Abstract
Bacteria of the genus Bacillus and related genera (e.g., Paenibacillus, Alicyclobacillus or Brevibacillus) belong to the phylum Firmicutes. Taxonomically, it is a diverse group of bacteria that, to date, has not been well described phylogenetically. The group consists of aerobic and relatively anaerobic bacteria, capable of spore-forming. Bacillus spp. and related genera are widely distributed in the environment, with a particular role in soil. Their abundance in the agricultural environment depends mainly on fertilization, but can also depend on soil cultivated methods, meaning whether the plants are grown in monoculture or rotation systems. The highest abundance of the phylum Firmicutes is usually recorded in soil fertilized with manure. Due to the great abundance of cellulose in the environment, one of the most important physiological groups among these spore-forming bacteria are cellulolytic bacteria. Three key cellulases produced by Bacillus spp. and related genera are required for complete cellulose degradation and include endoglucanases, exoglucanases, and β-glucosidases. Due to probable independent evolution, cellulases are encoded by hundreds of genes, which results in a large structural diversity of these enzymes. The microbial degradation of cellulose depends on its type and environmental conditions such as pH, temperature, and various substances including metal ions. In addition, Bacillus spp. are among a few bacteria capable of producing multi-enzymatic protein complexes called cellulosomes. In conclusion, the taxonomy of Bacillus spp. and related bacteria needs to be reorganized based on, among other things, additional genetic markers. Also, the ecology of soil bacteria of the genus Bacillus requires additions, especially in the identification of physical and chemical parameters affecting the occurrence of the group of bacteria. Finally, it is worth adding that despite many spore-forming strains well-studied for cellulolytic activity, still few are used in industry, for instance for biodegradation or bioconversion of lignocellulosic waste into biogas or biofuel. Therefore, research aimed at optimizing the cellulolytic properties of spore-forming bacteria is needed for more efficient commercialization.
1. Introduction
Bacillus spp. and related genera, including Paenibacillus, Alicyclobacillus, or Brevibacillus, are mostly Gram-positive and have the ability to produce spores and display metabolic capabilities under aerobic as well as relatively anaerobic conditions (Figure 1). Due to their characteristics, bacteria of this group have high resistance to environmental stresses such as drought, water stress, UV radiation, or low nutrient content in the environment [1,2]. Bacillus spp. and related genera commonly populate the Earth and occur in a variety of environments of both natural and anthropogenic origin [2,3,4].
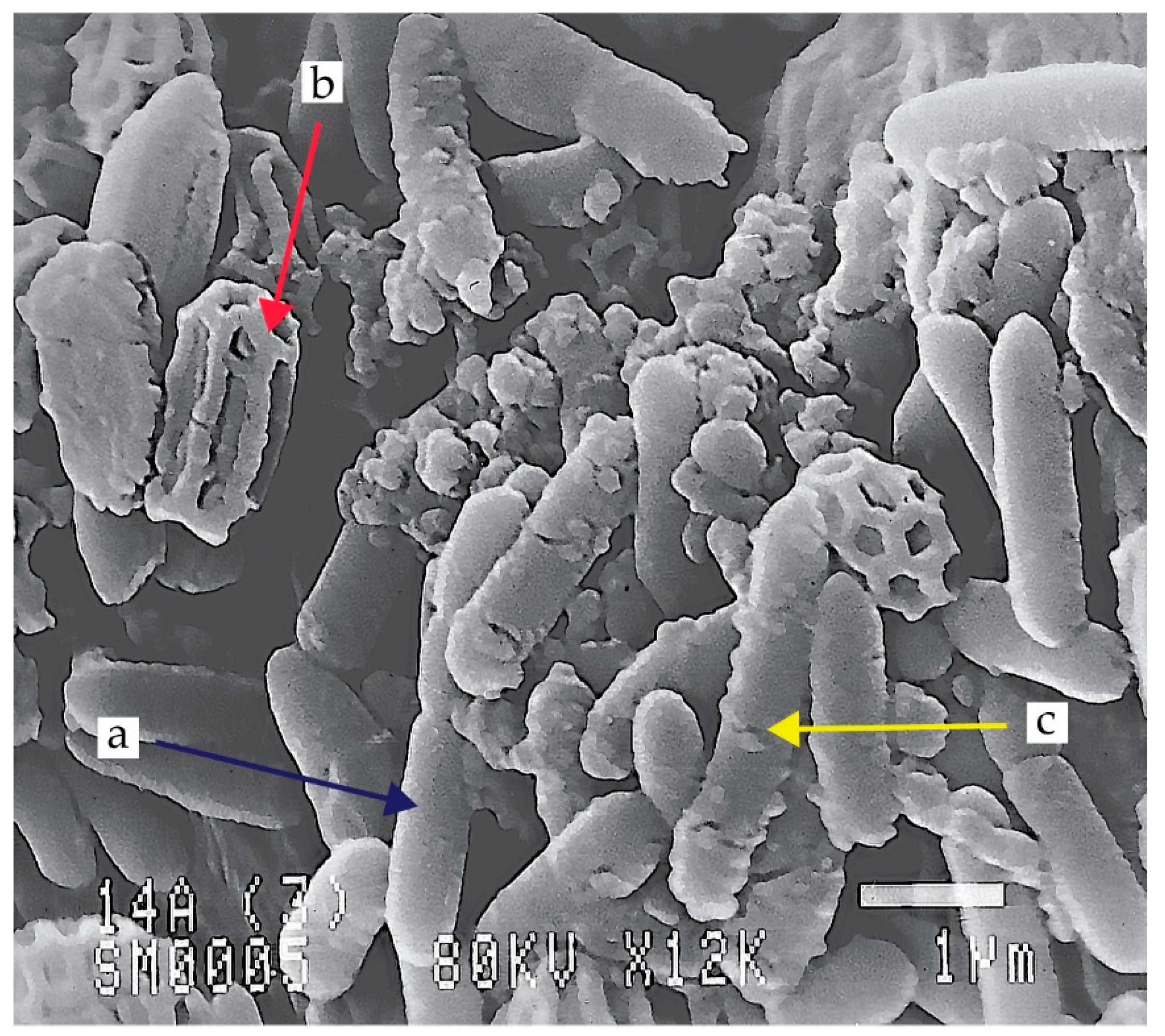
Figure 1.
Scanning electron microscope (SEM) image of Paenibacillus polymyxa EG14 cultivated on medium with 0.5% cellobiose: vegetative cell (a), endospores (b), cellulosomes (c) (own photo).
In recent decades, rapid population growth has resulted in a significant intensification of agriculture, which has contributed to environmental pollution affecting both community structure and physiology of most microbial groups in the soil [5,6,7]. Because of the high cellulose abundance, organisms that have cellulolytic activity gained importance. Cellulases are synthesized by almost all groups of systematic organisms including microorganisms such as bacteria, fungi, protists, plants, and nematodes [8]. The bacteria capable of producing cellulolytic enzymes include both aerobic bacteria, e.g., Butyrivibrio spp. and Cellulomonas spp., as well as anaerobic bacteria, e.g., Clostridium spp. or Ruminococcus spp. bacteria [9]. However, due to their resistance to unfavorable environmental conditions, aerobic and relatively anaerobic, spore-forming bacteria of the phylum Firmicutes (i.e., bacterial strains of the genus Bacillus and related genera) are the most interesting [10]. So far, hundreds of cellulolytic spore-forming strains belonging to the phylum Firmicutes were isolated, including the genus Bacillus (e.g., B. subtilis) [11]; the genus Alicyclobacillus (e.g., A. cellulosilyticus [12] and A. acidocaldarius [13]); the genus Geobacillus (e.g., Geobacillus sp. HTA426) [14], or the genus Lysinibacillus (e.g., Lysinibacillus fusiformis) [15]. Despite the large number of isolated strains, still only a small part of them is commercialized, e.g., in the biodegradation of lignocellulosic waste.
The aim of the review is to summarize current knowledge of the taxonomy, ecology and properties of cellulolytic bacteria and to find gaps, the filling of which may lead to a better understanding of the ecology of Bacillus spp. and related genera, improving taxonomy and to a better exploitation of the cellulolytic potential of the bacteria group.
2. Taxonomy of the Genus Bacillus and Related Genera
The genus Bacillus and related genera (e.g., Paenibacillus or Alicylobacillus) are a very diverse group of bacteria that belongs to the phylum Firmicutes. The phylum includes several classes such as Bacilli, Clostridia, Mollicutes, and Erysipelotrichia. The group of bacteria which is the subject of the review belongs to the class Bacilli. Currently, it is classified into several families including Bacillaceae, Paenibacillaceae, etc. [16,17]. However, earlier, there was only the genus Bacillus which was first described in 1874. One of the first species specified in the genus Bacillus is type species-B. subtilis. The species is also one of the best-studied organisms belonging to the prokaryota and thus is extensively used as a model microorganism for Gram-positive bacteria. Also, in the past, B. subtilis was the model organism in studies conducted to understand spore formation mechanisms [18,19]. However, despite numerous and extensive studies on Bacillus and related bacteria, the overall phylogenetic and evolutionary history of these genera remains unclear and relatively unexplored.
Initially, bacteria were identified by phenotypic methods using light microscopy and staining techniques including Gram staining [20]. Other older techniques that remain helpful nowadays include evaluation of bacterial biochemical properties, for example determining the metabolic profile, which can be used to differentiate between bacterial species. An example of such identification methods is the API® 50 CHB/E system, which is based on 50 biochemical tests that test the carbohydrate metabolism of the data from Bacillus spp. and related genera [21]. On the other hand, an improved version for bacterial identification using rapid tests is the Biolog OmniLog System. In addition to carbon source metabolism, the method also includes 23 chemical tests that determine, for example, the bacteria’s tolerance to salinity or sensitivity to other chemicals [22]. In both cases, the obtained results can be compared with databases and, to some extent, determine the taxonomic affiliation of the studied bacteria. However, it was not until the development of sequencing techniques in the 1990s that major changes in the taxonomy of spore-forming bacteria occurred. Then, other genera began to be separated from the genus Bacillus. Most phylogenetic studies are based on 16S rRNA gene sequences [23]. Based on branching in phylogenetic trees, initial phylogenetic studies delineated and identified five clusters of Bacillus species [24]. One of these clusters including B. subtilis was named Bacillus sensu stricto [24], while bacterial species from the other clusters were subsequently reclassified to form the following genera: Paenibacillus, Lysinibacillus Brevibacillus, and Geobacillus [23,25,26]. In subsequent years, based on phylogenetic and phenotypic results, many other Bacillus species were reclassified to form several new genera, for instance Aneurinibacillus, Alicyclobacillus, Alkalicoccus, Sporosarcina, Gracilibacillus, Virgibacillus, Hydrogenibacillus, Ureibacillus, Solibacillus. [23,25,27,28]. The bacterial genera listed above belong to different families, as shown in Table 1. For instance, Alicylobacillus, along with Tumebacillus, Effusibacillus, Kyrpidia, and Sulfobacillus, have been assigned to the family Alicylobacillace; a particularly important genus among those listed is Alicylobacillus [29,30]. On the other hand, the Paenibacillace family includes 14 genera of spore-forming bacteria of which Paenibacillus and Brevibacillus are the most interesting in terms of potential industrial use [31].

Table 1.
List of families assigned to the order Bacillales [17,23,29,32,33,34,35,36,37,38,39].
Although studies using sequences coding 16S rRNA have led to the reclassification of many species to new genera, according to many researchers, analyzes based on this variable gene are not fully sufficient to correctly distinguish taxa at the species level [40,41,42,43]. Similarly to other taxa, the previous classification of order Bacillales and other related orders was mostly based on 16S rRNA gene sequences. Moreover, research based on this type of analysis contributes to the formation of various types of anomalies. The occurrence of anomalies among the order is confirmed by the fact that several families and genera forming spores and non-spores were placed in it. Such patterns suggest that one gene marker is not sufficient to determine the phylogenetic structure of the Bacillales order [41]. Phylogenetic analyses have also been carried out using several other gene or protein sequences [44,45,46]. However, due to the relatively small number of Bacillus species studied in these researches, the analysis is insufficient to elucidate species relationships within this large genus. Consequently, Bacillus spp. is still a highly heterogeneous genus characterized by extensive polyphyletic branching with other genera of the family Bacillaceae [47,48]. Furthermore, as a result of the diverse branching of current species in the genus Bacillus, it was difficult to limit the addition of new species to this genus, even despite the large differences between the new species and the type species. Therefore, more valid methods should be studied and used to delineate the genus Bacillus and limit the placement of unrelated species within it [23]. For instance, comparative analysis of whole genomes (based on NCBI available sequences/genomes) makes it possible to study the evolutionary relations of species, and thus provide opportunities to identify molecular markers (molecular synpomorphies) [23,49]. For example, molecular synapomorphies that contain conserved insertions and signature deletions in protein sequences are good means of differentiating species from the two major clades of the genus Bacillus, i.e., the “Subtilis clade” and the “Cereus clade”. According to ICNP rule 56a, the transfer of a species from the Cereus clade to a new genus may play some part in human health; therefore, transfer to another species is not advisable. As evidenced by a comprehensive genomic analysis of Bacillaceae species, 36 new genetic markers (i.e., conserved signature indels (CSIs)) were detected [23]. Importantly, based on new CSIs, the monophyletic groups found in all reconstructed or new phylogenetic trees were named as follows: Simplex, Firmus, Alcalophilus, Niacini, Fastidiosus, and Jeotgali clades, and collectively included from 5 to 23 Bacillus species. In addition, researchers also performed a phylogenomic analysis on various Firmicutes proteins including core and conserved proteins. Moreover, the combined sequences of highly conserved proteins such as GyrB, GyrA, RpoC, RpoB, UvrD, or PolA were also studied, and confirmed by an extended comparative analysis of the genome of the above-mentioned protein sequences [23]. The authors of this study, based on robust evidence from many lines of research (conducted in parallel) confirming the existence of six distinct Bacillus clades, propose the transfer of species from these clades to six novel genera of Bacillaceae family, namely Alkalihalobacillus gen. nov., Cytobacillus gen. nov., Mesobacillus gen. nov., Neobacillus gen. nov., Metabacillus gen. nov., and Peribacillus gen. nov. [23]. Moreover, as a result of the creation of these new genera, 103 erroneously assigned species, that were insufficiently related to the genus Bacillus, were assigned to the new genera. The results above constitute an important step in elucidating the taxonomy of the Bacillus spp. and related genera. However, as indicated above, comprehensive studies are still needed for the correct classification of Bacillus spp. and related species.
3. Occurrence of Spore-Forming Bacteria in Arable Soils
Bacteria of the genus Bacillus and related genera are widely distributed in the environment, e.g., in soil, air, water, animals, plants, or sediments [50,51,52,53]. This group of bacteria plays a particularly important role in the soil, including the decay of matter [54], promotion of plant growth, and protection against phytopathogens.
A very good and widely used tool for assessing the abundance of bacteria is the next-generation sequencing (NGS), including 16S rRNA genes sequencing. However, due to the still existing limitations of sequencing technologies, most studies present the abundance of bacteria at high taxonomic levels, i.e., phyla or orders, and rarely present the abundance of bacteria at the genus level, which is a subject to much greater error [55,56].
The phylum Firmicutes is one of the dominant phyla in cultivation soils. Its relative abundance in the soil ranges from 2% to about 20% depending on agrotechnical practices used, including crop rotation systems and fertilization type [57,58,59,60,61]. In general, the Firmicutes type is more abundant in soils from crop rotation than in soils from continuous cropping [62,63,64]. The reason for these patterns is probably a greater influence of crop residues and decomposing roots in the soil from crop rotation compared to monoculture soil. For instance, in a greenhouse experiment, Li et al. [59] detected a higher number of sequences assigned to the phylum Firmicutes in soil (Mollisol with sandy loam texture) derived from rotation (tomato/potato-onion) compared to monoculture (tomato). The same patterns were noted for the genus Bacillus. The abundance values obtained by the authors at the level of the phylum Firmicutes and the genus Bacillus did not exceed 10%. However, there are also cases where more Firmicutes are detected in monocultures than in rotations, or in longer monocultures than shorter ones. For example, in the soil from the Morrow Plots experiment (USA), the relative abundance of the phylum Firmicutes ranged from a few to a maximum of 14%. Its abundance was dependent on soil management; in this case, the highest value was recorded in soil from a maize monoculture, while the lowest abundance of sequences assigned to the phylum Firmicutes was noted in soil from a maize-soybean rotation [6]. Similarly, Zhao et al. [60] observed a significantly increased number of sequences belonging to phylum Firmicutes in soil from 15- and 22-year continuous cropping of cucumber in comparison with cucumber grown for only one year. Earlier, Zhao et al. [65] noted similar patterns in continuous cropping of coffee. However, these authors did not find specific reasons for this phenomenon [60,65]. Hence, further studies are needed to find parameters that have a considerable role in shaping the abundance of the phylum Firmicutes including Bacillus spp. and related genera, e.g., identifying detailed correlations between physical and chemical properties of the soil and the abundance of bacteria belonging to the phylum Firmicutes in differently managed soils. For example, Alami et al. [66] observed robust correlations between the phylum Firmicutes and the physicochemical properties of arable soil including continuous cropping of maize and cabbage continuous cropping of cabbage (Hubei province, China); total phosphorus, available potassium, and available boron contents were positively correlated with the phylum Firmicutes. Furthermore, a study on the effect of continuous cotton cultivation (20 years) on the bacterial communities of the soil showed a positive correlation between the number of the OTUs of the phylum Firmicutes and the EC of the soil [66].
In addition, fertilization also affects the abundance of the phylum Firmicutes in the soil. Particularly because most members of the phylum Firmicutes are considered copiotrophs which are fast-growing microorganisms that prefer environments rich in C and N [67]. For instance, Li et al. [68] also found a several percent abundance of the phylum Firmicutes in fertilized soil (rice-rape rotation), and the highest number of OTUs belonging to the phylum was found in soil fertilized with NPKS (NPK + straw). Similar values were also found by Zeng et al. [58] who observed an abundance of the phylum Firmicutes at an average of 7% (the highest value was 10%) in soil fertilized with nitrogen fertilizer. Dang et al. [69] observed a significant increase in the abundance of Firmicutes in soil fertilized with manure (compared to the controls) across the globe, and detected a positive correlation between the SOC content and the abundance of the phylum. Furthermore, Francioli et al. [70] noted more OTUs assigned to the phylum Firmicutes in farmyard manure (FYM) fertilized soils compared to mineral fertilization (in a long-term fertilization trial). Hartmann et al. [71] also observed higher abundance of the phylum Firmicutes in long-term FYM fertilization in comparison with mineral fertilization. Similar findings were noted in a study on the effects of various treatments on the microbial community of bulk and rhizosphere soil [72]. Importantly, it was also found that manure fertilization is a factor influencing the bacterial community (including the abundance of Firmicutes) more strongly than the method of cultivation, including monoculture and crop rotation [31,73].
In conclusion, it should be noted that the abundance of bacteria of the phylum Firmicutes in soil may also be influenced by other agronomic treatments such as the use of plant protection agents. Thus, the study results may also have been caused by the heterogeneity of agricultural practices, as previously recorded by Soman et al. [6]. Moreover, the discrepancies in studies in this aspect may be an effect of the diversity of soils around the world, e.g., in terms of physical properties.
4. Cellulolytic Properties of Bacillus Spp. and Related Genera
4.1. Cellulases
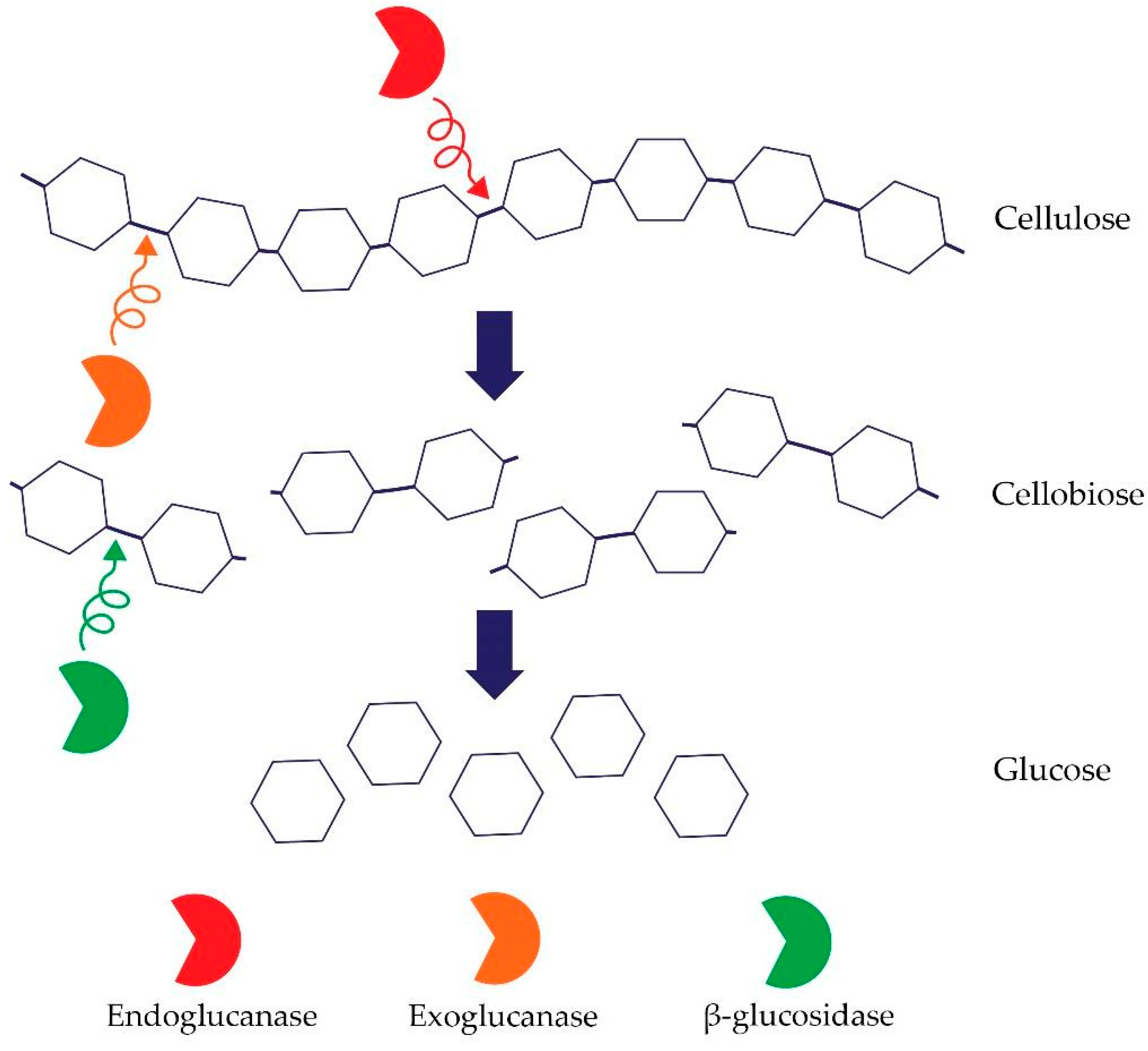
Cellulose is the most common (bio)polymer on earth, made of glucose linked by β-1,4-glycosidic bonds. It contains two types of regions—crystalline and amorphous regions [74]. Hence, an important group of microbes that are participating in the element’s circulation in the soil are microorganisms that decompose cellulose [75]. Soil properties such as pH, organic carbon content, nitrogen content, and moisture impact microbial cellulose degradation. The process of cellulose degradation depends on the presence of a complex of enzymes belonging to the class of O-glycoside hydrolases, including the three main cellulases [74]. Cellulolytic enzymes include: (i) endo-β-1,4-glucanases (EC 3.2.1.4) whose mechanism of action is based on random degradation of β-1,4-glycosidic bonds in amorphous regions of cellulose–endoglucanase activity is measured using cellulose derivatives, for instance, semi-soluble carboxymethylcellulose (CMC); the enzyme that degrades CMC is carboxymethylcellulase (CMCase); (ii) exo-1,4-β-glucanases (EC 3.2.1.91) that separate single molecules of glucose and cellobiose from reducing or non-reducing ends of the cellulose. Exoglucanases include e.g., avicelase–microcrystalline–cellulose (Avicel) degrading enzyme; and (iii) β-glucosidase whose mechanism of action is the conversion of cellobiose into glucose (EC 3.2.1.21) [76,77]. The synergistic cooperation of the above-mentioned enzymes and, in particular, the presence of a processive exoglucanase is required for cellulose degradation [Figure 2].
Figure 2.
Mechanism of action of cellulases (own figure).
Cellulases have a modular structure that contains catalytic modules (CM) that act synergistically and/or with non-catalytic modules, i.e., substrate-binding modules [78]. In the case of cellulases, substrate-binding modules are called cellulose–binding modules (also called carbohydrate-binding module) (CBMs) and they can have affinity for amorphous or crystalline cellulose as well as binding to other similar polymers composed of carbon chains [79]. Due to the enzyme’s ability to bind to cellulase, the local concentration of the enzyme increases, resulting in better substrate degradation efficiency. Some CBMs also have a structural function of stabilizing the catalytic module or altering its activity, for instance by inserting a substrate molecule into a substrate pocket [79]. Although the binding of cellulase by CBM is very stable, the enzyme can still diffuse across the substrate surface and, in some cases, CBM can also catalyze the breaking of non-covalent bonds located between the cellulose chains of crystalline cellulose [80].
4.2. Structural Diversity of Cellulases
Cellulases belong to the glycoside hydrolases (GH). The classification of GH is based on similarities in amino acid sequence and is included in the carbohydrate-active enzymes (CAZy) database. The CAZy database contains the CAZy families and subfamilies and is very dynamically updated. Due to the large differences in amino acid sequences, the GH group is remarkably heterogeneous and is divided into as many as 165 families [81]. The enzymes involved in cellulose degradation are classified in the following families of glycoside hydrolases: endoglucanases in families 5, 6, 7, 8, 12, 44, 45, 48, 51, 64, 71, 74, 81, 87, 124, 128, exoglucanases 5, 6, 7 and 4, and β-glucosidases in families 1, 3, 4, 17, 30, and 116 [82,83].
Sequence diversity can be related to a distinct modular architecture. It has been shown that the domain architecture in fungi is not very complex. However, in bacterial cellulases, there are many combinations of domain architectures, even though most sequences consist of a single catalytic domain [84]. In terms of carbohydrate-binding modules, CBM2 is related to the cellulolytic GH families and is found in the following families–GH5, GH12, GH44, GH45, GH48, GH51, and GH74. So far, it has been shown that the common CBM2 domain (in bacterial cellulases) in most cases binds cellulose, and less often chitin and xylan. Importantly, CBM2 is often found together with other accessory domains including CBM3 and CBM4, as well as catalytic domains [85]. In the terms of bacteria of the genus Bacillus, B. licheniformis possesses the H1AD14 gene encoding an endoglucanase belonging to the GH9 family and the cellulase has a CBM3 domain that is attached to the C-terminal end and plays a significant role in substrate degradation [86]. CBM3 has also been detected in a cellulase belonging to the GH9 family in B. pumilus [87]. Interestingly, Honda et al. [88] found that a unique chitinase domain in B. thuringiensis enabled binding to both crystalline chitin and cellulose, indicating that CBMs with affinity to multiple substrates could contribute to the increased occurrence of multifunctional hydrolytic enzymes [88]. On the other hand, previously, in the B. subtilis IFO 3034, an endoglucanase was detected that possessed a microcrystalline cellulose-binding domain but was unable to degrade microcrystalline cellulose [89]. Also in species related to Bacillus spp. CBMs were detected; in the Paenibacillus lautus BHU3 as many as four domains were detected, including CBM6, CBM46, CBM56, and CBM9, showing affinity for amorphous cellulose [90]. The CBM9 was also found in the genome of P. dendritiformis CRN1 [91].
Furthermore, cellulases belonging to different families have various protein fold structures, including the (β/α)8 barrel fold, which is found in the GH5, GH44 and GH51 families, modified α/β barrel in family GH6, β-jelly roll—GH7 and GH12, the 7-fold β-propeller (GH74), (α/α)6 barrel—GH8, GH9 and GH48, the superhelical fold—GH124, and modified β barrel (GH45) [81]. Importantly, within a single GH family, structures are globally conserved, but sequences can be remarkably different. For example, GH5, one of the largest GH families, is currently divided into 166 subfamilies on the basis of sequence similarity, with only eight residues conserved across the family, including two catalytic glutamic acid residues [92]. In conclusion, GH families exhibiting different classes of protein folds have evolved to bind and degrade the same substrate, indicating that cellulolytic enzymes may have evolved independently and may be derived from many evolutionary origins, but have converged functionally [81]. Similar patterns regarding the evolution of cellulases can be inferred from the large number of cellulose-binding domains.
4.3. Cellulases Genes
Referring to the number of cellulases, it can be concluded that cellulolytic enzymes are highly diverse, which is further manifested in the large number of genes that are responsible for encoding these hydrolases. The number of genes encoding cellulases exceeds 100 [83,93]. As mentioned earlier, the reason for such a large number of cellulase-encoding genes may be due to independent evolution [81]. In fungi, the genes encoding cellulases in bacteria are located on a chromosome [94]. The spatial organization of these genes may differ between microorganism species, for example, in the bacterial species Clostridium thermocellum there is a random distribution, whereas in C. cellulovorans “clustered” distribution in a cluster occurs [95,96]. The cellulosome gene cluster in C. cellulovorans is about 22 kpz in size and contains nine genes encoding cellulosome domains with a putative transposon gene in the flanking region. A similar organization was also detected in the chromosome of the bacterial species C. acetobutylicum and C. cellulolyticum, suggesting the presence of a common bacterial ancestor of the clostridia [97]. In contrast, in fungi the genes encoding cellulases are usually distributed randomly, in which case each gene has its own transcriptional regulation. Only in exceptional cases, e.g., in Phanerochaete chrysosporium (fungus), the cellulase genes form a three-gene cluster [94].
In terms of Bacillus spp., in the genome of B. licheniformis [98] detected two clusters of genes involved in the cellulose decomposition. For instance, in the genome of the strain B. subtilis 168 no equivalents of the cluster were found. The enzymes encoded by the first gene cluster are likely endoglucanases belonging to the GH9 and GH5 families, and the probable cellulase–1,4-β-cellobiosidase belonging to GH48 and the potential β-mannanase belonging to GH5. Importantly, β-mannanase (GH5) and endoglucanase (GH9) contain carbohydrate-binding modules. In addition, with the exception of 1,4-β-cellobiosidase belonging to GH48, all gene proteins encoded have secretory signal peptides and all have homologs with Bacillus spp. but other than B. subtilis [98]. Researchers also detected a second cluster—encoding a probable β-glucosidase (from the family GH1). In addition, a second β-glucosidase gene (from the family GH3) was found at an unrelated locus in the genome. Importantly, the presence of these genes in the B. licheniformis genome indicate the possibility of complete degradation of cellulose [98]. Furthermore, 4 genes responsible for encoding β-glucosidase and 1 gene encoding endoglucanase were noted in strain B. amyloliquefaciens TL106. The β-glucosidases encoded by these genes belong to the GH1 and GH73 families, and the endoglucanase belongs to the GH5 family [99]. Moreover, Carbonaro et al. [100] analyzed the genome of Alicyclobacillus mali FL18 to find new cellulose–degrading enzymes. The analysis revealed four genes belonging to the GH1, GH9, GH51, and GH94 families, of which GH1 and GH94 legitimately hydrolyse short oligosaccharides, and a gene from GH1 encodes a β-glucosidase. In addition, the A. mali FL-18 genome also contained genes encoding two probable arabinofuranosidases, which belong to GH51. Interestingly, A. acidocaldarius, which is a close relative of the aforementioned species, also possesses two endoglucanases–CelA belonging to the GH9 family and CelB from the GH51 family [101,102]. At the same time, other authors have detected a large number of genes encoding various GH enzymes in the P. polymyxa genome, including cellulases belonging to GH 1, 3, and 5 [103].
Finally, it is worth adding that most of the cellulases described in metagenomic studies (different environments) have less than 70% homology with known cellulolytic enzymes, and some of them have no significant similarity to other glycosyl hydrolases, indicating that large numbers of new cellulolytic enzymes are still being found [104]. Moreover, approximately 40% of sequenced bacterial genomes contain at least one cellulase gene, but only 4% of these bacteria are known as true cellulase bacteria due to low cellulase diversity or a lack of gene expression [83].
4.4. Cellulosomes
Some bacteria exhibiting cellulolytic activity are capable of synthesizing and secreting enzyme multicomplexes called cellulosomes; the secreted proteins outside the bacterial cell take the form of spherical structures. Sometimes, individual cellulosomes are joined together to form so-called polycellulosomes. A single complex may contain up to a dozen proteins with different activities, including endoglucanase, cellobiase or hemicellulase, and lichenase [105,106]. Interestingly, most of the research on cellulosomes concerns the cellulosomes of the phylum Firmicutes [83]. First studies on cellulosomes were carried out on the anaerobic Clostridium thermocellum. As shown, C. thermocellum is capable of synthesizing an enzyme complex of more than 2000 kDa, which consists of fourteen different proteins with molecular weights ranging from 45 kDa to 210 kDa [107]. For instance, 15 genes encoding the presence of endoglucanases, two genes responsible for the expression of xylanases, two genes encoding cellobiase, and one gene encoding lichenase were detected in C. thermocellum strain NC1B 10682 [108]. Cellulosomes have also been detected in Bacillus spp. and related genera. However, to date, little research has been conducted on these genera. For instance, B. megaterium was found to be capable of producing a cellulosome (celluloxylanosomes) exhibiting avicelase, CMCase, and xylanase activity. In addition, van Dyk et al. [105] noted that the B. licheniformis SVD1 strain was capable of synthesizing multi-enzyme complex (MEC) with hemi-cellulolytic activity. The total molecular mass of the complex was about 2000 kDa. The enzymes included in the MEC hydrolyzed such compounds as xylan, mannose, pectin, and carboxymethylcellulose. However, the MEC was not able to bind Avicel cellulose and, despite several similarities to the cellulosome, was ultimately not identified as such [105,109]. Waeonukul et al. [110] studied an enzymatic complex from P. curdolanolyticus B-6 (in culture on Avicel microcrystalline cellulose). A single cellulosome had the ability to hydrolyze the Avicel cellulose and insoluble xylan. The researchers noted that the complex included such enzymes as avicelase, CMCase, cellobiohydrolase, β-glucosidase, xylanase, α-L-arabinofuranosidase, and β-xylosidase. The total mass of the multicomplex was about 1600 kDa. Importantly, the isolated cellulosome degraded lignocellulose efficiently. In terms of the genus Paenibacillus, using transmission microscopy, cellulosome production was detected in P. polymyxa strains EG2 and EG14 [111,112]. Besides, using scanning electron microscopy, protuberances were observed indicating cellulosome production on the cell surface in the thermophilic strain Brevibacillus sp. JXL [113].
4.5. Cellulase Activity
Cellulose decomposition starts when cellulase adsorbs to cellulose. Referring to previous subsections, it should be stated that bacteria of the genus Bacillus and related genera are capable of producing several types of cellulases including CMCase, FPase, or Avicelase. Different bacterial species have distinct activities of cellulolytic enzymes, and significant differences within the same species or strains may also occur due to discrepancies in culture conditions of the studies conducted on the topic. For instance, Acharya and Chaudhary [114] observed a CMCase activity of 0.300 U mL−1 in Bacillus licheniformis MVS1 (medium with beef extract). While Shajahan et al. [115], using response surface methodology in Bacillus licheniformis NCIM 5556, recorded a CMCase activity of 42.99 U mL−1 (medium contained CMC—19.21 g L−1, CaCl2—25.06 mg L−1, Tween 20—2.96 mL L−1, and temperature 43.35 °C).
The type of cellulose, medium composition, temperature and pH are most important for cellulase activity [2]. So far, depending on the strains, it has been found that the type of cellulose used as substrate induces cellulolytic activity to a different extent. For instance, Sadhu et al. [116] observed that carboxymethylcellulose better induced Avicelase and CMCase production by Bacillus sp. MTCC10046 compared to other substrates including sucrose, starch, glucose, or maltose. Also, Akaracharanya et al. [117] recorded higher cellulase activity of Bacillus sp. P3–1 and P4–6 in culture based on CMC medium, compared to culture with cellulose powder-containing medium. Similar patterns were also reported by Thomas et al. [118] who observed that CMCase activity by Bacillus sp. SV1 was higher in the CMC medium, compared to Avicel cellulose-containing medium and other carbon sources, including mannitol, glycerol, lactose, or chitin. In addition, CMCase and Avicelase activities were also obtained by Dobrzynski et al. [2] who noted the highest activity of the two enzymes in the cultures of Bacillus sp. 8E1A with CMC. However, in the case of FPase (cellulose saccharifying enzyme), the highest activity value was recorded for the culture of the studied strain with Avicel cellulose. Mihajlovski et al. [119] also reported slightly higher FPase activity in P. chitinolyticus CKS1 in a medium supplemented with Avicel compared to cultures with CMC. Similarly, in the case of thermophilic Bacillus sp. K-12, Kim and Kim [120] noted that FPase activity was higher when the strain studied by the authors was cultured in Avicel microcrystalline cellulose medium compared to other carbon sources. Interestingly, in contrast to previously cited reports, the strain Bacillus sp. K-12 also had high CMCase and Avicelase activity in cultures with Avicel cellulose. It is worth mentioning that the differences between studies may result from a number of factors including culture conditions.
Another important factor that affects the activity of cellulases produced by Bacillus spp. and related genera is temperature. According to the studies cited below, the optimum temperature range for cellulase activity ranges from 20 °C to 80 °C, depending on the strain and type of enzyme. For instance, Kazeem et al. [121] observed that a temperature of 20 °C is optimal for the production of FPases in the strain B. licheniformis 2D55. Cellulases produced by B. pseudomycoides (grown on sugarcane bagasse medium) have a slightly higher optimal temperature −40 °C (within 72 h of incubation) [122]. Interestingly, Li et al. [123] detected optimal cellulase activity in the thermophilic strain at 50 °C, and below this value the activity of enzymes significantly decreased. On the other hand, optimum temperature values for cellulase activity exceeding 70 °C have been recorded for activity of CMCase and Avicelase produced by Geobacillus thermoleovorans T4 (70 °C) and CMCase produced by Bacillus sp. DUSELR13 (75 °C) [124,125]. Similar patterns for Bacillus sp. 8E1A were observed by Dobrzyński et al. [2]. Importantly, thermophilic cellulases can potentially be used in various industries including textile, biofuel, and agriculture [2].
In terms of the optimal pH for cellulase activity, the range of values is as wide as for temperature; according to current reports, the highest activity of cellulases produced by Bacillus spp. and related genera is recorded in the pH range from 3 to 10. For example, Mihajlovski et al. [119] observed that the avicelase produced by the strain was most active at about pH 5. Similar results were reported by Seo et al. [126] whose B. licheniformis strain produced cellulases with high activity in the pH range of 4.0–6.0. While, in a study by Dobrzynski et al. [2], the highest CMCase and Avicelase activities were noted at pH 7.0 and FPase at 6.0. Interestingly, the highest cellulase activities produced by the bacteria of the genus Bacillus were also detected at pH 9.0 [127]. Previously, similar patterns were also obtained, as shown in Table 2.

Table 2.
Optimum temperature and pH for celullolytic activity.
Importantly, the differences between the optimal conditions for the activity of cellulolytic enzymes result from the large variety of cellulases produced by the spore-forming bacteria of the genus Bacillus and related genera.
Moreover, the activity of cellulases is also affected by other parameters of the media or solutions. Gaur and Tiwari [135] found that the cellulase activity of B. vallismortis RG-07 was stimulated by Tween-60, Ca2+, mercaptoethanol, and NaClO. While the cellulase activity of Lysinibacillus xylanilyticus was stimulated by the presence of CaCl2 nanoparticles in medium [136].
Importantly, some of the spore-forming strains of cellulolytic bacteria are already being used to convert lignocellulosic waste. For instance, the activity of P. polymyxa ND24 was studied in a 5-L laboratory bioreactor where the cellulosic substrate in the medium was sugarcane bagasse; the strain showed the highest endoglucanase activity after 72 h of incubation. The sugarcane hydrolysate was then used for biogas production; the authors suggest that the obtained results support the use of P. polymyxa ND24 for cost-effective bioprocessing of lignocellulosic biomass [137]. In turn, other authors have used strains from the genus Bacillus to treat rice straw in order to increase the biomethane fermentation efficiency. The study, using multiple strains, demonstrated that the use of mixtures of different bacterial strains was more effective than the use of single bacterial strains, due to an increase in the pool of cellulases present in the process. Finally, the authors concluded that the choice of a mixture of strains from the genus Bacillus, which decompose lignocelluloses, can be robust catalysts for the processing of biomass from these wastes [138].
However, despite such a large number of bacterial strains of the genus Bacillus and related ones that produce cellulases, there is still little research on the practical aspect of their use, including the utilization and conversion of lignocellulosic biomass. Nevertheless, potentially, cellulolytic bacteria of the genus Bacillus spp. and their cellulases can be used: (i) in the textile industry (for instance for biostoning of jeans); (ii) in biorefining; (iii) in biogas and biofuel production; (iv) in agriculture including biodegradation of lignocellulosic waste and biocontrol of fungal phytopathogens; (v) in the paper industry (coadditive in pulp bleaching); (vi) in detergents (cellulose-based detergents); (vii) in the food industry including release of the antioxidants from fruit and vegetables, and improved texture and quality of bakery products; (viii) and for improving carotenoids extraction or improving olive oil extraction [139,140].
5. Promoting Plant Growth by the Bacteria of the Genus Bacillus and Related Genera
Bacteria of the genus Bacillus and related genera are also classified as plant growth-stimulating bacteria [53,141,142,143]. Bacteria from this group are capable of promoting plant growth either directly or indirectly. Mechanisms of direct promotion of plant growth include i.a. production of phytohormones including indole-3-acetic acid (IAA), cytokinins, and gibberellins, production of nitrogenase thanks to which bacteria fix atmospheric nitrogen (N) and make it available to plants, and the possibility of solubilizing phosphorus. Indirect mechanisms, on the other hand, include for instance production of antibiotics including cyclic lipopeptides, and enzymes degrading fungal cell walls [144,145,146,147].
So far, plant growth-promoting abilities have been detected in a very large number of bacteria belonging to the genus Bacillus or related genera. Bacteria from this group have promoted plant growth both under controlled and field conditions. Because of the greater value of studies under field conditions, several examples of such studies are presented in the review. For instance, inoculation of rice seedlings with B. pumilus TUAT-1 supplemented with N fertilizer led to an increase in height, biomass, and chlorophyll content of rice plants [148]. Besides, Ali et al. [149] showed that B. cereus (potassium solubilizing strain) increased the plant’s height and shoots’ dry weight. Importantly, compared to plants that were not inoculated, the application of the strain resulted in an increase of about 20% in potato yield. Moreover, the application of Paenibacillus triticisoli BJ-18 led to an increase in N, P, and organic matter contents in soil and enhanced nitrogenase activity and wheat yield [150]. Interestingly, in comparison to the control, the application with the strain also increased the biodiversity of rhizosphere bacterial communities and led to an increase in the abundance of the genus Paenibacillus in the inoculated soil, which also resulted in a high abundance of genes encoding nitrogenases. Furthermore, the inoculation with P. triticisoli BJ-18 also increased the abundance of native plant growth-stimulating bacteria of the genera Bacillus and Podospora [150].
Besides Okoroafor et al. [151], after applying B. velezensis FZB42 (formerly B. amyloliquefaciens FZB4) in maize and common sunflower cultivations, detected over 20% increase in biomass production in each of the crops. Moreover, inoculation with the tested preparation increased the bioavailability of soil elements. Interestingly, the study on winter wheat cultivation by Stepien et al. [152] is an example of a field experiment with Bacillus and related bacteria. The researchers demonstrated that the combination of mineral fertilization and three bacteria-Paenibacillus azotofixans, B. megaterium, and B. subtilis-significantly increased wheat grain yield compared to the application of mineral fertilization alone. In addition, the bacteria significantly increased the leaf greenness index SPAD at two time points, and together with NPK fertilization, significantly increased the content of two forms of nitrogen (N-NO3 and N-NH4) and phosphorus in the soil.
Another example of research using a bacterial consortium with Bacillus spp. is an experiment using B. cereus AR156, B. subtilis SM21, and Serratia sp. XY21 (BBS) strains applied to phytophthora-infested sweet pepper [153]. Compared to the control, the application of BBS reduced the occurrence of phytophthora blight and enhanced the fruit quality and soil properties. BBS also significantly increased the abundance of the bacterial genera Burkholderia, Comamonas, and Ramlibacter, which were negatively correlated with disease severity; moreover, the abundance of these genera were associated with organic carbon, ammonia nitrogen, potassium, and available phosphorus. These patterns suggest that changing the bacterial community improved the soil properties and reduced the phytopathogen development.
Importantly, there are still not enough studies in field conditions, especially those showing the effect of the inoculants used on the native microbiota whose biodiversity and taxonomic composition have the greatest influence on the biochemical processes of the soil. Finally, field studies with a wide range of parameters will bring inoculants closer to commercialization. However, there are already quite a number of commercial preparations containing Bacillus and related bacteria, for example biofertilizers, biofungicides, or biopesticides (listed in Table 3).

Table 3.
Commercial preparations containing Bacillus and related bacteria.
6. Conclusions
In summary, bacteria of the genus Bacillus and related genera constitute an important group of bacteria that populate soil and other environments in large numbers, but their taxonomy is still inadequately defined, due to, among other things, their great diversity and the selection of insufficiently suitable molecular and biochemical techniques to determine their relationship. Among this group of bacteria, cellulolytic bacteria are one of the most important, but knowledge about their occurrence in the soil environment is still limited, which is caused by methodological difficulties faced by scientists studying it. Most studies on the presence of cellulolytic bacteria in the soil are limited to determining the abundance of genes encoding cellulase, which, due to the diversity of these genes, makes it impossible to determine the abundance of individual groups of cellulolytic bacteria.
Moreover, despite dozens of isolates of Bacillus and related bacteria showing cellulolytic activity, still few of these bacterial strains are used, for example, to degrade lignocellulosic waste. Importantly, the amount of lignocellulosic waste generated by agriculture and other industries is steadily increasing, which, in an era of progressive agriculture and other industries generating large amounts of such waste, poses a huge environmental problem. Therefore, researchers should focus on studying the cellulolytic bacteria, e.g., in biogasification processes or other conversions, which could contribute to the commercialization of these bacteria.
Author Contributions
Conceptualization, J.D.; methodology, J.D.; software, B.W.; writing—original draft preparation, J.D. and E.B.G.; writing—review and editing, J.D., B.W. and E.B.G.; visualization, B.W.; supervision, J.D. and E.B.G.; project administration, J.D.; funding acquisition, B.W. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Acknowledgments
Many thanks to Katarzyna Rafalska for help in revising the English language and Aleksandra Wróbel for help in data visualization.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Nicholson, W.L.; Munakata, N.; Horneck, G.; Melosh, H.J.; Setlow, P. Resistance of bacillus endospores to extreme terrestrial and extraterrestrial environments. Mol. Biol. Rev. 2000, 64, 548–557. [Google Scholar] [CrossRef] [PubMed]
- Dobrzyński, J.; Wróbel, B.; Górska, E.B. Cellulolytic Properties of a Potentially Lignocellulose-Degrading Bacillus sp. 8E1A Strain Isolated from Bulk Soil. Agronomy 2022, 12, 665. [Google Scholar] [CrossRef]
- Yakovleva, G.; Kurdy, W.; Gorbunova, A.; Khilyas, I.; Lochnit, G.; Ilinskaya, O. Bacillus pumilus proteome changes in response to 2, 4, 6-trinitrotoluene-induced stress. Biodegradation 2022, 33, 593–607. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Lin, Y.; Shen, G.Z.; Zhang, H.H.; Lyu, S.X. Siderophores of Bacillus pumilus promote 2-keto-L-gulonic acid production in a vitamin C microbial fermentation system. J. Basic Microbiol. 2022, 62, 833–842. [Google Scholar] [CrossRef]
- Gamrat, R.; Gałczyńska, M.; Brysiewicz, A. Preliminary assessment of the impact of soil microorganisms on greenhouse gas emissions expressed in CO2 equivalent and grass biomass. J. Water Land Dev. 2022, 53, 108–117. [Google Scholar] [CrossRef]
- Soman, C.; Li, D.; Wander, M.M.; Kent, A.D. Long-Term Fertilizer and Crop-Rotation Treatments Differentially Affect Soil Bacterial Community Structure. Plant Soil 2017, 413, 145–159. [Google Scholar] [CrossRef]
- Zielewicz, W.; Swędrzyński, A.; Dobrzyński, J.; Swędrzyńska, D.; Kulkova, I.; Wierzchowski, P.S.; Wróbel, B. Effect of Forage Plant Mixture and Biostimulants Application on the Yield, Changes of Botanical Composition, and Microbiological Soil Activity. Agronomy 2021, 11, 1786. [Google Scholar] [CrossRef]
- Zhang, X.-Z.; Zhang, Y.-H.P. Cellulases: Characteristics, sources, production, and applications. In Bioprocessing Technologies in Biorefinery for Sustainable Production of Fuels, Chemicals, and Polymers; Yang, S.-T., El-Enshasy, H.A., Thongchul, N., Eds.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2013; pp. 131–146. [Google Scholar]
- Poulsen, H.V.; Willink, F.W.; Ingvorsen, K. Aerobic and anaerobic cellulase production by Cellulomonas uda. Arch. Microbiol. 2016, 198, 725–735. [Google Scholar] [CrossRef] [PubMed]
- Deka, D.; Bhargavi, P.; Sharma, A.; Goyal, D.; Jawed, M.; Goyal, A. Enhancement of cellulase activity from a new strain of Bacillus subtilis by medium optimization and analysis with various cellulosic substrates. Enzym. Res. 2011, 2011, 151656. [Google Scholar] [CrossRef]
- Irfan, M.; Mushtaq, Q.; Tabssum, F.; Shakir, H.A.; Qazi, J.I. Carboxymethyl cellulase production optimization from newly isolated thermophilic Bacillus subtilis K-18 for saccharification using response surface methodology. AMB Express 2017, 7, 29. [Google Scholar] [CrossRef] [PubMed]
- Matsubara, H.; Goto, K.; Matsumura, T.; Mochida, K.; Iwaki, M.; Niwa, M.; Yamasato, K. Alicyclobacillus acidophilus sp. nov., a novel thermo-acidophilic, omega-alicyclic fatty acid-containing bacterium isolated from acidic beverages. Int. J. Syst. Evol. Microbiol. 2005, 55, 1681–1685, Erratum in Int. J. Syst. Evol. Microbiol. 2005, 55 Pt 5, 2233. [Google Scholar] [CrossRef][Green Version]
- Morana, A.; Esposito, A.; Maurelli, L.; Ruggiero, G.; Ionata, E.; Rossi, M.; Cara, F. A novel thermoacidophilic cellulase from Alicyclobacillus acidocaldarius. Protein Pept. Lett. 2008, 15, 1017–1021. [Google Scholar] [CrossRef] [PubMed]
- Potprommanee, L.; Wang, X.-Q.; Han, Y.-J.; Nyobe, D.; Peng, Y.-P.; Huang, Q.; Liu, J.; Liao, Y.-L.; Chang, K.-L. Characterization of a thermophilic cellulase from Geobacillus sp. HTA426, an efficient cellulase-producer on alkali pre-treated of lignocellulosic biomass. PLoS ONE 2017, 12, e0175004. [Google Scholar] [CrossRef] [PubMed]
- Khianngam, S.; Pootaeng-on, Y.; Techakriengkrai, T.; Tanasupawat, S. Screening and identification of cellulase producing bacteria isolated from oil palm meal. J. Appl. Pharm. Sci. 2014, 4, 90. [Google Scholar] [CrossRef]
- Fritze, D. Taxonomy of the genus Bacillus and related genera: The aerobic endospore-forming bacteria. Phytopathology 2004, 94, 1245–1248. [Google Scholar] [CrossRef] [PubMed]
- Harirchi, S.; Sar, T.; Ramezani, M.; Aliyu, H.; Etemadifar, Z.; Nojoumi, S.A.; Yazdian, F.; Awasthi, M.K.; Taherzadeh, M.J. Bacillales: From Taxonomy to Biotechnological and Industrial Perspectives. Microorganisms 2022, 10, 2355. [Google Scholar] [CrossRef] [PubMed]
- Harwood, C.R. Bacillus subtilis and its relatives: Molecular biological and industrial workhorses. Trends Biotechnol. 1992, 10, 247–256. [Google Scholar] [CrossRef]
- Zeigler, D.R. The genome sequence of Bacillus subtilis subsp. spizizenii W23: Insights into speciation within the B. subtilis complex and into the history of B. subtilis genetics. Microbiology 2011, 157, 2033–2041. [Google Scholar] [CrossRef] [PubMed]
- Franco-Duarte, R.; Černáková, L.; Kadam, S.; Kaushik, K.S.; Salehi, B.; Bevilacqua, A.; Corbo, M.R.; Antolak, H.; Dybka-Stępień, K.; Leszczewicz, M.; et al. Advances in Chemical and Biological Methods to Identify Microorganisms—From Past to Present. Microorganisms 2019, 7, 130. [Google Scholar] [CrossRef] [PubMed]
- Boyd, M.A.; Antonio, M.A.; Hillier, S.L. Comparison of API 50 CH strips to whole-chromosomal DNA probes for identification of Lactobacillus species. J. Clin. Microbiol. 2005, 43, 5309–5311. [Google Scholar] [CrossRef]
- Klingler, J.M.; Stowe, R.P.; Obenhuber, D.C.; Groves, T.O.; Mishra, S.K.; Pierson, D.L. Evaluation of the Biolog automated microbial identification system. Appl. Environ. Microbiol. 1992, 58, 2089. [Google Scholar] [CrossRef]
- Patel, S.; Gupta, R.S. A phylogenomic and comparative genomic framework for resolving the polyphyly of the genus Bacillus: Proposal for six new genera of Bacillus species, Peribacillus gen. nov., Cytobacillus gen. nov., Mesobacillus gen. nov., Neobacillus gen. nov., Metabacillus gen. nov. and Alkalihalobacillus gen. nov. Int. J. Syst. Evol. Microbiol. 2020, 70, 406–438. [Google Scholar] [CrossRef] [PubMed]
- Ash, C.; Farrow, J.A.E.; Wallbanks, S.; Collins, M.D. Phylogenetic heterogeneity of the genus Bacillus revealed by comparative analysis of small-subunit-ribosomal RNA sequences. Lett. Appl. Microbiol. 1991, 13, 202–206. [Google Scholar] [CrossRef]
- Shida, O.; Takagi, H.; Kadowaki, K.; Komagata, K. Proposal for two new genera, Brevibacillus gen. nov. and Aneurinibacillus gen. nov. Int. J. Syst. Evol. Microbiol. 1996, 46, 939–946. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, I.; Yokota, A.; Yamazoe, A.; Fujiwara, T. Proposal of Lysinibacillus boronitolerans gen. nov. sp. nov., and transfer of Bacillus fusiformis to Lysini bacillus fusiformis comb. nov. and Bacillus sphaericus to Lysini bacillus sphaericus comb. nov. Int. J. Syst. Evol. Microbiol. 2007, 57, 1117–1125. [Google Scholar] [CrossRef] [PubMed]
- Wainø, M.; Tindall, B.J.; Schumann, P.; Ingvorsen, K. Gracilibacillus gen. nov., with description of Gracilibacillus halotolerans gen. nov., sp. nov.; transfer of Bacillus dipsosauri to Gracilibacillus dipsosauri comb. nov., and Bacillus salexigens to the genus Salibacillus gen. nov., as Sali bacillus salexigens comb. nov. Int. J. Syst. Bacteriol. 1999, 49, 821–831. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.H.; Lee, K.C.; Weiss, N.; Kho, Y.H.; Kang, K.H.; Park, Y.H. Sporosarcina aquimarina sp. nov., a bacterium isolated from seawater in Korea, and transfer of Bacillus globisporus (Larkin and Stokes 1967), Bacillus psychrophilus (Nakamura 1984) and Bacillus pasteurii (Chester 1898) to the genus Sporosarcina as Sporosarcina globispora comb. nov., Sporosarcina psychrophila comb. nov. and Sporosarcina pasteurii comb. nov., and emended description of th. Int. J. Syst. Evol. Microbiol. 2001, 51, 1079–1086. [Google Scholar] [CrossRef]
- Vos, P.; Garrity, G.M.; Jones, D.; Krieg, N.R.; Ludwig, W.; Rainey, F.A.; Schleifer, K.; Whitman, W.B. (Eds.) Bergey’s Manual of Systematic Bacteriology: Volume 3: The Firmicutes; Springer: New York, NY, USA, 2010. [Google Scholar]
- Yokota, A.; Fujii, T.; Goto, K. Alicyclobacillus: Thermophilic Acidophilic Bacilli; Springer: Tokyo, Japan, 2007. [Google Scholar]
- Dobrzyński, J.; Wierzchowski, P.S.; Stępień, W.; Górska, E.B. The Reaction of Cellulolytic and Potentially Cellulolytic Spore-Forming Bacteria to Various Types of Crop Management and Farmyard Manure Fertilization in Bulk Soil. Agronomy 2021, 11, 772. [Google Scholar] [CrossRef]
- De Maayer, P.; Aliyu, H.; Cowan, D.A. Reorganising the order Bacillales through phylogenomics. Syst. Appl. Microbiol. 2018, 42, 178–189. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.S.; Patel, S. Robust Demarcation of the Family Caryophanaceae (Planococcaceae) and Its Different Genera Including Three Novel Genera Based on Phylogenomics and Highly Specific Molecular Signatures. Front. Microbiol. 2020, 10, 2821. [Google Scholar] [CrossRef]
- Gupta, R.S.; Patel, S.; Saini, N.; Chen, S. Robust Demarcation of 17 Distinct Bacillus Species Clades, Proposed as Novel Bacillaceae Genera, by Phylogenomics and Comparative Genomic Analyses: Description of Robertmurraya kyonggiensis sp. Nov. and Proposal for an Emended Genus Bacillus Limiting It Only to the Members of the Subtilis and Cereus Clades of Species. Int. J. Syst. Evol. Microbiol. 2020, 70, 5753–5798. [Google Scholar] [PubMed]
- Liu, Y.; Lai, Q.; Du, J.; Shao, Z. Genetic diversity and population structure of the Bacillus cereus group bacteria from diverse marine environments. Sci. Rep. 2017, 7, 689. [Google Scholar] [CrossRef] [PubMed]
- NamesforLife Bacterial and Archaeal Nomenclature. Available online: https://www.namesforlife.com/search (accessed on 20 August 2022).
- Parte, A.C.; Carbasse, J.S.; Meier-Kolthoff, j.; Reimer, L.C.; Göker, M. List of Prokaryotic Names with Standing in Nomenclature (Lpsn) Moves to the Dsmz. Int. J. Syst. Evol. Microbiol. 2020, 70, 5607–5612. [Google Scholar] [CrossRef] [PubMed]
- Tindall, B.J. Names at the rank of class, subclass and order, their typification and current status: Supplementary information to Opinion 79. Judicial Commission of the International Committee on Systematics of Prokaryotes. Int. J. Syst. Evol. Microbiol. 2014, 64, 3599–3602. [Google Scholar] [CrossRef] [PubMed]
- Huang, W.C.; Hu, Y.; Zhang, G.; Li, M. Comparative genomic analysis reveals metabolic diversity of different Paenibacillus groups. Appl. Microbiol. Biotechnol. 2020, 104, 10133–10143. [Google Scholar] [CrossRef] [PubMed]
- Logan, N.A.; De Vos, P. Genus Bacillus Cohn 1872. In Bergey’s Manual of Systematic Bacteriology; De Vos, P., Garrity, M., Jones, D., Krieg, R., Ludwig, W., Eds.; Springer: New York, NY, USA, 2009; pp. 21–128. [Google Scholar]
- Logan, N.A.; Berge, O.; Bishop, A.H.; Busse, H.J.; De Vos, P.; Fritze, D.; Heyndrickx, M.; Kämpfer, P.; Rabinovitch, L.; Salkinoja-Salonen, M.S.; et al. Proposed minimal standards for describing new taxa of aerobic, endospore-forming bacteria. Int. J. Syst. Evol. Microbiol. 2009, 59, 2114–2121. [Google Scholar] [CrossRef]
- Patel, S.; Gupta, R.S. Robust demarcation of fourteen different species groups within the genus Streptococcus based on genome-based phylogenies and molecular signatures. Infect. Genet. Evol. 2018, 66, 130–151. [Google Scholar] [CrossRef]
- Rooney, A.P.; Price, N.P.J.; Ehrhardt, C.; Swezey, J.L.; Bannan, J.D. Phylogeny and molecular taxonomy of the Bacillus subtilis species complex and description of Bacillus subtilis subsp. inaquosorum subsp. nov. Int. J. Syst. Evol. Microbiol. 2009, 59, 2429–2436. [Google Scholar] [CrossRef]
- La Duc, M.T.; Satomi, M.; Agata, N.; Venkateswaran, K. gyrB as a phylogenetic discriminator for members of the Bacillus anthracis-cereus-thuringiensis group. J. Microbiol. Methods 2004, 56, 83–394. [Google Scholar] [CrossRef]
- Wang, L.T.; Lee, F.L.; Tai, C.J.; Kasai, H. Comparison of gyrB gene sequences, 16S rRNA gene sequences and DNA-DNA hybridization in the Bacillus subtilis group. Int. J. Syst. Evol. Microbiol. 2007, 57, 1846–1850. [Google Scholar] [CrossRef]
- Verma, A.; Ojha, A.K.; Pal, Y.; Kumari, P.; Schumann, P.; Gruber-Vodicka, H.; Dastager, D.G.; Natarajan, R.K.; Mayilraj, S.; Krishnamurthi, S. An investigation into the taxonomy of “Bacillus aminovorans” and its reclassification to the genus Domibacillus as Domibacillus aminovorans sp. nov. Syst. Appl. Microbiol. 2017, 40, 458–467. [Google Scholar] [CrossRef] [PubMed]
- Rössler, D.; Ludwig, W.; Schleifer, K.H.; Lin, C.; McGill, T.J.; Wisotzkey, J.D.; Jurtshuk, P., Jr.; Fox, G.E. Phylogenetic diversity in the genus Bacillus as seen by 16S rRNA sequencing studies. Syst. Appl. Microbiol. 1991, 14, 266–269. [Google Scholar] [CrossRef]
- Klenk, H.P.; Lapidus, A.; Chertkov, O.; Copeland, A.; Del Rio, T.G.; Nolan, M.; Lucas, S.; Chen, F.; Tice, H.; Cheng, J.-F.; et al. Complete genome sequence of the thermophilic, hydrogen-oxidizing Bacillus tusciae type strain (T2) and reclassification in the new genus, Kyrpidia gen. nov. as Kyrpidia tusciae comb. nov. and emendation of the family Alicyclobacillaceae dA costa and rainey, 2010. Stand. Genomic Sci. 2011, 5, 121–134. [Google Scholar] [CrossRef]
- Harwood, C.R.; Mouillon, J.-M.; Pohl, S.; Arnau, J. Secondary metabolite production and the safety of industrially important members of the Bacillus subtilis group. FEMS Microbiol. Rev. 2018, 42, 721–738. [Google Scholar] [CrossRef] [PubMed]
- Kovács, Á.T. Bacillus subtilis. Trends Microbiol. 2019, 27, 724–725. [Google Scholar] [CrossRef]
- Sansinenea, E.; Ortiz, A. Secondary metabolites of soil Bacillus spp. Biotechnol. Lett. 2011, 33, 1523–1538. [Google Scholar] [CrossRef]
- Zhang, L.; Jin, M.; Shi, X.; Jin, L.; Hou, X.; Yu, Y.; Liu, B.; Cao, J.; Quan, C. Macrolactin metabolite production by Bacillus sp. ZJ318 isolated from marine sediment. Appl. Biochem. Biotechnol. 2022, 194, 2581–2593. [Google Scholar] [CrossRef]
- Dobrzyński, J.; Jakubowska, Z.; Dybek, B. Potential of Bacillus pumilus to Directly Promote Plant Growth. Front. Microbiol. 2022, 13, 1069053. [Google Scholar] [CrossRef] [PubMed]
- Smagacz, J.; Martyniuk, S. Soil properties and crop yields as influenced by the frequency of straw incorporation in a rape-wheat-triticale rotation. J. Water Land Dev. 2023, 56, 1–6. [Google Scholar] [CrossRef]
- Mignard, S.; Flandrois, J.P. 16S rRNA sequencing in routine bacterial identification: A 30-month experiment. J. Microbiol. Methods 2006, 67, 574–581. [Google Scholar] [CrossRef] [PubMed]
- Winand, R.; Bogaerts, B.; Hoffman, S.; Lefevre, L.; Delvoye, M.; Van Braekel, J.; Fu, Q.; Roosens, N.H.; De Keersmaecker, S.C.; Vanneste, K. Targeting the 16S rRNA Gene for Bacterial Identification in Complex Mixed Samples: Comparative Evaluation of Second (Illumina) and Third (Oxford Nanopore Technologies) Generation Sequencing Technologies. Int. J. Mol. Sci. 2020, 21, 298. [Google Scholar] [CrossRef]
- Hartmann, M.; Widmer, F. Community Structure Analyses Are More Sensitive to Differences in Soil Bacterial Communities than Anonymous Diversity Indices. Appl. Environ. Microbiol. 2006, 72, 7804–7812. [Google Scholar] [CrossRef] [PubMed]
- Zeng, J.; Liu, X.; Song, L.; Lin, X.; Zhang, H.; Shen, C.; Chu, H. Nitrogen fertilization directly affects soil bacterial diversity and indirectly affects bacterial community composition. Soil Biol. Biochem. 2016, 92, 41–49. [Google Scholar] [CrossRef]
- Li, N.; Gao, D.; Zhou, X.; Chen, S.; Li, C.; Wu, F. Intercropping with Potato-Onion Enhanced the Soil Microbial Diversity of Tomato. Microorganisms 2020, 8, 834. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Mao, X.; Zhang, M.; Yang, W.; Di, H.J.; Ma, L.; Liu, W.; Li, B. Response of Soil Microbial Communities to Continuously Mono-Cropped Cucumber under Greenhouse Conditions in a Calcareous Soil of North China. J. Soils Sediments 2020, 20, 2446–2459. [Google Scholar] [CrossRef]
- Wierzchowski, P.S.; Dobrzyński, J.; Mazur, K.; Kierończyk, M.; Wardal, W.J.; Sakowski, T.; Barszczewski, J. Chemical Properties and Bacterial Community Reaction to Acidified Cattle Slurry Fertilization in Soil from Maize Cultivation. Agronomy 2021, 11, 601. [Google Scholar] [CrossRef]
- Li, H.; Wang, J.; Liu, Q.; Zhou, Z.; Chen, F.; Xiang, D. Effects of Consecutive Monoculture of Sweet Potato on Soil Bacterial Community as Determined by Pyrosequencing. J. Basic Microbiol. 2019, 59, 181–191. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.; Wang, Y.; Xu, X.; Liu, T.; Wu, D.; Zheng, X.; Tang, S.; Dai, Q. Potential Use of High-Throughput Sequencing of Soil Microbial Communities for Estimating the Adverse Effects of Continuous Cropping on Ramie (Boehmeria nivea L. Gaud). PLoS ONE 2018, 13, e0197095. [Google Scholar] [CrossRef]
- Yang, L.; Tan, L.; Zhang, F.; Gale, W.J.; Cheng, Z.; Sang, W. Duration of Continuous Cropping with Straw Return Affects the Composition and Structure of Soil Bacterial Communities in Cotton Fields. Can. J. Microbiol. 2018, 64, 167–181. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Q.; Xiong, W.; Xing, Y.; Sun, Y.; Lin, X.; Dong, Y. Long-Term Coffee Monoculture Alters Soil Chemical Properties and Microbial Communities. Sci. Rep. 2018, 8, 6116. [Google Scholar] [CrossRef] [PubMed]
- Alami, M.M.; Pang, Q.; Gong, Z.; Yang, T.; Tu, D.; Zhen, O.; Yu, W.; Alami, M.J.; Wang, X. Continuous Cropping Changes the Composition and Diversity of Bacterial Communities: A Meta-Analysis in Nine Different Fields with Different Plant Cultivation. Agriculture 2021, 11, 1224. [Google Scholar] [CrossRef]
- Lienhard, P.; Terrat, S.; Prévost-Bouré, N.C.; Nowak, V.; Régnier, T.; Sayphoummie, S.; Panyasiri, K.; Tivet, F.; Mathieu, O.; Levêque, J.; et al. Pyrosequencing Evidences the Impact of Cropping on Soil Bacterial and Fungal Diversity in Laos Tropical Grassland. Agron. Sustain. Dev. 2014, 34, 525–533. [Google Scholar] [CrossRef]
- Li, J.; Gan, G.; Chen, X.; Zou, J. Effects of Long-Term Straw Management and Potassium Fertilization on Crop Yield, Soil Properties, and Microbial Community in a Rice–Oilseed Rape Rotation. Agriculture 2021, 11, 1233. [Google Scholar] [CrossRef]
- Dang, P.; Li, C.; Lu, C.; Zhang, M.; Huang, T.; Wan, C.; Wang, H.; Chen, Y.; Qin, X.; Liao, Y.; et al. Effect of fertilizer management on the soil bacterial community in agroecosystems across the globe. Agric. Ecosyst. Environ. 2021, 326, 107795. [Google Scholar] [CrossRef]
- Francioli, D.; Schulz, E.; Lentendu, G.; Wubet, T.; Buscot, F.; Reitz, T. Mineral vs. Organic Amendments: Microbial Community Structure, Activity and Abundance of Agriculturally Relevant Microbes Are Driven by Long-Term Fertilization Strategies. Front. Microbiol. 2016, 7, 1446. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, M.; Frey, B.; Mayer, J.; Mäder, P.; Widmer, F. Distinct Soil Microbial Diversity under Long-Term Organic and Conventional Farming. ISME J. 2015, 9, 1177–1194. [Google Scholar] [CrossRef] [PubMed]
- Hartman, K.; van der Heijden, M.G.A.; Wittwer, R.A.; Banerjee, S.; Walser, J.-C.; Schlaeppi, K. Cropping Practices Manipulate Abundance Patterns of Root and Soil Microbiome Members Paving the Way to Smart Farming. Microbiome 2018, 6, 14. [Google Scholar]
- Guo, Z.; Wan, S.; Hua, K.; Yin, Y.; Chu, H.; Wang, D.; Guo, X. Fertilization regime has a greater effect on soil microbial community structure than crop rotation and growth stage in an agroecosystem. Appl. Soil Ecol. 2020, 149, 103510. [Google Scholar] [CrossRef]
- Bhardwaj, N.; Kumar, B.; Agrawal, K.; Verma, P. Current perspective on production and applications of microbial cellulases: A review. Bioresour. Bioprocess. 2021, 8, 95. [Google Scholar] [CrossRef]
- Thapa, S.; Mishra, J.; Arora, N.; Mishra, P.; Li, H.; O’Hair, J.; Bhatti, S.; Zhou, S. Microbial cellulolytic enzymes: Diversity and biotechnology with reference to lignocellulosic biomass degradation. Rev. Environ. Sci. Biol. Technol. 2020, 19, 621–648. [Google Scholar]
- Horn, S.; Vaaje-Kolstad, G.; Westereng, B.; Eijsink, V.G. Novel enzymes for the degradation of cellulose. Biotechnol. Biofuels 2012, 5, 45. [Google Scholar] [CrossRef] [PubMed]
- Singhania, R.R.; Ruiz, H.A.; Awasthi, M.K.; Dong, C.D.; Chen, C.W.; Patel, A.K. Challenges in cellulase bioprocess for biofuel applications. Renew. Sustain. Energy Rev. 2021, 151, 111622. [Google Scholar] [CrossRef]
- Zverlov, V.V.; Schwarz, W.H. Bacterial cellulose hydrolysis in anaerobic environmental subsystems—Clostridium thermocellum and Clostridium stercorarium, thermophilic plant-fiber degraders. Ann. N. Y. Acad. Sci. 2008, 1125, 298–307. [Google Scholar] [CrossRef]
- dos Santos, J.R.; de Souza Moreira, L.R.; Filho, E.X.F. Cellulose-degrading enzymes: Key players in biorefinery development. Biologia 2023, 78, 1759–1772. [Google Scholar] [CrossRef]
- Maki, M.; Leung, K.T.; Qin, W. The prospects of cellulase-producing bacteria for the bioconversion of lignocellulosic biomass. Int. J. Biol. Sci. 2009, 5, 500. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Huang, W.C.; Liu, Y.; Li, M. Diversity of cellulolytic microorganisms and microbial cellulases. Int. Biodeterior. Biodegrad. 2021, 163, 105277. [Google Scholar] [CrossRef]
- Himmel, M.E.; Xu, Q.; Luo, Y.; Ding, S.Y.; Lamed, R.; Bayer, E.A. Microbial enzyme systems for biomass conversion: Emerging paradigms. Biofuels 2010, 1, 323–341. [Google Scholar] [CrossRef]
- Koeck, D.E.; Pechtl, A.; Zverlov, V.V.; Schwarz, W.H. Genomics of cellulolytic bacteria. Curr. Opin. Biotechnol. 2014, 29, 171–183. [Google Scholar] [CrossRef]
- Nguyen, S.N.; Flores, A.; Talamantes, D.; Dar, F.; Valdez, A.; Schwans, J.; Berlemont, R. GeneHunt for rapid domain-specific annotation of glycoside hydrolases. Sci. Rep. 2019, 9, 10137. [Google Scholar] [CrossRef] [PubMed]
- Sidar, A.; Albuquerque, E.D.; Voshol, G.P.; Ram, A.F.; Vijgenboom, E.; Punt, P.J. Carbohydrate binding modules: Diversity of domain architecture in amylases and cellulases from filamentous microorganisms. Front. Bioeng. Biotechnol. 2020, 8, 871. [Google Scholar] [CrossRef] [PubMed]
- Konar, A.; Aich, S.; Katakojwala, R.; Datta, S.; Mohan, S.V. A processive GH9 family endoglucanase of Bacillus licheniformis and the role of its carbohydrate-binding domain. Appl. Microbiol. Biotechnol. 2022, 106, 6059–6075. [Google Scholar] [CrossRef] [PubMed]
- Ogonda, L.A.; Saumonneau, A.; Dion, M.; Muge, E.K.; Wamalwa, B.M.; Mulaa, F.J.; Tellier, C. Characterization and engineering of two new GH9 and GH48 cellulases from a Bacillus pumilus isolated from Lake Bogoria. Biotechnol. Lett. 2021, 43, 691–700. [Google Scholar] [CrossRef] [PubMed]
- Honda, S.; Kunii, T.; Nohara, K.; Wakita, S.; Sugahara, Y.; Kawakita, M.; Oyama, F.; Sakaguchi, M. Characterization of a Bacillus thuringiensis chitinase that binds to cellulose and chitin. AMB Express 2017, 7, 51. [Google Scholar] [CrossRef] [PubMed]
- Park, J.S.; Nakamura, A.; Horinouchi, S.; Beppu, T. Identification of the cellulose-binding domain of a Bacillus subtilis endoglucanase distinct from its catalytic domain. Biosci. Biotechnol. Biochem. 1993, 57, 260–264. [Google Scholar] [CrossRef] [PubMed]
- Yadav, S.; Dubey, S.K. Cellulose degradation potential of Paenibacillus lautus strain BHU3 and its whole genome sequence. Bioresour. Technol. 2018, 262, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, S.; Dafale, N.A.; Purohit, H.J. Functional genomics assessment of lytic polysaccharide mono-oxygenase with glycoside hydrolases in Paenibacillus dendritiformis CRN18. Int. J. Biol. Macromol. 2020, 164, 3729–3738. [Google Scholar] [CrossRef] [PubMed]
- Bianchetti, C.M.; Brumm, P.; Smith, R.W.; Dyer, K.; Hura, G.L.; Rutkoski, T.J.; Phillips, G.N., Jr. Structure, dynamics, and specificity of endoglucanase D from Clostridium cellulovorans. J. Mol. Biol. 2013, 425, 4267–4285. [Google Scholar] [CrossRef]
- Duan, C.J.; Feng, J.X. Mining metagenomes for novel cellulase genes. Biotechnol. Lett. 2010, 32, 1765–1775. [Google Scholar] [CrossRef] [PubMed]
- Tomme, P.R.; Warren, A.J.; Gilkes, N.R. Cellulose hydrolysis by bacteria and fungi. Adv. Microb. Physiol. 1995, 37, 1–81. [Google Scholar]
- Guglielmi, G.; Beguin, P. Cellulase and hemicellulase genes of Clostridium thermocellum from five independent collections contain few overlaps and are widely scattered across the chromosome. FEMS Microbiol. Lett. 1998, 161, 209–215. [Google Scholar] [CrossRef] [PubMed]
- Belaich, J.P.; Belaich, A.; Fierobe, H.P.; Gal, L.; Gaudin, C.; Pagès, S.; Reverbel-Leroy, C.; Tardif, C. The cellulolytic system of Clostridium cellulolyticum. In Genetics, Biochemistry and Ecology of Cellulose Degradation; Ohmiya, K., Hayashi, K., Sakka, Y., Kobayashi, S., Karita, T., Eds.; Kimura Uni Publishers: Tokyo, Japan, 1999; pp. 479–487. [Google Scholar]
- Tamaru, Y.; Karita, S.; Ibrahim, A.; Chan, H.; Doi, R.H. A large gene cluster for the Clostridium cellulovorans cellulosome. J. Bacteriol. 2000, 182, 5906–5910. [Google Scholar] [CrossRef] [PubMed]
- Rey, M.W.; Ramaiya, P.; Nelson, B.A.; Brody-Karpin, S.D.; Zaretsky, E.J.; Tang, M.; Lopez de Leon, A.; Xiang, H.; Gusti, V.; Clausen, I.G.; et al. Complete genome sequence of the industrial bacterium Bacillus licheniformis and comparisons with closely related Bacillus species. Genome Biol. 2004, 5, R77. [Google Scholar] [CrossRef] [PubMed]
- Shang, Z.; Liu, S.; Duan, Y.; Bao, C.; Wang, J.; Dong, B.; Cao, Y. Complete genome sequencing and investigation on the fiber-degrading potential of Bacillus amyloliquefaciens strain TL106 from the tibetan pig. BMC Microbiol. 2022, 22, 186. [Google Scholar] [CrossRef] [PubMed]
- Carbonaro, M.; Aulitto, M.; Gallo, G.; Contursi, P.; Limauro, D.; Fiorentino, G. Insight into CAZymes of Alicyclobacillus mali FL18: Characterization of a New Multifunctional GH9 Enzyme. Int. J. Mol. Sci. 2023, 24, 243. [Google Scholar] [CrossRef]
- Eckert, K.; Schneider, E. A Thermoacidophilic Endoglucanase (CelB) from Alicyclobacillus acidocaldarius Displays High Sequence Similarity to Arabinofuranosidases Belonging to Family 51 of Glycoside Hydrolases. Eur. J. Biochem. 2003, 270, 3593–3602. [Google Scholar] [CrossRef]
- Eckert, K.; Zielinski, F.; lo Leggio, L.; Schneider, E. Gene Cloning, Sequencing, and Characterization of a Family 9 Endoglucanase (CelA) with an Unusual Pattern of Activity from the Thermoacidophile Alicyclobacillus acidocaldarius ATCC27009. Appl. Microbiol. Biotechnol. 2002, 60, 428–436. [Google Scholar]
- Eastman, A.W.; Heinrichs, D.E.; Yuan, Z.C. Comparative and genetic analysis of the four sequenced Paenibacillus polymyxa genomes reveals a diverse metabolism and conservation of genes relevant to plant-growth promotion and competitiveness. BMC Genomics 2014, 15, 581. [Google Scholar] [CrossRef]
- Kuhad, R.C.; Deswal, D.; Sharma, S.; Bhattacharya, A.; Jain, K.K.; Kaur, A.; Pletschke, B.I.; Singh, A.K.; Karp, M. Revisiting cellulase production and redefining current strategies based on major challenges. Renew. Sustain. Energy Rev. 2016, 55, 249–272. [Google Scholar] [CrossRef]
- Van Dyk, J.S.; Sakka, M.; Sakka, K.; Pletschke, B.I. The cellulolytic and hemi-cellulolytic system of Bacillus licheniformis SVD1 and the evidence for production of a large multi-enzyme complex. Enzyme Microb. Technol. 2009, 45, 372–378. [Google Scholar] [CrossRef]
- Orencio-Trejo, M.; De la Torre-Zavala, S.; Rodriguez-Garcia, A.; Avilés-Arnaut, H.; Gastelum-Arellanez, A. Assessing the performance of bacterial cellulases: The use of Bacillus and Paenibacillus strains as enzyme sources for lignocellulose saccharification. BioEnergy Res. 2016, 9, 1023–1033. [Google Scholar] [CrossRef]
- Lamed, R.; Setter, E.; Kenig, R.; Bayer, E.A. Cellulosome: A discrete cell surface organelle of Clostridium thermocellum which exhibits separate antigenic, cellulose-binding and various cellulolytic activities. Biotechnol. Bioeng. Symp. 1983, 13, 163–181. [Google Scholar]
- Hazlewood, G.P.; Romaniec, M.P.M.; Davidson, K.; Grepinet, O.; Beguin, P.; Millet, J.; Raynaud, O.; Aubert, J.P. A catalogue of Clostridium thermocellum endoglucanase, B-glucosidase, and xylanase genes cloned in Escherichia coli. FEMS Microbiol. Lett. 1988, 51, 231–236. [Google Scholar] [CrossRef]
- Muras, A.; Romero, M.; Mayer, C.; Otero, A. Biotechnological applications of Bacillus licheniformis. Crit. Rev. Biotechnol. 2021, 41, 609–627. [Google Scholar] [CrossRef] [PubMed]
- Waeonukul, R.; Kyu, K.J.; Sakka, K.; Ratanakhanokchai, K. Isolation and characterization of a multienzyme complex (cellulosome) of the Paenibacillus curdlanolyticus B-6 grown on Avicel under aerobic conditions. J. Biosci. Bioeng. 2009, 107, 610–614. [Google Scholar] [CrossRef] [PubMed]
- Górska, E.; Tudek, B.; Russel, S. Degradation of cellulose by nitrogen-fixing strain of Bacillus polymyxa. Acta Microbiol. Pol. 2001, 50, 129–137. [Google Scholar] [PubMed]
- Górska, E.B.; Jankiewicz, U.; Dobrzynski, J.; Russel, S.; Pietkiewicz, S.; Kalaji, H.; Gozdowski, D. Degradation and colonization of cellulose by diazotrophic strains of Paenibacillus polymyxa isolated from soil. J. Bioremediat. Biodegrad. 2015, 6, 271. [Google Scholar] [CrossRef]
- Liang, Y.; Yesuf, J.; Schmitt, S.; Bender, K.; Bozzola, J. Study of cellulases from a newly isolated thermophilic and cellulolytic Brevibacillus sp. strain JXL. J. Ind. Microbiol. Biotechnol. 2009, 36, 961–970. [Google Scholar] [CrossRef] [PubMed]
- Acharya, S.; Chaudhary, A. Optimization of fermentation conditions for cellulases production by Bacillus licheniformis MVS1 and Bacillus sp. MVS3 isolated from Indian hot spring. Braz. Arch. Biol. Technol. 2012, 55, 497–503. [Google Scholar] [CrossRef]
- Shajahan, S.; Moorthy, I.G.; Sivakumar, N.; Selvakumar, G. Statistical modeling and optimization of cellulase production by Bacillus licheniformis NCIM 5556 isolated from the hot spring, Maharashtra, India. King Saud Univ. Sci. 2017, 29, 302–310. [Google Scholar] [CrossRef]
- Sadhu, S.; Ghosh, P.K.; Aditya, G.; Maiti, T.K. Optimization and strain improvement by mutation for enhanced cellulase production by Bacillus sp. (MTCC10046) isolated from cow dung. J. King Saud Univ. Sci. 2014, 26, 323–332. [Google Scholar] [CrossRef]
- Akaracharanya, A.; Taprig, T.; Sitdhipol, J.; Tanasupawat, S. Characterization of cellulase producing Bacillus and Paenibacillus strains from Thai soils. J. Appl. Pharm. Sci. 2014, 4, 6. [Google Scholar] [CrossRef]
- Thomas, L.; Ram, H.; Singh, V.P. Inducible cellulase production from an organic solvent tolerant Bacillus sp. SV1 and evolutionary divergence of endoglucanase in different species of the genus Bacillus. Braz. J. Microbiol. 2018, 49, 429–442. [Google Scholar] [CrossRef]
- Mihajlovski, K.R.; Carević, M.B.; Dević, M.L.; Šiler-Marinković, S.; Rajilić-Stojanović, M.D.; Dimitrijević-Branković, S. Lignocellulosic waste material as substrate for Avicelase production by a new strain of Paenibacillus chitinolyticus CKS1. Int. Biodeterior. Biodegrad. 2015, 104, 426–434. [Google Scholar] [CrossRef]
- Kim, D.S.; Kim, C.H. Production and characterization of crystalline cellulose-degrading cellulase components from a thermophilic and moderately alkalophilic bacterium. J. Microbiol. Biotechnol. 1992, 2, 7–13. [Google Scholar]
- Kazeem, M.O.; Shah, U.K.M.; Baharuddin, A.S.; Abdul Rahman, N.A. Enhanced cellulase production by a novel thermophilic Bacillus licheniformis 2D55: Characterization and application in lignocellulosic saccharification. Bioresources 2016, 11, 5404–5423. [Google Scholar] [CrossRef]
- Pramanik, S.K.; Mahmud, S.; Paul, G.K.; Jabin, T.; Naher, K.; Uddin, M.S.; Zaman, S.; Saleh, M.A. Fermentation optimization of cellulase production from sugarcane bagasse by Bacillus pseudomycoides and molecular modeling study of cellulase. Curr. Res. Microb. Sci. 2021, 2, 100013. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Zhang, W.-W.; Yang, M.-M.; Chen, Y.-L. Cloning of the thermostable cellulase gene from newly isolated Bacillus subtilis and its expression in Escherichia coli. Mol. Biotechnol. 2008, 40, 195–201. [Google Scholar] [CrossRef] [PubMed]
- Rastogi, G.; Bhalla, A.; Adhikari, A.; Bischoff, K.M.; Hughes, S.R.; Christopher, L.P.; Sani, R.K. Characterization of thermostable cellulases produced by Bacillus and Geobacillus strains. Bioresour. Technol. 2010, 101, 8798–8806. [Google Scholar] [CrossRef] [PubMed]
- Tai, S.-K.; Lin, H.-P.P.; Kuo, J.; Liu, J.-K. Isolation and characterization of a cellulolytic Geobacillus thermoleovorans T4 strain from sugar refinery wastewater. Extremophiles 2004, 8, 345–349. [Google Scholar] [CrossRef] [PubMed]
- Seo, J.K.; Park, T.S.; Kwon, I.H.; Piao, M.Y.; Lee, C.H.; Ha, J.K. Characterization of cellulolytic and xylanolytic enzymes of Bacillus licheniformis JK7 isolated from the rumen of a native Korean goat. Asian-Australas. J. Anim. Sci. 2013, 26, 50–58. [Google Scholar] [CrossRef] [PubMed]
- Tabssum, F.; Irfan, M.; Shakir, H.A.; Qazi, J.I. RSM based optimization of nutritional conditions for cellulase mediated Saccharification by Bacillus cereus. J. Biol. Eng. 2018, 12, 7. [Google Scholar] [CrossRef]
- Paudel, Y.P.; Qin, W. Characterization of novel cellulase-producing bacteria isolated from rotting wood samples. Appl. Biochem. Biotechnol. 2015, 177, 1186–1198. [Google Scholar] [CrossRef]
- Ozaki, K.; Ito, S. Purification and properties of an acid endo-1, 4-β-glucanase from Bacillus sp. KSM-330. Microbiology 1991, 137, 41–48. [Google Scholar] [CrossRef] [PubMed]
- Fukumori, F.; Kudo, T.; Horikoshi, K. Purification and properties of a cellulase from alkalophilic Bacillus sp. no. 1139. Microbiology 1985, 131, 3339–3345. [Google Scholar] [CrossRef][Green Version]
- Bischoff, K.M.; Rooney, A.P.; Li, X.L.; Liu, S.; Hughes, S.R. Purification and characterization of a family 5 endoglucanase from a moderately thermophilic strain of Bacillus licheniformis. Biotech. Lett. 2006, 28, 1761–1765. [Google Scholar] [CrossRef]
- Yin, L.J.; Lin, H.H.; Xiao, Z.R. Purification and characterization of a cellulase from Bacillus subtilis YJ1. J. Mar. Sci. Technol. 2010, 18, 19. [Google Scholar] [CrossRef]
- Wang, C.M.; Shyu, C.L.; Ho, S.P.; Chiou, S.H. Characterization of a novel thermophilic, cellulose-degrading bacterium Paenibacillus sp. strain B39. Lett. Appl. Microb. 2008, 47, 46–53. [Google Scholar] [CrossRef]
- Liang, Y.L.; Zhang, Z.; Wu, M.; Wu, Y.; Feng, J.X. Isolation, screening, and identification of cellulolytic bacteria from natural reserves in the subtropical region of China and optimization of cellulase production by Paenibacillus terrae ME27-1. BioMed Res. Int. 2014, 2014, 512497. [Google Scholar] [CrossRef] [PubMed]
- Gaur, R.; Tiwari, S. Isolation, production, purification and characterization of an organic-solvent-thermostable alkalophilic cellulase from Bacillus vallismortis RG-07. BMC Biotechnol. 2015, 15, 19. [Google Scholar] [CrossRef] [PubMed]
- Yousef, N.; Mawad, A.; Abeed, A. Enhancement the Cellulase Activity Induced by Endophytic Bacteria Using Calcium Nanoparticles. Curr. Microbiol. 2019, 76, 346–354. [Google Scholar] [CrossRef] [PubMed]
- Bohra, V.; Tikariha, H.; Dafale, N.A. Genomically Defined Paenibacillus polymyxa ND24 for Efficient Cellulase Production Utilizing Sugarcane Bagasse as a Substrate. Appl. Biochem. Biotechnol. 2019, 187, 266–281. [Google Scholar] [CrossRef] [PubMed]
- Shah, T.A.; Charles, C.; Orts, W.J.; Tabassum, R. Biological pretreatment of rice straw by ligninolytic Bacillus s strains for enhancing biogas production. Environ. Prog. Sustain. Energy 2018, 38, e13036. [Google Scholar] [CrossRef]
- Kuhad, R.C.; Gupta, R.; Singh, A. Microbial cellulases and their industrial applications. Enzyme Res. 2011, 2011, 280696. [Google Scholar] [CrossRef] [PubMed]
- Ejaz, U.; Sohail, M.; Ghanemi, A. Cellulases: From Bioactivity to a Variety of Industrial Applications. Biomimetics 2021, 6, 44. [Google Scholar] [CrossRef]
- Kaushal, M.; Kumar, A.; Kaushal, R. Bacillus pumilus strain YSPMK11 as plant growth promoter and biocontrol agent against Sclerotinia sclerotiorum. 3 Biotech 2017, 7, 90–99. [Google Scholar] [CrossRef] [PubMed]
- Wróbel, M.; Śliwakowski, W.; Kowalczyk, P.; Kramkowski, K.; Dobrzyński, J. Bioremediation of Heavy Metals by the Genus Bacillus. Int. J. Environ. Res. Public Health 2023, 20, 4964. [Google Scholar] [CrossRef]
- Kulkova, I.; Dobrzyński, J.; Kowalczyk, P.; Bełżecki, G.; Kramkowski, K. Plant Growth Promotion Using Bacillus cereus. Int. J. Mol. Sci. 2023, 24, 9759. [Google Scholar] [CrossRef] [PubMed]
- Lipková, N.; Cinkocki, R.; Maková, J.; Medo, J.; Javoreková, S. Characterization of endophytic bacteria of the genus bacillus and their influence on the growth of maize (Zea mays) in vivo. J. Microbiol. Biotechnol. Food Sci. 2021, 10, e3602. [Google Scholar] [CrossRef]
- Shahid, I.; Han, J.; Hanooq, S.; Malik, K.A.; Borchers, C.H.; Mehnaz, S. Profiling of metabolites of bacillus spp. and their application in sustainable plant growth promotion and biocontrol. Front. Sustain. Food Syst. 2021, 5, 605195. [Google Scholar] [CrossRef]
- Mirskaya, G.V.; Khomyakov, Y.V.; Rushina, N.A.; Vertebny, V.E.; Chizhevskaya, E.P.; Chebotar, V.K.; Chesnokov, Y.V.; Pishchik, V.N. Plant Development of Early-Maturing Spring Wheat (Triticum aestivum L.) under Inoculation with Bacillus sp. V2026. Plants 2022, 11, 1817. [Google Scholar] [CrossRef] [PubMed]
- Dobrzyński, J.; Jakubowska, Z.; Kulkova, I.; Kowalczyk, P.; Kramkowski, K. Biocontrol of fungal phytopathogens by Bacillus pumilus. Front. Microb. 2023, 14, 1194606. [Google Scholar] [CrossRef]
- Win, K.T.; Oo, A.Z.; Ohkama-Ohtsu, N.; Yokoyama, T. Bacillus pumilus Strain TUAT-1 and Nitrogen Application in Nursery Phase Promote Growth of Rice Plants under Field Conditions. Agronomy 2018, 8, 216. [Google Scholar] [CrossRef]
- Ali, A.M.; Awad, M.Y.M.; Hegab, S.A.; Gawad, A.M.A.E.; Eissa, M.A. Effect of Potassium Solubilizing Bacteria (Bacillus cereus) on Growth and Yield of Potato. J. Plant Nutr. 2021, 44, 411–420. [Google Scholar] [CrossRef]
- Li, Y.; Wang, M.; Chen, S. Application of N2-fixing Paenibacillus triticisoli BJ-18 changes the compositions and functions of the bacterial, diazotrophic, and fungal microbiomes in the rhizosphere and root/shoot endosphere of wheat under field conditions. Biol. Fertil. Soils 2021, 57, 347–362. [Google Scholar] [CrossRef]
- Okoroafor, P.U.; Ikwuka, G.; Zaffar, N.; Ngalle Epede, M.; Mensah, M.K.; Haupt, J.; Golde, A.; Heilmeier, H.; Wiche, O. Field Studies on the Effect of Bioaugmentation with Bacillus amyloliquefaciens FZB42 on Plant Accumulation of Rare Earth Elements and Selected Trace Elements. Minerals 2022, 12, 409. [Google Scholar] [CrossRef]
- Stępień, A.; Wojtkowiak, K.; Kolankowska, E. Effect of Commercial Microbial Preparations Containing Paenibacillus azotofixans, Bacillus megaterium and Bacillus subtilis on the Yield and Photosynthesis of Winter Wheat and the Nitrogen and Phosphorus Content in the Soil. Appl. Sci. 2022, 12, 12541. [Google Scholar] [CrossRef]
- Zhang, L.N.; Wang, D.C.; Hu, Q.; Dai, X.Q.; Xie, Y.S.; Li, Q.; Liu, H.M.; Guo, J.H. Consortium of plant growth-promoting rhizobacteria strains suppresses sweet pepper disease by altering the rhizosphere microbiota. Front. Microbiol. 2019, 10, 1668. [Google Scholar] [CrossRef]
- Jamily, A.S.; Koyama, Y.; Win, T.A.; Toyota, K.; Chikamatsu, S.; Shirai, T.; Yasuhara, T. Effects of inoculation with a commercial microbial inoculant Bacillus subtilis C-3102 mixture on rice and barley growth and its possible mechanism in the plant growth stimulatory effect. J. Plant Protect. Res. 2019, 59, 193–205. [Google Scholar] [CrossRef]
- Qiu, F.; Liu, W.; Chen, L.; Wang, Y.; Ma, Y.; Lyu, Q.; Zheng, Y. Bacillus subtilis biofertilizer application reduces chemical fertilization and improves fruit quality in fertigated Tarocco blood orange groves. Sci. Hortic. 2021, 281, 110004. [Google Scholar] [CrossRef]
- Artyszak, A.; Gozdowski, D. The Effect of Growth Activators and Plant Growth-Promoting Rhizobacteria (PGPR) on the Soil Properties, Root Yield, and Technological Quality of Sugar Beet. Agronomy 2020, 10, 1262. [Google Scholar] [CrossRef]
- Ngalimat, M.S.; Yahaya, R.S.R.; Baharudin, M.M.A.-a.; Yaminudin, S.M.; Karim, M.; Ahmad, S.A.; Sabri, S. A Review on the Biotechnological Applications of the Operational Group Bacillus amyloliquefaciens. Microorganisms 2021, 9, 614. [Google Scholar] [CrossRef] [PubMed]
- Matzen, N.; Heick, T.M.; Jørgensen, L.N. Control of powdery mildew (Blumeria graminis spp.) in cereals by Serenade® ASO (Bacillus amyloliquefaciens (former subtilis) strain QST 713). Biol. Control 2019, 139, 104067. [Google Scholar] [CrossRef]
- Montes-Bazurto, L.G.; Bustillo-Pardey, A.E.; Morales, A. New Alternative to Control Stenoma impressella (Lepidoptera: Elachistidae) Using Bacillus thuringiensis Commercial Formulations in Oil Palm Crops. Agronomy 2022, 12, 883. [Google Scholar] [CrossRef]
- Ghahremani, Z.; Escudero, N.; Beltrán-Anadón, D.; Saus, E.; Cunquero, M.; Andilla, J.; Sorribas, F.J. Bacillus firmus strain I-1582, a nematode antagonist by itself and through the plant. Front. Plant Sci. 2020, 11, 796. [Google Scholar] [CrossRef]
- Mendis, H.C.; Thomas, V.P.; Schwientek, P.; Salamzade, R.; Chien, J.-T.; Waidyarathne, P.; Kloepper, J.; De La Fuente, L. Strain-specific quantification of root colonization by plant growth promoting rhizobacteria Bacillus firmus I-1582 and Bacillus amyloliquefaciens QST713 in non-sterile soil and field conditions. PLoS ONE 2018, 13, e0193119. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).