Carbapenem-Resistant Klebsiella pneumoniae: Virulence Factors, Molecular Epidemiology and Latest Updates in Treatment Options
Abstract
1. Introduction
2. Genomic Pool
3. Virulence Factors and Molecular Pathogenesis
4. Mechanisms of Antimicrobial Resistance
- (a)
- (b)
- decreased cell permeability through loss of Omps [81].
- (c)
- overexpression of efflux pumps, which are transmembrane proteins, with the antimicrobial agent being usually excreted out of the bacterial cell through an energy-consuming process. For example, an efflux pump called KpnGH contributes to antimicrobial resistance in K. pneumoniae [82].
- (d)
- modification of the target of the antimicrobial agent [83].
4.1. B-Lactams—Ambler Classification of β-Lactamases
4.2. Decreased Cell Permeability through Loss of Omps
4.3. Transport of Antimicrobial Resistance Genes
5. Trends in Molecular Epidemiology
6. Trends in Antimicrobial Treatment
6.1. Colistin
6.2. Tigecycline
6.3. Fosfomycin
| Antimicrobial | PubChem CID | Molecular Formula | Synonyms | Structure | Mode of Action |
|---|---|---|---|---|---|
| Colistin | 44144393 | C52H98N16O13 | Polymyxin E | 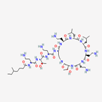 | Polycationic peptides which targets bacterial (in particular, Gram-negative bacteria) cell membrane to disrupt it through detergent-like mechanism. |
| Tigecycline | 54686904 | C29H39N5O8 | Tygacil |  | A member of the new class of glycylcyclines. Indeed, glycylcyclines are derived from tetracyclines. Tigecycline targets the ribosomal small subunit of 30S (with higher affinity than tetracyclines) to prevent bacterial protein translation. The attachment of tigecycline to the amino-acyl tRNA molecule inhibits the entrance of the amino-acyl tRNA molecule into the A site of the ribosome to stop the elongation process of the bacterial peptide biosynthesis |
| Fosfomycin | 446987 | C3H7O4P | Phosphomycin, Phosphonomycin | 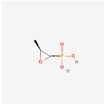 | Fosfomycin is used against bacterial strains with the property of difficult-to-treat. This antibiotic is the first option against UTIs. Fosfomycin inactivates the UDP-N-acetylglucosamine enolpyruvyl transferase (MurA) enzyme via binding to a cysteine residue of the enzyme’s active site. This process results in prevention of peptidoglycan precursor UDP N-acetylmuramic acid (UDP-MurNAc) biosynthesis and in consequence may lead to stopping bacterial cell wall biosynthesis. Therefore, fosfomycin has a bactericidal effect on pathogens. |
| Plazomicin | 42613186 | C25H48N6O10 | Zemdri | 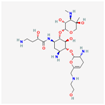 | Plazomicin, as a member of aminoglycoside antibiotics, has bactericidal effect through binding to ribosomal small subunit of 30S. This antibiotic changes the spatial structure of the ribosomal A-site (aminoacyl-tRNA site), which may lead to attachment of antibiotic to rRNA molecule. This feature results in mistranslation of mRNA molecules within the process of protein biosynthesis. |
| Eravacycline | 54726192 | C27H31FN4O8 | Xerava |  | Eravacycline as a fluorocycline antibacterial pertaining to tetracycline class has disruptive effect on bacterial pathogens through targeting their protein biosynthesis processes. This effect is achieved via targeting the ribosomal small subunit of 30S. |
| Cefiderocol | 77843966 | C30H34ClN7O10S2 | SZ34OMG6E8 | 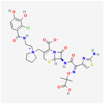 | As a cephalosporin drug, has bactericidal effect on aerobic Gram-negative bacteria such as K.pneumoniae. Cefiderocol binds to penicillin-binding proteins (PBPs) (in particular with PBP3 and in general with PBP1a, PBP1b, PBP2 and PBP4), inactivating their activities which may lead to inhibition of bacterial cell wall biosynthesis. |
| Temocillin | 171758 | C16H18N2O7S2 | Negaban | 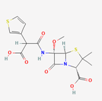 | Temocillin acts as inhibitor against bacterial reproduction and growth processes. |
| Ceftolozan–-tazobactam | 86291594 | C33H42N16O13S3 | Zerbaxa | 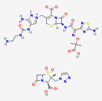 | Zerbaxa prevents the growth and reproduction process in bacterial pathogens and has bactericidal effect on UTIs’ bacterial causative agents. |
| Imipenem–cilastatin | 17756656 | C28H43N5O9S2 | Thienam | 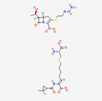 | Imipenem–cilastatin prevents the growth and reproduction process in bacterial pathogens. Thienam prevents/antagonizes the process of biosynthesis/actions of the enzymes of bacterial proteases. |
| Meropenem–vaborbactam | 86298703 | C29H41BN4O10S2 | Carbavance | 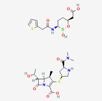 | Vaborbactam acts as bacterial serine-ß-lactamases to support the antibacterial effect of penem drugs, e.g., meropenem against CRCKP strains of K.pneumoniae, etc. |
| Ceftazidime–avibactam | 90643431 | C29H33N9O13S3 | Avycaz | 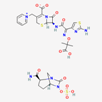 | Avycaz prevents the growth and reproduction process in bacterial pathogens. Avibactam prevents/blocks the activities of bacterial ß-lactamases. |
6.4. Plazomicin
6.5. Eravacycline
6.6. Cefiderocol
6.7. Temocillin
6.8. Ceftolozane–Tazobactam
6.9. Imipenem–Cilastatin/Relebactam
6.10. Meropenem–Vaborbactam
6.11. Ceftazidime–Avibactam
6.12. Aztreonam–Avibactam
6.13. Guidelines for the Treatment of CRKP Infections
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Katsiari, M.; Panagiota, G.; Likousi, S.; Roussou, Z.; Polemis, M.; Vatopoulos, C.A.; Platsouka, D.E.; Maguina, A. Carbapenem-resistant Klebsiella pneumoniae infections in a Greek intensive care unit: Molecular characterisation and treatment challenges. J. Glob. Antimicrob. Resist. 2015, 3, 123–127. [Google Scholar] [CrossRef] [PubMed]
- Zhen, X.; Stalsby Lundborg, C.; Sun, X.; Gu, S.; Dong, H. Clinical and Economic Burden of Carbapenem-Resistant Infection or Colonization Caused by Klebsiella pneumoniae, Pseudomonas aeruginosa, Acinetobacter baumannii: A Multicenter Study in China. Antibiotics 2020, 9, 514. [Google Scholar] [CrossRef] [PubMed]
- Ahmadi, Z.; Noormohammadi, Z.; Behzadi, P.; Ranjbar, R. Molecular Detection of gyrA Mutation in Clinical Strains of Klebsiella pneumoniae. Iran. J. Public Health 2022, 51, 2334–2339. [Google Scholar] [CrossRef] [PubMed]
- Sarshar, M.; Behzadi, P.; Ambrosi, C.; Zagaglia, C.; Palamara, A.T.; Scribano, D. FimH and Anti-Adhesive Therapeutics: A Disarming Strategy Against Uropathogens. Antibiotics 2020, 9, 397. [Google Scholar] [CrossRef] [PubMed]
- Martin, R.M.; Bachman, M.A. Colonization, Infection, and the Accessory Genome of Klebsiella pneumoniae. Front. Cell. Infect. Microbiol. 2018, 8, 4. [Google Scholar] [CrossRef]
- Cassini, A.; Högberg, L.D.; Plachouras, D.; Quattrocchi, A.; Hoxha, A.; Simonsen, G.S.; Colomb-Cotinat, M.; Kretzschmar, M.E.; Devleesschauwer, B.; Cecchini, M.; et al. Attributable deaths and disability-adjusted life-years caused by infections with antibiotic-resistant bacteria in the EU and the European Economic Area in 2015: A population-level modelling analysis. Lancet Infect. Dis. 2019, 19, 56–66. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control (2019) Surveillance Atlas of Infectious Diseases. Available online: https://atlas.ecdc.europa.eu/public/index.aspx?Dataset=27&HealthTopic=4 (accessed on 30 June 2022).
- Tesfa, T.; Mitiku, H.; Edae, M.; Assefa, N. Prevalence and incidence of carbapenem-resistant K. pneumoniae colonization: Systematic review and meta-analysis. Syst. Rev. 2022, 11, 240. [Google Scholar] [CrossRef]
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olsson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Petrosillo, N.; Giannella, M.; Lewis, R.; Viale, P. Treatment of carbapenem-resistant Klebsiella pneumoniae: The state of the art. Expert Rev. Anti Infect. Ther. 2013, 11, 159–177. [Google Scholar] [CrossRef]
- Gonzalez-Ferrer, S.; Peñaloza, H.F.; Budnick, J.A.; Bain, W.G.; Nordstrom, H.R.; Lee, J.S.; Van Tyne, D. Finding Order in the Chaos: Outstanding Questions in Klebsiella pneumoniae Pathogenesis. Infect. Immun. 2021, 89, e00693-20. [Google Scholar] [CrossRef]
- Baker, J.L.; Hendrickson, E.L.; Tang, X.; Lux, R.; He, X.; Edlund, A.; McLean, J.S.; Shi, W. Klebsiella and Providencia emerge as lone survivors following long-term starvation of oral microbiota. Proc. Natl. Acad. Sci. USA 2019, 116, 8499–8504. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, P.F.; Lu, Y.R.; Lin, T.L.; Lai, L.Y.; Wang, J.T. Klebsiella pneumoniae Type VI Secretion System Contributes to Bacterial Competition, Cell Invasion, Type-1 Fimbriae Expression, and In Vivo Colonization. J. Infect. Dis. 2019, 219, 637–647. [Google Scholar] [CrossRef] [PubMed]
- Behzadi, P. Classical chaperone-usher (CU) adhesive fimbriome: Uropathogenic Escherichia coli (UPEC) and urinary tract infections (UTIs). Folia Microbiol. 2020, 65, 45–65. [Google Scholar] [CrossRef]
- Dunn, S.J.; Connor, C.; McNally, A. The evolution and transmission of multi-drug resistant Escherichia coli and Klebsiella pneumoniae: The complexity of clones and plasmids. Curr. Opin. Microbiol. 2019, 51, 51–56. [Google Scholar] [CrossRef] [PubMed]
- Holt, K.E.; Wertheim, H.; Zadoks, R.N.; Baker, S.; Whitehouse, C.A.; Dance, D.; Jenney, A.; Connor, T.R.; Hsu, L.Y.; Severin, J.; et al. Genomic analysis of diversity, population structure, virulence, and antimicrobial resistance in Klebsiella pneumoniae, an urgent threat to public health. Proc. Natl. Acad. Sci. USA 2015, 112, E3574–E3581. [Google Scholar] [CrossRef]
- Wyres, K.L.; Lam, M.M.C.; Holt, K.E. Population genomics of Klebsiella pneumoniae. Nat. Rev. Microbiol. 2020, 18, 344–359. [Google Scholar] [CrossRef]
- Jahandeh, N.; Ranjbar, R.; Behzadi, P.; Behzadi, E. Uropathogenic Escherichia coli virulence genes: Invaluable approaches for designing DNA microarray probes. Cent. Eur. J. Urol. 2015, 68, 452–458. [Google Scholar]
- Hennart, M.; Guglielmini, J.; Bridel, S.; Maiden, M.C.; Jolley, K.A.; Criscuolo, A.; Brisse, S. A Dual Barcoding Approach to Bacterial Strain Nomenclature: Genomic Taxonomy of Klebsiella pneumoniae Strains. Mol. Biol. Evol. 2022, 39, msac135. [Google Scholar] [CrossRef]
- Maiden, M.C.J.; Van Rensburg, M.J.J.; Bray, J.; Earle, S.G.; Ford, S.A.; Jolley, K.; McCarthy, N.D. MLST revisited: The gene-by-gene approach to bacterial genomics. Nat. Rev. Microbiol. 2013, 11, 728–736. [Google Scholar] [CrossRef]
- Moura, A.; Criscuolo, A.; Pouseele, H.; Maury, M.M.; Leclercq, A.; Tarr, C.; Björkman, J.T.; Dallman, T.; Reimer, A.; Enouf, V.; et al. Whole genome-based population biology and epidemiological surveillance of Listeria monocytogenes. Nat. Microbiol. 2016, 2, 16185. [Google Scholar] [CrossRef]
- Lam, M.M.C.; Wick, R.R.; Watts, S.C.; Cerdeira, L.T.; Wyres, K.L.; Holt, K.E. A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex. Nat. Commun. 2021, 12, 4188. [Google Scholar] [CrossRef] [PubMed]
- Ali, S.; Alam, M.; Hasan, G.M.; Hassan, M.I. Potential therapeutic targets of Klebsiella pneumoniae: A multi-omics review perspective. Brief. Funct. Genom. 2022, 21, 63–77. [Google Scholar] [CrossRef] [PubMed]
- Issakhanian, L.; Behzadi, P. Antimicrobial Agents and Urinary Tract Infections. Curr. Pharm. Des. 2019, 25, 1409–1423. [Google Scholar] [CrossRef] [PubMed]
- Behzadi, P.; Garcia-Perdomo, H.A.; Karpinski, T.M.; Issakhanian, L. Metallo-ss-lactamases: A review. Mol. Biol. Rep. 2020, 47, 6281–6294. [Google Scholar] [CrossRef]
- Doorduijn, D.J.; Rooijakkers, S.H.; van Schaik, W.; Bardoel, B.W. Complement resistance mechanisms of Klebsiella pneumoniae. Immunobiology 2016, 221, 1102–1109. [Google Scholar] [CrossRef]
- Paczosa, M.K.; Mecsas, J. Klebsiella pneumoniae: Going on the Offense with a Strong Defense. Microbiol. Mol. Biol. Rev. MMBR 2016, 80, 629–661. [Google Scholar] [CrossRef]
- Koren, J.; Andrezal, M.; Drahovska, H.; Hubenakova, Z.; Liptakova, A.; Maliar, T. Next-Generation Sequencing of Carbapenem-Resistant Klebsiella pneumoniae Strains Isolated from Patients Hospitalized in the University Hospital Facilities. Antibiotics 2022, 11, 1538. [Google Scholar] [CrossRef]
- Foldes, A.; Oprea, M.; Szekely, E.; Usein, C.R.; Dobreanu, M. Characterization of Carbapenemase-Producing Klebsiella pneumoniae Isolates from Two Romanian Hospitals Co-Presenting Resistance and Heteroresistance to Colistin. Antibiotics 2022, 11, 1171. [Google Scholar] [CrossRef]
- Conceicao-Neto, O.C.; da Costa, B.S.; Pontes, L.D.S.; Silveira, M.C.; Justo-da-Silva, L.H.; de Oliveira Santos, I.C.; Teixeira, C.B.T.; Oliveira, T.R.T.e; Hermes, F.S.; Galvão, T.C.; et al. Polymyxin Resistance in Clinical Isolates of K. pneumoniae in Brazil: Update on Molecular Mechanisms, Clonal Dissemination and Relationship With KPC-Producing Strains. Front. Cell. Infect. Microbiol. 2022, 12, 898125. [Google Scholar] [CrossRef]
- Dong, N.; Yang, X.; Chan, E.W.; Zhang, R.; Chen, S. Klebsiella species: Taxonomy, hypervirulence and multidrug resistance. EBioMedicine 2022, 79, 103998. [Google Scholar] [CrossRef]
- Wyres, K.L.; Holt, K.E. Klebsiella pneumoniae Population Genomics and Antimicrobial-Resistant Clones. Trends Microbiol. 2016, 24, 944–956. [Google Scholar] [CrossRef] [PubMed]
- Dong, N.; Zhang, R.; Liu, L.; Li, R.; Lin, D.; Chan, E.W.-C.; Chen, S. Genome analysis of clinical multilocus sequence Type 11 Klebsiella pneumoniae from China. Microb. Genom. 2018, 4, e000149. [Google Scholar] [CrossRef] [PubMed]
- Wyres, K.L.; Wick, R.R.; Judd, L.M.; Froumine, R.; Tokolyi, A.; Gorrie, C.L.; Lam, M.M.C.; Duchêne, S.; Jenney, A.; Holt, K.E. Distinct evolutionary dynamics of horizontal gene transfer in drug resistant and virulent clones of Klebsiella pneumoniae. PLoS Genet. 2019, 15, e1008114. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.; Mathema, B.; Chavda, K.D.; DeLeo, F.R.; Bonomo, R.A.; Kreiswirth, B.N. Carbapenemase-producing Klebsiella pneumoniae: Molecular and genetic decoding. Trends Microbiol. 2014, 22, 686–696. [Google Scholar] [CrossRef]
- Russo, T.A.; Olson, R.; Fang, C.-T.; Stoesser, N.; Miller, M.; MacDonald, U.; Hutson, A.; Barker, J.H.; La Hoz, R.M.; Johnson, J.R.; et al. Identification of Biomarkers for Differentiation of Hypervirulent Klebsiella pneumoniae from Classical K. pneumoniae. J. Clin. Microbiol. 2018, 56, e00776-18. [Google Scholar] [CrossRef] [PubMed]
- Tian, D.; Wang, M.; Zhou, Y.; Hu, D.; Ou, H.Y.; Jiang, X. Genetic diversity and evolution of the virulence plasmids encoding aerobactin and salmochelin in Klebsiella pneumoniae. Virulence 2021, 12, 1323–1333. [Google Scholar] [CrossRef]
- Tian, D.; Liu, X.; Chen, W.; Zhou, Y.; Hu, D.; Wang, W.; Wu, J.; Mu, Q.; Jiang, X. Prevalence of hypervirulent and carbapenem-resistant Klebsiella pneumoniae under divergent evolutionary patterns. Emerg. Microbes Infect. 2022, 11, 1936–1949. [Google Scholar] [CrossRef]
- Lam, M.M.C.; Wyres, K.L.; Duchêne, S.; Wick, R.R.; Judd, L.M.; Gan, Y.-H.; Hoh, C.-H.; Archuleta, S.; Molton, J.S.; Kalimuddin, S.; et al. Population genomics of hypervirulent Klebsiella pneumoniae clonal-group 23 reveals early emergence and rapid global dissemination. Nat. Commun. 2018, 9, 2703. [Google Scholar] [CrossRef]
- Lan, P.; Jiang, Y.; Zhou, J.; Yu, Y. A global perspective on the convergence of hypervirulence and carbapenem resistance in Klebsiella pneumoniae. J. Glob. Antimicrob. Resist. 2021, 25, 26–34. [Google Scholar] [CrossRef]
- Zhang, N.; Qi, L.; Liu, X.; Jin, M.; Jin, Y.; Yang, X.; Chen, J.; Qin, S.; Liu, F.; Tang, Y.; et al. Clinical and Molecular Characterizations of Carbapenem-Resistant Klebsiella pneumoniae Causing Bloodstream Infection in a Chinese Hospital. Microbiol. Spectr. 2022, 10, e0169022. [Google Scholar] [CrossRef]
- Wyres, K.L.; Nguyen, T.N.T.; Lam, M.M.C.; Judd, L.M.; van Vinh Chau, N.; Dance, D.A.B.; Ip, M.; Karkey, A.; Ling, C.L.; Miliya, T.; et al. Genomic surveillance for hypervirulence and multi-drug resistance in invasive Klebsiella pneumoniae from South and Southeast Asia. Genome Med. 2020, 12, 11. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Jiang, X.; Yang, T.; Ju, Y.; Yin, Z.; Yue, L.; Ma, G.; Wang, X.; Jing, Y.; Luo, X.; et al. Genomic epidemiology of carbapenemase-producing Klebsiella pneumoniae in china. Genom. Proteom. Bioinform. 2022. [Google Scholar] [CrossRef] [PubMed]
- Munoz-Price, L.S.; Poirel, L.; Bonomo, R.A.; Schwaber, M.J.; Daikos, G.L.; Cormican, M.; Cornaglia, G.; Garau, J.; Gniadkowski, M.; Hayden, M.K.; et al. Clinical epidemiology of the global expansion of Klebsiella pneumoniae carbapenemases. Lancet Infect. Dis. 2013, 13, 785–796. [Google Scholar] [CrossRef] [PubMed]
- Naas, T.; Cuzon, G.; Villegas, M.V.; Lartigue, M.F.; Quinn, J.P.; Nordmann, P. Genetic structures at the origin of acquisition of the beta-lactamase bla KPC gene. Antimicrob. Agents Chemother. 2008, 52, 1257–1263. [Google Scholar] [CrossRef]
- Lee, I.R.; Molton, J.; Wyres, K.; Gorrie, C.; Wong, J.; Hoh, C.H.; Teo, J.; Kalimuddin, S.; Lye, D.; Archuleta, S.; et al. Differential host susceptibility and bacterial virulence factors driving Klebsiella liver abscess in an ethnically diverse population. Sci. Rep. 2016, 6, 29316. [Google Scholar] [CrossRef]
- Marr, C.M.; Russo, T.A. Hypervirulent Klebsiella pneumoniae: A new public health threat. Expert Rev. Anti-Infect. Ther. 2019, 17, 71–73. [Google Scholar] [CrossRef]
- Struve, C.; Roe, C.C.; Stegger, M.; Stahlhut, S.G.; Hansen, D.S.; Engelthaler, D.M.; Andersen, P.S.; Driebe, E.M.; Keim, P.; Krogfelt, K.A. Mapping the Evolution of Hypervirulent Klebsiella pneumoniae. mBio 2015, 6, e00630. [Google Scholar] [CrossRef]
- Huang, X.; Li, X.; An, H.; Wang, J.; Ding, M.; Wang, L.; Li, L.; Ji, Q.; Qu, F.; Wang, H.; et al. Capsule type defines the capability of Klebsiella pneumoniae in evading Kupffer cell capture in the liver. PLoS Pathog. 2022, 18, e1010693. [Google Scholar] [CrossRef]
- Wang, G.; Zhao, G.; Chao, X.; Xie, L.; Wang, H. The Characteristic of Virulence, Biofilm and Antibiotic Resistance of Klebsiella pneumoniae. Int. J. Environ. Res. Public Health 2020, 17, 6278. [Google Scholar] [CrossRef]
- Cubero, M.; Grau, I.; Tubau, F.; Pallares, R.; Dominguez, M.A.; Linares, J.; Ardanuy, C. Hypervirulent Klebsiella pneumoniae clones causing bacteraemia in adults in a teaching hospital in Barcelona, Spain (2007–2013). Clin. Microbiol. Infect. 2016, 22, 154–160. [Google Scholar] [CrossRef]
- Russo, T.A.; Marr, C.M. Hypervirulent Klebsiella pneumoniae. Clin. Microbiol. Rev. 2019, 32, e00001-19. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.H.Y.; Porto, W.F.; Jr, C.d.F.; Dias, S.C.; Alencar, S.A.; Pickard, D.J.; Hancock, R.E.W.; Franco, O.L. Genomic insights into the diversity, virulence and resistance of Klebsiella pneumoniae extensively drug resistant clinical isolates. Microb. Genom. 2021, 7, 000613. [Google Scholar] [CrossRef]
- Dai, P.; Hu, D. The making of hypervirulent Klebsiella pneumoniae. J. Clin. Lab. Anal. 2022, 36, e24743. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Zhao, Y.; Liu, C.; Chen, Z.; Zhou, D. Molecular pathogenesis of Klebsiella pneumoniae. Future Microbiol. 2014, 9, 1071–1081. [Google Scholar] [CrossRef]
- Ahmadi, M.; Ranjbar, R.; Behzadi, P.; Mohammadian, T. Virulence factors, antibiotic resistance patterns, and molecular types of clinical isolates of Klebsiella Pneumoniae. Expert Rev. Anti-Infect. Ther. 2022, 20, 463–472. [Google Scholar] [CrossRef] [PubMed]
- Modified. Available online: Biorender.com (accessed on 20 December 2022).
- Pan, Y.-J.; Lin, T.-L.; Chen, C.-T.; Chen, Y.-Y.; Hsieh, P.-F.; Hsu, C.-R.; Wu, M.-C.; Wang, J.-T. Genetic analysis of capsular polysaccharide synthesis gene clusters in 79 capsular types of Klebsiella spp. Sci. Rep. 2015, 5, 15573. [Google Scholar] [CrossRef] [PubMed]
- Whitfield, C.; Wear, S.S.; Sande, C. Assembly of Bacterial Capsular Polysaccharides and Exopolysaccharides. Annu. Rev. Microbiol. 2020, 74, 521–543. [Google Scholar] [CrossRef]
- Chuang, Y.P.; Fang, C.T.; Lai, S.Y.; Chang, S.C.; Wang, J.T. Genetic determinants of capsular serotype K1 of Klebsiella pneumoniae causing primary pyogenic liver abscess. J. Infect. Dis. 2006, 193, 645–654. [Google Scholar] [CrossRef]
- Walker, K.A.; Miller, V.L. The intersection of capsule gene expression, hypermucoviscosity and hypervirulence in Klebsiella pneumoniae. Curr. Opin. Microbiol. 2020, 54, 95–102. [Google Scholar] [CrossRef]
- Wyres, K.L.; Wick, R.R.; Gorrie, C.; Jenney, A.; Follador, R.; Thomson, N.R.; Holt, K.E. Identification of Klebsiella capsule synthesis loci from whole genome data. Microb. Genom. 2016, 2, e000102. [Google Scholar] [CrossRef]
- Liu, Y.; Bai, J.; Kang, J.; Song, Y.; Yin, D.; Wang, J.; Li, H.; Duan, J. Three Novel Sequence Types Carbapenem-Resistant Klebsiella pneumoniae Strains ST5365, ST5587, ST5647 Isolated from Two Tertiary Teaching General Hospitals in Shanxi Province, in North China: Molecular Characteristics, Resistance and Virulence Factors. Infect. Drug Resist. 2022, 15, 2551–2563. [Google Scholar] [CrossRef] [PubMed]
- Behzadi, P.; Garcia-Perdomo, H.A.; Karpinski, T.M. Toll-Like Receptors: General Molecular and Structural Biology. J. Immunol. Res. 2021, 2021, 9914854. [Google Scholar] [CrossRef] [PubMed]
- Behzadi, E.; Behzadi, P. The role of toll-like receptors (TLRs) in urinary tract infections (UTIs). Cent. Eur. J. Urol. 2016, 69, 404–410. [Google Scholar]
- Behzadi, P.; Behzadi, E.; Pawlak-Adamska, E.A. Urinary tract infections (UTIs) or genital tract infections (GTIs)? It’s the diagnostics that count. GMS Hyg. Infect. Control 2019, 14, Doc14. [Google Scholar]
- Behzadi, P.; Sameer, A.S.; Nissar, S.; Banday, M.Z.; Gajdács, M.; García-Perdomo, H.A.; Akhtar, K.; Pinheiro, M.; Magnusson, P.; Sarshar, M.; et al. The Interleukin-1 (IL-1) Superfamily Cytokines and Their Single Nucleotide Polymorphisms (SNPs). J. Immunol. Res. 2022, 2022, 2054431. [Google Scholar] [CrossRef]
- Soares, M.P.; Weiss, G. The Iron age of host-microbe interactions. EMBO Rep. 2015, 16, 1482–1500. [Google Scholar] [CrossRef]
- Jaeggi, T.; Kortman, G.A.M.; Moretti, D.; Chassard, C.; Holding, P.; Dostal, A.; Boekhorst, J.; Timmerman, H.M.; Swinkels, D.W.; Tjalsma, H.; et al. Iron fortification adversely affects the gut microbiome, increases pathogen abundance and induces intestinal inflammation in Kenyan infants. Gut 2015, 64, 731–742. [Google Scholar] [CrossRef]
- Aksoyalp, Z.S.; Temel, A.; Erdogan, B.R. Iron in infectious diseases friend or foe?: The role of gut microbiota. J. Trace Elem. Med. Biol. 2022, 75, 127093. [Google Scholar] [CrossRef]
- Shini, V.S.; Udayarajan, C.T.; Nisha, P. A comprehensive review on lactoferrin: A natural multifunctional glycoprotein. Food Funct. 2022, 13, 11954–11972. [Google Scholar] [CrossRef]
- Hu, D.; Li, Y.; Ren, P.; Tian, D.; Chen, W.; Fu, P.; Wang, W.; Li, X.; Jiang, X. Molecular Epidemiology of Hypervirulent Carbapenemase-Producing Klebsiella pneumoniae. Front. Cell. Infect. Microbiol. 2021, 11, 661218. [Google Scholar] [CrossRef]
- Murdoch, C.C.; Skaar, E.P. Nutritional immunity: The battle for nutrient metals at the host-pathogen interface. Nat. Rev. Microbiol. 2022, 20, 657–670. [Google Scholar] [CrossRef] [PubMed]
- Lopez, A.; Cacoub, P.; Macdougall, I.C.; Peyrin-Biroulet, L. Iron deficiency anaemia. Lancet 2016, 387, 907–916. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, P.F.; Lin, T.L.; Lee, C.Z.; Tsai, S.F.; Wang, J.T. Serum-induced iron-acquisition systems and TonB contribute to virulence in Klebsiella pneumoniae causing primary pyogenic liver abscess. J. Infect. Dis. 2008, 197, 1717–1727. [Google Scholar] [CrossRef] [PubMed]
- Saha, P.; Xiao, X.; Yeoh, B.S.; Chen, Q.; Katkere, B.; Kirimanjeswara, G.S.; Vijay-Kumar, M. The bacterial siderophore enterobactin confers survival advantage to Salmonella in macrophages. Gut Microbes 2019, 10, 412–423. [Google Scholar] [CrossRef] [PubMed]
- Raymond, K.N.; Dertz, E.A.; Kim, S.S. Enterobactin: An archetype for microbial iron transport. Proc. Natl. Acad. Sci. USA 2003, 100, 3584–3588. [Google Scholar] [CrossRef]
- Brock, J.H.; Williams, P.H.; Liceaga, J.; Wooldridge, K.G. Relative availability of transferrin-bound iron and cell-derived iron to aerobactin-producing and enterochelin-producing strains of Escherichia coli and to other microorganisms. Infect. Immun. 1991, 59, 3185–3190. [Google Scholar] [CrossRef]
- Nordmann, P.; Naas, T.; Poirel, L. Global spread of Carbapenemase-producing Enterobacteriaceae. Emerg. Infect. Dis. 2011, 17, 1791–1798. [Google Scholar] [CrossRef]
- Foudraine, D.E.; Strepis, N.; Klaassen, C.H.W.; Raaphorst, M.N.; Verbon, A.; Luider, T.M.; Goessens, W.H.F.; Dekker, L.J.M. Rapid and Accurate Detection of Aminoglycoside-Modifying Enzymes and 16S rRNA Methyltransferases by Targeted Liquid Chromatography-Tandem Mass Spectrometry. J. Clin. Microbiol. 2021, 59, e0046421. [Google Scholar] [CrossRef]
- Doménech-Sánchez, A.; Martínez-Martínez, L.; Hernández-Allés, S.; Conejo, M.D.C.; Pascual, A.; Tomás, J.M.; Albertí, S.; Benedí, V.J. Role of Klebsiella pneumoniae OmpK35 porin in antimicrobial resistance. Antimicrob. Agents Chemother. 2003, 47, 3332–3335. [Google Scholar] [CrossRef]
- Srinivasan, V.B.; Singh, B.B.; Priyadarshi, N.; Chauhan, N.K.; Rajamohan, G. Role of novel multidrug efflux pump involved in drug resistance in Klebsiella pneumoniae. PLoS ONE 2014, 9, e96288. [Google Scholar] [CrossRef]
- Nordmann, P.; Dortet, L.; Poirel, L. Carbapenem resistance in Enterobacteriaceae: Here is the storm! Trends Mol. Med. 2012, 18, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Essack, S.Y. The development of beta-lactam antibiotics in response to the evolution of beta-lactamases. Pharm. Res. 2001, 18, 1391–1399. [Google Scholar] [CrossRef] [PubMed]
- Bush, K.; Jacoby, G.A.; Medeiros, A.A. A functional classification scheme for beta-lactamases and its correlation with molecular structure. Antimicrob. Agents Chemother. 1995, 39, 1211–1233. [Google Scholar] [CrossRef] [PubMed]
- Ambler, R.P. The structure of beta-lactamases. Philos. Trans. R Soc. Lond. B Biol. Sci. 1980, 289, 321–331. [Google Scholar]
- Hall, B.G.; Barlow, M. Revised Ambler classification of {beta}-lactamases. J. Antimicrob. Chemother. 2005, 55, 1050–1051. [Google Scholar] [CrossRef]
- Akinci, E.; Vahaboglu, H. Minor extended-spectrum beta-lactamases. Expert Rev. Anti Infect. Ther. 2010, 8, 1251–1258. [Google Scholar] [CrossRef]
- Naas, T.; Poirel, L.; Nordmann, P. Minor extended-spectrum beta-lactamases. Clin. Microbiol. Infect. 2008, 14 (Suppl. 1), 42–52. [Google Scholar] [CrossRef]
- Landman, D.; Bratu, S.; Kochar, S.; Panwar, M.; Trehan, M.; Doymaz, M.; Quale, J. Evolution of antimicrobial resistance among Pseudomonas aeruginosa, Acinetobacter baumannii and Klebsiella pneumoniae in Brooklyn, NY. J. Antimicrob. Chemother. 2007, 60, 78–82. [Google Scholar] [CrossRef]
- Toleman, M.A.; Simm, A.M.; Murphy, T.A.; Gales, A.; Biedenbach, D.J.; Jones, R.N.; Walsh, T.R. Molecular characterization of SPM-1, a novel metallo-beta-lactamase isolated in Latin America: Report from the SENTRY antimicrobial surveillance programme. J. Antimicrob. Chemother. 2002, 50, 673–679. [Google Scholar] [CrossRef]
- Senda, K.; Arakawa, Y.; Nakashima, K.; Ito, H.; Ichiyama, S.; Shimokata, K.; Kato, N.; Ohta, M. Multifocal outbreaks of metallo-beta-lactamase-producing Pseudomonas aeruginosa resistant to broad-spectrum beta-lactams, including carbapenems. Antimicrob. Agents Chemother. 1996, 40, 349–353. [Google Scholar] [CrossRef]
- Poirel, L.; Naas, T.; Nicolas, D.; Collet, L.; Bellais, S.; Cavallo, J.D.; Nordmann, P. Characterization of VIM-2, a carbapenem-hydrolyzing metallo-beta-lactamase and its plasmid- and integron-borne gene from a Pseudomonas aeruginosa clinical isolate in France. Antimicrob. Agents Chemother. 2000, 44, 891–897. [Google Scholar] [CrossRef] [PubMed]
- Castanheira, M.; Toleman, M.A.; Jones, R.N.; Schmidt, F.J.; Walsh, T.R. Molecular characterization of a beta-lactamase gene, blaGIM-1, encoding a new subclass of metallo-beta-lactamase. Antimicrob. Agents Chemother. 2004, 48, 4654–4661. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Yum, J.H.; Yong, D.; Lee, H.M.; Kim, H.D.; Docquier, J.D.; Rossolini, G.M.; Chong, Y. Novel acquired metallo-beta-lactamase gene, bla(SIM-1), in a class 1 integron from Acinetobacter baumannii clinical isolates from Korea. Antimicrob. Agents Chemother. 2005, 49, 4485–4491. [Google Scholar] [CrossRef] [PubMed]
- Tada, T.; Shimada, K.; Satou, K.; Hirano, T.; Pokhrel, B.M.; Sherchand, J.B.; Kirikae, T. Pseudomonas aeruginosa Clinical Isolates in Nepal Coproducing Metallo-beta-Lactamases and 16S rRNA Methyltransferases. Antimicrob. Agents Chemother. 2017, 61, e00694-17. [Google Scholar] [CrossRef] [PubMed]
- Yong, D.; Toleman, M.A.; Bell, J.; Ritchie, B.; Pratt, R.; Ryley, H.; Walsh, T.R. Genetic and biochemical characterization of an acquired subgroup B3 metallo-beta-lactamase gene, blaAIM-1, and its unique genetic context in Pseudomonas aeruginosa from Australia. Antimicrob. Agents Chemother. 2012, 56, 6154–6159. [Google Scholar] [CrossRef]
- El Salabi, A.; Borra, P.S.; Toleman, M.A.; Samuelsen, O.; Walsh, T.R. Genetic and biochemical characterization of a novel metallo-beta-lactamase, TMB-1, from an Achromobacter xylosoxidans strain isolated in Tripoli, Libya. Antimicrob. Agents Chemother. 2012, 56, 2241–2245. [Google Scholar] [CrossRef]
- Pollini, S.; Maradei, S.; Pecile, P.; Olivo, G.; Luzzaro, F.; Docquier, J.D.; Rossolini, G.M. FIM-1, a new acquired metallo-beta-lactamase from a Pseudomonas aeruginosa clinical isolate from Italy. Antimicrob. Agents Chemother. 2013, 57, 410–416. [Google Scholar] [CrossRef]
- Jacoby, G.A. AmpC beta-lactamases. Clin. Microbiol. Rev. 2009, 22, 161–182, Table of Contents. [Google Scholar] [CrossRef]
- Antunes, N.T.; Lamoureaux, T.L.; Toth, M.; Stewart, N.K.; Frase, H.; Vakulenko, S.B. Class D beta-lactamases: Are they all carbapenemases? Antimicrob. Agents Chemother. 2014, 58, 2119–2125. [Google Scholar] [CrossRef]
- Shi, W.; Li, K.; Ji, Y.; Jiang, Q.; Wang, Y.; Shi, M.; Mi, Z. Carbapenem and cefoxitin resistance of Klebsiella pneumoniae strains associated with porin OmpK36 loss and DHA-1 beta-lactamase production. Braz. J. Microbiol. 2013, 44, 435–442. [Google Scholar] [CrossRef]
- Llobet, E.; March, C.; Gimenez, P.; Bengoechea, J.A. Klebsiella pneumoniae OmpA confers resistance to antimicrobial peptides. Antimicrob. Agents Chemother. 2009, 53, 298–302. [Google Scholar] [CrossRef] [PubMed]
- Partridge, S.R.; Ginn, A.N.; Wiklendt, A.M.; Ellem, J.; Wong, J.S.; Ingram, P.; Guy, S.; Garner, S.; Iredell, J.R. Emergence of blaKPC carbapenemase genes in Australia. Int. J. Antimicrob. Agents 2015, 45, 130–136. [Google Scholar] [CrossRef] [PubMed]
- Tang, H.J.; Ku, Y.H.; Lee, M.F.; Chuang, Y.C.; Yu, W.L. In Vitro Activity of Imipenem and Colistin against a Carbapenem-Resistant Klebsiella pneumoniae Isolate Coproducing SHV-31, CMY-2, and DHA-1. Biomed Res. Int. 2015, 2015, 568079. [Google Scholar] [CrossRef] [PubMed]
- Kaczmarek, F.M.; Dib-Hajj, F.; Shang, W.; Gootz, T.D. High-level carbapenem resistance in a Klebsiella pneumoniae clinical isolate is due to the combination of bla(ACT-1) beta-lactamase production, porin OmpK35/36 insertional inactivation, and down-regulation of the phosphate transport porin phoe. Antimicrob. Agents Chemother. 2006, 50, 3396–3406. [Google Scholar] [CrossRef] [PubMed]
- Norman, A.; Hansen, L.H.; Sorensen, S.J. Conjugative plasmids: Vessels of the communal gene pool. Philos. Trans. R Soc. Lond. B Biol. Sci. 2009, 364, 2275–2289. [Google Scholar] [CrossRef] [PubMed]
- San Millan, A.; Toll-Riera, M.; Escudero, J.A.; Canton, R.; Coque, T.M.; MacLean, R.C. Sequencing of plasmids pAMBL1 and pAMBL2 from Pseudomonas aeruginosa reveals a blaVIM-1 amplification causing high-level carbapenem resistance. J. Antimicrob. Chemother. 2015, 70, 3000–3003. [Google Scholar] [CrossRef]
- Munoz-Price, L.S.; Quinn, J.P. The spread of Klebsiella pneumoniae carbapenemases: A tale of strains, plasmids, and transposons. Clin. Infect. Dis. 2009, 49, 1739–1741. [Google Scholar] [CrossRef]
- Labbate, M.; Case, R.J.; Stokes, H.W. The integron/gene cassette system: An active player in bacterial adaptation. Methods Mol. Biol. 2009, 532, 103–125. [Google Scholar]
- Carattoli, A. Importance of integrons in the diffusion of resistance. Vet. Res. 2001, 32, 243–259. [Google Scholar] [CrossRef]
- Koh, T.H.; Babini, G.S.; Woodford, N.; Sng, L.H.; Hall, L.M.; Livermore, D.M. Carbapenem-hydrolysing IMP-1 beta-lactamase in Klebsiella pneumoniae from Singapore. Lancet 1999, 353, 2162. [Google Scholar] [CrossRef]
- Li, B.; Xu, X.H.; Zhao, Z.C.; Wang, M.H.; Cao, Y.P. High prevalence of metallo-beta-lactamase among carbapenem-resistant Klebsiella pneumoniae in a teaching hospital in China. Can. J. Microbiol. 2014, 60, 691–695. [Google Scholar] [CrossRef] [PubMed]
- Limbago, B.M.; Rasheed, J.K.; Anderson, K.F.; Zhu, W.; Kitchel, B.; Watz, N.; Munro, S.; Gans, H.; Banaei, N.; Kallen, A.J. IMP-producing carbapenem-resistant Klebsiella pneumoniae in the United States. J. Clin. Microbiol. 2011, 49, 4239–4245. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kassis-Chikhani, N.; Decré, D.; Gautier, V.; Burghoffer, B.; Saliba, F.; Mathieu, D.; Samuel, D.; Castaing, D.; Petit, J.-C.; Dussaix, E.; et al. First outbreak of multidrug-resistant Klebsiella pneumoniae carrying blaVIM-1 and blaSHV-5 in a French university hospital. J. Antimicrob. Chemother. 2006, 57, 142–145. [Google Scholar] [CrossRef] [PubMed]
- Karampatakis, T.; Antachopoulos, C.; Iosifidis, E.; Tsakris, A.; Roilides, E. Molecular epidemiology of carbapenem-resistant Klebsiella pneumoniae in Greece. Future Microbiol. 2016, 11, 809–823. [Google Scholar] [CrossRef]
- Hasan, C.M.; Turlej-Rogacka, A.; Vatopoulos, A.C.; Giakkoupi, P.; Maatallah, M.; Giske, C.G. Dissemination of blaVIM in Greece at the peak of the epidemic of 2005-2006: Clonal expansion of Klebsiella pneumoniae clonal complex 147. Clin. Microbiol. Infect. 2014, 20, 34–37. [Google Scholar] [CrossRef] [PubMed]
- Arca-Suárez, J.; Rodiño-Janeiro, B.K.; Pérez, A.; Guijarro-Sánchez, P.; Vázquez-Ucha, J.C.; Cruz, F.; Gómez-Garrido, J.; Alioto, T.S.; Álvarez-Tejado, M.; Gut, M.; et al. Emergence of 16S rRNA methyltransferases among carbapenemase-producing Enterobacterales in Spain studied by whole-genome sequencing. Int. J. Antimicrob. Agents 2022, 59, 106456. [Google Scholar] [CrossRef] [PubMed]
- Voulgari, E.; Gartzonika, C.; Vrioni, G.; Politi, L.; Priavali, E.; Levidiotou-Stefanou, S.; Tsakris, A. The Balkan region: NDM-1-producing Klebsiella pneumoniae ST11 clonal strain causing outbreaks in Greece. J. Antimicrob. Chemother. 2014, 69, 2091–2097. [Google Scholar] [CrossRef]
- Johnson, A.P.; Woodford, N. Global spread of antibiotic resistance: The example of New Delhi metallo-beta-lactamase (NDM)-mediated carbapenem resistance. J. Med. Microbiol. 2013, 62, 499–513. [Google Scholar] [CrossRef]
- Yong, D.; Toleman, M.A.; Giske, C.G.; Cho, H.S.; Sundman, K.; Lee, K.; Walsh, T.R. Characterization of a new metallo-beta-lactamase gene, bla(NDM-1), and a novel erythromycin esterase gene carried on a unique genetic structure in Klebsiella pneumoniae sequence type 14 from India. Antimicrob. Agents Chemother. 2009, 53, 5046–5054. [Google Scholar] [CrossRef]
- Moellering, R.C., Jr. NDM-1—A cause for worldwide concern. N. Engl. J. Med. 2010, 363, 2377–2379. [Google Scholar] [CrossRef]
- Izdebski, R.; Biedrzycka, M.; Urbanowicz, P.; Papierowska-Kozdój, W.; Dominiak, M.; Żabicka, D.; Gniadkowski, M. Multiple secondary outbreaks of NDM-producing Enterobacter hormaechei in the context of endemic NDM-producing Klebsiella pneumoniae. J. Antimicrob. Chemother. 2022, 77, 1561–1569. [Google Scholar] [CrossRef] [PubMed]
- Maltezou, H.; Giakkoupi, P.; Maragos, A.; Bolikas, M.; Raftopoulos, V.; Papahatzaki, H.; Vrouhos, G.; Liakou, V.; Vatopoulos, A. Outbreak of infections due to KPC-2-producing Klebsiella pneumoniae in a hospital in Crete (Greece). J. Infect. 2009, 58, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Pournaras, S.; Protonotariou, E.; Voulgari, E.; Kristo, I.; Dimitroulia, E.; Vitti, D.; Tsalidou, M.; Maniatis, A.N.; Tsakris, A.; Sofianou, D. Clonal spread of KPC-2 carbapenemase-producing Klebsiella pneumoniae strains in Greece. J. Antimicrob. Chemother. 2009, 64, 348–352. [Google Scholar] [CrossRef] [PubMed]
- Giakkoupi, P.; Maltezou, H.; Polemis, M.; Pappa, O.; Saroglou, G.; Vatopoulos, A. KPC-2-producing Klebsiella pneumoniae infections in Greek hospitals are mainly due to a hyperepidemic clone. Eurosurveillance 2009, 14, 19218. [Google Scholar] [CrossRef]
- Kitchel, B.; Rasheed, J.K.; Patel, J.B.; Srinivasan, A.; Navon-Venezia, S.; Carmeli, Y.; Brolund, A.; Giske, C.G. Molecular epidemiology of KPC-producing Klebsiella pneumoniae isolates in the United States: Clonal expansion of multilocus sequence type 258. Antimicrob. Agents Chemother. 2009, 53, 3365–3370. [Google Scholar] [CrossRef]
- Giakkoupi, P.; Papagiannitsis, C.C.; Miriagou, V.; Pappa, O.; Polemis, M.; Tryfinopoulou, K.; Tzouvelekis, L.S.; Vatopoulos, A.C. An update of the evolving epidemic of blaKPC-2-carrying Klebsiella pneumoniae in Greece (2009-10). J. Antimicrob. Chemother. 2011, 66, 1510–1513. [Google Scholar] [CrossRef]
- Tzouvelekis, L.S.; Miriagou, V.; Kotsakis, S.D.; Spyridopoulou, K.; Athanasiou, E.; Karagouni, E.; Tzelepi, E.; Daikos, G.L. KPC-producing, multidrug-resistant Klebsiella pneumoniae sequence type 258 as a typical opportunistic pathogen. Antimicrob. Agents Chemother. 2013, 57, 5144–5146. [Google Scholar] [CrossRef]
- Borer, A.; Saidel-Odes, L.; Eskira, S.; Nativ, R.; Riesenberg, K.; Livshiz-Riven, I.; Schlaeffer, F.; Sherf, M.; Peled, N. Risk factors for developing clinical infection with carbapenem-resistant Klebsiella pneumoniae in hospital patients initially only colonized with carbapenem-resistant K pneumoniae. Am. J. Infect. Control 2012, 40, 421–425. [Google Scholar] [CrossRef]
- Räisänen, K.; Koivula, I.; Ilmavirta, H.; Puranen, S.; Kallonen, T.; Lyytikäinen, O.; Jalava, J. Emergence of ceftazidime-avibactam-resistant Klebsiella pneumoniae during treatment, Finland, December 2018. Eurosurveillance 2019, 24, 1900256. [Google Scholar] [CrossRef]
- Papagiannitsis, C.C.; Giakkoupi, P.; Kotsakis, S.D.; Tzelepi, E.; Tzouvelekis, L.S.; Vatopoulos, A.C.; Miriagou, V. OmpK35 and OmpK36 porin variants associated with specific sequence types of Klebsiella pneumoniae. J. Chemother. 2013, 25, 250–254. [Google Scholar] [CrossRef]
- Karampatakis, T.; Zarras, C.; Pappa, S.; Vagdatli, E.; Iosifidis, E.; Roilides, E.; Papa, A. Emergence of ST39 carbapenem-resistant Klebsiella pneumoniae producing VIM-1 and KPC-2. Microb. Pathog. 2022, 162, 105373. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Heritier, C.; Tolun, V.; Nordmann, P. Emergence of oxacillinase-mediated resistance to imipenem in Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2004, 48, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Voulgari, E.; Zarkotou, O.; Ranellou, K.; Karageorgopoulos, D.E.; Vrioni, G.; Mamali, V.; Themeli-Digalaki, K.; Tsakris, A. Outbreak of OXA-48 carbapenemase-producing Klebsiella pneumoniae in Greece involving an ST11 clone. J. Antimicrob. Chemother. 2013, 68, 84–88. [Google Scholar] [CrossRef]
- Kasap, M.; Torol, S.; Kolayli, F.; Dundar, D.; Vahaboglu, H. OXA-162, a novel variant of OXA-48 displays extended hydrolytic activity towards imipenem, meropenem and doripenem. J. Enzyme. Inhib. Med. Chem. 2013, 28, 990–996. [Google Scholar] [CrossRef] [PubMed]
- Pereira, P.S.; Borghi, M.; de Araújo, C.F.M.; Aires, C.A.M.; Oliveira, J.C.R.; Asensi, M.D.; Carvalho-Assef, A.P.D. Clonal Dissemination of OXA-370-Producing Klebsiella pneumoniae in Rio de Janeiro, Brazil. Antimicrob. Agents Chemother. 2015, 59, 4453–4456. [Google Scholar] [CrossRef] [PubMed]
- Loli, A.; Tzouvelekis, L.S.; Tzelepi, E.; Carattoli, A.; Vatopoulos, A.C.; Tassios, P.T.; Miriagou, V. Sources of diversity of carbapenem resistance levels in Klebsiella pneumoniae carrying blaVIM-1. J. Antimicrob. Chemother. 2006, 58, 669–672. [Google Scholar] [CrossRef]
- Poulou, A.; Voulgari, E.; Vrioni, G.; Koumaki, V.; Xidopoulos, G.; Chatzipantazi, V.; Markou, F.; Tsakris, A. Outbreak caused by an ertapenem-resistant, CTX-M-15-producing Klebsiella pneumoniae sequence type 101 clone carrying an OmpK36 porin variant. J. Clin. Microbiol. 2013, 51, 3176–3182. [Google Scholar] [CrossRef]
- Biswas, S.; Brunel, J.M.; Dubus, J.C.; Reynaud-Gaubert, M.; Rolain, J.M. Colistin: An update on the antibiotic of the 21st century. Expert Rev. Anti Infect. Ther. 2012, 10, 917–934. [Google Scholar] [CrossRef]
- Cassir, N.; Rolain, J.M.; Brouqui, P. A new strategy to fight antimicrobial resistance: The revival of old antibiotics. Front. Microbiol. 2014, 5, 551. [Google Scholar] [CrossRef]
- Pournaras, S.; Vrioni, G.; Neou, E.; Dendrinos, J.; Dimitroulia, E.; Poulou, A.; Tsakris, A. Activity of tigecycline alone and in combination with colistin and meropenem against Klebsiella pneumoniae carbapenemase (KPC)-producing Enterobacteriaceae strains by time-kill assay. Int. J. Antimicrob. Agents 2011, 37, 244–247. [Google Scholar] [CrossRef]
- Zarkotou, O.; Pournaras, S.; Tselioti, P.; Dragoumanos, V.; Pitiriga, V.; Ranellou, K.; Prekates, A.; Themeli-Digalaki, K.; Tsakris, A. Predictors of mortality in patients with bloodstream infections caused by KPC-producing Klebsiella pneumoniae and impact of appropriate antimicrobial treatment. Clin. Microbiol. Infect. 2011, 17, 1798–1803. [Google Scholar] [CrossRef]
- Rafailidis, P.I.; Falagas, M.E. Options for treating carbapenem-resistant Enterobacteriaceae. Curr. Opin. Infect. Dis. 2014, 27, 479–483. [Google Scholar] [CrossRef]
- Falagas, M.E.; Kasiakou, S.K. Colistin: The revival of polymyxins for the management of multidrug-resistant gram-negative bacterial infections. Clin. Infect. Dis. 2005, 40, 1333–1341. [Google Scholar] [CrossRef]
- Tascini, C.; Tagliaferri, E.; Giani, T.; Leonildi, A.; Flammini, S.; Casini, B.; Lewis, R.; Ferranti, S.; Rossolini, G.M.; Menichetti, F. Synergistic activity of colistin plus rifampin against colistin-resistant KPC-producing Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2013, 57, 3990–3993. [Google Scholar] [CrossRef]
- Tuon, F.F.; Santos, T.A.; Almeida, R.; Rocha, J.L.; Cieslinsk, J.; Becker, G.N.; Arend, L.N. Colistin-resistant Enterobacteriaceae bacteraemia: Real-life challenges and options. Clin. Microbiol. Infect. 2016, 22, e9–e10. [Google Scholar] [CrossRef][Green Version]
- Paul, M.; Carrara, E.; Retamar, P.; Tängdén, T.; Bitterman, R.; Bonomo, R.A.; de Waele, J.; Daikos, G.L.; Akova, M.; Harbarth, S.; et al. European Society of Clinical Microbiology and Infectious Diseases (ESCMID) guidelines for the treatment of infections caused by multidrug-resistant Gram-negative bacilli (endorsed by European society of intensive care medicine). Clin. Microbiol. Infect. 2022, 28, 521–547. [Google Scholar] [CrossRef]
- Tamma, P.D.; Aitken, S.L.; Bonomo, R.A.; Mathers, A.J.; van Duin, D.; Clancy, C.J. Infectious Diseases Society of America 2022 Guidance on the Treatment of Extended-Spectrum beta-lactamase Producing Enterobacterales (ESBL-E), Carbapenem-Resistant Enterobacterales (CRE), and Pseudomonas aeruginosa with Difficult-to-Treat Resistance (DTR-P. aeruginosa). Clin. Infect. Dis. 2022, 75, 187–212. [Google Scholar]
- Entenza, J.M.; Moreillon, P. Tigecycline in combination with other antimicrobials: A review of in vitro, animal and case report studies. Int. J. Antimicrob. Agents 2009, 34, 8.e1–8.e9. [Google Scholar] [CrossRef]
- Hirsch, E.B.; Tam, V.H. Detection and treatment options for Klebsiella pneumoniae carbapenemases (KPCs): An emerging cause of multidrug-resistant infection. J. Antimicrob. Chemother. 2010, 65, 1119–1125. [Google Scholar] [CrossRef]
- Mezzatesta, M.L.; La Rosa, G.; Maugeri, G.; Zingali, T.; Caio, C.; Novelli, A.; Stefani, S. In vitro activity of fosfomycin trometamol and other oral antibiotics against multidrug-resistant uropathogens. Int. J. Antimicrob. Agents 2017, 49, 763–766. [Google Scholar] [CrossRef]
- Sharma, R.; Garcia, E.; Diep, J.K.; Lee, V.H.; Minhaj, F.; Jermain, B.; Ellis-Grosse, E.J.; Abboud, C.S.; Rao, G.G. Pharmacodynamic and immunomodulatory effects of polymyxin B in combination with fosfomycin against KPC-2-producing Klebsiella pneumoniae. Int. J. Antimicrob. Agents 2022, 59, 106566. [Google Scholar] [CrossRef]
- Yusuf, E.; Bax, H.I.; Verkaik, N.J.; van Westreenen, M. An Update on Eight “New” Antibiotics against Multidrug-Resistant Gram-Negative Bacteria. J. Clin. Med. 2021, 10, 1068. [Google Scholar] [CrossRef]
- PubChem Compounds. Available online: https://pubchem.ncbi.nlm.nih.gov/docs/compounds (accessed on 16 December 2022).
- Castanheira, M.; Sader, H.S.; Mendes, R.E.; Jones, R.N. Activity of Plazomicin Tested against Enterobacterales Isolates Collected from U.S. Hospitals in 2016–2017: Effect of Different Breakpoint Criteria on Susceptibility Rates among Aminoglycosides. Antimicrob. Agents Chemother. 2020, 64, e02418-19. [Google Scholar] [CrossRef]
- Livermore, D.M.; Mushtaq, S.; Warner, M.; Woodford, N. In Vitro Activity of Eravacycline against Carbapenem-Resistant Enterobacteriaceae and Acinetobacter baumannii. Antimicrob. Agents Chemother. 2016, 60, 3840–3844. [Google Scholar] [CrossRef]
- Maraki, S.; Mavromanolaki, V.E.; Magkafouraki, E.; Moraitis, P.; Stafylaki, D.; Kasimati, A.; Scoulica, E. Epidemiology and in vitro activity of ceftazidime-avibactam, meropenem-vaborbactam, imipenem-relebactam, eravacycline, plazomicin, and comparators against Greek carbapenemase-producing Klebsiella pneumoniae isolates. Infection 2022, 50, 467–474. [Google Scholar] [CrossRef]
- Zheng, J.X.; Lin, Z.W.; Sun, X.; Lin, W.H.; Chen, Z.; Wu, Y.; Qi, G.B.; Deng, Q.W.; Qu, D.; Yu, Z.J. Overexpression of OqxAB and MacAB efflux pumps contributes to eravacycline resistance and heteroresistance in clinical isolates of Klebsiella pneumoniae. Emerg. Microbes Infect. 2018, 7, 139. [Google Scholar] [CrossRef]
- Zhanel, G.G.; Golden, A.R.; Zelenitsky, S.; Wiebe, K.; Lawrence, C.K.; Adam, H.J.; Idowu, T.; Domalaon, R.; Schweizer, F.; Zhanel, M.A.; et al. Cefiderocol: A Siderophore Cephalosporin with Activity Against Carbapenem-Resistant and Multidrug-Resistant Gram-Negative Bacilli. Drugs 2019, 79, 271–289. [Google Scholar] [CrossRef]
- Falcone, M.; Tiseo, G.; Nicastro, M.; Leonildi, A.; Vecchione, A.; Casella, C.; Forfori, F.; Malacarne, P.; Guarracino, F.; Barnini, S.; et al. Cefiderocol as Rescue Therapy for Acinetobacter baumannii and Other Carbapenem-resistant Gram-negative Infections in Intensive Care Unit Patients. Clin. Infect. Dis. 2021, 72, 2021–2024. [Google Scholar] [CrossRef]
- Yao, J.; Wang, J.; Chen, M.; Cai, Y. Cefiderocol: An Overview of Its in-vitro and in-vivo Activity and Underlying Resistant Mechanisms. Front. Med. 2021, 8, 741940. [Google Scholar] [CrossRef]
- Poirel, L.; Sadek, M.; Kusaksizoglu, A.; Nordmann, P. Co-resistance to ceftazidime-avibactam and cefiderocol in clinical isolates producing KPC variants. Eur. J. Clin. Microbiol. Infect. Dis. 2022, 41, 677–680. [Google Scholar] [CrossRef]
- Adams-Haduch, J.M.; Potoski, B.A.; Sidjabat, H.E.; Paterson, D.L.; Doi, Y. Activity of temocillin against KPC-producing Klebsiella pneumoniae and Escherichia coli. Antimicrob. Agents Chemother. 2009, 53, 2700–2701. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Kuch, A.; Zieniuk, B.; Żabicka, D.; Van de Velde, S.; Literacka, E.; Skoczyńska, A.; Hryniewicz, W. Activity of temocillin against ESBL-, AmpC-, and/or KPC-producing Enterobacterales isolated in Poland. Eur. J. Clin. Microbiol. Infect. Dis. 2020, 39, 1185–1191. [Google Scholar] [CrossRef]
- Tsakris, A.; Koumaki, V.; Politi, L.; Balakrishnan, I. Activity of temocillin against KPC-producing Enterobacteriaceae clinical isolates. Int. J. Antimicrob. Agents 2020, 55, 105843. [Google Scholar] [CrossRef] [PubMed]
- Woodford, N.; Pike, R.; Meunier, D.; Loy, R.; Hill, R.; Hopkins, K.L. In vitro activity of temocillin against multidrug-resistant clinical isolates of Escherichia coli, Klebsiella spp. and Enterobacter spp., and evaluation of high-level temocillin resistance as a diagnostic marker for OXA-48 carbapenemase. J. Antimicrob. Chemother. 2014, 69, 564–567. [Google Scholar] [CrossRef]
- Karlowsky, J.A.; Lob, S.H.; Khan, A.; Chen, W.-T.; Woo, P.C.Y.; Seto, W.H.; Ip, M.; Leung, S.; Wong, Q.W.-L.; Chau, R.W.Y.; et al. Activity of ceftolozane/tazobactam against Gram-negative isolates among different infections in Hong Kong: SMART 2017–2019. J. Med. Microbiol. 2022, 71, 001487. [Google Scholar] [CrossRef] [PubMed]
- Mansour, H.; Ouweini, A.E.L.; Chahine, E.B.; Karaoui, L.R. Imipenem/cilastatin/relebactam: A new carbapenem beta-lactamase inhibitor combination. Am. J. Health Syst. Pharm. 2021, 78, 674–683. [Google Scholar] [CrossRef]
- Haidar, G.; Clancy, C.J.; Chen, L.; Samanta, P.; Shields, R.K.; Kreiswirth, B.N.; Nguyen, M.H. Identifying Spectra of Activity and Therapeutic Niches for Ceftazidime-Avibactam and Imipenem-Relebactam against Carbapenem-Resistant Enterobacteriaceae. Antimicrob. Agents Chemother. 2017, 61, e00642-17. [Google Scholar] [CrossRef] [PubMed]
- Gaibani, P.; Bovo, F.; Bussini, L.; Lazzarotto, T.; Amadesi, S.; Bartoletti, M.; Viale, P.; Ambretti, S. Dynamic evolution of imipenem/relebactam resistance in a KPC-producing Klebsiella pneumoniae from a single patient during ceftazidime/avibactam-based treatments. J. Antimicrob. Chemother. 2022, 77, 1570–1577. [Google Scholar] [CrossRef]
- Mouktaroudi, M.; Kotsaki, A.; Giamarellos-Bourboulis, E.J. Meropenem-vaborbactam: A critical positioning for the management of infections by Carbapenem-resistant Enterobacteriaceae. Expert Rev. Anti Infect. Ther. 2022, 20, 809–818. [Google Scholar] [CrossRef]
- Castanheira, M.; Doyle, T.B.; Kantro, V.; Mendes, R.E.; Shortridge, D. Meropenem-Vaborbactam Activity against Carbapenem-Resistant Enterobacterales Isolates Collected in U.S. Hospitals during 2016 to 2018. Antimicrob. Agents Chemother. 2020, 64, e01951-19. [Google Scholar] [CrossRef]
- Gaibani, P.; Lombardo, D.; Bussini, L.; Bovo, F.; Munari, B.; Giannella, M.; Bartoletti, M.; Viale, P.; Lazzarotto, T.; Ambretti, S. Epidemiology of Meropenem/Vaborbactam Resistance in KPC-Producing Klebsiella pneumoniae Causing Bloodstream Infections in Northern Italy, 2018. Antibiotics 2021, 10, 536. [Google Scholar] [CrossRef] [PubMed]
- Zhanel, G.G.; Lawson, C.D.; Adam, H.; Schweizer, F.; Zelenitsky, S.; Lagacé-Wiens, P.R.; Denisuik, A.; Rubinstein, E.; Gin, A.S.; Hoban, D.J.; et al. Ceftazidime-avibactam: A novel cephalosporin/beta-lactamase inhibitor combination. Drugs 2013, 73, 159–177. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Zhang, F.; Zhao, C.; Wang, Z.; Nichols, W.W.; Testa, R.; Li, H.; Chen, H.; He, W.; Wang, Q.; et al. In vitro activities of ceftazidime-avibactam and aztreonam-avibactam against 372 Gram-negative bacilli collected in 2011 and 2012 from 11 teaching hospitals in China. Antimicrob. Agents Chemother. 2014, 58, 1774–1778. [Google Scholar] [CrossRef] [PubMed]
- Sader, H.S.; Castanheira, M.; Flamm, R.K.; Farrell, D.J.; Jones, R.N. Antimicrobial activity of ceftazidime-avibactam against Gram-negative organisms collected from U.S. medical centers in 2012. Antimicrob. Agents Chemother. 2014, 58, 1684–1692. [Google Scholar] [CrossRef]
- Flamm, R.K.; Farrell, D.J.; Sader, H.S.; Jones, R.N. Ceftazidime/avibactam activity tested against Gram-negative bacteria isolated from bloodstream, pneumonia, intra-abdominal and urinary tract infections in US medical centres (2012). J. Antimicrob. Chemother. 2014, 69, 1589–1598. [Google Scholar] [CrossRef]
- Yu, F.; Lv, J.; Niu, S.; Du, H.; Tang, Y.-W.; Bonomo, R.A.; Kreiswirth, B.N.; Chen, L. In Vitro Activity of Ceftazidime-Avibactam against Carbapenem-Resistant and Hypervirulent Klebsiella pneumoniae Isolates. Antimicrob. Agents Chemother. 2018, 62, e01031-18. [Google Scholar] [CrossRef]
- Benchetrit, L.; Mathy, V.; Armand-Lefevre, L.; Bouadma, L.; Timsit, J.F. Successful treatment of septic shock due to NDM-1-producing Klebsiella pneumoniae using ceftazidime/avibactam combined with aztreonam in solid organ transplant recipients: Report of two cases. Int. J. Antimicrob. Agents 2020, 55, 105842. [Google Scholar] [CrossRef]
- Bocanegra-Ibarias, P.; Camacho-Ortiz, A.; Garza-Gonzalez, E.; Flores-Trevino, S.; Kim, H.; Perez-Alba, E. Aztreonam plus ceftazidime-avibactam as treatment of NDM-1-producing Klebsiella pneumoniae bacteraemia in a neutropenic patient: Last resort therapy? J. Glob. Antimicrob. Resis.t 2020, 23, 417–419. [Google Scholar] [CrossRef]
- Perrotta, F.; Perrini, M.P. Successful Treatment of Klebsiella pneumoniae NDM Sepsis and Intestinal Decolonization with Ceftazidime/Avibactam Plus Aztreonam Combination in a Patient with TTP Complicated by SARSCoV-2 Nosocomial Infection. Medicina 2021, 57, 424. [Google Scholar] [CrossRef]
- De la Calle, C.; Rodríguez, O.; Morata, L.; Marco, F.; Cardozo, C.; García-Vidal, C.; Del Río, A.; Feher, C.; Pellicé, M.; Puerta-Alcalde, P.; et al. Clinical characteristics and prognosis of infections caused by OXA-48 carbapenemase-producing Enterobacteriaceae in patients treated with ceftazidime-avibactam. Int. J. Antimicrob. Agents 2019, 53, 520–524. [Google Scholar] [CrossRef]
- Mawal, Y.; Critchley, I.A.; Riccobene, T.A.; Talley, A.K. Ceftazidime-avibactam for the treatment of complicated urinary tract infections and complicated intra-abdominal infections. Expert Rev. Clin. Pharmacol. 2015, 8, 691–707. [Google Scholar] [CrossRef] [PubMed]
- Sousa, A.; Pérez-Rodríguez, M.T.; Soto, A.; Rodríguez, L.; Perez-Landeiro, A.; Martínez-Lamas, L.; Nodar, A.; Crespo, M. Effectiveness of ceftazidime/avibactam as salvage therapy for treatment of infections due to OXA-48 carbapenemase-producing Enterobacteriaceae. J. Antimicrob. Chemother. 2018, 73, 3170–3175. [Google Scholar] [CrossRef] [PubMed]
- Thompson, C.A. Ceftazidime with beta-lactamase inhibitor approved for complicated infections. Am. J. Health Syst. Pharm. 2015, 72, 511. [Google Scholar] [CrossRef] [PubMed]
- Stein, G.E.; Smith, C.L.; Scharmen, A.; Kidd, J.M.; Cooper, C.; Kuti, J.; Mitra, S.; Nicolau, D.P.; Havlichek, D.H. Pharmacokinetic and Pharmacodynamic Analysis of Ceftazidime/Avibactam in Critically Ill Patients. Surg. Infect. 2019, 20, 55–61. [Google Scholar] [CrossRef]
- Dietl, B.; Martinez, L.M.; Calbo, E.; Garau, J. Update on the role of ceftazidime-avibactam in the management of carbapenemase-producing Enterobacterales. Future Microbiol. 2020, 15, 473–484. [Google Scholar] [CrossRef]
- Chen, F.; Zhong, H.; Yang, T.; Shen, C.; Deng, Y.; Han, L.; Chen, X.; Zhang, H.; Qian, Y. Ceftazidime-Avibactam as Salvage Treatment for Infections Due to Carbapenem-Resistant Klebsiella pneumoniae in Liver Transplantation Recipients. Infect. Drug Resist. 2021, 14, 5603–5612. [Google Scholar] [CrossRef] [PubMed]
- Sader, H.S.; Castanheira, M.; Flamm, R.K.; Mendes, R.E.; Farrell, D.J.; Jones, R.N. Ceftazidime/avibactam tested against Gram-negative bacteria from intensive care unit (ICU) and non-ICU patients, including those with ventilator-associated pneumonia. Int. J. Antimicrob. Agents 2015, 46, 53–59. [Google Scholar] [CrossRef]
- Liapikou, A.; Torres, A. Emerging drugs for nosocomial pneumonia. Expert Opin. Emerg. Drugs 2016, 21, 331–341. [Google Scholar] [CrossRef]
- Torres, A.; Rank, D.; Melnick, D.; Rekeda, L.; Chen, X.; Riccobene, T.; Critchley, I.A.; Lakkis, H.D.; Taylor, D.; Talley, A.K. Randomized Trial of Ceftazidime-Avibactam vs Meropenem for Treatment of Hospital-Acquired and Ventilator-Associated Bacterial Pneumonia (REPROVE): Analyses per US FDA-Specified End Points. Open Forum. Infect. Dis. 2019, 6, ofz149. [Google Scholar] [CrossRef]
- Wu, G.; Abraham, T.; Lee, S. Ceftazidime-Avibactam for Treatment of Carbapenem-Resistant Enterobacteriaceae Bacteremia. Clin. Infect. Dis. 2016, 63, 1147–1148. [Google Scholar] [CrossRef]
- Hachem, R.; Reitzel, R.; Rolston, K.; Chaftari, A.M.; Raad, I. Antimicrobial Activities of Ceftazidime-Avibactam and Comparator Agents against Clinical Bacteria Isolated from Patients with Cancer. Antimicrob. Agents Chemother. 2017, 61, e02106-16. [Google Scholar] [CrossRef] [PubMed]
- Castón, J.J.; Lacort-Peralta, I.; Martín-Dávila, P.; Loeches, B.; Tabares, S.; Temkin, L.; Torre-Cisneros, J.; Paño-Pardo, J.R. Clinical efficacy of ceftazidime/avibactam versus other active agents for the treatment of bacteremia due to carbapenemase-producing Enterobacteriaceae in hematologic patients. Int. J. Infect. Dis. 2017, 59, 118–123. [Google Scholar] [CrossRef] [PubMed]
- Shields, R.K.; Nguyen, M.H.; Chen, L.; Press, E.G.; Potoski, B.A.; Marini, R.V.; Doi, Y.; Kreiswirth, B.N.; Clancy, C.J. Ceftazidime-Avibactam Is Superior to Other Treatment Regimens against Carbapenem-Resistant Klebsiella pneumoniae Bacteremia. Antimicrob. Agents Chemother. 2017, 61, e00883-17. [Google Scholar] [CrossRef]
- Papp-Wallace, K.M.; Winkler, M.L.; Taracila, M.A.; Bonomo, R.A. Variants of beta-lactamase KPC-2 that are resistant to inhibition by avibactam. Antimicrob. Agents Chemother. 2015, 59, 3710–3717. [Google Scholar] [CrossRef] [PubMed]
- Humphries, R.M.; Yang, S.; Hemarajata, P.; Ward, K.W.; Hindler, J.A.; Miller, S.A.; Gregson, A. First Report of Ceftazidime-Avibactam Resistance in a KPC-3-Expressing Klebsiella pneumoniae Isolate. Antimicrob. Agents Chemother. 2015, 59, 6605–6607. [Google Scholar] [CrossRef]
- Livermore, D.M.; Warner, M.; Jamrozy, D.; Mushtaq, S.; Nichols, W.W.; Mustafa, N.; Woodford, N. In vitro selection of ceftazidime-avibactam resistance in Enterobacteriaceae with KPC-3 carbapenemase. Antimicrob. Agents Chemother. 2015, 59, 5324–5330. [Google Scholar] [CrossRef]
- Shields, R.K.; Chen, L.; Cheng, S.; Chavda, K.D.; Press, E.G.; Snyder, A.; Pandey, R.; Doi, Y.; Kreiswirth, B.N.; Nguyen, M.H.; et al. Emergence of Ceftazidime-Avibactam Resistance Due to Plasmid-Borne blaKPC-3 Mutations during Treatment of Carbapenem-Resistant Klebsiella pneumoniae Infections. Antimicrob. Agents Chemother. 2017, 61, e02097-16. [Google Scholar] [CrossRef]
- Tsivkovski, R.; Lomovskaya, O. Potency of Vaborbactam Is Less Affected than That of Avibactam in Strains Producing KPC-2 Mutations That Confer Resistance to Ceftazidime-Avibactam. Antimicrob. Agents Chemother. 2020, 64, e01936-19. [Google Scholar] [CrossRef]
- Belati, A.; Bavaro, D.F.; Diella, L.; De Gennaro, N.; Di Gennaro, F.; Saracino, A. Meropenem/Vaborbactam Plus Aztreonam as a Possible Treatment Strategy for Bloodstream Infections Caused by Ceftazidime/Avibactam-Resistant Klebsiella pneumoniae: A Retrospective Case Series and Literature Review. Antibiotics 2022, 11, 373. [Google Scholar] [CrossRef]
- Dulyayangkul, P.; Douglas, E.J.A.; Lastovka, F.; Avison, M.B. Resistance to Ceftazidime/Avibactam plus Meropenem/Vaborbactam When Both Are Used Together Is Achieved in Four Steps in Metallo-beta-Lactamase-Negative Klebsiella pneumoniae. Antimicrob. Agents Chemother. 2020, 64, e00409-20. [Google Scholar] [CrossRef]
- Bianco, G.; Boattini, M.; Bondi, A.; Comini, S.; Zaccaria, T.; Cavallo, R.; Costa, C. Outbreak of ceftazidime-avibactam resistant Klebsiella pneumoniae carbapenemase (KPC)-producing Klebsiella pneumoniae in a COVID-19 intensive care unit, Italy: Urgent need for updated diagnostic protocols of surveillance cultures. J. Hosp. Infect. 2022, 122, 217–219. [Google Scholar] [CrossRef] [PubMed]
- Ransom, E.; Bhatnagar, A.; Patel, J.B.; Machado, M.-J.; Boyd, S.; Reese, N.; Lutgring, J.D.; Lonsway, D.; Anderson, K.; Brown, A.C.; et al. Validation of Aztreonam-Avibactam Susceptibility Testing Using Digitally Dispensed Custom Panels. J. Clin. Microbiol. 2020, 58, e01944-19. [Google Scholar] [CrossRef] [PubMed]
- Yu, W.; Luo, Q.; Shen, P.; Chen, Y.; Xu, H.; Xiao, Y.; Qiu, Y. New options for bloodstream infections caused by colistin- or ceftazidime/avibactam-resistant Klebsiella pneumoniae. Int. J. Antimicrob. Agents 2021, 58, 106458. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Karampatakis, T.; Tsergouli, K.; Behzadi, P. Carbapenem-Resistant Klebsiella pneumoniae: Virulence Factors, Molecular Epidemiology and Latest Updates in Treatment Options. Antibiotics 2023, 12, 234. https://doi.org/10.3390/antibiotics12020234
Karampatakis T, Tsergouli K, Behzadi P. Carbapenem-Resistant Klebsiella pneumoniae: Virulence Factors, Molecular Epidemiology and Latest Updates in Treatment Options. Antibiotics. 2023; 12(2):234. https://doi.org/10.3390/antibiotics12020234
Chicago/Turabian StyleKarampatakis, Theodoros, Katerina Tsergouli, and Payam Behzadi. 2023. "Carbapenem-Resistant Klebsiella pneumoniae: Virulence Factors, Molecular Epidemiology and Latest Updates in Treatment Options" Antibiotics 12, no. 2: 234. https://doi.org/10.3390/antibiotics12020234
APA StyleKarampatakis, T., Tsergouli, K., & Behzadi, P. (2023). Carbapenem-Resistant Klebsiella pneumoniae: Virulence Factors, Molecular Epidemiology and Latest Updates in Treatment Options. Antibiotics, 12(2), 234. https://doi.org/10.3390/antibiotics12020234








