Abstract
Lower respiratory tract infections (LRTIs) remain a major cause of global morbidity and mortality, yet accurate pathogen identification and risk stratification continue to pose clinical challenges. Lipidomics—the comprehensive analysis of lipid species within biological systems—has emerged as a promising tool to unravel host–pathogen interactions and reveal novel diagnostic and prognostic biomarkers. This systematic review synthesizes evidence from nine original studies applying mass spectrometry-based lipidomic profiling in human LRTIs, including community-acquired pneumonia (CAP), ventilator-associated pneumonia (VAP), and coronavirus disease 2019 (COVID-19). Across diverse study designs, sample types, and analytical platforms, consistent alterations in lipid metabolism were observed. Perturbations in phospholipid classes, particularly phosphatidylcholines (PCs) and lysophosphatidylcholines (LPCs), were frequently associated with disease severity and immune activation. The ratios of PC to LPC and phosphatidylethanolamine (PE) to lysophosphatidylethanolamine (LPE) emerged as markers of inflammatory remodeling. Sphingolipids—including sphingomyelins (SMs) and sphingosine-1-phosphate (S1P)—were identified as key modulators of monocyte and neutrophil activation. Fatty acid–derived lipid mediators such as oxylipins (e.g., 12,13-epoxyoctadecenoic acid and 15-hydroxyeicosatetraenoic acid) and acylcarnitines reflected pathogen-specific immune responses and mitochondrial dysfunction. Several lipid-based classifiers demonstrated superior diagnostic and prognostic performance compared to conventional clinical scores, including the CURB-65 and pneumonia severity index. However, significant heterogeneity in experimental design, lipid identification workflows, and reporting standards limits inter-study comparability. While preliminary findings support the integration of lipidomics into infectious disease research, larger multi-omic and longitudinal studies are required. This review provides the first comprehensive synthesis of lipidomic alterations in human LRTIs and highlights their emerging translational relevance.
1. Introduction
Lower respiratory tract infections (LRTIs) continue to pose a significant public health burden worldwide, encompassing a broad spectrum of conditions ranging from viral illnesses such as influenza, respiratory syncytial virus (RSV), and severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), to bacterial pneumonias caused by pathogens like Streptococcus pneumoniae, Haemophilus influenzae, and Klebsiella pneumoniae [1,2,3,4]. These infections can be classified based on their anatomical location—upper or lower respiratory tract—as well as by the setting in which they are acquired, distinguishing community-acquired pneumonia (CAP) from nosocomial infections such as hospital-acquired pneumonia (HAP) and ventilator-associated pneumonia (VAP) [5]. Despite advances in diagnostic microbiology and host biomarker discovery, challenges remain in early detection, pathogen-specific diagnosis, and prognostication of disease severity in both adult and pediatric populations.
These diagnostic and prognostic limitations are particularly critical given that LRTIs remain among the leading causes of global morbidity and mortality, disproportionately affecting young children, older adults, and immunocompromised patients [3]. Conventional diagnostic tools, such as cultures, antigen-based assays, or commonly used biomarkers including C-reactive protein (CRP) and procalcitonin (PCT), often lack sensitivity and specificity, delaying appropriate treatment decisions. Similarly, clinical severity scores such as the pneumonia severity index (PSI) and CURB-65 may not consistently predict outcomes across heterogeneous patient populations [5]. This highlights an urgent need for novel biomarkers and molecular tools that can improve early pathogen identification, refine risk stratification, and guide therapeutic interventions.
In recent years, the emergence of lipidomics as a subfield of metabolomics has offered new opportunities for understanding the pathophysiological basis of infectious diseases [6]. Lipidomics entails the comprehensive characterization of lipid species within biological systems using mass spectrometry–based approaches such as liquid chromatography–mass spectrometry (LC-MS), gas chromatography–mass spectrometry (GC-MS), and shotgun lipidomics [7]. These technologies allow the quantification of hundreds to thousands of lipid molecules, including glycerophospholipids, sphingolipids, fatty acids, and sterols, across various biological matrices [8,9,10]. By providing a systems-level overview of lipid remodeling during infection, lipidomics offers a unique opportunity to link clinical phenotypes with underlying host–pathogen biology, thereby enhancing both diagnostic precision and prognostic assessment in respiratory infections.
Lipidomics has become an increasingly valuable tool for studying host–pathogen interactions, given the essential roles of lipids in membrane structure, energy balance, and immune signaling [11]. Bioactive lipid mediators such as lysophosphatidylcholines (LPCs), sphingolipids, oxylipins, and bile acids are often dysregulated in LRTIs, potentially affecting immune responses and disease progression. Integrated omics studies have shown that plasma lipidomic profiles can differentiate viral from bacterial CAP, offering promising diagnostic insights [12]. Specific phosphatidylcholines (PCs) (e.g., PC 16:0_18:1, 36:4, 38:6), identified via ultra-high performance liquid chromatography–tandem mass spectrometry (UHPLC-MS/MS), have been associated with inflammation, oxygenation, and 30-day outcomes, outperforming clinical scores like pneumonia severity index (PSI) and CURB-65 in predicting severe CAP [13].
Evidence from RSV and SARS-CoV-2 studies further supports lipidomics’ diagnostic and prognostic potential. RSV-induced pneumonia alters lipid metabolism in lungs and plasma, especially affecting phospholipids and acylcarnitines [14]. In COVID-19 acute respiratory distress syndrome (ARDS), lipidomic analysis of tracheal aspirates revealed distinct lipid changes associated with VAP, with sphingomyelin (34:1) and PC (O-34:1) outperforming C-reactive protein (CRP) and procalcitonin (PCT) as biomarkers. Similarly, pediatric CAP studies have identified reduced LPCs and increased sphinganine, reflecting inflammation-related metabolic changes [15].
Notably, the clinical value of lipidomics has already been demonstrated in several other fields of medicine. In cardiovascular research, plasma lipid signatures have been integrated into risk prediction models for atherosclerotic disease and heart failure [8]. In oncology, lipidomics has contributed to cancer subtyping and treatment monitoring, particularly in breast and colorectal cancer [9]. In metabolic disorders, altered lipid pathways have been used to identify biomarkers of insulin resistance and non-alcoholic fatty liver disease [10]. These advances illustrate the translational potential of lipidomics and provide a compelling rationale for its application to respiratory tract infections, where reliable diagnostic and prognostic tools remain limited. Despite these promising findings, lipidomic applications in LRTIs remain underexplored relative to other ‘omics’ disciplines. Heterogeneity in lipidomic platforms, study designs, and clinical endpoints has limited cross-study comparability and translational relevance. Nonetheless, the growing body of literature suggests that lipid profiling may yield novel biomarkers for early diagnosis, disease stratification, and mechanistic understanding of LRTIs across diverse patient populations.
The aim of this systematic review is to synthesize existing evidence on the use of lipidomics in respiratory tract infections. Specifically, the review explores lipidomic alterations associated with various pathogens, the biological functions of dysregulated lipids in infection pathophysiology, and the diagnostic and prognostic potential of lipid-based biomarkers in both viral and bacterial LRTIs. Furthermore, this work identifies methodological gaps and provides recommendations for future studies aimed at integrating lipidomic tools into clinical infectious disease research.
2. Methods
This systematic review was designed in accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) 2020 statement [16]. It has also been registered in the International Prospective Register of Systematic Reviews (PROSPERO) with ID number CRD420251103189.
2.1. Search Strategy
A systematic and comprehensive literature search was performed across four major electronic bibliographic databases—PubMed, Scopus, Web of Science, and Google Scholar—from their respective inceptions through August 2025. The search strategy was constructed using a combination of controlled vocabulary terms (e.g., Medical Subject Headings [MeSH]) and relevant free-text terms to maximize sensitivity and capture a broad spectrum of relevant studies.
The core search concepts included lipidomics and respiratory infections. Key search terms encompassed: “lipidomics”, “respiratory tract infection”, “community-acquired pneumonia”, “hospital-acquired pneumonia”, “ventilator-associated pneumonia”, “viral pneumonia”, “bacterial pneumonia”, “COVID-19”, “SARS-CoV-2”, “RSV”, “influenza”, “mass spectrometry”, “tuberculosis” “ultra-high-performance liquid chromatography”, “liquid chromatography–mass spectrometry”, “gas chromatography–mass spectrometry” “sphingolipids”, “phospholipids”, “oxylipins”, “lysophospholipids”, “bronchoalveolar lavage”, “tracheal aspirate”, “serum”, and “plasma”.
Boolean operators (AND, OR), truncation symbols, and field-specific tags (e.g., [MeSH Terms], [Title/Abstract]) were utilized to enhance the precision and breadth of the search. The full search syntax was adapted to the indexing system of each database. Additionally, the reference lists of all included full-text articles were manually screened to identify potentially eligible studies not captured by electronic search.
2.2. Inclusion and Exclusion Criteria
Studies were eligible for inclusion if they employed any form of lipidomic analysis, whether targeted or untargeted, and presented original research data relevant to LRTIs. Eligible study designs included both observational studies, such as cohort, case–control, and cross-sectional studies, and interventional studies, including clinical trials or controlled experimental models. The review considered studies involving human participants (patients or healthy controls), provided the respiratory infection was clearly defined and lipidomic data were reported. Acceptable sample types included serum, plasma, sputum, bronchoalveolar lavage fluid, tracheal aspirates, or other biologically relevant specimens collected in the context of LRTIs. To be included, studies had to utilize recognized lipidomic technologies such as LC-MS, GC-MS, nuclear magnetic resonance spectroscopy (NMR), or shotgun lipidomics, and report specific findings on lipid species, lipid class alterations, or lipid pathway perturbations linked to the infection.
Studies were excluded if they did not involve LRTIs or if lipidomic analysis was not performed. Publications that focused solely on other omics approaches (e.g., proteomics or transcriptomics) without presenting lipid-specific results were not considered eligible. Secondary literature including reviews, editorials, commentaries, expert opinions, and methodological papers without primary lipidomic data were also excluded. Additionally, case reports, small case series with fewer than five patients, animal studies and conference abstracts that lacked sufficient information on methods or results were not included. Only studies published in English and accessible in full-text format were considered for final inclusion.
2.3. PRISMA Process
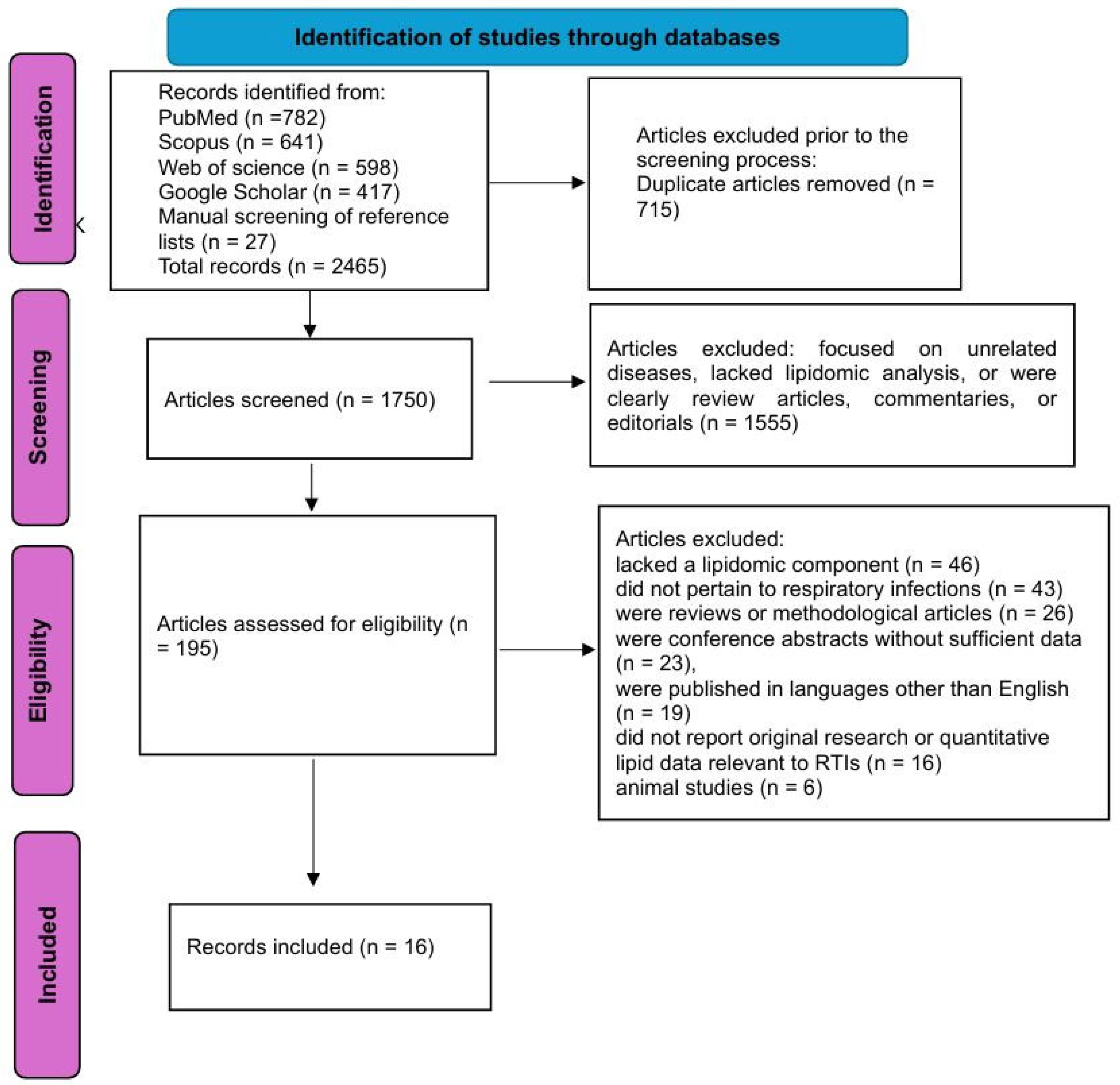
The literature search yielded a total of 2438 records across four major databases: PubMed (782), Scopus (641), Web of Science (598), and Google Scholar (417). An additional 27 articles were identified through manual screening of reference lists from relevant reviews and included studies, bringing the total to 2465 records.
After the removal of 715 duplicate entries, 1750 unique records remained for title and abstract screening. This initial screening was conducted independently by two reviewers, resulting in the exclusion of 1555 records that did not meet the inclusion criteria. These excluded records primarily focused on unrelated diseases, lacked lipidomic analysis, or were review articles, commentaries, or editorials.
A total of 195 full-text articles were retrieved and assessed for eligibility. During full-text screening, 179 articles were excluded for the following reasons: 46 studies lacked a lipidomic component, 43 did not pertain to respiratory infections or tuberculosis, 26 were reviews or methodological articles, 23 were conference abstracts without sufficient data, 19 were published in languages other than English, 16 did not report original research or quantitative lipid data relevant to LRTIs/tuberculosis (TB), and 6 were animal studies.
Following this selection process, 16 studies met all predefined inclusion criteria and were included in the final qualitative synthesis. These encompassed diverse respiratory pathogens, including community-acquired pneumonia, COVID-19, ventilator-associated pneumonia, Mycoplasma pneumoniae infection, and tuberculosis.
Figure 1 illustrates the study selection process.
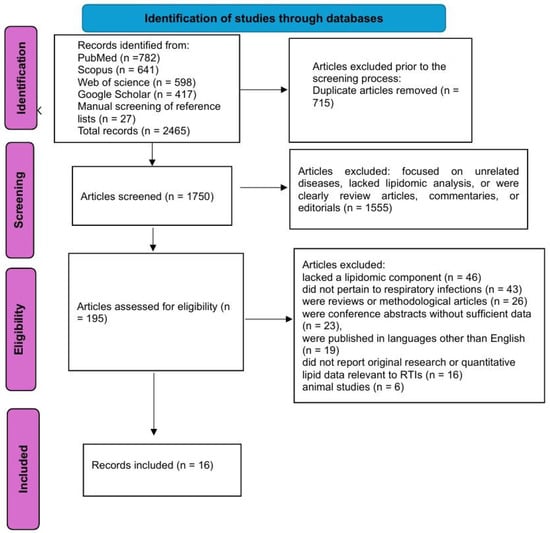
Figure 1.
The flowchart of the study selection process.
2.4. Quality Assessment
The methodological quality of the included studies was assessed using the Newcastle–Ottawa Scale (NOS), a validated tool specifically designed for evaluating the risk of bias in non-randomized observational studies, including cohort and case–control designs [17].
2.5. Data Extraction
A standardized data extraction form was used to record study characteristics, including author, year of publication, country, study design, infection type, sample source, analytical platform (e.g., LC-MS, GC-MS), number and classes of lipid species identified, and major lipidomic findings. Outcomes of interest included the identification of specific lipids associated with infection, the use of lipidomics for differential diagnosis or prognosis, and mechanistic insights into host–pathogen interactions mediated by lipids.
2.6. Data Synthesis
Given the heterogeneity in study populations, lipidomic techniques, and outcome measures, a meta-analysis was not performed. Instead, findings were synthesized narratively, with particular attention to patterns of lipid dysregulation across different pathogens, biological plausibility of identified lipids, and consistency of results across studies.
3. Results
3.1. Study Characteristics and Analytical Platforms
The included studies [12,13,15,18,19,20,21,22,23,24,25,26,27,28,29,30] span diverse geographic regions and clinical settings, with cohorts recruited from Spain, China, Germany, France, Greece, the Netherlands, the United States, and Brazil. Study designs encompassed prospective observational cohorts, retrospective post hoc analyses, cross-sectional investigations, and pilot studies. Patient populations covered a wide clinical spectrum, including adults and children, hospitalized and ICU patients, and individuals with varying degrees of severity across conditions such as CAP, COVID-19, VAP, sepsis, and tuberculosis. Sampling strategies were heterogeneous, drawing on serum, plasma, BALF, tracheal aspirates, urine, nasopharyngeal swabs, and isolated immune cells. Several studies incorporated validation cohorts (e.g., CAPSOD and EARLI) or longitudinal sampling (e.g., Days 1, 3, 6, 12). Most employed untargeted lipidomic approaches on LC-MS/MS or UHPLC-HRMS platforms, ensuring broad coverage of lipid classes, while others adopted targeted analyses for specific phospholipid ratios. Notably, integrative multi-omics frameworks combining lipidomics with proteomics, transcriptomics, or metabolomics provided additional mechanistic insights into host–pathogen interactions. Summary of the studies’ characteristics is displayed in Table 1.

Table 1.
Summary of the studies’ characteristics.
3.2. Identified Lipid Biomarkers in LRTIs
PCs and LPCs were the most frequently investigated lipid classes, with multiple studies reporting decreased LPC and variable PC levels in patients with CAP and COVID-19. Notably, specific phosphatidylcholine species such as PC(18:2_20:4), PC(36:4), and PC(38:6) were found to be significantly reduced in severe CAP and were independently associated with disease severity, oxygen requirements, and procalcitonin levels. In particular, PC(18:2_20:4) demonstrated high diagnostic accuracy with an area under the curve (AUC) of 0.954 [13].
Similarly, decreased LPC(22:6-sn2) and increased PC(36:1) were among the most discriminatory lipid alterations in hospitalized COVID-19 patients, with the LPC/PC ratio achieving an AUC > 0.95 for disease differentiation [18]. Other studies reported that the PC/LPC and phosphatidylethanolamine/lysophosphatidylethanolamine (PE/LPE) ratios were positively correlated with systemic inflammation markers and predictive of mortality [22].
Sphingolipids, particularly sphingomyelins and ceramides, were also frequently altered. Sphingomyelin (SM) (34:1) and PC(O-34:1) emerged as strong predictive biomarkers for VAP in COVID-19-associated ARDS, with area under the receiver operating characteristic curve (AUROC) values of 0.85 and 0.83, respectively [21]. In a complementary study, Schuurman et al. (2024) [23] identified significant upregulation of the sphingosine-1-phosphate (S1P) signaling pathway and its regulatory enzymes—sphingosine kinase 1 (SPHK1), UDP-glucose ceramide glucosyltransferase (UGCG), and sphingomyelin phosphodiesterase 1 (SMPD1)—in monocytes and neutrophils of patients with CAP, thereby linking sphingolipid metabolic alterations to pro-inflammatory immune responses.
In terms of fatty acid-derived lipids, oxylipins such as 9- and 13-hydroxyoctadecadienoic acid (9/13-HODE), 15-hydroxyeicosatetraenoic acid (15-HETE), 12,13-epoxyoctadecenoic acid (12,13-EpOME) and 9,10-dihydroxyoctadecenoic acid (9,10-DiHOME) were differentially expressed across viral and bacterial etiologies. Viral CAP was characterized by elevated oxylipins and FA 18:2–containing triglycerides (TGs) and diacylglycerols (DGs), while bacterial CAP showed enrichment in PCs and PC ethers [12].
Several studies identified cholesterol esters as inversely correlated with severity scores and systemic inflammation. In the study by Chouchane et al. [19], a 10% increase in cholesteryl ester (CE) levels by Day 4 in the ICU was associated with a reduced 30-day mortality [odds ration (OR) = 0.84], supporting their prognostic significance.
Notably, lipidomic analyses of tracheal aspirates revealed local lipid remodeling at the infection site. In patients with high VAP suspicion, multiple PC and SM species were upregulated, and these lipid signatures correlated with pathogen type and inflammation [15].
Humes et al. [24] showed that acylcarnitines, DGs, and LPCs defined pathogen-specific clusters, with high TG/DG profiles associated with influenza A H3 and low-lipid profiles with rhinovirus. Zheng et al. [25] reported >10-fold increases in lactosylceramides in CAP, while decreased sphingosine and lysolipids correlated with severity. In tuberculosis, Sun et al. [26] identified bile acid and fatty acid pathway dysregulation, with Angiotensin IV achieving diagnostic AUCs > 0.99. Zhang et al. [27] highlighted three lipid species (CER(24:0) H, HCER(d18:0/22:0) H, PE(18:1/18:1)) as strong TB classifiers. Chen et al. [28] found perturbations in lysoPE, LPC, and PI as discriminators of pneumonia severity in children. Tomalka et al. [29] demonstrated that AA, EPA, DHA, and DPA correlated strongly with WHO COVID-19 severity scores. De Almeida et al. [30] identified key lipid classes (PS, PI, PA vs. PC, SM, DG) that, combined with proteomic profiles, achieved >95% accuracy in COVID-19 diagnosis.
Table 2 displays the main lipid biomarkers identified across studies, their putative biological functions in the context of respiratory infections, and their reported diagnostic or prognostic performance.

Table 2.
Main lipid biomarkers, biological functions, and diagnostic/prognostic performance.
3.3. Role of Lipidomics in Pathophysiological Insights
One of the most consistent findings across studies is the dysregulation of phospholipids, particularly PCs and LPCs, which are integral to membrane integrity, surfactant composition, and inflammatory signaling. Reduced LPC levels, frequently reported in severe CAP and COVID-19, may reflect increased consumption during acute-phase inflammation and impaired resolution of infection [13,18,19,20]. Furthermore, the altered PC/LPC and PE/LPE ratios, which correlated with pro-inflammatory cytokines, CRP levels, and disease severity, suggest that lipid remodeling enzymes such as lysophosphatidylcholine acyltransferase 1/2 (LPCAT1/2) are tightly regulated in infection-induced stress responses [22].
Lipidomics has also shed light on bioactive lipid mediators such as oxylipins, which include hydroxyoctadecadienoic acids (HODEs) and epoxyoctadecenoic acids (EpOMEs). These metabolites, derived from linoleic acid and other polyunsaturated fatty acids via enzymatic oxidation, are known to modulate vascular permeability, leukocyte recruitment, and immune cell activation. Their reduction or elevation in viral versus bacterial infections points to divergent lipid signaling pathways depending on the pathogenic trigger [12,18].
Another key contribution of lipidomics is the elucidation of sphingolipid metabolism, particularly the role of SMs and S1P in immune regulation. Sphingolipids are not only structural membrane components but also active regulators of neutrophil migration, cytokine release, and lymphocyte trafficking. In CAP patients, upregulation of SPHK1, the enzyme responsible for S1P synthesis, was linked to enhanced monocyte and neutrophil activation, as confirmed by transcriptomic integration and functional inhibition assays [23].
Elevated acylcarnitines indicate disrupted mitochondrial β-oxidation and energy stress, while increased circulating triglycerides—particularly in viral CAP and COVID-19—are consistent with hepatic lipogenesis driven by inflammatory cytokines such as interleukin -6 (IL-6) and tumor necrosis factor alpha (TNF-α) [12,18].
Lipidomic signatures derived from localized samples such as tracheal aspirates further illuminate tissue-specific processes. In VAP, differential expression of surfactant-associated lipids and ether-linked phospholipids pointed to alveolar epithelial injury rather than direct bacterial lipid contribution. These changes were correlated with pathogen-specific profiles, offering insights into pulmonary lipid metabolism at the site of infection [15,21].
These studies further illuminate disease mechanisms. Humes et al. [24] demonstrated that sputum lipid clusters reflected viral subtype–specific metabolic remodeling. Zheng et al. [25] linked BALF lipid patterns with immune cell infiltration, suggesting a role in local host–pathogen interactions. In tuberculosis, Sun et al. [26] and Zhang et al. [27] both implicated bile acid and sphingolipid pathways, consistent with chronic inflammation and immune evasion. Chen et al. [28] showed that Mycoplasma pneumoniae infection perturbed lysophospholipid and PI metabolism, reflecting membrane damage. Tomalka et al. [29] revealed that pro-resolving mediators and eicosanoid precursors not only tracked severity but also mapped to interferon suppression in COVID-19. De Almeida et al. [30] demonstrated that systemic lipid and protein signatures serve as robust diagnostic fingerprints of infection.
3.4. Lipidomic Alterations by Pathogen Type
In the prospective cohort study by Rischke et al. [12], lipidomic profiling of patients with viral versus bacterial CAP revealed substantial differences. Bacterial CAP was characterized by elevated levels of PCs and ether-linked PCs, consistent with enhanced membrane biosynthesis and neutrophilic activation. In contrast, viral CAP showed increased concentrations of TGs and DGs, particularly those containing linoleic acid (18:2), as well as higher levels of pro-inflammatory oxylipins, including 12,13-EpOME and 9,10-DiHOME.
Similarly, in a pediatric ICU cohort, Virgiliou et al. [15] demonstrated that lipidomic profiles in VAP varied by pathogen. Blood lipid signatures—including specific PCs, SMs, and TGs—differentiated infections due to Staphylococcus aureus and Klebsiella pneumoniae, supporting the concept of pathogen-specific metabolic remodeling in critically ill children. Notably, PCs (e.g., PC 32:2) and TGs (e.g., TG 48:3) showed good discriminatory performance (AUC > 0.75).
Further evidence for pathogen-specific lipid responses was observed in the study by Kassa-Sombo et al. [21], which examined tracheal aspirate lipidomes in COVID-19 ARDS patients with or without VAP. While not explicitly focused on microbial taxonomy, the study identified eight significantly upregulated lipids [including SM(34:1) and PC(O-34:1)] in VAP patients. These lipids were associated with epithelial cell membrane breakdown rather than direct bacterial lipid production, but their expression patterns reflected the presence and severity of superimposed bacterial infection.
Humes et al. [24] provided clear evidence of pathogen-specific signatures among respiratory viruses, with influenza A H3 enriched in TG/DG and rhinovirus in lipid-depleted profiles. Zheng et al. [25] highlighted CAP-specific increases in lactosylceramides compared to controls. In tuberculosis, both Sun et al. [26] and Zhang et al. [27] revealed distinct lipid dysregulation patterns compared to healthy individuals. Chen et al. [28] distinguished M. pneumoniae pneumonia from bronchitis through plasma/urine metabolite panels. Tomalka et al. [29] observed conserved lipid mediator dysregulation across SARS-CoV-2 variants, while de Almeida et al. [30] identified COVID-19–specific lipid fingerprints distinct from seronegative controls.
3.5. Temporal Dynamics and Prognostic Implications
In a multicenter study of critically ill CAP patients, Chouchane et al. [19] documented time-dependent recovery of specific lipid classes, most notably CE and LPC. More than half (58%) of all quantified lipid species were decreased upon ICU admission. However, restoration of CE levels by Day 4 was independently associated with improved outcomes, with a 10% increase corresponding to a 16% reduction in 30-day mortality risk (OR = 0.84).
Similarly, Schuurman et al. [23] assessed monocyte and neutrophil lipidomes at baseline and one month following CAP. While lipid perturbations in monocytes largely resolved during convalescence, neutrophils exhibited sustained alterations, including persistent elevations in polyunsaturated TGs and DGs.
In pediatric populations, Virgiliou et al. [15] conducted serial lipidomic analyses in patients with suspected VAP across four timepoints (Days 1, 3, 6, and 12). A gradual increase in PCs and TGs was observed in the high-suspicion group, parallel with microbiological confirmation and clinical deterioration. Specific lipids such as TG(48:3) and SM(40:1) showed increasing discrimination power over time (AUC > 0.75).
Prognostically, several studies reported lipid-based models that outperformed traditional clinical scores. In the study by Chen et al. [13], phosphatidylcholine species PC(18:2_20:4) and PC(38:6) achieved AUCs of 0.954 and 0.959, respectively, for predicting CAP severity—superior to CURB-65 and the PSI. Likewise, Ma et al. [22] identified that elevated PC/LPC and PE/LPE ratios were positively associated with disease severity, inflammatory markers, and prolonged hospital stay, and independently predicted 30-day mortality (AUC = 0.838 for PC/LPC ratio).
Longitudinal patterns were also evident. Zheng et al. [25] linked BALF lipid clusters to CAP severity, with certain fatty acids discriminating severe vs. non-severe disease. Sun et al. [26] validated lipid-driven TB signatures across training and validation cohorts with very high reproducibility. Chen et al. [28] demonstrated that plasma metabolite panels predicted not only infection vs. health but also disease phenotype, correlating with neutrophil–lymphocyte ratios. Tomalka et al. [29] showed that eicosanoid precursors correlated with WHO severity scores and innate immune suppression, suggesting prognostic utility. De Almeida et al. [30] reported near-perfect reproducibility of lipidomic/proteomic classifiers across independent patient sets.
3.6. Integrative Models and Clinical Translation
Several included studies employed multivariate statistical and machine learning techniques to build predictive models based on lipidomic features. In a retrospective cohort of hospitalized COVID-19 patients, Castané et al. [18] used Monte Carlo simulations, principal component analysis (PCA), and partial least squares discriminant analysis (PLS-DA) to identify a panel of discriminatory lipids, including LPC(22:6-sn2) and PC(36:1). The derived ratio LPC(22:6-sn2)/PC(36:1) yielded an AUC exceeding 0.95 for distinguishing COVID-19 from other infectious or inflammatory conditions.
Similarly, Chen et al. [13] developed a ROC-based classifier using individual phosphatidylcholines (e.g., PC(38:6) and PC(18:2_20:4)) that outperformed traditional clinical severity scores such as CURB-65 and PSI, with AUCs > 0.95. These models were not only statistically robust but also biologically interpretable, as the lipid species selected were implicated in inflammation, cellular stress, and membrane dynamics.
The integration of lipidomics with other omics platforms further enhanced discriminatory power and mechanistic understanding. Rischke et al. [12] combined lipidomics with proteomics and metabolomics to differentiate viral from bacterial CAP, demonstrating that pathogen-specific clusters could be generated based on co-expression networks of linoleic acid–derived oxylipins, triglycerides, and host immune markers such as TNF-related apoptosis-inducing ligand (TRAIL) and Lymphocyte-activation gene 3 (LAG-3). Such multi-omic integration supports not only diagnostic classification but also insights into immune-metabolic crosstalk.
Machine learning approaches further advanced translation. Humes et al. [24] applied Bayesian regression and multinomial models to identify pathogen-specific sputum clusters. Zheng et al. [25] used clustering and ROC analyses to link lipid subsets to CAP severity. Sun et al. [26] leveraged multiple algorithms (LASSO, Random Forest, XGBoost) to derive reproducible TB classifiers. Chen et al. [28] developed a 13-metabolite panel (AUC = 0.927–1.0) for pediatric infection stratification. Tomalka et al. [29] combined lipidomics with RNA-seq to map lipid-mediated immune dysregulation in COVID-19. De Almeida et al. [30] integrated serum lipidomics with MALDI-based proteomics, achieving >98% diagnostic accuracy and near-zero false negatives in COVID-19 screening.
Table 3 displays lipidomics findings by clinical category.

Table 3.
Cross-study tabulation of lipidomics findings by clinical category.
4. Discussion
This systematic review synthesized evidence from nine original studies that employed lipidomic analyses in the context of LRTIs, including CAP, VAP, and COVID-19. Across diverse populations, analytical platforms, and sample types, a consistent pattern of lipid alterations emerged. Dysregulation of PCs and LPCs was the most commonly reported finding, with decreased LPC levels frequently associated with severe disease. Changes in lipid profiles also demonstrated pathogen-specific patterns, distinguishing viral from bacterial etiologies. Furthermore, temporal dynamics of lipid species were linked to clinical outcomes, with early normalization of certain lipid classes (e.g., cholesterol esters) associated with reduced mortality. Importantly, lipidomic biomarkers often outperformed conventional clinical scores in predictive models, underscoring their potential clinical utility.
Lipidomic analyses have provided critical mechanistic insights into how LRTIs disrupt host lipid metabolism. One of the most consistent findings across studies is the downregulation of LPCs and the alteration of PC/LPC and PE/LPE ratios, particularly in severe CAP and COVID-19 cases. These changes suggest involvement of phospholipid remodeling pathways regulated by LPCAT1/2, enzymes known to be modulated in response to inflammatory stress, potentially reflecting the host’s attempt to maintain membrane integrity and immune regulation [13,18,19,22].
Oxylipins, which are oxygenated derivatives of polyunsaturated fatty acids, were also differentially expressed across pathogen types. Elevated concentrations of 12,13-EpOME and 9,10-DiHOME in viral CAP highlight increased cytochrome P450 and epoxide hydrolase activity, which may mediate pro-inflammatory or vasoactive responses. Conversely, reduced levels of 9/13-HODE and 15-HETE observed in bacterial pneumonia may reflect suppressed resolution-phase signaling and altered lipid mediator balance [12,18].
Additionally, sphingolipid metabolism has emerged as a key regulatory axis in the host response to LRTIs. The upregulation of S1P signaling and its biosynthetic enzyme SPHK1, along with changes in specific SM species (e.g., SM 34:1), suggest a mechanistic link to monocyte and neutrophil activation, as well as cytokine production [21,23]. These alterations likely contribute to broader immunometabolic rewiring and may perpetuate inflammation or immune exhaustion during infection.
Finally, multiple studies reported elevated acylcarnitines and TGs in patients with viral pneumonia, indicating disrupted mitochondrial β-oxidation and enhanced hepatic lipogenesis, respectively—hallmarks of systemic metabolic stress and energy imbalance during acute infection [12,18].
Recent evidence highlights the role of sphingosine-1-phosphate (S1P) signaling in stabilizing the alveolar endothelial–epithelial barrier. Activation of S1PR1 promotes actin cytoskeleton remodeling and adherens-junction assembly through Rac1/Cdc42 signaling, thereby reducing vascular permeability, while S1PR2 activation exerts opposing barrier-disruptive effects [31].
Epithelial damage markers could further contextualize the present findings. Circulating sRAGE reflects type I alveolar epithelial injury and predicts worse outcomes in ARDS [32]. Surfactant protein D (SP-D) is a sensitive marker of type II pneumocyte injury [33], while KL-6/MUC1 is released from regenerating pneumocytes and is correlated with severity of epithelial damage [34]. More recent data suggest that lung tissue expression of RAGE and elafin, but not SP-D, is closely related to histopathological injury severity [35].
Induction of pro-inflammatory cytokines such as TNF-α, IL-1β, IL-6, IL-8, and CCL2 has profound effects on lipid signaling. These cytokines activate sPLA2, COX, and LOX pathways, leading to oxylipin generation [36]. In parallel, inflammatory stress increases ceramide accumulation, which promotes alveolar leaks and apoptosis [37].
Despite the promising results, several technical limitations warrant consideration. First, there is significant heterogeneity in analytical platforms (e.g., LC-MS/MS, UHPLC-HRMS, NMR), lipid extraction protocols, and bioinformatic pipelines, which limits direct comparison across studies. Second, many studies utilized untargeted lipidomics, which, while comprehensive, may suffer from challenges in lipid identification, quantification, and inter-laboratory reproducibility. Additionally, normalization techniques, internal standards, and statistical thresholds varied widely across studies, potentially influencing results.
Sample types and timing also varied considerably—ranging from serum to tracheal aspirates and from single to multi-timepoint sampling—which complicates harmonization. Finally, few studies validated their findings in independent cohorts, and even fewer integrated lipidomic data with clinical decision-making tools or prospective outcome measures.
This systematic review has several limitations that should be acknowledged. First, the number of eligible studies was relatively small (n = 9), reflecting the early and still emerging nature of lipidomic applications in LRTIs. As a result, some conclusions are based on limited data, and findings may not be generalizable across all LRTI subtypes, geographic regions, or patient populations. Second, the heterogeneity in study designs, sample types, disease severity, and lipidomic platforms precluded quantitative synthesis through meta-analysis. Variations in analytical methods—including differences in mass spectrometry instrumentation, lipid extraction protocols, normalization strategies, and data processing workflows—introduce methodological variability that complicates direct comparisons across studies.
Third, while most studies achieved high scores on the Newcastle–Ottawa Scale, many lacked external validation cohorts, used small sample sizes, or did not control for potential confounders such as comorbidities, medication use, or nutritional status, all of which can influence the lipidome. Beyond technical heterogeneity, methodological limitations of the included studies should also be considered. Several cohorts may have been subject to selection bias, as recruitment was often restricted to hospitalized or ICU patients, thereby limiting generalizability to milder disease. Moreover, potential confounders such as comorbidities, medication use (e.g., statins, corticosteroids), nutritional status, and prior infections were not consistently controlled for, despite their known influence on the circulating lipidome. These factors may have impacted both the validity and reproducibility of lipidomic associations reported. Future studies should therefore incorporate rigorous adjustment for such confounders and aim to stratify analyses by comorbidity burden and baseline characteristics.
Additionally, few studies conducted longitudinal sampling or integrated lipidomic data with real-time clinical decision-making, limiting insights into the temporal utility of lipid biomarkers in routine care. Fourth, publication bias may have influenced the findings, as studies with null or non-significant results may be underrepresented in the published literature. Moreover, only English-language articles were included, potentially excluding relevant non-English studies. Finally, while this review included both adult and pediatric populations, age-specific differences in lipid metabolism were not separately analyzed due to the limited number of pediatric studies, which may have masked age-related variability in lipidomic signatures.
Despite these limitations, this review provides the first comprehensive synthesis of the current evidence on the role of lipidomics in LRTIs. Future studies addressing the above limitations are needed to validate and expand upon these preliminary findings.
Several gaps remain in the current literature. Most notably, large-scale, multi-center studies are lacking, and many of the available data are derived from pilot or single-center cohorts with modest sample sizes. Future research should prioritize validation of lipid-based biomarkers in diverse patient populations and clinical settings. There is also a need for standardized protocols and reporting frameworks for lipidomic studies, akin to the Minimum Information About a Metabolomics Experiment (MIAME) guidelines used in transcriptomics.
Moreover, integrative multi-omics approaches—including transcriptomics, proteomics, and metabolomics—should be employed more consistently to contextualize lipidomic changes within broader biological networks. Functional studies exploring how specific lipid mediators (e.g., oxylipins, sphingolipids) influence immune cell behavior during infection would also enhance mechanistic understanding.
Finally, efforts should be directed toward translating lipidomic findings into clinically applicable tools, such as rapid point-of-care tests or algorithm-based risk stratification models. Collaboration between clinicians, analytical chemists, and computational biologists will be critical to realizing the translational potential of lipidomics in LRTIs.
5. Conclusions
This systematic review highlights the emerging role of lipidomics in unraveling the complex host response to respiratory tract infections. Across diverse clinical settings and infection types, lipidomic profiling has revealed consistent perturbations in key lipid classes—particularly phospholipids, sphingolipids, oxylipins, and acylcarnitines—reflecting alterations in membrane remodeling, immune activation, and systemic metabolic stress. The diagnostic and prognostic potential of lipid-based biomarkers, including specific lipid ratios and composite classifiers, appears promising and, in some studies, exceeds the predictive performance of conventional clinical scoring systems. Moreover, the integration of lipidomics with proteomic and transcriptomic data offers mechanistic insights into pathogen-specific immune pathways and points toward precision medicine approaches in infectious disease management. Nevertheless, substantial methodological heterogeneity, limited validation cohorts, and the lack of standardization across platforms currently constrain clinical translation. Future research should focus on multi-center, longitudinal studies with standardized protocols, age- and pathogen-stratified analyses, and integration with other omics technologies. Overall, lipidomics holds substantial potential not only as a tool for biomarker discovery but also for advancing our understanding of host–pathogen interactions in LRTIs. To bridge discovery with practice, translational studies are urgently required, including prospective clinical trials that test the integration of lipid panels into clinical decision-making algorithms for diagnosis, risk stratification, and treatment guidance. Such efforts will determine whether lipidomics can move beyond biomarker discovery toward actionable tools in real-world patient management.
Author Contributions
Conceptualization, V.E.G. and K.D.; methodology, V.E.G.; software, K.D.; validation, V.E.G., K.D. and V.C.P.; formal analysis, V.E.G.; investigation, K.D.; resources, V.C.P.; data curation, V.E.G.; writing—original draft preparation, V.E.G.; writing—review and editing, K.D. and V.C.P.; visualization, V.E.G.; supervision, V.C.P.; project administration, K.D.; funding acquisition, V.C.P. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable, no new data were created or analyzed in this study.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Georgakopoulou, V.E. Insights from respiratory virus co-infections. World J. Virol. 2024, 13, 98600. [Google Scholar] [CrossRef] [PubMed]
- Georgakopoulou, V.E.; Lempesis, I.G.; Tarantinos, K.; Sklapani, P.; Trakas, N.; Spandidos, D.A. Atypical pneumonia (Review). Exp. Ther. Med. 2024, 28, 424. [Google Scholar] [CrossRef]
- Hanage, W.P.; Schaffner, W. Burden of Acute Respiratory Infections Caused by Influenza Virus, Respiratory Syncytial Virus, and SARS-CoV-2 with Consideration of Older Adults: A Narrative Review. Infect. Dis. Ther. 2025, 14 (Suppl. S1), 5–37. [Google Scholar] [CrossRef]
- Liapikou, A.; Torres, A. The clinical management of lower respiratory tract infections. Expert Rev. Respir. Med. 2016, 10, 441–452. [Google Scholar] [CrossRef] [PubMed]
- Lim, W.S. Pneumonia—Overview. Encycl. Respir. Med. 2022, 185–197. [Google Scholar] [CrossRef]
- Wenk, M.R. Lipidomics of host-pathogen interactions. FEBS Lett. 2006, 580, 5541–5551. [Google Scholar] [CrossRef]
- Han, X.; Gross, R.W. Shotgun lipidomics: Multidimensional MS analysis of cellular lipidomes. Expert Rev. Proteom. 2005, 2, 253–264. [Google Scholar] [CrossRef]
- Anh, N.K.; Thu, N.Q.; Tien, N.T.N.; Long, N.P.; Nguyen, H.T. Advancements in Mass Spectrometry-Based Targeted Metabolomics and Lipidomics: Implications for Clinical Research. Molecules 2024, 29, 5934. [Google Scholar] [CrossRef]
- Cajka, T.; Fiehn, O. Comprehensive analysis of lipids in biological systems by liquid chromatography-mass spectrometry. TrAC Trends Anal. Chem. 2014, 61, 192–206. [Google Scholar] [CrossRef]
- Wu, Z.; Shon, J.C.; Liu, K.H. Mass Spectrometry-based Lipidomics and Its Application to Biomedical Research. J. Lifestyle Med. 2014, 4, 17–33. [Google Scholar] [CrossRef] [PubMed]
- Schenck, E.J.; Plataki, M.; Wheelock, C.E. A Lipid Map for Community-acquired Pneumonia with Sepsis: Observation Is the First Step in Scientific Progress. Am. J. Respir. Crit. Care Med. 2024, 209, 903–904. [Google Scholar] [CrossRef]
- Rischke, S.; Gurke, R.; Zielbauer, A.S.; Ziegler, N.; Hahnefeld, L.; Köhm, M.; Kannt, A.; Vehreschild, M.J.; Geisslinger, G.; Rohde, G.; et al. Proteomic, metabolomic and lipidomic profiles in community acquired pneumonia for differentiating viral and bacterial infections. Sci. Rep. 2025, 15, 1922. [Google Scholar] [CrossRef]
- Chen, L.; Zheng, Y.; Zhao, L.; Zhang, Y.; Yin, L.; He, Y.; Ma, X.; Xu, Y.; Gao, Z. Lipid profiles and differential lipids in serum related to severity of community-acquired pneumonia: A pilot study. PLoS ONE 2021, 16, e0245770. [Google Scholar] [CrossRef]
- Shan, J.; Qian, W.; Shen, C.; Lin, L.; Xie, T.; Peng, L.; Xu, J.; Yang, R.; Ji, J.; Zhao, X. High-resolution lipidomics reveals dysregulation of lipid metabolism in respiratory syncytial virus pneumonia mice. RSC Adv. 2018, 8, 29368–29377. [Google Scholar] [CrossRef]
- Virgiliou, C.; Begou, O.; Ftergioti, A.; Simitsopoulou, M.; Sdougka, M.; Roilides, E.; Theodoridis, G.; Gika, H.; Iosifidis, E. Untargeted Blood Lipidomics Analysis in Critically Ill Pediatric Patients with Ventilator-Associated Pneumonia: A Pilot Study. Metabolites 2024, 14, 466. [Google Scholar] [CrossRef]
- Page, M.J.; McKenzie, J.E.; Bossuyt, P.M.; Boutron, I.; Hoffmann, T.C.; Mulrow, C.D.; Shamseer, L.; Tetzlaff, J.M.; Akl, E.A.; Brennan, S.E.; et al. The PRISMA 2020 statement: An updated guideline for reporting systematic reviews. BMJ 2021, 372, n71. [Google Scholar] [CrossRef] [PubMed]
- Wells, G.A.; Shea, B.; O’Connell, D.; Peterson, J.; Welch, V.; Losos, M.; Tugwell, P. The Newcastle-Ottawa Scale (NOS) for Assessing the Quality of Nonrandomised Studies in Meta-Analyses. Available online: https://www.ohri.ca/programs/clinical_epidemiology/oxford.asp (accessed on 18 September 2025).
- Castañé, H.; Iftimie, S.; Baiges-Gaya, G.; Rodríguez-Tomàs, E.; Jiménez-Franco, A.; López-Azcona, A.F.; Garrido, P.; Castro, A.; Camps, J.; Joven, J. Machine learning and semi-targeted lipidomics identify distinct serum lipid signatures in hospitalized COVID-19-positive and COVID-19-negative patients. Metabolism 2022, 131, 155197. [Google Scholar] [CrossRef]
- Chouchane, O.; Schuurman, A.R.; Reijnders, T.D.Y.; Peters-Sengers, H.; Butler, J.M.; Uhel, F.; Schultz, M.J.; Bonten, M.J.; Cremer, O.L.; Calfee, C.S.; et al. The Plasma Lipidomic Landscape in Patients with Sepsis due to Community-acquired Pneumonia. Am. J. Respir. Crit. Care Med. 2024, 209, 973–986. [Google Scholar] [CrossRef] [PubMed]
- Saballs, M.; Parra, S.; Martínez, N.; Amigo, N.; Cabau, L.; Iftimie, S.; Pavon, R.; Gabaldó, X.; Correig, X.; Paredes, S.; et al. Lipidomic and metabolomic changes in community-acquired and COVID-19 pneumonia. J. Lipid Res. 2024, 65, 100622. [Google Scholar] [CrossRef] [PubMed]
- Kassa-Sombo, A.; Verney, C.; Pasquet, A.; Vaidie, J.; Brea, D.; Vasseur, V.; Cezard, A.; Lefevre, A.; David, C.; Piver, E.; et al. Lipidomic signatures of ventilator-associated pneumonia in COVID-19 ARDS patients: A new frontier for diagnostic biomarkers. Ann. Intensive Care 2025, 15, 78. [Google Scholar] [CrossRef]
- Ma, X.; Chen, L.; He, Y.; Zhao, L.; Yu, W.; Xie, Y.; Yu, Y.; Xu, Y.; Zheng, Y.; Li, R.; et al. Targeted lipidomics reveals phospholipids and lysophospholipids as biomarkers for evaluating community-acquired pneumonia. Ann. Transl. Med. 2022, 10, 395. [Google Scholar] [CrossRef]
- Schuurman, A.R.; Chouchane, O.; Butler, J.M.; Peters-Sengers, H.; Joosten, S.; Brands, X.; Haak, B.W.; Otto, N.A.; Uhel, F.; Klarenbeek, A.; et al. The shifting lipidomic landscape of blood monocytes and neutrophils during pneumonia. JCI Insight 2024, 9, e164400. [Google Scholar] [CrossRef]
- Humes, S.T.; Iovine, N.; Prins, C.; Garrett, T.J.; Lednicky, J.A.; Coker, E.S.; Sabo-Attwood, T. Association between lipid profiles and viral respiratory infections in human sputum samples. Respir. Res. 2022, 23, 177. [Google Scholar] [CrossRef]
- Zheng, Y.; Ning, P.; Luo, Q.; He, Y.; Yu, X.; Liu, X.; Chen, Y.; Wang, X.; Kang, Y.; Gao, Z. Inflammatory responses relate to distinct bronchoalveolar lavage lipidome in community-acquired pneumonia patients: A pilot study. Respir. Res. 2019, 20, 82. [Google Scholar] [CrossRef]
- Sun, G.; Wang, Q.; Shan, X.; Kuerbanjiang, M.; Ma, R.; Zhou, W.; Sun, L.; Li, Q. Metabolomics and lipidomics of plasma biomarkers for tuberculosis diagnostics using UHPLC-HRMS. Front. Cell. Infect. Microbiol. 2025, 15, 1526740. [Google Scholar] [CrossRef]
- Zhang, C.; Cai, M.; Cai, H.; Chen, X. Plasma lipidomic analysis reveals distinct lipid alterations in patients with pulmonary tuberculosis. Eur. J. Med. Res. 2025, 30, 566. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Yu, Q.; Wei, H.; Hu, X.; Wang, H.; Wei, M.; Xu, J.; Zhao, J. Integration of lipidomics and metabolomics reveals plasma and urinary profiles in children with Mycoplasma pneumoniae pneumonia. Biomed. Chromatogr. 2023, 37, e5702. [Google Scholar] [CrossRef]
- Tomalka, J.A.; Owings, A.; Galeas-Pena, M.; Ziegler, C.G.K.; Robinson, T.O.; Wichman, T.G.; Laird, H.; Williams, H.B.; Ghaliwal, N.S.; Everman, S.; et al. Enhanced production of lipid mediators in plasma and activation of DNA damage pathways in PBMCs are correlated with the severity of ancestral SARS-CoV-2 infection. FASEB J. 2025, 39, e70600. [Google Scholar] [CrossRef] [PubMed]
- de Almeida, R.A.M.; Souza, L.A.; de Oliveira, M.A.; do Carmo, A.P.; Oliveira, L.L.; Arêas, G.P.T.; Ferreira, C.R.; Catharino, R.R. Diagnostic screening of COVID-19 based on multiomics data by high-resolution mass spectrometry. Sci. Rep. 2025, 15, 11849. [Google Scholar] [CrossRef]
- Al-Kuraishy, H.M.; Al-Gareeb, A.I.; Alexiou, A.; Papadakis, M.; Batiha, G.E. Receptor-Dependent Effects of Sphingosine-1-Phosphate (S1P) in COVID-19: The Black Side of the Moon. Mol. Cell. Biochem. 2023, 478, 2271–2279. [Google Scholar] [CrossRef]
- Uchida, T.; Shirasawa, M.; Ware, L.B.; Kojima, K.; Hata, Y.; Makita, K.; Mednick, G.; Matthay, M.A. Receptor for Advanced Glycation End-Products Is a Marker of Type I Cell Injury in Acute Lung Injury. Am. J. Respir. Crit. Care Med. 2006, 173, 1008–1015. [Google Scholar] [CrossRef] [PubMed]
- Ware, L.B.; Koyama, T.; Zhao, Z.; Janz, D.R.; Wickersham, N.; Bernard, G.R.; Matthay, M.A. Biomarkers of Lung Epithelial Injury and Inflammation Distinguish Severe Sepsis Patients with Acute Respiratory Distress Syndrome. Crit. Care 2013, 17, R253. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, N.; Hattori, N.; Yokoyama, A.; Kohno, N. Utility of KL-6/MUC1 in the Clinical Management of Interstitial Lung Diseases. Respir. Investig. 2012, 50, 3–13. [Google Scholar] [CrossRef]
- de Souza Xavier Costa, N.; da Costa Sigrist, G.; Schalch, A.S.; Belotti, L.; Dolhnikoff, M.; da Silva, L.F.F. Lung Tissue Expression of Epithelial Injury Markers Is Associated with Acute Lung Injury Severity but Does Not Discriminate Sepsis from ARDS. Respir. Res. 2024, 25, 129. [Google Scholar] [CrossRef]
- Dennis, E.A.; Cao, J.; Hsu, Y.H.; Magrioti, V.; Kokotos, G. Phospholipase A2 Enzymes: Physical Structure, Biological Function, Disease Implication, Chemical Inhibition, and Therapeutic Intervention. Chem. Rev. 2011, 111, 6130–6185. [Google Scholar] [CrossRef]
- Petrache, I.; Natarajan, V.; Zhen, L.; Medler, T.R.; Richter, A.T.; Cho, C.; Hubbard, W.C.; Berdyshev, E.V. Ceramide Upregulation Causes Pulmonary Cell Apoptosis and Emphysema-Like Disease in Mice. Nat. Med. 2005, 11, 491–498. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).