Multi-Omics Feature Selection to Identify Biomarkers for Hepatocellular Carcinoma
Abstract
1. Introduction
2. Materials and Methods
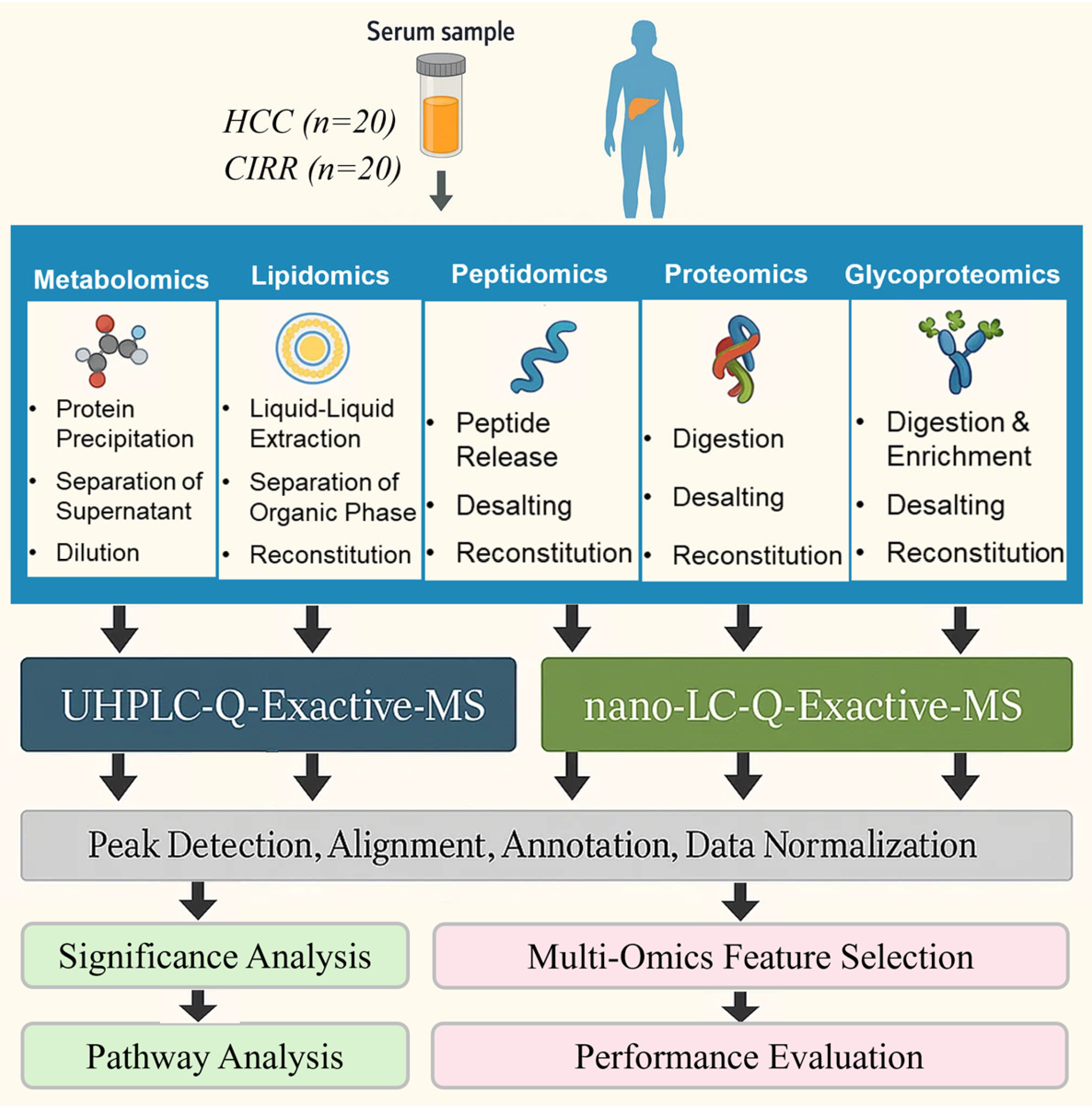
2.1. Untargeted Multi-Omics Studies
2.2. Targeted Multi-Omics Studies
3. Results
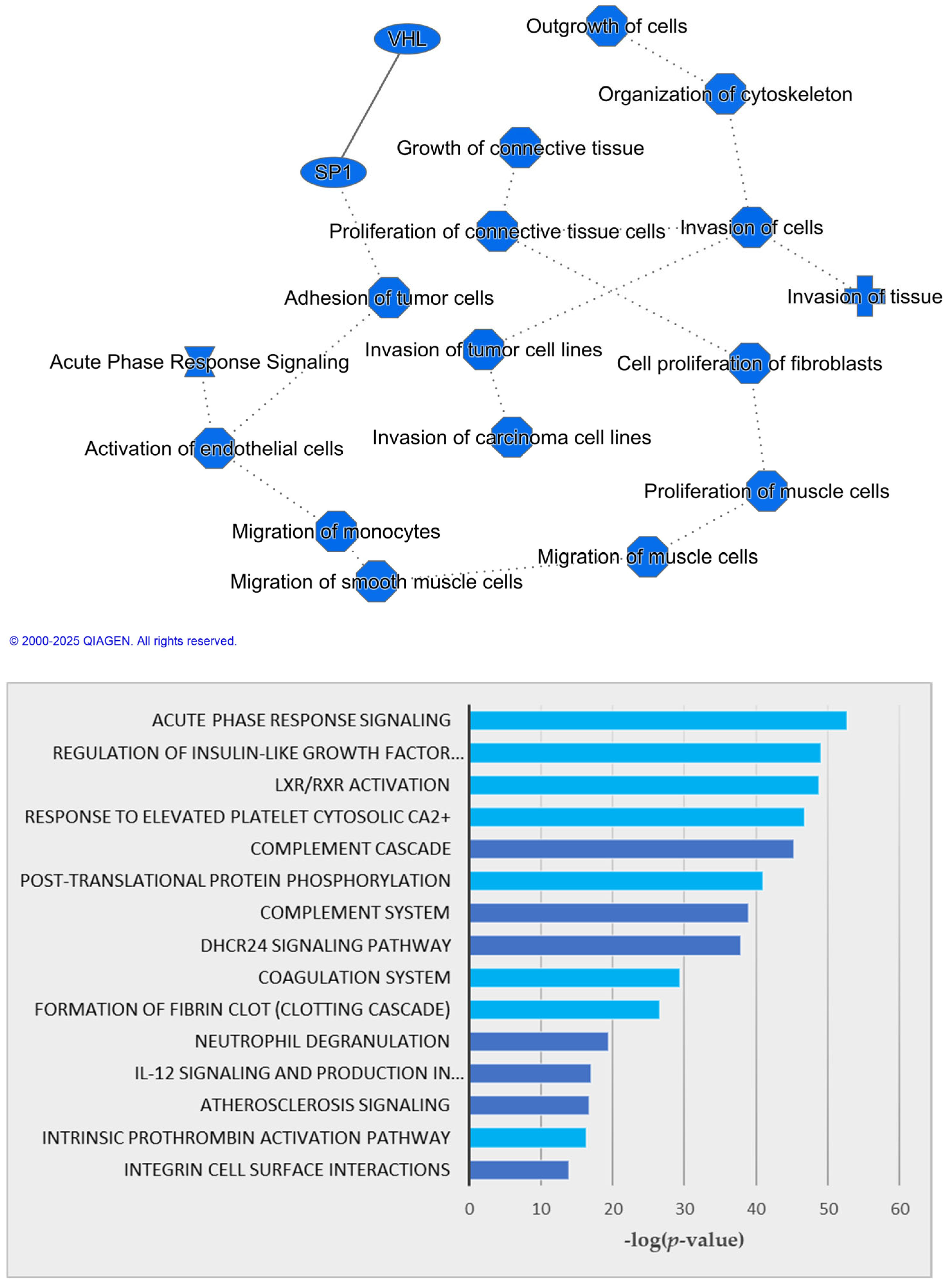
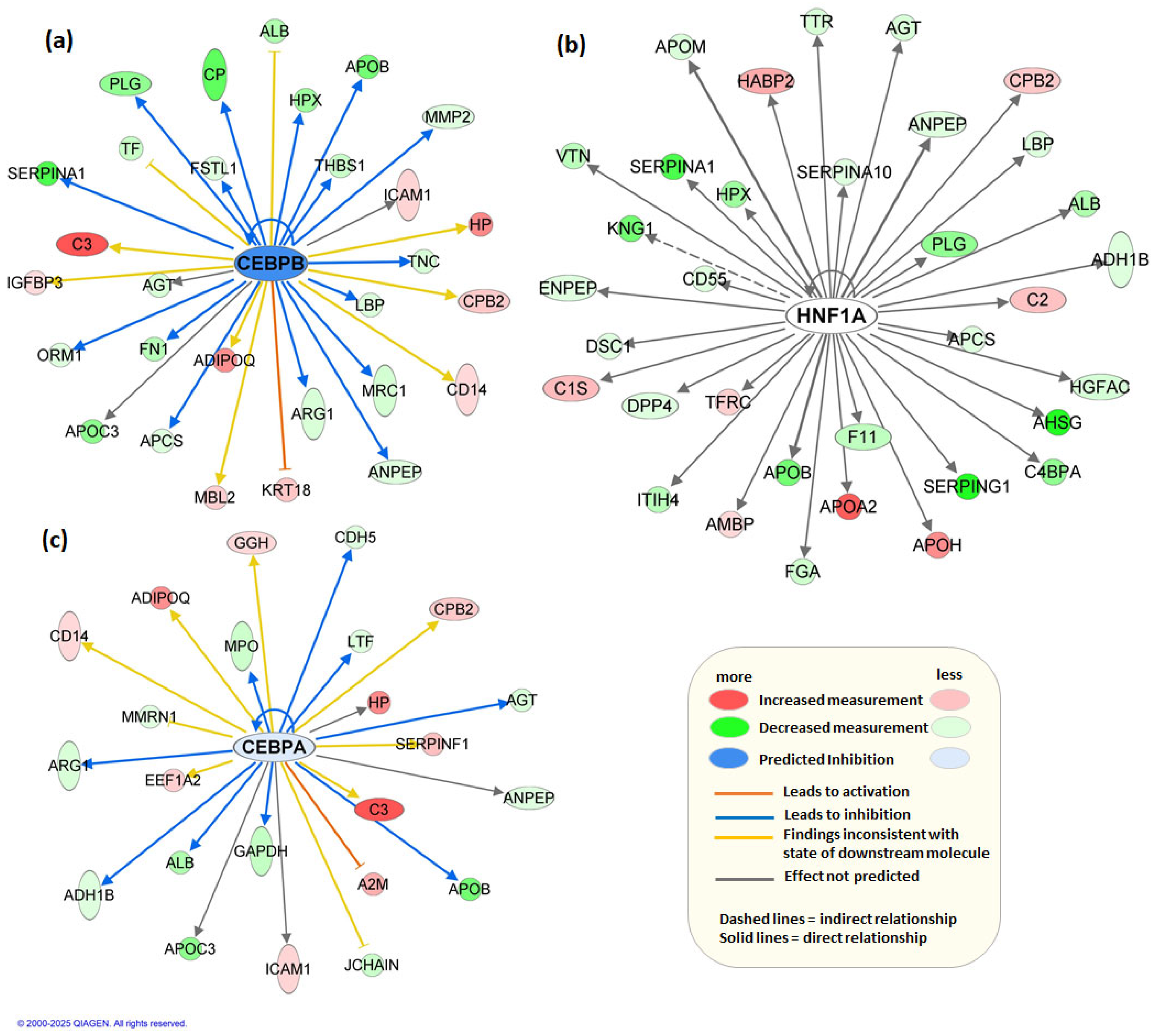
3.1. Untargeted Multi-Omics Studies
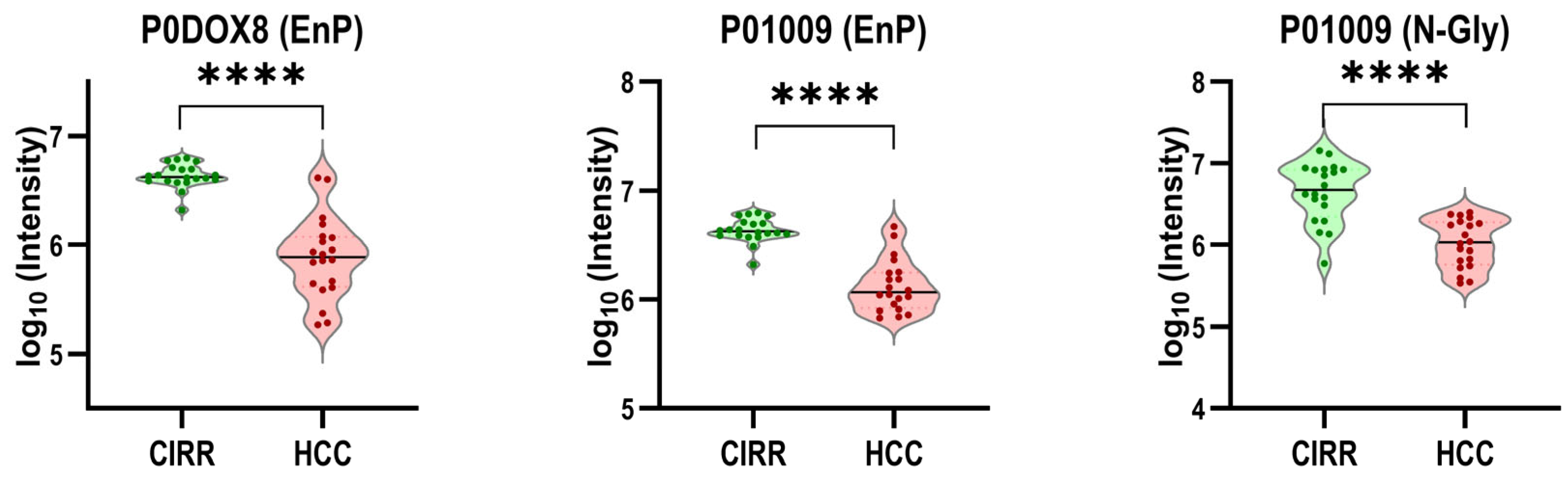
3.2. Targeted Multi-Omics Studies
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kim, D.Y. Changing etiology and epidemiology of hepatocellular carcinoma: Asia and worldwide. J. Liver Cancer 2024, 24, 622–670. [Google Scholar] [CrossRef]
- McCurdy, H.; Nobbe, A.; Scott, D.; Patton, H.; Morgan, T.R.; Bajaj, J.S.; Yakovchenko, V.; Merante, M.; Gibson, S.; Lamorte, C.; et al. Organizational and Implementation Factors Associated with Cirrhosis Care in the Veterans Health Administration. Dig. Dis. Sci. 2024, 69, 2008–2017. [Google Scholar] [CrossRef] [PubMed]
- Hammad, R.; Eldosoky, M.A.; Elmadbouly, A.A.; Aglan, R.B.; AbdelHamid, S.G.; Zaky, S.; Ali, E.; El Hakam, F.E.-Z.A.; Mosaad, A.M.; Abdelmageed, N.A.; et al. Monocytes subsets altered distribution and dysregulated plasma hsa-miR-21-5p and hsa-miR-155-5p in HCV-linked liver cirrhosis progression to hepatocellular carcinoma. J. Cancer Res. Clin. Oncol. 2023, 149, 15349–15364. [Google Scholar] [CrossRef]
- Youssef, S.S.; Hamdy, N.M. SOCS1 and pattern recognition receptors: TLR9 and RIG-I; novel haplotype associations in Egyptian fibrotic/cirrhotic patients with HCV genotype 4. Arch. Virol. 2017, 162, 3347–3354. [Google Scholar] [CrossRef] [PubMed]
- Llovet, J.M.; Kelley, R.K.; Villanueva, A.; Singal, A.G.; Pikarsky, E.; Roayaie, S.; Lencioni, R.; Koike, K.; Zucman-Rossi, J.; Finn, R.S. Hepatocellular carcinoma. Nat. Rev. Dis. Primers 2021, 7, 6. [Google Scholar] [CrossRef]
- Abaza, T.; El-Aziz, M.K.A.; Daniel, K.A.; Karousi, P.; Papatsirou, M.; Fahmy, S.A.; Hamdy, N.M.; Kontos, C.K.; Youness, R.A. Emerging Role of Circular RNAs in Hepatocellular Carcinoma Immunotherapy. Int. J. Mol. Sci. 2023, 24, 16484. [Google Scholar] [CrossRef]
- Reguram, R.; Ghonge, A.; Tse, J.; Dhanasekaran, R. Practical approach to diagnose and manage benign liver masses. Hepatol. Commun. 2024, 8, e0560. [Google Scholar] [CrossRef]
- Kim, E.; Viatour, P. Hepatocellular carcinoma: Old friends and new tricks. Exp. Mol. Med. 2020, 52, 1898–1907. [Google Scholar] [CrossRef]
- Hanif, H.; Ali, M.J.; Susheela, A.T.; Khan, I.W.; Luna-Cuadros, M.A.; Khan, M.M.; Lau, D.T.-Y. Update on the applications and limitations of alpha-fetoprotein for hepatocellular carcinoma. World J. Gastroenterol. 2022, 28, 216–229. [Google Scholar] [CrossRef] [PubMed]
- Parikh, N.D.; Mehta, A.S.; Singal, A.G.; Block, T.; Marrero, J.A.; Lok, A.S. Biomarkers for the Early Detection of Hepatocellular Carcinoma. Cancer Epidemiol. Biomark. Prev. 2020, 29, 2495–2503. [Google Scholar] [CrossRef]
- Parikh, N.D.; Tayob, N.; Singal, A.G. Blood-based biomarkers for hepatocellular carcinoma screening: Approaching the end of the ultrasound era? J. Hepatol. 2023, 78, 207–216. [Google Scholar] [CrossRef] [PubMed]
- Misra, B.B.; Langefeld, C.D.; Olivier, M.; Cox, L.A. Integrated Omics: Tools, Advances, and Future Approaches. J. Mol. Endocrinol. 2019, 62, R21–R45. [Google Scholar] [CrossRef]
- Hasin, Y.; Seldin, M.; Lusis, A. Multi-omics approaches to disease. Genome Biol. 2017, 18, 83. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, I.; Verma, S.; Kumar, S.; Jere, A.; Anamika, K. Multi-Omics Data Integration, Interpretation, and Its Application. Bioinform. Biol. Insights 2020, 14, 1177932219899051. [Google Scholar] [CrossRef] [PubMed]
- Ewald, J.D.; Zhou, G.; Lu, Y.; Kolic, J.; Ellis, C.; Johnson, J.D.; Macdonald, P.E.; Xia, J. Web-based multi-omics integration using the Analyst software suite. Nat. Protoc. 2024, 19, 1467–1497. [Google Scholar] [CrossRef]
- Uppal, K.; Ma, C.; Go, Y.-M.; Jones, D.P.; Wren, J. xMWAS: A data-driven integration and differential network analysis tool. Bioinformatics 2018, 34, 701–702. [Google Scholar] [CrossRef]
- Kuo, T.-C.; Tian, T.-F.; Tseng, Y.J. 3Omics: A web-based systems biology tool for analysis, integration and visualization of human transcriptomic, proteomic and metabolomic data. BMC Syst. Biol. 2013, 7, 64. [Google Scholar] [CrossRef]
- Zhou, G.; Pang, Z.; Lu, Y.; Ewald, J.; Xia, J. OmicsNet 2.0: A web-based platform for multi-omics integration and network visual analytics. Nucleic Acids Res. 2022, 50, W527–W533. [Google Scholar] [CrossRef]
- Goecks, J.; Nekrutenko, A.; Taylor, J. The Galaxy Team Galaxy: A comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 2010, 11, R86. [Google Scholar] [CrossRef]
- Sheynkman, G.M.; Johnson, J.E.; Jagtap, P.D.; Shortreed, M.R.; Onsongo, G.; Frey, B.L.; Griffin, T.J.; Smith, L.M. Using Galaxy-P to leverage RNA-Seq for the discovery of novel protein variations. BMC Genom. 2014, 15, 703. [Google Scholar] [CrossRef]
- Davidson, R.L.; Weber, R.J.M.; Liu, H.; Sharma-Oates, A.; Viant, M.R. Galaxy-M: A Galaxy workflow for processing and analyzing direct infusion and liquid chromatography mass spectrometry-based metabolomics data. GigaScience 2016, 5, 10. [Google Scholar] [CrossRef] [PubMed]
- Berthold, M.; Cebron, N.; Dill, F.; Gabriel, T.; Kötter, T.; Meinl, T.; Ohl, P.; Thiel, K.; Wiswedel, B. KNIME-The konstanz information miner: Version 2.0 and beyond. SIGKDD Explor. 2009, 11, 26–31. [Google Scholar] [CrossRef]
- Rohart, F.; Gautier, B.; Singh, A.; Lê Cao, K.-A. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS Comput. Biol. 2017, 13, e1005752. [Google Scholar] [CrossRef]
- Chalise, P.; Kwon, D.; Fridley, B.L.; Mo, Q. Statistical methods for integrative clustering of multi-omics data. In Statistical Genomics. Methods in Molecular Biology; Humana: New York, NY, USA, 2023; Volume 2629. [Google Scholar] [CrossRef]
- Argelaguet, R.; Velten, B.; Arnol, D.; Dietrich, S.; Zenz, T.; Marioni, J.C.; Buettner, F.; Huber, W.; Stegle, O. Multi-Omics Factor Analysis—A framework for unsupervised integration of multi-omics data sets. Mol. Syst. Biol. 2018, 14, e8124. [Google Scholar] [CrossRef]
- Luo, W.; Pant, G.; Bhavnasi, Y.K.; Blanchard, S.G., Jr.; Brouwer, C. Pathview Web: User friendly pathway visualization and data integration. Nucleic Acids Res. 2017, 45, W501–W508. [Google Scholar] [CrossRef] [PubMed]
- Dong, X.; Vegesna, K.; Brouwer, C.; Luo, W.; Mathelier, A. SBGNview: Towards data analysis, integration and visualization on all pathways. Bioinformatics 2022, 38, 1473–1476. [Google Scholar] [CrossRef]
- Luo, W.; Brouwer, C. Pathview: An R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics 2013, 29, 1830–1831. [Google Scholar] [CrossRef]
- Lock, E.F.; Hoadley, K.A.; Marron, J.S.; Nobel, A.B. Joint and individual variation explained (JIVE) for integrated analysis of multiple data types. Ann. Appl. Stat. 2013, 7, 523–542. [Google Scholar] [CrossRef]
- Devarajan, K.; Bryant, B. Nonnegative Matrix Factorization: An Analytical and Interpretive Tool in Computational Biology. PLoS Comput. Biol. 2008, 4, e1000029. [Google Scholar] [CrossRef]
- Chalise, P.; Fridley, B.L.; Peddada, S.D. Integrative clustering of multi-level ‘omic data based on non-negative matrix factorization algorithm. PLoS ONE 2017, 12, e0176278. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Ma, J.; Leng, L.; Han, M.; Li, M.; He, F.; Zhu, Y. MoGCN: A Multi-Omics Integration Method Based on Graph Convolutional Network for Cancer Subtype Analysis. Front. Genet. 2022, 13, 806842. [Google Scholar] [CrossRef]
- Kinney, W.K.; Egorshin, E.V.; Podratz, K.C. Wertheim hysterectomy in the geriatric population. Gynecol. Oncol. 1988, 31, 227–232. [Google Scholar] [CrossRef]
- Cho, H.J.; Shu, M.; Bekiranov, S.; Zang, C.; Zhang, A.; Wren, J. Interpretable meta-learning of multi-omics data for survival analysis and pathway enrichment. Bioinformatics 2023, 39, btad113. [Google Scholar] [CrossRef]
- Cai, Z.; Poulos, R.C.; Aref, A.; Robinson, P.J.; Reddel, R.R.; Zhong, Q. DeePathNet: A Transformer-Based Deep Learning Model Integrating Multiomic Data with Cancer Pathways. Cancer Res. Commun. 2024, 4, 3151–3164. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Tao, Y.; Cai, Z.; Bao, P.; Ma, H.; Li, K.; Li, M.; Zhu, Y.; Lu, Z.J.; Wren, J. Pathformer: A biological pathway informed transformer for disease diagnosis and prognosis using multi-omics data. Bioinformatics 2024, 40, btae316. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Mou, M.; Hu, W.; Lu, M.; Zhang, H.; Zhang, H.; Luo, Y.; Xu, H.; Tao, L.; Dai, H.; et al. MOINER: A Novel Multiomics Early Integration Framework for Biomedical Classification and Biomarker Discovery. J. Chem. Inf. Model. 2024, 64, 2720–2732. [Google Scholar] [CrossRef] [PubMed]
- Rashid, M.; Varghese, R.S.; Ding, Y.; Ressom, H.W. Biomarker Discovery for Hepatocellular Carcinoma in Patients with Liver Cirrhosis Using Untargeted Metabolomics and Lipidomics Studies. Metabolites 2023, 13, 1047. [Google Scholar] [CrossRef]
- Krämer, A.; Green, J.; Pollard, J., Jr.; Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 2014, 30, 523–530. [Google Scholar] [CrossRef]
- Lundberg, S. A unified approach to interpreting model predictions. arXiv 2017, arXiv:1705.07874. [Google Scholar] [CrossRef]
- Geron, A. Hands-On Machine Learning with Scikit-Learn, Keras, and TensorFlow: Concepts, Tools, and Techniques to Build Intelligent Systems (2nd ed.). 2022. Available online: http://bvbr.bib-bvb.de:8991/F?func=service&doc_library=BVB01&local_base=BVB01&doc_number=033747193&sequence=000001&line_number=0001&func_code=DB_RECORDS&service_type=MEDIA (accessed on 25 March 2025).
- Wang, L.; Wang, Y.; Chang, Q. Feature selection methods for big data bioinformatics: A survey from the search perspective. Methods 2016, 111, 21–31. [Google Scholar] [CrossRef]
- Hu, J.; Szymczak, S. A review on longitudinal data analysis with random forest. Briefings Bioinform. 2023, 24, bbad002. [Google Scholar] [CrossRef]
- Wang, T.; Shao, W.; Huang, Z.; Tang, H.; Zhang, J.; Ding, Z.; Huang, K. MOGONET integrates multi-omics data using graph convolutional networks allowing patient classification and biomarker identification. Nat. Commun. 2021, 12, 3445. [Google Scholar] [CrossRef]
- Ranjbar, M.R.N.; Luo, Y.; Di Poto, C.; Varghese, R.S.; Ferrarini, A.; Zhang, C.; Sarhan, N.I.; Soliman, H.; Tadesse, M.G.; Ziada, D.H.; et al. GC-MS Based Plasma Metabolomics for Identification of Candidate Biomarkers for Hepatocellular Carcinoma in Egyptian Cohort. PLoS ONE 2015, 10, e0127299. [Google Scholar] [CrossRef]
- Ranjbar, M.R.N.; Di Poto, C.; Wang, Y.; Ressom, H.W. SIMAT: GC-SIM-MS data analysis tool. BMC Bioinform. 2015, 16, 259. [Google Scholar] [CrossRef]
- Tsai, T.; Song, E.; Zhu, R.; Di Poto, C.; Wang, M.; Luo, Y.; Varghese, R.S.; Tadesse, M.G.; Ziada, D.H.; Desai, C.S.; et al. LC-MS/MS-based serum proteomics for identification of candidate biomarkers for hepatocellular carcinoma. Proteomics 2015, 15, 2369–2381. [Google Scholar] [CrossRef]
- Tsai, T.-H.; Wang, M.; Di Poto, C.; Hu, Y.; Zhou, S.; Zhao, Y.; Varghese, R.S.; Luo, Y.; Tadesse, M.G.; Ziada, D.H.; et al. LC–MS Profiling of N-Glycans Derived from Human Serum Samples for Biomarker Discovery in Hepatocellular Carcinoma. J. Proteome Res. 2014, 13, 4859–4868. [Google Scholar] [CrossRef]
- MacLean, B.; Tomazela, D.M.; Shulman, N.; Chambers, M.; Finney, G.L.; Frewen, B.; Kern, R.; Tabb, D.L.; Liebler, D.C.; MacCoss, M.J. Skyline: An open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 2010, 26, 966–968. [Google Scholar] [CrossRef]
- Wang, M.; Yu, G.; Ressom, H.W. Integrative Analysis of Proteomic, Glycomic, and Metabolomic Data for Biomarker Discovery. IEEE J. Biomed. Health Inform. 2016, 20, 1225–1231. [Google Scholar] [CrossRef] [PubMed]
- Barefoot, M.E.; Varghese, R.S.; Zhou, Y.; Di Poto, C.; Ferrarini, A.; Ressom, H.W. Multi-omic Pathway and Network Analysis to Identify Biomarkers for Hepatocellular Carcinoma. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; pp. 1350–1354. [Google Scholar] [CrossRef]
- Ricklin, D.; Hajishengallis, G.; Yang, K.; Lambris, J.D. Complement: A key system for immune surveillance and homeostasis. Nat. Immunol. 2010, 11, 785–797. [Google Scholar] [CrossRef] [PubMed]
- Greenbaum, L.E.; Cressman, D.E.; Haber, B.A.; Taub, R. Coexistence of C/EBP alpha, beta, growth-induced proteins and DNA synthesis in hepatocytes during liver regeneration. Implications for maintenance of the differentiated state during liver growth. J. Clin. Investig. 1995, 96, 1351–1365. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Jakobsen, J.S.; Waage, J.; Rapin, N.; Bisgaard, H.C.; Larsen, F.S.; Porse, B.T. Temporal mapping of CEBPA and CEBPB binding during liver regeneration reveals dynamic occupancy and specific regulatory codes for homeostatic and cell cycle gene batteries. Genome Res. 2013, 23, 592–603. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.; Cheung, S.T. STAT3: An Emerging Therapeutic Target for Hepatocellular Carcinoma. Cancers 2019, 11, 1646. [Google Scholar] [CrossRef] [PubMed]
- Sajid, M.S.; Ding, Y.; Varghese, R.S.; Kroemer, A.; Ressom, H.W. Unveiling Endogenous Serum Peptides as Potential Biomarkers for Hepatocellular Carcinoma in Patients with Liver Cirrhosis. J. Proteome Res. 2024, 23, 3974–3983. [Google Scholar] [CrossRef]
- Mouillot, T.; Rizk, M.; de Barros, J.P.; Gilloteau, A.; Busson, A.; Bernard-Chabert, B.; Thiefin, G.; Barraud, H.; Bronowicki, J.; Richou, C.; et al. Fatty acid composition of the erythrocyte membrane and risk of hepatocellular carcinoma in cirrhotic patients. Aliment. Pharmacol. Ther. 2020, 52, 1503–1515. [Google Scholar] [CrossRef]
- O’Connell, T.M. The Complex Role of Branched Chain Amino Acids in Diabetes and Cancer. Metabolites 2013, 3, 931–945. [Google Scholar] [CrossRef] [PubMed]


| HCC (n = 20) | CIRR (n= 20) | p-Value | ||
|---|---|---|---|---|
| Age | Mean (SD) | 59 (6) | 57 (6) | 0.487 |
| Sex | Male | 60% | 65% | 1 |
| Race | AA | 50% | 40% | 0.5231 |
| EA | 50% | 60% | ||
| HCV Serology | HCV Ab+ | 80% | 75% | 0.6948 |
| HBV Serology | anti HBC+ | 45% | 40% | 0.7431 |
| HBs Ag+ | 5% | 0 | 1 | |
| Smoking | Current | 25% | 25% | 1 |
| Former | 55% | 50% | ||
| Alcohol | Current | 25% | 20% | 0.6851 |
| Former | 55% | 60% | ||
| MELD * | Median (IQR) | 10.5 (5.2) | 13.5 (9.2) | 0.0475 |
| AFP | Median (IQR) | 29.1 (60.8) | 7 (35.1) | 0.113 |
| HCC Stage | Stage I | 30% | ||
| Stage II | 65% | |||
| Stage III | 5% |
| HCC (n = 20) | CIRR (n = 24) | p-Value | ||
|---|---|---|---|---|
| Age | Mean (SD) | 59.7 (6) | 57.8 (7) | 0.346 |
| Sex | Male | 80% | 79% | 1 |
| Race * | AA | 35% | 17% | 0.045 |
| EA | 45% | 70% | ||
| HCV Serology | HCV Ab+ | 70% | 42% | 0.15 |
| HBV Serology | anti HBC+ | 60% | 21% | 0.18 |
| HBs Ag+ | 10% | 0 | 0.386 | |
| Smoking | Current | 15% | 22% | 0.862 |
| Former | 50% | 48% | 1 | |
| Alcohol | Current | 20% | 13% | 0.84 |
| Former | 50% | 57% | 0.904 | |
| MELD * | Median (IQR) | 10 (4.3) | 14.5 (6.3) | 0.0004 |
| AFP | Median (IQR) | 29.1 (74.3) | 4.2 (7.3) | 0.0438 |
| HCC Stage | Stage I | 60% | ||
| Stage II | 40% |
| Omics Study | No. of Features Detected | No. of Features (p < 0.05) | No. of Features (FDR < 0.05) |
|---|---|---|---|
| Metabolomics (Met) | 7174 | 345 | - |
| Lipidomics (Lip) | 2252 | 397 | 117 |
| Endogenous Peptidomics (EnP) | 2355 | 438 | 70 |
| N-linked Glycoproteomics (N-Gly) | 750 | 164 | 49 |
| O-linked Glycoproteomics (O-Gly) | 244 | 74 | 67 |
| Proteomics (Prot) | 3530 | 1339 | 975 |
| SelectKBest | SVM-RFE | Transformer–RFE | RF | MOINER | MOGONET | |
|---|---|---|---|---|---|---|
| Multi-Omics Features | P80748 | P01009 | P01009 | P80748 | P01042 | P0DOX5 |
| P0DOX8 | P80748 | P02656 | P04275 | P01009 (N-Gly) | P0DOX8 | |
| P01009 (EnP) | Q53H89 | heptacosanoic acid | P01009 | heptacosanoic acid | P02656 | |
| P01009 (N-Gly) | P0DOX8 | PC(18:0/20:0) | P0DOX2 | P05154 | A0A075B6R2 | |
| P0DOX5 | Q8WZ75 | Q06033 | O75882 | P02768 | P01009 | |
| Accuracy | 100% | 97.5% | 97.5% | 100% | 87.5% | 97.5% |
| AUC | 1.00 | 1.00 | 1.00 | 1.00 | 0.938 | 0.988 |
| Omics Dataset | No. of Features | No. of Features (p < 0.05) |
|---|---|---|
| Metabolomics | 53 | 5 |
| Proteomics | 101 | 43 |
| Glycomics | 82 | 8 |
| SelectKBest | SVM-RFE | Transformer–RFE | RF | MOINER | MOGONET | |
|---|---|---|---|---|---|---|
| Multi-Omics Features | O75636 | P03952 | tyramine | P01023 | isoleucine | threitol |
| P03952 | P05156 | P01876 | leucine | P02652 | ethanolamine | |
| Q6EMK4 | O75636 | P02774 | O75636 | leucine | sorbose | |
| P22891 | 6-hydroxy caproic acid | P05156 | Q6EMK4 | 43211 | leucine | |
| P05156 | P01876 | 25000 | isoleucine | P22891 | creatinine | |
| Accuracy | 77.2% | 75.0% | 77.2% | 72.8% | 70.0% | 68.0% |
| AUC | 0.848 | 0.825 | 0.798 | 0.853 | 0.756 | 0.741 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Varghese, R.S.; Zhang, X.; Giridharan, S.; Sajid, M.S.; Rashid, M.M.; Kroemer, A.; Ressom, H.W. Multi-Omics Feature Selection to Identify Biomarkers for Hepatocellular Carcinoma. Metabolites 2025, 15, 575. https://doi.org/10.3390/metabo15090575
Varghese RS, Zhang X, Giridharan S, Sajid MS, Rashid MM, Kroemer A, Ressom HW. Multi-Omics Feature Selection to Identify Biomarkers for Hepatocellular Carcinoma. Metabolites. 2025; 15(9):575. https://doi.org/10.3390/metabo15090575
Chicago/Turabian StyleVarghese, Rency S., Xinran Zhang, Sarada Giridharan, Muhammad Salman Sajid, Md Mamunur Rashid, Alexander Kroemer, and Habtom W. Ressom. 2025. "Multi-Omics Feature Selection to Identify Biomarkers for Hepatocellular Carcinoma" Metabolites 15, no. 9: 575. https://doi.org/10.3390/metabo15090575
APA StyleVarghese, R. S., Zhang, X., Giridharan, S., Sajid, M. S., Rashid, M. M., Kroemer, A., & Ressom, H. W. (2025). Multi-Omics Feature Selection to Identify Biomarkers for Hepatocellular Carcinoma. Metabolites, 15(9), 575. https://doi.org/10.3390/metabo15090575







