Similarity Downselection: Finding the n Most Dissimilar Molecular Conformers for Reference-Free Metabolomics
Abstract
1. Introduction
2. Application: Molecular Conformer Sampling
3. Similarity Downselection Python Module
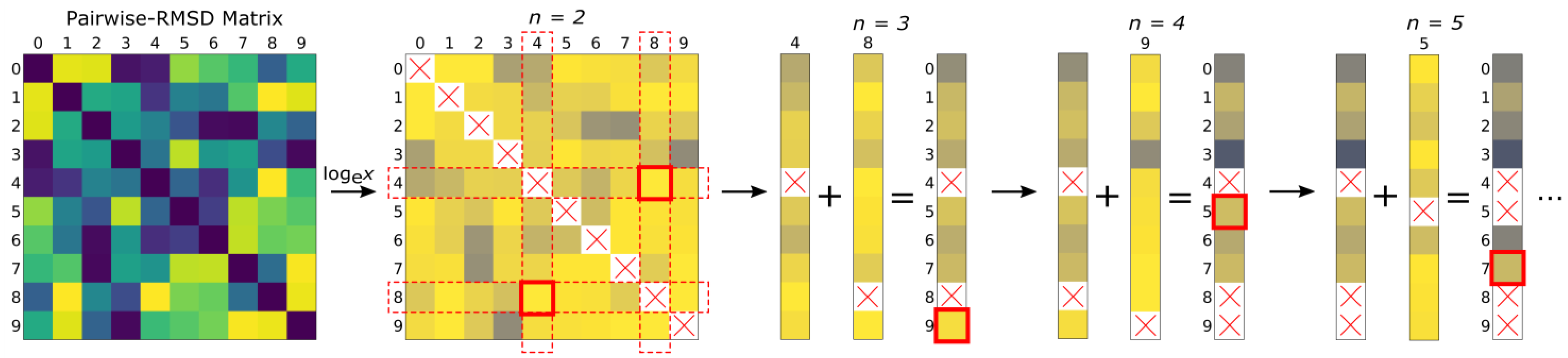
3.1. Algorithm Description
3.2. Problem and Algorithm Description Using Graph Theory (Nodes and Edges)
4. Benchmarking
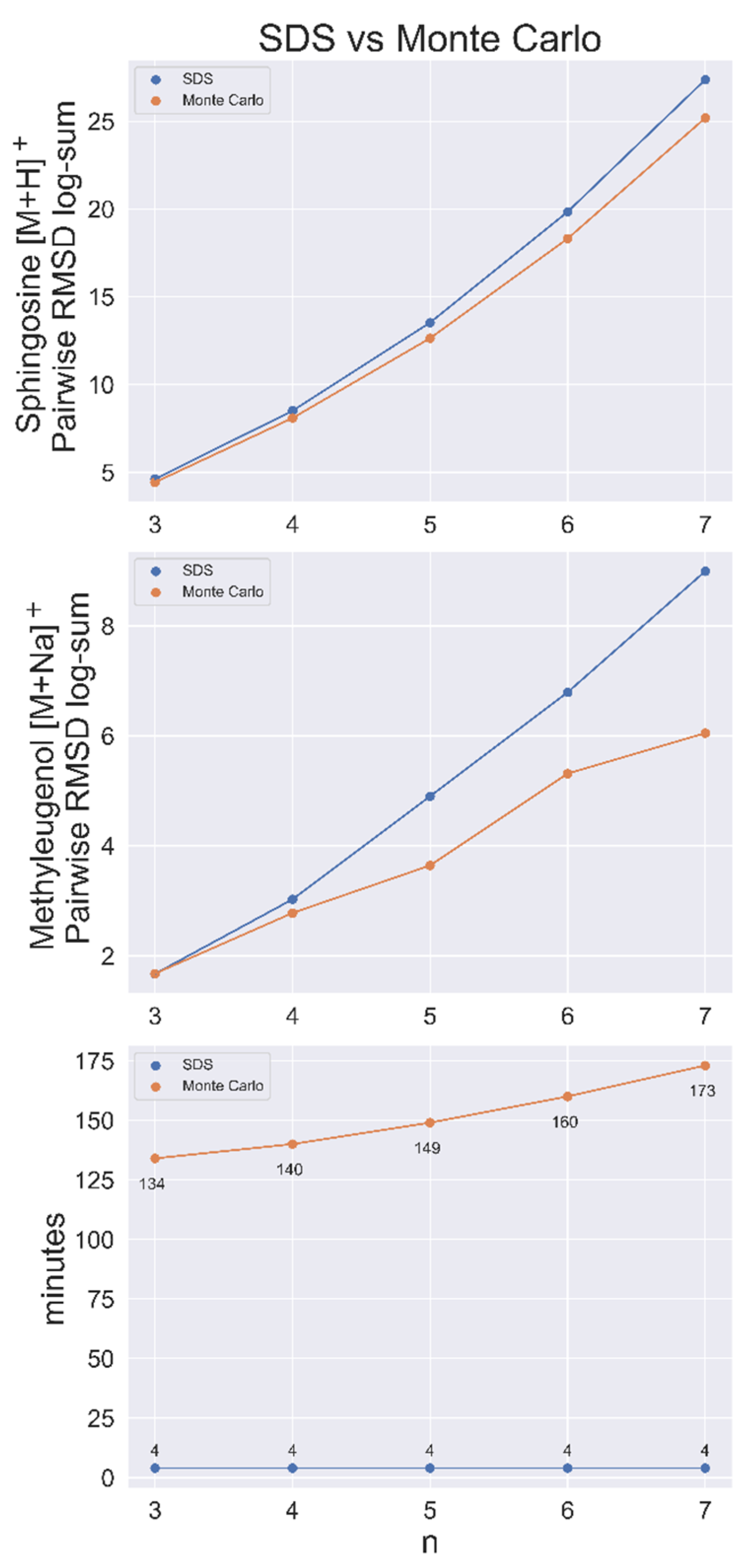
4.1. Performance against a Monte Carlo Method
4.2. Performance against the Exact Solution
4.3. Comparing Computational Costs of Calculating Pairwise Relations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Yongye, A.B.; Bender, A.; Martinez-Mayorga, K. Dynamic clustering threshold reduces conformer ensemble size while maintaining a biologically relevant ensemble. J. Comput.-Aided Mol. Des. 2010, 24, 675–686. [Google Scholar] [CrossRef] [PubMed]
- Colby, S.M.; Thomas, D.G.; Nunez, J.R.; Baxter, D.J.; Glaesemann, K.R.; Brown, J.M.; Pirrung, M.A.; Govind, N.; Teeguarden, J.G.; Metz, T.O.; et al. ISiCLE: A Quantum Chemistry Pipeline for Establishing in Silico Collision Cross Section Libraries. Anal. Chem. 2019, 91, 4346–4356. [Google Scholar] [CrossRef] [PubMed]
- Ebejer, J.P.; Morris, G.M.; Deane, C.M. Freely available conformer generation methods: How good are they? J. Chem. Inf. Model 2012, 52, 1146–1158. [Google Scholar] [CrossRef] [PubMed]
- Pearlman, D.; Case, D.; Caldwell, J.; Seibel, G.; Singh, U.C.; Weiner, P.; Kollman, P. AMBER 2017; Unversity of California: San Francisco, CA, USA, 2017. [Google Scholar]
- Pracht, P.; Bohle, F.; Grimme, S. Automated exploration of the low-energy chemical space with fast quantum chemical methods. Phys. Chem. Chem. Phys. 2020, 22, 7169–7192. [Google Scholar] [CrossRef] [PubMed]
- Nielson, F.F.; Colby, S.M.; Thomas, D.G.; Renslow, R.S.; Metz, T.O. Exploring the Impacts of Conformer Selection Methods on Ion Mobility Collision Cross Section Predictions. Anal. Chem. 2021, 93, 3830–3838. [Google Scholar] [CrossRef] [PubMed]
- Sabuncuoglu, I.; Bayiz, M. Job shop scheduling with beam search. Eur. J. Oper. Res. 1999, 118, 390–412. [Google Scholar] [CrossRef]
- Alsabti, K.; Ranka, S.; Singh, V. An efficient k-means clustering algorithm. Electr. Eng. Comput. Sci. 1997, 43. [Google Scholar]
- Kanungo, T.; Mount, D.M.; Netanyahu, N.S.; Piatko, C.D.; Silverman, R.; Wu, A.Y. An efficient k-means clustering algorithm: Analysis and implementation. IEEE Trans. Pattern Anal. Mach. Intell. 2002, 24, 881–892. [Google Scholar] [CrossRef]
- Khanmohammadi, S.; Adibeig, N.; Shanehbandy, S. An improved overlapping k-means clustering method for medical applications. Expert Syst. Appl. 2017, 67, 12–18. [Google Scholar] [CrossRef]
- Clark, R.D. OptiSim: An Extended Dissimilarity Selection Method for Finding Diverse Representative Subsets. J. Chem. Inf. Comput. Sci. 1997, 37, 1181–1188. [Google Scholar] [CrossRef]
- Elhamifar, E.; Sapiro, G.; Sastry, S.S. Dissimilarity-Based Sparse Subset Selection. IEEE Trans. Pattern Anal. Mach. Intell. 2016, 38, 2182–2197. [Google Scholar] [CrossRef] [PubMed]
- Willett, P. Dissimilarity-based algorithms for selecting structurally diverse sets of compounds. J. Comput. Biol. 1999, 6, 447–457. [Google Scholar] [CrossRef] [PubMed]
- Tanemura, K.A.; Das, S.; Merz, K.M. AutoGraph: Autonomous Graph-Based Clustering of Small-Molecule Conformations. J. Chem. Inf. Modeling 2021, 61, 1647–1656. [Google Scholar] [CrossRef] [PubMed]
- Ermanis, K.; Parkes, K.E.B.; Agback, T.; Goodman, J.M. The optimal DFT approach in DP4 NMR structure analysis-pushing the limits of relative configuration elucidation. Org. Biomol. Chem. 2019, 17, 5886–5890. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Jang, C.; Yadav, D.K.; Kim, M.H. The comparison of automated clustering algorithms for resampling representative conformer ensembles with RMSD matrix. J. Cheminform. 2017, 9, 21. [Google Scholar] [CrossRef] [PubMed]
- O’Boyle, N.M.; Banck, M.; James, C.A.; Morley, C.; Vandermeersch, T.; Hutchison, G.R. Open Babel: An open chemical toolbox. J. Cheminform. 2011, 3, 33. [Google Scholar] [CrossRef] [PubMed]
- O’Boyle, N.M.; Morley, C.; Hutchison, G.R. Pybel: A Python wrapper for the OpenBabel cheminformatics toolkit. Chem. Cent. J. 2008, 2, 5. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, S.; Yamaguchi, K.; Masuda, S. A maximum edge-weight clique extraction algorithm based on branch-and-bound. Discret. Optim. 2020, 37, 100583. [Google Scholar] [CrossRef]
- Martí, R.; Gallego, M.; Duarte, A. A branch and bound algorithm for the maximum diversity problem. Eur. J. Oper. Res. 2010, 200, 36–44. [Google Scholar] [CrossRef]
- Ghosh, J.B. Computational aspects of the maximum diversity problem. Oper. Res. Lett. 1996, 19, 175–181. [Google Scholar] [CrossRef]
- Sørensen, M.M. New facets and a branch-and-cut algorithm for the weighted clique problem. Eur. J. Oper. Res. 2004, 154, 57–70. [Google Scholar] [CrossRef]
- Glover, F. Improved linear integer programming formulations of nonlinear integer problems. Manag. Sci. 1975, 22, 455–460. [Google Scholar] [CrossRef]
- Gouveia, L.; Martins, P. Solving the maximum edge-weight clique problem in sparse graphs with compact formulations. EURO J. Comput. Optim. 2015, 3, 1–30. [Google Scholar] [CrossRef]
- Hosseinian, S.; Fontes, D.; Butenko, S. A nonconvex quadratic optimization approach to the maximum edge weight clique problem. J. Glob. Optim. 2018, 72, 219–240. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nielson, F.F.; Kay, B.; Young, S.J.; Colby, S.M.; Renslow, R.S.; Metz, T.O. Similarity Downselection: Finding the n Most Dissimilar Molecular Conformers for Reference-Free Metabolomics. Metabolites 2023, 13, 105. https://doi.org/10.3390/metabo13010105
Nielson FF, Kay B, Young SJ, Colby SM, Renslow RS, Metz TO. Similarity Downselection: Finding the n Most Dissimilar Molecular Conformers for Reference-Free Metabolomics. Metabolites. 2023; 13(1):105. https://doi.org/10.3390/metabo13010105
Chicago/Turabian StyleNielson, Felicity F., Bill Kay, Stephen J. Young, Sean M. Colby, Ryan S. Renslow, and Thomas O. Metz. 2023. "Similarity Downselection: Finding the n Most Dissimilar Molecular Conformers for Reference-Free Metabolomics" Metabolites 13, no. 1: 105. https://doi.org/10.3390/metabo13010105
APA StyleNielson, F. F., Kay, B., Young, S. J., Colby, S. M., Renslow, R. S., & Metz, T. O. (2023). Similarity Downselection: Finding the n Most Dissimilar Molecular Conformers for Reference-Free Metabolomics. Metabolites, 13(1), 105. https://doi.org/10.3390/metabo13010105






