Abstract
Exploring the spatiotemporal dynamic patterns of multi-channel electroencephalography (EEG) is crucial for interpreting dementia and related cognitive decline. Spatiotemporal patterns of EEG can be described through microstate analysis, which provides a discrete approximation of the continuous electric field patterns generated by the brain cortex. Here, we propose a novel microstate spatiotemporal dynamic indicator, termed the microstate sequence non-randomness index (MSNRI). The essence of the method lies in initially generating a sequence of microstate transition patterns through state space compression of EEG data using microstate analysis. Following this, we assess the non-randomness of these microstate patterns using information-based similarity analysis. The results suggest that this MSNRI metric is a potential marker for distinguishing between health control (HC) and frontotemporal dementia (FTD) (HC vs. FTD: 6.958 vs. 5.756, p < 0.01), as well as between HC and populations with Alzheimer’s disease (AD) (HC vs. AD: 6.958 vs. 5.462, p < 0.001). Healthy individuals exhibit more complex macroscopic structures and non-random spatiotemporal patterns of microstates, whereas dementia disorders lead to more random spatiotemporal patterns. Additionally, we extend the proposed method by integrating the Complementary Ensemble Empirical Mode Decomposition (CEEMD) method to explore spatiotemporal dynamic patterns of microstates at specific frequency scales. Moreover, we assessed the effectiveness of this innovative method in predicting cognitive scores. The results demonstrate that the incorporation of CEEMD-enhanced microstate dynamic indicators significantly improved the prediction accuracy of Mini-Mental State Examination (MMSE) scores (R2 = 0.940). The CEEMD-enhanced MSNRI method not only aids in the exploration of large-scale neural changes in populations with dementia but also offers a robust tool for characterizing the dynamics of EEG microstate transitions and their impact on cognitive function.
1. Introduction
Global population aging is rapidly intensifying, leading to a growing number of individuals experiencing cognitive decline. According to estimates by the Global Burden of Disease study, there are currently 57 million people worldwide living with dementia, and it is projected to increase to 153 million in 2050 [1]. Alzheimer’s disease (AD) and frontotemporal dementia (FTD) are progressive neurodegenerative conditions that primarily afflict elderly individuals. AD, the most prevalent form of dementia, constitutes 60–80% of cases, whereas FTD, though less common, accounts for 5–10% of cases [2,3]. Cognitive decline and behavioral changes represent characteristic features of both AD and FTD, albeit affecting the brain in distinct manners.
For an extended period, researchers have shared a common goal of exploring biomarkers or other specific indicators for dementia screening and diagnosis, aiming to replace diagnostic methods based solely on scales or tests. Methods such as positron emission tomography (PET) and magnetic resonance imaging (MRI) can identify certain pathological changes associated with diseases, but they are expensive and have low acceptance rates among patients with non-severe conditions. There is an urgent need for faster and more cost-effective biomarker alternatives to aid in the early detection of dementia.
As technology has advanced, electroencephalography (EEG) has emerged as a crucial tool for investigating and comprehending brain function. Previous studies have highlighted three typical effects on resting-state EEG signals in patients with dementia compared to healthy individuals: diffuse slowing, reduced complexity, and decreased synchrony [4,5]. Changes in the power spectrum, shifting from high-frequency components (alpha, beta, and gamma bands) to low-frequency components (delta and theta bands), are frequently observed in patients with dementia [6]. Reduced complexity refers to a decline in the complexity of brain electrical activity observed in patients with dementia compared to healthy individuals [7]. Additionally, decreased connectivity between cortical areas is seen in many patients with dementia [8].
EEG microstates consider the spatiotemporal evolution of cortical EEG signals, describing the dynamic characteristics of the entire brain neural networks. Renowned for its remarkable resolution in both space and time, microstate analysis finds extensive utility across various domains including dementia [9,10], depression [11], cognitive psychology [12], and sleep [13], facilitating deeper understanding and exploration within these fields. Previous studies predominantly focused on temporal statistical characteristics of microstates, such as duration, occurrence rate, coverage rate, and transition probabilities, while overlooking the broader patterns and characteristics of large-scale changes in brain activity. Recent research efforts have shifted towards investigating dynamic information related to microstate transition patterns. These include analyzing scale-free properties of microstate time series [14], conducting Lempel–Ziv complexity analysis on microstate transitions [15], and exploring microstate entropy rate analysis [16], among others. From the perspective of the brain’s complex systems, there is an urgent need to further investigate analytical methods that can effectively encapsulate the spatio-temporal dynamics of microstates, thereby revealing the dynamic transition patterns inherent in microstate sequences. We intend to employ an information-based similarity analysis tool [17] to detect and quantify certain fundamental patterns embedded in EEG microstates.
Additionally, exploring the spatiotemporal characteristics of brain electrical signals at different frequency scales is also meaningful. Traditional microstate analysis only analyzes the spatio-temporal dynamics of microstates in a wide range of frequency bands and lacks the mining of the spatio-temporal dynamics of EEG microstates in different frequency bands. Researchers have attempted to explicitly decompose the spatiotemporal dynamics of microstates within four discrete, narrowband frequency bands (delta, theta, alpha, and beta) and compare them with classical analyses of broadband signals [18]. Victor Férat et al. argued that microstate analysis with specific frequency decomposition can provide finer-grained spectral information not observable with standard broadband analysis. Although their study was validated in healthy subjects, it is anticipated that this analytical approach will contribute to biomarker research in populations with dementia. Further exploration is required to investigate independent spatiotemporal dynamics indicators across various EEG frequency domain scales, which can be applied to biomarker research in patients with dementia and predictive studies of cognitive function. We intend to investigate the spatio-temporal patterns of EEG microstates across various frequency band scales based on adaptive frequency domain decomposition.
While some prior studies have compared EEG biomarkers between patients with dementia or mild cognitive impairment and healthy controls [15,19,20,21], this study, in addition to investigating EEG microstate dynamic markers in studying health and disease, also focuses on observing the predictive capacity of EEG markers for Mini-Mental State Examination (MMSE) scores.
In light of these challenges, we propose three hypotheses: (1) dementia causes a disruption in the normal transition patterns of EEG microstates, (2) the non-randomness quantification of EEG microstates’ transition patterns can be used to predict cognitive decline scores, and (3) cross-frequency bands microstate analysis can offer more comprehensive insights into dementia disease. To explore these concepts, we introduce a novel method for microstate sequence similarity analysis based on information theory, which captures the microstate sequence non-randomness index (MSNRI) within the underlying patterns of microstates transitions. The effectiveness of this novel approach is evaluated using a publicly available EEG datasets including health controls (HC), FTD, and AD participants [2]. Furthermore, we explored microstate dynamics information across multiple frequencies. Based on the complementary ensemble empirical mode decomposition (CEEMD) [22] method, we introduce a CEEMD-enhanced MSNRI method to provide finer-grained pattern variation information that conventional broadband analysis cannot capture. This research contributes to a more comprehensive understanding of how the dementia brain works, representing a crucial step towards understanding and characterizing the complexity of brain systems.
2. Materials and Methods
2.1. Dataset
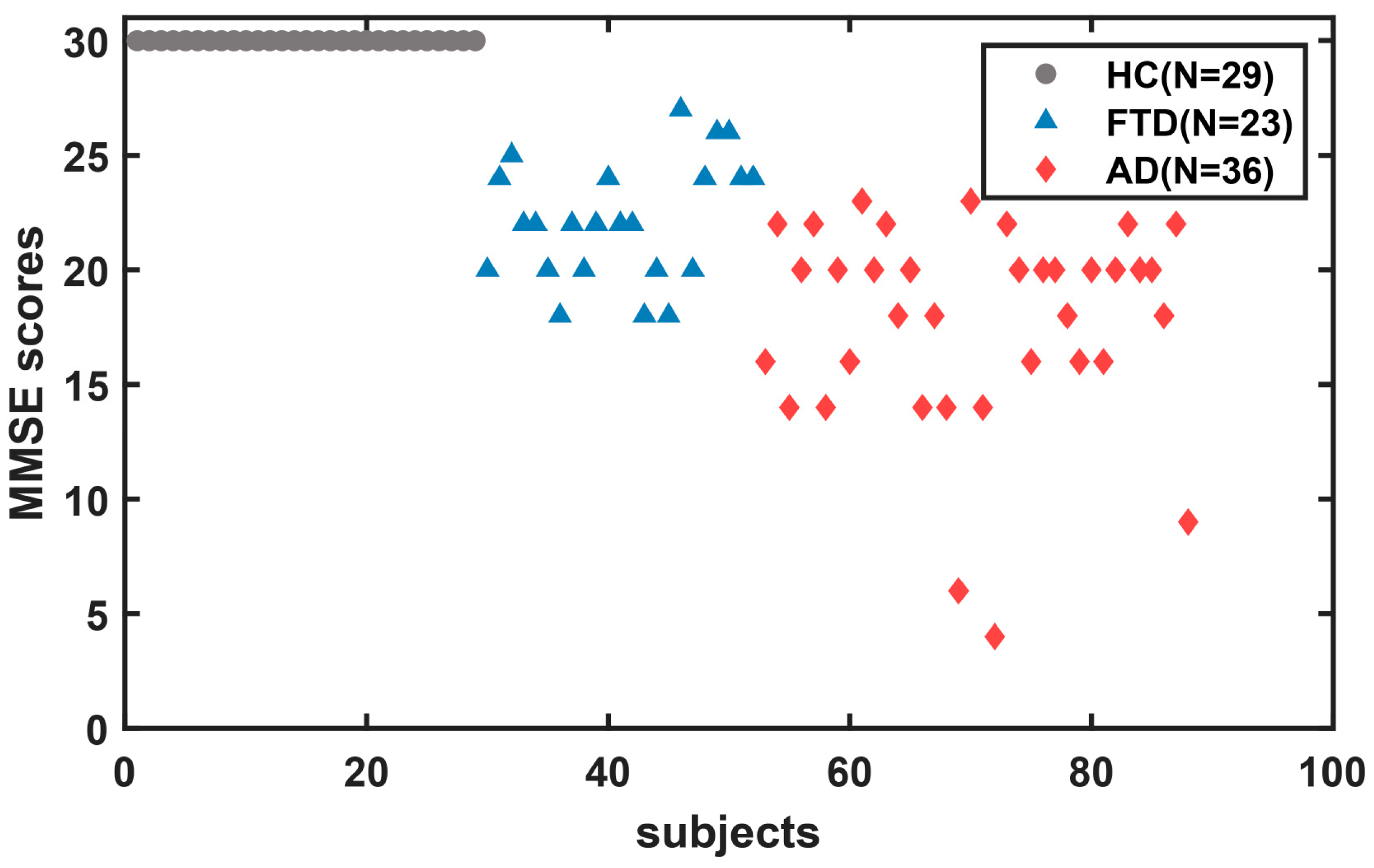
The dataset contains closed-eye resting-state EEG recordings of 88 participants obtained from the Second Neurology Department of the AHEPA General University Hospital in Thessaloniki [2]. Detailed protocol and inclusion criteria are reported in the literature [2]. Thirty-six of them (13 males) were diagnosed with AD, 23 (14 males) were diagnosed with FTD, and 29 (11 males) were healthy controls (HC). The initial diagnosis of patients with AD and FTD was made in accordance with the Diagnostic and Statistical Manual of Mental Disorders, Third Revised Edition (DSM-IIIR, DSM IV, ICD-10) [23], and the criteria provided by the National Institute of Neurological, Communication Disorders and Stroke—Alzheimer’s Disease and Related Disorders Association (NINCDS-ADRDA) [24]. The demographic characteristics are evaluated in Table 1. Cognitive and neuropsychological status was evaluated using the MMSE; the MMSE includes multiple cognitive subfields, such as orientation, memory, attention and calculation, recall ability, and language ability. Total MMSE scores range from 0 to 30, with higher scores indicating better cognitive functioning. MMSE scores of 27–30 indicate normal function, 21–26 indicate mild impairment, 10–20 suggest moderate, and 0–9 suggest severe cognitive impairment. It is important to note that MMSE scores were not the sole criterion for diagnosing cognitive disorders in this study. The scatter plot of total MMSE scores for all subjects is shown in Figure 1, with a mean total MMSE of 17.75 (SD = 4.5) in the AD group, 22.17 (SD = 8.22) in the FTD group, and 30 in the HC group. And the detailed MMSE scores for each subject are shown in Table S1. This study was approved by the Scientific and Ethical Committee of the AHEPA University Hospital of the Aristotle University of Thessaloniki under the protocol number 142/12-04-2023.

Table 1.
Demographic characterization and database description.
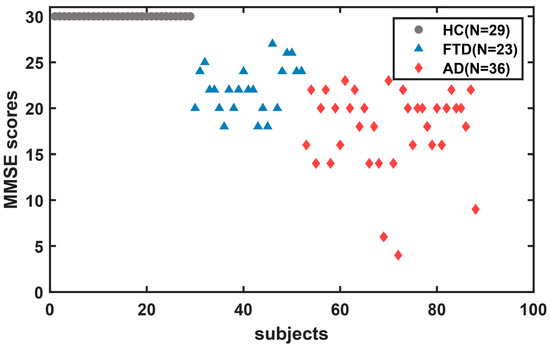
Figure 1.
Scatter plot of total MMSE scores for all subjects.
The EEG recordings encompass signals from 19 scalp electrodes (Fp1, Fp2, F7, F3, Fz, F4, F8, T3, C3, Cz, C4, T4, T5, P3, Pz, P4, T6, O1, and O2) and 2 reference electrodes (A1, A2), arranged in accordance with the 10–20 international system. Each recording session adhered to a standardized clinical protocol, with participants seated and their eyes closed. Prior to the onset of recording, impedance levels for each electrode were meticulously maintained below 5 kΩ. The recordings were conducted at a sampling rate of 500 Hz, with a resolution of 10 uV/mm. For the AD group, recording sessions lasted approximately 13.5 min each, ranging from 5.1 to 21.3 min. In the FTD group, sessions lasted around 12 min per session, ranging from 7.9 to 16.9 min. For the HC group, recording sessions lasted approximately 13.8 min each, with durations ranging from 12.5 to 16.5 min. Overall, the dataset comprised a cumulative duration of 485.5 min for AD recordings, 276.5 min for FTD recordings, and 402 min for HC recordings.
2.2. Preprocessing
The preprocessing of EEG signals was conducted using the Fieldtrip toolbox (version 20220215) implemented in Matlab R2023a [25]. The EEG signals were filtered using a notch filter with a frequency range of 49.8–50.2 Hz, along with a 5th order high-pass filter with a cutoff frequency of 0.3 Hz and a 5th order low-pass filter with a cutoff frequency of 70 Hz.
The EEG data were subsequently down-sampled to 250 Hz and re-referenced to the average reference. After filtering, we conducted a visual inspection of signal quality and removed segments with significant artifacts. The eye movements, eye blinks, or heartbeat-related signal contaminants were removed through infomax independent component analysis (ICA) with the “fastica” method [26]. The preprocessed data for each participant, comprising 180 s of data, were selected for subsequent analysis.
2.3. EEG Microstate Analysis
We performed the EEG microstate analysis using the functions provided by Microstate EEGlab Toolbox (version 1.0) [27]. We first extracted the peak of the Global Field Power (GFP), setting the minimum peak distance to 10 ms, and configuring the number of GFP peaks for segmentation for each participant to be 2000. Additionally, any GFP peaks exceeding one standard deviation of the GFPs across all maps were excluded from consideration. For microstate segmentation, we employed a modified K-means algorithm, disregarding the polarity of EEG topographies during clustering [28,29]. Here, we set the number of microstates to four based on previous findings from methods employing K-means clustering, which demonstrated, through cross-validation, that this number yielded optimal clustering with high reproducibility [30,31]. We configured the algorithm with 50 random initializations and a maximum of 1000 iterations. Additionally, the threshold of convergence based on relative change in noise variance was set to 1 × 10−6. The original toolkit documentation includes a complete set of sample code for microstate analysis.
Then, the clustered stable-state templates were reverse-fitted onto the EEG or IMFs of each participant, marking the microstate categories for segments of EEG or IMFs. Additionally, to mitigate short-term continuous changes in microstate labeling caused by noise, microstates with durations of less than 30 ms were classified as the next most probable microstate class based on global map dissimilarity (GMD). Next, we computed classical microstate dynamic characteristics, including duration (the average duration of a microstate activity in milliseconds), occurrence (the frequency of occurrence of a particular microstate), and coverage (the percentage of time that each microstate category occupies throughout the entire period of analysis).
When evaluating the microstate templates for health and disease, we created group-level microstate templates separately for the HC group, FTD group, and AD group, and global maps between groups were aligned by calculating correlation coefficients of the maps and visual inspection. However, in subsequent analyses of microstate indicators, we uniformly employed the microstate template of all participants to reverse-fit the data of each individual participant.
2.4. CEEMD-Enhanced Microstate Sequence Non-Randomness Analysis
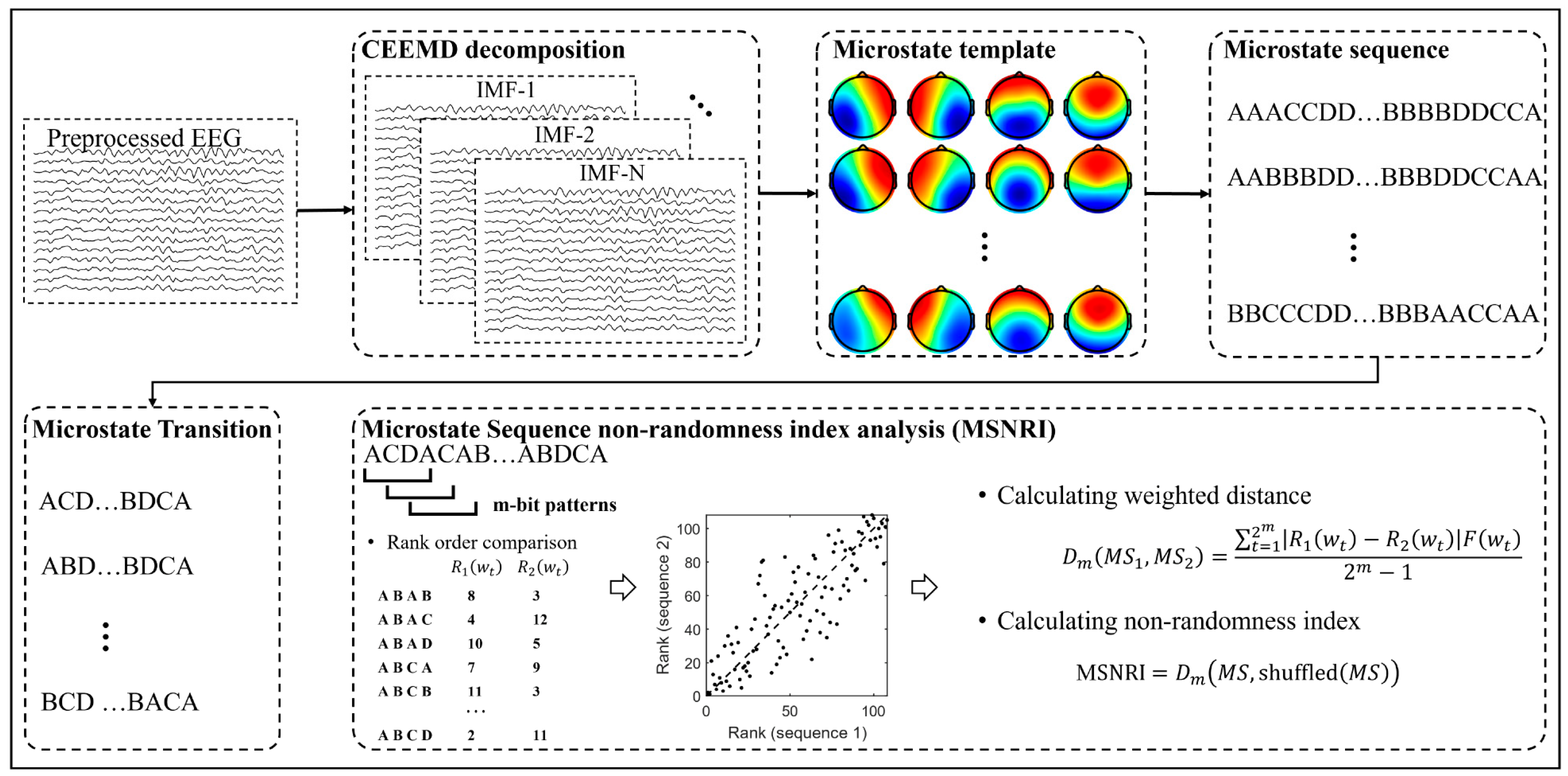
Here we propose a novel measure of EEG microstate transitioning, called microstate sequence non-randomness analysis. In addition to studying the spatio-temporal patterns of microstates at a broadband scale or at narrowband frequency bands (delta, theta, alpha, and beta), this study explores the dynamic patterns of microstates at multiple adaptive frequency scales applying CEEMD. The schematic flow of CEEMD-enhanced microstate sequences non-randomness analysis is shown in Figure 2.
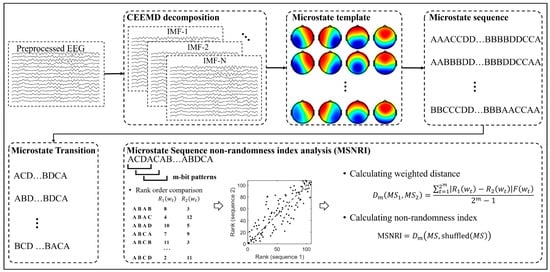
Figure 2.
Schematic flow of CEEMD-enhanced microstate sequences non-randomness index analysis. Step 1: decompose broadband EEG signals into a sequence of Intrinsic Mode Functions (IMFs) using CEEMD. Step 2: perform the microstate analysis on different IMFs and reconstruct the microstate sequence based on the microstate topographic template at each IMF (Red and blue indicate positive and negative values, respectively). Step 3: construct microstate transition sequences from microstate sequences. Step 4: calculate the microstate sequence non-randomness index according to the proposed methodology.
Prior to conducting microstate analysis, broadband EEG signals are decomposed into a sequence of Intrinsic Mode Functions (IMFs) using CEEMD. These IMFs elucidate the oscillatory patterns of EEG across various frequency scales, enabling microstate analysis to be conducted at different IMF scales thereafter.
The CEEMD proposed by Yeh et al. [22] effectively addresses the mode mixing issue of the Empirical Mode Decomposition (EMD) algorithm and the noise residue problem of the Ensemble Empirical Mode Decomposition (EEMD) algorithm. CEEMD and EEMD have similar computational processes, with the difference being that EEMD adds white noise only once each time, while the CEEMD algorithm adds white noise to the original signal and subtracts white noise from the original signal. Both signals undergo EMD decomposition simultaneously, and the average is found to counteract the noise added to the signal.
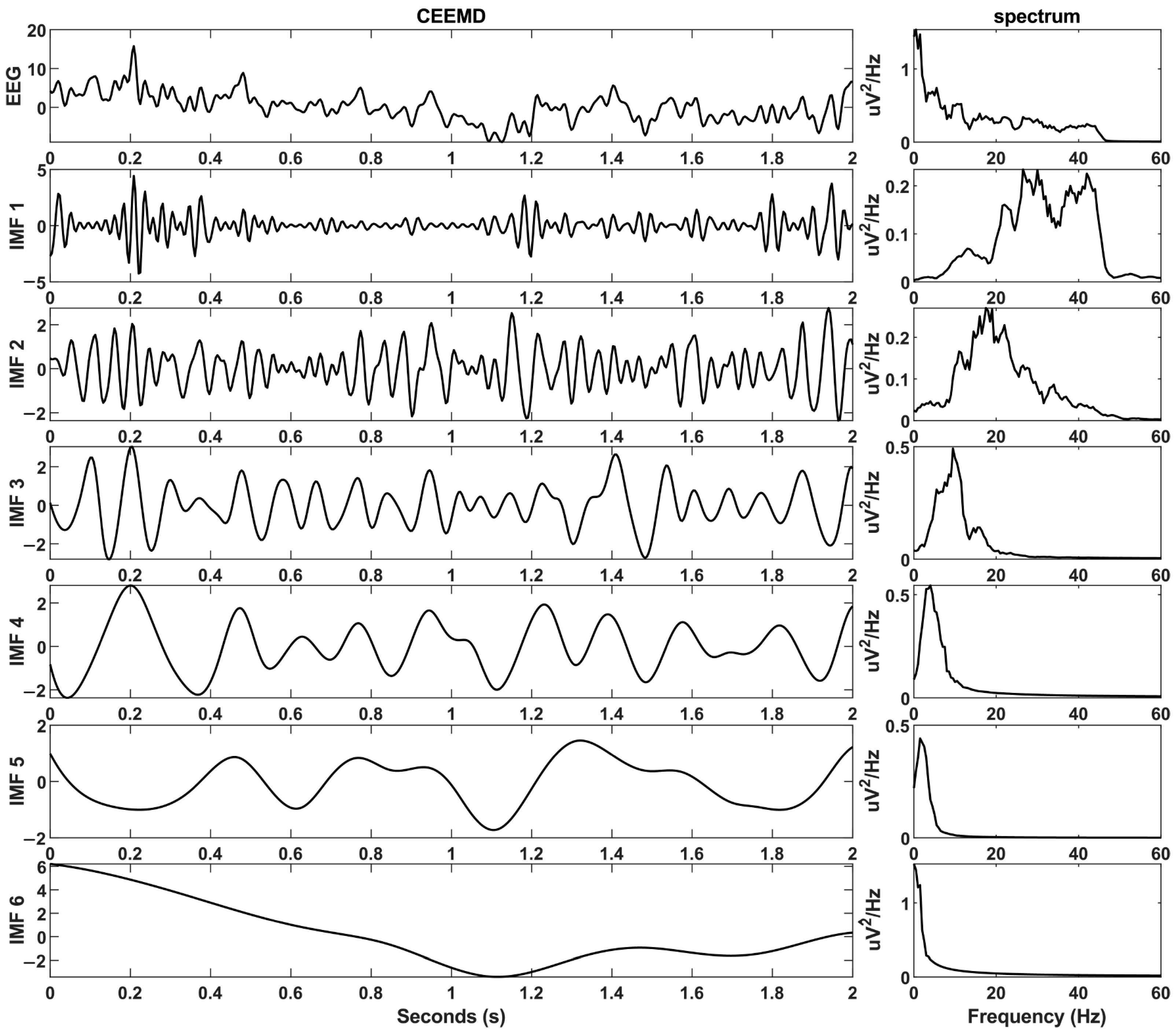
where, represents the original signal, represents the added white noise, is the original data with added white noise, and is the original signal with subtracted white noise. Then, EMD decomposition is performed separately on the signals with added noise and subtracted noise. The resulting two sets of IMFs are averaged to obtain the final IMF component. In this study, the standard deviation of the added white noise is set to 0.1, and the number of iterations is set to 100. The waveform and corresponding spectrum results of the IMF components after CEEMD decomposition are displayed in Figure 3.
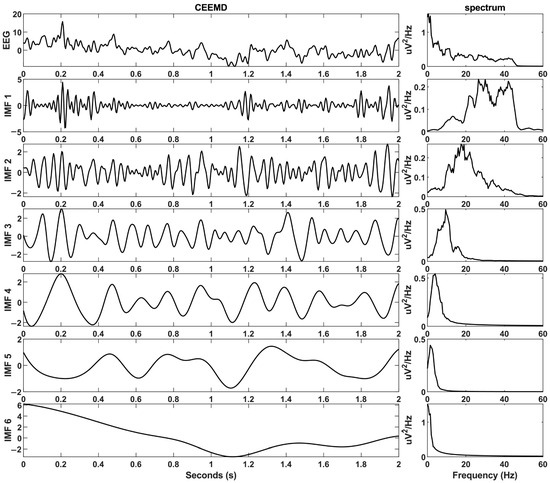
Figure 3.
Waveform and corresponding spectrum graphs of IMF components after CEEMD decomposition.
EEG IMF frames are labeled based on their best matching group template using a “winner takes all” strategy (frames with correlations below 0.5 are not labeled), thereby generating microstate sequences. We first transformed the microstate sequence into a sequence without duplicate labels, for example, transforming a sequence like AACCDDBBBBDDCCA into ACDBDCA. This transformation overlooks the duration of microstates, rooted in the notion that transitions between different brain states measured through EEG microstates convey fundamental information about brain activity [28].
Following that, we utilized the Information-based Similarity (IBS) analysis, a tool rooted in physics and statistical linguistics [32], to detect and quantify the inherent patterns of microstate dynamic transitions. Subsequently, the microstate transition sequence was partitioned into numerous sets of symbolic sequences with length m, called m-bit “word”. Each m-bit word, denoted as , signifies a particular change pattern within cortical EEG IMFs. Through the incremental shifting of a single symbolic point, a sequence of m-bit words is generated, resulting in distinct change patterns for each microstate transition sequence. The frequency of occurrence for each word is tabulated, and all m-bit words are then arranged in descending order based on their respective frequencies. Subsequently, the change pattern sequences (“words”) present in both microstate symbolic sequences are singled out for scrutinizing the information-based similarity between two EEG signals.
and represent the probability and rank of the m-bit word in the symbolic transition sequence , and and represent the probability and rank of the same word in the sequence , respectively. The rank of each m-bit word in the first symbolic sequence is plotted against its rank in the second symbolic sequence, creating a rank order comparison graph. When two symbolic series exhibit a similar arrangement, the scatter points will cluster around the diagonal line. We defined the microstate sequence IBS distance between and as follows:
represent the weight of each m-bit word, computed through Shannon’s entropy and normalized with the normalization factor . Given that not all words carry equal weight in the rank frequency distribution, a weight function must be factored in when computing IBS distance.
Finally, we proposed a method to estimate the degree of complexity or structural richness of spatiotemporal changes in EEG microstate. To this end, we generated alternative symbolic transition sequences by randomly shuffling the microstate transition sequence . Random shuffling of the data produces exactly the same distribution as the original sequence but destroys the sequential ordering. The microstate sequence non-randomness index (MSNRI) was defined as the distance between an EEG spatiotemporal symbolization sequence and its random alternatives,
The closer the distance between the original microstate dynamic sequence and its disordered sequence, the smaller the MSNRI. On the contrary, the farther the distance, the greater the MSNRI. The different IMFs were analyzed separately to obtain the CEEMD-enhanced MSNRI.
2.5. Microstate Sequence Lempel–Ziv Complexity
Calculating the nonlinear complexity of transition sequences using the Lempel–Ziv compression algorithm has been a commonly used method in previous studies [15,16]. The Lempel–Ziv complexity measurement quantifies the number of sub-words that can be found in the entire sequence, assessing the richness of a sequence with a finite number of generating elements. In this study, the calculation of microstate sequence Lempel–Ziv complexity (MSLZC) follows the method proposed by Michael Lassi et al. [9] The MSLZC is computed by measuring the length of the encoded dictionary in the microstate sequence using the zlib Python package (version 1.3.1), after excluding the Huffman coding component. To address variations in sequence length, normalization by the total length of the signal is conducted.
2.6. Statistical Analysis
To explore the potential disparity between the EEG microstate features of health and patients, as well as to prove the probable significance of our proposed methodology, we employ the Mann–Whitney U test on our datasets. The Mann–Whitney U test is a nonparametric method enabling us to ascertain whether the statistics under consideration, in our case, synchrony results, exhibit differing values across two distinct populations. Significant difference was defined as a p-value < 0.05. All statistical analysis methods were performed in MATLAB R2023a.
2.7. Predictive Models for the MMSE Score
We developed a predictive model for MMSE total scores based on resting EEG microstate metrics to explore the possibility of resting-state EEG microstate dynamics characterization to replace or supplement MMSE screening. For feature preprocessing, all predictor variables were normalized from 0 to 1. To fully capture the contribution of microstate dynamics features to predict MMSE scores, we used multiple linear regression to estimate MMSE scores.
In order to demonstrate the effectiveness of the predictive model in real-world scenarios and to ensure the stability and accuracy of the prediction outcomes, we employed the Leave-One-Subject-Out (LOSO) cross validation method. For the LOSO classification experiment, all data from all subjects except one was used for training, with the excluded subject’s data reserved for testing. For a dataset containing N samples, LOSO performed N iterations. Specifically, for the MMSE score prediction model for all subjects (N = 88), the LOSO approach required 88 iterations for validation, with 87 subjects as the training set and the remaining 1 subject as the test set for each iteration. Finally, the coefficient of determination (R-squared), mean squared error (MSE), root mean squared error (RMSE), and mean absolute error (MAE) metrics were utilized to assess the consistency between the actual MMSE scores and the predicted MMSE scores by the prediction model.
3. Results
3.1. Microstate Maps Analysis
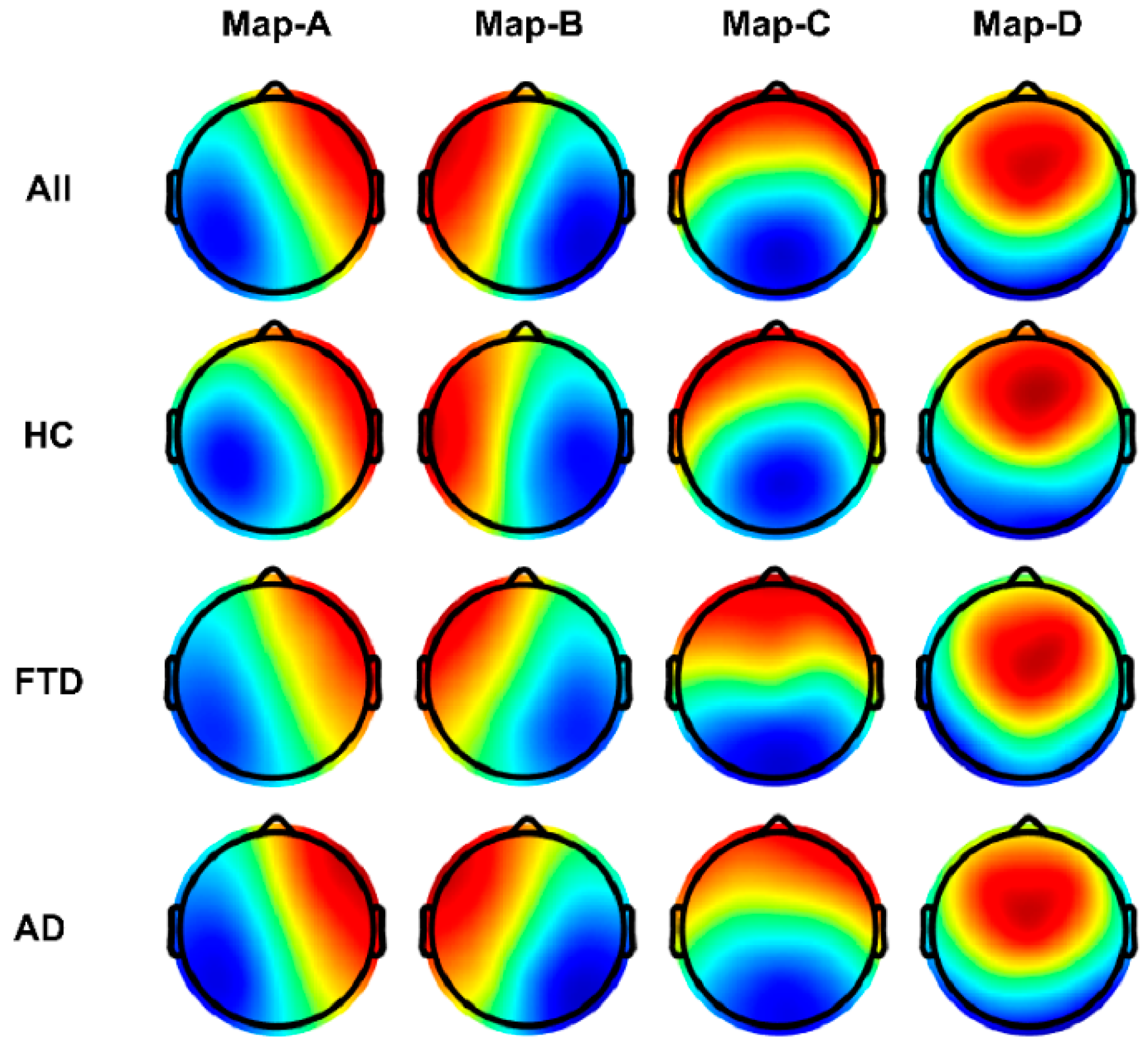
The topology of each of the four microstate categories in the data separately for all subjects, the healthy group, the FTD group, and the AD group is shown in Figure 4. In the HC group, these microstate classes closely correspond to the classical categories A–D, which are linked to wakeful rest and represent electrophysiological manifestations of the auditory (Map-A), visual (Map-B), salience (Map-C), and frontal working memory/attention (Map-D) resting state networks [28].
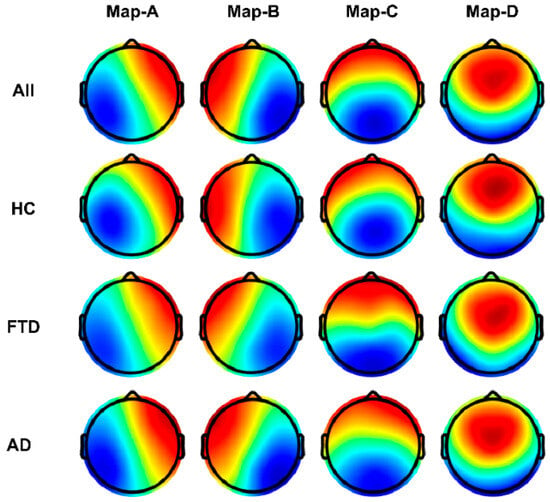
Figure 4.
Microstate template topographies for the four classes. (Top) Globally clustered maps for all cohorts for maps A–D from left to right. (Second) As above, but for the HC group. (Third) For the FTD group. (Bottom) For the AD group. Red and blue indicate positive and negative values, respectively. The polarity is ignored during microstate analysis.
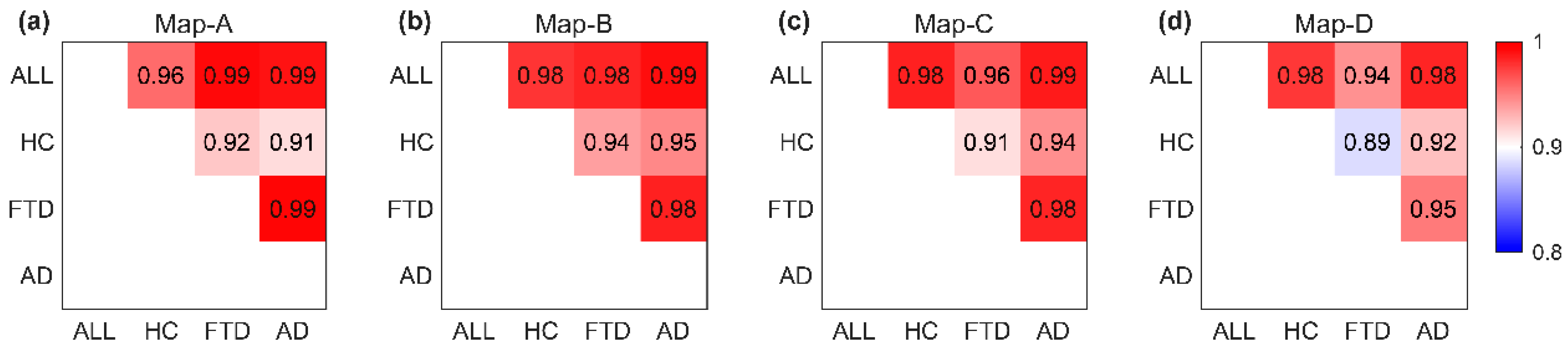
Spatial correlation between microstate topographies measured by the absolute value of spatial cross-correlation coefficients reveals subtle differences between healthy and diseased groups. Figure 5 illustrates the topological correlation comparison matrix between microstate template maps of the same category across different groups (all groups, HC, FTD, AD). The largest difference was observed for Map-D between HC and disease groups (Table 2, average correlation of 0.921 for Map-D).
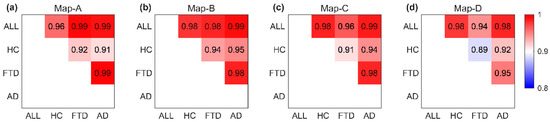
Figure 5.
Spatial correlation between microstate template maps of the same class across groups. The allocation of microstate maps assigned to the same grade (a–d) was compared across all subject groups (All, HC, FTD, AD). Each element of every matrix displays the spatial correlation coefficient between two maps (numeric values within the squares, represented by colors on a color bar).

Table 2.
Spatial correlation between microstate template maps across groups.
Finally, spatial correlation between the microstate templates identified for all people (All) and those identified for each subject was calculated using a segment temporal smoothing constraint of 5 samples (20 ms). EEG frames are labeled based on their best matching group template using a “winner takes all” strategy (frames with correlations below 0.5 are not labeled), thereby generating microstate sequences for each participant, each comprising 45,000 symbols (180 s multiplied by 250 Hz) for further analysis.
3.2. EEG Microstate Sequence Analysis
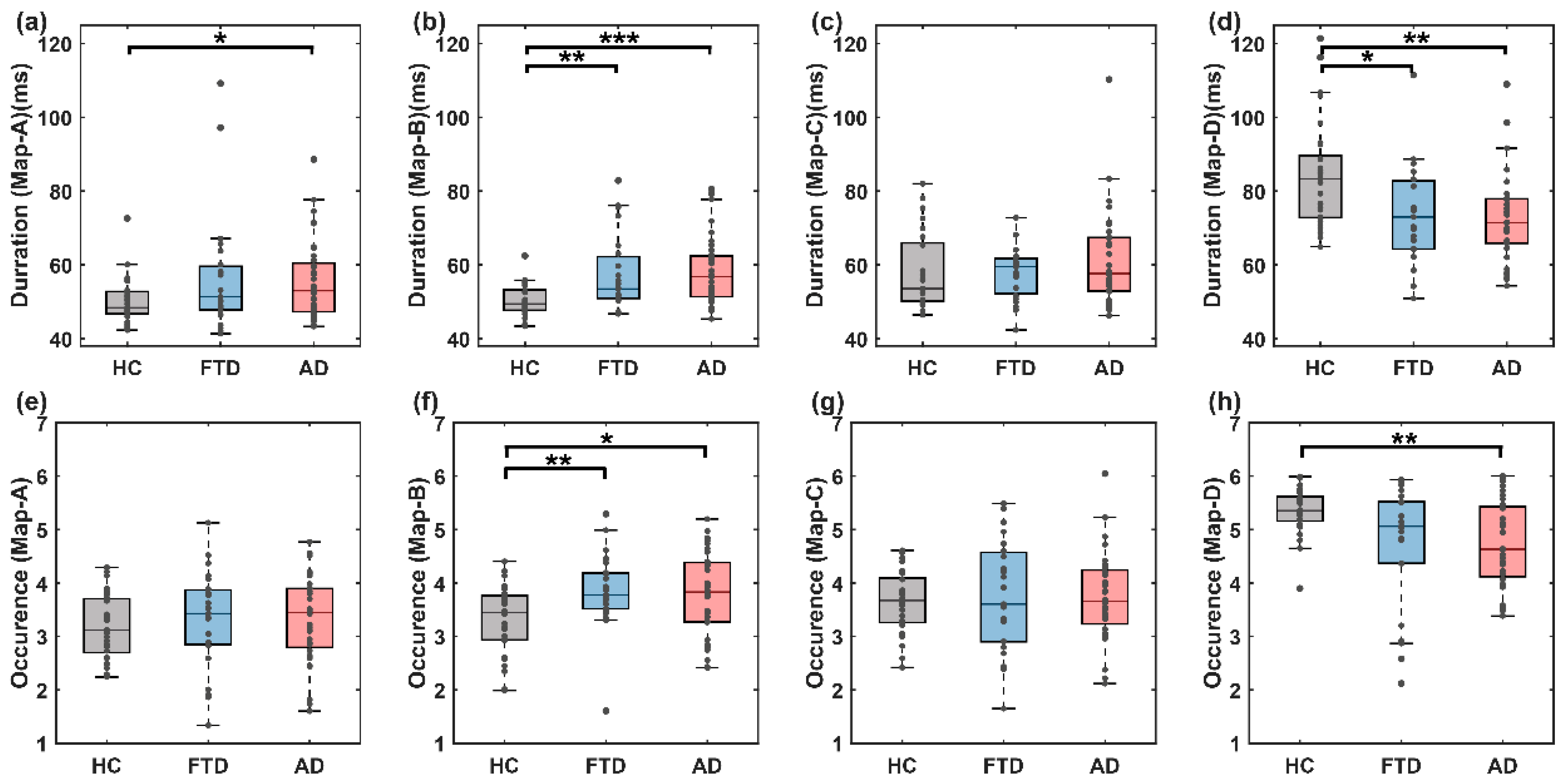
We meticulously examined the temporal dynamics of EEG microstates, specifically focusing on the mean duration (Figure 6a–d) and occurrence (Figure 6e–h) of each microstate among different groups. Our results, detailed in Figure 6, revealed significant disparities in the temporal properties of the microstates when comparing the HC group to both the FTD and AD groups.
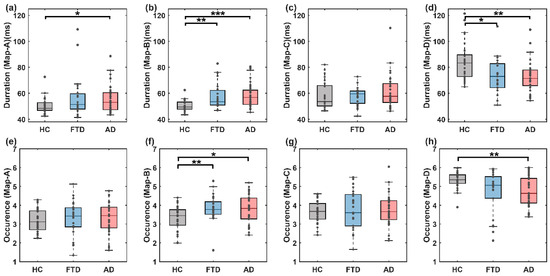
Figure 6.
Classical dynamic characteristics of microstates. (a–d) Duration of microstates in the HC, FTD and AD groups; (e–h) occurrence of microstates in the HC, FTD and AD groups. In all panels, the significance of the corresponding statistical test is represented as follows: * p < 0.05, ** p < 0.01, *** p < 0.001.
For microstate B, the disease groups exhibited a pronounced increase in both the mean duration and occurrence, as illustrated by the following statistics: HC (50.180 ms) versus FTD (57.485 ms) with a non-parametric Mann–Whitney U test (z = −3.058, p = 0.002), and HC versus AD with a duration of 50.180 ms versus 58.223 ms (z = −4.058, p < 0.001). Similarly, the occurrence parameters were significantly higher in the disease groups, with 3.318 for HC compared to 3.862 for FTD (z = −2.763, p = 0.006) and 3.318 for HC compared to 3.777 for AD (z = −2.408, p = 0.016).
Conversely, microstate D showed a significant decrease in both metrics within the disease groups compared to the HC group: HC versus FTD with a duration of 84.160 ms versus 75.397 ms (z = 2.506, p = 0.012), and HC (84.160 ms) versus AD (75.348 ms) (z = 2989, p = 0.003). The occurrence parameters were only significantly reduced in the AD group (HC vs. AD: 5.333 vs. 4.725, z = 3.082, p = 0.002). No significant differences in the time-domain characteristics between the FTD and AD groups were found in any of the microstate categories.
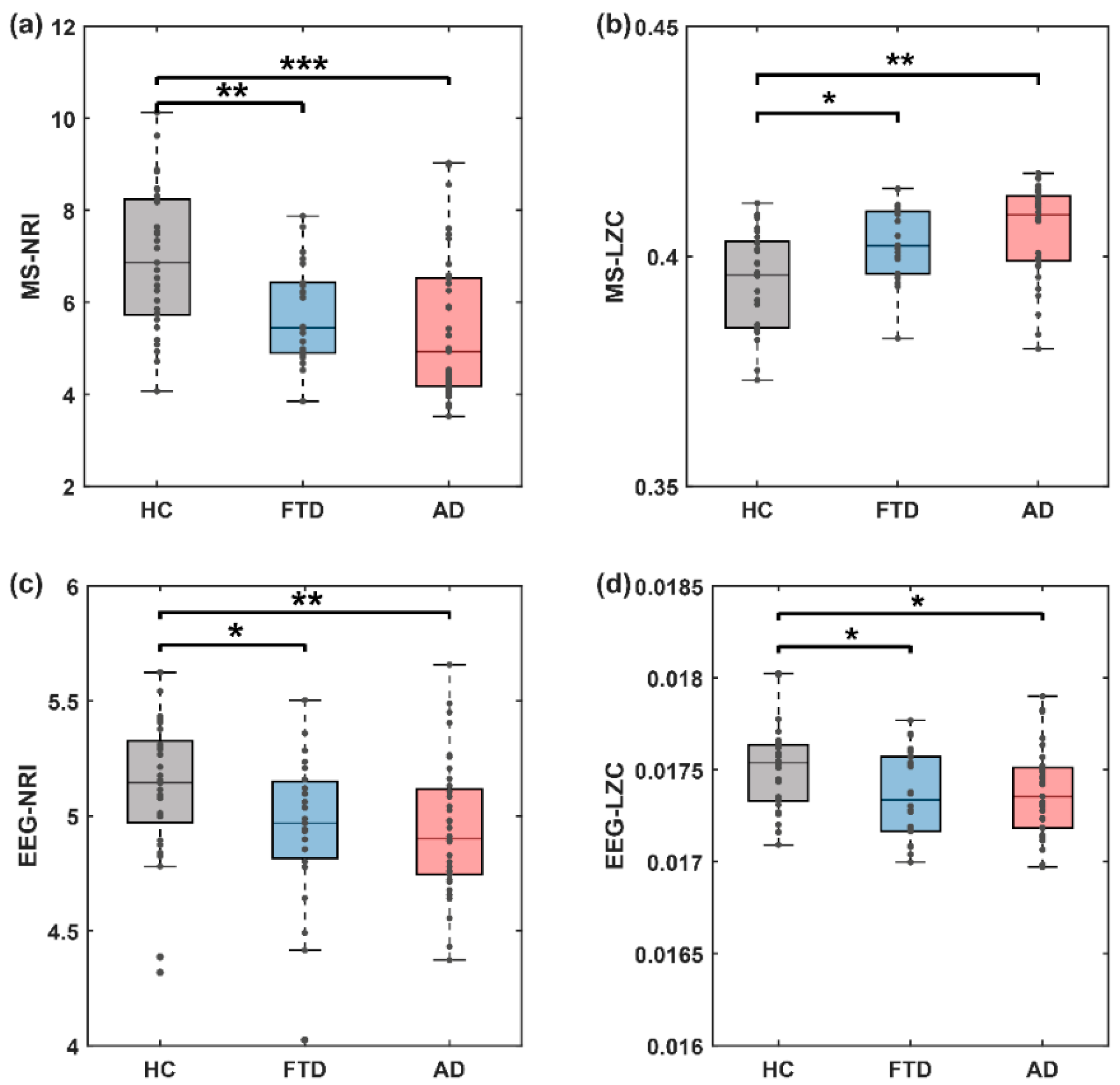
We next explored whether the transition behavior of microstate sequences was altered. Researches have shown that the transitions of microstate sequence are non-smooth, non-Markovian, and scale-free [14]. We therefore calculated the MSNRI and MSLZC of the microstate transitioning sequenc. Compared with the HC group, the MSNRI was markedly reduced in both the FTD group (6.958 vs. 5.756, z = 2.805, p = 0.005, Figure 7a) and AD group (6.958 vs. 5.462, z = 3.426, p = 0.0006, Figure 7a), suggesting the presence of abnormal microstate pattern transitions in the EEGs of patients with cognitive impairment. Furthermore, this effect size was notably larger compared to other used measures of EEG microstate complexity utilized in previous studies on AD or MCI (Mild Cognitive Impairment), such as MSLZC (HC vs. FTD: 0.394 vs. 0.405, z = −2.414, p = 0.016, HC vs. AD: 0.394 vs. 0.403, z = −3.200, p = 0.001, Figure 7a). Meanwhile, we also compared the results of NRI and LZC computed on the EEG time series (EEG-NRI and EEG-LZC), both of which had weaker intergroup differences than those analyzed on the microstate transition series (Figure 7c,d).
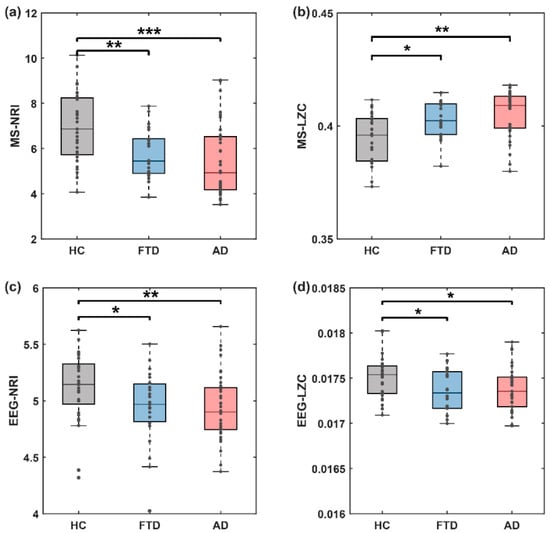
Figure 7.
Microstate sequence dynamic statistics are significantly altered in disease condition (both FTD and AD). (a) Microstate sequence NRI, (b) microstate sequence LZC, (c) EEG time series NRI, (d) EEG time series LZC. Stars denote effect size of Mann–Whitney U test: * p < 0.05, ** p < 0.01, *** p < 0.001. Points in the boxplots show values for each participant. (MSNRI with parameter m = 8).
3.3. CEEMD-Enhanced Non-Randomness Analysis of Microstate Dynamic Patterns
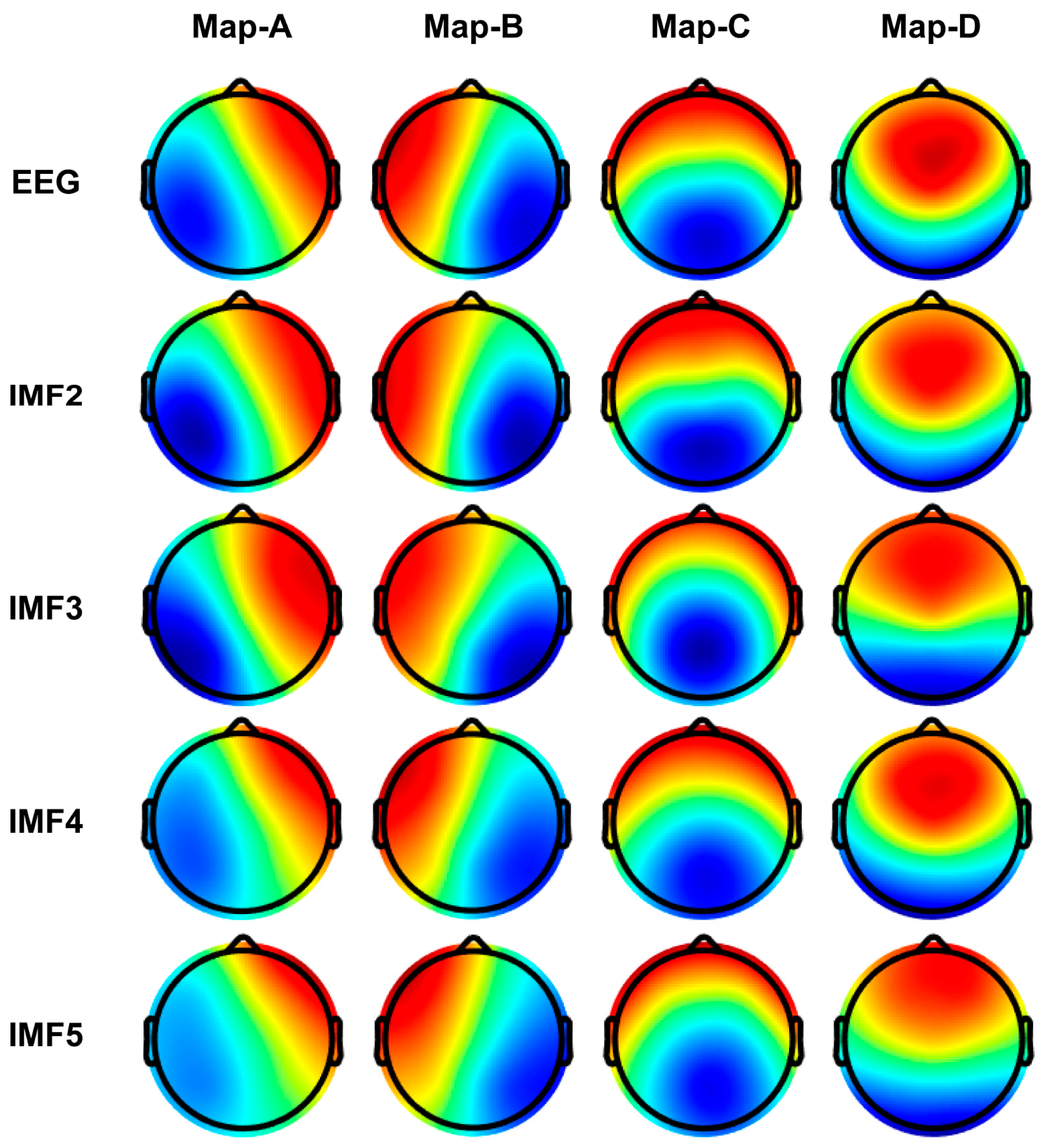
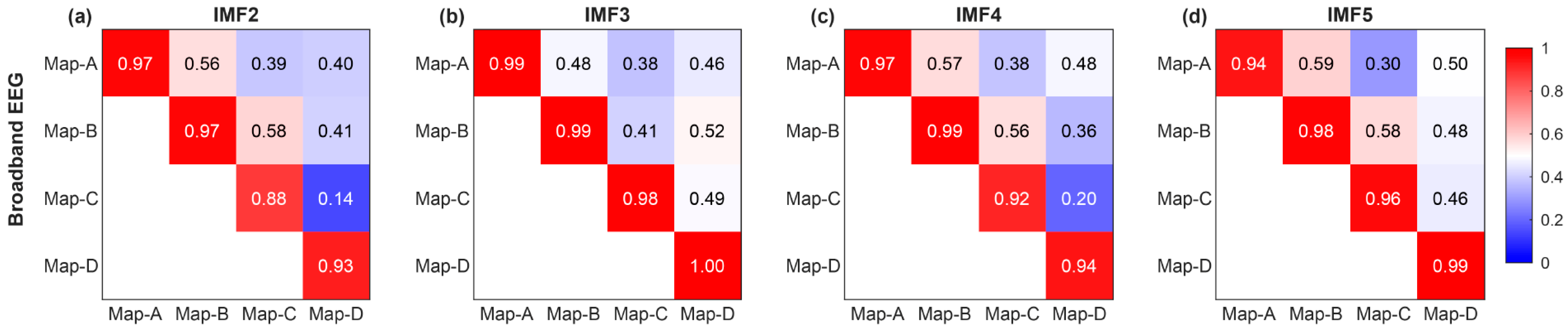
Next, we expanded the EEG into various IMFs using CEEMD decomposition, aiming to gain insights into the microstate spatio-temporal dynamics of different IMFs. Figure 8 illustrates the topographic results of microstate segmentations in the different IMF bands (IMF2, IMF3, IMF4, and IMF5). We found that these microstate topologies on different IMFs are very similar to those on the full-band EEG, all conforming to previously specified prototypes in the literature. As shown in Figure 9, all spatial correlations are very high when comparing the topology of the microstate templates between the broadband EEG and each IMFs (i.e., diagonal entries in the correlation matrix), which provides a rationale for reconstructing the sequence of microstate transitions within the different IMFs using their respective microstate templates.
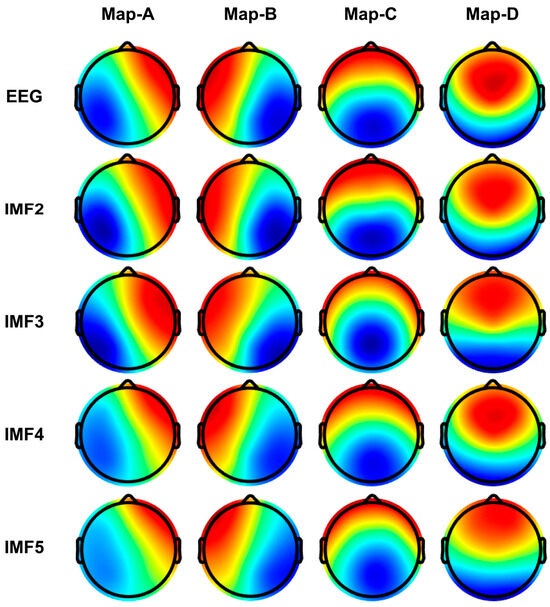
Figure 8.
Microstate template topographies for each IMF after CEEMD decomposition. Red and blue indicate positive and negative values, respectively.
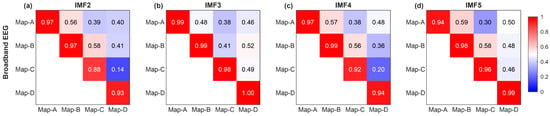
Figure 9.
Spatial correlation between microstate topographies across IMFs. Spatial correlation of microstate template topographies (a) between the broadband EEG and IMF 2, (b) between the broadband EEG and IMF 3, (c) between the broadband EEG and IMF 4, (d) between the broadband EEG and IMF 5. Numeric values within the squares, represented by colors on a color bar.
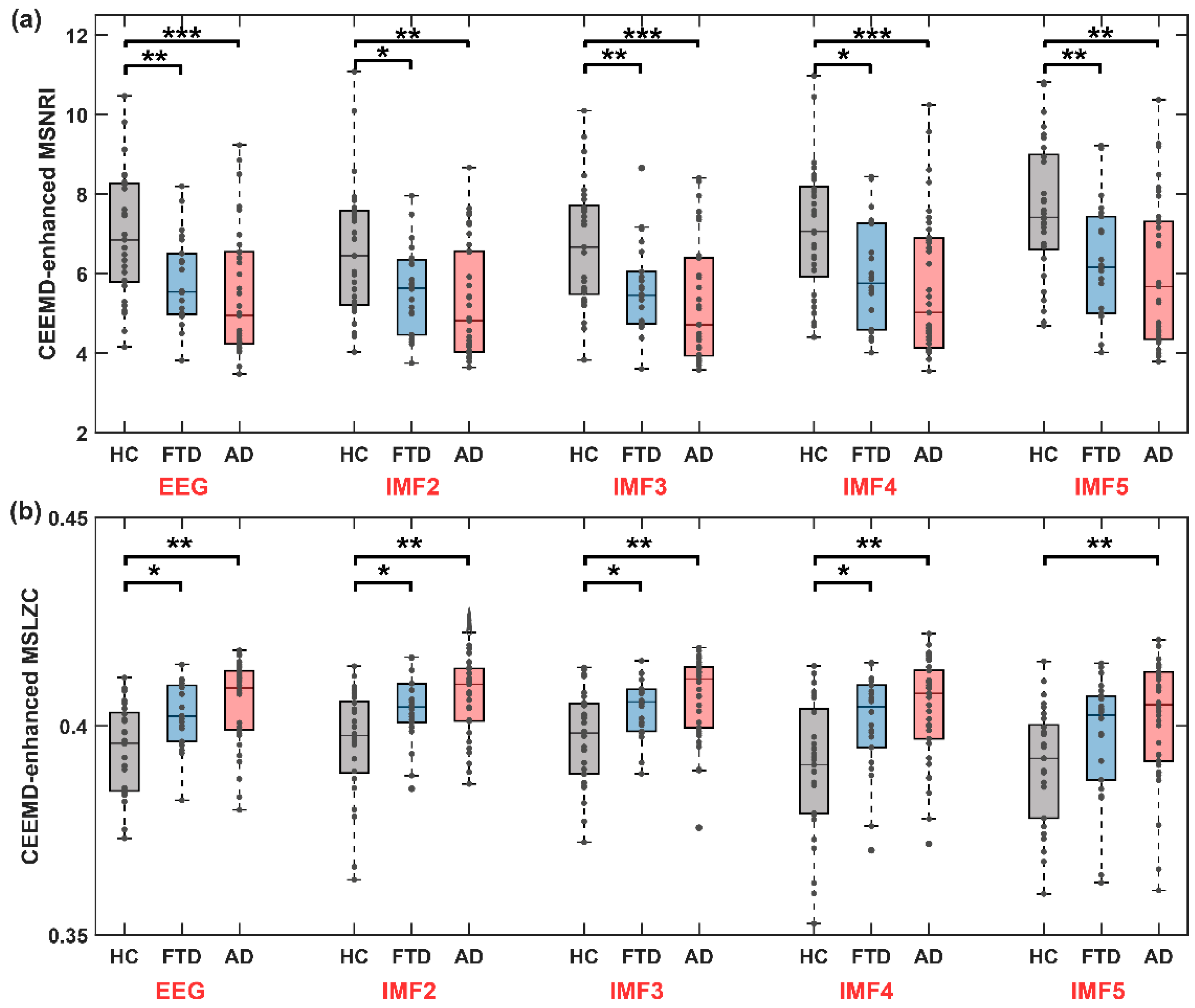
Here, we obtained the sequence of microstate transitions specific unique to each IMF through a conventional microstate analysis conducted on individual IMF components. Subsequently, we obtained MSNRI and MSLZC for each distinct IMF to discern variations in microstate dynamic patterns among the HC, FTD, and AD groups across different IMF components. The analysis results of the MSNRI and MSLZC indices across IMF components for the HC, FTD, and AD groups are illustrated in Figure 10. The detailed results of MSNRI and MSLZC for the HC, FTD, and AD groups across different IMF scales are shown in Table 3.
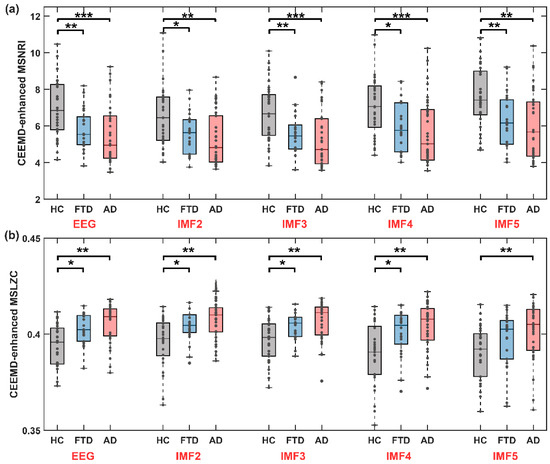
Figure 10.
Microstate sequence dynamic statistics are significantly altered in disease condition (both FTD and AD) in all IMFs after CEEMD decomposition. (a) The CEEMD-enhanced MSNRI among the HC, FTD, and AD groups (MSNRI with parameter m = 8). (b) The CEEMD-enhanced MSLZC among the HC, FTD, and AD groups. Stars denote effect size of Mann–Whitney U test: * p < 0.05, ** p < 0.01, *** p < 0.001.

Table 3.
The detailed results of MS-NRI and MS-LZC for the HC, FTD, and AD groups across different IMFs.
Across all IMF components, the HC group consistently exhibited the highest MS-NRI, followed by the FTD group, and the lowest was observed in the AD group. Notably, in all the IMF components included in the analysis, the MS-NRI index for the AD group was significantly lower than that for the HC group (IMF2: z-value = 3.196, p = 0.001, IMF3: z-value = 3.507, p < 0.001, IMF4: z-value = 3.304, p = 0.001, IMF5: z-value = 3.264, p = 0.001). For the FTD group, there was a significant decrease in MS-NRI compared to the HC group across the scales of IMF2 to IMF5.
It is noteworthy that the trend in MSLZC contrasted with that of MSNRI. Across all IMF components, the AD group consistently demonstrated the highest MSLZC, followed by the FTD group, while the HC group consistently displayed the lowest. Although MSLZC across different IMF scales also exhibited significant intergroup differences, the MSNRI index proposed in this study statistically demonstrated more significant distinctions in characterizing differences between healthy individuals and those with cognitive impairment disorders. The findings imply that conducting cross-frequency microstate dynamic analysis could provide further valuable insights into diagnosing and predicting cognitive impairment disorders.
3.4. Microstate Sequence Non-Randomness Has Predictive Power for MMSE Scores
To better understand how well the CEEMD-enhanced non-randomness analysis method can aiding diagnosis and cognitive prediction of Alzheimer’s disease, we developed a predictive model for MMSE scores based on EEG microstate metrics to explore the possibility of resting-state EEG to replace or supplement MMSE screening. The baseline method incorporates temporal statistical features of microstates extracted from resting-state EEG, along with MSLZC and MSNRI (with parameter m from 4 to 8). By contrast, the proposed method in this paper includes CEEMD-enhanced MSLZC and MSNRI (with parameter m from 4 to 8), along with the integration of microstate temporal statistical indicators.
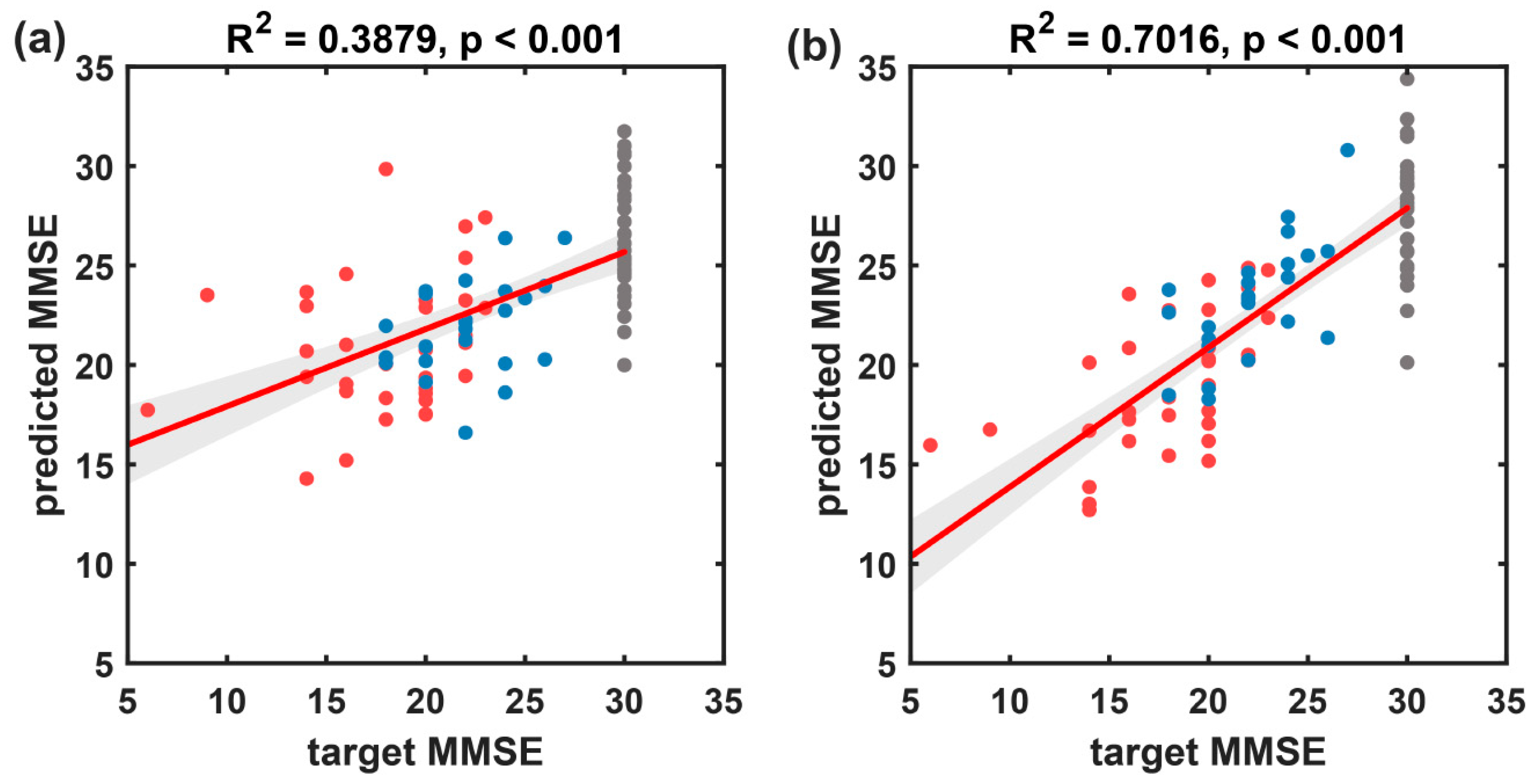
The results of the multivariate linear regression prediction model based on MMSE scores for all subject samples are shown in Figure 11 and Table 4. Compared to the baseline method (R2 = 0.388, rMSE of 4.784, and MAE of 3.511), the proposed method in this paper achieved superior predictive performance (R2 = 0.702, rMSE of 3.340, and MAE of 2.555).
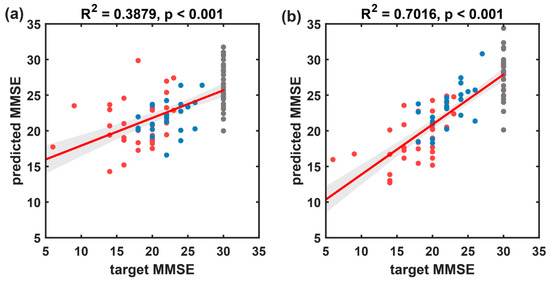
Figure 11.
The results of the multivariate linear regression model predicting MMSE scores using resting-state EEG indices for all participants, with target MMSE scores on the x-axis and predicted MMSE scores on the y-axis. (a) Scatter plot and fitted line of the baseline model’s predicted values and true values based on microstate temporal metrics. (b) Scatter plot and fitted line of the prediction model’s predicted values and true values based on the microstate sequence dynamic metrics proposed in this paper. Black circles represent the HC group, blue circles represent the FTD group, and red circles represent the AD group. The solid red line represents the scatter fit curve, and the shaded area indicates the 95% confidence interval.

Table 4.
Evaluation metrics for the multivariate linear regression model predicting MMSE scores using resting-state EEG indices for all participants.
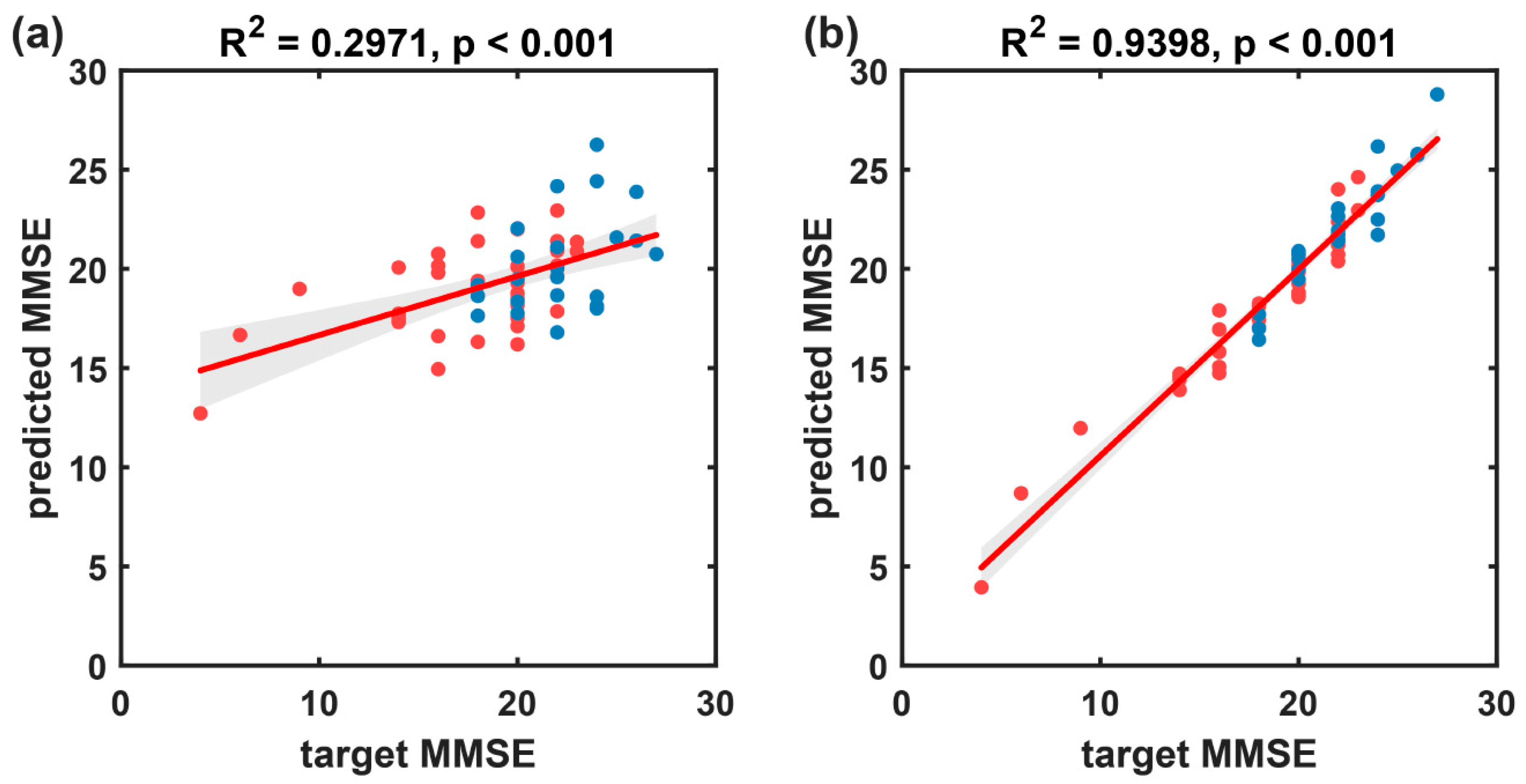
From the scatter plot of predicted MMSE scores against target MMSE scores (Figure 11), it is evident that the MMSE scores for the HC group are all 30, forming a cluster on the plot, which might have influenced the overall performance of the predictive model. Therefore, this subsection further establishes a multivariate linear regression prediction model for MMSE scores based on participants with cognitive impairment disorders. The results are illustrated in Figure 12 and summarized in Table 5. On the disease group, the proposed method in this paper exhibited significantly improved predictive performance (R2 = 0.940, rMSE of 1.077, and MAE of 0.807) compared to the baseline method (R2 = 0.297, rMSE of 3.680, and MAE of 2.877).
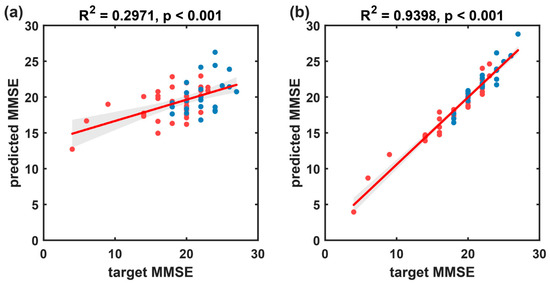
Figure 12.
The prediction results of the multivariate linear regression model, which predicts MMSE scores based on resting-state EEG indices, for the cognitive impairment group (FTD and AD). (a) Scatter plot and fitted line of the baseline model’s predicted values and true values based on microstate temporal metrics. (b) Scatter plot and fitted line of the prediction model’s predicted values and true values based on the microstate sequence dynamic metrics proposed in this paper. Blue circles represent the FTD group, and red circles represent the AD group. The solid red line represents the scatter fit curve, and the shaded area indicates the 95% confidence interval.

Table 5.
Evaluation metrics for the multivariate linear regression model predicting MMSE scores based on resting-state EEG indices for the cognitive impairment group.
The results of the multivariate linear regression prediction model demonstrate that utilizing resting-state EEG microstate indicators can achieve high accuracy in the cross-subject prediction of MMSE scores, confirming the potential of EEG biomarkers as substitutes or complements to MMSE. Additionally, the exploration of non-randomness and complexity indices of microstate spatio-temporal patterns enhanced by CEEMD holds significant importance in assessing the severity of cognitive impairment.
4. Discussion
In this study, our core hypothesis is that the disruption in the equilibrium between spatiotemporal pattern transitions in brain microstates constitutes a crucial characteristic of the dementia process. Furthermore, we postulate that the patterns of transitions between EEG microstate may be biomarkers to predict cognitive abilities. To test this hypothesis, we developed a microstate sequence non-randomness index and gauged its efficacy in distinguishing HC and dementia diseases (FTD and AD). Our research findings indicate that this method exhibits highly significant differences in distinguishing between healthy subjects and patients with dementia, providing compelling evidence that spatiotemporal patterns of microstates in resting-state EEG signals can serve as valuable biomarkers for dementia.
This investigation uncovers another significant revelation. Cross-frequency microstate dynamic analysis, CEEMD-enhanced MSNRI, expands the differences between healthy and dementia states, significantly enhancing cognitive function prediction performance, making it a reasonable complement to broadband microstate analysis. These findings carry practical significance, suggesting that EEG can offer an alternative avenue for assessing cognitive diseases or cognitive abilities.
In this work, we present a novel method of non-randomness to microstate transition sequences since microstate transitioning is non-linear, non-stationary, and non-Markovian [14]. As anticipated [9], maintaining the balance and stability of brain spatiotemporal pattern transitions plays a crucial role in human cognitive health. We found significant differences in the non-randomness of spatiotemporal patterns of microstates between healthy individuals and patients with cognitive impairment disorders. The MSNRI of patients with dementia decreases, indicating a shift towards more randomness in the spatiotemporal patterns of their brain’s complex system (Figure 7a, HC vs. FTD vs. AD: 6.958 vs. 5.756 vs. 5.462). Interestingly, MSNRI will decline more in AD than in FTD. In addition, the CEEMD-enhanced MSNRI produced consistent results at every IMF scale (Figure 10a, all IMF scales). In a healthy state, the spatiotemporal patterns of brain microstates exhibit a more complex macroscopic structure and demonstrate the lowest level of randomness. Conversely, dementia diseases lead to increased randomness in brain spatiotemporal patterns. This understanding contributes to a deeper comprehension of the neural mechanisms underlying cognitive impairment disorders and provides crucial clues for related research and clinical diagnosis.
MSLZC is a previously reported measurement for studying microstate transitions [9,15,16]. In this study, we also observed differences in MSLZC between the healthy group and the group with the disease, but the significance of the differences was weaker compared to MSNRI. We observed a significant increase in MSLZC tendency in the AD group (Figure 7 and Figure 10). It is worth noting that, in comparison with another MSLZC-based analysis of AD patients, our results for the LZC analysis of the EEG time series were consistent, but the results for the MSLZC analysis of the microstate series were reversed (MSLZC was reduced in AD, p = 0.0023) [15]. Possible reasons for this discrepancy include (1) differences in the length of data included in the analysis (we used 180 s, while they used 20 s), (2) differences in microstate segmentation methods, and (3) variations in the severity of AD; in their study, the average MMSE score for AD was 23, whereas ours was 17.75. The U-shaped relationship between complexity and disease severity might lead to such differences in results [33]. Furthermore, the biological significance of LZC may differ from that indicated by other microstate indicators [34]. For instance, in other studies, MSLZC was found to be higher in patients with depression than in controls [35], and LZC in patients with thalamic ischemic stroke has been found to be higher compared to the controls [36].
Meanwhile, the CEEMD-enhanced MSNRI algorithm is more effective compared to broad frequency domain scale analysis (Figure 10 and Figure 12 and Table 3). In this study, CEEMD was used to refine the frequency domain of EEG signals into multiple scales. Compared to Fourier transform, CEEMD can analyze nonlinear and non-stationary signals. Unlike wavelet transform, CEEMD does not select basis functions. It adaptively extracts the intrinsic mode functions of the signal while simultaneously separating trend components and noise, which helps capture patterns in the specific frequency scales of EEG signals from patients with cognitive impairment disorders.
After the introduction of the CEEMD-based MSNRI and MSLZC indicators, compared to traditional microstate temporal features (Figure 12a, R2 = 0.297, rMSE of 3.680, and MAE of 2.877), the method integrating CEEMD-enhanced MSNRI and MSLZC shows a significant improvement in the accuracy of predicting MMSE scores (Figure 12b, R2 = 0.940, rMSE of 1.077, and MAE of 0.807). As shown in Table 6, this prediction performance exceeds the results of previous studies of MMSE prediction using microstate or other EEG characterization methods [20,37,38,39,40,41], while also confirming the potential of resting-state EEG biomarkers as substitutes or supplements for MMSE.

Table 6.
Comparison with previous studies on the EEG-based prediction of MMSE scores.
This research also has several limitations. One notable limitation concerns the necessary length of the data. Since we were analyzing pattern information within EEG microstate transition sequences, the microstate sequences were reduced to transition sequences during the analysis process. As a result, a relatively longer data length is required to obtain sufficient information. In this article, we used a data length of 180 s. This is in contrast to other methods that may only require several seconds or tens of seconds of EEG data to complete feature extraction. Secondly, we employed a relatively small clinical dataset (29 healthy individuals, 59 patients with dementia) for screening diseases. While this validated the effectiveness of the algorithm to some extent, further validation of its robustness in cognitive disease screening applications requires larger datasets, which will be pursued in future work. Additionally, in predicting cognitive test scores, we only utilized EEG markers to predict MMSE scores. While MMSE exhibits acceptable accuracy for dementia screening, its specificity and sensitivity are lower for milder cognitive impairments such as mild cognitive impairment because some individuals with early cognitive decline can still score within the normal range, meaning that it may not be sensitive enough to detect subtle cognitive changes that occur in the early stages of MCI. Therefore, future work needs to validate the ability of EEG markers to predict scores from other computerized tests assessing early cognitive decline.
5. Conclusions
The proposed novel CEEMD-based MSNRI approach employs topographic space compression, physics, and statistical linguistic tools to detect and quantify certain fundamental patterns embedded in EEG microstates beyond frequency bands. By leveraging EEG datasets from HC, FTD, and AD participants, we validated our hypotheses. Our findings revealed that healthy individuals exhibit more complex macroscopic structures and non-random spatiotemporal patterns of microstates, whereas dementia disorders lead to the generation of more random spatiotemporal patterns of microstates.
Supplementary Materials
The following supporting information can be downloaded at: https://www.mdpi.com/article/10.3390/brainsci14050487/s1, Table S1: Detailed MMSE scores for each subject.
Author Contributions
Conceptualization, W.W. and X.C.; methodology, W.W. and X.C.; software, W.W.; validation, X.C.; formal analysis, W.W.; investigation, W.W.; resources, Z.G. and X.C.; data curation, W.W.; writing—original draft preparation, W.W.; writing—review and editing, X.C. and C.-K.P.; visualization, W.W.; supervision, Z.G., X.C. and C.-K.P.; project administration, X.C.; funding acquisition, X.C. and Z.G. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Natural Science Foundation of China, grant number 82274631 and 62077013.
Institutional Review Board Statement
The data in the experiment were obtained from publicly available databases whose study was conducted in accordance with the Declaration of Helsinki and approved by the Scientific and Ethics Committee of AHEPA University Hospital, Aristotle University of Thessaloniki, under protocol number 142/12 April 2023 [2].
Informed Consent Statement
Informed consent was obtained from all subjects involved in this study [2].
Data Availability Statement
These data were derived from the following resources available in the public domain: https://openneuro.org/datasets/ds004504, accessed on 17 October 2023 [2].
Acknowledgments
We extend our gratitude to the researchers who provided the open EEG dataset for their valuable contribution.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Nichols, E.; Steinmetz, J.D.; Vollset, S.E.; Fukutaki, K.; Chalek, J.; Abd-Allah, F.; Abdoli, A.; Abualhasan, A.; Abu-Gharbieh, E.; Akram, T.T.; et al. Estimation of the Global Prevalence of Dementia in 2019 and Forecasted Prevalence in 2050: An Analysis for the Global Burden of Disease Study 2019. Lancet Public Health 2022, 7, e105–e125. [Google Scholar] [CrossRef]
- Miltiadous, A.; Tzimourta, K.D.; Afrantou, T.; Ioannidis, P.; Grigoriadis, N.; Tsalikakis, D.G.; Angelidis, P.; Tsipouras, M.G.; Glavas, E.; Giannakeas, N.; et al. A Dataset of Scalp EEG Recordings of Alzheimer’s Disease, Frontotemporal Dementia and Healthy Subjects from Routine EEG. Data 2023, 8, 95. [Google Scholar] [CrossRef]
- Miltiadous, A.; Tzimourta, K.D.; Giannakeas, N.; Tsipouras, M.G.; Afrantou, T.; Ioannidis, P.; Tzallas, A.T. Alzheimer’s Disease and Frontotemporal Dementia: A Robust Classification Method of Eeg Signals and a Comparison of Validation Methods. Diagnostics 2021, 11, 1437. [Google Scholar] [CrossRef]
- Zheng, X.; Wang, B.; Liu, H.; Wu, W.; Sun, J.; Fang, W.; Jiang, R.; Hu, Y.; Jin, C.; Wei, X.; et al. Diagnosis of Alzheimer’s Disease via Resting-State EEG: Integration of Spectrum, Complexity, and Synchronization Signal Features. Front. Aging Neurosci. 2023, 15, 1288295. [Google Scholar] [CrossRef]
- Cassani, R.; Estarellas, M.; San-Martin, R.; Fraga, F.J.; Falk, T.H. Systematic Review on Resting-State EEG for Alzheimer’s Disease Diagnosis and Progression Assessment. Dis. Markers 2018, 2018, 5174815. [Google Scholar] [CrossRef]
- Wang, R.; Wang, J.; Yu, H.; Wei, X.; Yang, C.; Deng, B. Power Spectral Density and Coherence Analysis of Alzheimer’s EEG. Cogn. Neurodyn. 2015, 9, 291–304. [Google Scholar] [CrossRef]
- Şeker, M.; Özbek, Y.; Yener, G.; Özerdem, M.S. Complexity of EEG Dynamics for Early Diagnosis of Alzheimer’s Disease Using Permutation Entropy Neuromarker. Comput. Methods Programs Biomed. 2021, 206, 106116. [Google Scholar] [CrossRef]
- Briels, C.T.; Briels, C.T.; Schoonhoven, D.N.; Schoonhoven, D.N.; Stam, C.J.; De Waal, H.; Scheltens, P.; Gouw, A.A. Reproducibility of EEG Functional Connectivity in Alzheimer’s Disease. Alzheimer’s Res. Ther. 2020, 12, 68. [Google Scholar] [CrossRef]
- Lassi, M.; Fabbiani, C.; Mazzeo, S.; Burali, R.; Vergani, A.A.; Giacomucci, G.; Moschini, V.; Morinelli, C.; Emiliani, F.; Scarpino, M.; et al. Degradation of EEG Microstates Patterns in Subjective Cognitive Decline and Mild Cognitive Impairment: Early Biomarkers along the Alzheimer’s Disease Continuum? NeuroImage Clin. 2023, 38, 103407. [Google Scholar] [CrossRef] [PubMed]
- Smailovic, U.; Koenig, T.; Laukka, E.J.; Kalpouzos, G.; Andersson, T.; Winblad, B.; Jelic, V. EEG Time Signature in Alzheimer’s Disease: Functional Brain Networks Falling Apart. NeuroImage Clin. 2019, 24, 102046. [Google Scholar] [CrossRef] [PubMed]
- Lei, L.; Liu, Z.; Zhang, Y.; Guo, M.; Liu, P.; Hu, X.; Yang, C.; Zhang, A.; Sun, N.; Wang, Y.; et al. EEG Microstates as Markers of Major Depressive Disorder and Predictors of Response to SSRIs Therapy. Prog. Neuro-Psychopharmacol. Biol. Psychiatry 2022, 116, 110514. [Google Scholar] [CrossRef]
- Kim, K.; Duc, N.T.; Choi, M.; Lee, B. EEG Microstate Features According to Performance on a Mental Arithmetic Task. Sci. Rep. 2021, 11, 343. [Google Scholar] [CrossRef]
- Wiemers, M.C.; Laufs, H.; von Wegner, F. Frequency Analysis of EEG Microstate Sequences in Wakefulness and NREM Sleep. Brain Topogr. 2023, 37, 312–328. [Google Scholar] [CrossRef]
- Van De Ville, D.; Britz, J.; Michel, C.M. EEG Microstate Sequences in Healthy Humans at Rest Reveal Scale-Free Dynamics. Proc. Natl. Acad. Sci. USA 2010, 107, 18179–18184. [Google Scholar] [CrossRef]
- Tait, L.; Tamagnini, F.; Stothart, G.; Barvas, E.; Monaldini, C.; Frusciante, R.; Volpini, M.; Guttmann, S.; Coulthard, E.; Brown, J.T.; et al. EEG Microstate Complexity for Aiding Early Diagnosis of Alzheimer’s Disease. Sci. Rep. 2020, 10, 17627. [Google Scholar] [CrossRef]
- von Wegner, F.; Wiemers, M.; Hermann, G.; Tödt, I.; Tagliazucchi, E.; Laufs, H. Complexity Measures for EEG Microstate Sequences: Concepts and Algorithms. Brain Topogr. 2024, 37, 296–311. [Google Scholar] [CrossRef]
- Yang, A.C.C.; Goldberger, A.L.; Peng, C.K. Genomic Classification Using an Information-Based Similarity Index: Application to the SARS Coronavirus. J. Comput. Biol. 2005, 12, 1103–1116. [Google Scholar] [CrossRef]
- Férat, V.; Seeber, M.; Michel, C.M.; Ros, T. Beyond Broadband: Towards a Spectral Decomposition of EEG Microstates. bioRxiv 2020. [Google Scholar] [CrossRef]
- Houmani, N.; Vialatte, F.; Gallego-Jutglà, E.; Dreyfus, G.; Nguyen-Michel, V.H.; Mariani, J.; Kinugawa, K. Diagnosis of Alzheimer’s Disease with Electroencephalography in a Differential Framework. PLoS ONE 2018, 13, e0193607. [Google Scholar] [CrossRef] [PubMed]
- Jiao, B.; Li, R.; Zhou, H.; Qing, K.; Liu, H.; Pan, H.; Lei, Y.; Fu, W.; Wang, X.; Xiao, X.; et al. Neural Biomarker Diagnosis and Prediction to Mild Cognitive Impairment and Alzheimer’s Disease Using EEG Technology. Alzheimer’s Res. Ther. 2023, 15, 32. [Google Scholar] [CrossRef] [PubMed]
- Vicchietti, M.L.; Ramos, F.M.; Betting, L.E.; Campanharo, A.S.L.O. Computational Methods of EEG Signals Analysis for Alzheimer’s Disease Classification. Sci. Rep. 2023, 13, 8184. [Google Scholar] [CrossRef]
- Yeh, J.R.; Shieh, J.S.; Huang, N.E. Complementary Ensemble Empirical Mode Decomposition: A Novel Noise Enhanced Data Analysis Method. Adv. Adapt. Data Anal. 2010, 2, 135–156. [Google Scholar] [CrossRef]
- Bell, C.C. DSM-IV: Diagnostic and Statistical Manual of Mental Disorders. JAMA J. Am. Med. Assoc. 1994, 272, 828–829. [Google Scholar] [CrossRef]
- McKhann, G.; Drachman, D.; Folstein, M.; Katzman, R.; Price, D.; Stadlan, E.M. Clinical Diagnosis of Alzheimer’s Disease: Report of the NINCDS-ADRDA Work Group⋆ under the Auspices of Department of Health and Human Services Task Force on Alzheimer’s Disease. Neurology 1984, 34, 939. [Google Scholar] [CrossRef] [PubMed]
- Oostenveld, R.; Fries, P.; Maris, E.; Schoffelen, J.M. FieldTrip: Open Source Software for Advanced Analysis of MEG, EEG, and Invasive Electrophysiological Data. Comput. Intell. Neurosci. 2011, 2011, 156869. [Google Scholar] [CrossRef]
- Vigário, R.N. Extraction of Ocular Artefacts from EEG Using Independent Component Analysis. Electroencephalogr. Clin. Neurophysiol. 1997, 103, 395–404. [Google Scholar] [CrossRef]
- Poulsen, A.T.; Pedroni, A.; Langer, N.; Hansen, L.K. Microstate EEGlab Toolbox: An Introductory Guide. bioRxiv 2018. [Google Scholar] [CrossRef]
- Michel, C.M.; Koenig, T. EEG Microstates as a Tool for Studying the Temporal Dynamics of Whole-Brain Neuronal Networks: A Review. Neuroimage 2018, 180, 577–593. [Google Scholar] [CrossRef] [PubMed]
- Khanna, A.; Pascual-Leone, A.; Michel, C.M.; Farzan, F. Microstates in Resting-State EEG: Current Status and Future Directions. Neurosci. Biobehav. Rev. 2015, 49, 105–113. [Google Scholar] [CrossRef] [PubMed]
- Britz, J.; Van De Ville, D.; Michel, C.M. BOLD Correlates of EEG Topography Reveal Rapid Resting-State Network Dynamics. Neuroimage 2010, 52, 1162–1170. [Google Scholar] [CrossRef]
- Musaeus, C.S.; Nielsen, M.S.; Høgh, P. Microstates as Disease and Progression Markers in Patients with Mild Cognitive Impairment. Front. Neurosci. 2019, 13, 563. [Google Scholar] [CrossRef]
- Yang, A.C.C.; Hseu, S.S.; Yien, H.W.; Goldberger, A.L.; Peng, C.K. Linguistic Analysis of the Human Heartbeat Using Frequency and Rank Order Statistics. Phys. Rev. Lett. 2003, 90, 108103. [Google Scholar] [CrossRef]
- Gaubert, S.; Raimondo, F.; Houot, M.; Corsi, M.C.; Naccache, L.; Sitt, J.D.; Hermann, B.; Oudiette, D.; Gagliardi, G.; Habert, M.O.; et al. EEG Evidence of Compensatory Mechanisms in Preclinical Alzheimer’s Disease. Brain 2019, 142, 2096–2112. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, Z.; Luo, L.; Tong, H.; Chen, F.; Hou, S.T. 40 Hz Light Flicker Alters Human Brain Electroencephalography Microstates and Complexity Implicated in Brain Diseases. Front. Neurosci. 2021, 15, 777183. [Google Scholar] [CrossRef]
- Zhao, Z.; Niu, Y.; Zhao, X.; Zhu, Y.; Shao, Z.; Wu, X.; Wang, C.; Gao, X.; Wang, C.; Xu, Y.; et al. EEG Microstate in First-Episode Drug-Naive Adolescents with Depression. J. Neural Eng. 2022, 19, 056016. [Google Scholar] [CrossRef]
- Liu, S.; Guo, J.; Meng, J.; Wang, Z.; Yao, Y.; Yang, J.; Qi, H.; Ming, D. Abnormal EEG Complexity and Functional Connectivity of Brain in Patients with Acute Thalamic Ischemic Stroke. Comput. Math. Methods Med. 2016, 2016, 2582478. [Google Scholar] [CrossRef]
- Wei, R.; Ganglberger, W.; Sun, H.; Hadar, P.N.; Gollub, R.L.; Pieper, S.; Billot, B.; Au, R.; Iglesias, J.E.; Cash, S.S.; et al. Linking Brain Structure, Cognition, and Sleep: Insights from Clinical Data. Sleep 2024, 47, zsad294. [Google Scholar] [CrossRef]
- Doan, D.N.T.; Ku, B.; Choi, J.; Oh, M.; Kim, K.; Cha, W.; Kim, J.U. Predicting Dementia with Prefrontal Electroencephalography and Event-Related Potential. Front. Aging Neurosci. 2021, 13, 659817. [Google Scholar] [CrossRef]
- Jesus, B.; Cassani, R.; McGeown, W.J.; Cecchi, M.; Fadem, K.C.; Falk, T.H. Multimodal Prediction of Alzheimer’s Disease Severity Level Based on Resting-State EEG and Structural MRI. Front. Hum. Neurosci. 2021, 15, 700627. [Google Scholar] [CrossRef]
- Zorick, T.; Landers, J.; Leuchter, A.; Mandelkern, M.A. EEG Multifractal Analysis Correlates with Cognitive Testing Scores and Clinical Staging in Mild Cognitive Impairment. J. Clin. Neurosci. 2020, 76, 195–200. [Google Scholar] [CrossRef]
- Si, Y.; He, R.; Jiang, L.; Yao, D.; Zhang, H.; Xu, P.; Ma, X.; Yu, L.; Li, F. Differentiating between Alzheimer’s Disease and Frontotemporal Dementia Based on the Resting-State Multilayer EEG Network. IEEE Trans. Neural Syst. Rehabil. Eng. 2023, 31, 4521–4527. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).