Volleth, M. Chromosomal Evolution in Chiroptera. Genes 2017, 8, 272. [Google Scholar] [CrossRef] [PubMed]
- Simmons, N.; Cirranello, A. Bat Species of the World: A Taxonomic and Geographic Database. 2024. Available online: https://batnames.org/ (accessed on 27 July 2025).
- Cirranello, A.; Simmons, N.B.; Solari, S.; Baker, R.J. Morphological Diagnoses of Higher-Level Phyllostomid Taxa (Chiroptera: Phyllostomidae). Acta Chiropterologica 2016, 18, 39–71. [Google Scholar] [CrossRef]
- Castillo-Figueroa, D. Ecological Morphology of Neotropical Bat Wing Structures. Zool. Stud. 2020, 59, e60. [Google Scholar] [CrossRef] [PubMed]
- Baker, R.J.; Hood, C.S.; Honeycutt, R.L. Phylogenetic Relationships and Classification of the Higher Categories of the New World Bat Family Phyllostomidae. Syst. Zool. 1989, 38, 228. [Google Scholar] [CrossRef]
- Varella-Garcia, M.; Taddei, V.A. Citogenética de Quirópteros: Métodos e Aplicações. Rev. Bras. Zool. 1989, 6, 297–323. [Google Scholar] [CrossRef]
- Ortêncio-Filho, H.; de Lima, I.P.; Fogaça, F.N.O. Subfamília Caroliinae. In Morcegos do Brasil; Adriano, L.P., Wagner, A.P., Isaac, P.d.L., Eds.; Nelio R. dos Reis: Londrina, Brazil, 2007; p. 253. ISBN 978-85-906395-1-0. [Google Scholar]
- Hsu, T.C.; Baker, R.J.; Utakoji, T. The Multiple Sex Chromosome System of American Leaf-Nosed Bats (Chiroptera, Phyllostomidae). Cytogenet. Genome Res. 1968, 7, 27–38. [Google Scholar] [CrossRef]
- Stock, D. Chromosome Banding Pattern Homology and Its Phylogenetic Implications in the Bat Genera Carollia and Choeroniscus. Cytogenet. Genome Res. 1975, 14, 34–41. [Google Scholar] [CrossRef]
- Tucker, P.K. Sex Chromosome-Autosome Translocations in the Leaf-Nosed Bats, Family Phyllostomidae. Cytogenet. Genome Res. 1986, 43, 19–27. [Google Scholar] [CrossRef]
- Faria, K.C.; Morielle-Versute, E. Genetic Relationships Between Brazilian Species of Molossidae and Phyllostomidae (Chiroptera, Mammalia). Genetica 2006, 126, 215–225. [Google Scholar] [CrossRef]
- Tucker, P.K.; Bickham, J.W. Sex Chromosome-Autosome Translocations in the Leaf-Nosed Bats, Family Phyllostomidae. Cytogenet. Genome Res. 1986, 43, 28–37. [Google Scholar] [CrossRef]
- Calixto, M.D.S.; De Andrade, I.S.; Cabral-de-Mello, D.C.; Santos, N.; Martins, C.; Loreto, V.; De Souza, M.J. Patterns of rDNA and Telomeric Sequences Diversification: Contribution to Repetitive DNA Organization in Phyllostomidae Bats. Genetica 2014, 142, 49–58. [Google Scholar] [CrossRef]
- Baker, R.J.; Hamilton, M.J.; Parish, D.A. Preparations of Mammalian Karyotypes Under Field Conditions; Occasional papers; Museum of Texas Tech University: Lubbock, TX, USA, 2003. [Google Scholar]
- Moratelli, R.; Andrade, C.M.; Armada, J.L.A. A Technique to Obtain Fibroblast Cells from Skin Biopsies of Living Bats (Chiroptera) for Cytogenetic Studies. Genet. Mol. Res. 2002, 1, 128–130. [Google Scholar]
- Sambrook, J.; Russell, D.W. Isolation of High-molecular-weight DNA from Mammalian Cells Using Proteinase K and Phenol. CSH Protocol. 2006, 2006, pdb.prot4036. [Google Scholar] [CrossRef]
- Ruiz-Ruano, F.J.; López-León, M.D.; Cabrero, J.; Camacho, J.P.M. High-Throughput Analysis of the Satellitome Illuminates Satellite DNA Evolution. Sci. Rep. 2016, 6, 28333. [Google Scholar] [CrossRef]
- Utsunomia, R.; Silva, D.M.Z.D.A.; Ruiz-Ruano, F.J.; Goes, C.A.G.; Melo, S.; Ramos, L.P.; Oliveira, C.; Porto-Foresti, F.; Foresti, F.; Hashimoto, D.T. Satellitome Landscape Analysis of Megaleporinus macrocephalus (Teleostei, Anostomidae) Reveals Intense Accumulation of Satellite Sequences on the Heteromorphic Sex Chromosome. Sci. Rep. 2019, 9, 5856. [Google Scholar] [CrossRef]
- Novák, P.; Neumann, P.; Pech, J.; Steinhaisl, J.; Macas, J. RepeatExplorer: A Galaxy-Based Web Server for Genome-Wide Characterization of Eukaryotic Repetitive Elements from next-Generation Sequence Reads. Bioinformatics 2013, 29, 792–793. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A Flexible Trimmer for Illumina Sequence Data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Schmieder, R.; Edwards, R. Fast Identification and Removal of Sequence Contamination from Genomic and Metagenomic Datasets. PLoS ONE 2011, 6, e17288. [Google Scholar] [CrossRef]
- Kearse, M.; Moir, R.; Wilson, A.; Stones-Havas, S.; Cheung, M.; Sturrock, S.; Buxton, S.; Cooper, A.; Markowitz, S.; Duran, C.; et al. Geneious Basic: An Integrated and Extendable Desktop Software Platform for the Organization and Analysis of Sequence Data. Bioinformatics 2012, 28, 1647–1649. [Google Scholar] [CrossRef]
- Smit, A.; Hubley, R.; Green, P. RepeatMasker Open-4.0. 2013–2015. Available online: http://www.repeatmasker.org (accessed on 15 May 2024).
- IJdo, J.W.; Wells, R.A.; Baldini, A.; Reeders, S.T. Improved Telomere Detection Using a Telomere Repeat Probe (TTAGGG)n Generated by PCR. Nucl. Acids Res. 1991, 19, 4780. [Google Scholar] [CrossRef] [PubMed]
- Yano, C.F.; Bertollo, L.A.C.; Cioffi, M.D.B. Fish-FISH: Molecular Cytogenetics in Fish Species. In Fluorescence In Situ Hybridization (FISH); Liehr, T., Ed.; Springer Protocols Handbooks; Springer: Berlin/Heidelberg, Germany, 2017; pp. 429–443. ISBN 978-3-662-52957-7. [Google Scholar]
- Seabright, M. A rapid banding technique for human chromosomes. Lancet 1971, 298, 971–972. [Google Scholar] [CrossRef] [PubMed]
- Sumner, A.T. A Simple Technique for Demonstrating Centromeric Heterochromatin. Exp. Cell Res. 1972, 75, 304–306. [Google Scholar] [CrossRef]
- Traut, W.; Eickhoff, U.; Schorch, J.-C. Identification and Analysis of Sex Chromosomes by Comparative Genomic Hybridization (CGH). In Chromosome Painting; Sharma, A.K., Sharma, A., Eds.; Springer: Dordrecht, The Netherland, 2001; pp. 155–161. ISBN 978-94-010-3840-9. [Google Scholar]
- Valeri, M.P.; Dias, G.B.; Do Espírito Santo, A.A.; Moreira, C.N.; Yonenaga-Yassuda, Y.; Sommer, I.B.; Kuhn, G.C.S.; Svartman, M. First Description of a Satellite DNA in Manatees’ Centromeric Regions. Front. Genet. 2021, 12, 694866. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, J.; Aleix-Mata, G.; Montiel, E.E.; Cabral-de-Mello, D.C.; Marchal, J.A.; Sánchez, A. Satellitome Analysis on Talpa aquitania Genome and Inferences about the satDNAs Evolution on Some Talpidae. Genes 2022, 14, 117. [Google Scholar] [CrossRef]
- Aleix Mata, G.; Montiel, E.E.; Mora-Ruiz, P.; Yurchenko, A.; Rico-Porras, J.M.; Anguita, F.; Palomo, F.; Marchal, J.A.; Rovatsos, M.; Sánchez Baca, A. Satellitome Analysis on Microtus thomasi (Arvicolinae) Genome, a Mammal Species with High Karyotype and Sex Chromosome Variations. Genome 2025, 68, gen-2024-0141. [Google Scholar] [CrossRef] [PubMed]
- Pagan, H.J.T.; Macas, J.; Novak, P.; McCulloch, E.S.; Stevens, R.D.; Ray, D.A. Survey Sequencing Reveals Elevated DNA Transposon Activity, Novel Elements, and Variation in Repetitive Landscapes among Vesper Bats. Genome Biol. Evol. 2012, 4, 575–585. [Google Scholar] [CrossRef] [PubMed]
- Kretschmer, R.; Goes, C.A.G.; Bertollo, L.A.C.; Ezaz, T.; Porto-Foresti, F.; Toma, G.A.; Utsunomia, R.; De Bello Cioffi, M. Satellitome Analysis Illuminates the Evolution of ZW Sex Chromosomes of Triportheidae Fishes (Teleostei: Characiformes). Chromosoma 2022, 131, 29–45. [Google Scholar] [CrossRef] [PubMed]
- Vozdova, M.; Kubickova, S.; Martínková, N.; Galindo, D.J.; Bernegossi, A.M.; Cernohorska, H.; Kadlcikova, D.; Musilová, P.; Duarte, J.M.; Rubes, J. Satellite DNA in Neotropical Deer Species. Genes 2021, 12, 123. [Google Scholar] [CrossRef]
- Camacho, J.P.M.; Cabrero, J.; López-León, M.D.; Martín-Peciña, M.; Perfectti, F.; Garrido-Ramos, M.A.; Ruiz-Ruano, F.J. Satellitome Comparison of Two Oedipodine Grasshoppers Highlights the Contingent Nature of Satellite DNA Evolution. BMC Biol. 2022, 20, 36. [Google Scholar]
- Yurchenko, A.; Pšenička, T.; Mora, P.; Ortega, J.A.M.; Baca, A.S.; Rovatsos, M. Cytogenetic Analysis of Satellitome of Madagascar Leaf-Tailed Geckos. Genes 2024, 15, 429. [Google Scholar] [CrossRef]
- Fry, K.; Salser, W. Nucleotide Sequences of HS-α Satellite DNA from Kangaroo Rat Dipodomys ordii and Characterization of Similar Sequences in Other Rodents. Cell 1977, 12, 1069–1084. [Google Scholar] [CrossRef]
- Garrido-Ramos, M.A. Satellite DNA in Plants: More than Just Rubbish. Cytogenet. Genome Res. 2015, 146, 153–170. [Google Scholar] [CrossRef]
- Šatović-Vukšić, E.; Plohl, M. Satellite DNAs—From Localized to Highly Dispersed Genome Components. Genes 2023, 14, 742. [Google Scholar] [CrossRef]
- Kumar, S.; Suleski, M.; Craig, J.M.; Kasprowicz, A.E.; Sanderford, M.; Li, M.; Stecher, G.; Hedges, S.B. TimeTree 5: An Expanded Resource for Species Divergence Times. Mol. Biol. Evol. 2022, 39, msac174. [Google Scholar] [CrossRef] [PubMed]
- Henikoff, S.; Ahmad, K.; Malik, H.S. The Centromere Paradox: Stable Inheritance with Rapidly Evolving DNA. Science 2001, 293, 1098–1102. [Google Scholar] [CrossRef]
- Zhang, H.; Koblížková, A.; Wang, K.; Gong, Z.; Oliveira, L.; Torres, G.A.; Wu, Y.; Zhang, W.; Novák, P.; Buell, C.R.; et al. Boom-Bust Turnovers of Megabase-Sized Centromeric DNA in Solanum Species: Rapid Evolution of DNA Sequences Associated with Centromeres. Plant Cell 2014, 26, 1436–1447. [Google Scholar] [CrossRef]
- Huang, Y.; Ding, W.; Zhang, M.; Han, J.; Jing, Y.; Yao, W.; Hasterok, R.; Wang, Z.; Wang, K. The Formation and Evolution of Centromeric Satellite Repeats in Saccharum Species. Plant J. 2021, 106, 616–629. [Google Scholar] [CrossRef]
- Sena, R.S.; Heringer, P.; Valeri, M.P.; Pereira, V.S.; Kuhn, G.C.S.; Svartman, M. Identification and Characterization of Satellite DNAs in Two-Toed Sloths of the Genus Choloepus (Megalonychidae, Xenarthra). Sci. Rep. 2020, 10, 19202. [Google Scholar] [CrossRef]
- Kejnovsky, E.; Jedlicka, P.; Lexa, M.; Kubat, Z. Factors Determining Chromosomal Localization of Transposable Elements in Plants. Plant Biol. J. 2025. [Google Scholar] [CrossRef]
- Sotero-Caio, C.G.; Cabral-de-Mello, D.C.; Calixto, M.D.S.; Valente, G.T.; Martins, C.; Loreto, V.; De Souza, M.J.; Santos, N. Centromeric Enrichment of LINE-1 Retrotransposons and Its Significance for the Chromosome Evolution of Phyllostomid Bats. Chromosome Res. 2017, 25, 313–325. [Google Scholar] [CrossRef]
- Thakur, J.; Packiaraj, J.; Henikoff, S. Sequence, Chromatin and Evolution of Satellite DNA. IJMS 2021, 22, 4309. [Google Scholar] [CrossRef] [PubMed]
- Santos, N.; Fagundes, V.; Yonenaga-Yassuda, Y.; Jose, M. Comparative Karyology of Brazilian Vampire Bats Desmodus rotundus and Diphylla ecaudata (Phyllostomidae, Chiroptera): Banding Patterns, Base-Specific Fluorochromes and FISH of Ribosomal Genes. Hereditas 2001, 134, 189–194. [Google Scholar] [CrossRef]
- Silva, A.M.D.; Marques-Aguiar, S.A.; Barros, R.M.D.S.; Nagamachi, C.Y.; Pieczarka, J.C. Comparative Cytogenetic Analysis in the Species Uroderma magnirostrum and U. bilobatum (Cytotype 2n = 42) (Phyllostomidae, Stenodermatinae) in the Brazilian Amazon. Genet. Mol. Biol. 2005, 28, 248–253. [Google Scholar] [CrossRef]
- Faria, K.D.C.; Morielle-Versute, E. In Situ Hybridization of Bat Chromosomes with Human (TTAGGG)n Probe, after Previous Digestion with Alu I. Genet. Mol. Biol. 2002, 25, 365–371. [Google Scholar] [CrossRef][Green Version]
- Cabral-de-Mello, D.C.; Palacios-Gimenez, O.M. Repetitive DNAs: The ‘Invisible’ Regulators of Insect Adaptation and Speciation. Curr. Opin. Insect Sci. 2025, 67, 101295. [Google Scholar] [CrossRef]
- Gotoh, T.; Suzuki, H.; Moriyama, M.; Futahashi, R.; Osanai-Futahashi, M. Acquisition and Repeated Alteration of (TTGGG)n Telomeric Repeats in Odonata (Dragonflies and Damselflies). Insect. Biochem. Mol. Biol. 2025, 182, 104353. [Google Scholar] [CrossRef]
- Ingles, E.D.; Deakin, J.E. Global DNA Methylation Patterns on Marsupial and Devil Facial Tumour Chromosomes. Mol. Cytogenet. 2015, 8, s13039–s14015. [Google Scholar] [CrossRef]
- Liehr, T.; Starke, H.; Senger, G.; Melotte, C.; Weise, A.; Vermeesch, J.R. Overrepresentation of small supernumerary marker chromosomes (sSMC) from chromosome 6 origin in cases with multiple sSMC. Am. J. Med. Genet.-Part A 2006, 140, 46–51. [Google Scholar] [CrossRef] [PubMed]
- Garrido-Ramos, M.A.; De La Herrán, R.; Ruiz Rejón, C.; Ruiz Rejón, M. A Satellite DNA of the Sparidae Family (Pisces, Perciformes) Associated with Telomeric Sequences. Cytogenet. Genome Res. 1998, 83, 3–9. [Google Scholar] [CrossRef]
- Navajas-Pérez, R.; Quesada Del Bosque, M.E.; Garrido-Ramos, M.A. Effect of Location, Organization, and Repeat-Copy Number in Satellite-DNA Evolution. Mol. Genet Genom. 2009, 282, 395–406. [Google Scholar] [CrossRef]
- Rovatsos, M.T.; Marchal, J.A.; Romero-Fernández, I.; Fernández, F.J.; Giagia-Athanosopoulou, E.B.; Sánchez, A. Rapid, Independent, and Extensive Amplification of Telomeric Repeats in Pericentromeric Regions in Karyotypes of Arvicoline Rodents. Chromosome Res. 2011, 19, 869–882. [Google Scholar] [CrossRef] [PubMed]
- Rovatsos, M.; Marchal, J.A.; Giagia-Athanasopoulou, E.; Sánchez, A. Molecular Composition of Heterochromatin and Its Contribution to Chromosome Variation in the Microtus thomasi/Microtus atticus Species Complex. Genes 2021, 12, 807. [Google Scholar] [CrossRef] [PubMed]
- Glugoski, L.; Nogaroto, V.; Deon, G.A.; Azambuja, M.; Moreira-Filho, O.; Vicari, M.R. Enriched Tandem Repeats in Chromosomal Fusion Points of Rineloricaria latirostris (Boulenger, 1900) (Siluriformes: Loricariidae). Genome 2022, 65, 479–489. [Google Scholar] [CrossRef]
- Ferreira, A.M.V.; Viana, P.F.; Marajó, L.; Feldberg, E. First Karyotypic Insights into Potamotrygon schroederi Fernández-Yépez, 1958: Association of Different Classes of Repetitive DNA. Cytogenet. Genome Res. 2024, 164, 60–68. [Google Scholar] [CrossRef]
- Kim, T.-S.; Hong, C.-Y.; Oh, S.-J.; Choe, Y.-H.; Hwang, T.-S.; Kim, J.; Lee, S.-L.; Yoon, H.; Bok, E.-Y.; Cho, A.; et al. RNA Sequencing Provides Novel Insights into the Pathogenesis of Naturally Occurring Myxomatous Mitral Valve Disease Stage B1 in Beagle Dogs. PLoS ONE 2024, 19, e0300813. [Google Scholar] [CrossRef]
- Rodrigues, L.R.R.; Barros, R.M.S.; Marques-Aguiar, S.; Assis, M.D.F.L.; Pieczarka, J.C.; Nagamachi, C.Y. Comparative Cytogenetics of Two Phyllostomids Bats. A New Hypothesis to the Origin of the Rearranged X Chromosome from Artibeus Lituratus (Chiroptera, Phyllostomidae). Caryologia 2003, 56, 413–419. [Google Scholar] [CrossRef]
- Noronha, R.C.R.; Nagamachi, C.Y.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; Pieczarka, J.C. Meiotic Analysis of XX/XY and Neo-XX/XY Sex Chromosomes in Phyllostomidae by Cross-Species Chromosome Painting Revealing a Common Chromosome 15-XY Rearrangement in Stenodermatinae. Chromosome Res. 2010, 18, 667–676. [Google Scholar] [CrossRef] [PubMed]
- Pieczarka, J.C.; Gomes, A.J.B.; Nagamachi, C.Y.; Rocha, D.C.C.; Rissino, J.D.; O’Brien, P.C.M.; Yang, F.; Ferguson-Smith, M.A. A Phylogenetic Analysis Using Multidirectional Chromosome Painting of Three Species (Uroderma magnirostrum, U. bilobatum and Artibeus obscurus) of Subfamily Stenodermatinae (Chiroptera-Phyllostomidae). Chromosome Res. 2013, 21, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Gomes, A.J.B.; Nagamachi, C.Y.; Rodrigues, L.R.R.; Benathar, T.C.M.; Ribas, T.F.A.; O’Brien, P.C.M.; Yang, F.; Ferguson-Smith, M.A.; Pieczarka, J.C. Chromosomal Phylogeny of Vampyressine Bats (Chiroptera, Phyllostomidae) with Description of Two New Sex Chromosome Systems. BMC Evol. Biol. 2016, 16, 119. [Google Scholar] [CrossRef]
- Noronha, R.C.R.; Nagamachi, C.Y.; O’Brien, P.C.M.; Ferguson-Smith, M.A.; Pieczarka, J.C. Neo-XY Body: An Analysis of XY1Y2 Meiotic Behavior in Carollia (Chiroptera, Phyllostomidae) by Chromosome Painting. Cytogenet. Genome Res. 2009, 124, 37–43. [Google Scholar] [CrossRef]
- Milioto, V.; Arizza, V.; Vizzini, A.; Perelman, P.L.; Roelke-Parker, M.E.; Dumas, F. Comparative Genomic Hybridization (CGH) in New World Monkeys (Primates) Reveals the Distribution of Repetitive Sequences in Cebinae and Callitrichinae. Biology 2024, 14, 22. [Google Scholar] [CrossRef]
- Palacios-Gimenez, O.M.; Marti, D.A.; Cabral-de-Mello, D.C. Neo-Sex Chromosomes of Ronderosia bergi: Insight into the Evolution of Sex Chromosomes in Grasshoppers. Chromosoma 2015, 124, 353–365. [Google Scholar] [CrossRef]
- Palacios-Gimenez, O.M.; Dias, G.B.; De Lima, L.G.; Kuhn, G.C.E.S.; Ramos, É.; Martins, C.; Cabral-de-Mello, D.C. High-Throughput Analysis of the Satellitome Revealed Enormous Diversity of Satellite DNAs in the Neo-Y Chromosome of the Cricket Eneoptera surinamensis. Sci. Rep. 2017, 7, 6422. [Google Scholar] [CrossRef] [PubMed]
- Palacios-Gimenez, O.M.; Milani, D.; Lemos, B.; Castillo, E.R.; Martí, D.A.; Ramos, E.; Martins, C.; Cabral-de-Mello, D.C. Uncovering the Evolutionary History of Neo-XY Sex Chromosomes in the Grasshopper Ronderosia bergii (Orthoptera, Melanoplinae) through Satellite DNA Analysis. BMC Evol. Biol. 2018, 18, 2. [Google Scholar] [CrossRef] [PubMed]
- Ferretti, A.B.S.M.; Milani, D.; Palacios-Gimenez, O.M.; Ruiz-Ruano, F.J.; Cabral-de-Mello, D.C. High Dynamism for Neo-Sex Chromosomes: Satellite DNAs Reveal Complex Evolution in a Grasshopper. Heredity 2020, 125, 124–137. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).