Changepoint Detection in Noisy Data Using a Novel Residuals Permutation-Based Method (RESPERM): Benchmarking and Application to Single Trial ERPs
Abstract
:1. Introduction
1.1. The Changepoint Regression Model
1.2. SEGMENTED Method
1.3. Application to ERPs
2. Materials and Methods
2.1. The Residuals Permutation-Based Method (RESPERM)
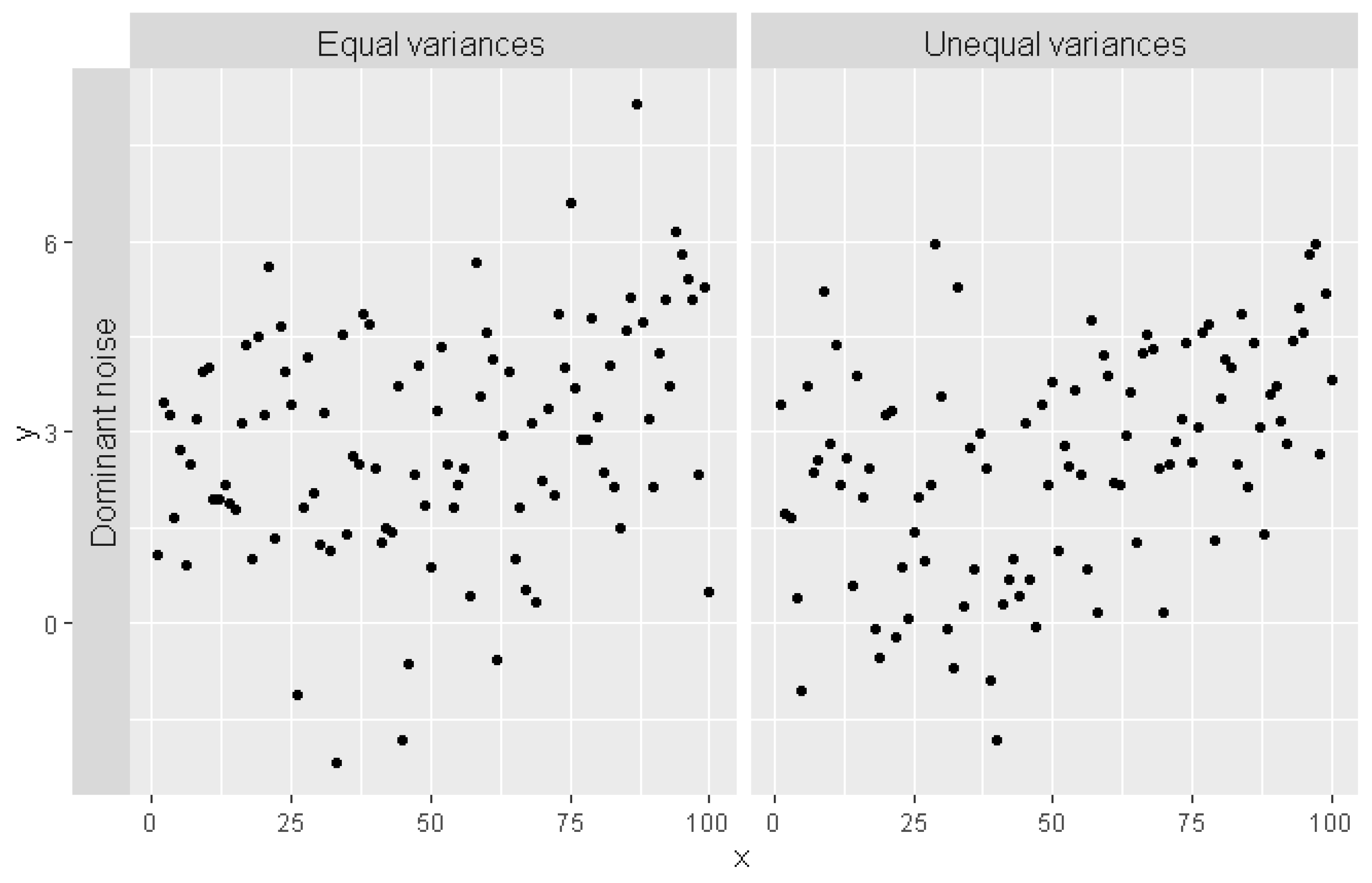
2.2. Monte Carlo Verification Setup
- (1)
- ,
- (2)
- ,
- (3)
- ,
- (4)
- ).
2.3. Single-Trial ERP Data
3. Results
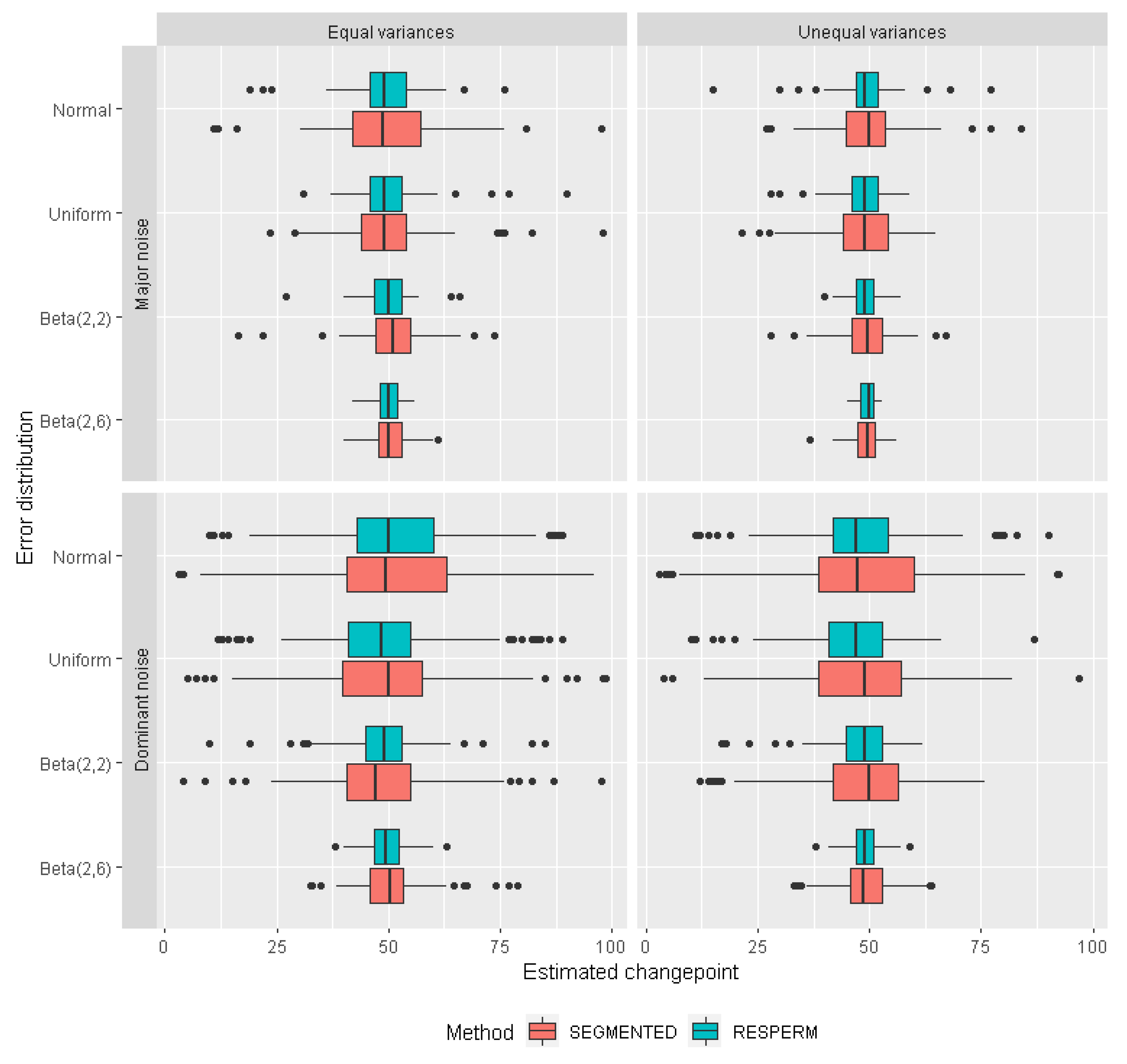
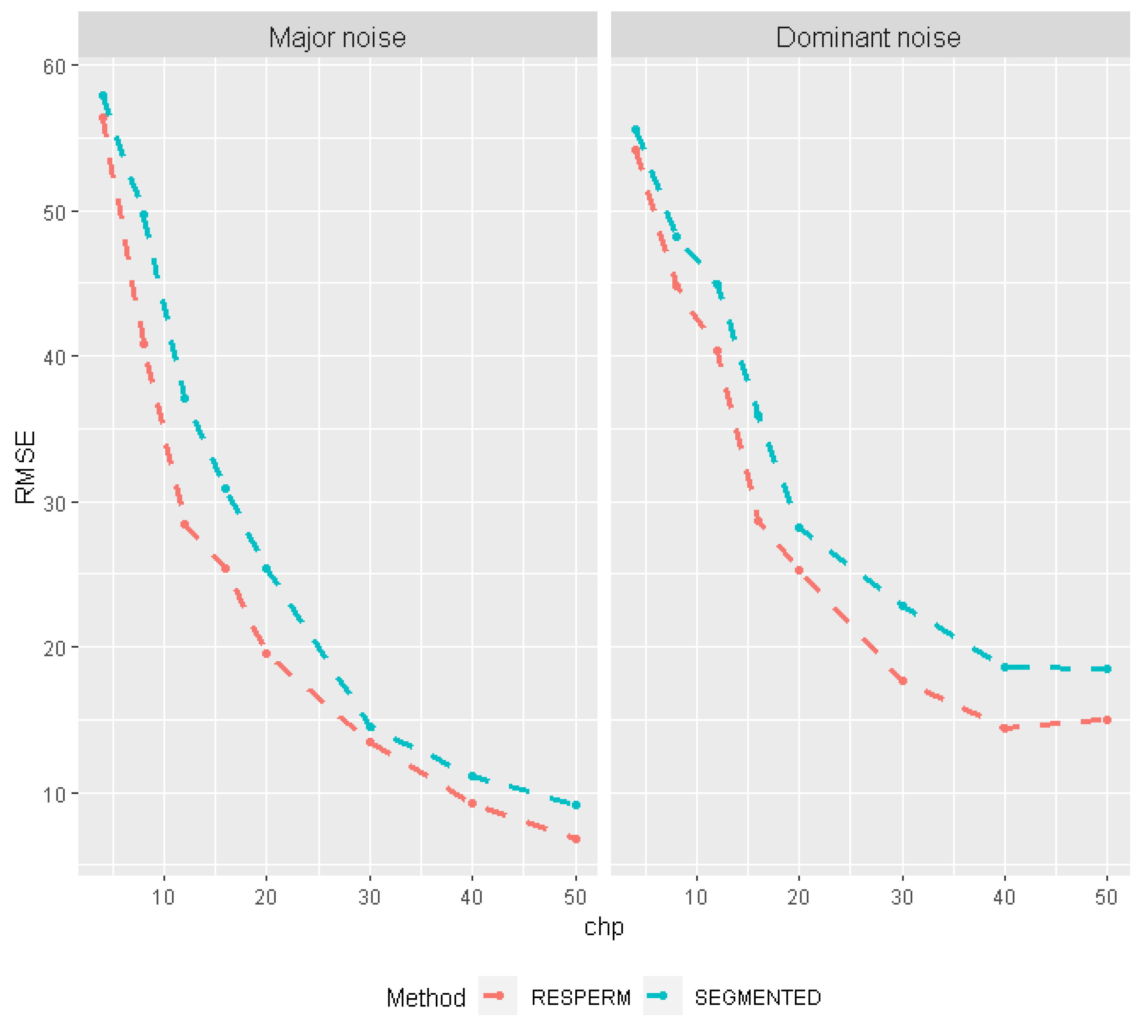
3.1. Monte Carlo Simulations
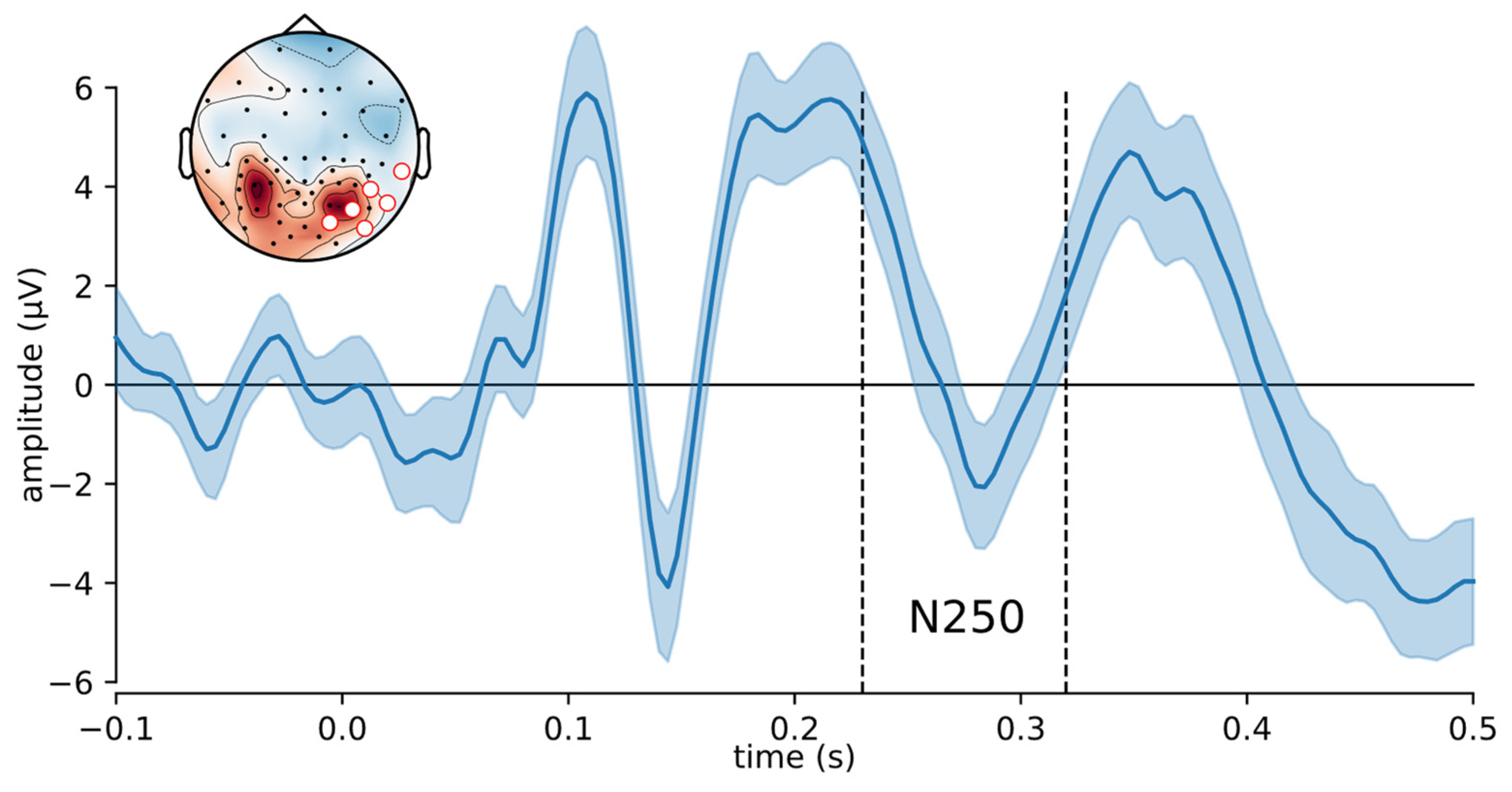
3.2. Application to Single Trial ERP Data from a Face Memory Task
4. Discussion and Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Appendix A
| Algorithm A1: The code of RESPERM implemented in R. |
| res.perm <- function(x,y,N_perm=100) { n = length(x) first_k = 10 last_k = n − 10 Cohen_d = rep(NA,n) simple.fit = lm(y~x) res = simple.fit$residuals yf = simple.fit$fitted if (N_perm < 100) stop(“Too few permutations”) if (n < 50) stop(“Too few observations”) ### Finding the greatest value of the vector Cohen_d for (k in first_k:last_k) { simple1.fit = lm(y[1:k]~x[1:k]) simple2.fit = lm(y[(k+1):n]~x[(k+1):n]) b1 = simple1.fit$coefficients[2] b2 = simple2.fit$coefficients[2] b1s = c(); b2s = c() for (i in 1:N_perm) { ys = yf + sample(res) b1s[i] = lm(ys[1:k]~x[1:k])$coefficients[2] b2s[i] = lm(ys[(k + 1):n]~x[(k + 1):n])$coefficients[2] } Cohen_d[k] = (b2 − b1)/sqrt(((k − 1)*var(b1s)+(n − k − 1)*var(b2s))/(n − 2)) } d = max(Cohen_d[first_k:last_k], na.rm =T) k_star = order(Cohen_d[first_k:last_k],decreasing = T)[1] + first_k−1 return(list(“k_star” = k_star, “chp” = x[k_star], “d” = d)) } |
References
- Truong, C.; Oudre, L.; Vayatis, N. Selective Review of Offline Change Point Detection Methods. Signal Process. 2020, 167, 107299. [Google Scholar] [CrossRef] [Green Version]
- Chow, G.C. Tests of Equality Between Sets of Coefficients in Two Linear Regressions. Econometrica 1960, 28, 591–605. [Google Scholar] [CrossRef]
- Aminikhanghahi, S.; Cook, D.J. A Survey of Methods for Time Series Change Point Detection. Knowl. Inf. Syst. 2017, 51, 339–367. [Google Scholar] [CrossRef] [Green Version]
- Lerman, P.M. Fitting Segmented Regression Models by Grid Search. J. R. Stat. Soc. Ser. C (Appl. Stat.) 1980, 29, 77–84. [Google Scholar] [CrossRef]
- Ulm, K. A Statistical Method for Assessing a Threshold in Epidemiological Studies. Stat. Med. 1991, 10, 341–349. [Google Scholar] [CrossRef]
- Muggeo, V.M.R. Estimating Regression Models with Unknown Break-Points. Stat. Med. 2003, 22, 3055–3071. [Google Scholar] [CrossRef]
- Tanaka, J.W.; Curran, T.; Porterfield, A.L.; Collins, D. Activation of Preexisting and Acquired Face Representations: The N250 Event-Related Potential as an Index of Face Familiarity. J. Cogn. Neurosci. 2006, 18, 1488–1497. [Google Scholar] [CrossRef]
- Sommer, W.; Stapor, K.; Kończak, G.; Kotowski, K.; Fabian, P.; Ochab, J.; Bereś, A.; Ślusarczyk, G. The N250 Event-Related Potential as an Index of Face Familiarity: A Replication Study. R. Soc. Open Sci. 2021, 8, 202356. [Google Scholar] [CrossRef]
- Muggeo, V.M. Segmented: An R Package to Fit Regression Models with Broken-Line Relationships. R News 2008, 8, 20–25. [Google Scholar]
- Doerr, E.M.; Carretti, B.; Toffalini, E.; Lanfranchi, S.; Meneghetti, C. Developmental Trajectories in Spatial Visualization and Mental Rotation in Individuals with Down Syndrome. Brain Sci. 2021, 11, 610. [Google Scholar] [CrossRef]
- Küchenhoff, H.; Günther, F.; Höhle, M.; Bender, A. Analysis of the Early COVID-19 Epidemic Curve in Germany by Regression Models with Change Points. Epidemiol. Infect. 2021, 149, e68. [Google Scholar] [CrossRef]
- Rhodes, G.; Calder, A.; Johnson, M.; Haxby, J.V.; Eimer, M. The Face-Sensitive N170 Component of the Event-Related Brain Potential. In Oxford Handbook of Face Perception; Oxford University Press: Oxford, UK, 2012; ISBN 978-0-19-955905-3. [Google Scholar]
- Cecotti, H.; Rivet, B. Subject Combination and Electrode Selection in Cooperative Brain-Computer Interface Based on Event Related Potentials. Brain Sci. 2014, 4, 335–355. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Puce, A.; Hämäläinen, M.S. A Review of Issues Related to Data Acquisition and Analysis in EEG/MEG Studies. Brain Sci. 2017, 7, 58. [Google Scholar] [CrossRef] [PubMed]
- Luck, S.J. An Introduction to the Event-Related Potential Technique, 2nd ed.; The MIT Press: Cambridge, MA, USA, 2014; ISBN 978-0-262-52585-5. [Google Scholar]
- Ouyang, G.; Sommer, W.; Zhou, C. Reconstructing ERP Amplitude Effects after Compensating for Trial-to-Trial Latency Jitter: A Solution Based on a Novel Application of Residue Iteration Decomposition. Int. J. Psychophysiol. 2016, 109, 9–20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ouyang, G.; Hildebrandt, A.; Sommer, W.; Zhou, C. Exploiting the Intra-Subject Latency Variability from Single-Trial Event-Related Potentials in the P3 Time Range: A Review and Comparative Evaluation of Methods. Neurosci. Biobehav. Rev. 2017, 75, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J. Statistical Power Analysis for the Behavioral Sciences, 2nd ed.; Routledge: New York, NY, USA, 1988. [Google Scholar]
- Davison, A.C.; Hinkley, D.V. Cambridge Series in Statistical and Probabilistic Mathematics. In Bootstrap Methods and Their Application; Cambridge University Press: Cambridge, MA, USA, 1997; ISBN 978-0-521-57471-6. [Google Scholar]
- Good, P.I. Permutation, Parametric, and Bootstrap Tests of Hypotheses, 3rd ed.; Springer: New York, NY, USA, 2005. [Google Scholar]
- Grootswagers, T.; Wardle, S.G.; Carlson, T.A. Decoding Dynamic Brain Patterns from Evoked Responses: A Tutorial on Multivariate Pattern Analysis Applied to Time Series Neuroimaging Data. J. Cogn. Neurosci. 2017, 29, 677–697. [Google Scholar] [CrossRef] [PubMed]

| Error Distribution | Noise Level | Equal Variances | Unequal Variances | ||
|---|---|---|---|---|---|
| SEGMENTED | RESPERM | SEGMENTED | RESPERM | ||
| Normal | Major | 12.96 | 7.88 | 9.16 | 6.88 |
| Dominant | 20.48 | 17.38 | 18.51 | 14.94 | |
| Uniform | Major | 11.09 | 7.71 | 9.04 | 5.89 |
| Dominant | 19.55 | 15.35 | 16.42 | 14.16 | |
| Beta (2,2) | Major | 8.12 | 4.63 | 6.12 | 3.30 |
| Dominant | 15.43 | 10.39 | 12.51 | 8.79 | |
| Beta (2,6) | Major | 4.10 | 2.75 | 3.52 | 2.06 |
| Dominant | 8.10 | 4.17 | 6.59 | 3.62 | |
| Errors Distribution | Noise Level | Equal Variances | Unequal Variances | ||
|---|---|---|---|---|---|
| SEGMENTED | RESPERM | SEGMENTED | RESPERM | ||
| RB/SD | RB/SD | RB/SD | RB/SD | ||
| Normal | Major | 0.12/12.96 | −1.26/7.86 | −1.60/9.13 | −2.08/6.80 |
| Dominant | 1.02/20.47 | 0.64/17.38 | −3.84/18.41 | −5.78/14.66 | |
| Uniform | Major | −0.66/11.08 | −0.20/7.71 | −3.16/8.90 | −3.84/5.57 |
| Dominant | −1.08/19.54 | −1.72/15.32 | −5.28/16.21 | −9.68/13.30 | |
| Beta (2,2) | Major | 1.66/8.07 | 0.32/4.63 | −1.32/6.09 | −2.20/3.11 |
| Dominant | −3.02/15.36 | −1.88/10.35 | −3.30/12.40 | −4.42/8.51 | |
| Beta (2,6) | Major | 0.68/4.09 | 0.42/2.74 | −1.58/3.43 | −1.44/1.93 |
| Dominant | 1.80/8.05 | −0.54/4.16 | −2.56/6.46 | −1.80/3.51 | |
| Error Distribution Type | Major Noise | Dominant Noise | ||
|---|---|---|---|---|
| eV | ueV | eV | ueV | |
| Normal | 0.59 | 0.75 | 0.83 | 0.46 |
| Uniform | 0.82 | 0.61 | 0.66 | 0.42 |
| Beta (2,2) | 0.81 | 0.79 | 0.80 | 0.69 |
| Beta (2,6) | 0.84 | 0.67 | 0.77 | 0.87 |
| RESPERM | SEGMENTED | ||||
|---|---|---|---|---|---|
| Participant Number | d | kres | chpres | kseg | chpseg |
| 3 | 3.556 | 14 | 122 | 13 | 110 |
| 6 | 6.250 | 12 | 139 | 10 | 114 |
| 2 | 4.636 | 16 | 179 | 15 | 165 |
| 15 | 3.791 | 17 | 188 | 12 | 136 |
| 17 | 4.512 | 17 | 208 | 14 | 172 |
| 9 | 5.340 | 21 | 235 | 20 | 226 |
| 20 | 2.088 | 24 | 282 | 29 | 334 |
| 14 | 1.358 | 10 | 284 | 27 | 486 |
| 18 | 3.631 | 29 | 319 | 29 | 319 |
| 5 | 4.520 | 28 | 335 | 26 | 305 |
| 13 | 3.120 | 30 | 365 | 48 | 572 |
| 19 | 4.563 | 45 | 370 | 22 | 177 |
| 7 | 5.781 | 35 | 389 | 34 | 378 |
| 11 | 5.089 | 48 | 569 | 52 | 613 |
| 12 | 4.029 | 50 | 578 | 57 | 657 |
| 4 | 2.058 | 57 | 673 | - | - |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sommer, W.; Stapor, K.; Kończak, G.; Kotowski, K.; Fabian, P.; Ochab, J.; Bereś, A.; Ślusarczyk, G. Changepoint Detection in Noisy Data Using a Novel Residuals Permutation-Based Method (RESPERM): Benchmarking and Application to Single Trial ERPs. Brain Sci. 2022, 12, 525. https://doi.org/10.3390/brainsci12050525
Sommer W, Stapor K, Kończak G, Kotowski K, Fabian P, Ochab J, Bereś A, Ślusarczyk G. Changepoint Detection in Noisy Data Using a Novel Residuals Permutation-Based Method (RESPERM): Benchmarking and Application to Single Trial ERPs. Brain Sciences. 2022; 12(5):525. https://doi.org/10.3390/brainsci12050525
Chicago/Turabian StyleSommer, Werner, Katarzyna Stapor, Grzegorz Kończak, Krzysztof Kotowski, Piotr Fabian, Jeremi Ochab, Anna Bereś, and Grażyna Ślusarczyk. 2022. "Changepoint Detection in Noisy Data Using a Novel Residuals Permutation-Based Method (RESPERM): Benchmarking and Application to Single Trial ERPs" Brain Sciences 12, no. 5: 525. https://doi.org/10.3390/brainsci12050525
APA StyleSommer, W., Stapor, K., Kończak, G., Kotowski, K., Fabian, P., Ochab, J., Bereś, A., & Ślusarczyk, G. (2022). Changepoint Detection in Noisy Data Using a Novel Residuals Permutation-Based Method (RESPERM): Benchmarking and Application to Single Trial ERPs. Brain Sciences, 12(5), 525. https://doi.org/10.3390/brainsci12050525








