Tunable-Q Wavelet Transform Based Multiscale Entropy Measure for Automated Classification of Epileptic EEG Signals
Abstract
:1. Introduction
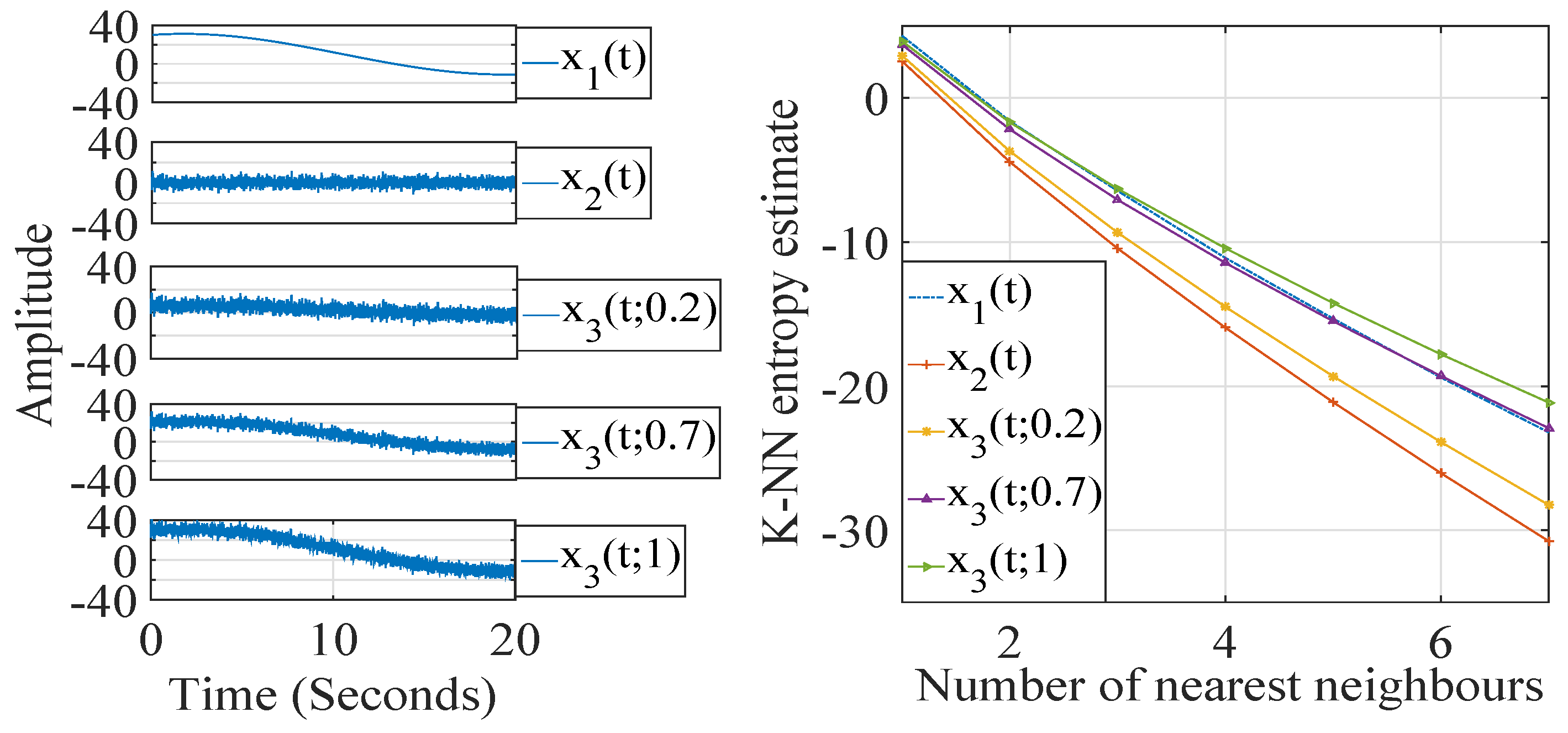
2. K-Nearest Neighbor Entropy Estimation
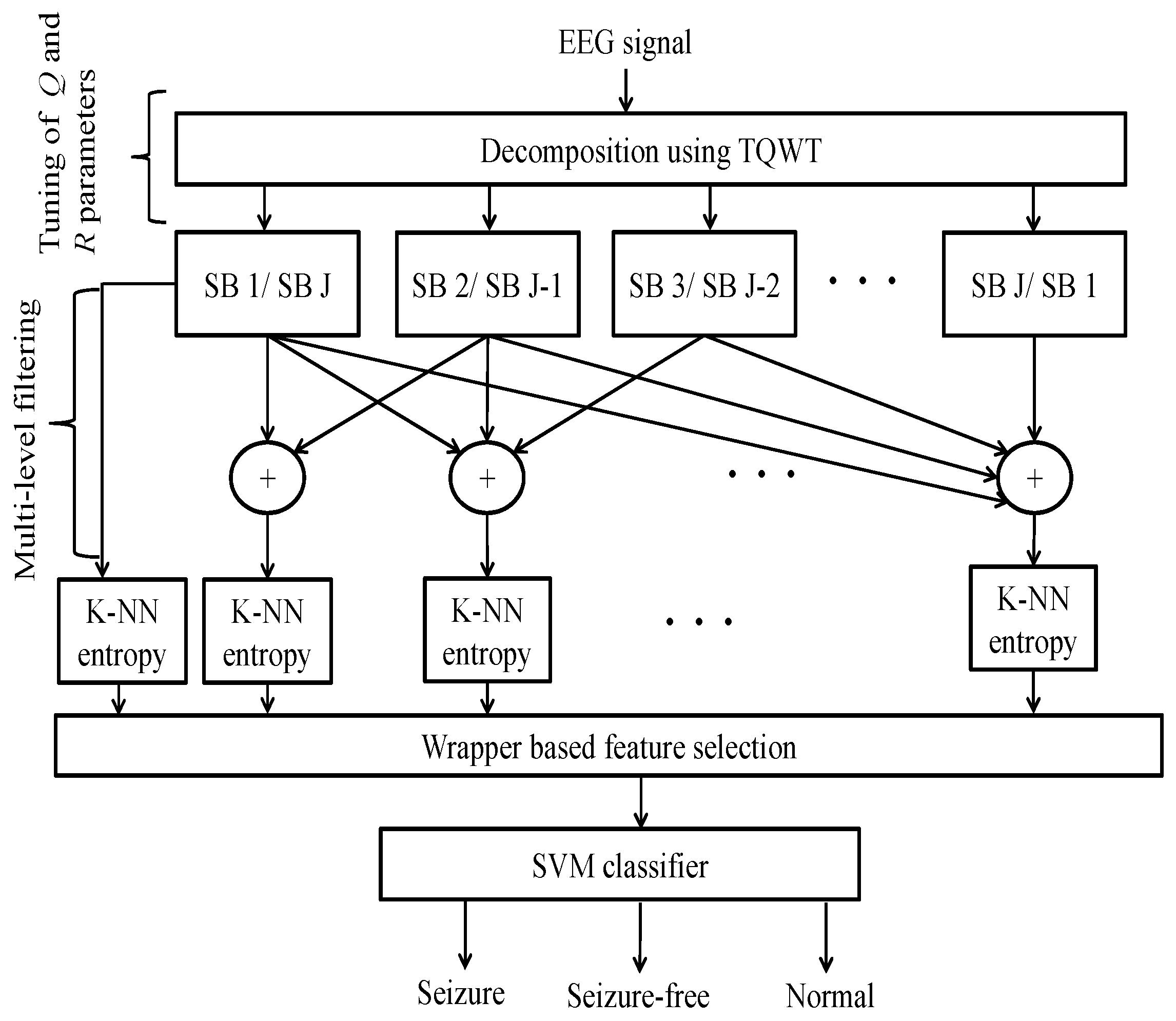
3. TQWT-Based K-NN Entropy
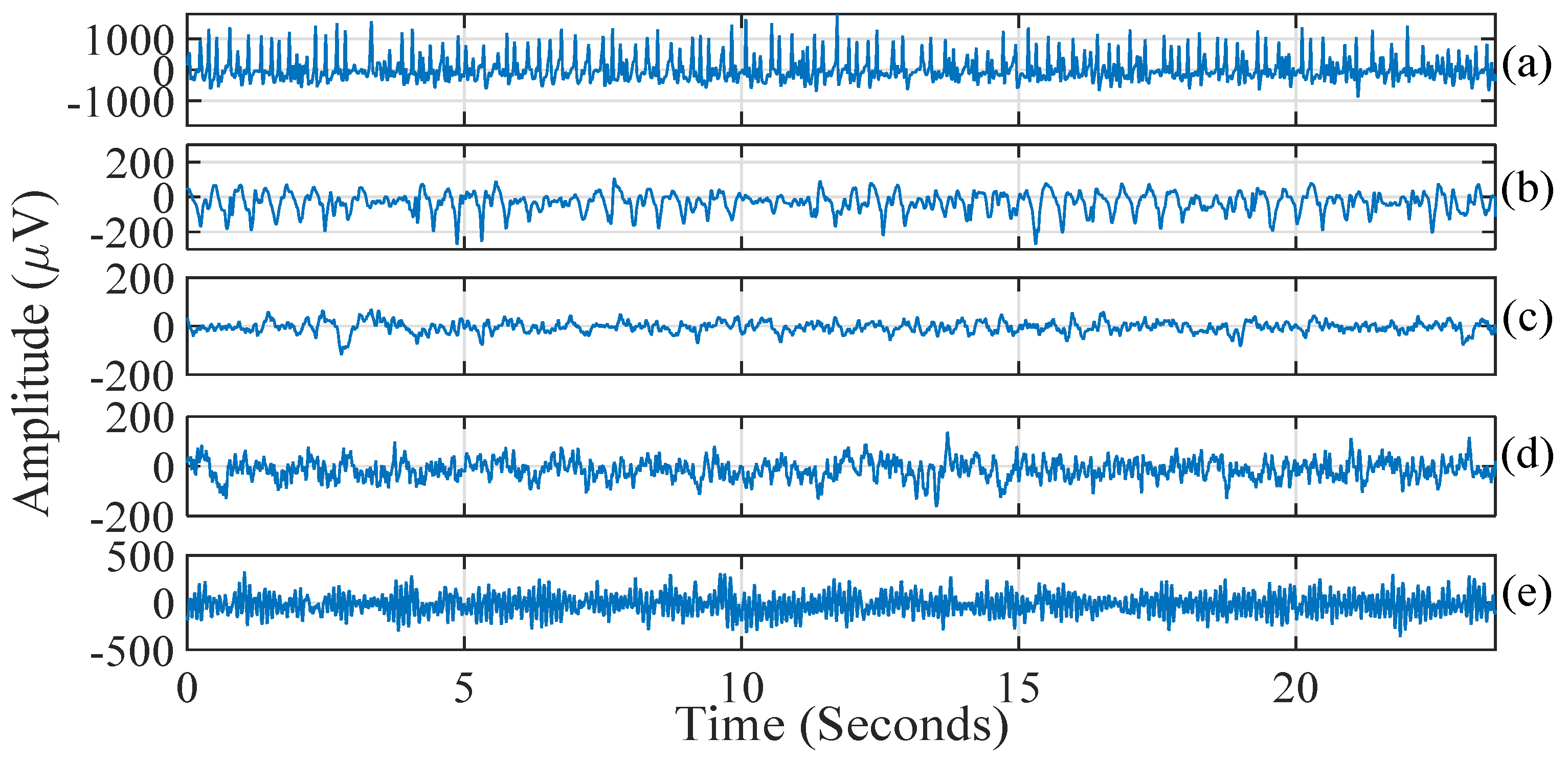
- Sub-band signals denoted as are reconstructed by applying the inverse TQWT operation.
- To measure the complexity at multiple oscillatory levels, K-NN entropy has been computed on the signals, generated by cumulatively summing the reconstructed sub-band signals. We have formulated two kinds of multilevel filtering approaches as follows:
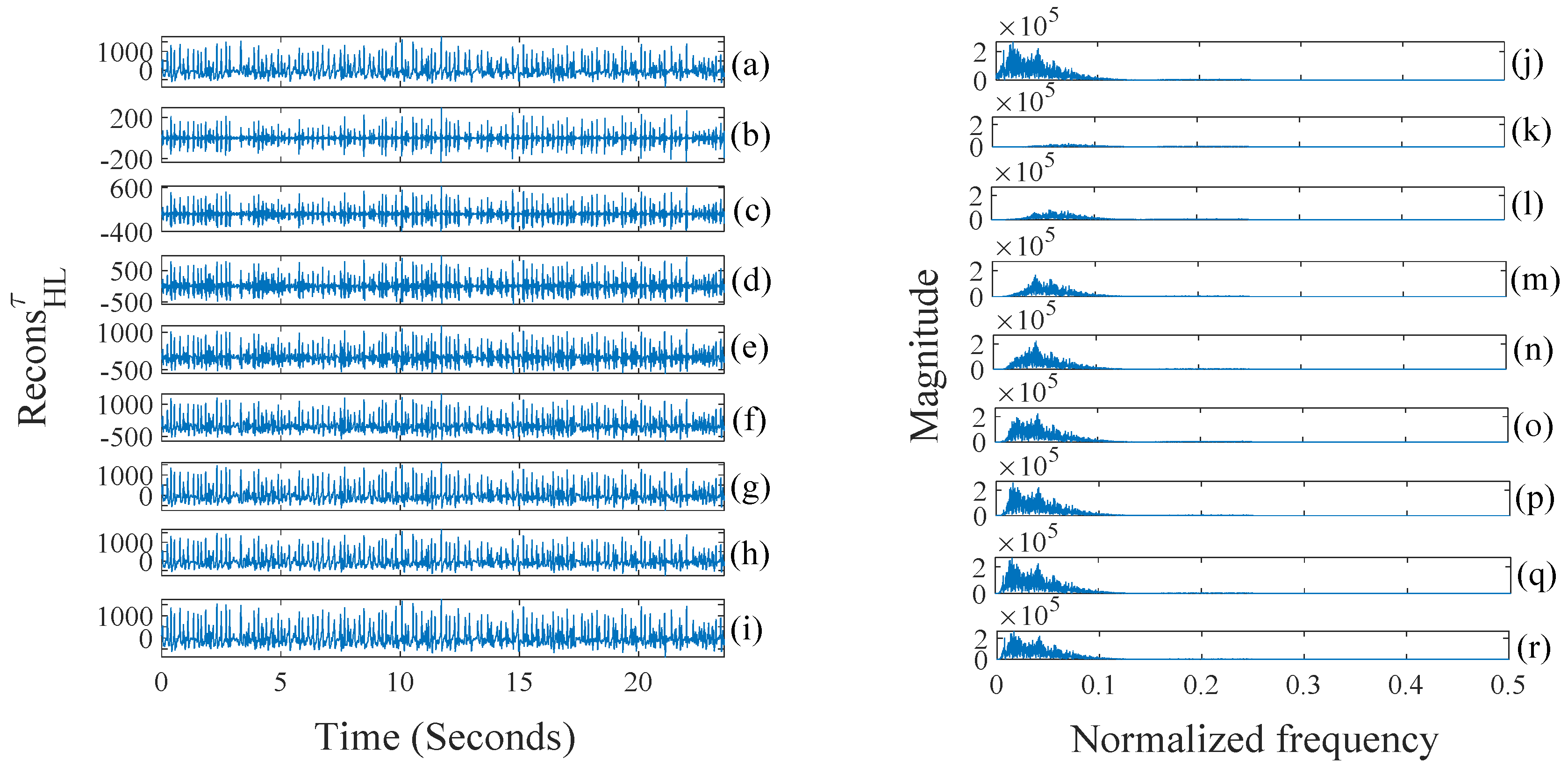
- (a)
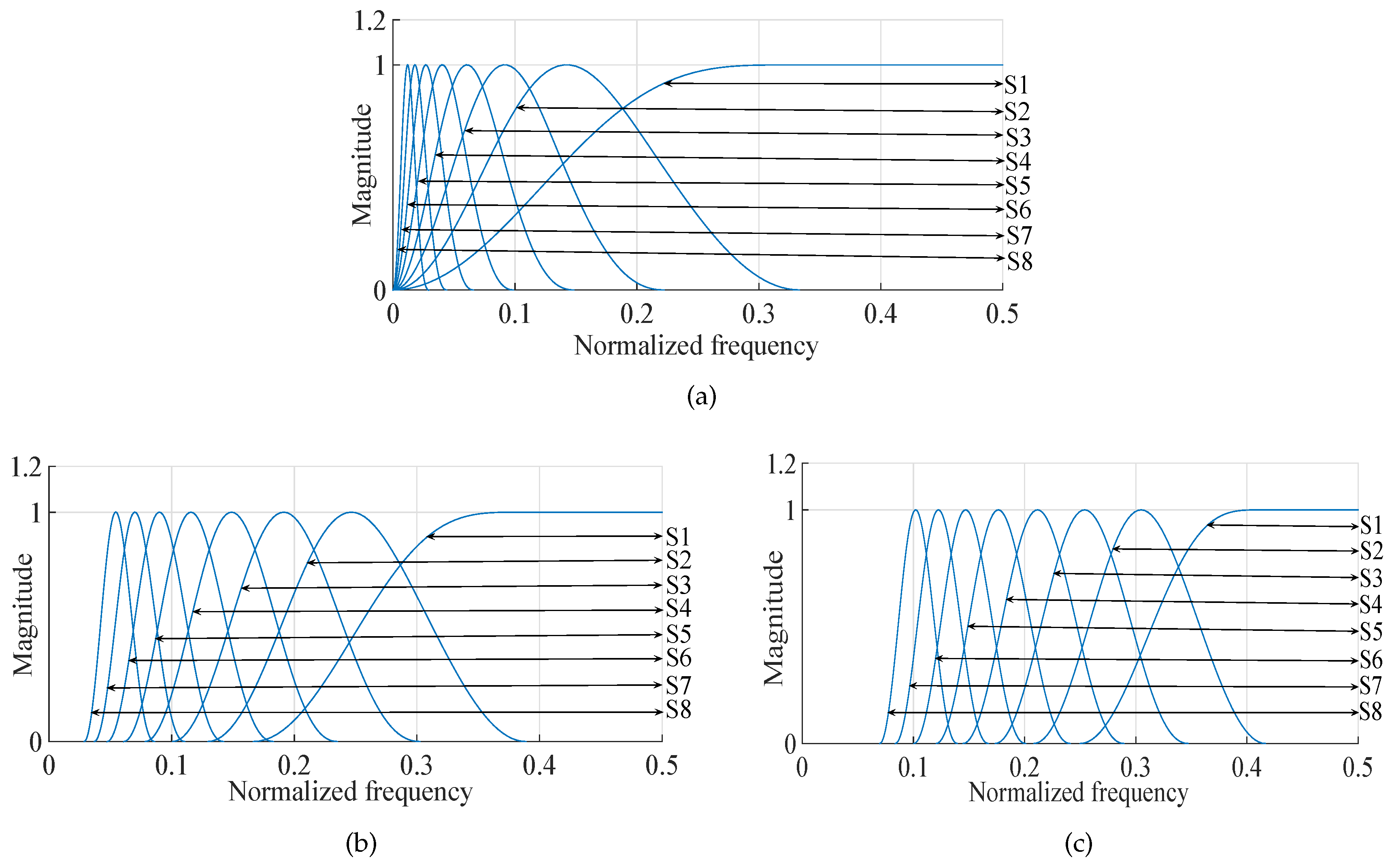
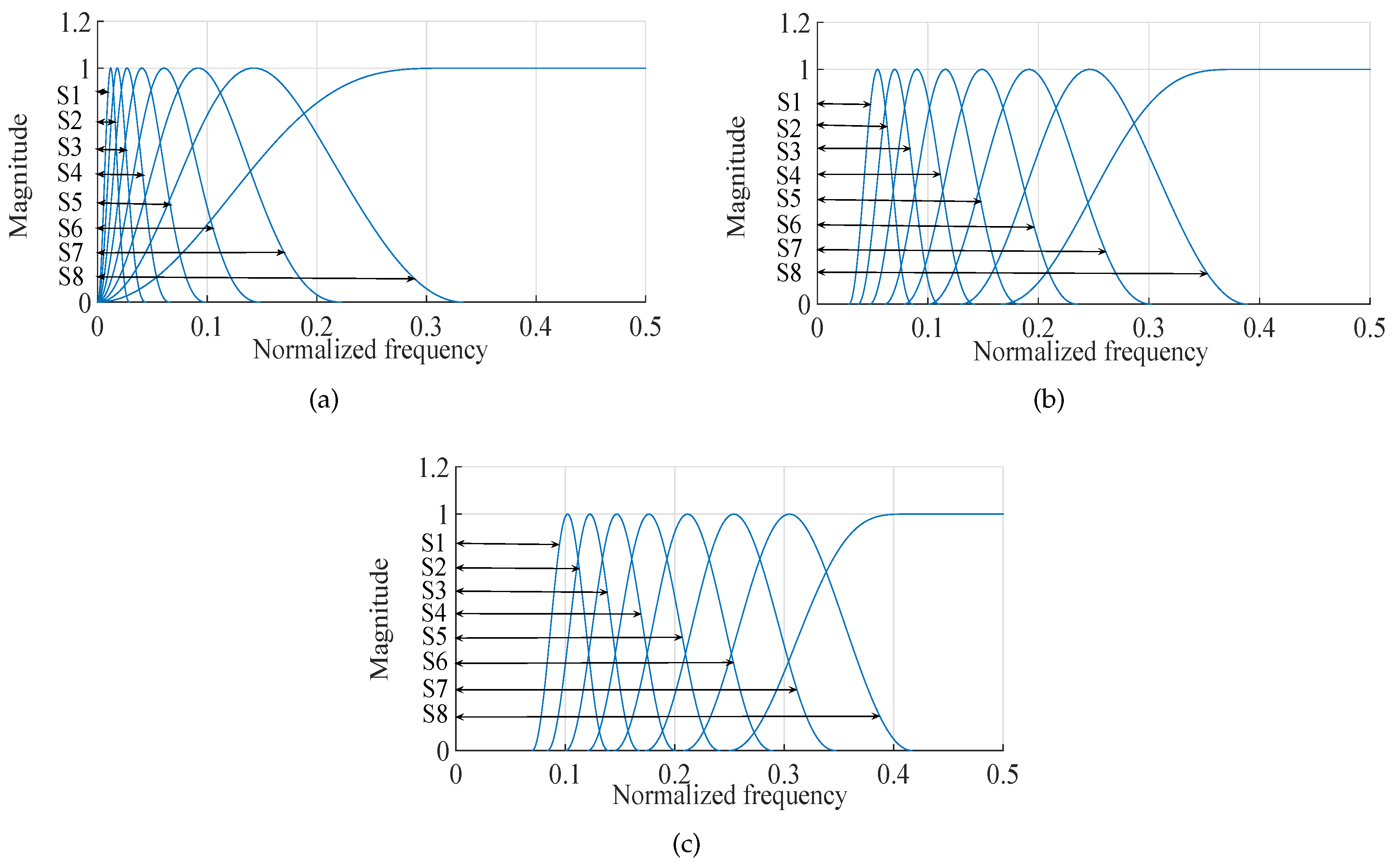
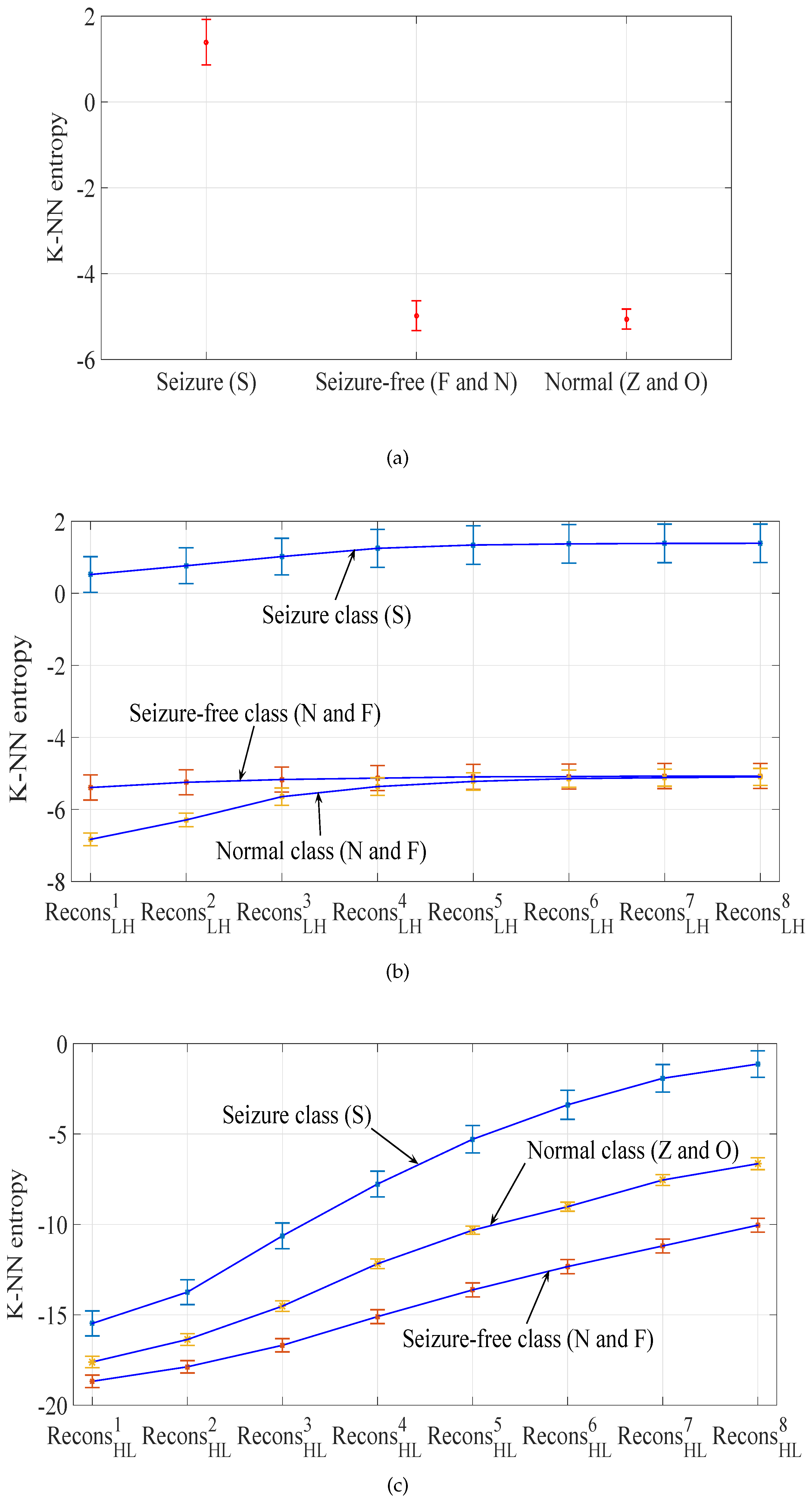
- Detailed sub-band to approximate sub-band: The multilevel filtering starts from the highest oscillatory level sub-band to the dominant low-frequency trend sub-band . Finally, can be mathematically formulated as follows:where corresponds to highest oscillatory sub-band and corresponds to the dominant low-frequency trend, is the scale factor and l is the number of nearest neighbors.
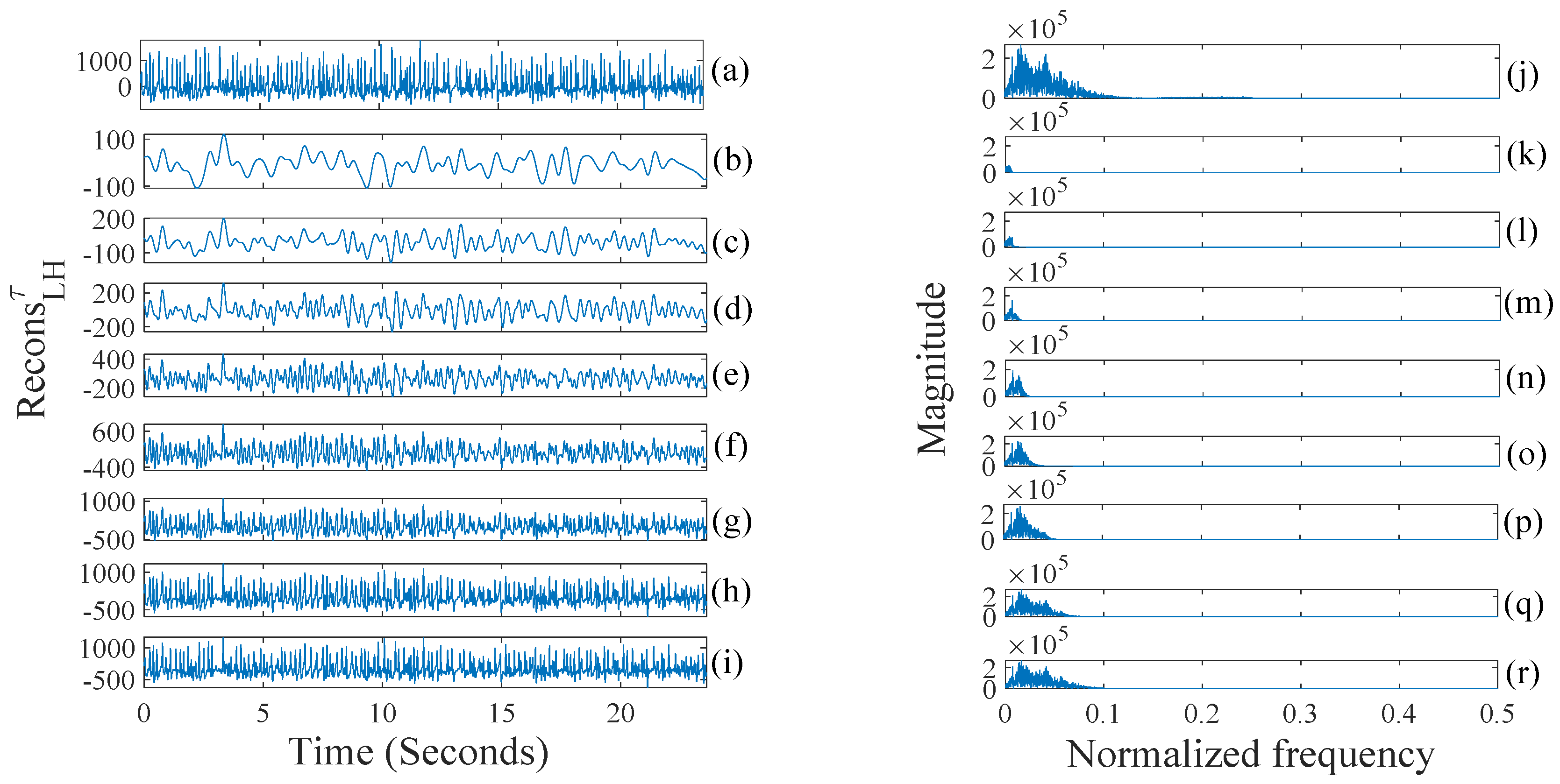
- (b)
- Approximate sub-band to the detailed sub-band: The multilevel filtering starts from the dominant low-frequency trend sub-band to the highest oscillatory level sub-band . Finally, can be expressed as follows:where corresponds to the dominant low-frequency trend and corresponds to the highest oscillatory sub-band, denotes the scale factor and l represents number of nearest neighbors. In this work, we have set for K-NN entropy estimation.
4. EEG Dataset
- (S-Z): classification of two classes, namely seizure (S) EEG signals and normal eyes-open (Z) EEG signals.
- (S-O): classification of two classes, namely seizure (S) EEG signals and normal eyes-closed (O) EEG signals.
- (S-N): classification of two classes, namely seizure (S) EEG signals and seizure-free (N) EEG signals.
- (S-F): classification of two classes, namely seizure (S) EEG signals and seizure-free (F) EEG signals.
- (S-FNZO): classification of two classes, namely seizure (S) EEG signals and non-seizure (F, N, Z and O) EEG signals.
- (S-FN-ZO): classification of three classes, namely seizure (S) EEG signals, seizure-free (F and N) EEG signals and normal (Z and O) EEG signals.
5. Classification of EEG Records
6. Experimental Results
7. Discussion
8. Conclusions
Author Contributions
Conflicts of Interest
References
- Witte, H.; Iasemidis, L.D.; Litt, B. Special issue on epileptic seizure prediction. IEEE Trans. Biomed. Eng. 2003, 50, 537–539. [Google Scholar] [CrossRef]
- Sharma, R.; Pachori, R.B. Classification of epileptic seizures in EEG signals based on phase space representation of intrinsic mode functions. Expert Syst. Appl. 2015, 42, 1106–1117. [Google Scholar] [CrossRef]
- Bajaj, V.; Pachori, R.B. Classification of seizure and nonseizure EEG signals using empirical mode decomposition. IEEE Trans. Inf. Technol. Biomed. 2012, 16, 1135–1142. [Google Scholar] [CrossRef] [PubMed]
- Pachori, R.B.; Sharma, R.; Patidar, S. Classification of normal and epileptic seizure EEG signals based on empirical mode decomposition. In Complex System Modelling and Control through Intelligent Soft Computations; Zhu, Q., Azar, A.T., Eds.; Springer International Publishing: Cham, Switzerland, 2015; pp. 367–388. [Google Scholar]
- Tzallas, A.T.; Tsipouras, M.G.; Fotiadis, D.I. Epileptic seizure detection in EEGs using time-frequency analysis. IEEE Trans. Inf. Technol. Biomed. 2009, 13, 703–710. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Yanti, R.; Zheng, J.W.; Krishnan, M.M.R.; TAN, J.H.; Martis, R.J.; Lim, C.M. Automated diagnosis of epilepsy using CWT, HOS and texture parameters. Int. J. Neural Syst. 2013, 23, 1350009. [Google Scholar] [CrossRef] [PubMed]
- Subasi, A.; Gursoy, M.I. EEG signal classification using PCA, ICA, LDA and support vector machines. Expert Syst. Appl. 2010, 37, 8659–8666. [Google Scholar] [CrossRef]
- Uthayakumar, R.; Easwaramoorthy, D. Epileptic seizure detection in EEG signals using multifractal analysis and wavelet transform. Fractals 2013, 21, 1350011. [Google Scholar] [CrossRef]
- Patidar, S.; Panigrahi, T. Detection of epileptic seizure using Kraskov entropy applied on tunable-Q wavelet transform of EEG signals. Biomed. Signal Process. Control 2017, 34, 74–80. [Google Scholar] [CrossRef]
- Hassan, A.R.; Siuly, S.; Zhang, Y. Epileptic seizure detection in EEG signals using tunable-Q factor wavelet transform and bootstrap aggregating. Comput. Methods Progr. Biomed. 2016, 137, 247–259. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Sree, S.V.; Chattopadhyay, S.; Yu, W.; Ang, P.C.A. Application of recurrence quantification analysis for the automated identification of epileptic EEG signals. Int. J. Neural Syst. 2011, 21, 199–211. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Chua, C.K.; Lim, T.C.; Dorithy; Suri, J.S. Automatic identification of epileptic EEG signals using nonlinear parameters. J. Mech. Med. Biol. 2009, 9, 539–553. [Google Scholar] [CrossRef]
- Srinivasan, V.; Eswaran, C.; Sriraam, N. Approximate entropy-based epileptic EEG detection using artificial neural networks. IEEE Trans. Inf. Technol. Biomed. 2007, 11, 288–295. [Google Scholar] [CrossRef] [PubMed]
- Kaya, Y.; Uyar, M.; Tekin, R.; Yıldırım, S. 1D-local binary pattern based feature extraction for classification of epileptic EEG signals. Appl. Math. Comput. 2014, 243, 209–219. [Google Scholar] [CrossRef]
- Kumar, T.S.; Kanhangad, V.; Pachori, R.B. Classification of seizure and seizure-free EEG signals using local binary patterns. Biomed. Signal Process. Control 2015, 15, 33–40. [Google Scholar] [CrossRef]
- Tiwari, A.K.; Pachori, R.B.; Kanhangad, V.; Panigrahi, B. Automated diagnosis of epilepsy using key-point based local binary pattern of EEG signals. IEEE J. Biomed. Health Inform. 2016, in press. [Google Scholar] [CrossRef] [PubMed]
- Altunay, S.; Telatar, Z.; Erogul, O. Epileptic EEG detection using the linear prediction error energy. Expert Syst. Appl. 2010, 37, 5661–5665. [Google Scholar] [CrossRef]
- Joshi, V.; Pachori, R.B.; Vijesh, A. Classification of ictal and seizure-free EEG signals using fractional linear prediction. Biomed. Signal Process. Control 2014, 9, 1–5. [Google Scholar] [CrossRef]
- Ghosh-Dastidar, S.; Adeli, H.; Dadmehr, N. Principal component analysis-enhanced cosine radial basis function neural network for robust epilepsy and seizure detection. IEEE Trans. Biomed. Eng. 2008, 55, 512–518. [Google Scholar] [CrossRef] [PubMed]
- Mammone, N.; Morabito, F.C.; Principe, J.C. Visualization of the short term maximum Lyapunov exponent topography in the epileptic brain. In Proceedings of the 28th Annual International Conference of the IEEE Engineering in Medicine and Biology Society, New York, NY, USA, 30 August–3 September 2006; pp. 4257–4260. [Google Scholar]
- Mammone, N.; Principe, J.C.; Morabito, F.C.; Shiau, D.S.; Sackellares, J.C. Visualization and modeling of STLmax topographic brain activity maps. J. Neurosci. Methods 2010, 189, 281–294. [Google Scholar] [CrossRef] [PubMed]
- Mammone, N.; Morabito, F.C. Analysis of absence seizure EEG via Permutation entropy spatio-temporal clustering. Proceedingd of the 2011 International Joint Conference on Neural Networks (IJCNN), San Jose, CA, USA, 31 July–5 August 2011; pp. 1417–1422. [Google Scholar]
- Bandt, C.; Pompe, B. Permutation entropy: A natural complexity measure for time series. Phys. Rev. Lett. 2002, 88, 174102. [Google Scholar] [CrossRef] [PubMed]
- Mammone, N.; Duun-Henriksen, J.; Kjaer, T.W.; Morabito, F.C. Differentiating interictal and ictal states in childhood absence epilepsy through permutation Rényi entropy. Entropy 2015, 17, 4627–4643. [Google Scholar] [CrossRef]
- Kumar, M.; Pachori, R.B.; Acharya, U.R. An efficient automated technique for CAD diagnosis using flexible analytic wavelet transform and entropy features extracted from HRV signals. Expert Syst. Appl. 2016, 63, 165–172. [Google Scholar] [CrossRef]
- Sharma, R.; Pachori, R.B.; Acharya, U.R. Application of entropy measures on intrinsic mode functions for the automated identification of focal electroencephalogram signals. Entropy 2015, 17, 669–691. [Google Scholar] [CrossRef]
- Patidar, S.; Pachori, R.B.; Acharya, U.R. Automated diagnosis of coronary artery disease using tunable-Q wavelet transform applied on heart rate signals. Knowl. Based Syst. 2015, 82, 1–10. [Google Scholar] [CrossRef]
- Zhang, Z.; Zhou, Y.; Chen, Z.; Tian, X.; Du, S.; Huang, R. Approximate entropy and support vector machines for electroencephalogram signal classification. Neural Regen. Res. 2013, 8, 1844. [Google Scholar] [PubMed]
- Lin, J. Divergence measures based on the Shannon entropy. IEEE Trans. Inf. Theory 1991, 37, 145–151. [Google Scholar] [CrossRef]
- Grassberger, P.; Procaccia, I. Estimation of the Kolmogorov entropy from a chaotic signal. Phys. Rev. A 1983, 28, 2591. [Google Scholar] [CrossRef]
- Pincus, S. Approximate entropy as a measure of system complexity. Proc. Natl. Acad. Sci. USA 1991, 88, 2297–2301. [Google Scholar] [CrossRef] [PubMed]
- Richman, J.; Moorman, J. Physiological time-series analysis using approximate entropy and sample entropy. Am. J. Physiol. Heart Circ. Physiol. 2000, 278, H2039–H2049. [Google Scholar] [PubMed]
- Costa, M.; Goldberger, A.; Peng, C. Multiscale entropy analysis of complex physiologic time series. Phys. Rev. Lett. 2002, 89, 068102. [Google Scholar] [CrossRef] [PubMed]
- Labate, D.; La Foresta, F.; Morabito, G.; Palamara, I.; Morabito, F.C. Entropic measures of EEG complexity in Alzheimer’s disease through a multivariate multiscale approach. IEEE Sens. J. 2013, 13, 3284–3292. [Google Scholar] [CrossRef]
- Labate, D.; Palamara, I.; Mammone, N.; Morabito, G.; La Foresta, F.; Morabito, F.C. SVM classification of epileptic EEG recordings through multiscale permutation entropy. In Proceedings of the International Joint Conference on Neural Networks (IJCNN), Dallas, TX, USA, 4–9 August 2013; pp. 1–5. [Google Scholar]
- Valencia, J.; Porta, A.; Vallverdú, M.; Claria, F.; Baranowski, R.; Orlowska-Baranowska, E.; Caminal, P. Refined multiscale entropy: Application to 24-h holter recordings of heart period variability in healthy and aortic stenosis subjects. IEEE Trans. Biomed. Eng. 2009, 56, 2202–2213. [Google Scholar] [CrossRef] [PubMed]
- Amoud, H.; Snoussi, H.; Hewson, D.; Doussot, M.; Duchêne, J. Intrinsic mode entropy for nonlinear discriminant analysis. IEEE Signal Process. Lett. 2007, 14, 297–300. [Google Scholar] [CrossRef]
- Pachori, R.B.; Hewson, D.; Snoussi, H.; Duchêne, J. Postural time-series analysis using empirical mode decomposition and second-order difference plots. In Proceedings of the 2009 IEEE International Conference on Acoustics, Speech and Signal Processing, Taipei, Taiwan, 19–24 April 2009; pp. 537–540. [Google Scholar]
- Bayram, I. An analytic wavelet transform with a flexible time-frequency covering. IEEE Trans. Signal Process. 2013, 61, 1131–1142. [Google Scholar] [CrossRef]
- Kumar, M.; Pachori, R.B.; Acharya, U.R. Use of accumulated entropies for automated detection of congestive heart failure in flexible analytic wavelet transform framework based on short-term HRV signals. Entropy 2017, 19, 92. [Google Scholar] [CrossRef]
- Peng, Z.; Peter, W.T.; Chu, F. A comparison study of improved Hilbert–Huang transform and wavelet transform: Application to fault diagnosis for rolling bearing. Mech. Syst. Signal Process. 2005, 19, 974–988. [Google Scholar] [CrossRef]
- Gupta, A.; Joshi, S. On the concept of intrinsic wavelet functions. In Proceedings of the International Conference on Signal Processing and Communications (SPCOM), Bangalore, India, 22–25 July 2014; pp. 1–5. [Google Scholar]
- Selesnick, I. Wavelet transform with tunable Q-factor. IEEE Trans. Signal Process. 2011, 59, 3560–3575. [Google Scholar] [CrossRef]
- Bhattacharyya, A.; Pachori, R.B.; Acharya, U.R. Tunable-Q wavelet transform based multivariate sub-band fuzzy entropy with application to focal EEG signal analysis. Entropy 2017, 19, 99. [Google Scholar] [CrossRef]
- Veselkov, K.A.; Pahomov, V.I.; Lindon, J.C.; Volynkin, V.S.; Crockford, D.; Osipenko, G.S.; Davies, D.B.; Barton, R.H.; Bang, J.W.; Holmes, E.; et al. A metabolic entropy approach for measurements of systemic metabolic disruptions in patho-physiological states. J. Proteome Res. 2010, 9, 3537–3544. [Google Scholar] [CrossRef] [PubMed]
- Kraskov, A.; Stögbauer, H.; Grassberger, P. Estimating mutual information. Phys. Rev. E 2004, 69, 066138. [Google Scholar] [CrossRef] [PubMed]
- Andrzejak, R.; Lehnertz, K.; Mormann, F.; Rieke, C.; David, P.; Elger, C. Indications of nonlinear deterministic and finite-dimensional structures in time series of brain electrical activity: Dependence on recording region and brain state. Phys. Rev. E 2001, 64, 061907. [Google Scholar] [CrossRef] [PubMed]
- Kohavi, R.; John, G.H. Wrappers for feature subset selection. Artif. Intell. 1997, 97, 273–324. [Google Scholar] [CrossRef]
- Cortes, C.; Vapnik, V. Support-vector networks. Mach. Learn. 1995, 20, 273–297. [Google Scholar] [CrossRef]
- Bhattacharyya, A.; Sharma, M.; Pachori, R.B.; Sircar, P.; Acharya, U.R. A novel approach for automated detection of focal EEG signals using empirical wavelet transform. Neural Comput. Appl. 2016, in press. [Google Scholar] [CrossRef]
- Bhattacharyya, A.; Pachori, R.B. A multivariate approach for patient specific EEG seizure detection using empirical wavelet transform. IEEE Trans. Biomed. Eng. 2017, in press. [Google Scholar] [CrossRef] [PubMed]
- Kohavi, R. A Study of Cross-Validation and Bootstrap for Accuracy Estimation And Model Selection; International Joint Conference on Artificial Intelligence: Montreal, QC, Canada, 1995; Volume 14, pp. 1137–1145. [Google Scholar]
- Hall, M.; Frank, E.; Holmes, G.; Pfahringer, B.; Reutemann, P.; Witten, I.H. The WEKA data mining software: An update. ACM SIGKDD Explor. Newsl. 2009, 11, 10–18. [Google Scholar] [CrossRef]
- Tzallas, A.; Tsipouras, M.; Fotiadis, D. Automatic seizure detection based on time-frequency analysis and artificial neural networks. Comput. Intell. Neurosci. 2007, 2007, 80510. [Google Scholar] [CrossRef] [PubMed]
- Peker, M.; Sen, B.; Delen, D. A novel method for automated diagnosis of epilepsy using complex-valued classifiers. IEEE J. Biomed. Health Inform. 2016, 20, 108–118. [Google Scholar] [CrossRef] [PubMed]
- Chen, G. Automatic EEG seizure detection using dual-tree complex wavelet-Fourier features. Expert Syst. Appl. 2014, 41, 2391–2394. [Google Scholar] [CrossRef]
- Orhan, U.; Hekim, M.; Ozer, M. EEG signals classification using the K-means clustering and a multilayer perceptron neural network model. Expert Syst. Appl. 2011, 38, 13475–13481. [Google Scholar] [CrossRef]
- Samiee, K.; Kovács, P.; Gabbouj, M. Epileptic seizure classification of EEG time-series using rational discrete short-time Fourier transform. IEEE Trans. Biomed. Eng. 2015, 62, 541–552. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Rivero, D.; Dorado, J.; Rabunal, J.R.; Pazos, A. Automatic epileptic seizure detection in EEGs based on line length feature and artificial neural networks. J. Neurosci. Methods 2010, 191, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Acharya, U.R.; Molinari, F.; Sree, S.V.; Chattopadhyay, S.; Ng, K.H.; Suri, J.S. Automated diagnosis of epileptic EEG using entropies. Biomed. Signal Process. Control 2012, 7, 401–408. [Google Scholar] [CrossRef]
- Yuan, Q.; Zhou, W.; Li, S.; Cai, D. Epileptic EEG classification based on extreme learning machine and nonlinear features. Epilepsy Res. 2011, 96, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Al Ghayab, H.R.; Li, Y.; Abdulla, S.; Diykh, M.; Wan, X. Classification of epileptic EEG signals based on simple random sampling and sequential feature selection. Brain Inform. 2016, 3, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Bhati, D.; Sharma, M.; Pachori, R.B.; Gadre, V.M. Time–frequency localized three-band biorthogonal wavelet filter bank using semidefinite relaxation and nonlinear least squares with epileptic seizure EEG signal classification. Digit. Signal Process. 2017, 62, 259–273. [Google Scholar] [CrossRef]
- Ahirwal, M.K.; Kumar, A.; Singh, G.K. Analysis and testing of PSO variants through application in EEG/ERP adaptive filtering approach. Biomed. Eng. Lett. 2012, 2, 186–197. [Google Scholar] [CrossRef]
| Dataset | Type of Recording | Subjects | Total Number of Signals | Predetermined Class |
|---|---|---|---|---|
| S | Intracranial | 5 patients | 100 | Seizure |
| F | Intracranial (Epileptogenic zone) | 5 patients | 100 | Seizure-free |
| N | Intracranial (Hippocampal formation opposite hemisphere of the brain) | 5 patients | 100 | Seizure-free |
| Z | Surface (with eyes open) | 5 healthy | 100 | Normal |
| volunteers | ||||
| O | Surface (with eyes closed) | 5 healthy | 100 | Normal |
| volunteers |
| Scale No. | ||||||
|---|---|---|---|---|---|---|
| Seizure | Seizure-Free | Normal | Seizure | Seizure-Free | Normal | |
| 1 | ||||||
| 2 | ||||||
| 3 | ||||||
| 4 | ||||||
| 5 | ||||||
| 6 | ||||||
| 7 | ||||||
| 8 | ||||||
| Feature | Experiment Type | Acc (%) | Sens (%) | Spec (%) |
|---|---|---|---|---|
| K-NN entropy computed | S-Z | 100 | 100 | 100 |
| S-O | 96.5 | 93 | 100 | |
| S-N | 98.5 | 100 | 97 | |
| S-F | 93 | 97 | 89 | |
| S-FNZO | 96.4 | 91 | 97.8 | |
| S-FN-ZO | 64.8 | 93 | 96.33 | |
| 47 | 79.67 | |||
| 68.5 | 65.33 |
| Filtering Type | Parameters (Q and R) | Experiment Type | Number of Levels | Number of Levels | ||||
|---|---|---|---|---|---|---|---|---|
| Acc (%) | Sens (%) | Spec (%) | Acc (%) | Sens (%) | Spec (%) | |||
| Approximation to detail | ; | S-Z | 100 | 100 | 100 | 100 | 100 | 100 |
| S-O | 99 | 99 | 99 | 99 | 99 | 99 | ||
| S-N | 98.5 | 99 | 98 | 99 | 100 | 98 | ||
| S-F | 95 | 96 | 94 | 97.5 | 97 | 98 | ||
| S-FNZO | 98 | 96 | 98.5 | 99 | 96 | 99.8 | ||
| S-FN-ZO | 93.4 | 95 | 98.25 | 95.6 | 95 | 99.25 | ||
| 94 | 93.33 | 96.5 | 95 | |||||
| 92 | 98 | 95 | 98.67 | |||||
| ; | S-Z | 100 | 100 | 100 | 100 | 100 | 100 | |
| S-O | 99 | 99 | 99 | 99.5 | 100 | 99 | ||
| S-N | 98.5 | 100 | 97 | 99 | 99 | 99 | ||
| S-F | 92.5 | 96 | 89 | 96 | 96 | 96 | ||
| S-FNZO | 97 | 92 | 98.3 | 98.2 | 96 | 98.8 | ||
| S-FN-ZO | 93.4 | 95 | 97.25 | 95.8 | 97 | 98.5 | ||
| 91 | 95 | 95.5 | 96 | |||||
| 95 | 97.67 | 95.5 | 99 | |||||
| ; | S-Z | 100 | 100 | 100 | 100 | 100 | 100 | |
| S-O | 96.5 | 95 | 98 | 99.5 | 99 | 100 | ||
| S-N | 98 | 99 | 97 | 98 | 99 | 97 | ||
| S-F | 92.5 | 96 | 89 | 96 | 98 | 94 | ||
| S-FNZO | 97 | 93 | 98 | 97.8 | 97 | 98 | ||
| S-FN-ZO | 74.6 | 93 | 97.25 | 94.2 | 95 | 98.25 | ||
| 77.5 | 76.33 | 93.5 | 94.67 | |||||
| 62.5 | 85 | 94.5 | 98 | |||||
| Detail to approximation | ; | S-Z | 100 | 100 | 100 | 100 | 100 | 100 |
| S-O | 99.5 | 99 | 100 | 99.5 | 99 | 100 | ||
| S-N | 98.5 | 98 | 99 | 99 | 99 | 99 | ||
| S-F | 97.5 | 97 | 98 | 97.5 | 97 | 98 | ||
| S-FNZO | 98.4 | 95 | 99.3 | 98.8 | 95 | 99.8 | ||
| S-FN-ZO | 97.2 | 95 | 99.25 | 97.2 | 95 | 99 | ||
| 95.5 | 98.66 | 95.5 | 98.67 | |||||
| 100 | 97.66 | 100 | 98 | |||||
| ; | S-Z | 98.5 | 98 | 99 | 100 | 100 | 100 | |
| S-O | 95.5 | 94 | 97 | 100 | 100 | 100 | ||
| S-N | 99.5 | 100 | 99 | 99.5 | 99 | 100 | ||
| S-F | 97 | 96 | 98 | 97.5 | 97 | 98 | ||
| S-FNZO | 98.2 | 92 | 99.8 | 98.8 | 96 | 99.5 | ||
| S-FN-ZO | 92.2 | 94 | 99.75 | 98.6 | 96 | 99.75 | ||
| 86.5 | 98.33 | 98.5 | 98.67 | |||||
| 97 | 89 | 100 | 99.33 | |||||
| ; | S-Z | 97 | 95 | 99 | 98.5 | 98 | 99 | |
| S-O | 96 | 92 | 100 | 97 | 94 | 100 | ||
| S-N | 99.5 | 99 | 100 | 99.5 | 99 | 100 | ||
| S-F | 97 | 96 | 98 | 98 | 98 | 98 | ||
| S-FNZO | 98.2 | 91 | 100 | 98.8 | 96 | 99.5 | ||
| S-FN-ZO | 92.2 | 89 | 100 | 97.4 | 96 | 99.5 | ||
| 88.5 | 98.33 | 96 | 99 | |||||
| 97.5 | 88.66 | 99.5 | 97.33 | |||||
| Authors | Method | Training and Testing (Data Selection) | Experiment Type | Accuracy (%) |
|---|---|---|---|---|
| Tzallas et al. [54] (2007) | Time-frequency analysis and artificial neural network | 50% training and 50% testing | S-Z | 100 |
| S-FNZO | 97.73 | |||
| S-FN-ZO | 97.72 | |||
| Tiwari et al. [16] (2016) | Key-point-based LBP and SVM | 10-fold cross-validation | S-FNZO | 99.31 |
| S-FN-ZO | 98.80 | |||
| Peker et al. [55] (2016) | Dual tree complex wavelet transform (DTCWT) and complex valued neural networks | 10-fold cross-validation | S-Z | 100 |
| S-FNZO | 99.15 | |||
| S-FN-ZO | 98.28 | |||
| Chen [56] (2014) | DTCWT and Fourier features with nearest neighbor classifier | First half of the signals for training and the rest for testing | S-Z | 100 |
| S-FNZO | 100 | |||
| Orhan et al. [57] (2011) | K-means clustering and multilayer perceptron (MLP) neural network model | Randomly selected | S-Z | 100 |
| S-FNZO | 99.60 | |||
| S-FN-ZO | 95.60 | |||
| Samiee et al. [58] (2015) | Rational discrete STFT and MLP classifier | Randomly selected 50% data for training | S-Z | 99.80 |
| S-O | 99.30 | |||
| S-N | 98.50 | |||
| S-F | 94.90 | |||
| S-FNZO | 98.100 | |||
| Bajaj and Pachori [3] (2012) | Amplitude and frequency modulation bandwidths of IMFs and least-squares SVM (LS-SVM) | 10-fold cross-validation | S-FNZO | 99.50-100 |
| Guo et al. [59] (2010) | Line length feature and artificial neural networks | Randomly selected 50% data for training | S-Z | 99.6 |
| S-FNZO | 97.77 | |||
| Kaya et al. [14] (2014) | 1D LBP and functional tree 1D LBP and BayesNet | 10-fold cross-validation | S-Z | 99.50 |
| S-F | 95.50 | |||
| Acharya et al. [11] (2009) | RQA features, SVM classifier | 3-fold cross-validation | S-FN-ZO | 95.6 |
| Acharya et al. [60] (2012) | ApEn, SEn, phase entropy features, and fuzzy classifier | 3-fold cross-validation | S-FN-ZO | 98.1 |
| Yuan et al. [61] (2011) | ApEn, Hurst exponent, scaling exponents of EEG, and extreme learning machine (ELM) algorithm | 50% data for training | S-F | 96.5 |
| Ghayab et al. [62] (2016) | Simple random sampling, sequential feature selection and LS-SVM | Not specified | S-Z | 99.9 |
| Our work | TQWT-based multi-scale K-NN entropy | 10-fold cross-validation | S-Z | 100 |
| S-O | 100 | |||
| S-N | 99.50 | |||
| S-F | 98 | |||
| S-FNZO | 99 | |||
| S-FN-ZO | 98.60 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bhattacharyya, A.; Pachori, R.B.; Upadhyay, A.; Acharya, U.R. Tunable-Q Wavelet Transform Based Multiscale Entropy Measure for Automated Classification of Epileptic EEG Signals. Appl. Sci. 2017, 7, 385. https://doi.org/10.3390/app7040385
Bhattacharyya A, Pachori RB, Upadhyay A, Acharya UR. Tunable-Q Wavelet Transform Based Multiscale Entropy Measure for Automated Classification of Epileptic EEG Signals. Applied Sciences. 2017; 7(4):385. https://doi.org/10.3390/app7040385
Chicago/Turabian StyleBhattacharyya, Abhijit, Ram Bilas Pachori, Abhay Upadhyay, and U. Rajendra Acharya. 2017. "Tunable-Q Wavelet Transform Based Multiscale Entropy Measure for Automated Classification of Epileptic EEG Signals" Applied Sciences 7, no. 4: 385. https://doi.org/10.3390/app7040385
APA StyleBhattacharyya, A., Pachori, R. B., Upadhyay, A., & Acharya, U. R. (2017). Tunable-Q Wavelet Transform Based Multiscale Entropy Measure for Automated Classification of Epileptic EEG Signals. Applied Sciences, 7(4), 385. https://doi.org/10.3390/app7040385






