A De Novo DNM1L Mutation in Twins with Variable Symptoms, Including Paraparesis and Optic Neuropathy
Abstract
1. Introduction
2. Materials and Methods
2.1. Genetic Studies
2.2. Cell Culture
2.3. RNAseq Analysis
2.4. Immunoblot
2.5. Fluorescence Microscopy
3. Results
3.1. Clinical Features
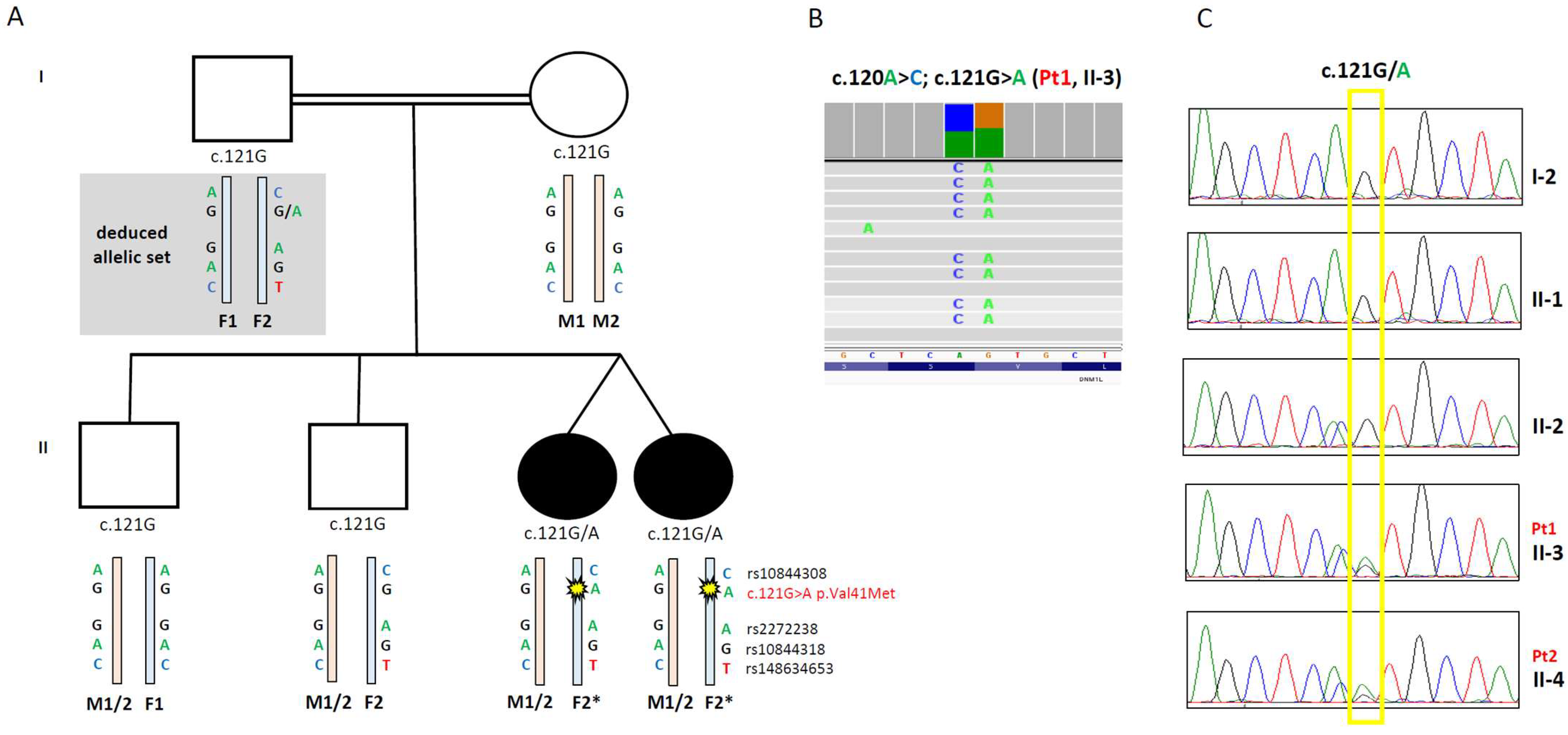
3.2. Genetic Analysis
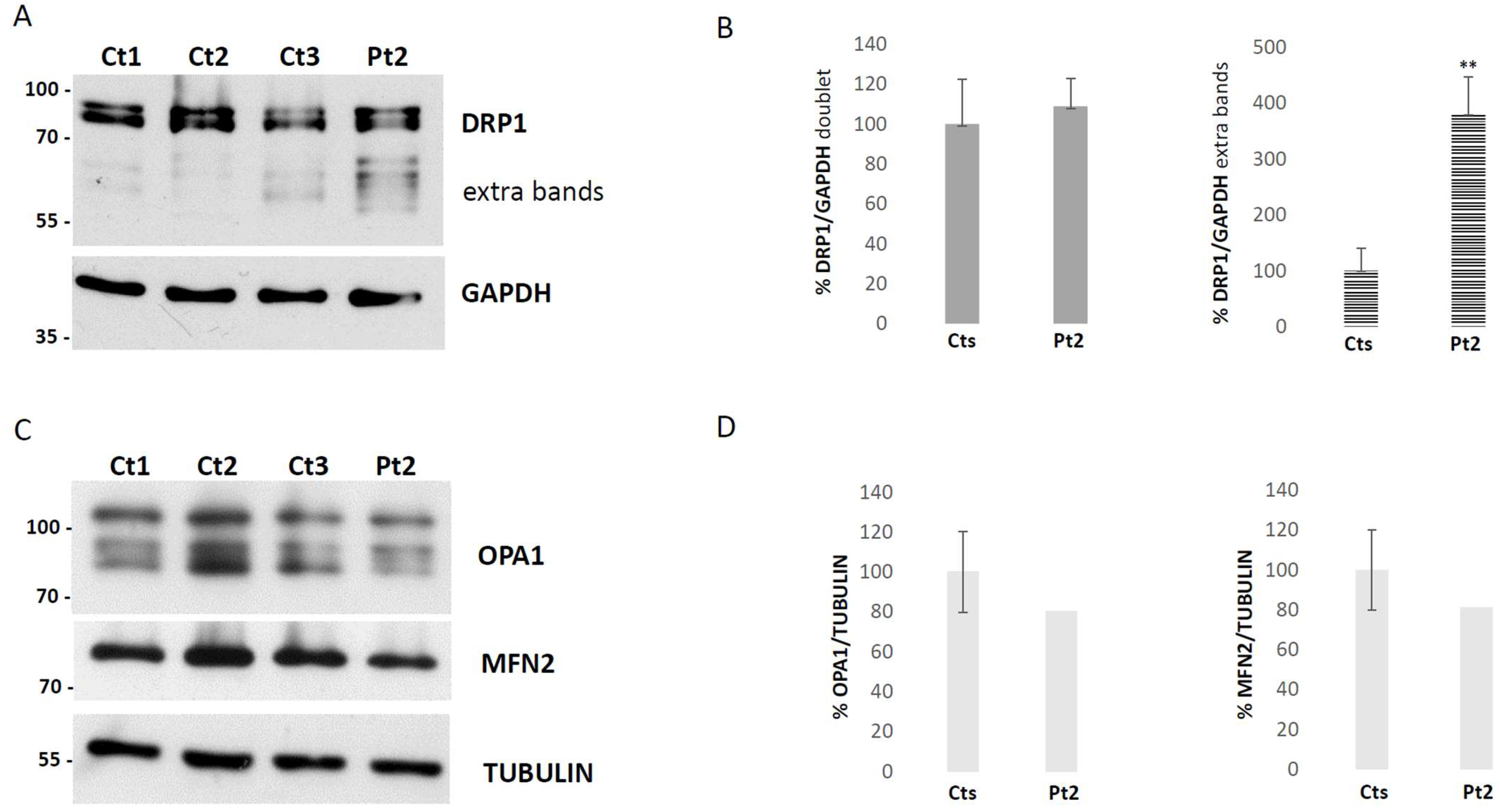
3.3. Immunostaining and Imaging in Fibroblasts
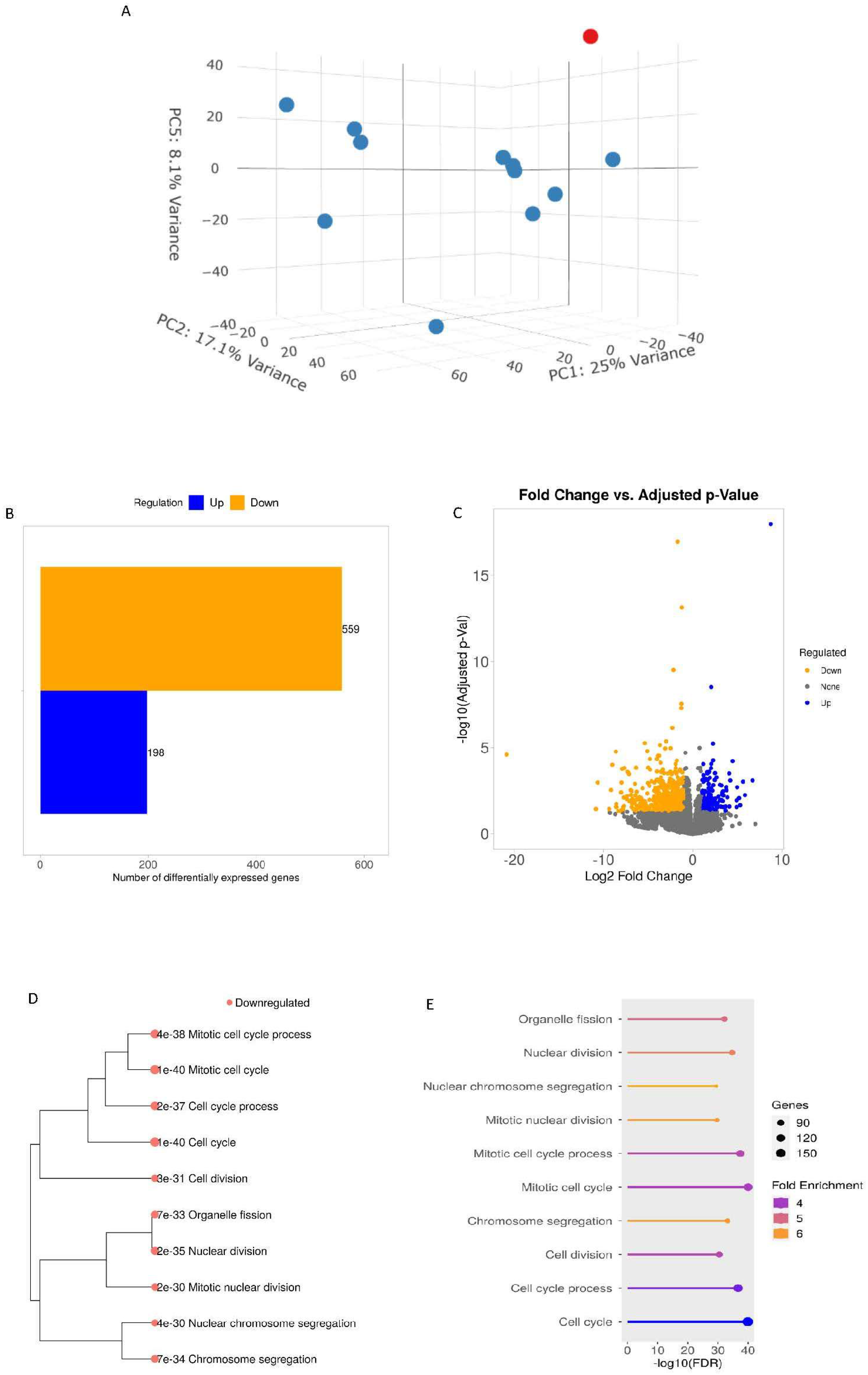
3.4. Transcriptomics Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wai, T.; Langer, T. Mitochondrial Dynamics and Metabolic Regulation. Trends Endocrinol. Metab. 2016, 27, 105–117. [Google Scholar] [CrossRef]
- Chen, W.; Zhao, H.; Li, Y. Mitochondrial Dynamics in Health and Disease: Mechanisms and Potential Targets. Sig. Transduct. Target. Ther. 2023, 8, 333. [Google Scholar] [CrossRef]
- Le, H.T.; Yu, J.; Ahn, H.S.; Kim, M.-J.; Chae, I.G.; Cho, H.-N.; Kim, J.; Park, H.-K.; Kwon, H.N.; Chae, H.-J.; et al. EIF2α Phosphorylation-ATF4 Axis-Mediated Transcriptional Reprogramming Mitigates Mitochondrial Impairment during ER Stress. Mol. Cells 2025, 48, 100176. [Google Scholar] [CrossRef]
- Vásquez-Trincado, C.; García-Carvajal, I.; Pennanen, C.; Parra, V.; Hill, J.A.; Rothermel, B.A.; Lavandero, S. Mitochondrial Dynamics, Mitophagy and Cardiovascular Disease. J. Physiol. 2016, 594, 509–525. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Alford, J.; Qiu, H. Structural and Functional Remodeling of Mitochondria in Cardiac Diseases. Int. J. Mol. Sci. 2021, 22, 4167. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.-A.; Wei, Y.; Sowers, J.R. Role of Mitochondrial Dysfunction in Insulin Resistance. Circ. Res. 2008, 102, 401–414. [Google Scholar] [CrossRef] [PubMed]
- Arun, S.; Liu, L.; Donmez, G. Mitochondrial Biology and Neurological Diseases. Curr. Neuropharmacol. 2016, 14, 143–154. [Google Scholar] [CrossRef]
- Park, S.; Won, J.-H.; Hwang, I.; Hong, S.; Lee, H.K.; Yu, J.-W. Defective Mitochondrial Fission Augments NLRP3 Inflammasome Activation. Sci. Rep. 2015, 5, 15489. [Google Scholar] [CrossRef]
- Moindjie, H.; Rodrigues-Ferreira, S.; Nahmias, C. Mitochondrial Metabolism in Carcinogenesis and Cancer Therapy. Cancers 2021, 13, 3311. [Google Scholar] [CrossRef]
- Tilokani, L.; Nagashima, S.; Paupe, V.; Prudent, J. Mitochondrial Dynamics: Overview of Molecular Mechanisms. Essays Biochem. 2018, 62, 341–360. [Google Scholar] [CrossRef]
- Zhang, Z.; Bie, X.; Chen, Z.; Liu, J.; Xie, Z.; Li, X.; Xiao, M.; Zhang, Q.; Zhang, Y.; Yang, Y.; et al. A Novel Variant of DNM1L Expanding the Clinical Phenotypic Spectrum: A Case Report and Literature Review. BMC Pediatr. 2024, 24, 104. [Google Scholar] [CrossRef]
- Kamerkar, S.C.; Kraus, F.; Sharpe, A.J.; Pucadyil, T.J.; Ryan, M.T. Dynamin-Related Protein 1 Has Membrane Constricting and Severing Abilities Sufficient for Mitochondrial and Peroxisomal Fission. Nat. Commun. 2018, 9, 5239. [Google Scholar] [CrossRef]
- Zhu, P.-P.; Patterson, A.; Stadler, J.; Seeburg, D.P.; Sheng, M.; Blackstone, C. Intra-and Intermolecular Domain Interactions of the C-Terminal GTPase Effector Domain of the Multimeric Dynamin-like GTPase Drp1*. J. Biol. Chem. 2004, 279, 35967–35974. [Google Scholar] [CrossRef]
- Smirnova, E.; Griparic, L.; Shurland, D.L.; van der Bliek, A.M. Dynamin-Related Protein Drp1 Is Required for Mitochondrial Division in Mammalian Cells. Mol. Biol. Cell 2001, 12, 2245–2256. [Google Scholar] [CrossRef] [PubMed]
- Gerber, S.; Charif, M.; Chevrollier, A.; Chaumette, T.; Angebault, C.; Kane, M.S.; Paris, A.; Alban, J.; Quiles, M.; Delettre, C.; et al. Mutations in DNM1L, as in OPA1, Result in Dominant Optic Atrophy despite Opposite Effects on Mitochondrial Fusion and Fission. Brain 2017, 140, 2586–2596. [Google Scholar] [CrossRef]
- Waterham, H.R.; Koster, J.; van Roermund, C.W.T.; Mooyer, P.A.W.; Wanders, R.J.A.; Leonard, J.V. A Lethal Defect of Mitochondrial and Peroxisomal Fission. N. Engl. J. Med. 2007, 356, 1736–1741. [Google Scholar] [CrossRef] [PubMed]
- Longo, F.; Benedetti, S.; Zambon, A.A.; Sora, M.G.N.; Di Resta, C.; De Ritis, D.; Quattrini, A.; Maltecca, F.; Ferrari, M.; Previtali, S.C. Impaired Turnover of Hyperfused Mitochondria in Severe Axonal Neuropathy Due to a Novel DRP1 Mutation. Hum. Mol. Genet. 2020, 29, 177–188. [Google Scholar] [CrossRef]
- Verrigni, D.; Di Nottia, M.; Ardissone, A.; Baruffini, E.; Nasca, A.; Legati, A.; Bellacchio, E.; Fagiolari, G.; Martinelli, D.; Fusco, L.; et al. Clinical-Genetic Features and Peculiar Muscle Histopathology in Infantile DNM1L-Related Mitochondrial Epileptic Encephalopathy. Hum. Mutat. 2019, 40, 601–618. [Google Scholar] [CrossRef]
- Berti, B.; Verrigni, D.; Nasca, A.; Di Nottia, M.; Leone, D.; Torraco, A.; Rizza, T.; Bellacchio, E.; Legati, A.; Palermo, C.; et al. De Novo DNM1L Mutation in a Patient with Encephalopathy, Cardiomyopathy and Fatal Non-Epileptic Paroxysmal Refractory Vomiting. Int. J. Mol. Sci. 2024, 25, 7782. [Google Scholar] [CrossRef] [PubMed]
- Nasca, A.; Legati, A.; Baruffini, E.; Nolli, C.; Moroni, I.; Ardissone, A.; Goffrini, P.; Ghezzi, D. Biallelic Mutations in DNM1L Are Associated with a Slowly Progressive Infantile Encephalopathy. Hum. Mutat. 2016, 37, 898–903. [Google Scholar] [CrossRef]
- Magistrati, M.; Zupin, L.; Lamantea, E.; Baruffini, E.; Ghezzi, D.; Legati, A.; Celsi, F.; Murru, F.M.; Capaci, V.; Pinamonti, M.; et al. De Novo DNM1L Pathogenic Variant Associated with Lethal Encephalocardiomyopathy—Case Report and Literature Review. Int. J. Mol. Sci. 2025, 26, 846. [Google Scholar] [CrossRef] [PubMed]
- Schmid, S.J.; Wagner, M.; Goetz, C.; Makowski, C.; Freisinger, P.; Berweck, S.; Mall, V.; Burdach, S.; Juenger, H. A De Novo Dominant Negative Mutation in DNM1L Causes Sudden Onset Status Epilepticus with Subsequent Epileptic Encephalopathy. Neuropediatrics 2019, 50, 197–201. [Google Scholar] [CrossRef] [PubMed]
- Minghetti, S.; Giorda, R.; Mastrangelo, M.; Tassi, L.; Zanotta, N.; Galbiati, S.; Bassi, M.T.; Zucca, C. Epilepsia Partialis Continua Associated with the p.Arg403Cys Variant of the DNM1L Gene: An Unusual Clinical Progression with Two Episodes of Super-Refractory Status Epilepticus with a 13-Year Remission Interval. Epileptic Disord. 2022, 24, 176–182. [Google Scholar] [CrossRef] [PubMed]
- Vanstone, J.R.; Smith, A.M.; McBride, S.; Naas, T.; Holcik, M.; Antoun, G.; Harper, M.-E.; Michaud, J.; Sell, E.; Chakraborty, P.; et al. DNM1L-Related Mitochondrial Fission Defect Presenting as Refractory Epilepsy. Eur. J. Hum. Genet. 2016, 24, 1084–1088. [Google Scholar] [CrossRef]
- Patro, R.; Duggal, G.; Love, M.I.; Irizarry, R.A.; Kingsford, C. Salmon Provides Fast and Bias-Aware Quantification of Transcript Expression. Nat. Methods 2017, 14, 417–419. [Google Scholar] [CrossRef]
- Ge, S.X.; Son, E.W.; Yao, R. IDEP: An Integrated Web Application for Differential Expression and Pathway Analysis of RNA-Seq Data. BMC Bioinform. 2018, 19, 534. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef]
- The Gene Ontology Consortium. The Gene Ontology Resource: 20 Years and Still GOing Strong. Nucleic Acids Res. 2019, 47, D330–D338. [Google Scholar] [CrossRef]
- Ge, S.X.; Jung, D.; Yao, R. ShinyGO: A Graphical Gene-Set Enrichment Tool for Animals and Plants. Bioinformatics 2020, 36, 2628–2629. [Google Scholar] [CrossRef]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An Open-Source Platform for Biological-Image Analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef]
- Liu, J.; Huang, J.; Zhao, Y.; Pan, H.; Wang, Y.; Liu, Z.; Xu, Q.; Sun, Q.; Tan, J.; Yan, X.; et al. Evaluating the Association between DNM1L Variants and Parkinson’s Disease in the Chinese Population. Front. Neurol. 2023, 14, 1133449. [Google Scholar] [CrossRef]
- Thorvaldsdóttir, H.; Robinson, J.T.; Mesirov, J.P. Integrative Genomics Viewer (IGV): High-Performance Genomics Data Visualization and Exploration. Brief. Bioinform. 2013, 14, 178–192. [Google Scholar] [CrossRef]
- Yoon, G.; Malam, Z.; Paton, T.; Marshall, C.R.; Hyatt, E.; Ivakine, Z.; Scherer, S.W.; Lee, K.-S.; Hawkins, C.; Cohn, R.D.; et al. Lethal Disorder of Mitochondrial Fission Caused by Mutations in DNM1L. J. Pediatr. 2016, 171, 313–316.e1-2. [Google Scholar] [CrossRef]
- Hogarth, K.A.; Costford, S.R.; Yoon, G.; Sondheimer, N.; Maynes, J.T. DNM1L Variant Alters Baseline Mitochondrial Function and Response to Stress in a Patient with Severe Neurological Dysfunction. Biochem. Genet. 2018, 56, 56–77. [Google Scholar] [CrossRef]
- Assia Batzir, N.; Bhagwat, P.K.; Eble, T.N.; Liu, P.; Eng, C.M.; Elsea, S.H.; Robak, L.A.; Scaglia, F.; Goldman, A.M.; Dhar, S.U.; et al. De Novo Missense Variant in the GTPase Effector Domain (GED) of DNM1L Leads to Static Encephalopathy and Seizures. Cold Spring Harb. Mol. Case Stud. 2019, 5, a003673. [Google Scholar] [CrossRef]
- Nolden, K.A.; Egner, J.M.; Collier, J.J.; Russell, O.M.; Alston, C.L.; Harwig, M.C.; Widlansky, M.E.; Sasorith, S.; Barbosa, I.A.; Douglas, A.G.; et al. Novel DNM1L Variants Impair Mitochondrial Dynamics through Divergent Mechanisms. Life Sci. Alliance 2022, 5, e202101284. [Google Scholar] [CrossRef]
- Westermann, B.; Neupert, W. Mitochondria-Targeted Green Fluorescent Proteins: Convenient Tools for the Study of Organelle Biogenesis in Saccharomyces Cerevisiae. Yeast 2000, 16, 1421–1427. [Google Scholar] [CrossRef]
- Bleazard, W.; McCaffery, J.M.; King, E.J.; Bale, S.; Mozdy, A.; Tieu, Q.; Nunnari, J.; Shaw, J.M. The Dynamin-Related GTPase Dnm1 Regulates Mitochondrial Fission in Yeast. Nat. Cell Biol. 1999, 1, 298–304. [Google Scholar] [CrossRef]
- Piccoli, C.; Scrima, R.; D’Aprile, A.; Chetta, M.; Cela, O.; Pacelli, C.; Ripoli, M.; D’Andrea, G.; Margaglione, M.; Bukvic, N.; et al. Pathogenic DNM1L Variant (1085G > A) Linked to Infantile Progressive Neurological Disorder: Evidence of Maternal Transmission by Germline Mosaicism and Influence of a Contemporary in Cis Variant (1535T > C). Genes 2021, 12, 1295. [Google Scholar] [CrossRef]
- Wang, A.S.; Lemire, G.; VanNoy, G.E.; Austin-Tse, C.; O’Donnell-Luria, A.; Kilbane, C. DNM1L Variant Presenting as Adolescent-Onset Sensory Neuronopathy, Spasticity, Dystonia, and Ataxia. J. Pediatr. Neurol. 2023, 21, 475–478. [Google Scholar] [CrossRef]
- Ryan, C.S.; Fine, A.L.; Cohen, A.L.; Schiltz, B.M.; Renaud, D.L.; Wirrell, E.C.; Patterson, M.C.; Boczek, N.J.; Liu, R.; Babovic-Vuksanovic, D.; et al. De Novo DNM1L Variant in a Teenager with Progressive Paroxysmal Dystonia and Lethal Super-Refractory Myoclonic Status Epilepticus. J. Child. Neurol. 2018, 33, 651–658. [Google Scholar] [CrossRef]
- Zemet, R.; Van den Veyver, I.B.; Stankiewicz, P. Parental Mosaicism for Apparent de Novo Genetic Variants: Scope, Detection, and Counseling Challenges. Prenat. Diagn. 2022, 42, 811–821. [Google Scholar] [CrossRef] [PubMed]
- Mohiuddin, M.; Kooy, R.F.; Pearson, C.E. De Novo Mutations, Genetic Mosaicism and Human Disease. Front. Genet. 2022, 13, 983668. [Google Scholar] [CrossRef] [PubMed]
- Whitley, B.N.; Lam, C.; Cui, H.; Haude, K.; Bai, R.; Escobar, L.; Hamilton, A.; Brady, L.; Tarnopolsky, M.A.; Dengle, L.; et al. Aberrant Drp1-Mediated Mitochondrial Division Presents in Humans with Variable Outcomes. Hum. Mol. Genet. 2018, 27, 3710–3719. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nasca, A.; Catania, A.; Legati, A.; Izzo, R.; D’onofrio, C.; Ciavattini, T.; Lamantea, E.; Lamperti, C.; Ghezzi, D. A De Novo DNM1L Mutation in Twins with Variable Symptoms, Including Paraparesis and Optic Neuropathy. Biomolecules 2025, 15, 1230. https://doi.org/10.3390/biom15091230
Nasca A, Catania A, Legati A, Izzo R, D’onofrio C, Ciavattini T, Lamantea E, Lamperti C, Ghezzi D. A De Novo DNM1L Mutation in Twins with Variable Symptoms, Including Paraparesis and Optic Neuropathy. Biomolecules. 2025; 15(9):1230. https://doi.org/10.3390/biom15091230
Chicago/Turabian StyleNasca, Alessia, Alessia Catania, Andrea Legati, Rossella Izzo, Carola D’onofrio, Teresa Ciavattini, Eleonora Lamantea, Costanza Lamperti, and Daniele Ghezzi. 2025. "A De Novo DNM1L Mutation in Twins with Variable Symptoms, Including Paraparesis and Optic Neuropathy" Biomolecules 15, no. 9: 1230. https://doi.org/10.3390/biom15091230
APA StyleNasca, A., Catania, A., Legati, A., Izzo, R., D’onofrio, C., Ciavattini, T., Lamantea, E., Lamperti, C., & Ghezzi, D. (2025). A De Novo DNM1L Mutation in Twins with Variable Symptoms, Including Paraparesis and Optic Neuropathy. Biomolecules, 15(9), 1230. https://doi.org/10.3390/biom15091230







