Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia: Host, Pathogen, and Treatment
Abstract
1. Introduction
Persistent MRSAB
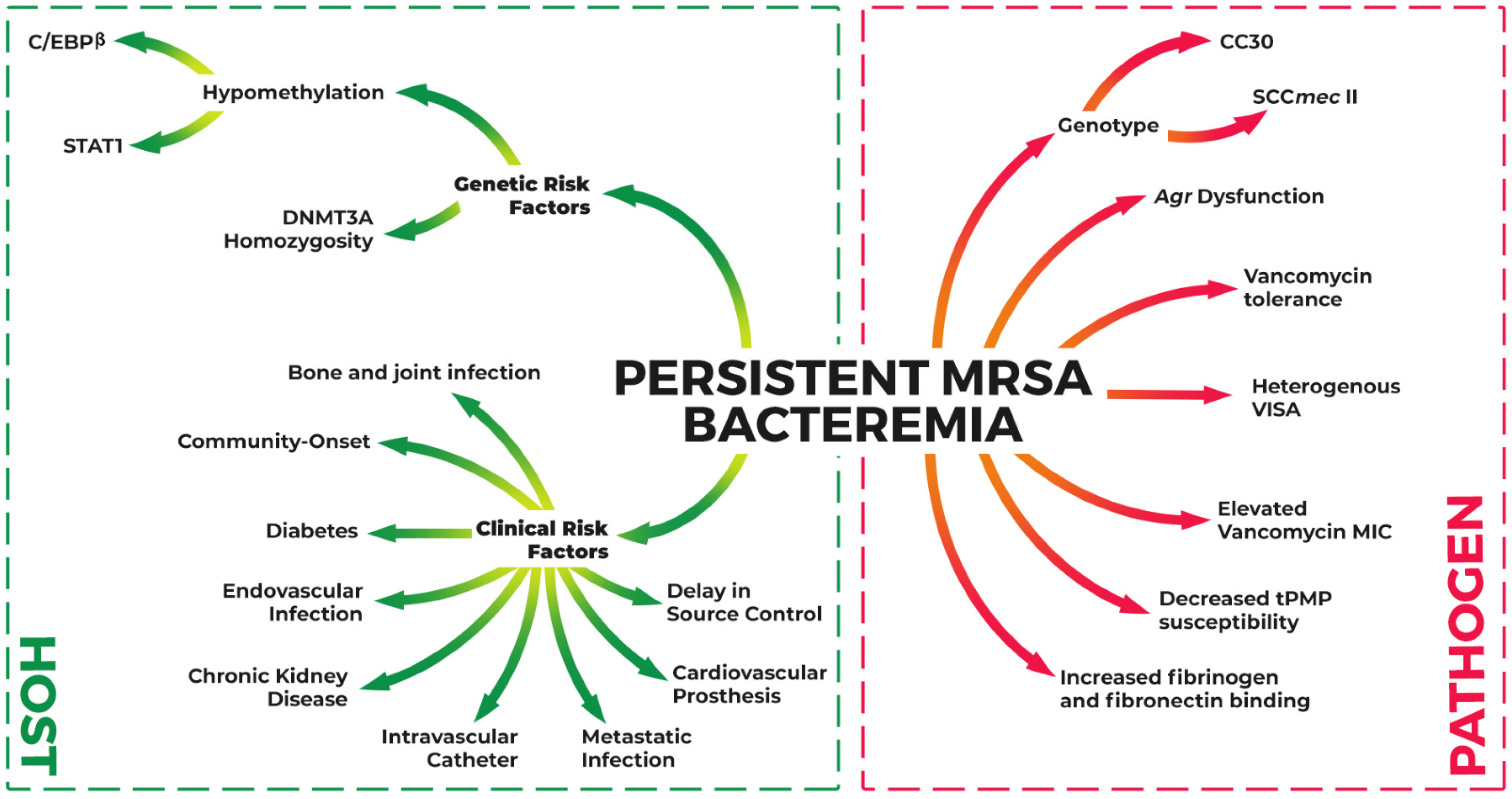
2. Host Factors Associated with Persistent MRSAB
2.1. Clinical Risk Factors
2.2. Host Genetic Variation and SAB
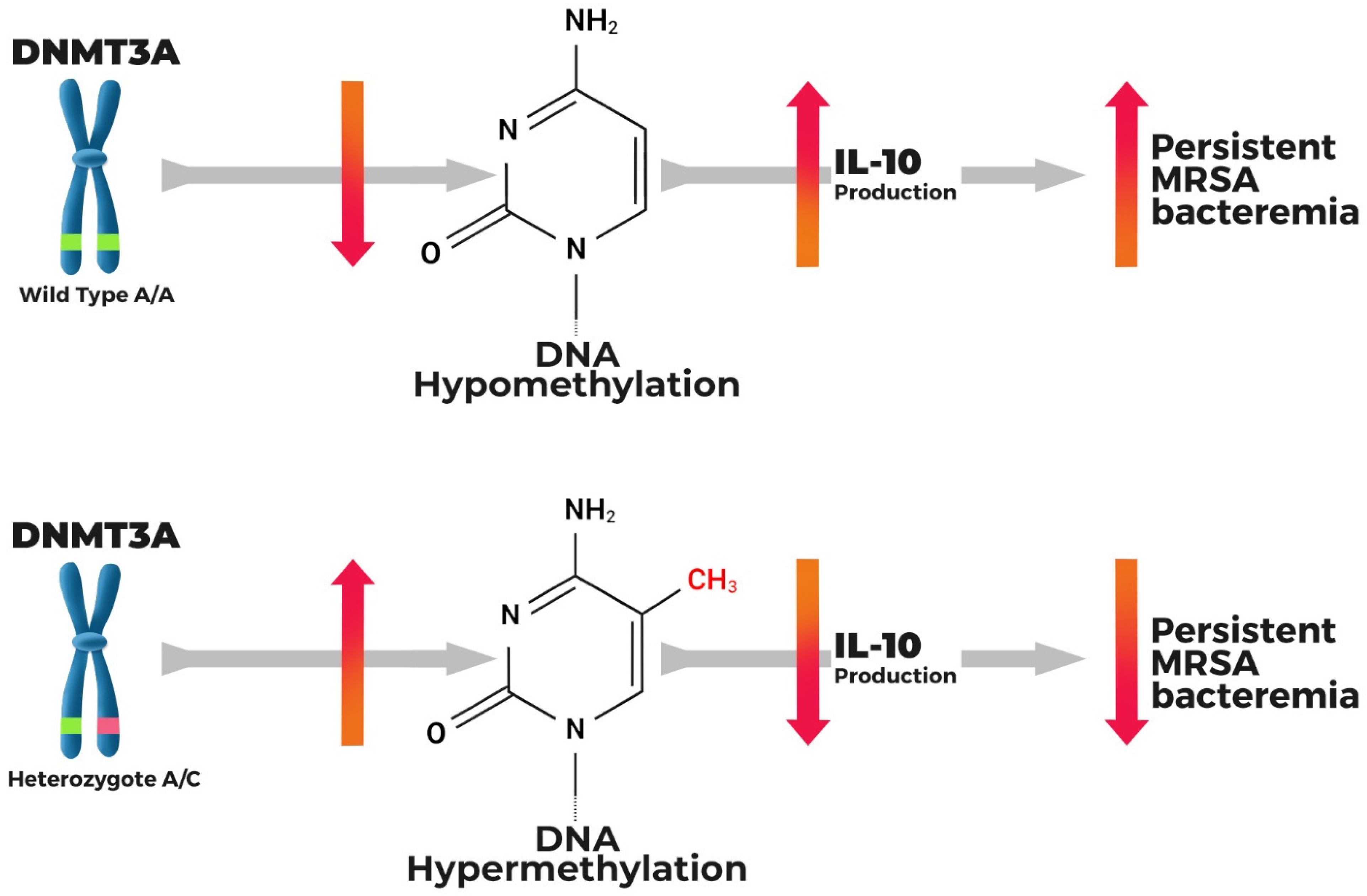
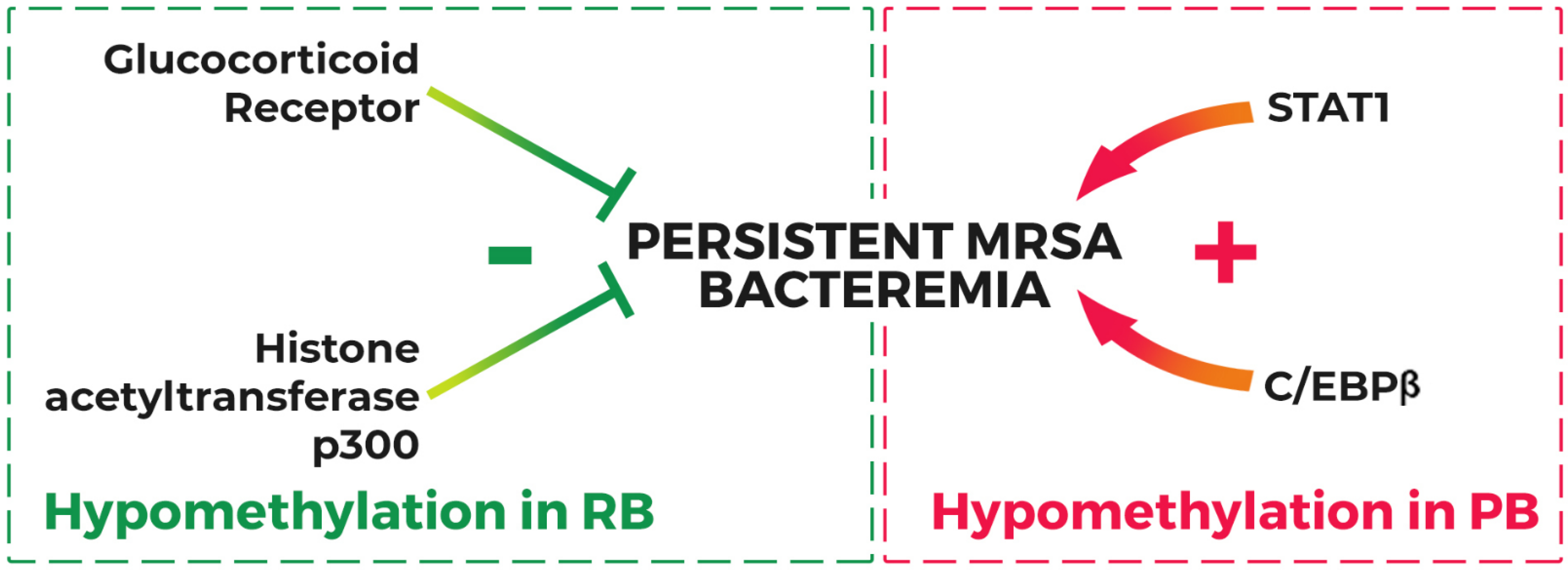
2.3. Host Genetic Variation and Persistent MRSAB
2.4. Biomarkers for Persistent SAB
3. Pathogen-Associated Risk Factors for Persistent S. aureus Bacteremia
3.1. Accessory Gene Regulator Dysfunction
3.2. Variability in Virulence Factor Production
3.3. Phenotypic Variability of SAB Isolates
3.4. Antibiotic Tolerance
3.5. Reduced Vancomycin Susceptibility and Heterogenous Vancomycin-Intermediate S. aureus
4. Treatment of Persistent MRSAB
4.1. The Past
4.2. The Present
4.3. The Future
5. Conclusions
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Kourtis, A.P.; Hatfield, K.; Baggs, J.; Mu, Y.; See, I.; Epson, E.; Nadle, J.; Kainer, M.A.; Dumyati, G.; Petit, S.; et al. Vital Signs: Epidemiology and Recent Trends in Methicillin-Resistant and in Methicillin-Susceptible Staphylococcus aureus Bloodstream Infections—United States. MMWR Morb. Mortal. Wkly. Rep. 2019, 68, 214–219. [Google Scholar] [CrossRef] [PubMed]
- Jernigan, J.A.; Hatfield, K.M.; Wolford, H.; Nelson, R.E.; Olubajo, B.; Reddy, S.C.; McCarthy, N.; Paul, P.; McDonald, L.C.; Kallen, A.; et al. Multidrug-Resistant Bacterial Infections in U.S. Hospitalized Patients 2012–2017. N. Engl. J. Med. 2020, 382, 1309–1319. [Google Scholar] [CrossRef] [PubMed]
- van Hal, S.J.; Jensen, S.O.; Vaska, V.L.; Espedido, B.A.; Paterson, D.L.; Gosbell, I.B. Predictors of mortality in Staphylococcus aureus Bacteremia. Clin. Microbiol. Rev. 2012, 25, 362–386. [Google Scholar] [CrossRef] [PubMed]
- Bai, A.D.; Lo, C.K.; Komorowski, A.S.; Suresh, M.; Guo, K.; Garg, A.; Tandon, P.; Senecal, J.; Del Corpo, O.; Stefanova, I.; et al. Staphylococcus aureus bacteraemia mortality: A systematic review and meta-analysis. Clin. Microbiol. Infect. 2022, 28, 1076–1084. [Google Scholar] [CrossRef]
- Yousaf, A.; Baird, G.L.; Mermel, L. Association of Infectious Disease Consultation with Clinical Outcomes in Patients with Staphylococcus aureus Bacteremia at Low Risk for Endocarditis. Open Forum Infect. Dis. 2018, 5. [Google Scholar] [CrossRef]
- Goto, M.; Jones, M.P.; Schweizer, M.L.; Livorsi, D.J.; Perencevich, E.N.; Richardson, K.; Beck, B.F.; Alexander, B.; Ohl, M.E. Association of Infectious Diseases Consultation with Long-term Postdischarge Outcomes among Patients with Staphylococcus aureus Bacteremia. JAMA Netw. Open 2020, 3, e1921048. [Google Scholar] [CrossRef]
- Bai, A.D.; Showler, A.; Burry, L.; Steinberg, M.; Ricciuto, D.R.; Fernandes, T.; Chiu, A.; Raybardhan, S.; Science, M.; Fernando, E.; et al. Impact of Infectious Disease Consultation on Quality of Care, Mortality, and Length of Stay in Staphylococcus aureus Bacteremia: Results from a Large Multicenter Cohort Study. Clin. Infect. Dis. 2015, 60, 1451–1461. [Google Scholar] [CrossRef]
- Pragman, A.A.; Kuskowski, M.A.; Abraham, J.; Filice, G.A. Infectious Disease Consultation for Staphylococcus aureus Bacteremia Improves Patient Management and Outcomes. Infect. Dis. Clin. Pract. 2012, 20, 261–267. [Google Scholar] [CrossRef]
- Holland, T.L.; Arnold, C.J.; Fowler, V.G. Clinical Management of Staphylococcus aureus Bacteremia: A review. JAMA 2014, 312, 1330–1341. [Google Scholar] [CrossRef]
- Liu, C.; Bayer, A.; Cosgrove, S.E.; Daum, R.S.; Fridkin, S.K.; Gorwitz, R.J.; Kaplan, S.L.; Karchmer, A.W.; Levine, D.P.; Murray, B.E.; et al. Clinical Practice Guidelines by the Infectious Diseases Society of America for the Treatment of Methicillin-Resistant Staphylococcus aureus Infections in Adults and Children: Executive Summary. Clin. Infect. Dis. 2011, 52, 285–292. [Google Scholar] [CrossRef]
- Fowler, V.G.; Olsen, M.K.; Corey, G.R.; Woods, C.W.; Cabell, C.H.; Reller, L.B.; Cheng, A.C.; Dudley, T.; Oddone, E.Z. Clinical Identifiers of Complicated Staphylococcus aureus Bacteremia. Arch. Intern. Med. 2003, 163, 2066–2072. [Google Scholar] [CrossRef]
- Minejima, E.; Mai, N.; Bui, N.; Mert, M.; Mack, W.J.; She, R.C.; Nieberg, P.; Spellberg, B.; Wong-Beringer, A. Defining the Breakpoint Duration of Staphylococcus aureus Bacteremia Predictive of Poor Outcomes. Clin. Infect. Dis. 2019, 70, 566–573. [Google Scholar] [CrossRef]
- Minejima, E.; Bensman, J.; She, R.C.; Mack, W.J.; Tran, M.T.; Ny, P.; Lou, M.; Yamaki, J.; Nieberg, P.; Ho, J.; et al. A Dysregulated Balance of Proinflammatory and Anti-Inflammatory Host Cytokine Response Early during Therapy Predicts Persistence and Mortality in Staphylococcus aureus Bacteremia. Crit. Care Med. 2016, 44, 671–679. [Google Scholar] [CrossRef]
- Kuehl, R.; Morata, L.; Boeing, C.; Subirana, I.; Seifert, H.; Rieg, S.; Kern, W.V.; Bin Kim, H.; Kim, E.S.; Liao, C.-H.; et al. Defining persistent Staphylococcus aureus bacteraemia: Secondary analysis of a prospective cohort study. Lancet Infect. Dis. 2020, 20, 1409–1417. [Google Scholar] [CrossRef]
- Souli, M.; Ruffin, F.; Choi, S.-H.; Park, L.P.; Gao, S.; Lent, N.C.; Sharma-Kuinkel, B.K.; Thaden, J.T.; Maskarinec, S.; Wanda, L.; et al. Changing Characteristics of Staphylococcus aureus Bacteremia: Results from a 21-Year, Prospective, Longitudinal Study. Clin. Infect. Dis. 2019, 69, 1868–1877. [Google Scholar] [CrossRef]
- Fowler, J.V.G.; Sakoulas, G.; McIntyre, L.M.; Meka, V.G.; Arbeit, R.D.; Cabell, C.H.; Stryjewski, M.E.; Eliopoulos, G.M.; Reller, L.B.; Corey, G.R.; et al. Persistent Bacteremia Due to Methicillin-Resistant Staphylococcus aureus Infection Is Associated with agr Dysfunction and Low-Level In Vitro Resistance to Thrombin-Induced Platelet Microbicidal Protein. J. Infect. Dis. 2004, 190, 1140–1149. [Google Scholar] [CrossRef]
- Levine, D.P.; Fromm, B.S.; Reddy, B.R. Slow Response to Vancomycin or Vancomycin plus Rifampin in Methicillin-resistant Staphylococcus aureus Endocarditis. Ann. Intern. Med. 1991, 115, 674–680. [Google Scholar] [CrossRef] [PubMed]
- Markowitz, N.; Quinn, E.L.; Saravolatz, L.D. Trimethoprim-Sulfamethoxazole Compared with Vancomycin for the Treatment of Staphylococcus aureus Infection. Ann. Intern. Med. 1992, 117, 390–398. [Google Scholar] [CrossRef] [PubMed]
- Fowler, V.G., Jr.; Boucher, H.W.; Corey, G.R.; Abrutyn, E.; Karchmer, A.W.; Rupp, M.E.; Levine, D.P.; Chambers, H.F.; Tally, F.P.; Vigliani, G.A.; et al. Daptomycin versus Standard Therapy for Bacteremia and Endocarditis Caused by Staphylococcus aureus. N. Engl. J. Med. 2006, 355, 653–665. [Google Scholar] [CrossRef]
- Pani, A.; Colombo, F.; Agnelli, F.; Frantellizzi, V.; Baratta, F.; Pastori, D.; Scaglione, F. Off-label use of ceftaroline fosamil: A systematic review. Int. J. Antimicrob. Agents 2019, 54, 562–571. [Google Scholar] [CrossRef] [PubMed]
- Holland, T.L.; Bayer, A.S.; Fowler, V.G. Persistent methicillin-Resistant Staphylococcus aureus Bacteremia: Resetting the Clock for Optimal Management. Clin. Infect. Dis. 2022, 75, 1668–1674. [Google Scholar] [CrossRef]
- Chong, Y.P.; Park, S.-J.; Kim, H.S.; Kim, E.S.; Kim, M.-N.; Park, K.-H.; Kim, S.-H.; Lee, S.-O.; Choi, S.-H.; Jeong, J.-Y.; et al. Persistent Staphylococcus aureus Bacteremia. Medicine 2013, 92, 98–108. [Google Scholar] [CrossRef] [PubMed]
- Khatib, R.; Johnson, L.B.; Sharma, M.; Fakih, M.G.; Ganga, R.; Riederer, K. Persistent Staphylococcus aureus bacteremia: Incidence and outcome trends over time. Scand. J. Infect. Dis. 2009, 41, 4–9. [Google Scholar] [CrossRef]
- Yoon, Y.K.; Kim, J.Y.; Park, D.W.; Sohn, J.W.; Kim, M.J. Predictors of persistent methicillin-resistant Staphylococcus aureus bacteraemia in patients treated with vancomycin. J. Antimicrob. Chemother. 2010, 65, 1015–1018. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, C.; Huang, J.; Jin, N.; Noskin, G.A.; Zembower, T.R.; Bolon, M. Persistent Staphylococcus aureus Bacteremia: An Analysis of Risk Factors and Outcomes. Arch. Intern. Med. 2007, 167, 1861–1867. [Google Scholar] [CrossRef] [PubMed]
- Khatib, R.; Johnson, L.B.; Fakih, M.G.; Riederer, K.; Khosrovaneh, A.; Tabriz, M.S.; Sharma, M.; Saeed, S. Persistence in Staphylococcus aureus bacteremia: Incidence, characteristics of patients and outcome. Scand. J. Infect. Dis. 2006, 38, 7–14. [Google Scholar] [CrossRef]
- Chung, H.; Kim, E.; Yang, E.; Lee, Y.W.; Park, J.H.; Bae, S.; Jung, J.; Kim, M.J.; Chong, Y.P.; Kim, S.-H.; et al. C-reactive protein predicts persistent bacteremia caused by community-acquired methicillin-resistant Staphylococcus aureus strain. Eur. J. Clin. Microbiol. Infect. Dis. 2021, 40, 2497–2504. [Google Scholar] [CrossRef]
- Ganga, R.; Riederer, K.; Sharma, M.; Fakih, M.G.; Johnson, L.B.; Shemes, S.; Khatib, R. Role of SCC mec Type in Outcome of Staphylococcus aureus Bacteremia in a Single Medical Center. J. Clin. Microbiol. 2009, 47, 590–595. [Google Scholar] [CrossRef]
- Kwok, A.J.; Mentzer, A.; Knight, J.C. Host genetics and infectious disease: New tools, insights and translational opportunities. Nat. Rev. Genet. 2020, 22, 137–153. [Google Scholar] [CrossRef]
- Sørensen, T.I.; Nielsen, G.G.; Andersen, P.K.; Teasdale, T.W. Genetic and Environmental Influences on Premature Death in Adult Adoptees. N. Engl. J. Med. 1988, 318, 727–732. [Google Scholar] [CrossRef]
- Komiyama, A.; Saitoh, H.; Yamazaki, M.; Kawai, H.; Miyagawa, Y.; Akabane, T.; Ichikawa, M.; Shigematsu, H. Hyperactive phagocytosis by circulating neutrophils and monocytes in Chèdiak-Higashi syndrome. Scand. J. Haematol. 1986, 37, 162–167. [Google Scholar] [CrossRef] [PubMed]
- Holland, S.; DeLeo, F.R.; Elloumi, H.Z.; Hsu, A.P.; Uzel, G.; Brodsky, N.; Freeman, A.F.; Demidowich, A.; Davis, J.; Turner, M.L.C.; et al. STAT3 Mutations in the Hyper-IgE Syndrome. N. Engl. J. Med. 2007, 357, 1608–1619. [Google Scholar] [CrossRef]
- Ben-Ari, J.; Wolach, O.; Gavrieli, R.; Wolach, B. Infections associated with chronic granulomatous disease: Linking genetics to phenotypic expression. Expert Rev. Anti-Infect. Ther. 2012, 10, 881–894. [Google Scholar] [CrossRef]
- Messina, J.A.; Thaden, J.T.; Sharma-Kuinkel, B.K.; Fowler, V.G. Impact of Bacterial and Human Genetic Variation on Staphylococcus aureus Infections. PLoS Pathog. 2016, 12, e1005330. [Google Scholar] [CrossRef] [PubMed]
- Oestergaard, L.B.; Christiansen, M.N.; Schmiegelow, M.D.; Skov, R.L.; Andersen, P.S.; Petersen, A.; Aasbjerg, K.; Gerds, T.A.; Andersen, P.K.; Torp-Pedersen, C. Familial Clustering of Staphylococcus aureus Bacteremia in First-Degree Relatives. Ann. Intern. Med. 2016, 165, 390–398. [Google Scholar] [CrossRef] [PubMed]
- Eye, Z.; Vasco, D.A.; Carter, T.C.; Brilliant, M.H.; Schrodi, S.; Shukla, S.K. Genome wide association study of SNP-, gene-, and pathway-based approaches to identify genes influencing susceptibility to Staphylococcus aureus infections. Front. Genet. 2014, 5, 125. [Google Scholar] [CrossRef]
- Nelson, C.L.; Pelak, K.; Podgoreanu, M.V.; Ahn, S.H.; Scott, W.K.; Allen, A.S.; Cowell, L.G.; Rude, T.H.; Zhang, Y.; Tong, A.; et al. A genome-wide association study of variants associated with acquisition of Staphylococcus aureus bacteremia in a healthcare setting. BMC Infect. Dis. 2014, 14, 833. [Google Scholar] [CrossRef]
- DeLorenze, G.N.; Nelson, C.L.; Scott, W.K.; Allen, A.S.; Ray, G.T.; Tsai, A.-L.; Quesenberry, C.P.; Fowler, V.G. Polymorphisms in HLA Class II Genes Are Associated with Susceptibility to Staphylococcus aureus Infection in a White Population. J. Infect. Dis. 2015, 213, 816–823. [Google Scholar] [CrossRef]
- Cyr, D.D.; Allen, A.S.; Du, G.-J.; Ruffin, F.; Adams, C.; Thaden, J.T.; Maskarinec, S.A.; Souli, M.; Guo, S.; Dykxhoorn, D.M.; et al. Evaluating genetic susceptibility to Staphylococcus aureus bacteremia in African Americans using admixture mapping. Genes Immun. 2017, 18, 95–99. [Google Scholar] [CrossRef]
- Kotb, M.; Norrby-Teglund, A.; McGeer, A.; El-Sherbini, H.; Dorak, M.T.; Khurshid, A.; Green, K.; Peeples, J.; Wade, J.; Thomson, G.; et al. An immunogenetic and molecular basis for differences in outcomes of invasive group A streptococcal infections. Nat. Med. 2002, 8, 1398–1404. [Google Scholar] [CrossRef]
- Nooh, M.M.; El-Gengehi, N.; Kansal, R.; David, C.S.; Kotb, M. HLA Transgenic Mice Provide Evidence for a Direct and Dominant Role of HLA Class II Variation in Modulating the Severity of Streptococcal Sepsis. J. Immunol. 2007, 178, 3076–3083. [Google Scholar] [CrossRef]
- Llewelyn, M.; Sriskandan, S.; Peakman, M.; Ambrozak, D.R.; Douek, D.C.; Kwok, W.W.; Cohen, J.; Altmann, D.M. HLA Class II Polymorphisms Determine Responses to Bacterial Superantigens. J. Immunol. 2004, 172, 1719–1726. [Google Scholar] [CrossRef]
- Kim, J.; Urban, R.G.; Strominger, J.L.; Wiley, D.C. Toxic Shock Syndrome Toxin-1 Complexed with a Class II Major Histocompatibility Molecule HLA-DR1. Science 1994, 266, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Lavoie, P.M.; Thibodeau, J.; Cloutier, I.; Busch, R.; Sékaly, R.-P. Selective binding of bacterial toxins to major histocompatibility complex class II-expressing cells is controlled by invariant chain and HLA-DM. Proc. Natl. Acad. Sci. USA 1997, 94, 6892–6897. [Google Scholar] [CrossRef] [PubMed]
- Salgado-Pabon, W.; Breshears, L.; Spaulding, A.R.; Merriman, J.A.; Stach, C.S.; Horswill, A.R.; Peterson, M.L.; Schlievert, P.M. Superantigens Are Critical for Staphylococcus aureus Infective Endocarditis, Sepsis, and Acute Kidney Injury. Mbio 2013, 4, e00494-13. [Google Scholar] [CrossRef] [PubMed]
- Medie, F.M.; Sharma-Kuinkel, B.K.; Ruffin, F.; Chan, L.C.; Rossetti, M.; Chang, Y.-L.; Park, L.P.; Bayer, A.S.; Filler, S.G.; Ahn, R.; et al. Genetic variation of DNA methyltransferase-3A contributes to protection against persistent MRSA bacteremia in patients. Proc. Natl. Acad. Sci. USA 2019, 116, 20087–20096. [Google Scholar] [CrossRef]
- Guimaraes, A.O.; Cao, Y.; Hong, K.; Mayba, O.; Peck, M.C.; Gutierrez, J.; Ruffin, F.; Carrasco-Triguero, M.; Dinoso, J.B.; Clemenzi-Allen, A.; et al. A Prognostic Model of Persistent Bacteremia and Mortality in Complicated Staphylococcus aureus Bloodstream Infection. Clin. Infect. Dis. 2018, 68, 1502–1511. [Google Scholar] [CrossRef]
- Redford, P.S.; Murray, P.J.; O’Garra, A. The role of IL-10 in immune regulation during M. tuberculosis infection. Mucosal Immunol. 2011, 4, 261–270. [Google Scholar] [CrossRef]
- Gazzinelli, R.T.; Oswald, I.P.; James, S.L.; Sher, A. IL-10 inhibits parasite killing and nitrogen oxide production by IFN-gamma-activated macrophages. J. Immunol. 1992, 148, 1792–1796. [Google Scholar] [CrossRef]
- O’Leary, S.; O’Sullivan, M.P.; Keane, J. IL-10 Blocks Phagosome Maturation in Mycobacterium tuberculosis–Infected Human Macrophages. Am. J. Respir. Cell Mol. Biol. 2011, 45, 172–180. [Google Scholar] [CrossRef]
- Redpath, S.; Ghazal, P.; Gascoigne, N.R. Hijacking and exploitation of IL-10 by intracellular pathogens. Trends Microbiol. 2001, 9, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Dokka, S.; Shi, X.; Leonard, S.; Wang, L.; Castranova, V.; Rojanasakul, Y. Interleukin-10-mediated inhibition of free radical generation in macrophages. Am. J. Physiol. Cell. Mol. Physiol. 2001, 280, L1196–L1202. [Google Scholar] [CrossRef]
- Chang, Y.-L.; Rossetti, M.; Gjertson, D.W.; Rubbi, L.; Thompson, M.; Montoya, D.J.; Morselli, M.; Ruffin, F.; Hoffmann, A.; Pellegrini, M.; et al. Human DNA methylation signatures differentiate persistent from resolving MRSA bacteremia. Proc. Natl. Acad. Sci. USA 2021, 118, e2000663118. [Google Scholar] [CrossRef] [PubMed]
- Hirai, H.; Zhang, P.; Dayaram, T.; Hetherington, C.J.; Mizuno, S.-I.; Imanishi, J.; Akashi, K.; Tenen, D. C/EBPβ is required for ‘emergency’ granulopoiesis. Nat. Immunol. 2006, 7, 732–739. [Google Scholar] [CrossRef] [PubMed]
- Petta, I.; Dejager, L.; Ballegeer, M.; Lievens, S.; Tavernier, J.; De Bosscher, K.; Libert, C. The Interactome of the Glucocorticoid Receptor and Its Influence on the Actions of Glucocorticoids in Combatting Inflammatory and Infectious Diseases. Microbiol. Mol. Biol. Rev. 2016, 80, 495–522. [Google Scholar] [CrossRef] [PubMed]
- Cao, Y.; Guimaraes, A.O.; Peck, M.C.; Mayba, O.; Ruffin, F.; Hong, K.; Carrasco-Triguero, M.; Fowler, V.G.; Maskarinec, S.A.; Rosenberger, C.M. Risk stratification biomarkers for Staphylococcus aureus bacteraemia. Clin. Transl. Immunol. 2020, 9, e1110. [Google Scholar] [CrossRef]
- Zhao, H.; Xu, S.; Yang, H.; He, C.; Xu, X.; Hu, F.; Shu, W.; Gong, F.; Zhang, C.; Liu, Q. Molecular Typing and Variations in Amount of tst Gene Expression of TSST-1-Producing Clinical Staphylococcus aureus Isolates. Front. Microbiol. 2019, 10, 1388. [Google Scholar] [CrossRef]
- Loughman, J.A.; Fritz, S.A.; Storch, G.A.; Hunstad, D.A. Virulence Gene Expression in Human Community-Acquired Staphylococcus aureus Infection. J. Infect. Dis. 2009, 199, 294–301. [Google Scholar] [CrossRef]
- Lacoma, A.; Laabei, M.; Sánchez-Herrero, J.F.; Young, B.; Godoy-Tena, G.; Gomes-Fernandes, M.; Sumoy, L.; Plans, O.; Arméstar, F.; Prat, C. Genotypic and Phenotypic Characterization of Staphylococcus aureus Isolates from the Respiratory Tract in Mechanically-Ventilated Patients. Toxins 2021, 13, 122. [Google Scholar] [CrossRef]
- Afzal, M.; Vijay, A.K.; Stapleton, F.; Willcox, M.D.P. Genomics of Staphylococcus aureus Strains Isolated from Infectious and Non-Infectious Ocular Conditions. Antibiotics 2022, 11, 1011. [Google Scholar] [CrossRef]
- Traber, K.E.; Lee, E.; Benson, S.; Corrigan, R.; Cantera, M.; Shopsin, B.; Novick, R.P. agr function in clinical Staphylococcus aureus isolates. Microbiology 2008, 154, 2265–2274. [Google Scholar] [CrossRef] [PubMed]
- Sakoulas, G.; Eliopoulos, G.M.; Moellering, R.C., Jr.; Wennersten, C.; Venkataraman, L.; Novick, R.P.; Gold, H.S. Accessory Gene Regulator (agr) Locus in Geographically Diverse Staphylococcus aureus Isolates with Reduced Susceptibility to Vancomycin. Antimicrob. Agents Chemother. 2002, 46, 1492–1502. [Google Scholar] [CrossRef] [PubMed]
- Chong, Y.P.; Kim, E.S.; Park, S.-J.; Park, K.-H.; Kim, T.; Kim, M.-N.; Kim, S.-H.; Lee, S.-O.; Choi, S.-H.; Woo, J.H.; et al. Accessory Gene Regulator (agr) Dysfunction in Staphylococcus aureus Bloodstream Isolates from South Korean Patients. Antimicrob. Agents Chemother. 2013, 57, 1509–1512. [Google Scholar] [CrossRef] [PubMed]
- Tuchscherr, L.; Pöllath, C.; Siegmund, A.; Deinhardt-Emmer, S.; Hoerr, V.; Svensson, C.-M.; Figge, M.T.; Monecke, S.; Löffler, B. Clinical S. aureus Isolates Vary in Their Virulence to Promote Adaptation to the Host. Toxins 2019, 11, 135. [Google Scholar] [CrossRef] [PubMed]
- Midorikawa, K.; Ouhara, K.; Komatsuzawa, H.; Kawai, T.; Yamada, S.; Fujiwara, T.; Yamazaki, K.; Sayama, K.; Taubman, M.A.; Kurihara, H.; et al. Staphylococcus aureus Susceptibility to Innate Antimicrobial Peptides, β-Defensins and CAP18, Expressed by Human Keratinocytes. Infect. Immun. 2003, 71, 3730–3739. [Google Scholar] [CrossRef] [PubMed]
- Dhawan, V.K.; Yeaman, M.R.; Cheung, A.L.; Kim, E.; Sullam, P.M.; Bayer, A.S. Phenotypic resistance to thrombin-induced platelet microbicidal protein in vitro is correlated with enhanced virulence in experimental endocarditis due to Staphylococcus aureus. Infect. Immun. 1997, 65, 3293–3299. [Google Scholar] [CrossRef]
- Dhawan, V.K.; Bayer, A.S.; Yeaman, M.R. In Vitro Resistance to Thrombin-Induced Platelet Microbicidal Protein Is Associated with Enhanced Progression and Hematogenous Dissemination in Experimental Staphylococcus aureus Infective Endocarditis. Infect. Immun. 1998, 66, 3476–3479. [Google Scholar] [CrossRef]
- Wu, T.; Yeaman, M.R.; Bayer, A.S. In vitro resistance to platelet microbicidal protein correlates with endocarditis source among bacteremic staphylococcal and streptococcal isolates. Antimicrob. Agents Chemother. 1994, 38, 729–732. [Google Scholar] [CrossRef]
- Jenul, C.; Horswill, A.R. Regulation of Staphylococcus aureus Virulence. Microbiol. Spectr. 2019, 7. [Google Scholar] [CrossRef]
- Painter, K.L.; Krishna, A.; Wigneshweraraj, S.; Edwards, A.M. What role does the quorum-sensing accessory gene regulator system play during Staphylococcus aureus bacteremia? Trends Microbiol. 2014, 22, 676–685. [Google Scholar] [CrossRef]
- Butterfield, J.M.; Tsuji, B.T.; Brown, J.; Ashley, E.D.; Hardy, D.; Brown, K.; Forrest, A.; Lodise, T.P. Predictors of agr Dysfunction in Methicillin-Resistant Staphylococcus aureus (MRSA) Isolates among Patients with MRSA Bloodstream Infections. Antimicrob. Agents Chemother. 2011, 55, 5433–5437. [Google Scholar] [CrossRef] [PubMed]
- Schweizer, M.L.; Furuno, J.P.; Sakoulas, G.; Johnson, J.K.; Harris, A.D.; Shardell, M.D.; McGregor, J.C.; Thom, K.A.; Perencevich, E.N. Increased Mortality with Accessory Gene Regulator (agr) Dysfunction in Staphylococcus aureus among Bacteremic Patients. Antimicrob. Agents Chemother. 2011, 55, 1082–1087. [Google Scholar] [CrossRef] [PubMed]
- Jang, H.-C.; Kang, S.-J.; Choi, S.-M.; Park, K.-H.; Shin, J.-H.; Choy, H.E.; Jung, S.-I.; BIN Kim, H. Difference in agr Dysfunction and Reduced Vancomycin Susceptibility between MRSA Bacteremia Involving SCCmec Types IV/IVa and I–III. PLoS ONE 2012, 7, e49136. [Google Scholar] [CrossRef] [PubMed]
- Park, S.-Y.; Chong, Y.P.; Park, H.J.; Park, K.-H.; Moon, S.M.; Jeong, J.-Y.; Kim, M.-N.; Kim, S.-H.; Lee, S.-O.; Choi, S.-H.; et al. agr dysfunction and persistent methicillin-resistant Staphylococcus aureus bacteremia in patients with removed eradicable foci. Infection 2012, 41, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Kang, C.K.; Kim, Y.K.; Jung, S.I.; Park, W.B.; Song, K.H.; Park, K.H.; Choe, P.G.; Jang, H.C.; Lee, S.; Kim, Y.S.; et al. agr functionality affects clinical outcomes in patients with persistent methicillin-resistant Staphylococcus aureus bacteraemia. Eur. J. Clin. Microbiol. Infect. Dis. 2017, 36, 2187–2191. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.Q.; Fowler, J.V.G.; Yeaman, M.R.; Perdreau-Remington, F.; Kreiswirth, B.N.; Bayer, A.S. Phenotypic and Genotypic Characteristics of Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia In Vitro and in an Experimental Endocarditis Model. J. Infect. Dis. 2009, 199, 201–208. [Google Scholar] [CrossRef] [PubMed]
- Seidl, K.; Bayer, A.S.; Fowler, V.G., Jr.; McKinnell, J.A.; Abdel Hady, W.; Sakoulas, G.; Yeaman, M.R.; Xiong, Y.Q. Combinatorial Phenotypic Signatures Distinguish Persistent from Resolving Methicillin-Resistant Staphylococcus aureus Bacteremia Isolates. Antimicrob. Agents Chemother. 2011, 55, 575–582. [Google Scholar] [CrossRef] [PubMed]
- Yeaman, M.R. Platelets in defense against bacterial pathogens. Cell. Mol. Life Sci. 2009, 67, 525–544. [Google Scholar] [CrossRef]
- Dhawan, V.K.; Yeaman, M.R.; Bayer, A.S. Influence of In Vitro Susceptibility Phenotype against Thrombin?Induced Platelet Microbicidal Protein on Treatment and Prophylaxis Outcomes of Experimental Staphylococcus aureus Endocarditis. J. Infect. Dis. 1999, 180, 1561–1568. [Google Scholar] [CrossRef][Green Version]
- Brauner, A.; Fridman, O.; Gefen, O.; Balaban, N.Q. Distinguishing between resistance, tolerance and persistence to antibiotic treatment. Nat. Rev. Microbiol. 2016, 14, 320–330. [Google Scholar] [CrossRef]
- Surewaard, B.G.; Deniset, J.F.; Zemp, F.J.; Amrein, M.; Otto, M.; Conly, J.; Omri, A.; Yates, R.M.; Kubes, P. Identification and treatment of the Staphylococcus aureus reservoir in vivo. J. Exp. Med. 2016, 213, 1141–1151. [Google Scholar] [CrossRef]
- Lehar, S.M.; Pillow, T.; Xu, M.; Staben, L.; Kajihara, K.K.; Vandlen, R.; DePalatis, L.; Raab, H.; Hazenbos, W.L.; Morisaki, J.H.; et al. Novel antibody–antibiotic conjugate eliminates intracellular S. aureus. Nature 2015, 527, 323–328. [Google Scholar] [CrossRef] [PubMed]
- Rowe, S.E.; Wagner, N.J.; Li, L.; Beam, J.E.; Wilkinson, A.D.; Radlinski, L.C.; Zhang, Q.; Miao, E.A.; Conlon, B.P. Reactive oxygen species induce antibiotic tolerance during systemic Staphylococcus aureus infection. Nat. Microbiol. 2019, 5, 282–290. [Google Scholar] [CrossRef] [PubMed]
- Beam, J.E.; Rowe, S.E.; Conlon, B.P. Shooting yourself in the foot: How immune cells induce antibiotic tolerance in microbial pathogens. PLoS Pathog. 2021, 17, e1009660. [Google Scholar] [CrossRef] [PubMed]
- Ledger, E.V.K.; Mesnage, S.; Edwards, A.M. Human serum triggers antibiotic tolerance in Staphylococcus aureus. Nat. Commun. 2022, 13, 2041. [Google Scholar] [CrossRef] [PubMed]
- Huemer, M.; Shambat, S.M.; Brugger, S.D.; Zinkernagel, A.S. Antibiotic resistance and persistence—Implications for human health and treatment perspectives. EMBO Rep. 2020, 21, e51034. [Google Scholar] [CrossRef] [PubMed]
- Rose, W.E.; Fallon, M.; Moran, J.J.M.; Vanderloo, J.P. Vancomycin Tolerance in Methicillin-Resistant Staphylococcus aureus: Influence of Vancomycin, Daptomycin, and Telavancin on Differential Resistance Gene Expression. Antimicrob. Agents Chemother. 2012, 56, 4422–4427. [Google Scholar] [CrossRef] [PubMed]
- Britt, N.S.; Patel, N.; Shireman, T.I.; El Atrouni, W.I.; Horvat, R.T.; Steed, M.E. Relationship between vancomycin tolerance and clinical outcomes in Staphylococcus aureus bacteraemia. J. Antimicrob. Chemother. 2016, 72, 535–542. [Google Scholar] [CrossRef]
- Gonzalez, N.; Sevillano, D.; Alou, L.; Cafini, F.; Gimenez, M.-J.; Gomez-Lus, M.-L.; Prieto, J.; Aguilar, L. Influence of the MBC/MIC ratio on the antibacterial activity of vancomycin versus linezolid against methicillin-resistant Staphylococcus aureus isolates in a pharmacodynamic model simulating serum and soft tissue interstitial fluid concentrations reported in diabetic patients. J. Antimicrob. Chemother. 2013, 68, 2291–2295. [Google Scholar] [CrossRef]
- Safdar, A.; Rolston, K.V.I. Vancomycin tolerance, a potential mechanism for refractory gram-positive bacteremia observational study in patients with cancer. Cancer 2006, 106, 1815–1820. [Google Scholar] [CrossRef]
- Levin-Reisman, I.; Ronin, I.; Gefen, O.; Braniss, I.; Shoresh, N.; Balaban, N.Q. Antibiotic tolerance facilitates the evolution of resistance. Science 2017, 355, 826–830. [Google Scholar] [CrossRef] [PubMed]
- Sader, H.S.; Jones, R.N.; Rossi, K.L.; Rybak, M.J. Occurrence of vancomycin-tolerant and heterogeneous vancomycin-intermediate strains (hVISA) among Staphylococcus aureus causing bloodstream infections in nine USA hospitals. J. Antimicrob. Chemother. 2009, 64, 1024–1028. [Google Scholar] [CrossRef] [PubMed]
- Moise, P.A.; Sakoulas, G.; Forrest, A.; Schentag, J.J. Vancomycin In Vitro Bactericidal Activity and Its Relationship to Efficacy in Clearance of Methicillin-Resistant Staphylococcus aureus Bacteremia. Antimicrob. Agents Chemother. 2007, 51, 2582–2586. [Google Scholar] [CrossRef]
- Elgrail, M.M.; Chen, E.; Shaffer, M.G.; Srinivasa, V.; Griffith, M.P.; Mustapha, M.M.; Shields, R.K.; Van Tyne, D.; Culyba, M.J. Convergent Evolution of Antibiotic Tolerance in Patients with Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia. Infect. Immun. 2022, 90, e00001-22. [Google Scholar] [CrossRef]
- Michaux, C.; Ronneau, S.; Giorgio, R.T.; Helaine, S. Antibiotic tolerance and persistence have distinct fitness trade-offs. PLoS Pathog. 2022, 18, e1010963. [Google Scholar] [CrossRef]
- McCormick, M.H.; McGuire, J.M.; Pittenger, G.E.; Pittenger, R.C.; Stark, W.M. Vancomycin, a new antibiotic. I. Chemical and biologic properties. Antibiot. Annu. 1955, 3, 606–611. [Google Scholar] [PubMed]
- Cong, Y.; Yang, S.; Rao, X. Vancomycin resistant Staphylococcus aureus infections: A review of case updating and clinical features. J. Adv. Res. 2020, 21, 169–176. [Google Scholar] [CrossRef]
- Shariati, A.; Dadashi, M.; Moghadam, M.T.; van Belkum, A.; Yaslianifard, S.; Darban-Sarokhalil, D. Global prevalence and distribution of vancomycin resistant, vancomycin intermediate and heterogeneously vancomycin intermediate Staphylococcus aureus clinical isolates: A systematic review and meta-analysis. Sci. Rep. 2020, 10, 12689. [Google Scholar] [CrossRef]
- Holland, T.L.; Fowler, V.G. Vancomycin Minimum Inhibitory Concentration and Outcome in Patients with Staphylococcus aureus Bacteremia: Pearl or Pellet? J. Infect. Dis. 2011, 204, 329–331. [Google Scholar] [CrossRef]
- van Hal, S.J.; Lodise, T.P.; Paterson, D.L. The Clinical Significance of Vancomycin Minimum Inhibitory Concentration in Staphylococcus aureus Infections: A Systematic Review and Meta-analysis. Clin. Infect. Dis. 2012, 54, 755–771. [Google Scholar] [CrossRef]
- Holmes, N.E.; Turnidge, J.D.; Munckhof, W.J.; Robinson, J.O.; Korman, T.; O’Sullivan, M.; Anderson, T.L.; Roberts, S.A.; Gao, W.; Christiansen, K.J.; et al. Antibiotic Choice May Not Explain Poorer Outcomes in Patients with Staphylococcus aureus Bacteremia and High Vancomycin Minimum Inhibitory Concentrations. J. Infect. Dis. 2011, 204, 340–347. [Google Scholar] [CrossRef]
- Neuner, E.A.; Casabar, E.; Reichley, R.; McKinnon, P.S. Clinical, microbiologic, and genetic determinants of persistent methicillin-resistant Staphylococcus aureus bacteremia. Diagn. Microbiol. Infect. Dis. 2010, 67, 228–233. [Google Scholar] [CrossRef]
- Adani, S.; Bhowmick, T.; Weinstein, M.P.; Narayanan, N. Impact of Vancomycin MIC on Clinical Outcomes of Patients with Methicillin-Resistant Staphylococcus aureus Bacteremia Treated with Vancomycin at an Institution with Suppressed MIC Reporting. Antimicrob. Agents Chemother. 2018, 62, e02512-17. [Google Scholar] [CrossRef]
- van Hal, S.J.; Paterson, D.L. Systematic Review and Meta-Analysis of the Significance of Heterogeneous Vancomycin-Intermediate Staphylococcus aureus Isolates. Antimicrob. Agents Chemother. 2011, 55, 405–410. [Google Scholar] [CrossRef] [PubMed]
- Hiramatsu, K.; Hanaki, H.; Ino, T.; Yabuta, K.; Oguri, T.; Tenover, F.C. Methicillin-resistant Staphylococcus aureus clinical strain with reduced vancomycin susceptibility. J. Antimicrob. Chemother. 1997, 40, 135–136. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Chambers, H.F. Staphylococcus aureus with Heterogeneous Resistance to Vancomycin: Epidemiology, Clinical Significance, and Critical Assessment of Diagnostic Methods. Antimicrob. Agents Chemother. 2003, 47, 3040–3045. [Google Scholar] [CrossRef] [PubMed]
- Hiramatsu, K.; Kayayama, Y.; Matsuo, M.; Aiba, Y.; Saito, M.; Hishinuma, T.; Iwamoto, A. Vancomycin-intermediate resistance in Staphylococcus aureus. J. Glob. Antimicrob. Resist. 2014, 2, 213–224. [Google Scholar] [CrossRef]
- Howden, B.P.; Johnson, P.D.R.; Ward, P.B.; Stinear, T.P.; Davies, J.K. Isolates with Low-Level Vancomycin Resistance Associated with Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia. Antimicrob. Agents Chemother. 2006, 50, 3039–3047. [Google Scholar] [CrossRef]
- Gaillard, T.; Dupieux-Chabert, C.; Butin, M.; Dumitrescu, O.; Naceur, O.; Bouveyron, C.; Martra, A.; Bes, M.; Tristan, A.; Vandenesch, F.; et al. Heterogeneous vancomycin resistance in Staphylococcus aureus does not predict development of vancomycin resistance upon vancomycin pressure. J. Antimicrob. Chemother. 2022, 77, 1032–1035. [Google Scholar] [CrossRef] [PubMed]
- Casapao, A.M.; Leonard, S.N.; Davis, S.L.; Lodise, T.P.; Patel, N.; Goff, D.A.; LaPlante, K.L.; Potoski, B.A.; Rybak, M.J. Clinical Outcomes in Patients with Heterogeneous Vancomycin-Intermediate Staphylococcus aureus Bloodstream Infection. Antimicrob. Agents Chemother. 2013, 57, 4252–4259. [Google Scholar] [CrossRef]
- Hu, H.C.; Kao, K.C.; Chiu, L.C.; Chang, C.H.; Hung, C.Y.; Li, L.F.; Liu, T.P.; Lin, L.C.; Chen, N.-H.; Huang, C.C.; et al. Clinical outcomes and molecular typing of heterogenous vancomycin-intermediate Staphylococcus aureus bacteremia in patients in intensive care units. BMC Infect. Dis. 2015, 15, 444. [Google Scholar] [CrossRef] [PubMed]
- Charles, P.G.P.; Ward, P.B.; Johnson, P.D.R.; Howden, B.P.; Grayson, M.L. Clinical Features Associated with Bacteremia Due to Heterogeneous Vancomycin-Intermediate Staphylococcus aureus. Clin. Infect. Dis. 2004, 38, 448–451. [Google Scholar] [CrossRef]
- Bae, I.; Federspiel, J.J.; Miró, J.M.; Woods, C.W.; Park, L.; Rybak, M.J.; Rude, T.H.; Bradley, S.; Bukovski, S.; de la Maria, C.G.; et al. Heterogeneous Vancomycin-Intermediate Susceptibility Phenotype in Bloodstream Methicillin-Resistant Staphylococcus aureus Isolates from an International Cohort of Patients with Infective Endocarditis: Prevalence, Genotype, and Clinical Significance. J. Infect. Dis. 2009, 200, 1355–1366. [Google Scholar] [CrossRef]
- Maor, Y.; Hagin, M.; Belausov, N.; Keller, N.; Ben-David, D.; Rahav, G. Clinical Features of Heteroresistant Vancomycin-Intermediate Staphylococcus aureus Bacteremia versus Those of Methicillin-Resistant S. Aureus Bacteremia. J. Infect. Dis. 2009, 199, 619–624. [Google Scholar] [CrossRef]
- Fong, R.K.C.; Low, J.; Koh, T.H.; Kurup, A. Clinical features and treatment outcomes of vancomycin-intermediate Staphylococcus aureus (VISA) and heteroresistant vancomycin-intermediate Staphylococcus aureus (hVISA) in a tertiary care institution in Singapore. Eur. J. Clin. Microbiol. Infect. Dis. 2009, 28, 983–987. [Google Scholar] [CrossRef]
- Lin, S.Y.; Chen, T.C.; Chen, F.J.; Chen, Y.H.; Lin, Y.I.; Siu, L.K.; Lu, P.L. Molecular epidemiology and clinical characteristics of hetero-resistant vancomycin intermediate Staphylococcus aureus bacteremia in a Taiwan Medical Center. J. Microbiol. Immunol. Infect. 2012, 45, 435–441. [Google Scholar] [CrossRef]
- Park, K.H.; Kim, E.S.; Kim, H.S.; Park, S.J.; Bang, K.M.; Park, H.J.; Park, S.Y.; Moon, S.M.; Chong, Y.P.; Kim, S.H.; et al. Comparison of the clinical features, bacterial genotypes and outcomes of patients with bacteraemia due to heteroresistant vancomycin-intermediate Staphylococcus aureus and vancomycin-susceptible S. aureus. J. Antimicrob. Chemother. 2012, 67, 1843–1849. [Google Scholar] [CrossRef]
- Maor, Y.; Belausov, N.; Ben-David, D.; Smollan, G.; Keller, N.; Rahav, G. hVISA and MRSA endocarditis: An 8-year experience in a tertiary care centre. Clin. Microbiol. Infect. 2014, 20, O730. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Yang, C.C.; Sy, C.L.; Huang, Y.C.; Shie, S.S.; Shu, J.C.; Hsieh, P.H.; Hsiao, C.H.; Chen, C.J. Risk factors of treatment failure and 30-day mortality in patients with bacteremia due to MRSA with reduced vancomycin susceptibility. Sci. Rep. 2018, 8, 7868. [Google Scholar] [CrossRef] [PubMed]
- van Hal, S.; Jones, M.; Gosbell, I.B.; Paterson, D.L. Vancomycin Heteroresistance Is Associated with Reduced Mortality in ST239 Methicillin-Resistant Staphylococcus aureus Blood Stream Infections. PLoS ONE 2011, 6, e21217. [Google Scholar] [CrossRef] [PubMed]
- Musta, A.C.; Riederer, K.; Shemes, S.; Chase, P.; Jose, J.; Johnson, L.B.; Khatib, R. Vancomycin MIC plus Heteroresistance and Outcome of Methicillin-Resistant Staphylococcus aureus Bacteremia: Trends over 11 Years. J. Clin. Microbiol. 2009, 47, 1640–1644. [Google Scholar] [CrossRef] [PubMed]
- Davis, J.S.; Petersiel, N.; Tong, S.Y. How I manage a patient with MRSA bacteraemia. Clin. Microbiol. Infect. 2021, 28, 190–194. [Google Scholar] [CrossRef] [PubMed]
- Sullenberger, A.L.; Avedissian, L.S.; Kent, S.M. Importance of transesophageal echocardiography in the evaluation of Staphylococcus aureus bacteremia. J. Heart Valve Dis. 2005, 14, 23–28. [Google Scholar] [PubMed]
- Fowler, V.G.; Li, J.; Corey, G.R.; Boley, J.; Marr, K.A.; Gopal, A.K.; Ryan, T. Role of echocardiography in evaluation of patients with Staphylococcus aureus bacteremia: Experience in 103 patients. J. Am. Coll. Cardiol. 1997, 30, 1072–1078. [Google Scholar] [CrossRef]
- Buis, D.T.P.; Sieswerda, E.; Kouijzer, I.J.E.; Huynh, W.Y.; Burchell, G.L.; Berrevoets, M.A.H.; Prins, J.M.; Sigaloff, K.C.E. [18F]FDG-PET/CT in Staphylococcus aureus bacteremia: A systematic review. BMC Infect. Dis. 2022, 22, 282. [Google Scholar] [CrossRef] [PubMed]
- Duval, X.; Le Moing, V.; Tubiana, S.; Esposito-Farèse, M.; Ilic-Habensus, E.; Leclercq, F.; Bourdon, A.; Goehringer, F.; Selton-Suty, C.; Chevalier, E.; et al. Impact of Systematic Whole-body 18F-Fluorodeoxyglucose PET/CT on the Management of Patients Suspected of Infective Endocarditis: The Prospective Multicenter TEPvENDO Study. Clin. Infect. Dis. 2020, 73, 393–403. [Google Scholar] [CrossRef]
- Tissot, F.; Calandra, T.; Prod’Hom, G.; Taffe, P.; Zanetti, G.; Greub, G.; Senn, L. Mandatory infectious diseases consultation for MRSA bacteremia is associated with reduced mortality. J. Infect. 2014, 69, 226–234. [Google Scholar] [CrossRef][Green Version]
- Cosgrove, S.E.; Vigliani, G.A.; Campion, M.; Fowler, J.V.G.; Abrutyn, E.; Corey, G.R.; Levine, D.P.; Rupp, M.E.; Chambers, H.F.; Karchmer, A.W.; et al. Initial Low-Dose Gentamicin for Staphylococcus aureus Bacteremia and Endocarditis Is Nephrotoxic. Clin. Infect. Dis. 2009, 48, 713–721. [Google Scholar] [CrossRef]
- Paul, M.; Bishara, J.; Yahav, D.; Goldberg, E.; Neuberger, A.; Ghanem-Zoubi, N.; Dickstein, Y.; Nseir, W.; Dan, M.; Leibovici, L. Trimethoprim-sulfamethoxazole versus vancomycin for severe infections caused by meticillin resistant Staphylococcus aureus: Randomised controlled trial. BMJ 2015, 350, h2219. [Google Scholar] [CrossRef]
- Thwaites, G.E.; Scarborough, M.; Szubert, A.; Nsutebu, E.; Tilley, R.; Greig, J.; Wyllie, S.A.; Wilson, P.; Auckland, C.; Cairns, J.; et al. Adjunctive rifampicin for Staphylococcus aureus bacteraemia (ARREST): A multicentre, randomised, double-blind, placebo-controlled trial. Lancet 2018, 391, 668–678. [Google Scholar] [CrossRef]
- Hageman, J.C.; Liedtke, L.A.; Sunenshine, R.H.; Strausbaugh, L.J.; McDonald, L.C.; Tenover, F.C.; Network, I.D.S.O.A.E.I. Management of Persistent Bacteremia Caused by Methicillin-Resistant Staphylococcus aureus: A Survey of Infectious Diseases Consultants. Clin. Infect. Dis. 2006, 43, e42–e45. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gasch, O.; Camoez, M.; Dominguez, M.A.; Padilla, B.; Pintado, V.; Almirante, B.; Martin, C.; Lopez-Medrano, F.; de Gopegui, E.R.; Blanco, J.R.; et al. Emergence of resistance to daptomycin in a cohort of patients with methicillin-resistant Staphylococcus aureus persistent bacteraemia treated with daptomycin. J. Antimicrob. Chemother. 2013, 69, 568–571. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Riederer, K.; Chase, P.; Khatib, R. High rate of decreasing daptomycin susceptibility during the treatment of persistent Staphylococcus aureus bacteremia. Eur. J. Clin. Microbiol. Infect. Dis. 2008, 27, 433–437. [Google Scholar] [CrossRef] [PubMed]
- Brown, N.M.; Goodman, A.L.; Horner, C.; Jenkins, A.; Brown, E.M. Treatment of methicillin-resistant Staphylococcus aureus (MRSA): Updated guidelines from the UK. JAC-Antimicrob. Resist. 2021, 3, dlaa114. [Google Scholar] [CrossRef] [PubMed]
- Habib, G.; Lancellotti, P.; Antunes, M.J.; Bongiorni, M.G.; Casalta, J.-P.; Del Zotti, F.; Dulgheru, R.; El Khoury, G.; Erba, P.A.; Iung, B.; et al. 2015 ESC Guidelines for the management of infective endocarditis: The Task Force for the Management of Infective Endocarditis of the European Society of Cardiology (ESC). Endorsed by: European Association for Cardio-Thoracic Surgery (EACTS), the European Association of Nuclear Medicine (EANM). Eur. Heart J. 2015, 36, 3075–3128. [Google Scholar] [CrossRef] [PubMed]
- Tong, S.Y.C.; Lye, D.C.; Yahav, D.; Sud, A.; Robinson, J.O.; Nelson, J.; Archuleta, S.; Roberts, M.A.; Cass, A.; Paterson, D.L.; et al. Effect of Vancomycin or Daptomycin with vs. without an Antistaphylococcal β-Lactam on Mortality, Bacteremia, Relapse, or Treatment Failure in Patients with MRSA Bacteremia: A Randomized Clinical Trial. JAMA 2020, 323, 527–537. [Google Scholar] [CrossRef]
- Rose, W.; Fantl, M.; Geriak, M.; Nizet, V.; Sakoulas, G. Current Paradigms of Combination Therapy in Methicillin-Resistant Staphylococcus aureus (MRSA) Bacteremia: Does it Work, Which Combination, and for Which Patients? Clin. Infect. Dis. 2021, 73, 2353–2360. [Google Scholar] [CrossRef]
- Cheng, M.P.; Lawandi, A.; Butler-Laporte, G.; De L’Étoile-Morel, S.; Paquette, K.; Lee, T.C. Adjunctive Daptomycin in the Treatment of Methicillin-susceptible Staphylococcus aureus Bacteremia: A Randomized, Controlled Trial. Clin. Infect. Dis. 2020, 72, e196–e203. [Google Scholar] [CrossRef]
- Moisan, H.; Pruneau, M.; Malouin, F. Binding of ceftaroline to penicillin-binding proteins of Staphylococcus aureus and Streptococcus pneumoniae. J. Antimicrob. Chemother. 2010, 65, 713–716. [Google Scholar] [CrossRef]
- Geriak, M.; Haddad, F.; Rizvi, K.; Rose, W.; Kullar, R.; LaPlante, K.; Yu, M.; Vasina, L.; Ouellette, K.; Zervos, M.; et al. Clinical Data on Daptomycin plus Ceftaroline versus Standard of Care Monotherapy in the Treatment of Methicillin-Resistant Staphylococcus aureus Bacteremia. Antimicrob. Agents Chemother. 2019, 63, e02483-18. [Google Scholar] [CrossRef]
- McCreary, E.K.; Kullar, R.; Geriak, M.; Zasowski, E.J.; Rizvi, K.; Schulz, L.T.; Ouellette, K.; Vasina, L.; Haddad, F.; Rybak, M.J.; et al. Multicenter Cohort of Patients with Methicillin-Resistant Staphylococcus aureus Bacteremia Receiving Daptomycin Plus Ceftaroline Compared with Other MRSA Treatments. Open Forum Infect. Dis. 2019, 7, ofz538. [Google Scholar] [CrossRef]
- Huang, C.; Chen, I.; Lin, L. Comparing the Outcomes of Ceftaroline plus Vancomycin or Daptomycin Combination Therapy versus Vancomycin or Daptomycin Monotherapy in Adults with Methicillin-Resistant Staphylococcus aureus Bacteremia—A Meta-Analysis. Antibiotics 2022, 11, 1104. [Google Scholar] [CrossRef]
- Hornak, J.P.; Anjum, S.; Reynoso, D. Adjunctive ceftaroline in combination with daptomycin or vancomycin for complicated methicillin-resistant Staphylococcus aureus bacteremia after monotherapy failure. Ther. Adv. Infect. Dis. 2019, 6, 2049936119886504. [Google Scholar] [CrossRef] [PubMed]
- Burnett, Y.J.; Echevarria, K.; Traugott, K.A. Ceftaroline as Salvage Monotherapy for Persistent MRSA Bacteremia. Ann. Pharmacother. 2016, 50, 1051–1059. [Google Scholar] [CrossRef]
- Sakoulas, G.; Moise, P.A.; Casapao, A.M.; Nonejuie, P.; Olson, J.; Okumura, C.Y.; Rybak, M.J.; Kullar, R.; Dhand, A.; Rose, W.E.; et al. Antimicrobial Salvage Therapy for Persistent Staphylococcal Bacteremia Using Daptomycin Plus Ceftaroline. Clin. Ther. 2014, 36, 1317–1333. [Google Scholar] [CrossRef] [PubMed]
- Cortes-Penfield, N.; Oliver, N.T.; Hunter, A.; Rodriguez-Barradas, M. Daptomycin and combination daptomycin-ceftaroline as salvage therapy for persistent methicillin-resistant Staphylococcus aureus bacteremia. Infect. Dis. 2018, 50, 643–647. [Google Scholar] [CrossRef] [PubMed]
- Gritsenko, D.; Fedorenko, M.; Ruhe, J.J.; Altshuler, J. Combination Therapy with Vancomycin and Ceftaroline for Refractory Methicillin-resistant Staphylococcus aureus Bacteremia: A Case Series. Clin. Ther. 2016, 39, 212–218. [Google Scholar] [CrossRef] [PubMed]
- Tattevin, P.; Boutoille, D.; Vitrat, V.; Van Grunderbeeck, N.; Revest, M.; Dupont, M.; Alfandari, S.; Stahl, J.-P. Salvage treatment of methicillin-resistant staphylococcal endocarditis with ceftaroline: A multicentre observational study. J. Antimicrob. Chemother. 2014, 69, 2010–2013. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.T.; Cadena, J.; Childs, L.M.; Gonzalez-Velez, M.; Lewis, I.J.S. Methicillin-resistant Staphylococcus aureus bacteraemia and endocarditis treated with ceftaroline salvage therapy. J. Antimicrob. Chemother. 2012, 67, 1267–1270. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Strnad, L.; Beekmann, S.E.; Polgreen, P.M.; Chambers, H.F. Clinical Practice Variation Among Adult Infectious Disease Physicians in the Management of Staphylococcus aureus Bacteremia. Clin. Infect. Dis. 2019, 69, 530–533. [Google Scholar] [CrossRef] [PubMed]
- Rose, W.E.; Rybak, M.J.; Kaatz, G.W. Evaluation of daptomycin treatment of Staphylococcus aureus bacterial endocarditis: An in vitro and in vivo simulation using historical and current dosing strategies. J. Antimicrob. Chemother. 2007, 60, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Pfaller, M.; Flamm, R.K.; Duncan, L.R.; Shortridge, D.; Smart, J.I.; Hamed, K.; Mendes, R.; Sader, H. Ceftobiprole activity when tested against contemporary bacteria causing bloodstream infections in the United States (2016–2017). Diagn. Microbiol. Infect. Dis. 2019, 94, 304–313. [Google Scholar] [CrossRef] [PubMed]
- Tattevin, P.; Basuino, L.; Bauer, D.; Diep, B.A.; Chambers, H.F. Ceftobiprole Is Superior to Vancomycin, Daptomycin, and Linezolid for Treatment of Experimental Endocarditis in Rabbits Caused by Methicillin-Resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 2010, 54, 610–613. [Google Scholar] [CrossRef] [PubMed]
- Hamed, K.; Engelhardt, M.; Jones, M.E.; Saulay, M.; Holland, T.L.; Seifert, H.; Jr, V.G.F. Ceftobiprole versus daptomycin in Staphylococcus aureus bacteremia: A novel protocol for a double-blind, Phase III trial. Futur. Microbiol. 2020, 15, 35–48. [Google Scholar] [CrossRef] [PubMed]
- Boucher, H.W.; Wilcox, M.; Talbot, G.H.; Puttagunta, S.; Das, A.F.; Dunne, M.W. Once-Weekly Dalbavancin versus Daily Conventional Therapy for Skin Infection. N. Engl. J. Med. 2014, 370, 2169–2179. [Google Scholar] [CrossRef]
- Cooper, M.M.; Preslaski, C.R.; Shihadeh, K.C.; Hawkins, K.L.; Jenkins, T.C. Multiple-Dose Dalbavancin Regimens as the Predominant Treatment of Deep-Seated or Endovascular Infections: A Scoping Review. Open Forum Infect. Dis. 2021, 8, ofab486. [Google Scholar] [CrossRef]
- Turner, N.A.; Zaharoff, S.; King, H.; Evans, S.; Hamasaki, T.; Lodise, T.; Ghazaryan, V.; Beresnev, T.; Riccobene, T.; Patel, R.; et al. Dalbavancin as an option for treatment of S. aureus bacteremia (DOTS): Study protocol for a phase 2b, multicenter, randomized, open-label clinical trial. Trials 2022, 23, 1–15. [Google Scholar] [CrossRef]
- Fowler, V.G., Jr.; Das, A.F.; Lipka-Diamond, J.; Schuch, R.; Pomerantz, R.; Jáuregui-Peredo, L.; Bressler, A.; Evans, D.; Moran, G.J.; Rupp, M.E.; et al. Exebacase for patients with Staphylococcus aureus bloodstream infection and endocarditis. J. Clin. Investig. 2020, 130, 3750–3760. [Google Scholar] [CrossRef]
- Clinicaltrials.gov. Direct Lysis of Staph Aureus Resistant Pathogen Trial of Exebacase (DISRUPT). Available online: https://clinicaltrials.gov/ct2/show/NCT04160468 (accessed on 15 January 2023).
- Wire, M.B.; Jun, S.Y.; Jang, I.-J.; Lee, S.-H.; Hwang, J.G.; Huang, D.B. A Phase 1 Study To Evaluate Safety and Pharmacokinetics following Administration of Single and Multiple Doses of the Antistaphylococcal Lysin LSVT-1701 in Healthy Adult Subjects. Antimicrob. Agents Chemother. 2022, 66, e01842-21. [Google Scholar] [CrossRef]
- Petrovic Fabijan, A.; Lin, R.C.Y.; Ho, J.; Maddocks, S.; Ben Zakour, N.L.; Iredell, J.R. Safety of bacteriophage therapy in severe Staphylococcus aureus infection. Nat. Microbiol. 2020, 5, 465–472. [Google Scholar] [CrossRef]
- Clinicaltrials.gov. Study Evaluating Safety, Tolerability, and Efficacy of Intravenous AP-SA02 in Subjects with S. aureus Bacteremia (diSArm). Available online: https://clinicaltrials.gov/ct2/show/NCT05184764 (accessed on 15 January 2023).
| Study | Year | MSSA or MRSA | Definition of Persistent Bacteremia | Clinical Risk Factors Identified |
|---|---|---|---|---|
| Khatib et al. [26] | 2006 | MSSA and MRSA | 3 days |
|
| Hawkins et al. [25] | 2007 | MSSA and MRSA | 7 days |
|
| Khatib et al. [23] | 2009 | MSSA and MRSA | 7 days |
|
| Ganga et al. [28] | 2009 | MRSA and MSSA | 7 days |
|
| Yoon et al. [24] | 2010 | MRSA | 7 days |
|
| Chong et al. [22] | 2013 | MSSA and MRSA | 7 days |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Parsons, J.B.; Westgeest, A.C.; Conlon, B.P.; Fowler, V.G., Jr. Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia: Host, Pathogen, and Treatment. Antibiotics 2023, 12, 455. https://doi.org/10.3390/antibiotics12030455
Parsons JB, Westgeest AC, Conlon BP, Fowler VG Jr. Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia: Host, Pathogen, and Treatment. Antibiotics. 2023; 12(3):455. https://doi.org/10.3390/antibiotics12030455
Chicago/Turabian StyleParsons, Joshua B., Annette C. Westgeest, Brian P. Conlon, and Vance G. Fowler, Jr. 2023. "Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia: Host, Pathogen, and Treatment" Antibiotics 12, no. 3: 455. https://doi.org/10.3390/antibiotics12030455
APA StyleParsons, J. B., Westgeest, A. C., Conlon, B. P., & Fowler, V. G., Jr. (2023). Persistent Methicillin-Resistant Staphylococcus aureus Bacteremia: Host, Pathogen, and Treatment. Antibiotics, 12(3), 455. https://doi.org/10.3390/antibiotics12030455







