Abstract
Copy number variations (CNVs) are gains and losses of genomic sequence between two individuals of a species when compared to a reference genome. The data from single nucleotide polymorphism (SNP) microarrays are now routinely used for genotyping, but they also can be utilized for copy number detection. Substantial progress has been made in array design and CNV calling algorithms and at least 10 comparison studies in humans have been published to assess them. In this review, we first survey the literature on existing microarray platforms and CNV calling algorithms. We then examine a number of CNV calling tools to evaluate their impacts using bovine high-density SNP data. Large incongruities in the results from different CNV calling tools highlight the need for standardizing array data collection, quality assessment and experimental validation. Only after careful experimental design and rigorous data filtering can the impacts of CNVs on both normal phenotypic variability and disease susceptibility be fully revealed.
1. Introduction
Genomic structural variation, including copy number variation (CNV), has been extensively studied in humans [1,2,3,4,5] and rodents [6,7,8,9]. Initial CNV reports have also been released for domesticated animals, including dog [10,11,12], cattle [13,14], chicken [15,16], pig [17,18], sheep [19,20], and goat [21] amongst others. Recent bovine CNV studies have generated several cattle CNV maps based on the data from Illumina Bovine SNP50K microarrays [22,23,24,25].
CNVs can be identified using various approaches, including comparative genomic hybridization (CGH) arrays, SNP arrays, and DNA sequencing. In spite of the increasing adoption of next-generation sequencing, microarrays will continue to be the primary platform for CNV detection in the near future. Compared to other approaches, the advantages of SNP arrays include their relative low cost and high throughput. Substantial genotyping data have been produced from genome-wide association studies, which can be directly exploited for CNV analysis. Dozens of human and mouse CNV studies have demonstrated that some CNVs are associated with phenotypic traits and diseases [26,27,28,29]. Efforts to explore the association between cattle CNV and economical traits have been published [30,31,32], even though the actual functional mechanisms are not yet well defined.
2. CNV Detection Using SNP Arrays
SNP arrays were initially designed to genotype thousands of SNPs across the genome concurrently. Their applications have now expanded to include CNV detection using additional information such as the probe hybridization signal on each individual chip. The most well-known SNP microarrays are available from commercial vendors such as Illumina and Affymetrix [33,34]. Both companies sell competing arrays and continue to offer ever increasing coverage for detecting SNPs and CNVs simultaneously. However, one important consideration is the inherent bias of the SNP chip coverage against areas of the genome known to frequently harbor CNVs. For example, common copy number polymorphisms (CNPs) may cause a SNP to be rejected when the SNP fails standard inheritance checks and Hardy-Weinberg tests [35].
Segmental duplications (SDs), defined as >1 kb stretches of duplicated DNA with high sequence identity in a species, were shown to be one of the catalysts and hotspots for CNV formation [36,37,38]. Although the current microarray platforms offer some detection power in SD regions, calls within these regions are often affected by low probe density and cross-hybridization of repetitive sequence. In addition, only a relative copy number (CN) increase or decrease is reported with respect to the reference samples in SD regions. This poses a particular problem in the detection of CNVs in SD regions as the test individual’s copy number may differ from that of the reference by a smaller proportion than is detectable using array-based calling criteria. Although analyses of a subset of CNVs provided evidence of linkage disequilibrium with flanking SNPs [39], a significant portion of CNVs fell in genomic regions not well covered by SNP arrays, such as SD regions, and thus were not genotyped [40,41,42].
Since SNP chips are primarily designed for their use in SNP genotyping, some background noise that does not affect SNP calling may cause problems for CNV calling algorithms. For example, SNP data is typically normalized against a reference population in order to reduce between-array variations and probe-specific hybridization effects. The assumption that the large majority of reference samples have the same two copies does not hold for common CNV regions. At these regions, the normalization should be further optimized to derive correct parameters. Several new array designs have incorporated CNV detection, for example, monomorphic probes in common CNV regions are included on more recent Illumina and Affymetrix SNP array platforms.
3. Algorithms for CNV Detection
Undoubtedly, microarray development has spurred the advances in computational analysis methodology in quantitative fields of biology. A wide range of CNV discovery tools has been developed based on data derived from SNP arrays, such as cnvPartition [43], Birdsuite [44], PennCNV [45], and amongst others. In this section, we briefly introduce these CNV detection tools.
cnvPartition: Illumina data can be initially viewed, processed and exported using the proprietary GenomeStudio program (Illumina, CA, USA). In addition to quality checking and genotype calling, the program calculates several important input values for CNV discovery. The log R ratio (LRR), i.e., log2(Robserved/Rexpected), is calculated from the observed normalized intensity of a sample and expected normalized intensity, which is calculated from linear interpolation of canonical genotype clusters. The B allele frequency (BAF, normalized measure of relative signal intensity ratio of the B and A alleles) is calculated from the difference between the actual value and the expected position of the cluster group. LRR and BAF are used by many CNV detection algorithms. cnvPartition is offered as a plug-in for the GenomeStudio program, where it uses LRR and BAF to assess copy number using 14 different Gaussian distribution models between zero and four copies. cnvPartition also uses a likelihood-based method to compute the confidence score for each CNV call. Given the integration of cnvPartition into Illumina proprietary software (GenomeStudio), cnvPartition is currently unable to process and analyze Affymetrix chip data.
Birdsuite: Affymetrix SNP array data from older chips must first be analyzed in the Genotyping Console program provided by Affymetrix for initial quality checks and controls. Data from the newer Affymetrix chip can be processed by additional programs contained in the Birdsuite package [44]. The Canary module of Birdsuite genotypes the known common CNVs using an Expectation-Maximization (EM) algorithm while the Birdseye module detects novel CNVs by using a Hidden Markov Model (HMM) with a Viterbi algorithm calculating emission states. For Affymetrix SNP arrays, there are other freely available CNV detection programs, such as GADA [46], Cokgen [47], iPattern [26] in addition to Birdsuite. For details about these programs, please see these published reviews [35,48,49]. The developers of Birdsuite have mentioned future plans for Illumina platform support [50] but current options only include a beta version for Illumina 610 array platforms.
PennCNV and QuantiSNP: PennCNV and QuantiSNP are two freely available programs developed based on HMMs [45,51]. Both programs can process Illumina and Affymetrix SNP data. PennCNV incorporates multiple sources of information, including LRR and BAF at each SNP marker, the distance between neighboring SNPs and the allele frequency of SNPs. PennCNV also integrates a computational approach by fitting regression models with GC content to overcome “genomic waves” [52,53]. Additionally, PennCNV is capable of considering pedigree information (a parents-offspring trio) to improve call rates and accuracy of breakpoint prediction as well as to infer chromosome-specific SNP genotypes in CNVs. Finally, PennCNV also reports data quality control measurements for each CNV dataset.
QuantiSNP, by contrast, uses an Objective Bayes approach [51] to infer copy number states based on the LogR ratio and the B allele frequency for each SNP marker. Whereas the PennCNV algorithm uses a transition matrix to model realistic copy number transitions between SNP probes [45], QuantiSNP calculates Bayesian probabilities for each SNP marker pair and then uses a HMM to join markers to form CNVs. Another significant difference between the two programs is that PennCNV is an open-source project whereas QuantiSNP was written for MatLab, which may limit availability to users that may not have a MatLab license. Finally, QuantiSNP is no longer under active development as listed on its webpage [54].
Approaches originally developed for array CGH: Several tools for CNV detection, which were originally developed for array CGH CNV calling, have been modified for SNP array analysis. However, these methods normally do not consider BAF information, which is the preferred data source to use for CNV calling in SNP data. For example, the Circular Binary Segmentation (CBS) method was designed to convert noisy intensity values into neighboring segments of distinct assigned copy numbers using dynamic programming [55]. DNAcopy is a widely used R implementation of the CBS method.
Other commercial CNV detection tools: Other commercially available programs include Partek Genomics Suite, Nexus Copy Number software and Golden Helix SNP & Variation Suite (SVS). The strength of these commercial tools include their graphical user interfaces, streamlined pipelines for analysis and work flow, optimized computational speed as well as technical support. These factors are very important to labs with limited bioinformatics support; however, commercial companies often do not utilize some of the latest methods developed in the academic environment. For this study, we have chosen to look in detail at the Golden Helix SVS [56]. The SVS Copy Number Analysis Module (CNAM) employs a segmentation algorithm using only the signal intensity data to detect CNVs on either a per-sample (univariate) or multi-sample (multivariate) basis. According to its online manual, the univariate method, which considers only one sample at a time, is designed for detecting rare and/or large CNVs. The multivariate method, which considers all samples simultaneously, is designed for detecting small, common CNVs.
Comparing univariate and multivariate methods: Although the exact algorithm of each method is proprietary, Breheny et al. explored the strengths and weaknesses of two similar approaches using both simulations and real data [57]. In their study, the univariate method (the CNV-level testing, i.e., across markers within one sample) involves estimating, at the level of the individual genome, the underlying copy number at each location. Once this is completed, tests are performed to determine the association between copy number state and phenotype. The multivariate method (the pooled marker-level testing across samples) carries out association testing first between the phenotypes and raw intensities at the level of the individual marker, and then aggregates neighboring test results to identify CNVs associated with the phenotype. Accounting for multiple comparisons across SNP markers is more straightforward, as a multiple-comparison correction (e.g., Bonferroni, permutation) can directly control the family-wise error rate (FWER) of the overall procedure [58]. False discovery rates can be calculated to account for multiple comparisons with the CNV-level testing method [59]; however, this is more complicated and somewhat conservative. Partially overlapping CNVs across cell lines introduce dependence across the tests, thereby reducing the effective number of independent tests. Breheny et al. confirmed that that the univariate method/CNV-level testing has greater power to detect associations involving large, rare CNVs, while the multivariate method/pooled marker-level testing has greater power to detect associations involving small, common CNVs. It is important to understand these tradeoffs. Several recent papers have proposed to develop methods capable of simultaneously pooling information across both markers and samples for CNV detection and association studies [60,61,62,63,64].
CNV quality score: Many programs like cnvPartition, Birdsuite, PennCNV and QuantiSNP reported CNV quality scores, which are quantitative values indicating CNV confidences. Although their exact meanings and interpretations depend on each algorithm and they are often not reported in microarray studies. These CNV quality scores are important for constructing CNV regions, which can then be used in association studies.
4. Comparing the CNV Detection Algorithms Using Human Data
As shown in Table 1, at least 10 comparisons of the strengths and weaknesses of these array platforms and CNV calling tools have been published using human CNV data. Although published results are quickly outdated as new platforms and tools are introduced, a general theme is consistent across these comparisons. The first of these is the lack of a standard approach to collecting the data and the lack of standardized reference samples; this makes it difficult to compare CNV results across different studies [65]. The second is that CNV results also differ substantially depending on CNV detection methods [35,49]. For example, as the most comprehensive study on this topic, Pinto et al. have systematically compared CNV detection on 11 microarray platforms to evaluate data quality and CNV calling, reproducibility, concordance across array platforms and laboratories, breakpoint accuracy and analysis tool variability [49]. It is surprising that reproducibility in replicate experiments is <70% for most platforms and different analytic tools applied to the same raw data typically yield CNV calls with <50% concordance. The authors attributed these poor reproducibility observations to these facts: (1) large CNVs often overlap with SDs in complex genomic regions (as we described before) and (2) large CNVs also lead to call fragmentation (a single CNV is detected as multiple smaller variants). This led the authors to conclude that, “the striking differences between CNV calls from different platforms and analytic tools highlight the importance of careful assessment of experimental design in discovery and association studies and of strict data curation and filtering in diagnostics” [49].

Table 1.
Survey of recent comparison studies of copy number variation (CNV) detection.
| Authors | Year | Algorithm | Data | Platform | Vendor | Conclusion | Comment | |
|---|---|---|---|---|---|---|---|---|
| Lai [66] | 2005 | CGHseq, Quantreg, CLAC, GLAD, CBS, HMM, Wavelet, Lowess, ChARM, GA and ACE | Simulation and empirical samples for Glioblastoma | array CGH | Custom cDNA array | Several general characteristics of future program development were suggested. | Earlier programs for array CGH. | |
| Baross [67] | 2007 | CNAG, dChip, CNAT, GLAD | Simulation and empirical mental retardation 100K Affymetrix SNP array | SNP array | Affymetrix | Multiple programs were needed to find all real aberrations. | False positive deletions was substantial, but could be greatly reduced by using the SNP genotype information to confirm loss of heterozygosity. | |
| Winchester [35] | 2009 | Birdsuite, CNAT, GADA, PennCNV, QuantiSNP | NA12156, NA15510 | SNP array | Affymetrix, Illumina | Multiple predictions from different software. | Use software designed for the platform. | |
| Dellinger [68] | 2010 | CBS, cnvFinder, cnvPartition, GALD, Nexus, PennCNV and QuantiSNP | Simulation and empirical samples from Singapore cohort study of the risk factors for Myopia | SNP array | Illumina | QuantiSNP outperformed other methods based on ROC curve residuals over most datasets. Nexus Rank and SNPRank have low specificity and high power. Nexus Rank calls oversized CNVs. PennCNV detects one of the fewest numbers of CNVs. | The normalized singleton ratio (NSR) is proposed as a metric for parameter optimization. | |
| Tsuang [69] | 2010 | PennCNV, QuantiSNP, HMMSeg, and cnvPartition | 48 Schizophrenia samples | SNP array | Illumina | Both guidelines for the identification of CNVs inferred from high-density arrays and the establishment of a gold standard for validation of CNVs are needed. | Given the variety of methods used, there will be many false positives and false negatives. | |
| Zhang [70] | 2011 | Birdsuite, Partek Genomics Suite, HelixTree, and PennCNV-affy | ~1,000 Bipolar + 270 HapMap samples | SNP array | Affymetrix | Birdsuite and Partek had higher positive predictive values. | Poor overlap between 2 gold standards (Kidd et al. and Conrad et al.). | |
| Marenne [71] | 2011 | cnvPartition, PennCNV, and QuantiSNP | 96 pair samples from Spanish Bladder Cancer/EPICURO study | SNP array | Illumina | PennCNV was the most reliable algorithm when assessing the number of copies. | Current calling algorithms should be improved for high performance CNV analysis in genome-wide scans. | |
| Pinto [49] | 2011 | Birdsuite, cnvFinder, cnvPartition, dCHIP, ADM-2 (DNA Analytics), Genotyping Console (GTC), iPattern, Nexus Copy Number, Partek Genomics Suite, PennCNV, QuantiSNP | 6 samples in triplicate on 11 array platforms | array CGH, SNP array, and BAC array | Agilent, NimbleGen, Affymetrix, and Illumina | Different analytic tools applied to the same raw data typically yield CNV calls with <50% concordance. Moreover, reproducibility in replicate experiments is <70% for most platforms. | The CNV resource presented here allows independent data evaluation and provides a means to benchmark new algorithms. CNV calls are disproportionally affected by genome complexity as they tend to overlap SDs and a single CNV is detected as multiple smaller variants. | |
| Koike [48] | 2011 | Birdsuite, Birdseye, PennCNV, CGHseg, DNAcopy | HapMap samples | SNP array | Affymetrix | Hidden Markov model-based programs PennCNV and Birdseye (part of Birdsuite), or Birdsuite show better detection performance. | Segmental duplications and interspersed repeats (LINEs) are involved in CNVs. | |
| Eckel-Passow [72] | 2011 | Affymetrix Power Tools (APT), Aroma.Affymetrix, PennCNV and CRLMM | 1,418 GENOA (Genetic Epidemiology Network of Atherosclerosis)/FBPP (Family Blood Pressure Program) samples | SNP array | Affymetrix | Recommended trying multiple algorithms, evaluating concordance/discordance and subsequently consider the union of regions for downstream association tests. | Advocated that software developers need to provide guidance with respect to evaluating and choosing optimal settings in order to obtain optimal results for an individual dataset. | |
5. Comparing CNV Detection Algorithms Using Bovine High-Density SNP Data
We performed an analysis of CNVs based on Illumina BovineHD chips, which contain more than 750,000 SNP markers [73], using PennCNV. As a consequence of the higher SNP count, more CNVs were identified with higher resolution boundaries. In order to provide an additional comparison of CNV detection methods, we have tested three additional tools to call CNVs on the same BovineHD dataset: cnvPartition version 3.6.1, Golden Helix SVS 7.0 and DNAcopy [55]. These four tools were applicable to our dataset (Illumina bead array), available to us (due to existing commercial licensing or free availability) and were not designed specifically for human-based array studies.
In order to perform an accurate and fair comparison of calls across the different methods, our PennCNV calls were derived from the same 630 animals of 27 cattle breeds on the cattle reference assembly UMD3.1 without using trio information [73]. We carried out cnvPartion calling using the default parameters as recommended by Illumina. For the Golden Helix SVS7.0, we used the SVS DSF Export Plug-In 4.1 to export LRRs from the GenomeStudio project. We then utilized CNAM to process the DSF file under the univariate option (minimum 3 markers/segment, a significance level of p = 0.005 for 2,000 pairwise permutations). We also performed DNAcopy analysis based on LRR. Finally, CNV segments were then filtered with a minimum of 3 probes for all 4 tools and a minimum of absolute segment mean values of 0.3 for SVS and DNAcopy.

Table 2.
CNVs and CNVRs identified using PennCNV, cnvPartition, SVS, and DNAcopy.
| Tool | Event | Count | Gain | Loss | Average Length |
|---|---|---|---|---|---|
| PennCNV | CNV | 46,751 (74.2) | 17,796 (28.2) | 28,955 (46.0) | 2,334,244,479 (49,929) |
| CNVR | 3,364 a | 1,382 b | 2,376 c | 147,476,461 (43,840) | |
| cnvPartition | CNV | 16,566 (26.3) | 5,021 (8.0) | 11,545 (18.3) | 2,191,528,246 (132,291) |
| CNVR | 1,298 a | 541 b | 916 c | 172,378,730 (132,803) | |
| SVS | CNV | 92,463 (146.8) | 205 (0.3) | 92,258 (146.4) | 2,234,601,290 (24,168) |
| CNVR | 7,099 a | 78 b | 7,056 c | 151,471,634 (21,337) | |
| DNAcopy | CNV | 41,858 (66.4) | 4,469 (7.1) | 37,389 (59.3) | 1,863,930,368 (44,530) |
| CNVR | 5,961 a | 1,457 b | 5,284 c | 194,287,154 (32,593) |
Numbers in parentheses are values normalized by sample counts, except in the case of the parentheses values in the “Average Length” column, which are average lengths normalized by CNV counts. a These numbers represent non-redundant CNVR counts after merging both gain and loss CNVs identified across all 630 samples. b Gain CNV events were merged separately. c Loss CNV events were merged separately.
A summary of CNV and CNVR results derived from all 630 samples is shown in Table 2. Detailed results can be found in the four worksheets of Supplementary Table 1. Compared to PennCNV results, CNVs and CNVRs in cnvPartition results are fewer and ~3 times longer (45 kb vs. 130 kb, respectively). While PennCNV and cnvPartition have loss/gain ratios of ~1.7 and DNAcopy has a ratio of 3.6, SVS has a ratio over 90, suggesting SVS is more sensitive to loss events than to gain events. Additionally, both SVS and DNAcopy CNVRs (average length approximately 20 kb and 30 kb, respectively) are shorter than PennCNV (~40 kb), and significantly shorter than cnvPartition CNVRs (~132 kb). Similar observations were also obtained when each subspecies/group (i.e. taurine, indicine, composite (taurine × indicine) and African breeds) was processed separately, confirming the above results (data not shown). When we compared CNV calls across subspecies/groups for all four CNV calling methods, CNV counts per sample were higher in African and indicine breeds, intermediate in composite breeds, and lower in taurine breeds, agreeing with our previous results using PennCNV [73]. We then compared the CNVRs from the four datasets derived from our calling programs based on the UMD3.1 cattle reference assembly (Figure 1). Approximately 50 Mb of core CNVRs are shared among the four CNVR sets. We calculated concordances using the ratios of between intersections and unions for both counts and lengths (Table 3 and Figure 1). PennCNV shared more regions (108 Mb or 50.82%) by length and 43.80% by count with cnvPartition than with any other tools. Therefore, we have observed that tools based on similar algorithms and input data (both LRR and BAF) seem to share more common regions. By contrast, PennCNV and SVS shared 24.88% or 60 Mb in length and 13.74% by count. This comparison was consistent with a recent publication based on PennCNV and SVS using human autism samples [74]. When we applied different filtering criteria requiring a minimum of five or 12 probes, the overlap of calls from these two methods increased slightly, ranging from 24–52% by the number of nucleotides that overlapped. We also evaluated the overlaps between loss CNV events across four datasets for each individual sample. The number of bases overlapped by CNVs from each dataset ranged from 26–48%, which agreed with our CNVR overlapping results.
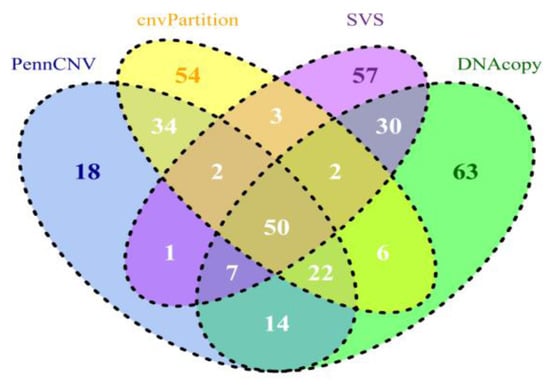
Figure 1.
Comparisons of CNVR results identified by PennCNV, cnvPartition, SVS, and DNAcopy based on genomic location in UMD3.1. The overlap lengths of CNVRs were indicated in Mb.
Figure 1.
Comparisons of CNVR results identified by PennCNV, cnvPartition, SVS, and DNAcopy based on genomic location in UMD3.1. The overlap lengths of CNVRs were indicated in Mb.
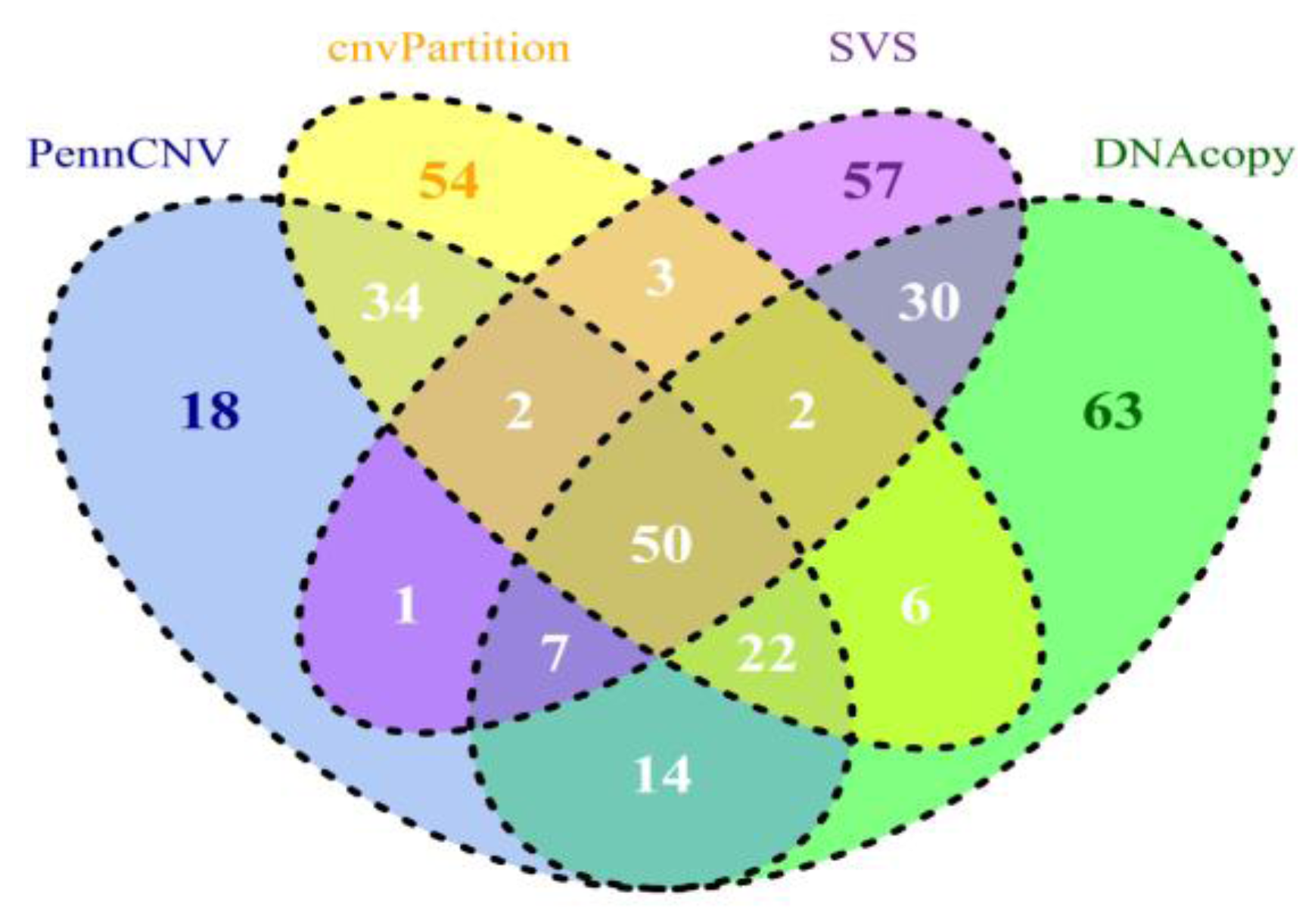

Table 3.
Overlaps among CNVRs across 4 CNV detection tools.
| Count | Length (base pair) | ||||||
|---|---|---|---|---|---|---|---|
| Tool1 | Tool2 | Intersection a | Union a | Percentage | Intersection b | Union b | Percentage |
| PennCNV | cnvPartition | 1,420 | 3,242 | 43.80% | 107,775,740 | 212,079,451 | 50.82% |
| PennCNV | DNAcopy | 2,355 | 6,970 | 33.79% | 93,149,061 | 248,614,554 | 37.47% |
| PennCNV | SVS | 1,264 | 9,199 | 13.74% | 59,557,597 | 239,390,498 | 24.88% |
| cnvPartition | DNAcopy | 1,284 | 5,975 | 21.49% | 79,825,624 | 286,840,260 | 27.83% |
| cnvPartition | SVS | 981 | 7,416 | 13.23% | 56,569,347 | 267,281,017 | 21.16% |
| DNAcopy | SVS | 2,332 | 10,728 | 21.74% | 88,864,805 | 256,893,983 | 34.59% |
a These numbers represent intersections and unions of two CNVR datasets by count. b These numbers represent intersections and unions of two CNVR datasets by length in base pair.
6. Conclusions
Like other published comparisons of CNV calling methods, we observed large variations in calls made by different programs. As pointed out previously, hybridization studies will generate both false positive and false negative results, regardless of how the data are analyzed [75]. As shown in Table 1, many authors recommended using multiple CNV calling algorithms instead of just one [35]; however, although the net effect of this strategy decreases the false negative rate, it also increases the false positive rate. With next generation sequencing projects producing better CNV calling standards, such as the 1,000 human genomes project [5] and our recent effort [76], we should be able to better estimate the false positive and false negative rates for each tool. Large incongruities in the results from different CNV calling tools highlight the need for standardizing array data collection, quality assessment and experimental validation. This is extremely true for other species like cattle, for which there is no gold standard of CNV calls to compare data against. Only after careful experimental design and rigorous data filtering can the impacts of CNVs on both normal phenotypic variability and disease susceptibility be fully revealed.
Acknowledgments
We thank members of the Illumina Bovine HD SNP Consortium for sharing their data. The bovine data will be available to scientists interested in non-commercial research upon signing a Materials Transfer Agreement (MTA). We would also like to thank Reuben Anderson and Alexandre Dimtchev for technical assistance. G.E.L. was supported by NRI/AFRI grants no. 2011-67015-30183 from the USDA NIFA and Project 1265-31000-098-00 from USDA-ARS. Mention of trade names or commercial products in this article is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the US Department of Agriculture. The USDA is an equal opportunity provider and employer.
Conflict of Interest
The authors declare no conflict of interest.
References
- Sebat, J.; Lakshmi, B.; Troge, J.; Alexander, J.; Young, J.; Lundin, P.; Maner, S.; Massa, H.; Walker, M.; Chi, M.; et al. Large-scale copy number polymorphism in the human genome. Science 2004, 305, 525–528. [Google Scholar] [CrossRef]
- Redon, R.; Ishikawa, S.; Fitch, K.R.; Feuk, L.; Perry, G.H.; Andrews, T.D.; Fiegler, H.; Shapero, M.H.; Carson, A.R.; Chen, W.; et al. Global variation in copy number in the human genome. Nature 2006, 444, 444–454. [Google Scholar] [CrossRef]
- Conrad, D.F.; Pinto, D.; Redon, R.; Feuk, L.; Gokcumen, O.; Zhang, Y.; Aerts, J.; Andrews, T.D.; Barnes, C.; Campbell, P.; et al. Origins and functional impact of copy number variation in the human genome. Nature 2009, 464, 704–712. [Google Scholar]
- Altshuler, D.M.; Gibbs, R.A.; Peltonen, L.; Dermitzakis, E.; Schaffner, S.F.; Yu, F.L.; Bonnen, P.E.; de Bakker, P.I.W.; Deloukas, P.; Gabriel, S.B.; et al. Integrating common and rare genetic variation in diverse human populations. Nature 2010, 467, 52–58. [Google Scholar] [CrossRef]
- Mills, R.E.; Walter, K.; Stewart, C.; Handsaker, R.E.; Chen, K.; Alkan, C.; Abyzov, A.; Yoon, S.C.; Ye, K.; Cheetham, R.K.; et al. Mapping copy number variation by population-scale genome sequencing. Nature 2011, 470, 59–65. [Google Scholar] [CrossRef]
- Graubert, T.A.; Cahan, P.; Edwin, D.; Selzer, R.R.; Richmond, T.A.; Eis, P.S.; Shannon, W.D.; Li, X.; McLeod, H.L.; Cheverud, J.M.; et al. A high-resolution map of segmental DNA copy number variation in the mouse genome. PLoS. Genet. 2007, 3, e3. [Google Scholar] [CrossRef]
- Guryev, V.; Saar, K.; Adamovic, T.; Verheul, M.; van Heesch, S.A.; Cook, S.; Pravenec, M.; Aitman, T.; Jacob, H.; Shull, J.D.; et al. Distribution and functional impact of DNA copy number variation in the rat. Nat. Genet. 2008, 40, 538–545. [Google Scholar] [CrossRef]
- She, X.; Cheng, Z.; Zollner, S.; Church, D.M.; Eichler, E.E. Mouse segmental duplication and copy number variation. Nat. Genet. 2008, 40, 909–914. [Google Scholar] [CrossRef]
- Yalcin, B.; Wong, K.; Agam, A.; Goodson, M.; Keane, T.M.; Gan, X.C.; Nellaker, C.; Goodstadt, L.; Nicod, J.; Bhomra, A.; et al. Sequence-based characterization of structural variation in the mouse genome. Nature 2011, 477, 326–329. [Google Scholar] [CrossRef]
- Chen, W.K.; Swartz, J.D.; Rush, L.J.; Alvarez, C.E. Mapping DNA structural variation in dogs. Genome Res. 2009, 19, 500–509. [Google Scholar]
- Nicholas, T.J.; Cheng, Z.; Ventura, M.; Mealey, K.; Eichler, E.E.; Akey, J.M. The genomic architecture of segmental duplications and associated copy number variants in dogs. Genome Res. 2009, 19, 491–499. [Google Scholar]
- Nicholas, T.J.; Baker, C.; Eichler, E.E.; Akey, J.M. A high-resolution integrated map of copy number polymorphisms within and between breeds of the modern domesticated dog. BMC Genomics 2011, 12, 414. [Google Scholar] [CrossRef]
- Liu, G.E.; van Tassell, C.P.; Sonstegard, T.S.; Li, R.W.; Alexander, L.J.; Keele, J.W.; Matukumalli, L.K.; Smith, T.P.; Gasbarre, L.C. Detection of germline and somatic copy number variations in cattle. Dev. Biol. 2008, 132, 231–237. [Google Scholar]
- Liu, G.E.; Hou, Y.; Zhu, B.; Cardone, M.F.; Jiang, L.; Cellamare, A.; Mitra, A.; Alexander, L.J.; Coutinho, L.L.; Dell’aquila, M.E.; et al. Analysis of copy number variations among diverse cattle breeds. Genome Res. 2010, 20, 693–703. [Google Scholar] [CrossRef]
- Volker, M.; Backstrom, N.; Skinner, B.M.; Langley, E.J.; Bunzey, S.K.; Ellegren, H.; Griffin, D.K. Copy number variation, chromosome rearrangement, and their association with recombination during avian evolution. Genome Res. 2010, 20, 503–511. [Google Scholar] [CrossRef]
- Wang, X.F.; Nahashon, S.; Feaster, T.K.; Bohannon-Stewart, A.; Adefope, N. An initial map of chromosomal segmental copy number variations in the chicken. BMC Genomics 2010, 11, 351. [Google Scholar] [CrossRef]
- Fadista, J.; Nygaard, M.; Holm, L.E.; Thomsen, B.; Bendixen, C. A snapshot of CNVs in the pig genome. PLoS ONE 2008, 3, e3916. [Google Scholar] [CrossRef]
- Ramayo-Caldas, Y.; Castelló, A.; Pena, R.N.; Alves, E.; Mercadé, A.; Souza, C.A.; Fernández, A.I.; Perez-Enciso, M.; Folch, J.M. Copy number variation in the porcine genome inferred from a 60 k SNP BeadChip. BMC Genomics 2010, 11, 593. [Google Scholar] [CrossRef]
- Fontanesi, L.; Beretti, F.; Martelli, P.L.; Colombo, M.; Dall’olio, S.; Occidente, M.; Portolano, B.; Casadio, R.; Matassino, D.; Russo, V. A first comparative map of copy number variations in the sheep genome. Genomics 2011, 97, 158–165. [Google Scholar] [CrossRef]
- Liu, J.; Zhang, L.; Xu, L.; Ren, H.; Lu, J.; Zhang, X.; Zhang, S.; Zhou, X.; Wei, C.; Zhao, F.; et al. Analysis of copy number variations in the sheep genome using 50 k SNP BeadChip array. BMC Genomics 2013, 14, 229. [Google Scholar] [CrossRef]
- Fontanesi, L.; Martelli, P.L.; Beretti, F.; Riggio, V.; Dall’olio, S.; Colombo, M.; Casadio, R.; Russo, V.; Portolano, B. An initial comparative map of copy number variations in the goat (Capra hircus) genome. BMC Genomics 2010, 11, 639. [Google Scholar] [CrossRef]
- Hou, Y.; Liu, G.E.; Bickhart, D.M.; Cardone, M.F.; Wang, K.; Kim, E.S.; Matukumalli, L.K.; Ventura, M.; Song, J.; Vanradan, P.M.; et al. Genomic characteristics of cattle copy number variations. BMC Genomics 2011, 12, 127. [Google Scholar] [CrossRef]
- Bae, J.S.; Cheong, H.S.; Kim, L.H.; NamGung, S.; Park, T.J.; Chun, J.Y.; Kim, J.Y.; Pasaje, C.F.; Lee, J.S.; Shin, H.D. Identification of copy number variations and common deletion polymorphisms in cattle. BMC Genomics 2010, 11, 232. [Google Scholar] [CrossRef]
- Fadista, J.; Thomsen, B.; Holm, L.E.; Bendixen, C. Copy number variation in the bovine genome. BMC Genomics 2010, 11, 284. [Google Scholar] [CrossRef]
- Seroussi, E.; Glick, G.; Shirak, A.; Yakobson, E.; Weller, J.I.; Ezra, E.; Zeron, Y. Analysis of copy loss and gain variations in Holstein cattle autosomes using BeadChip SNPs. BMC Genomics 2010, 11, 673. [Google Scholar] [CrossRef]
- Pinto, D.; Pagnamenta, A.T.; Klei, L.; Anney, R.; Merico, D.; Regan, R.; Conroy, J.; Magalhaes, T.R.; Correia, C.; Abrahams, B.S.; et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 2010, 466, 368–372. [Google Scholar] [CrossRef]
- Cook, E.H., Jr.; Scherer, S.W. Copy-number variations associated with neuropsychiatric conditions. Nature 2008, 455, 919–923. [Google Scholar] [CrossRef]
- Sebat, J.; Lakshmi, B.; Malhotra, D.; Troge, J.; Lese-Martin, C.; Walsh, T.; Yamrom, B.; Yoon, S.; Krasnitz, A.; Kendall, J.; et al. Strong association of de novo copy number mutations with autism. Science 2007, 316, 445–449. [Google Scholar] [CrossRef]
- Aitman, T.J.; Dong, R.; Vyse, T.J.; Norsworthy, P.J.; Johnson, M.D.; Smith, J.; Mangion, J.; Roberton-Lowe, C.; Marshall, A.J.; Petretto, E.; et al. Copy number polymorphism in Fcgr3 predisposes to glomerulonephritis in rats and humans. Nature 2006, 439, 851–855. [Google Scholar] [CrossRef]
- Liu, G.E.; Brown, T.; Hebert, D.A.; Cardone, M.F.; Hou, Y.L.; Choudhary, R.K.; Shaffer, J.; Amazu, C.; Connor, E.E.; Ventura, M.; et al. Initial analysis of copy number variations in cattle selected for resistance or susceptibility to intestinal nematodes. Mamm. Genome 2011, 22, 111–121. [Google Scholar] [CrossRef]
- Hou, Y.; Liu, G.E.; Bickhart, D.M.; Matukumalli, L.K.; Li, C.; Song, J.; Gasberre, L.C.; van Tassell, C.P.; Sonstegard, T.S. Genomic regions showing copy number variations associate with resistance or susceptibility to gastrointestinal nematodes in Angus cattle. Funct. Integr. Genomics 2011, 12, 81–92. [Google Scholar]
- Hou, Y.; Bickhart, D.M.; Chung, H.; Hutchison, J.L.; Norman, H.D.; Connor, E.E.; Liu, G.E. Analysis of copy number variations in Holstein cows identify potential mechanisms contributing to differences in residual feed intake. Funct. Integr. Genomics 2012, 12, 717–723. [Google Scholar] [CrossRef]
- LaFramboise, T. Single nucleotide polymorphism arrays: A decade of biological, computational and technological advances. Nucleic Acids Res. 2009, 37, 4181–4193. [Google Scholar] [CrossRef]
- Rincon, G.; Weber, K.L.; van Eenennaam, A.L.; Golden, B.L.; Medrano, J.F. Hot topic: Performance of bovine high-density genotyping platforms in Holsteins and Jerseys. J. Dairy Sci. 2011, 94, 6116–6121. [Google Scholar] [CrossRef]
- Winchester, L.; Yau, C.; Ragoussis, J. Comparing CNV detection methods for SNP arrays. Brief. Funct. Genomic Proteomic 2009, 8, 353–366. [Google Scholar] [CrossRef]
- Sharp, A.J.; Locke, D.P.; McGrath, S.D.; Cheng, Z.; Bailey, J.A.; Vallente, R.U.; Pertz, L.M.; Clark, R.A.; Schwartz, S.; Segraves, R.; et al. Segmental duplications and copy-number variation in the human genome. Am. J. Hum. Genet. 2005, 77, 78–88. [Google Scholar] [CrossRef]
- Marques-Bonet, T.; Girirajan, S.; Eichler, E.E. The origins and impact of primate segmental duplications. Trends Genet. 2009, 25, 443–454. [Google Scholar] [CrossRef]
- Alkan, C.; Kidd, J.M.; Marques-Bonet, T.; Aksay, G.; Antonacci, F.; Hormozdiari, F.; Kitzman, J.O.; Baker, C.; Malig, M.; Mutlu, O.; et al. Personalized copy number and segmental duplication maps using next-generation sequencing. Nat. Genet. 2009, 41, 1061–1067. [Google Scholar] [CrossRef]
- McCarroll, S.A.; Kuruvilla, F.G.; Korn, J.M.; Cawley, S.; Nemesh, J.; Wysoker, A.; Shapero, M.H.; de Bakker, P.I.; Maller, J.B.; Kirby, A.; et al. Integrated detection and population-genetic analysis of SNPs and copy number variation. Nat. Genet. 2008, 40, 1166–1174. [Google Scholar] [CrossRef]
- Estivill, X.; Armengol, L. Copy number variants and common disorders: Filling the gaps and exploring complexity in genome-wide association studies. PLoS Genet. 2007, 3, 1787–1799. [Google Scholar]
- Locke, D.P.; Sharp, A.J.; McCarroll, S.A.; McGrath, S.D.; Newman, T.L.; Cheng, Z.; Schwartz, S.; Albertson, D.G.; Pinkel, D.; Altshuler, D.M.; et al. Linkage disequilibrium and heritability of copy-number polymorphisms within duplicated regions of the human genome. Am. J. Hum. Genet. 2006, 79, 275–290. [Google Scholar] [CrossRef]
- Campbell, C.D.; Sampas, N.; Tsalenko, A.; Sudmant, P.H.; Kidd, J.M.; Malig, M.; Vu, T.H.; Vives, L.; Tsang, P.; Bruhn, L.; et al. Population-genetic properties of differentiated human copy-number polymorphisms. Am. J. Human Genet. 2011, 88, 317–332. [Google Scholar] [CrossRef]
- Illumina—Sequencing and Array-Based Solutions for Genetic Research. Available online: http://www.illumina.com (accessed on 6 June 2013).
- Korn, J.M.; Kuruvilla, F.G.; McCarroll, S.A.; Wysoker, A.; Nemesh, J.; Cawley, S.; Hubbell, E.; Veitch, J.; Collins, P.J.; Darvishi, K.; et al. Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat. Genet. 2008, 40, 1253–1260. [Google Scholar] [CrossRef]
- Wang, K.; Li, M.; Hadley, D.; Liu, R.; Glessner, J.; Grant, S.F.; Hakonarson, H.; Bucan, M. PennCNV: An integrated hidden Markov model designed for high-resolution copy number variation detection in whole-genome SNP genotyping data. Genome Res. 2007, 17, 1665–1674. [Google Scholar] [CrossRef]
- Pique-Regi, R.; Monso-Varona, J.; Ortega, A.; Seeger, R.C.; Triche, T.J.; Asgharzadeh, S. Sparse representation and Bayesian detection of genome copy number alterations from microarray data. Bioinformatics 2008, 24, 309–318. [Google Scholar] [CrossRef]
- Yavas, G.; Koyuturk, M.; Ozsoyoglu, M.; Gould, M.P.; LaFramboise, T. An optimization framework for unsupervised identification of rare copy number variation from SNP array data. Genome Biol. 2009, 10, R119. [Google Scholar] [CrossRef]
- Koike, A.; Nishida, N.; Yamashita, D.; Tokunaga, K. Comparative analysis of copy number variation detection methods and database construction. BMC Genet. 2011, 12, 29. [Google Scholar] [CrossRef]
- Pinto, D.; Darvishi, K.; Shi, X.H.; Rajan, D.; Rigler, D.; Fitzgerald, T.; Lionel, A.C.; Thiruvahindrapuram, B.; MacDonald, J.R.; Mills, R.; et al. Comprehensive assessment of array-based platforms and calling algorithms for detection of copy number variants. Nat. Biotechnol. 2011, 29, 512–520. [Google Scholar] [CrossRef]
- Birdsuite FAQ. Broad Institute of MIT and Harvard. Available online: http://www.broadinstitute.org/science/programs/medical-and-population-genetics/birdsuite/birdsuite-faq (accessed on 6 June 2013).
- Colella, S.; Yau, C.; Taylor, J.M.; Mirza, G.; Butler, H.; Clouston, P.; Bassett, A.S.; Seller, A.; Holmes, C.C.; Ragoussis, J. QuantiSNP: An objective bayes hidden-Markov model to detect and accurately map copy number variation using SNP genotyping data. Nucleic Acids Res. 2007, 35, 2013–2025. [Google Scholar] [CrossRef]
- Marioni, J.C.; Thorne, N.P.; Valsesia, A.; Fitzgerald, T.; Redon, R.; Fiegler, H.; Andrews, T.D.; Stranger, B.E.; Lynch, A.G.; Dermitzakis, E.T.; et al. Breaking the waves: Improved detection of copy number variation from microarray-based comparative genomic hybridization. Genome Biol. 2007, 8, R228. [Google Scholar] [CrossRef]
- Diskin, S.J.; Li, M.; Hou, C.; Yang, S.; Glessner, J.; Hakonarson, H.; Bucan, M.; Maris, J.M.; Wang, K. Adjustment of genomic waves in signal intensities from whole-genome SNP genotyping platforms. Nucleic Acids Res. 2008, 36, e126. [Google Scholar] [CrossRef]
- QuantiSNP. Available online: http://sites.google.com/site/quantisnp/ (accessed on 6 June 2013).
- Olshen, A.B.; Venkatraman, E.S.; Lucito, R.; Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 2004, 5, 557–572. [Google Scholar] [CrossRef]
- Genetic Association Software, Genome-Wide Association (GWAS) Software for SNP, CNV, and NGS. Available online: http://www.goldenhelix.com/SNP_Variation/ (accessed on 6 June 2013).
- Breheny, P.; Chalise, P.; Batzler, A.; Wang, L.; Fridley, B.L. Genetic association studies of copy-number variation: Should assignment of copy number states precede testing? PLoS ONE 2012, 7, e34262. [Google Scholar] [CrossRef]
- Storey, J.D.; Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 2003, 100, 9440–9445. [Google Scholar] [CrossRef]
- Benjamini, Y.; Yekutieli, D. The control of the false discovery rate in multiple testing under dependency. Ann. Stat. 2001, 29, 1165–1188. [Google Scholar] [CrossRef]
- Li, B.; Leal, S.M. Methods for detecting associations with rare variants for common diseases: Application to analysis of sequence data. Am. J. Hum. Genet. 2008, 83, 311–321. [Google Scholar] [CrossRef]
- Yang, H.C.; Hsieh, H.Y.; Fann, C.S. Kernel-based association test. Genetics 2008, 179, 1057–1068. [Google Scholar] [CrossRef]
- Baladandayuthapani, V.; Ji, Y.; Talluri, R.; Nieto-Barajas, L.E.; Morris, J.S. Bayesian random segmentation models to identify shared copy number aberrations for array CGH data. J. Am. Stat. Assoc. 2010, 105, 1358–1375. [Google Scholar] [CrossRef]
- Nowak, G.; Hastie, T.; Pollack, J.R.; Tibshirani, R. A fused lasso latent feature model for analyzing multi-sample aCGH data. Biostatistics 2011, 12, 776–791. [Google Scholar] [CrossRef]
- Glessner, J.T.; Li, J.; Hakonarson, H. ParseCNV integrative copy number variation association software with quality tracking. Nucleic Acids Res. 2013, 41, e64. [Google Scholar] [CrossRef]
- Scherer, S.W.; Lee, C.; Birney, E.; Altshuler, D.M.; Eichler, E.E.; Carter, N.P.; Hurles, M.E.; Feuk, L. Challenges and standards in integrating surveys of structural variation. Nat. Genet. 2007, 39, S7–S15. [Google Scholar] [CrossRef]
- Lai, W.R.; Johnson, M.D.; Kucherlapati, R.; Park, P.J. Comparative analysis of algorithms for identifying amplifications and deletions in array CGH data. Bioinformatics 2005, 21, 3763–3770. [Google Scholar] [CrossRef]
- Baross, A.; Delaney, A.D.; Li, H.I.; Nayar, T.; Flibotte, S.; Qian, H.; Chan, S.Y.; Asano, J.; Ally, A.; Cao, M.; et al. Assessment of algorithms for high throughput detection of genomic copy number variation in oligonucleotide microarray data. BMC Bioinformatics 2007, 8, 368. [Google Scholar] [CrossRef]
- Dellinger, A.E.; Saw, S.M.; Goh, L.K.; Seielstad, M.; Young, T.L.; Li, Y.J. Comparative analyses of seven algorithms for copy number variant identification from single nucleotide polymorphism arrays. Nucleic Acids Res. 2010, 38, e105. [Google Scholar] [CrossRef]
- Tsuang, D.W.; Millard, S.P.; Ely, B.; Chi, P.; Wang, K.; Raskind, W.H.; Kim, S.; Brkanac, Z.; Yu, C.E. The effect of algorithms on copy number variant detection. PLoS ONE 2010, 5, e14456. [Google Scholar] [CrossRef]
- Zhang, D.; Qian, Y.; Akula, N.; Alliey-Rodriguez, N.; Tang, J.; Gershon, E.S.; Liu, C. Accuracy of CNV detection from GWAS data. PLoS ONE 2011, 6, e14511. [Google Scholar] [CrossRef]
- Marenne, G.; Rodriguez-Santiago, B.; Closas, M.G.; Perez-Jurado, L.; Rothman, N.; Rico, D.; Pita, G.; Pisano, D.G.; Kogevinas, M.; Silverman, D.T.; et al. Assessment of copy number variation using the Illumina Infinium 1M SNP-array: A comparison of methodological approaches in the Spanish Bladder Cancer/EPICURO study. Hum. Mutat. 2011, 32, 240–248. [Google Scholar] [CrossRef]
- Eckel-Passow, J.E.; Atkinson, E.J.; Maharjan, S.; Kardia, S.L.; de Andrade, M. Software comparison for evaluating genomic copy number variation for Affymetrix 6.0 SNP array platform. BMC Bioinformatics 2011, 12, 220. [Google Scholar] [CrossRef]
- Hou, Y.; Bickhart, D.M.; Hvinden, M.L.; Li, C.; Song, J.; Boichard, D.A.; Fritz, S.; Eggen, A.; Denise, S.; Wiggans, G.R.; et al. Fine mapping of copy number variations on two cattle genome assemblies using high density SNP array. BMC Genomics 2012, 13, 376. [Google Scholar] [CrossRef]
- Matsunami, N.; Hadley, D.; Hensel, C.H.; Christensen, G.B.; Kim, C.; Frackelton, E.; Thomas, K.; da Silva, R.P.; Stevens, J.; Baird, L.; et al. Identification of rare recurrent copy number variants in high-risk autism families and their prevalence in a large ASD population. PLoS ONE 2013, 8, e52239. [Google Scholar] [CrossRef]
- Carter, N.P. Methods and strategies for analyzing copy number variation using DNA microarrays. Nat. Genet. 2007, 39, S16–S21. [Google Scholar] [CrossRef]
- Bickhart, D.M.; Hou, Y.; Schroeder, S.G.; Alkan, C.; Cardone, M.F.; Matukumalli, L.K.; Song, J.; Schnabel, R.D.; Ventura, M.; Taylor, J.F.; et al. Copy number variation of individual cattle genomes using next-generation sequencing. Genome Res. 2012, 22, 778–790. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
