Comprehensive Analysis of Immune Response and Transcriptome Profiling Reveals the Molecular Basis Underlying Breed-Specific Responses to Eimeria tenella Infection in Chickens with Differing Susceptibility Levels
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals and Parasites
2.2. Experimental Design and Sample Collection
2.3. Peripheral Blood Lymphocyte Isolation and Flow Cytometric Analysis
2.4. ELISA Analysis of the Serum Antibody
2.5. Total RNA Isolation, cDNA Library Construction, and Illumina Sequencing
2.6. Differentially Expressed Genes (DEGs) and Functional Enrichment Analysis
2.7. Construction of Gene–Gene Interaction Networks
2.8. Analysis of the Overlapping Genes at Three Time Points
2.9. Validation of the RNA−seq Data via qRT−PCR
2.10. Data Statistical Analysis
3. Results
3.1. Different Susceptibility to E. tenella Infection Between WCCs and RWFCs
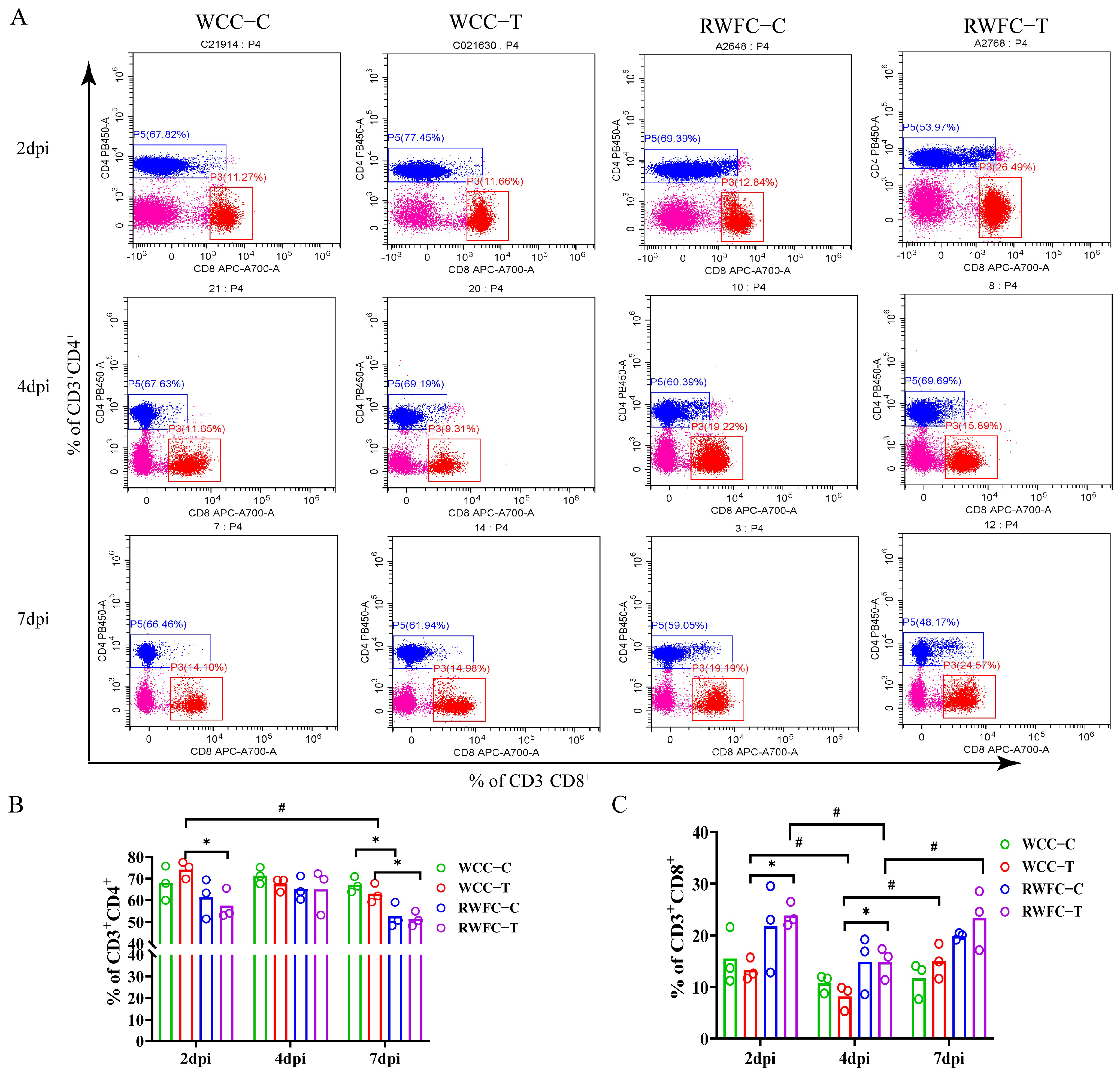
3.2. E. tenella-Induced Alterations of CD3+CD4+ and CD3+CD8α+ T lymphocyte Subsets in WCCs and RWFCs
3.3. E. tenella-Induced Alterations of Serum IgG and IgA in WCCs and RWFCs
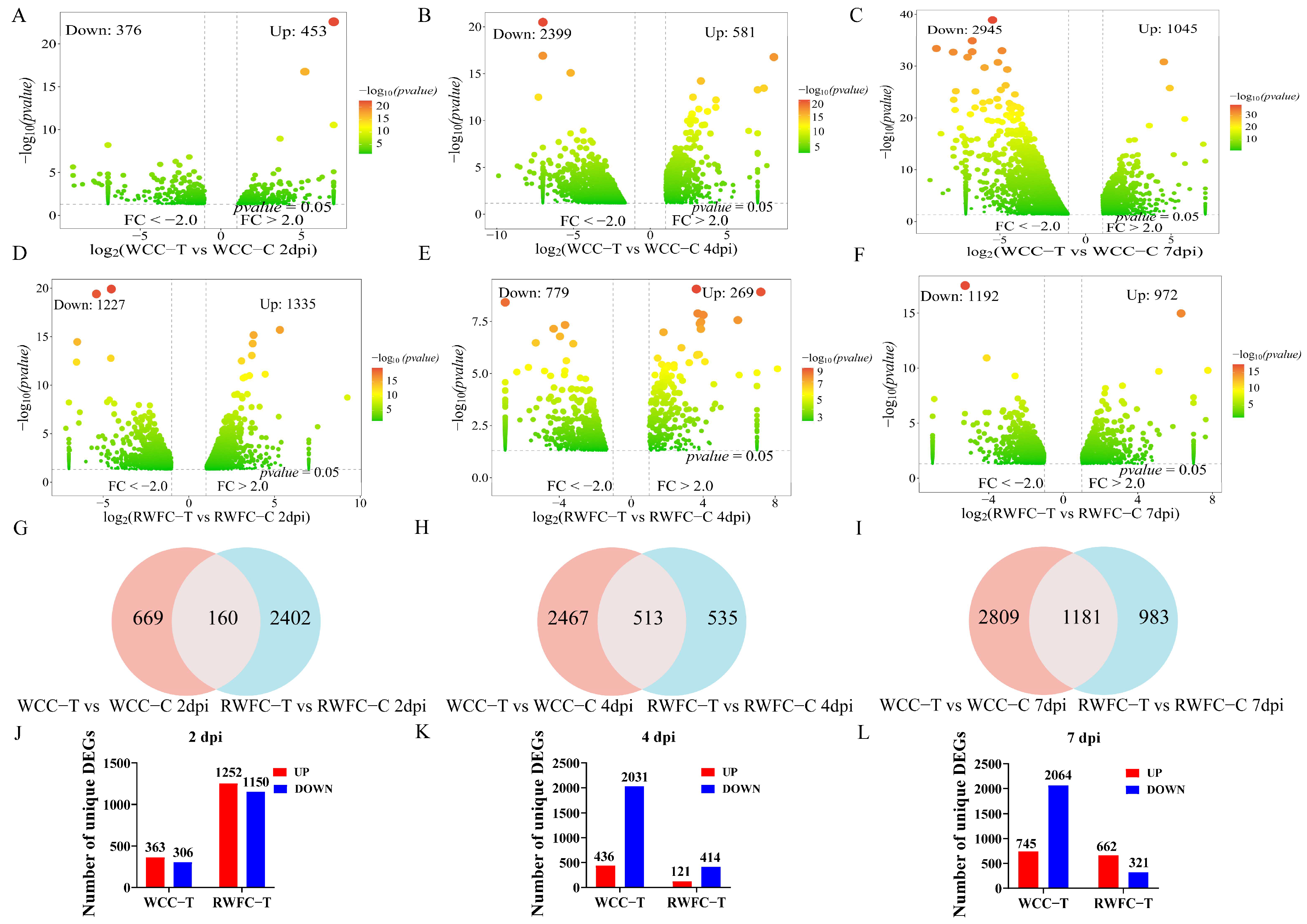
3.4. RNA−seq Profiles of WCC and RWFC Cecum Tissues Following E. tenella Infection
3.5. Analysis of Gene Expression in Cecal Tissues of WCCs and RWFCs Following E. tenella Infection
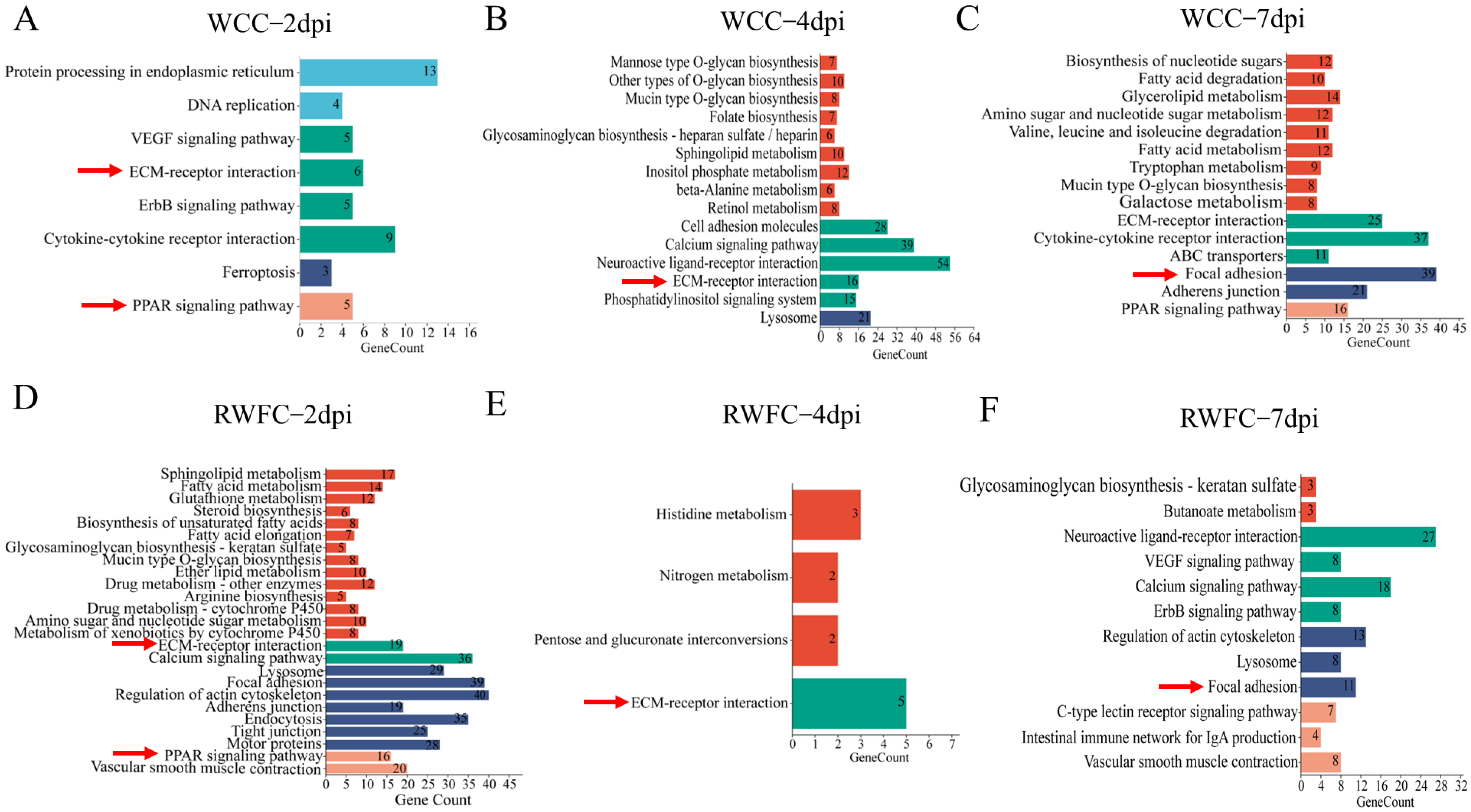
3.6. Functional Annotation of the Unique DEGs in WCCs and RWFCs Following E. tenella Infection
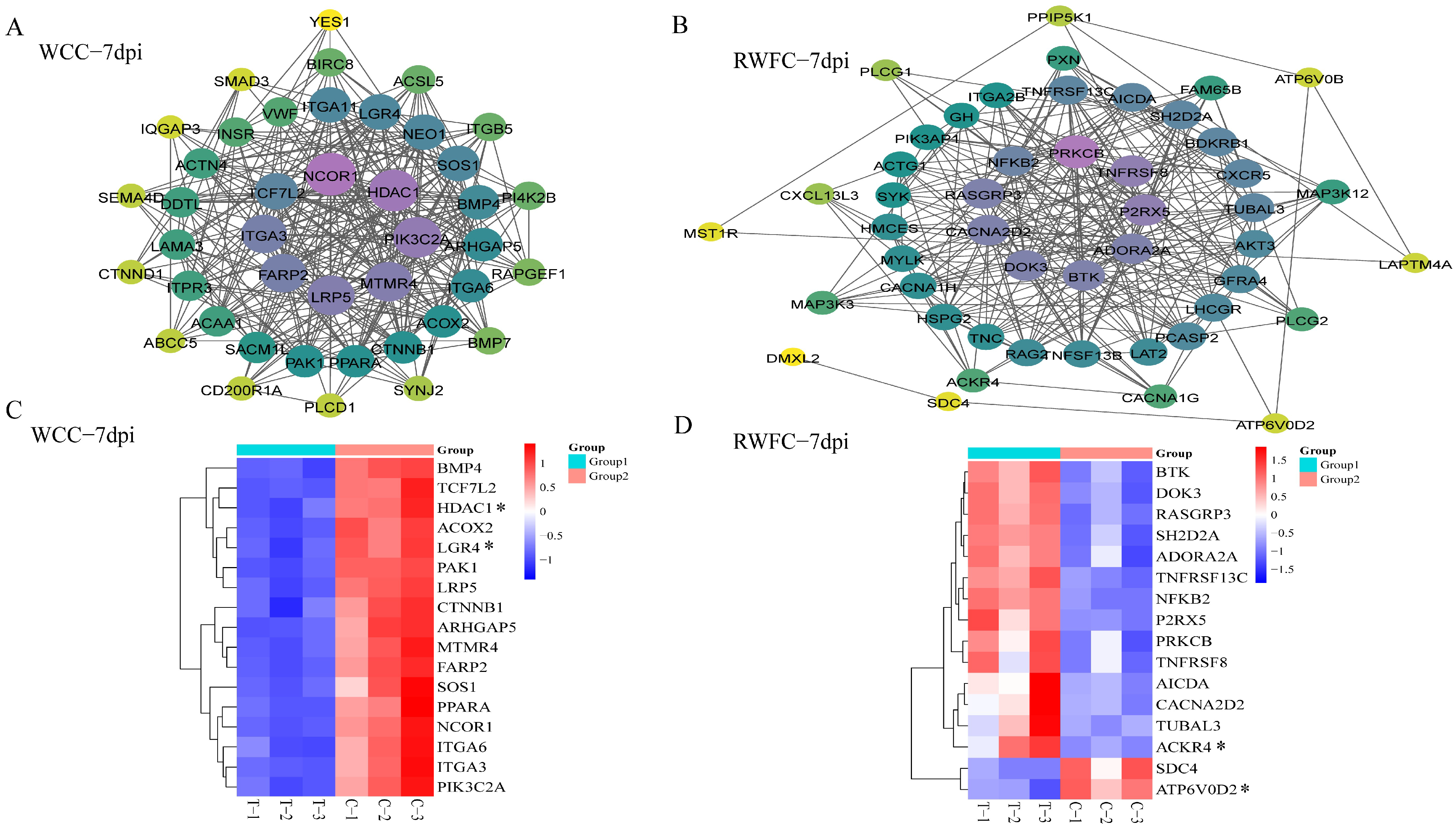
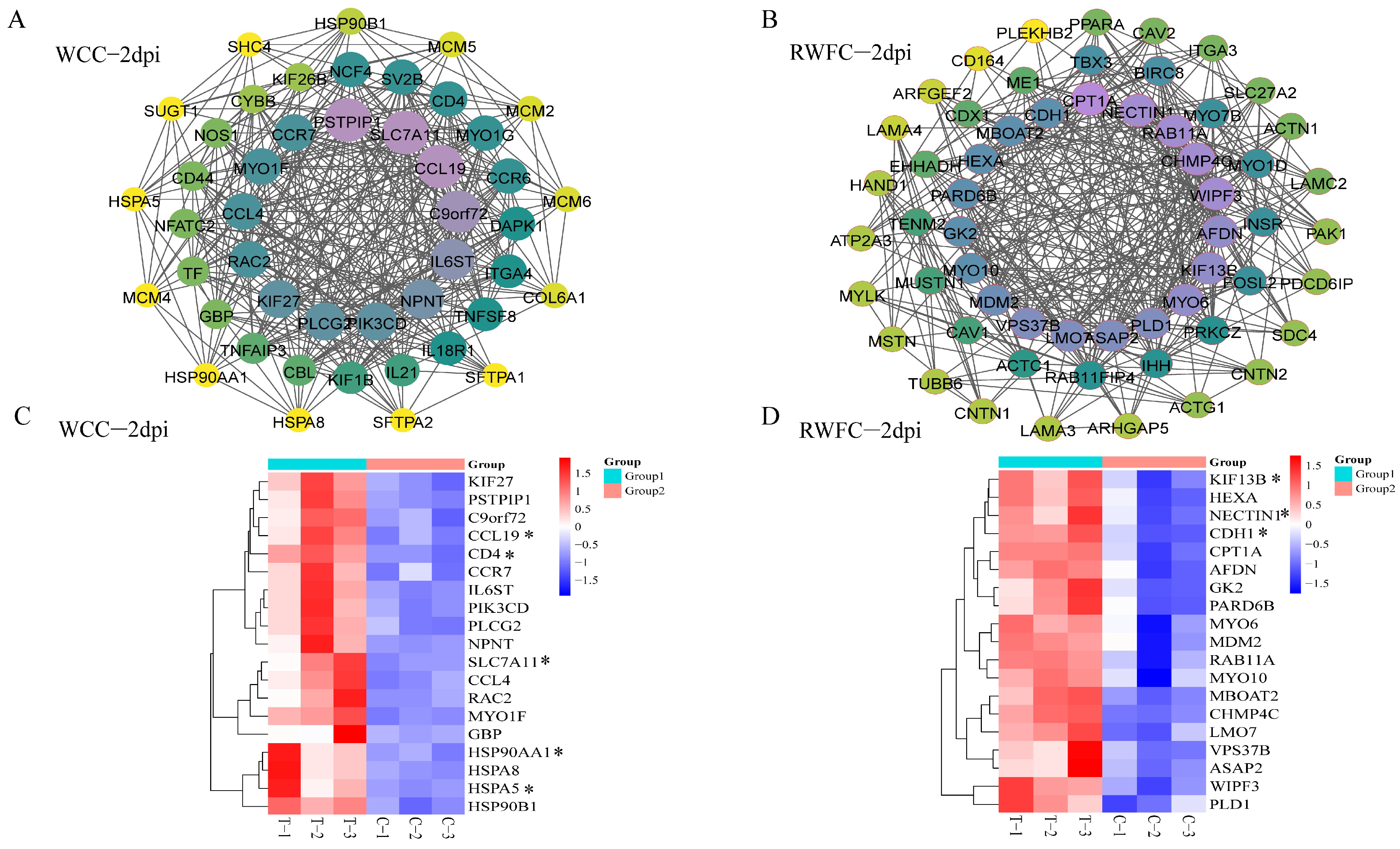
3.7. Gene–Gene Interaction Network of WCC–Specific and RWFC–Specific DEGs Following E. tenella Infection
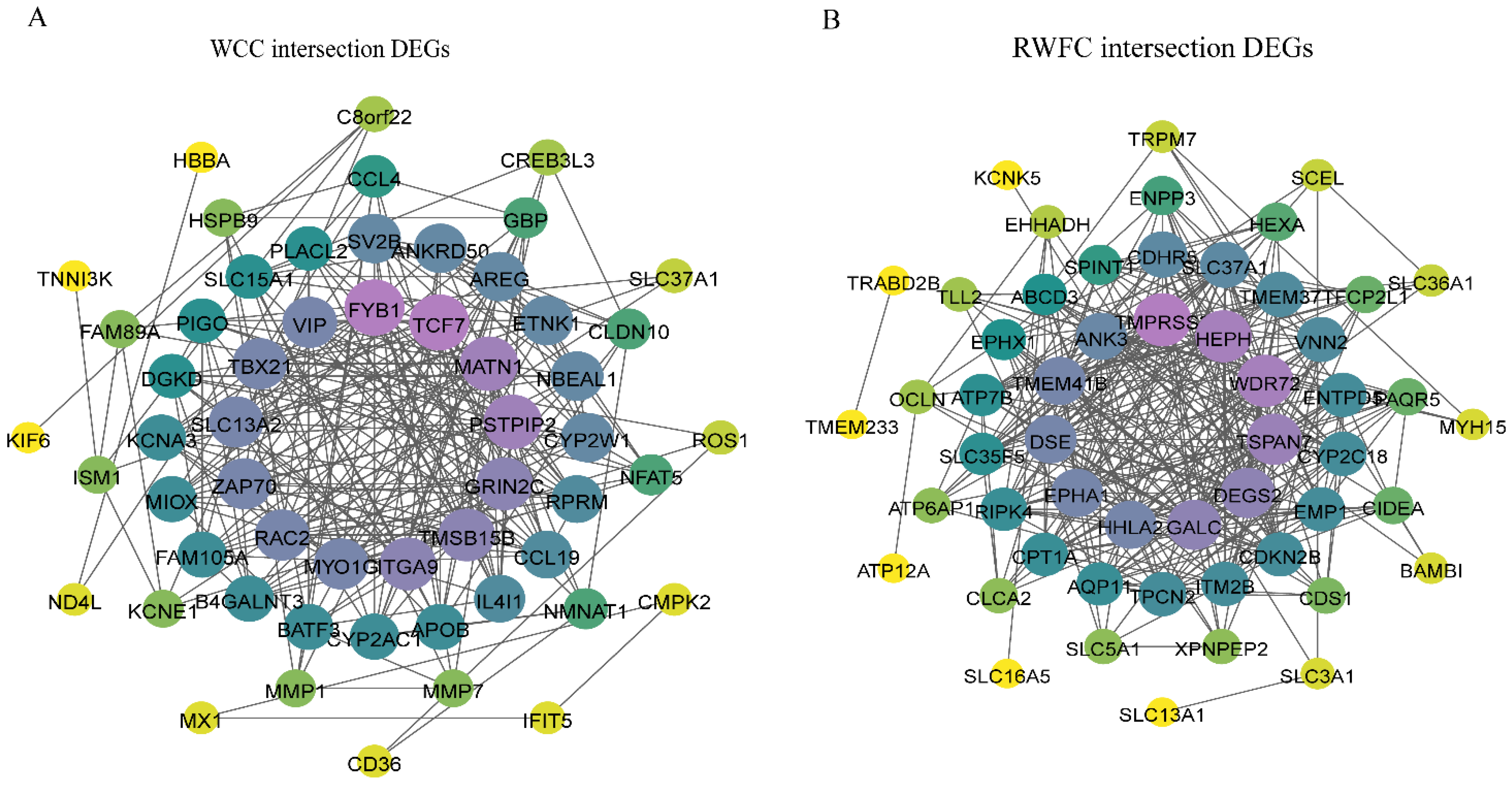
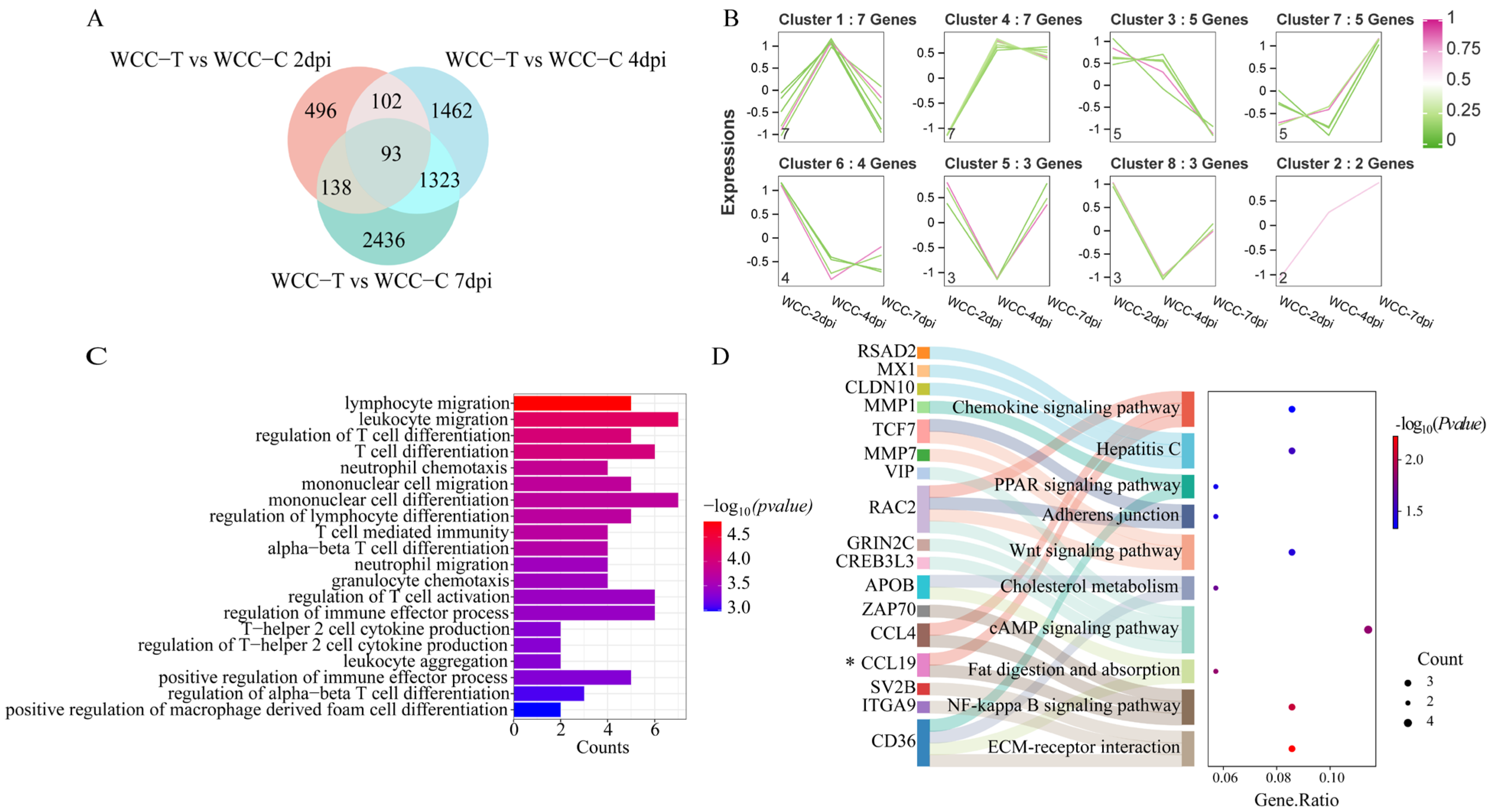
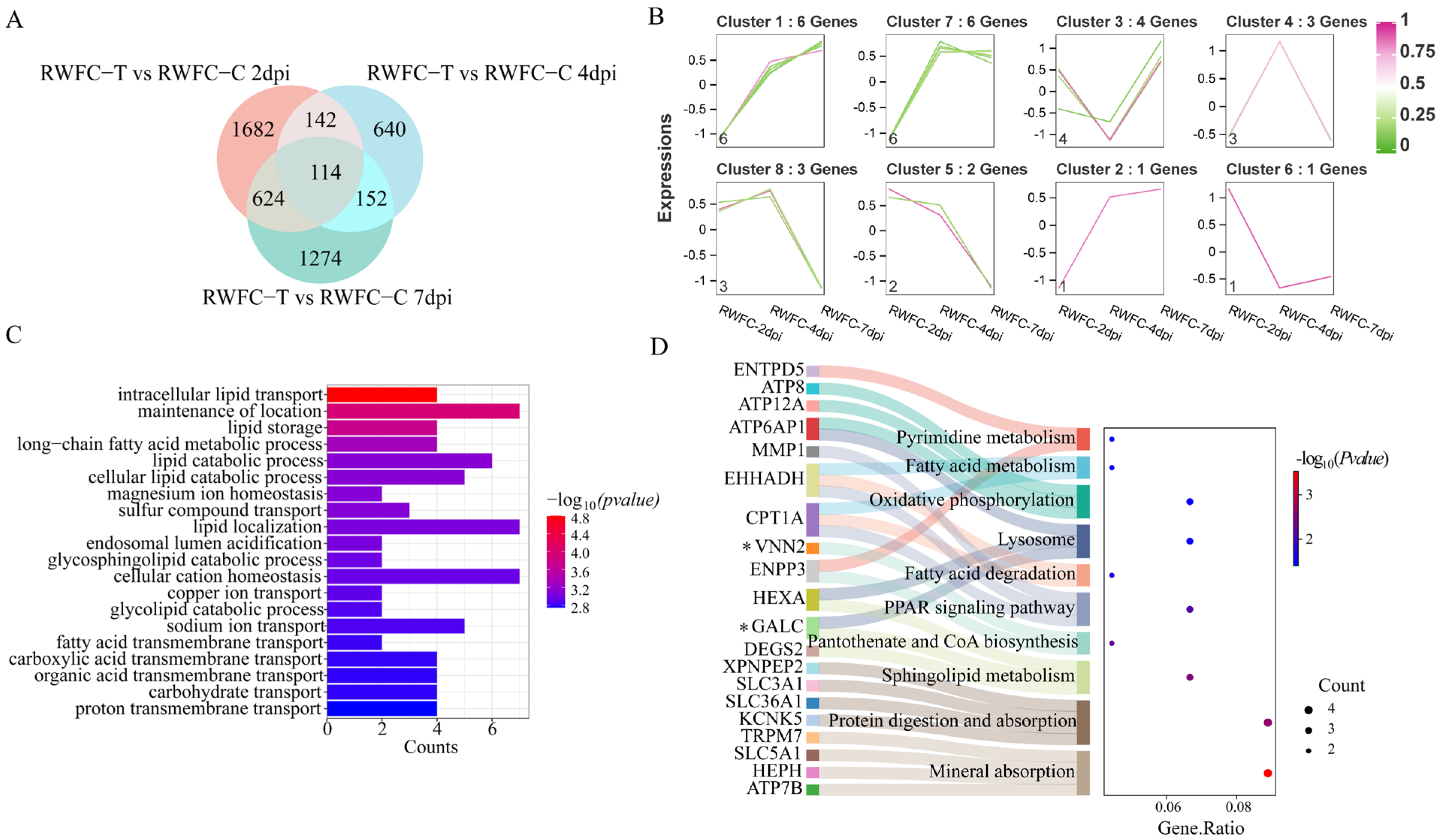
3.8. Analysis of Overlapping DEGs in the Cecum of WCCs and RWFCs Following E. tenella Infection
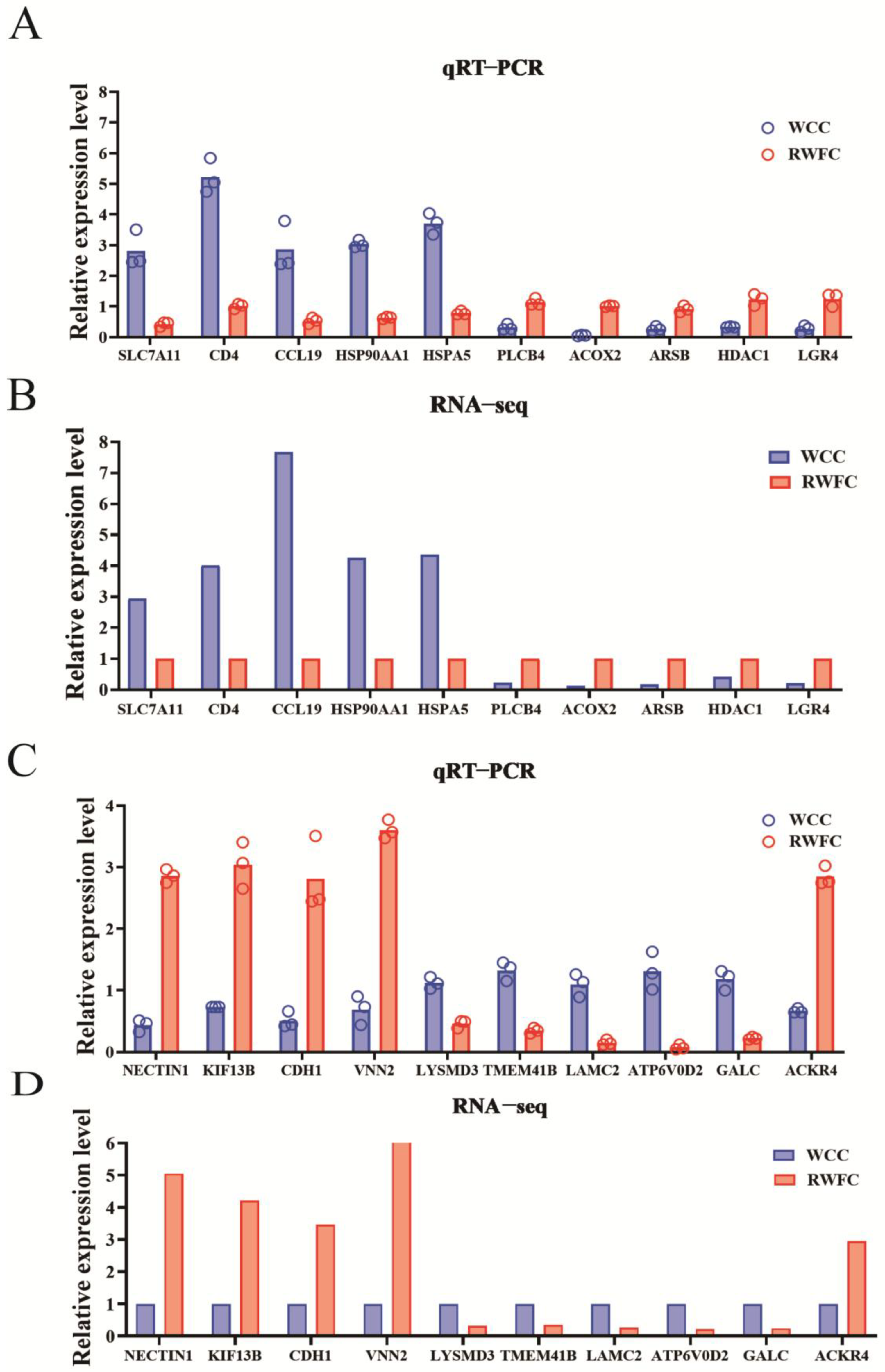
3.9. Validation of Some Unique DEGs in the RNA−sequencing Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| WCCs | Wenchang chickens |
| RWFCs | Recessive white feather chickens |
| DEGs | Differentially expressed genes |
| PBS | Phosphate-buffered saline |
| MR | Mortality rate |
| BWG | Body weight gain |
| OPG | Oocyst output per gram of feces |
| CLS | Cecal lesion score |
| dpi | Days post-infection |
| RIN | RNA integrity number |
| GO | Gene ontology |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| FDR | False discovery rate |
| qRT−PCR | Quantitative reverse transcription polymerase chain reaction |
| MF | Molecular function |
| CC | Cellular component |
| BP | Biological process |
| Th | T helper cell |
| CTL | Cytotoxic T lymphocyte |
| ER | Endoplasmic reticulum |
| HSPs | Heat shock proteins |
| SCFAs | Short-chain fatty acids |
| LCFAs | Long-chain fatty acids |
Appendix A
| No. | Sample | Sample No. | Residual Volume (μL) | Nanomolar Concentration (ng/μL) | Total Mass (μg) | OD260/280 | OD260/230 | RIN Value | Quality Inspection Conclusion |
|---|---|---|---|---|---|---|---|---|---|
| 1 | A4244 | JB22ZNO00396 | 155.0 | 308.6 | 47.83 | 1.89 | 2.37 | 8.9 | A |
| 2 | A4229 | JB22ZNO00397 | 155.0 | 541.8 | 83.98 | 1.93 | 2.36 | 7.3 | A |
| 3 | A3456 | JB22ZNO00398 | 155.0 | 493.2 | 76.45 | 1.94 | 2.34 | 7.4 | A |
| 4 | A3554 | JB22ZNO00400 | 155.0 | 544.6 | 84.41 | 1.92 | 2.34 | 10.0 | A |
| 5 | A2768 | JB22ZNO00401 | 155.0 | 442.2 | 68.54 | 1.92 | 2.35 | 10.0 | A |
| 6 | A3465 | JB22ZNO00402 | 155.0 | 428.7 | 66.45 | 1.92 | 2.34 | 10.0 | A |
| 7 | C021934 | JB22ZNO00404 | 155.0 | 357.4 | 55.4 | 1.91 | 2.35 | 8.1 | A |
| 8 | C021932 | JB22ZNO00405 | 155.0 | 396.5 | 61.46 | 1.92 | 2.31 | 10.0 | A |
| 9 | C021450 | JB22ZNO00406 | 155.0 | 392.5 | 60.84 | 1.91 | 2.34 | 7.2 | A |
| 10 | C021867 | JB22ZNO00408 | 155.0 | 507.1 | 78.6 | 1.91 | 2.28 | 9.6 | A |
| 11 | C022079 | JB22ZNO00409 | 155.0 | 422.0 | 65.41 | 1.92 | 2.32 | 7.6 | A |
| 12 | C021880 | JB22ZNO00410 | 155.0 | 997.2 | 154.57 | 1.98 | 2.33 | 10.0 | A |
| 13 | A4517 | JB22ZNO00421 | 155.0 | 658.4 | 102.05 | 1.96 | 2.33 | 7.8 | A |
| 14 | A4143 | JB22ZNO00422 | 155.0 | 584.8 | 90.64 | 1.95 | 2.36 | 9.9 | A |
| 15 | A4466 | JB22ZNO00454 | 300.0 | 473.2 | 141.96 | 1.96 | 2.40 | 7.2 | A |
| 16 | A2769 | JB22ZNO00425 | 200.0 | 564.5 | 112.9 | 1.91 | 2.37 | 8.1 | A |
| 17 | A4263 | JB22ZNO00426 | 155.0 | 704.1 | 109.14 | 1.97 | 2.40 | 7.3 | A |
| 18 | A4392 | JB22ZNO00427 | 155.0 | 949.0 | 147.1 | 1.97 | 2.33 | 8.9 | A |
| 19 | C021451 | JB22ZNO00429 | 60.0 | 459.11 | 27.55 | 2.07 | 1.98 | 9.9 | A |
| 20 | C021763 | JB22ZNO00430 | 60.0 | 345.01 | 20.7 | 2.05 | 2.16 | 10.0 | A |
| 21 | C021137 | JB22ZNO00431 | 60.0 | 195.18 | 11.71 | 2.02 | 2.20 | 7.7 | A |
| 22 | C021583 | JB22ZNO00437 | 300.0 | 593.7 | 178.11 | 1.98 | 2.39 | 7.9 | A |
| 23 | C021411 | JB22ZNO00435 | 60.0 | 635.4 | 38.12 | 2.06 | 2.38 | 9.0 | A |
| 24 | C022061 | JB22ZNO00436 | 60.0 | 361.84 | 21.71 | 2.06 | 2.30 | 8.5 | A |
| 25 | A4533 | JB22ZNO00447 | 300.0 | 393.0 | 117.9 | 1.93 | 2.23 | 10.0 | A |
| 26 | A4004 | JB22ZNO00448 | 300.0 | 377.8 | 113.34 | 1.90 | 2.37 | 10.0 | A |
| 27 | A3532 | JB22ZNO00449 | 300.0 | 559.6 | 167.88 | 1.96 | 2.21 | 8.7 | A |
| 28 | A4464 | JB22ZNO00423 | 155.0 | 652.7 | 101.17 | 1.95 | 2.35 | 9.6 | A |
| 29 | A4307 | JB22ZNO00452 | 300.0 | 379.9 | 113.97 | 1.91 | 2.37 | 7.4 | A |
| 30 | A3573 | JB22ZNO00455 | 300.0 | 491.9 | 147.57 | 1.97 | 2.38 | 7.5 | A |
| 31 | C021881 | JB22ZNO00456 | 300.0 | 289.5 | 86.85 | 1.87 | 2.21 | 9.8 | A |
| 32 | C022320 | JB22ZNO00457 | 300.0 | 403.1 | 120.93 | 1.91 | 2.35 | 8.5 | A |
| 33 | C021885 | JB22ZNO00458 | 300.0 | 518.3 | 155.49 | 1.94 | 2.40 | 9.9 | A |
| 34 | C021978 | JB22ZNO00460 | 250.0 | 455.9 | 113.98 | 1.91 | 2.25 | 9.2 | A |
| 35 | C021876 | JB22ZNO00461 | 300.0 | 397.4 | 119.22 | 1.91 | 2.33 | 9.0 | A |
| 36 | C021908 | JB22ZNO00462 | 300.0 | 593.3 | 177.99 | 1.96 | 2.30 | 8.2 | A |
| Primer Name | Primer Sequence (5′-3′) | Product Length (bp) | GenBank Accession No. |
|---|---|---|---|
| SLC7A11 | TCGTGACGGTGCCTAATG | 175 | MH760782.1 |
| TTGGGTGAGATGAAGATGC | |||
| CD4 | TCCCTTTGTCCTGTTTCG | 99 | Y12012.1 |
| TCTTGTCCATTGGCTCCT | |||
| CCL19 | TATGCTGGCAACAACGTCCT | 135 | NM001302168.1 |
| TGCAGTGATGAACACGGTGG | |||
| HSP90AA1 | CGCAGCGTTGAGGGTGTA | 85 | NM001109785.2 |
| CACAGCTTCTGGCATCTTGG | |||
| HSPA5 | CCATTAAGGTCTATGAAGGTGAACG | 224 | NM205491.2 |
| CTCTGGTGTTAGCCGATTCT | |||
| PLCB4 | ATTCAAGCCATTAAGGAAAC | 193 | NM_001398213.1 |
| CATTAGGAGAAGGCAAGG | |||
| ACOX2 | CCTCCTGGGGATCATTCGG | 165 | XM046926551.1 |
| TAGGCTGGGCTTATCTGCTTGTT | |||
| ARSB | TGGAGGACATACCAATGGCAC | 146 | XM_046936477.1 |
| TGTCAATGCCAAGACATGGGT | |||
| HDAC1 | GTCTGCTACTACTACGACGGTG | 212 | NM_204156.2 |
| AACAGCCCATCGAACACAGG | |||
| LGR4 | AGTCAGCCCACTCCGACTAT | 101 | XM_046919090.1 |
| TCGGCTTTGGTAGAAGCAGG | |||
| NECTIN1 | ATTTCAAGGGTCCTGTCTCC | 249 | XM_040690207.2 |
| GGTCATGGTCTGCCTCTGT | |||
| KIF13B | GGCTACGGATTGCTTGTA | 190 | XM_040668432.2 |
| CTGTGGGTCCTCCTTCTG | |||
| CDH1 | CTCGGCAGAGGCAGCG | 268 | NM_001039258.3 |
| CTGCAAAGCTCACTCGTCCC | |||
| VNN2 | GGCATTTATGGCTGGGTT | 126 | NM_001379258.2 |
| TTCCTGCACTGGTGTTCG | |||
| LYSMD3 | TTTCAGTATAGACAGGAGGGTA | 214 | NM_001031414.2 |
| TAATCACAGCGAAGTGCC | |||
| TMEM41B | TGGTCAGAACAGGTTGAGCG | 221 | NM_001008469.2 |
| TCCCCTGCTGTTGTAAGTTGG | |||
| LAMC2 | TGCTGCGTCTGATGAACC | 158 | XM_040677531.2 |
| TTTCTGTACCCGAACCTTCT | |||
| ATP6V0D2 | CAGCGACCCATAGCTGAACT | 129 | NM001008455.2 |
| AGCTAATGGCGTGTCAACCA | |||
| GALC | CGATGTCATTGGGGCTCACT | 140 | XM_040672942.2 |
| GGTTTAGGATCCGAGCCCAG | |||
| ACKR4 | TGGGAACCTTTGTGCCAAGA | 123 | XM_040662558.2 |
| TGTGTCTCTGAAGCGGGAAC |
| No. | Sample | Total Reads | Total Beas | Clean Reads | Clean Ratio | No rRNA | rRNA Ratio | Q20 | Q30 |
|---|---|---|---|---|---|---|---|---|---|
| 1 | A2768 | 41290886 | 6193632900 | 40788174 | 0.98782511 | 40639184 | 0.003652774 | 97.15979161 | 93.71256892 |
| 2 | A2769 | 43203532 | 6480529800 | 42829888 | 0.99135154 | 40363728 | 0.057580351 | 97.57117474 | 94.43186642 |
| 3 | A3456 | 47722774 | 7158416100 | 47036118 | 0.985611566 | 45621216 | 0.030081181 | 96.96813771 | 93.13143801 |
| 4 | A3465 | 42878956 | 6431843400 | 42423802 | 0.989385143 | 42311348 | 0.002650729 | 97.75753001 | 93.97439017 |
| 5 | A3532 | 48303236 | 7245485400 | 47802900 | 0.989641771 | 47477980 | 0.006797077 | 97.3906402 | 94.33633048 |
| 6 | A3554 | 47623442 | 7143516300 | 47073466 | 0.98845157 | 46927954 | 0.003091168 | 97.69168792 | 93.80712829 |
| 7 | A3573 | 56306766 | 8446014900 | 55602814 | 0.987497915 | 55405198 | 0.003554065 | 97.07041633 | 93.59152257 |
| 8 | A4004 | 44902512 | 6735376800 | 44461722 | 0.9901834 | 44281266 | 0.004058682 | 97.43871664 | 94.39259682 |
| 9 | A4143 | 42843266 | 6426489900 | 42440050 | 0.990588579 | 42260616 | 0.00422794 | 97.50059303 | 94.50548856 |
| 10 | A4229 | 47528730 | 7129309500 | 46951590 | 0.987857029 | 46520526 | 0.009181031 | 97.19146595 | 93.68076321 |
| 11 | A4244 | 38980140 | 5847021000 | 38323006 | 0.983141826 | 37490376 | 0.021726636 | 97.15792741 | 93.44682431 |
| 12 | A4263 | 39640364 | 5946054600 | 39172264 | 0.988191329 | 38236900 | 0.023878222 | 97.41623636 | 94.11710039 |
| 13 | A4307 | 44204974 | 6630746100 | 43689548 | 0.98834009 | 43078204 | 0.013992912 | 97.23425705 | 93.84134892 |
| 14 | A4392 | 42519718 | 6377957700 | 42093520 | 0.989976462 | 41502136 | 0.014049288 | 97.48222704 | 94.42762943 |
| 15 | A4464 | 40600416 | 6090062400 | 40112294 | 0.987977414 | 39673088 | 0.010949411 | 97.3925364 | 94.10944045 |
| 16 | A4466 | 46694404 | 7004160600 | 46013862 | 0.98542562 | 45845946 | 0.003649248 | 97.00504549 | 93.39408161 |
| 17 | A4517 | 46574380 | 6986157000 | 46047940 | 0.98869679 | 45648192 | 0.008681127 | 97.35008101 | 94.01003127 |
| 18 | A4533 | 42679370 | 6401905500 | 42265082 | 0.990293015 | 42086276 | 0.004230584 | 97.43959975 | 94.37898149 |
| 19 | C021137 | 49039190 | 7355878500 | 48484036 | 0.988679381 | 47655404 | 0.017090821 | 97.23522776 | 93.7609628 |
| 20 | C021411 | 43741574 | 6561236100 | 43264444 | 0.989092071 | 40508214 | 0.063706585 | 97.25531608 | 93.70296684 |
| 21 | C021450 | 47319258 | 7097888700 | 46746086 | 0.98788713 | 46167824 | 0.012370276 | 97.11530885 | 93.45446355 |
| 22 | C021451 | 43562000 | 6534300000 | 43142000 | 0.990358569 | 42339150 | 0.018609476 | 97.47991956 | 94.42757067 |
| 23 | C021583 | 47628328 | 7144249200 | 47094544 | 0.98879272 | 46711880 | 0.008125442 | 97.24236733 | 93.78820254 |
| 24 | C021763 | 44549006 | 6682350900 | 44073958 | 0.989336507 | 43712598 | 0.008198946 | 97.81448294 | 94.0684538 |
| 25 | C021867 | 52064670 | 7809700500 | 51472932 | 0.988634558 | 50792016 | 0.013228623 | 97.24649714 | 93.94987346 |
| 26 | C021876 | 47505956 | 7125893400 | 46826836 | 0.985704529 | 46494220 | 0.007103106 | 96.87017573 | 92.93656506 |
| 27 | C021880 | 46751736 | 7012760400 | 46262020 | 0.98952518 | 46124444 | 0.002973843 | 97.79330031 | 94.1210958 |
| 28 | C021881 | 46247662 | 6937149300 | 45677094 | 0.987662771 | 45528750 | 0.003247667 | 97.65677164 | 93.89365084 |
| 29 | C021885 | 45717196 | 6857579400 | 45198398 | 0.988652016 | 45034068 | 0.003635748 | 97.68713207 | 93.93626766 |
| 30 | C021908 | 46521638 | 6978245700 | 45906050 | 0.986767706 | 45574782 | 0.007216217 | 96.97610112 | 93.34343283 |
| 31 | C021932 | 42264136 | 6339620400 | 41762602 | 0.988133343 | 41624064 | 0.003317274 | 97.19962902 | 93.77237577 |
| 32 | C021934 | 46716960 | 7007544000 | 46144728 | 0.987751087 | 45945478 | 0.004317936 | 97.1564588 | 93.64715317 |
| 33 | C021978 | 46890136 | 7033520400 | 46252006 | 0.986390954 | 45999846 | 0.005451872 | 97.03469799 | 93.49602634 |
| 34 | C022061 | 37672330 | 5650849500 | 37146066 | 0.98603049 | 36569664 | 0.015517175 | 97.13091751 | 93.40916687 |
| 35 | C022079 | 49964930 | 7494739500 | 49291222 | 0.986516383 | 48962380 | 0.006671411 | 97.04056102 | 93.35227813 |
| 36 | C022320 | 42161842 | 6324276300 | 41699750 | 0.989040042 | 41553490 | 0.003507455 | 97.71758185 | 94.00582674 |
| No. | Sample | Total_Reads | Mapped_Reads | Pair_Mapped_Reads | Single_Mapped_Reads | Unique_Mapped_Reads | Multi_Mapped_Reads | Mapped_Ratio |
|---|---|---|---|---|---|---|---|---|
| 1 | A2768 | 40639184 | 37648050 | 36856456 | 791594 | 37648050 | 0 | 0.926397784 |
| 2 | A2769 | 40363728 | 37136457 | 36607972 | 528485 | 37136457 | 0 | 0.920045269 |
| 3 | A3456 | 45621216 | 42549740 | 41693572 | 856168 | 42549740 | 0 | 0.932674394 |
| 4 | A3465 | 42311348 | 39546101 | 38729866 | 816235 | 39546101 | 0 | 0.934645263 |
| 5 | A3532 | 47477980 | 44601659 | 43667408 | 934251 | 44601659 | 0 | 0.939417789 |
| 6 | A3554 | 46927954 | 43783548 | 42959934 | 823614 | 43783548 | 0 | 0.932995033 |
| 7 | A3573 | 55405198 | 51353274 | 50245204 | 1108070 | 51353274 | 0 | 0.926867439 |
| 8 | A4004 | 44281266 | 41224141 | 40452846 | 771295 | 41224141 | 0 | 0.930961211 |
| 9 | A4143 | 42260616 | 39444407 | 38666896 | 777511 | 39444407 | 0 | 0.93336091 |
| 10 | A4229 | 46520526 | 43670452 | 42921832 | 748620 | 43670452 | 0 | 0.93873513 |
| 11 | A4244 | 37490376 | 35070310 | 34471096 | 599214 | 35070310 | 0 | 0.93544834 |
| 12 | A4263 | 38236900 | 33282278 | 32793024 | 489254 | 33282278 | 0 | 0.870423021 |
| 13 | A4307 | 43078204 | 39293524 | 38376134 | 917390 | 39293524 | 0 | 0.91214397 |
| 14 | A4392 | 41502136 | 38637784 | 37964146 | 673638 | 38637784 | 0 | 0.930983022 |
| 15 | A4464 | 39673088 | 37165180 | 36472096 | 693084 | 37165180 | 0 | 0.936785662 |
| 16 | A4466 | 45845946 | 41188366 | 40215890 | 972476 | 41188366 | 0 | 0.898408029 |
| 17 | A4517 | 45648192 | 42749198 | 41948938 | 800260 | 42749198 | 0 | 0.936492687 |
| 18 | A4533 | 42086276 | 39455760 | 38808252 | 647508 | 39455760 | 0 | 0.93749706 |
| 19 | C021137 | 47655404 | 44757529 | 43960204 | 797325 | 44757529 | 0 | 0.939191052 |
| 20 | C021411 | 40508214 | 35861255 | 35372872 | 488383 | 35861255 | 0 | 0.885283538 |
| 21 | C021450 | 46167824 | 43457350 | 42595930 | 861420 | 43457350 | 0 | 0.941290844 |
| 22 | C021451 | 42339150 | 39501327 | 38742672 | 758655 | 39501327 | 0 | 0.93297402 |
| 23 | C021583 | 46711880 | 37963792 | 37300058 | 663734 | 37963792 | 0 | 0.812722417 |
| 24 | C021763 | 43712598 | 40986368 | 40234750 | 751618 | 40986368 | 0 | 0.937632854 |
| 25 | C021867 | 50792016 | 47449270 | 46505334 | 943936 | 47449270 | 0 | 0.93418757 |
| 26 | C021876 | 46494220 | 41612642 | 40456134 | 1156508 | 41612642 | 0 | 0.895006777 |
| 27 | C021880 | 46124444 | 43005919 | 42131352 | 874567 | 43005919 | 0 | 0.932388887 |
| 28 | C021881 | 45528750 | 42528020 | 41639618 | 888402 | 42528020 | 0 | 0.934091536 |
| 29 | C021885 | 45034068 | 41875304 | 40997744 | 877560 | 41875304 | 0 | 0.929858346 |
| 30 | C021908 | 45574782 | 41179602 | 40222118 | 957484 | 41179602 | 0 | 0.90356114 |
| 31 | C021932 | 41624064 | 38877768 | 38106328 | 771440 | 38877768 | 0 | 0.934021435 |
| 32 | C021934 | 45945478 | 42834059 | 42037960 | 796099 | 42834059 | 0 | 0.932280191 |
| 33 | C021978 | 45999846 | 41358882 | 40363442 | 995440 | 41358882 | 0 | 0.89910914 |
| 34 | C022061 | 36569664 | 32455229 | 31969348 | 485881 | 32455229 | 0 | 0.887490489 |
| 35 | C022079 | 48962380 | 45904961 | 44905224 | 999737 | 45904961 | 0 | 0.937555752 |
| 36 | C022320 | 41553490 | 39033181 | 38230696 | 802485 | 39033181 | 0 | 0.939347838 |


References
- Guo, J.; Zhao, Z.; Broadwater, C.; Tobin, I.; Liu, J.; Whitmore, M.; Zhang, G. Is intestinal microbiota fully restored after chickens have recovered from coccidiosis? Pathogens 2025, 14, 81. [Google Scholar] [CrossRef]
- Taha, S.; Nguyen-Ho-Bao, T.; Berberich, L.M.; Gawlowska, S.; Daugschies, A.; Rentería-Solís, Z. Interplay between Eimeria acervulina and Cryptosporidium parvum during in vitro infection of a chicken macrophage cell line (HD11). Life 2023, 13, 1267. [Google Scholar] [CrossRef]
- Zhang, L.; Song, Z.; Li, P.; Song, X.; Tang, X.; Hu, D. Molecular characterization and immune protective efficacy of 3 Eimeria tenella antigens. Poult. Sci. 2024, 103, 103234. [Google Scholar] [CrossRef]
- Bremner, A.; Kim, S.; Morris, K.M.; Nolan, M.J.; Borowska, D.; Wu, Z.; Tomley, F.; Blake, D.P.; Hawken, R.; Kaiser, P.; et al. Kinetics of the cellular and transcriptomic response to Eimeria maxima in relatively resistant and susceptible chicken lines. Front. Immunol. 2021, 12, 653085. [Google Scholar] [CrossRef] [PubMed]
- Heams, T.; Bed’hom, B.; Rebours, E.; Jaffrezic, F.; Pinard-van der Laan, M.H. Insights into gene expression profiling of natural resistance to coccidiosis in contrasting chicken lines. BMC Proc. 2011, 5 (Suppl. S4), S26. [Google Scholar] [CrossRef] [PubMed]
- Niraula, A.; Wolc, A.; Fulton, J.E.; Taylor, R.L., Jr.; Dalloul, R.A. The chicken major histocompatibility complex (MHC-B) and alloantigen systems A, D, E, and I impact resistance to coccidiosis. Poult. Sci. 2025, 104, 105151. [Google Scholar] [CrossRef]
- Du, S.; Song, Z.; Cen, Y.; Fan, J.; Li, P.; Si, H.; Hu, D. Susceptibility and cecal microbiota alteration to Eimeria-infection in Yellow-feathered broilers, Arbor Acres broilers and Lohmann pink layers. Poult. Sci. 2024, 103, 103824. [Google Scholar] [CrossRef]
- Broadwater, C.; Guo, J.; Liu, J.; Tobin, I.; Whitmore, M.A.; Kaiser, M.G.; Lamont, S.J.; Zhang, G. Breed-specific responses to coccidiosis in chickens: Identification of intestinal bacteria linked to disease resistance. J. Anim. Sci. Biotechnol. 2025, 16, 65. [Google Scholar] [CrossRef]
- Broom, L.J.; Kogut, M.H. Deciphering desirable immune responses from disease models with resistant and susceptible chickens. Poult. Sci. 2019, 98, 1634–1642. [Google Scholar] [CrossRef]
- Gul, H.; Habib, G.; Khan, I.M.; Rahman, S.U.; Khan, N.M.; Wang, H.; Khan, N.U.; Liu, Y. Genetic resilience in chickens against bacterial, viral and protozoal pathogens. Front. Vet. Sci. 2022, 9, 1032983. [Google Scholar] [CrossRef]
- Wang, X.; Zou, W.; Yu, H.; Lin, Y.; Dai, G.; Zhang, T.; Zhang, G.; Xie, K.; Wang, J.; Shi, H. RNA sequencing analysis of chicken cecum tissues following Eimeria tenella infection in vivo. Genes 2019, 10, 420. [Google Scholar] [CrossRef]
- Kim, M.; Chung, Y.; Manjula, P.; Seo, D.; Cho, S.; Cho, E.; Ediriweera, T.K.; Yu, M.; Nam, S.; Lee, J.H. Time-series transcriptome analysis identified differentially expressed genes in broiler chicken infected with mixed Eimeria species. Front. Genet. 2022, 13, 886781. [Google Scholar] [CrossRef] [PubMed]
- Tian, S.; Tang, W.; Zhong, Z.; Wang, Z.; Xie, X.; Liu, H.; Chen, F.; Liu, J.; Han, Y.; Qin, Y.; et al. Identification of runs of homozygosity islands and functional variants in Wenchang chicken. Animals 2023, 13, 1645. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Cao, L.; Zhu, Y.; Deng, C.; Su, S.; Xu, J.; Jin, W.; Li, J.; Wu, L.; Tao, J. Cloning and characterization of an Eimeria necatrix gene encoding a gametocyte protein and associated with oocyst wall formation. Parasites Vectors 2014, 7, 27. [Google Scholar] [CrossRef] [PubMed]
- Hodgson, J.N. Coccidiosis: Oocyst counting technique for coccidiostat evaluation. Exp. Parasitol. 1970, 28, 99–102. [Google Scholar] [CrossRef]
- Johnson, J.; Reid, W.M. Anticoccidial drugs: Lesion scoring techniques in battery and floor-pen experiments with chicken. Exp. Parasitol. 1970, 28, 30–36. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef]
- Kim, D.; Langmead, B.; Salzberg, S.L. HISAT: A fast spliced aligner with low memory requirements. Nat. Methods 2015, 12, 357–360. [Google Scholar] [CrossRef]
- Pertea, M.; Kim, D.; Pertea, G.M.; Leek, J.T.; Salzberg, S.L. Transcript-level expression analysis of RNA−seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 2016, 11, 1650–1667. [Google Scholar] [CrossRef]
- Yu, G.; Wang, L.G.; Han, Y.; He, Q.Y. Cluster Profiler: An R package for comparing biological themes among gene clusters. Omics J. Integr. Biol. 2012, 16, 284–287. [Google Scholar] [CrossRef]
- Mu, H.; Chen, J.; Huang, W.; Huang, G.; Deng, M.; Hong, S.; Ai, P.; Gao, C.; Zhou, H. OmicShare tools: A zero-code interactive online platform for biological data analysis and visualization. Imeta 2024, 3, e228. [Google Scholar] [CrossRef]
- Feng, Y.; Li, Y.; Zhao, Y.; Shi, H. Physiological, transcriptome and co-expression network analysis of chlorophyll-deficient mutants in flue-cured tobacco. BMC Plant Biol. 2023, 23, 153. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(T) (-Delta DeltaC) method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- Tang, D.; Chen, M.; Huang, X.; Zhang, G.; Zeng, L.; Zhang, G.; Wu, S.; Wang, Y. SRplot: A free online platform for data visualization and graphing. PLoS ONE 2023, 18, e0294236. [Google Scholar] [CrossRef] [PubMed]
- Mathis, G.F.; Lumpkins, B.; Cervantes, H.M.; Fitz-Coy, S.H.; Jenkins, M.C.; Jones, M.K.; Price, K.R.; Dalloul, R.A. Coccidiosis in poultry: Disease mechanisms, control strategies, and future directions. Poult. Sci. 2025, 104, 104663. [Google Scholar] [CrossRef]
- Tan, X.; Liu, R.; Zhao, D.; He, Z.; Li, W.; Zheng, M.; Li, Q.; Wang, Q.; Liu, D.; Feng, F.; et al. Large-scale genomic and transcriptomic analyses elucidate the genetic basis of high meat yield in chickens. J. Adv. Res. 2024, 55, 1–16. [Google Scholar] [CrossRef]
- Tian, S.; Li, W.; Zhong, Z.; Wang, F.; Xiao, Q. Genome-wide re-sequencing data reveals the genetic diversity and population structure of Wenchang chicken in China. Anim. Genet. 2023, 54, 328–337. [Google Scholar] [CrossRef]
- Xie, X.F.; Wang, Z.Y.; Zhong, Z.Q.; Pan, D.Y.; Hou, G.Y.; Xiao, Q. Genome-wide scans for selection signatures in indigenous chickens reveal candidate genes associated with local adaptation. Animal 2024, 18, 101151. [Google Scholar] [CrossRef]
- Zhang, T.; Qu, H.; Zheng, W.; Zhang, Y.; Li, Y.; Pan, T.; Li, J.; Yang, W.; Cao, X.; Jiang, Y.; et al. Oral vaccination with a recombinant Lactobacillus plantarum expressing the Eimeria tenella rhoptry neck 2 protein elicits protective immunity in broiler chickens infected with Eimeria tenella. Parasites Vectors 2024, 17, 277. [Google Scholar] [CrossRef]
- Wallach, M. Role of antibody in immunity and control of chicken coccidiosis. Trends Parasitol. 2010, 26, 382–387. [Google Scholar] [CrossRef]
- Kim, W.H.; Chaudhari, A.A.; Lillehoj, H.S. Involvement of T Cell immunity in avian coccidiosis. Front. Immunol. 2019, 10, 2732. [Google Scholar] [CrossRef] [PubMed]
- Dubey, J.P.; Jenkins, M.C. Re-evaluation of the life cycle of Eimeria maxima Tyzzer, 1929 in chickens (Gallus domesticus). Parasitology 2018, 145, 1051–1058. [Google Scholar] [CrossRef] [PubMed]
- Murphy, K.N.; Brinkworth, A.J. Manipulation of focal adhesion signaling by pathogenic microbes. Int. J. Mol. Sci. 2021, 22, 1358. [Google Scholar] [CrossRef] [PubMed]
- Liao, H.; Liu, S.; Ma, Q.; Huang, H.; Goel, A.; Torabian, P.; Mohan, C.D.; Duan, C. Endoplasmic reticulum stress induced autophagy in cancer and its potential interactions with apoptosis and ferroptosis. Biochim. Biophys. Acta Mol. Cell. Res. 2025, 1872, 119869. [Google Scholar] [CrossRef]
- Sadr, A.S.; Nassiri, M.; Ghaderi-Zefrehei, M.; Heidari, M.; Smith, J.; Muhaghegh Dolatabady, M. RNA−seq profiling between commercial and indigenous Iranian chickens highlights differences in innate immune gene expression. Genes 2023, 14, 793. [Google Scholar] [CrossRef]
- Xiang, S.; Zhang, W.; Wang, Z.; Chen, H.; Yang, C. Evaluating HSP90AA1 as a predictive biomarker for prognosis in lung adenocarcinoma. Transl. Cancer Res. 2025, 14, 2580–2593. [Google Scholar] [CrossRef]
- Liu, Z.; Liu, G.; Ha, D.P.; Wang, J.; Xiong, M.; Lee, A.S. ER chaperone GRP78/BiP translocates to the nucleus under stress and acts as a transcriptional regulator. Proc. Natl. Acad. Sci. USA 2023, 120, e2303448120. [Google Scholar] [CrossRef]
- Yang, H.; Jung, S.; Choi, E.Y. E3 ubiquitin ligase TRIM38 regulates macrophage polarization to reduce hepatic inflammation by interacting with HSPA5. Int. Immunopharmacol. 2025, 157, 114662. [Google Scholar] [CrossRef]
- Miao, C.; Zhang, Y.; Yu, M.; Wei, Y.; Dong, C.; Pei, G.; Xiao, Y.; Yang, J.; Yao, Z.; Wang, Q. HSPA8 regulates anti-bacterial autophagy through liquid-liquid phase separation. Autophagy 2023, 19, 2702–2718. [Google Scholar] [CrossRef]
- Deng, L.; He, S.; Guo, N.; Tian, W.; Zhang, W.; Luo, L. Molecular mechanisms of ferroptosis and relevance to inflammation. Inflamm. Res. 2023, 72, 281–299. [Google Scholar] [CrossRef]
- Hu, S.; Chu, Y.; Zhou, X.; Wang, X. Recent advances of ferroptosis in tumor: From biological function to clinical application. Biomed. Pharmacother. 2023, 166, 115419. [Google Scholar] [CrossRef]
- Yan, Y.; Chen, R.; Wang, X.; Hu, K.; Huang, L.; Lu, M.; Hu, Q. CCL19 and CCR7 expression, signaling pathways, and adjuvant functions in viral infection and prevention. Front. Cell Dev. Biol. 2019, 7, 212. [Google Scholar] [CrossRef] [PubMed]
- Hong, W.; Yang, B.; He, Q.; Wang, J.; Weng, Q. New insights of CCR7 signaling in dendritic cell migration and inflammatory diseases. Front. Pharmacol. 2022, 13, 841687. [Google Scholar] [CrossRef] [PubMed]
- Kawabe, T. Homeostasis and immunological function of self-driven memory-phenotype CD4+ T lymphocytes. Immunol. Med. 2023, 46, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Xie, Z.; Zhou, G.; Zhang, M.; Han, J.; Wang, Y.; Li, X.; Wu, Q.; Li, M.; Zhang, S. Recent developments on BMPs and their antagonists in inflammatory bowel diseases. Cell Death Discov. 2023, 9, 210. [Google Scholar] [CrossRef]
- Vu, T.H.; Hong, Y.; Truong, A.D.; Lee, J.; Lee, S.; Song, K.D.; Cha, J.; Dang, H.V.; Tran, H.T.T.; Lillehoj, H.S.; et al. Cytokine-cytokine receptor interactions in the highly pathogenic avian influenza H5N1 virus-infected lungs of genetically disparate Ri chicken lines. Anim. Biosci. 2022, 35, 367–376. [Google Scholar] [CrossRef]
- Weinberg, S.E.; Sun, L.Y.; Yang, A.L.; Liao, J.; Yang, G.Y. Overview of inositol and inositol phosphates on chemoprevention of colitis-induced carcinogenesis. Molecules 2020, 26, 31. [Google Scholar] [CrossRef]
- Du, Y.; He, C.; An, Y.; Huang, Y.; Zhang, H.; Fu, W.; Wang, M.; Shan, Z.; Xie, J.; Yang, Y.; et al. The role of short chain fatty acids in inflammation and body health. Int. J. Mol. Sci. 2024, 25, 7379. [Google Scholar] [CrossRef]
- Yan, D.; Ye, S.; He, Y.; Wang, S.; Xiao, Y.; Xiang, X.; Deng, M.; Luo, W.; Chen, X.; Wang, X. Fatty acids and lipid mediators in inflammatory bowel disease: From mechanism to treatment. Front. Immunol. 2023, 14, 1286667. [Google Scholar] [CrossRef]
- Su, J.; Cheng, F.; Yuan, W. Unraveling the cGAS/STING signaling mechanism: Impact on glycerolipid metabolism and diseases. Front. Med. 2024, 11, 1512916. [Google Scholar] [CrossRef]
- Sutherland, T.E.; Dyer, D.P.; Allen, J.E. The extracellular matrix and the immune system: A mutually dependent relationship. Science 2023, 379, eabp8964. [Google Scholar] [CrossRef]
- Mukherjee, T.; Kumar, N.; Chawla, M.; Philpott, D.J.; Basak, S. The NF-κB signaling system in the immunopathogenesis of inflammatory bowel disease. Sci. Signal. 2024, 17, eadh1641. [Google Scholar] [CrossRef] [PubMed]
- Tavares, L.P.; Negreiros-Lima, G.L.; Lima, K.M.; E Silva, P.M.R.; Pinho, V.; Teixeira, M.M.; Sousa, L.P. Blame the signaling: Role of cAMP for the resolution of inflammation. Pharmacol. Res. 2020, 159, 105030. [Google Scholar] [CrossRef]
- Kayhan, M.; Koyas, A.; Akdemir, I.; Savas, A.C.; Cekic, C. Adenosine receptor signaling Ttargets both PKA and Epac pathways to polarize dendritic cells to a suppressive phenotype. J. Immunol. 2019, 203, 3247–3255. [Google Scholar] [CrossRef]
- Zhao, C.; Wu, K.; Hao, H.; Zhao, Y.; Bao, L.; Qiu, M.; He, Y.; He, Z.; Zhang, N.; Hu, X.; et al. Gut microbiota-mediated secondary bile acid alleviates Staphylococcus aureus-induced mastitis through the TGR5-cAMP-PKA-NF-kappaB/NLRP3 pathways in mice. NPJ Biofilms Microbiomes 2023, 9, 8. [Google Scholar] [CrossRef]
- Jiang, Z.; Zhou, W.; Tian, X.; Zou, P.; Li, N.; Zhang, C.; Li, Y.; Liu, G. A protective role of canonical Wnt/β-Catenin pathway in pathogenic bacteria-induced inflammatory responses. Mediat. Inflamm. 2024, 2024, 8869510. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Li, J.; Shen, H.; Xu, M.; Han, W.; Dou, X. Comprehensive Analysis of Immune Response and Transcriptome Profiling Reveals the Molecular Basis Underlying Breed-Specific Responses to Eimeria tenella Infection in Chickens with Differing Susceptibility Levels. Animals 2025, 15, 2533. https://doi.org/10.3390/ani15172533
Li J, Shen H, Xu M, Han W, Dou X. Comprehensive Analysis of Immune Response and Transcriptome Profiling Reveals the Molecular Basis Underlying Breed-Specific Responses to Eimeria tenella Infection in Chickens with Differing Susceptibility Levels. Animals. 2025; 15(17):2533. https://doi.org/10.3390/ani15172533
Chicago/Turabian StyleLi, Jianmei, Haiyu Shen, Ming Xu, Wei Han, and Xinhong Dou. 2025. "Comprehensive Analysis of Immune Response and Transcriptome Profiling Reveals the Molecular Basis Underlying Breed-Specific Responses to Eimeria tenella Infection in Chickens with Differing Susceptibility Levels" Animals 15, no. 17: 2533. https://doi.org/10.3390/ani15172533
APA StyleLi, J., Shen, H., Xu, M., Han, W., & Dou, X. (2025). Comprehensive Analysis of Immune Response and Transcriptome Profiling Reveals the Molecular Basis Underlying Breed-Specific Responses to Eimeria tenella Infection in Chickens with Differing Susceptibility Levels. Animals, 15(17), 2533. https://doi.org/10.3390/ani15172533





