Abstract
Spontaneous intracerebral hemorrhage (ICH) is associated with substantial mortality and morbidity. Current management paradigms rely heavily on the rapid interpretation of neuroimaging and clinical data, yet are frequently constrained by limitations in processing speed, diagnostic accuracy, and prognostic precision. Artificial intelligence (AI), specifically machine learning (ML) and deep learning (DL), offers transformative potential to circumvent these challenges across the entire continuum of ICH care. This comprehensive review synthesizes the rapidly evolving landscape of AI applications in ICH management. Through a systematic evaluation of recent literature, we examine studies focused on the development, validation, or critical appraisal of AI-driven technologies for ICH care. Our analysis encompasses automated neuroimaging, computer-assisted surgical navigation, brain–computer interfaces (BCIs), prognostic modeling, and fundamental research into disease mechanisms. AI has demonstrated performance comparable to that of clinical experts in automating hematoma segmentation, predicting complications such as hematoma expansion, and refining surgical planning via augmented reality. Furthermore, BCIs present innovative therapeutic avenues for motor rehabilitation. However, the translation of these technological advances into routine clinical practice is impeded by substantial challenges, including data heterogeneity, model opacity (“black-box” issues), workflow integration barriers, regulatory ambiguities, and ethical concerns surrounding accountability and algorithmic bias. The integration of AI into ICH care signifies a paradigm shift from standardized treatment protocols toward dynamic, precision medicine. Realizing this vision necessitates interdisciplinary collaboration to engineer robust, generalizable, and interpretable AI systems. Key priorities include the establishment of large-scale multimodal data repositories, the advancement of explainable AI (XAI) frameworks, the execution of rigorous prospective clinical trials to validate efficacy, and the implementation of adaptive regulatory and ethical guidelines. By systematically addressing these barriers, AI can evolve from a mere analytical tool into an indispensable clinical partner, ultimately optimizing patient outcomes.
1. Introduction
Stroke remains a leading cause of mortality and permanent disability globally [1,2,3]. Among stroke subtypes, spontaneous intracerebral hemorrhage (ICH)—characterized by the acute rupture of a cerebral vessel and subsequent bleeding into the surrounding parenchyma—constitutes 10–15% of all strokes worldwide. Despite its lower incidence compared to ischemic stroke, ICH is disproportionately responsible for a substantial burden of stroke-related mortality and long-term morbidity [4,5,6]. Notwithstanding advancements in acute neurocritical care, the prognosis for patients with ICH remains dismal: one-month mortality rates approach 40%, and fewer than 20% of survivors achieve functional independence at six months [7,8]. This unfavorable clinical trajectory is driven by a complex pathophysiological cascade, primarily involving acute hematoma expansion, perihematomal edema (PHE) formation, and secondary neuroinflammatory injury [8,9].
Critical decision-making in ICH management—encompassing hematoma evacuation, hemodynamic control, prognostication, and rehabilitation planning—is heavily contingent upon the rapid and precise interpretation of neuroimaging findings and clinical markers. However, current clinical paradigms face inherent limitations. The manual volumetric segmentation of hematomas and PHE on computed tomography (CT) scans is labor-intensive, susceptible to inter-rater variability, and frequently unfeasible in time-sensitive emergency settings [10,11]. Furthermore, the prediction of clinical outcomes and secondary complications, such as hematoma expansion or stroke-associated pneumonia, traditionally relies on clinical grading scales that often lack the granularity required for individual-level precision [12,13].
In this context, AI, particularly machine learning (ML) and deep learning (DL), has emerged as a disruptive technology capable of addressing these long-standing clinical bottlenecks. Recent years have witnessed an explosive growth in AI applications for ICH, transitioning from basic task automation to generating data-driven insights for complex clinical dilemmas. AI algorithms now demonstrate expert-level proficiency in the automated segmentation of intraparenchymal and intraventricular hemorrhages, as well as PHE, facilitating rapid, quantitative, and reproducible volumetric analyses [14,15]. Beyond image processing, ML models are increasingly deployed to predict functional outcomes, in-hospital mortality, and specific sequelae with high fidelity by integrating multimodal datasets encompassing imaging, electronic health records, and genomic profiles [16]. Pioneering research is also exploring AI-augmented surgical planning, leveraging augmented reality (AR) navigation systems and optimizing stereotactic catheter puncture trajectories [11,17].
However, the integration of these powerful computational tools into routine clinical workflows is non-trivial. Challenges regarding model generalizability, data privacy, “black-box” opacity, and ethical accountability must be rigorously addressed before AI can be seamlessly translated to the bedside [18,19,20,21].
Therefore, this comprehensive review aims to synthesize the rapid advancements in AI-driven ICH management. We delineate the key opportunities AI presents across the clinical continuum—from initial neuroimaging interpretation to long-term outcome prediction. Subsequently, we critically examine the significant systemic and technical barriers that hinder widespread clinical adoption. By providing a balanced perspective on both the potential and the pitfalls of these technologies, this review seeks to inform clinicians and researchers, ultimately guiding the future development of robust, equitable, and clinically impactful AI solutions for ICH management.
2. Materials and Methods
This article is designed as a narrative review aimed at providing a conceptually oriented synthesis of recent developments in artificial intelligence (AI) applications for intracerebral hemorrhage (ICH).
To ensure broad coverage of relevant literature, a structured search of major biomedical and engineering databases, including PubMed, Web of Science, and IEEE Xplore, was conducted. The search focused on studies addressing AI-related methodologies and technologies relevant to ICH, such as machine learning, deep learning, neuroimaging analysis, prognostic modeling, surgical navigation, and brain–computer interfaces.
Articles were considered based on their relevance, methodological contribution, and conceptual significance to the evolving landscape of AI in ICH research and clinical practice. Emphasis was placed on representative studies that illustrate key technological directions, methodological challenges, and translational considerations.
Given the narrative nature of this review and the heterogeneity of the included literature, no formal systematic review framework, eligibility criteria, or risk-of-bias assessment was applied. Instead, the review prioritizes thematic integration, interpretative analysis, and critical discussion of emerging trends, limitations, and future perspectives.
3. AI-Powered Clinical and Translational Applications in ICH
3.1. Automated Quantification and Diagnostic Biomarker Discovery
Computed tomography (CT) remains the gold standard for the acute diagnostic evaluation of ICH, facilitating the rapid identification of acute hemorrhage in emergency settings [22]. Crucially, hematoma volume, morphological characteristics, and the extent of perihematomal edema (PHE) serve as primary determinants of patient prognosis [23,24,25]. Conventional techniques for quantifying these parameters—such as the manual ABC/2 volumetric estimation or slice-by-slice region-of-interest manual segmentation—are constrained by significant methodological limitations [26]. These approaches are labor-intensive, exhibit diminished accuracy when applied to irregularly shaped hematomas, and are subject to substantial inter-rater variability, thereby limiting their utility in time-sensitive acute care environments.
To circumvent these bottlenecks, deep learning (DL) architectures, notably U-Net and its derivatives, have been engineered for the automated and high-fidelity delineation of ICH compartments [15,16,27]. For instance, Kuang et al. introduced an uncertainty-aware DL framework employing cross-task supervision for PHE segmentation. By leveraging slice-level labels and pixel-wise structural ICH annotations to generate high-fidelity pseudo-labels, this model achieved Dice similarity coefficients commensurate with fully supervised algorithms [14]. Similarly, Zhang et al. developed a contrastive learning-based architecture featuring dynamic memory banks for the segmentation of minute “spot signs” on multiphase CT angiography (CTA). This model achieved a mean Dice score of 0.638, underscoring the viability of automated volumetric analysis in forecasting hematoma expansion [28]. Expanding beyond hematoma and spot sign analysis, Ma et al. curated the first public benchmark CT dataset (PHE-SICH-CT-IDS) dedicated to PHE segmentation, detection, and radiomic feature extraction. This repository facilitates the standardized benchmarking of DL models, thereby catalyzing subsequent methodological advancements [29].
The quantitative outputs derived from these segmentation networks function as pivotal inputs for downstream diagnostic and prognostic pipelines, enabling robust, data-driven clinical decision support [15,16]. Crucially, the predictive utility of AI extends beyond static anatomical mapping to the identification of dynamic imaging biomarkers. For example, Chen et al. deployed a 3D U-Net to co-segment the hematoma and PHE, formalizing the concept of “Delayed Perihematomal Edema Expansion” (DPE). They identified an absolute PHE expansion exceeding 3.34 mL between days 4–7 and 8–14 post-ictus as a robust predictor of poor functional outcomes [16]. Such model-derived biomarkers equip clinicians with objective criteria to stratify high-risk cohorts who may require targeted therapeutic interventions [14,16]. Concurrently, the rigorous quantification of PHE via uncertainty-aware DL networks introduces a critical dimension to surgical planning by estimating the trajectory of secondary injury and its impending impact on adjacent eloquent cortex [14]. Furthermore, Tong et al. illustrated the direct translation of segmentation accuracy into therapeutic action: their 3D U-Net model not only yielded high Dice scores for intraparenchymal and intraventricular hemorrhages but also facilitated the optimization of stereotactic catheter trajectories, achieving a 96% suitability rate (defined by a centroid deviation < 10 mm or Dice > 0.8) [15]. Together, these innovations delineate a closed-loop, AI-augmented workflow wherein automated segmentation seamlessly informs both prognostic stratification and neurosurgical intervention [15,28].
The convergence of AI-driven analytics with advanced neuroimaging is forging a transformative ecosystem in ICH management, concurrently elevating prognostic precision and democratizing clinical access. Automated segmentation algorithms yield indispensable quantitative biomarkers—ranging from spot sign volumes for predicting hematoma expansion to PHE expansion rates correlated with functional deficits [15,16,28]. These imaging-derived indices are increasingly fused with clinical metadata to engineer sophisticated prognostic models, such as those forecasting 90-day functional recovery or the risk of stroke-associated pneumonia, thereby expediting the triage of high-risk patients [30,31]. This analytical capacity is further augmented by the advent of portable, low-field magnetic resonance imaging (MRI) systems. Empowered by DL-based image reconstruction, these devices demonstrate robust diagnostic accuracy, rendering advanced neuroimaging viable in resource-constrained environments or at the bedside of hemodynamically unstable patients [32]. The synergistic interplay among specialized benchmark datasets, uncertainty-aware segmentation algorithms, and portable imaging modalities establishes a cohesive pipeline spanning from raw image acquisition to actionable clinical intelligence [14,29]. Underpinned by explainable AI (XAI) frameworks and cross-modal architectures, this integrated paradigm ensures that data-driven insights can be reliably deployed across diverse clinical settings, ultimately propelling equitable and personalized care for patients with ICH [33,34].
Beyond conventional data augmentation techniques, generative adversarial networks (GANs) have recently emerged as a powerful paradigm for synthesizing high-fidelity medical images to address data scarcity and class imbalance. In the neuro-oncology domain, Pix2Pix GAN-based frameworks have been shown to generate realistic contrast-enhanced MRI sequences, significantly improving brain tumour classification performance [35]. Similarly, conditional deep convolutional GANs (cDCGANs) have been successfully employed to produce synthetic brain tumour datasets that, when combined with real images, enhance the robustness and generalizability of DL classifiers [36]. These advances hold considerable promise for ICH imaging, where annotated datasets are often limited and hemorrhage or edema regions are highly imbalanced. Adapting such generative augmentation strategies to non-contrast CT or portable MRI could further improve segmentation accuracy and prognostic model stability, particularly in low-resource settings.
3.2. Intelligent Surgical Navigation and Brain–Computer Interface
The integration of AI into the surgical management of ICH is refining both decision-making and technical execution. By building upon automated segmentation, AI provides a dynamic foundation for real-time surgical navigation. Advanced visualization systems, particularly augmented reality (AR) platforms, create detailed 3D virtual models of the hemorrhage and critical brain structures, which are overlaid directly onto the surgeon’s field of view [17]. This integration offers sub-millimeter guidance for tasks ranging from macroscopic trajectory planning to microscopic endoscopic evacuation, effectively bridging preoperative planning with intraoperative action. This capability allows surgeons to navigate complex anatomy with enhanced precision, thereby potentially minimizing damage to eloquent brain regions. Furthermore, the synergy of AI with portable imaging technologies is broadening access to advanced surgical planning. Portable MRI systems, enhanced by DL reconstruction algorithms, have demonstrated high sensitivity and specificity for ICH detection at the bedside in early studies [32]. This innovation could make sophisticated neuroimaging more feasible in resource-limited settings or for critically ill patients who cannot be transported, suggesting a pathway toward more integrated care from point-of-care image acquisition to AI-driven analysis.
Brain–computer interface (BCI) technologies represent one of the most intriguing frontiers in neurorehabilitation; however, their application in intracerebral hemorrhage remains largely experimental. Unlike imaging-based AI tools that are increasingly approaching clinical integration, BCI systems are primarily supported by proof-of-concept studies, small-scale clinical investigations, and translational research. Therefore, the following discussion of BCIs should be interpreted within the context of emerging rather than established clinical practice.
The conceptual foundation for BCI in ICH care lies in addressing the limitations of conventional rehabilitation. While surgical evacuation addresses the macroscopic anatomical problem, a patient’s ultimate quality of life depends on neural functional remodeling—a process where BCI technology may eventually offer novel therapeutic avenues [37]. By establishing a direct communication pathway between the brain and external devices that bypasses peripheral nerves and muscles, BCI could provide a platform for real-time insight into brain states, quantification of neural function, and intervention in neural activity [38].
At the level of neurological function assessment, preliminary research suggests that multimodal BCIs integrating electroencephalography (EEG) and functional near-infrared spectroscopy (fNIRS) may be capable of non-invasively capturing aspects of brain function. For instance, a novel hybrid EEG-fNIRS BCI framework incorporating Wasserstein metric-based transfer learning has been developed, with early results suggesting it may help quantify differences in neurosignals between healthy individuals and ICH patients, potentially enhancing cross-subject decoding performance in small-scale studies [39]. Such approaches are being explored as possible tools for addressing individual variability and objectively assessing neurological function changes before and after surgery, though these applications remain investigational.
In the domain of motor function restoration, preliminary studies suggest that BCI could offer novel approaches for patients with severe paralysis who may not benefit adequately from conventional rehabilitation. Minimally invasive endovascular BCIs, such as the Stentrode device (Synchron, New York, NY, USA), represent a promising technological alternative to traditional implanted systems, with early feasibility studies demonstrating stable signal recording and potential for functional restoration in severely paralyzed patients in home environments [40]. However, these findings derive from highly selected patient populations and require confirmation in larger, more diverse cohorts. Similarly, BCI-driven neuromuscular electrical stimulation (NMES) has shown initial promise for chronic paresis after stroke in proof-of-concept investigations. The approach—utilizing the brain’s own motor intent to drive electrical stimulation—may theoretically promote functional reorganization of motor pathways, but robust evidence from adequately powered trials is currently lacking [41].
Looking forward, precision surgical techniques centered on AI and AR, increasingly accessible diagnostic tools represented by portable imaging, and emerging neurorehabilitation technologies grounded in BCI research collectively hint at an evolving therapeutic landscape for ICH [42]. If validated in future research, these approaches could one day contribute to a more comprehensive care continuum, though at present they remain largely within the domain of clinical investigation rather than routine practice [43].
3.3. Proactive Prognostication
In neurocritical care, the cornerstone of proactive prognostication is the utilization of multimodal clinical data to prospectively identify patients at an elevated risk of adverse trajectories, thereby facilitating targeted and preemptive interventions. The operationalization of this paradigm relies implicitly on the development and rigorous validation of robust predictive algorithms. Recently, ML and allied AI technologies have emerged as primary engines for constructing high-fidelity prognostic models. Their efficacy stems from an intrinsic capacity to process and map high-dimensional, non-linear biomedical data [44,45].
Throughout the acute phase of ICH, patients are vulnerable to a spectrum of severe complications—including nosocomial infections, deep vein thrombosis, and acute symptomatic seizures—all of which independently exacerbate morbidity and mortality. Regarding hospital-acquired infections, stroke-induced immunosuppression and subsequent gut microbiota dysbiosis render ICH patients uniquely susceptible to systemic pathogens [46,47]. Furthermore, the onset of severe infections, such as sepsis, initiates a deleterious feedback loop with the primary central nervous system injury. This exacerbates the systemic inflammatory response syndrome (SIRS) and precipitates multiple organ dysfunction, significantly amplifying mortality and long-term disability [48,49]. Consequently, the preemptive and accurate stratification of complication risk is an indispensable component of proactive neurocritical management.
Conventional risk stratification tools typically depend on isolated biomarkers or reductionist clinical scoring systems, which frequently suffer from suboptimal sensitivity and specificity. In contrast, ML architectures can seamlessly ingest vast, heterogeneous data matrices available at admission—encompassing demographics, continuous vital signs, comprehensive laboratory panels, comorbidities, and acute interventions—to autonomously identify latent prognostic patterns and generate early clinical alerts. Extensive literature confirms that ensemble learning algorithms (e.g., Random Forests, Gradient Boosting Machines) and Recurrent Neural Networks (RNNs) consistently outperform traditional logistic regression models in forecasting diverse clinical end-points [50,51].
A compelling demonstration of this is provided by Tang et al. [52], who systematically engineered and benchmarked multiple ML algorithms to predict sepsis risk among ICH patients utilizing expansive public intensive care unit (ICU) registries. Their analysis revealed the superiority of the Random Forest model, which maintained excellent discriminatory capacity and generalizability across independent internal and external validation cohorts. This performance aligns with the architectural strengths of Random Forests, specifically their resilience to missing data, their ability to model complex multimodal interactions, and their inherent robustness against overfitting in noisy biomedical datasets [53], establishing them as an optimal substrate for prognostic modeling.
Beyond binary risk stratification, ensuring model interpretability is paramount for translating predictions into actionable clinical interventions. The integration of post hoc explanatory frameworks, such as SHapley Additive exPlanations (SHAP), in the aforementioned study elucidated the primary drivers of sepsis risk, highlighting variables such as fluid and electrolyte imbalances, leukocytosis, supplemental oxygen requirements, depressed Glasgow Coma Scale (GCS) scores, and acute kidney injury. These algorithmically derived insights are corroborated by established pathophysiological literature: severe electrolyte derangements independently predict ICU mortality [54], concurrent acute pneumonia exponentially increases the risk of downstream post-stroke sequelae [55], and invasive monitoring or respiratory support are well-established vectors for nosocomial pathogens [56]. Thus, advanced ML models serve a dual imperative: they provide highly sensitive early warning systems while simultaneously isolating critical pathophysiological drivers, thereby directing clinicians toward precision, mechanism-targeted interventions.
Beyond automated image analysis, a critical clinical priority is the accurate prognostication of patient survival and long-term functional independence. Traditional risk stratification tools, such as the classic ICH Score, are clinically pragmatic but often lack the precision required for individualized long-term functional prediction. Recently, machine learning models have demonstrated superior discriminatory ability in forecasting both short-term mortality and long-term functional outcomes.
For instance, studies utilizing ensemble learning algorithms (e.g., Random Forest, XGBoost, and CatBoost) integrated with multimodal data—including admission clinical metrics, laboratory values, and radiomic features—have consistently outperformed traditional scores in predicting 90-day modified Rankin Scale (mRS) scores and mortality rates [57]. Research by Guo et al. highlighted that ML models could achieve an Area Under the Curve (AUC) of nearly 0.89 for predicting 90-day functional recovery, providing a significantly more robust prognostic profile than the conventional ICH Score. Furthermore, predictive models have been specifically tailored for vulnerable populations, such as elderly patients, where predicting the 6-month Glasgow Outcome Scale (GOS) is arguably more critical than mortality alone for establishing realistic goals of care [58]. Similarly, the integration of real-world Electronic Health Record (EHR) data with raw CT imaging has enabled models to achieve superior discrimination and calibration in predicting in-hospital mortality compared to standard clinical grading scales [59]. By uncovering complex, non-linear synergies among clinical variables that traditional regression models might miss, AI equips clinicians with dynamic prognostic tools that can continuously update as the patient’s condition evolves, thereby facilitating more personalized and transparent family counseling and therapeutic decision-making.
3.4. Decoding Disease Mechanisms in Fundamental Research
AI is catalyzing a paradigm shift in fundamental ICH research by empowering a multi-scale, data-driven approach to deciphering underlying pathophysiology. At the molecular echelon, AI-driven structural biology platforms, notably AlphaFold (Google DeepMind, London, UK) and RoseTTAFold (University of Washington, Seattle, WA, USA), generate unprecedented, atomic-resolution models of proteins implicated in ICH pathogenesis. In the context of cerebral amyloid angiopathy (CAA)—a predominant etiology of lobar ICH—these computational tools accelerate the rational design of novel therapeutics aimed at enhancing amyloid-β clearance and restoring cerebrovascular integrity [60,61]. Extending beyond static structural conformations, utilizing AI to model dynamic protein–protein interaction (PPI) networks is elucidating critical molecular cascades, such as the complement system activation that propagates secondary neuroinflammation following hemorrhage [62].
Within systems biology, ML architectures are increasingly deployed to interrogate high-dimensional “multi-omics” datasets to extract latent biological signatures. A paramount application involves the discovery of robust diagnostic and prognostic biomarkers from peripheral biofluids. Specifically regarding CAA, circulating microRNAs (miRNAs) have emerged as highly promising non-invasive biomarkers and potential therapeutic targets, mechanistically bridging the gap between progressive vascular amyloid deposition and acute hemorrhagic rupture [63]. Building upon this framework, ML algorithms have successfully identified distinct circulating miRNA signatures in patient serum that correlate strongly with the risk of hematoma expansion and long-term functional recovery profiles [64]. Furthermore, the fusion of multi-omics data (e.g., transcriptomics, proteomics) via deep learning models is unmasking novel pathogenic networks. For example, algorithmic identification of correlations between iron-metabolism gene expression and delayed PHE underscores ferroptosis as a highly viable therapeutic target for mitigating secondary brain injury [65].
A particularly potent methodological advancement is the integration of bulk and single-cell RNA sequencing (scRNA-seq) datasets utilizing specialized ML pipelines. This synergy not only isolates critical differentially expressed genes (DEGs)—such as ANXA2 and COTL1, which are upregulated post-ictus and mechanistically linked to microglial activation and oxidative stress—but also facilitates the deconvolution of immense cellular heterogeneity, mapping these molecular signatures precisely to their cellular origins (e.g., microglia versus reactive astrocytes) [66]. Advanced computational techniques, such as pseudotime trajectory inference algorithms applied to scRNA-seq matrices, map the dynamic temporal transition of microglia from a homeostatic resting state to a pro-inflammatory phenotype. This high-resolution temporal modeling meticulously characterizes the transcriptional drivers governing this pathogenic phenotypic switch [67]. Ultimately, this AI-powered, multi-scale analytical framework provides an indispensable mechanism for deconvoluting the intricate cellular kinetics and molecular pathways driving secondary neural injury, thereby laying the groundwork for the next generation of precision, cell-type-specific neurotherapeutics.
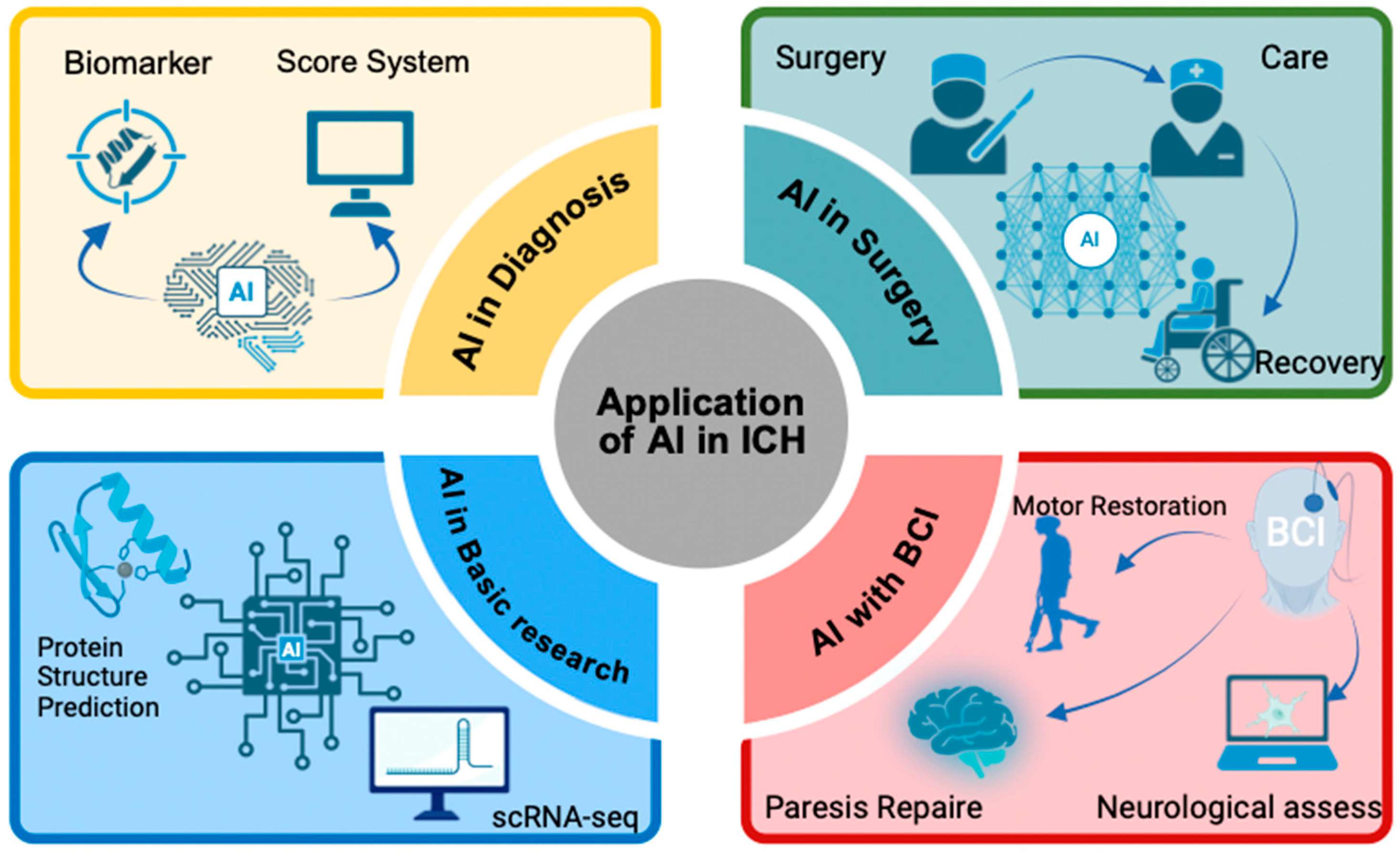
Figure 1 illustrates from four perspectives: diagnosis, treatment, brain–computer interfaces, and basic research.
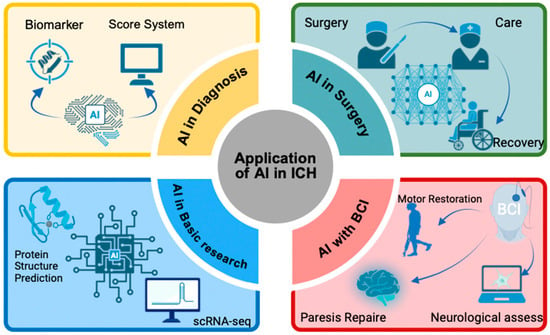
Figure 1.
The applications of AI in ICH.
3.5. Critical Appraisal of Current Evidence
The technological advances surveyed across Section 3.1, Section 3.2, Section 3.3 and Section 3.4 illustrate the remarkable breadth of AI applications in ICH research. However, a critical examination reveals substantial heterogeneity in methodological rigor and clinical readiness.
In automated image segmentation, while deep learning models frequently report Dice coefficients exceeding 0.9, most studies remain retrospective, single-center designs utilizing curated datasets that may not reflect real-world heterogeneity. Sample sizes vary considerably, with some studies training on fewer than 200 scans. External validation—the gold standard for assessing transportability—remains exceptional rather than routine. Crucially, few studies demonstrate that improved segmentation accuracy translates into meaningful changes in clinical decision-making or patient outcomes.
For prognostic prediction models, although machine learning approaches often outperform traditional regression in internal validation, performance degradation upon external validation is well-documented. Most models are developed on modest-sized datasets and lack temporal or geographic validation. Calibration—essential for clinical utility—is frequently underreported, and prospective studies demonstrating improved patient management remain absent.
Surgical navigation research is dominated by technical feasibility studies with sample sizes rarely exceeding 30 patients, precluding robust conclusions about safety or efficacy. BCI research in ICH remains at the proof-of-concept stage, involving highly selected patients under controlled conditions without adequate control groups.
Collectively, the field is characterized by exploratory research with limited confirmatory evidence. The path from technical proof-of-concept to clinically validated tool requires rigorous external validation, prospective evaluation against relevant comparators, and demonstration of meaningful impact on patient-centered outcomes.
6. Discussion
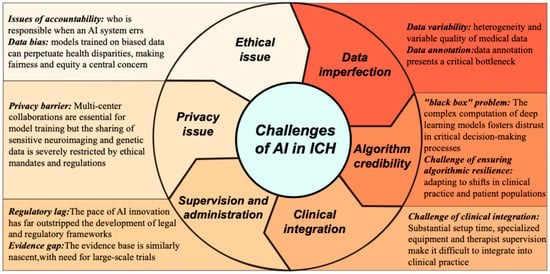
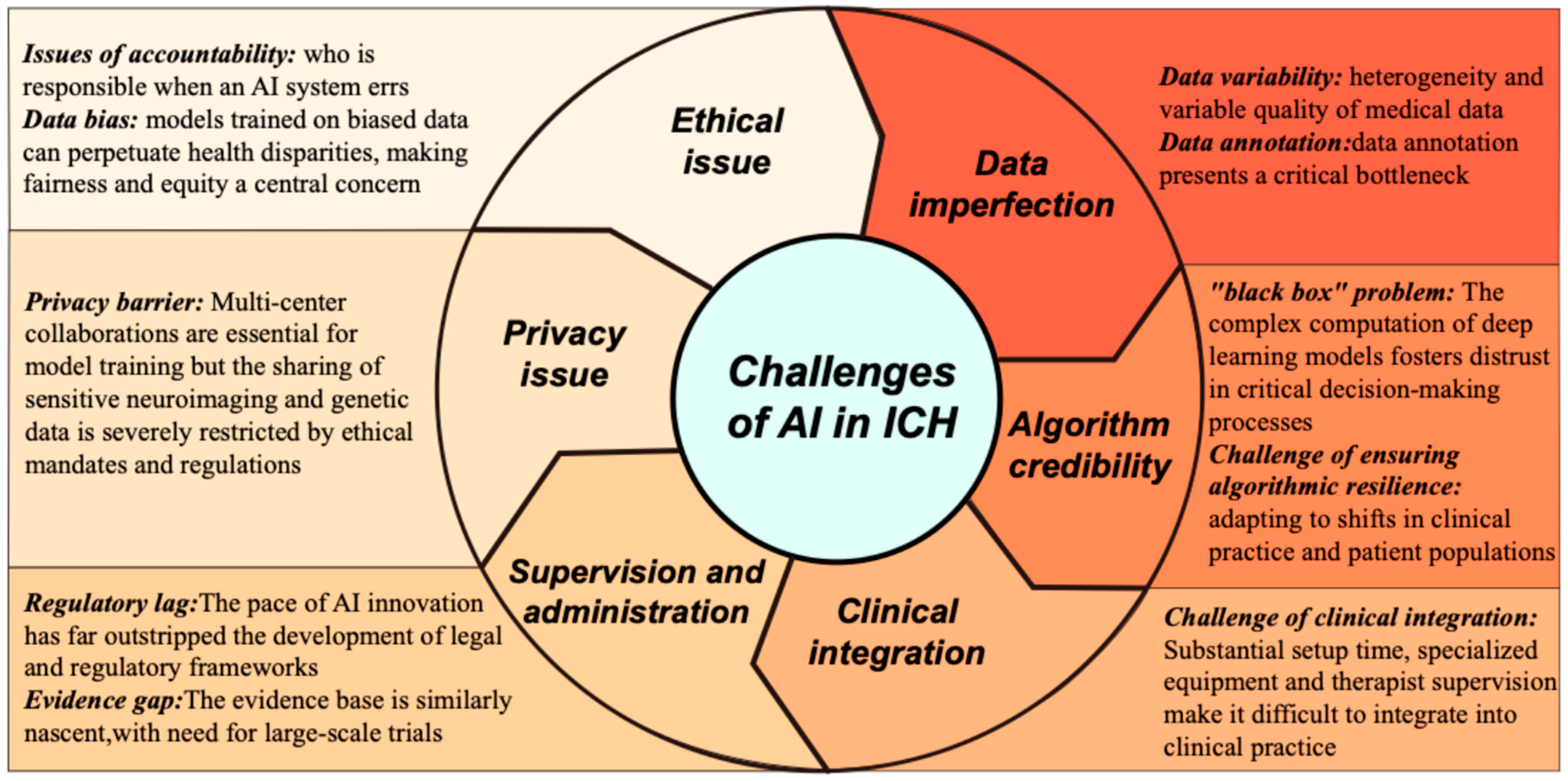
Despite its comprehensive scope, this review is subject to several limitations that should be acknowledged. First, although a systematic search methodology was employed for literature synthesis, no formal quality assessment or risk-of-bias analysis was conducted on the included studies or specific models discussed, which may affect the robustness of the conclusions drawn. Second, foundational research applying AI to investigate ICH pathogenesis or discover biological markers remains scarce, rendering our summary of these aspects necessarily incomplete. Third, the pronounced lack of prospective randomized controlled trials for AI tools in intracerebral hemorrhage—coupled with fundamental computer science challenges such as poor model generalization and high operational costs—forms a major barrier to translational application. This dual deficiency obscures not only the true clinical value but also the cost-effectiveness and practical feasibility of large-scale implementation for many models across diverse healthcare environments. Finally, while our review primarily focuses on the technical and clinical aspects of AI integration and touches upon ethical and regulatory challenges at a societal level, a deeper analysis of socio-technical barriers—including physician acceptance, workflow redesign, and health economic implications—lies beyond the scope of this article, though these factors are undeniably critical for successful and widespread implementation. Beyond these methodological considerations, a critical appraisal of the existing literature must also recognize the substantial heterogeneity in evidentiary maturity across studies. As illustrated in Table 1, AI research in ICH spans a spectrum from exploratory retrospective model-development studies—which remain the most prevalent—to externally validated predictive models, and, in rare instances, prospective clinical evaluations or real-world implementations. This distinction is not merely taxonomic; it carries direct implications for clinical interpretability and translational readiness. Models that have undergone rigorous external validation [31,33,34] warrant greater confidence than those confined to single-center retrospective derivation, while prospective evaluations [32,99]—though still exceptional—provide the most compelling evidence of real-world utility. Future research should explicitly position its contributions along this spectrum and prioritize efforts to advance models from exploratory derivation toward external validation and, ultimately, prospective assessment of clinical impact.

Table 1.
Representative AI Studies in ICH Categorized by Evidentiary Maturity.
Beyond the immediate context of intracerebral hemorrhage, the AI methodologies surveyed in this review—particularly deep learning-based segmentation, multimodal prognostic modeling, and brain–computer interfaces—carry substantial translational potential for other neurological and oncological conditions. For instance, automated hematoma segmentation frameworks such as U-Net and its variants have been successfully adapted to brain tumor delineation in MRI, while generative augmentation techniques originally developed for oncological imaging are poised to enhance ICH datasets. In real-world clinical settings, the integration of these tools into existing workflows could reduce manual measurement variability, expedite triage decisions, and enable precision monitoring across diverse patient populations. However, successful generalization requires careful validation against domain-specific data heterogeneity, imaging protocols, and outcome definitions. Prospective multicenter studies evaluating the portability of ICH-trained models to other hemorrhagic or space-occupying lesions—and vice versa—will be essential to establish robust, cross-disease AI platforms. Such efforts align with the broader vision of a learning healthcare system, where AI continuously refines its performance across institutions and indications, ultimately improving patient outcomes at scale.
7. Conclusions
The integration of artificial intelligence (AI) into the clinical management of intracerebral hemorrhage (ICH) transcends a mere technological upgrade; it heralds a fundamental paradigm shift toward proactive, personalized, and precision neurocritical care. This review has synthesized the rapid evolution of AI applications, from achieving expert-level proficiency in automated neuroimaging analysis to constructing high-fidelity prognostic models and pioneering novel therapeutic modalities via brain–computer interfaces (BCIs). These computational tools possess the undeniable potential to optimize the entire clinical continuum, spanning from hyperacute diagnosis to long-term functional rehabilitation.
Nevertheless, the translational trajectory from algorithmic development to routine bedside deployment remains fraught with formidable systemic barriers. The maturation of promising in silico prototypes into robust clinical instruments is currently impeded by the inherent fragility of models trained on heterogeneous datasets, the pervasive “black-box” opacity that undermines clinical trust, and the profound ethical and regulatory dilemmas precipitated by increasingly autonomous systems. Successfully navigating this translational chasm necessitates a concerted, multidisciplinary consensus among data scientists, clinicians, ethicists, and regulatory bodies.
Crucially, the ultimate objective of medical AI is not to supplant the clinician, but to cultivate a framework of “collaborative intelligence.” Within this paradigm, AI executes data-intensive computational workflows, thereby liberating physicians to concentrate on higher-order diagnostic synthesis, complex clinical decision-making, and the indispensable humanistic dimensions of patient care. By embracing this synergistic vision and systematically resolving extant methodological and regulatory bottlenecks with rigorous empirical validation, the biomedical community can fully leverage AI to redefine the standard of care and substantively improve clinical outcomes for patients afflicted by ICH.
Author Contributions
Conceptualization, Y.J.; methodology, M.J.; formal analysis, L.T.; data curation, Y.C. and Y.S. (Yutong She); writing—original draft preparation, Q.G.; writing—review and editing, M.L.; visualization, Y.S. (Yuxuan Sun). All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Scientific Research Fund of Zhejiang University [grant number XY2025074].
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
No new data were created or analyzed in this study. Data sharing is not applicable to this article.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Feigin, V.L.; Brainin, M.; Norrving, B.; Martins, S.O.; Pandian, J.; Lindsay, P.; Grupper, M.F.; Rautalin, I. World Stroke Organization: Global Stroke Fact Sheet 2025. Int. J. Stroke 2025, 20, 132–144. [Google Scholar] [CrossRef]
- Tu, W.-J.; Wang, L.-D.; Yan, F.; Peng, B.; Hua, Y.; Liu, M.; Ji, X.-M.; Ma, L.; Shan, C.-L.; Wang, Y.-L.; et al. China stroke surveillance report 2021. Mil. Med. Res. 2023, 10, 33. [Google Scholar] [CrossRef]
- Potter, T.B.H.; Tannous, J.; Vahidy, F.S. A Contemporary Review of Epidemiology, Risk Factors, Etiology, and Outcomes of Premature Stroke. Curr. Atheroscler. Rep. 2022, 24, 939–948. [Google Scholar] [CrossRef] [PubMed]
- Hemphill, J.C.; Bonovich, D.C.; Besmertis, L.; Manley, G.T.; Johnston, S.C. The ICH Score. Stroke 2001, 32, 891–897. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, A.I.; Tuhrim, S.; Broderick, J.P.; Batjer, H.H.; Hondo, H.; Hanley, D.F. Spontaneous Intracerebral Hemorrhage. N. Engl. J. Med. 2001, 344, 1450–1460. [Google Scholar] [CrossRef] [PubMed]
- Puy, L.; Parry-Jones, A.R.; Sandset, E.C.; Dowlatshahi, D.; Ziai, W.; Cordonnier, C. Intracerebral haemorrhage. Nat. Rev. Dis. Primers 2023, 9, 14. [Google Scholar] [CrossRef]
- Poon, M.T.; Fonville, A.F.; Al-Shahi Salman, R. Long-term prognosis after intracerebral haemorrhage: Systematic review and meta-analysis. J. Neurol. Neurosurg. Psychiatry 2014, 85, 660–667. [Google Scholar] [CrossRef]
- Magid-Bernstein, J.; Girard, R.; Polster, S.; Srinath, A.; Romanos, S.; Awad, I.A.; Sansing, L.H. Cerebral Hemorrhage: Pathophysiology, Treatment, and Future Directions. Circ. Res. 2022, 130, 1204–1229. [Google Scholar] [CrossRef]
- Zille, M.; Farr, T.D.; Keep, R.F.; Römer, C.; Xi, G.; Boltze, J. Novel targets, treatments, and advanced models for intracerebral haemorrhage. EBioMedicine 2022, 76, 103880. [Google Scholar] [CrossRef]
- Sarraj, A.; Pujara, D.K.; Campbell, B.C. Current State of Evidence for Neuroimaging Paradigms in Management of Acute Ischemic Stroke. Ann. Neurol. 2024, 95, 1017–1034. [Google Scholar] [CrossRef]
- Mead, G.E.; Sposato, L.A.; Sampaio Silva, G.; Yperzeele, L.; Wu, S.; Kutlubaev, M.; Cheyne, J.; Wahab, K.; Urrutia, V.C.; Sharma, V.K.; et al. A systematic review and synthesis of global stroke guidelines on behalf of the World Stroke Organization. Int. J. Stroke 2023, 18, 499–531. [Google Scholar] [CrossRef]
- Kaminogo, M.; Yonekura, M.; Shibata, S. Incidence and outcome of multiple intracranial aneurysms in a defined population. Stroke 2003, 34, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Etminan, N.; Chang, H.-S.; Hackenberg, K.; de Rooij, N.K.; Vergouwen, M.D.I.; Rinkel, G.J.E.; Algra, A. Worldwide Incidence of Aneurysmal Subarachnoid Hemorrhage According to Region, Time Period, Blood Pressure, and Smoking Prevalence in the Population. JAMA Neurol. 2019, 76, 588–597. [Google Scholar] [CrossRef] [PubMed]
- Kuang, Z.; Yan, Z.; Yu, L.; Deng, X.; Hua, Y.; Li, S. Uncertainty-Aware Deep Learning with Cross-Task Supervision for PHE Segmentation on CT Images. IEEE J. Biomed. Health Inform. 2022, 26, 2615–2626. [Google Scholar] [CrossRef]
- Tong, G.; Wang, X.; Jiang, H.; Wu, A.; Cheng, W.; Cui, X.; Bao, L.; Cai, R.; Cai, W. A Deep Learning Model for Automatic Segmentation of Intraparenchymal and Intraventricular Hemorrhage for Catheter Puncture Path Planning. IEEE J. Biomed. Health Inform. 2023, 27, 4454–4465. [Google Scholar] [CrossRef]
- Chen, Y.; Qin, C.; Chang, J.; Liu, Y.; Zhang, Q.; Ye, Z.; Li, Z.; Tian, F.; Ma, W.; Wei, J.; et al. Defining Delayed Perihematomal Edema Expansion in Intracerebral Hemorrhage: Segmentation, Time Course, Risk Factors and Clinical Outcome. Front. Immunol. 2022, 13, 911207. [Google Scholar] [CrossRef]
- Zhu, T.; Jiang, S.; Yang, Z.; Zhou, Z.; Li, Y.; Ma, S.; Zhuo, J. A neuroendoscopic navigation system based on dual-mode augmented reality for minimally invasive surgical treatment of hypertensive intracerebral hemorrhage. Comput. Biol. Med. 2022, 140, 105091. [Google Scholar] [CrossRef] [PubMed]
- Bako, A.T.; Vahidy, F.S. Machine Learning-Based Models for Intracerebral Hemorrhage In-Hospital Mortality Prediction. J. Am. Heart Assoc. 2025, 14, e039398. [Google Scholar] [CrossRef]
- Ironside, N.; Chen, C.-J.; Mutasa, S.; Sim, J.L.; Marfatia, S.; Roh, D.; Ding, D.; Mayer, S.A.; Lignelli, A.; Connolly, E.S. Fully Automated Segmentation Algorithm for Hematoma Volumetric Analysis in Spontaneous Intracerebral Hemorrhage. Stroke 2019, 50, 3416–3423. [Google Scholar] [CrossRef]
- Bonkhoff, A.K.; Grefkes, C. Precision medicine in stroke: Towards personalized outcome predictions using artificial intelligence. Brain 2022, 145, 457–475. [Google Scholar] [CrossRef]
- Gupta, N.S.; Kumar, P. Perspective of artificial intelligence in healthcare data management: A journey towards precision medicine. Comput. Biol. Med. 2023, 162, 107051. [Google Scholar] [CrossRef]
- Hillal, A.; Ullberg, T.; Ramgren, B.; Wassélius, J. Computed tomography in acute intracerebral hemorrhage: Neuroimaging predictors of hematoma expansion and outcome. Insights Imaging 2022, 13, 180. [Google Scholar] [CrossRef]
- Sporns, P.B.; Psychogios, M.-N.; Boulouis, G.; Charidimou, A.; Li, Q.; Fainardi, E.; Dowlatshahi, D.; Goldstein, J.N.; Morotti, A. Neuroimaging of Acute Intracerebral Hemorrhage. J. Clin. Med. 2021, 10, 1086. [Google Scholar] [CrossRef]
- Boulouis, G.; Morotti, A.; Brouwers, H.B.; Charidimou, A.; Jessel, M.J.; Auriel, E.; Pontes-Neto, O.; Ayres, A.; Vashkevich, A.; Schwab, K.M.; et al. Association Between Hypodensities Detected by Computed Tomography and Hematoma Expansion in Patients with Intracerebral Hemorrhage. JAMA Neurol. 2016, 73, 961–968. [Google Scholar] [CrossRef]
- Lv, X.-N.; Deng, L.; Yang, W.-S.; Wei, X.; Li, Q. Computed Tomography Imaging Predictors of Intracerebral Hemorrhage Expansion. Curr. Neurol. Neurosci. Rep. 2021, 21, 22. [Google Scholar] [CrossRef]
- Dsouza, L.B.; Pathan, S.A.; Bhutta, Z.A.; Thomas, S.A.; Momin, U.; Mirza, S.; Elanani, R.; Qureshi, R.; Khalaf, W.; Thomas, S.H. ABC/2 estimation in intracerebral hemorrhage: A comparison study between emergency radiologists and emergency physicians. Am. J. Emerg. Med. 2019, 37, 1818–1822. [Google Scholar] [CrossRef]
- Yu, N.; Yu, H.; Li, H.; Ma, N.; Hu, C.; Wang, J. A Robust Deep Learning Segmentation Method for Hematoma Volumetric Detection in Intracerebral Hemorrhage. Stroke 2022, 53, 167–176. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Horn, M.; Tanaka, K.; Bala, F.; Singh, N.; Benali, F.; Ganesh, A.; Demchuk, A.M.; Menon, B.K.; Qiu, W. Tiny-objective segmentation for spot signs on multi-phase CT angiography via contrastive learning with dynamic-updated positive-negative memory banks. Comput. Biol. Med. 2025, 196, 110613. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.; Li, C.; Du, T.; Qiao, L.; Tang, D.; Ma, Z.; Shi, L.; Lu, G.; Meng, Q.; Chen, Z.; et al. PHE-SICH-CT-IDS: A benchmark CT image dataset for evaluation semantic segmentation, object detection and radiomic feature extraction of perihematomal edema in spontaneous intracerebral hemorrhage. Comput. Biol. Med. 2024, 173, 108342. [Google Scholar] [CrossRef]
- Geng, Z.; Yang, C.; Zhao, Z.; Yan, Y.; Guo, T.; Liu, C.; Wu, A.; Wu, X.; Wei, L.; Tian, Y.; et al. Development and validation of a machine learning-based predictive model for assessing the 90-day prognostic outcome of patients with spontaneous intracerebral hemorrhage. J. Transl. Med. 2024, 22, 236. [Google Scholar] [CrossRef] [PubMed]
- Yan, J.; Zhai, W.; Li, Z.; Ding, L.; You, J.; Zeng, J.; Yang, X.; Wang, C.; Meng, X.; Jiang, Y.; et al. ICH-LR2S2: A new risk score for predicting stroke-associated pneumonia from spontaneous intracerebral hemorrhage. J. Transl. Med. 2022, 20, 193. [Google Scholar] [CrossRef]
- Mazurek, M.H.; Parasuram, N.R.; Peng, T.J.; Beekman, R.; Yadlapalli, V.; Sorby-Adams, A.J.; Lalwani, D.; Zabinska, J.; Gilmore, E.J.; Petersen, N.H.; et al. Detection of Intracerebral Hemorrhage Using Low-Field, Portable Magnetic Resonance Imaging in Patients with Stroke. Stroke 2023, 54, 2832–2841. [Google Scholar] [CrossRef]
- Xia, Y.; Li, J.; Deng, B.; Huang, Q.; Cai, F.; Xie, Y.; Sun, X.; Shi, Q.; Dan, W.; Zhan, Y.; et al. Knowledge Graph-Enhanced Deep Learning Model (H-SYSTEM) for Hypertensive Intracerebral Hemorrhage: Model Development and Validation. J. Med. Internet Res. 2025, 27, e66055. [Google Scholar] [CrossRef]
- Yu, X.; Elazab, A.; Ge, R.; Zhu, J.; Zhang, L.; Jia, G.; Wu, Q.; Wan, X.; Li, L.; Wang, C. ICH-PRNet: A cross-modal intracerebral haemorrhage prognostic prediction method using joint-attention interaction mechanism. Neural Netw. 2025, 184, 107096. [Google Scholar] [CrossRef]
- Onakpojeruo, E.P.; Mustapha, M.T.; Ozsahin, D.U.; Ozsahin, I. Enhanced MRI-based brain tumour classification with a novel Pix2pix generative adversarial network augmentation framework. Brain Commun. 2024, 6, fcae372. [Google Scholar] [CrossRef]
- Onakpojeruo, E.P.; Mustapha, M.T.; Ozsahin, D.U.; Ozsahin, I. A Comparative Analysis of the Novel Conditional Deep Convolutional Neural Network Model, Using Conditional Deep Convolutional Generative Adversarial Network-Generated Synthetic and Augmented Brain Tumor Datasets for Image Classification. Brain Sci. 2024, 14, 559. [Google Scholar] [CrossRef] [PubMed]
- Lebedev, M.A.; Nicolelis, M.A.L. Brain-Machine Interfaces: From Basic Science to Neuroprostheses and Neurorehabilitation. Physiol. Rev. 2017, 97, 767–837. [Google Scholar] [CrossRef] [PubMed]
- Soekadar, S.R.; Birbaumer, N.; Slutzky, M.W.; Cohen, L.G. Brain–machine interfaces in neurorehabilitation of stroke. Neurobiol. Dis. 2015, 83, 172–179. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.; Shi, J.; Tao, B.; Zhao, X.; Zhao, Z.; Li, S.; Xu, Y.; Ding, T.; Zhang, P.; Ye, Q.; et al. A Novel Transfer Learning-Based Hybrid EEG-fNIRS Brain-Computer Interface for Intracerebral Hemorrhage Rehabilitation. Adv. Sci. 2025, 12, e05426. [Google Scholar] [CrossRef]
- Brannigan, J.F.M.; Fry, A.; Opie, N.L.; Campbell, B.C.V.; Mitchell, P.J.; Oxley, T.J. Endovascular Brain-Computer Interfaces in Poststroke Paralysis. Stroke 2024, 55, 474–483. [Google Scholar] [CrossRef]
- Mukaino, M.; Ono, T.; Shindo, K.; Fujiwara, T.; Ota, T.; Kimura, A.; Liu, M.; Ushiba, J. Efficacy of brain-computer interface-driven neuromuscular electrical stimulation for chronic paresis after stroke. J. Rehabil. Med. 2014, 46, 378–382. [Google Scholar] [CrossRef]
- Chen, D.; Zhao, Z.; Zhang, S.; Chen, S.; Wu, X.; Shi, J.; Liu, N.; Pan, C.; Tang, Y.; Meng, C.; et al. Evolving Therapeutic Landscape of Intracerebral Hemorrhage: Emerging Cutting-Edge Advancements in Surgical Robots, Regenerative Medicine, and Neurorehabilitation Techniques. Transl. Stroke Res. 2025, 16, 975–989. [Google Scholar] [CrossRef]
- Biasiucci, A.; Leeb, R.; Iturrate, I.; Perdikis, S.; Al-Khodairy, A.; Corbet, T.; Schnider, A.; Schmidlin, T.; Zhang, H.; Bassolino, M.; et al. Brain-actuated functional electrical stimulation elicits lasting arm motor recovery after stroke. Nat. Commun. 2018, 9, 2421. [Google Scholar] [CrossRef]
- Hou, N.; Li, M.; He, L.; Xie, B.; Wang, L.; Zhang, R.; Yu, Y.; Sun, X.; Pan, Z.; Wang, K. Predicting 30-days mortality for MIMIC-III patients with sepsis-3: A machine learning approach using XGboost. J. Transl. Med. 2020, 18, 462. [Google Scholar] [CrossRef]
- Du, M.; Haag, D.G.; Lynch, J.W.; Mittinty, M.N. Comparison of the Tree-Based Machine Learning Algorithms to Cox Regression in Predicting the Survival of Oral and Pharyngeal Cancers: Analyses Based on SEER Database. Cancers 2020, 12, 2802. [Google Scholar] [CrossRef]
- Berger, B.; Gumbinger, C.; Steiner, T.; Sykora, M. Epidemiologic features, risk factors, and outcome of sepsis in stroke patients treated on a neurologic intensive care unit. J. Crit. Care 2014, 29, 241–248. [Google Scholar] [CrossRef] [PubMed]
- Cheng, Y.; Zan, J.; Song, Y.; Yang, G.; Shang, H.; Zhao, W. Evaluation of intestinal injury, inflammatory response and oxidative stress following intracerebral hemorrhage in mice. Int. J. Mol. Med. 2018, 42, 2120–2128. [Google Scholar] [CrossRef] [PubMed]
- Singer, M.; Deutschman, C.S.; Seymour, C.W.; Shankar-Hari, M.; Annane, D.; Bauer, M.; Bellomo, R.; Bernard, G.R.; Chiche, J.-D.; Coopersmith, C.M.; et al. The Third International Consensus Definitions for Sepsis and Septic Shock (Sepsis-3). JAMA 2016, 315, 801–810. [Google Scholar] [CrossRef] [PubMed]
- Kodali, M.C.; Chen, H.; Liao, F.-F. Temporal unsnarling of brain’s acute neuroinflammatory transcriptional profiles reveals panendothelitis as the earliest event preceding microgliosis. Mol. Psychiatry 2020, 26, 3905–3919. [Google Scholar] [CrossRef]
- Scherpf, M.; Gräßer, F.; Malberg, H.; Zaunseder, S. Predicting sepsis with a recurrent neural network using the MIMIC III database. Comput. Biol. Med. 2019, 113, 103395. [Google Scholar] [CrossRef]
- Aşuroğlu, T.; Oğul, H. A deep learning approach for sepsis monitoring via severity score estimation. Comput. Methods Programs Biomed. 2021, 198, 105816. [Google Scholar] [CrossRef] [PubMed]
- Tang, L.; Li, Y.; Zhang, J.; Zhang, F.; Tang, Q.; Zhang, X.; Wang, S.; Zhang, Y.; Ma, S.; Liu, R.; et al. Machine learning model to predict sepsis in ICU patients with intracerebral hemorrhage. Sci. Rep. 2025, 15, 16326. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Xin, Y.; Li, Q.; Ma, J.; Li, S.; Lv, X.; Lv, W. Empirical study of seven data mining algorithms on different characteristics of datasets for biomedical classification applications. Biomed. Eng. Online 2017, 16, 125. [Google Scholar] [CrossRef] [PubMed]
- Qureshi, A.I.; Huang, W.; Hanley, D.F.; Hsu, C.Y.; Martin, R.H.; Malhotra, K.; Steiner, T.; Suarez, J.I.; Yamamoto, H.; Toyoda, K. Early Hyperchloremia is Independently Associated with Death or Disability in Patients with Intracerebral Hemorrhage. Neurocritical Care 2022, 37, 487–496. [Google Scholar] [CrossRef]
- Koennecke, H.C.; Belz, W.; Berfelde, D.; Endres, M.; Fitzek, S.; Hamilton, F.; Kreitsch, P.; Mackert, B.M.; Nabavi, D.G.; Nolte, C.H.; et al. Factors influencing in-hospital mortality and morbidity in patients treated on a stroke unit. Neurology 2011, 77, 965–972. [Google Scholar] [CrossRef]
- Mehta, A.; Mohamed, A.S.; James, P. Predictors of mortality of severe sepsis among adult patients in the medical Intensive Care Unit. Lung India 2017, 34, 330–335. [Google Scholar] [CrossRef]
- Guo, R.; Zhang, R.; Liu, R.; Liu, Y.; Li, H.; Ma, L.; He, M.; You, C.; Tian, R. Machine Learning-Based Approaches for Prediction of Patients’ Functional Outcome and Mortality after Spontaneous Intracerebral Hemorrhage. J. Pers. Med. 2022, 12, 112. [Google Scholar] [CrossRef]
- Trevisi, G.; Caccavella, V.M.; Scerrati, A.; Signorelli, F.; Salamone, G.G.; Orsini, K.; Fasciani, C.; D’Arrigo, S.; Auricchio, A.M.; D’Onofrio, G.; et al. Machine learning model prediction of 6-month functional outcome in elderly patients with intracerebral hemorrhage. Neurosurg. Rev. 2022, 45, 2857–2867. [Google Scholar] [CrossRef]
- Matsumoto, K.; Ishihara, K.; Matsuda, K.; Tokunaga, K.; Yamashiro, S.; Soejima, H.; Nakashima, N.; Kamouchi, M. Machine Learning-Based Prediction for In-Hospital Mortality After Acute Intracerebral Hemorrhage Using Real-World Clinical and Image Data. J. Am. Heart Assoc. 2024, 13, e036447. [Google Scholar] [CrossRef]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; Ronneberger, O.; Willmore, L.; Ballard, A.J.; Bambrick, J.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Krishna, R.; Wang, J.; Ahern, W.; Sturmfels, P.; Venkatesh, P.; Kalvet, I.; Lee, G.R.; Morey-Burrows, F.S.; Anishchenko, I.; Humphreys, I.R.; et al. Generalized biomolecular modeling and design with RoseTTAFold All-Atom. Science 2024, 384, eadl2528. [Google Scholar] [CrossRef] [PubMed]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Weldon Furr, J.; Morales-Scheihing, D.; Manwani, B.; Lee, J.; McCullough, L.D. Cerebral Amyloid Angiopathy, Alzheimer’s Disease and MicroRNA: miRNA as Diagnostic Biomarkers and Potential Therapeutic Targets. Neuromol. Med. 2019, 21, 369–390. [Google Scholar] [CrossRef]
- Yuan, B.; Zhao, X.-D.; Shen, J.-D.; Chen, S.-J.; Huang, H.-Y.; Zhou, X.-M.; Han, Y.-L.; Zhou, L.-J.; Lu, X.-J.; Wu, Q.; et al. Accurate Activation of SIRT1 Alleviates Ferroptosis in the Early Brain Injury after Subarachnoid Hemorrhage. Oxidative Med. Cell. Longev. 2022, 2022, 9069825. [Google Scholar] [CrossRef]
- Zille, M.; Karuppagounder, S.S.; Chen, Y.; Gough, P.J.; Bertin, J.; Finger, J.; Milner, T.A.; Jonas, E.A.; Ratan, R.R. Neuronal Death After Hemorrhagic Stroke In Vitro and In Vivo Shares Features of Ferroptosis and Necroptosis. Stroke 2017, 48, 1033–1043. [Google Scholar] [CrossRef]
- Du, C.; Wang, C.; Liu, Z.; Xin, W.; Zhang, Q.; Ali, A.; Zeng, X.; Li, Z.; Ma, C. Machine learning algorithms integrate bulk and single-cell RNA data to unveil oxidative stress following intracerebral hemorrhage. Int. Immunopharmacol. 2024, 137, 112449. [Google Scholar] [CrossRef]
- Zhang, M.; Ning, J.; Liu, J.; Sun, Y.; Xiao, N.; Xu, H.; Chen, J. Peripheral blood immune landscape and NXPE3 as a novel biomarker for hypertensive intracerebral hemorrhage risk prediction and targeted therapy. iMeta 2025, 4, e70030. [Google Scholar] [CrossRef]
- Li, C.Y.; Chang, K.J.; Yang, C.F.; Wu, H.Y.; Chen, W.; Bansal, H.; Chen, L.; Yang, Y.P.; Chen, Y.C.; Chen, S.P.; et al. Towards a holistic framework for multimodal LLM in 3D brain CT radiology report generation. Nat. Commun. 2025, 16, 2258. [Google Scholar] [CrossRef]
- Gaastra, B.; Duncan, P.; Bakker, M.K.; Hostettler, I.C.; Alg, V.S.; Houlden, H.; Ruigrok, Y.M.; Galea, I.; Tapper, W.; Werring, D.; et al. Genetic variation in NFE2L2 is associated with outcome following aneurysmal subarachnoid haemorrhage. Eur. J. Neurol. 2023, 30, 116–124. [Google Scholar] [CrossRef] [PubMed]
- Taha, A.A.; Hanbury, A. Metrics for evaluating 3D medical image segmentation: Analysis, selection, and tool. BMC Med. Imaging 2015, 15, 29. [Google Scholar] [CrossRef] [PubMed]
- Mane, R.; Chouhan, T.; Guan, C. BCI for stroke rehabilitation: Motor and beyond. J. Neural Eng. 2020, 17, 041001. [Google Scholar] [CrossRef]
- Mongan, J.; Moy, L.; Kahn, C.E., Jr. Checklist for Artificial Intelligence in Medical Imaging (CLAIM): A Guide for Authors and Reviewers. Radiol. Artif. Intell. 2020, 2, e200029. [Google Scholar] [CrossRef]
- Yuste, R.; Goering, S.; Arcas, B.A.Y.; Bi, G.; Carmena, J.M.; Carter, A.; Fins, J.J.; Friesen, P.; Gallant, J.; Huggins, J.E.; et al. Four ethical priorities for neurotechnologies and AI. Nature 2017, 551, 159–163. [Google Scholar] [CrossRef]
- Svedbo Engstrom, M.; Johansson, U.B.; Leksell, J.; Linder, E.; Eeg-Olofsson, K. Implementing the Digital Diabetes Questionnaire as a Clinical Tool in Routine Diabetes Care: Focus Group Discussions with Patients and Health Care Professionals. JMIR Diabetes 2022, 7, e34561. [Google Scholar] [CrossRef]
- Rieke, N.; Hancox, J.; Li, W.; Milletari, F.; Roth, H.R.; Albarqouni, S.; Bakas, S.; Galtier, M.N.; Landman, B.A.; Maier-Hein, K.; et al. The future of digital health with federated learning. NPJ Digit. Med. 2020, 3, 119. [Google Scholar] [CrossRef] [PubMed]
- Hartman, E.; Scott, A.M.; Karlsson, C.; Mohanty, T.; Vaara, S.T.; Linder, A.; Malmstrom, L.; Malmstrom, J. Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis. Nat. Commun. 2023, 14, 5359. [Google Scholar] [CrossRef]
- Janizek, J.D.; Dincer, A.B.; Celik, S.; Chen, H.; Chen, W.; Naxerova, K.; Lee, S.I. Uncovering expression signatures of synergistic drug responses via ensembles of explainable machine-learning models. Nat. Biomed. Eng. 2023, 7, 811–829. [Google Scholar] [CrossRef]
- Adadi, A.; Berrada, M. Peeking Inside the Black-Box: A Survey on Explainable Artificial Intelligence (XAI). IEEE Access 2018, 6, 52138–52160. [Google Scholar] [CrossRef]
- Burwell, S.; Sample, M.; Racine, E. Ethical aspects of brain computer interfaces: A scoping review. BMC Med. Ethics 2017, 18, 60. [Google Scholar] [CrossRef] [PubMed]
- Perdikis, S.; Tonin, L.; Saeedi, S.; Schneider, C.; Millán, J.D.R. The Cybathlon BCI race: Successful longitudinal mutual learning with two tetraplegic users. PLoS Biol. 2018, 16, e2003787. [Google Scholar] [CrossRef]
- Reyes, M.; Meier, R.; Pereira, S.; Silva, C.A.; Dahlweid, F.M.; von Tengg-Kobligk, H.; Summers, R.M.; Wiest, R. On the Interpretability of Artificial Intelligence in Radiology: Challenges and Opportunities. Radiol. Artif. Intell. 2020, 2, e190043. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Huang, Z.; Guo, Y.; Li, Y.; Zhu, J.; Wen, J.; Gao, Y.; Liu, J. Identification and Validation of an Explainable Prediction Model of Sepsis in Patients With Intracerebral Hemorrhage: Multicenter Retrospective Study. J. Med. Internet Res. 2025, 27, e71413. [Google Scholar] [CrossRef]
- Müller-Putz, G.R.; Scherer, R.; Neuper, C.; Pfurtscheller, G. Steady-state somatosensory evoked potentials: Suitable brain signals for brain-computer interfaces? IEEE Trans. Neural Syst. Rehabil. Eng. 2006, 14, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Kelly, C.J.; Karthikesalingam, A.; Suleyman, M.; Corrado, G.; King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med. 2019, 17, 195. [Google Scholar] [CrossRef]
- Gilbert, F.; Cook, M.; O’Brien, T.; Illes, J. Embodiment and Estrangement: Results from a First-in-Human “Intelligent BCI” Trial. Sci. Eng. Ethics 2019, 25, 83–96. [Google Scholar] [CrossRef]
- Liu, X.; Cruz Rivera, S.; Moher, D.; Calvert, M.J.; Denniston, A.K.; Spirit, A.I.; Group, C.-A.W. Reporting guidelines for clinical trial reports for interventions involving artificial intelligence: The CONSORT-AI extension. Lancet Digit. Health 2020, 2, e537–e548. [Google Scholar] [CrossRef] [PubMed]
- Ben Elhadj, H.; Chaari, L.; Boudjit, S.; Kamoun, L. Node and Hub Data Gathering Architectures for Healthcare Applications based on IEEE 802.15.6 Standard. Int. J. E-Health Med. Commun. (IJEHMC) 2015, 6, 38–62. [Google Scholar] [CrossRef]
- Gao, M.; Mao, J. A Novel Active Rehabilitation Model for Stroke Patients Using Electroencephalography Signals and Deep Learning Technology. Front. Neurosci. 2021, 15, 780147. [Google Scholar] [CrossRef]
- Bobillo, S.; Nieto, J.C.; Barba, P. Use of checkpoint inhibitors in patients with lymphoid malignancies receiving allogeneic cell transplantation: A review. Bone Marrow Transplant. 2021, 56, 1784–1793. [Google Scholar] [CrossRef]
- Lewandowsky, S.; Oberauer, K. Publisher Correction: Low replicability can support robust and efficient science. Nat. Commun. 2020, 11, 4109. [Google Scholar] [CrossRef]
- Dong, Y.; Pachade, S.; Roberts, K.; Jiang, X.; Sheth, S.A.; Giancardo, L. Generalizable self-supervised learning for brain CTA in acute stroke. Comput. Biol. Med. 2025, 184, 109337. [Google Scholar] [CrossRef]
- Wang, Z.; Cao, C.; Chen, L.; Gu, B.; Liu, S.; Xu, M.; He, F.; Ming, D. Multimodal Neural Response and Effect Assessment During a BCI-Based Neurofeedback Training After Stroke. Front. Neurosci. 2022, 16, 884420. [Google Scholar] [CrossRef]
- Kim, M.; Rho, K.; Kim, Y.D.; Jung, K. Action-driven contrastive representation for reinforcement learning. PLoS ONE 2022, 17, e0265456. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Han, K. Methodologic Guide for Evaluating Clinical Performance and Effect of Artificial Intelligence Technology for Medical Diagnosis and Prediction. Radiology 2018, 286, 800–809. [Google Scholar] [CrossRef] [PubMed]
- Mrachacz-Kersting, N.; Jiang, N.; Stevenson, A.J.; Niazi, I.K.; Kostic, V.; Pavlovic, A.; Radovanovic, S.; Djuric-Jovicic, M.; Agosta, F.; Dremstrup, K.; et al. Efficient neuroplasticity induction in chronic stroke patients by an associative brain-computer interface. J. Neurophysiol. 2016, 115, 1410–1421. [Google Scholar] [CrossRef]
- Sutton, R.T.; Pincock, D.; Baumgart, D.C.; Sadowski, D.C.; Fedorak, R.N.; Kroeker, K.I. An overview of clinical decision support systems: Benefits, risks, and strategies for success. NPJ Digit. Med. 2020, 3, 17. [Google Scholar] [CrossRef]
- Topol, E.J. High-performance medicine: The convergence of human and artificial intelligence. Nat. Med. 2019, 25, 44–56. [Google Scholar] [CrossRef]
- Wolpaw, J.R.; Birbaumer, N.; McFarland, D.J.; Pfurtscheller, G.; Vaughan, T.M. Brain-computer interfaces for communication and control. Clin. Neurophysiol. 2002, 113, 767–791. [Google Scholar] [CrossRef]
- Holder, D.; Leeseberg, K.; Giles, J.A.; Lee, J.M.; Namazie, S.; Ford, A.L. Central Triage of Acute Stroke Patients Across a Distributive Stroke Network Is Safe and Reduces Transfer Denials. Stroke 2021, 52, 2671–2675. [Google Scholar] [CrossRef] [PubMed]
- He, J.; Baxter, S.L.; Xu, J.; Xu, J.; Zhou, X.; Zhang, K. The practical implementation of artificial intelligence technologies in medicine. Nat. Med. 2019, 25, 30–36. [Google Scholar] [CrossRef]
- Theruvath, J.; Menard, M.; Smith, B.A.H.; Linde, M.H.; Coles, G.L.; Dalton, G.N.; Wu, W.; Kiru, L.; Delaidelli, A.; Sotillo, E.; et al. Anti-GD2 synergizes with CD47 blockade to mediate tumor eradication. Nat. Med. 2022, 28, 333–344. [Google Scholar] [CrossRef]
- Price, W.N.; Cohen, I.G., 2nd. Privacy in the age of medical big data. Nat. Med. 2019, 25, 37–43. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Murguialday, A.; Broetz, D.; Rea, M.; Laer, L.; Yilmaz, O.; Brasil, F.L.; Liberati, G.; Curado, M.R.; Garcia-Cossio, E.; Vyziotis, A.; et al. Brain-machine interface in chronic stroke rehabilitation: A controlled study. Ann. Neurol. 2013, 74, 100–108. [Google Scholar] [CrossRef]
- Kamel, H.; Suarez, J.I.; Connolly, E.S.; Amin-Hanjani, S.; Mack, W.T.; Chou, S.H.; Busl, K.M.; Derdeyn, C.P.; Dangayach, N.S.; Elm, J.E.; et al. Addressing the Evidence Gap in Aneurysmal Subarachnoid Hemorrhage: The Need for a Pragmatic Randomized Trial Platform. Stroke 2024, 55, 2397–2400. [Google Scholar] [CrossRef] [PubMed]
- Ienca, M.; Andorno, R. Towards new human rights in the age of neuroscience and neurotechnology. Life Sci. Soc. Policy 2017, 13, 5. [Google Scholar] [CrossRef] [PubMed]
- Geng, Y.-J. Mental Stress Contributes to the Pathogenesis of Atherosclerotic Heart and Brain Diseases. Heart Mind 2023, 7, 157–163. [Google Scholar] [CrossRef]
- Obermeyer, Z.; Powers, B.; Vogeli, C.; Mullainathan, S. Dissecting racial bias in an algorithm used to manage the health of populations. Science 2019, 366, 447–453. [Google Scholar] [CrossRef]
- Váša, F.; Romero-Garcia, R.; Kitzbichler, M.G.; Seidlitz, J.; Whitaker, K.J.; Vaghi, M.M.; Kundu, P.; Patel, A.X.; Fonagy, P.; Dolan, R.J.; et al. Conservative and disruptive modes of adolescent change in human brain functional connectivity. Proc. Natl. Acad. Sci. USA 2020, 117, 3248–3253. [Google Scholar] [CrossRef]
- Pfohl, S.R.; Foryciarz, A.; Shah, N.H. An empirical characterization of fair machine learning for clinical risk prediction. J. Biomed. Inform. 2021, 113, 103621. [Google Scholar] [CrossRef]
- Sheller, M.J.; Edwards, B.; Reina, G.A.; Martin, J.; Pati, S.; Kotrotsou, A.; Milchenko, M.; Xu, W.; Marcus, D.; Colen, R.R.; et al. Federated learning in medicine: Facilitating multi-institutional collaborations without sharing patient data. Sci. Rep. 2020, 10, 12598. [Google Scholar] [CrossRef]
- Yang, Q.; Liu, Y.; Chen, T.; Tong, Y. Federated Machine Learning. ACM Trans. Intell. Syst. Technol. 2019, 10, 1–19. [Google Scholar] [CrossRef]
- Moon, I.; Lee, J.; Lee, S.-A.; Jeong, D.; Jeon, J.; Jang, Y.; Jeong, S.; Kim, J.; Choi, H.-M.; Hwang, I.-C.; et al. Artificial Intelligence-Enhanced Analysis of Echocardiography-Based Radiomic Features for Myocardial Hypertrophy Detection and Etiology Differentiation. Circ. Cardiovasc. Imaging 2025, 18, e017436. [Google Scholar] [CrossRef] [PubMed]
- Kuo, W.; Häne, C.; Mukherjee, P.; Malik, J.; Yuh, E.L. Expert-level detection of acute intracranial hemorrhage on head computed tomography using deep learning. Proc. Natl. Acad. Sci. USA 2019, 116, 22737–22745. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Xu, H.; Chen, Q.; Zhang, T.; Sheng, W.; Huang, Q.; Song, J.; Huang, D.; Lan, L.; Li, Y.; et al. Prediction of hematoma expansion in spontaneous intracerebral hemorrhage using support vector machine. EBioMedicine 2019, 43, 454–459. [Google Scholar] [CrossRef]
- Soun, J.E.; Chow, D.S.; Nagamine, M.; Takhtawala, R.S.; Filippi, C.G.; Yu, W.; Chang, P.D. Artificial Intelligence and Acute Stroke Imaging. Am. J. Neuroradiol. 2021, 42, 2–11. [Google Scholar] [CrossRef]
- Rudin, C. Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead. Nat. Mach. Intell. 2019, 1, 206–215. [Google Scholar] [CrossRef]
- Bica, I.; Alaa, A.M.; Lambert, C.; van der Schaar, M. From Real-World Patient Data to Individualized Treatment Effects Using Machine Learning: Current and Future Methods to Address Underlying Challenges. Clin. Pharmacol. Ther. 2020, 109, 87–100. [Google Scholar] [CrossRef]
- Corral-Acero, J.; Margara, F.; Marciniak, M.; Rodero, C.; Loncaric, F.; Feng, Y.; Gilbert, A.; Fernandes, J.F.; Bukhari, H.A.; Wajdan, A.; et al. The ‘Digital Twin’ to enable the vision of precision cardiology. Eur. Heart J. 2020, 41, 4556–4564. [Google Scholar] [CrossRef]
- Nagendran, M.; Chen, Y.; Lovejoy, C.A.; Gordon, A.C.; Komorowski, M.; Harvey, H.; Topol, E.J.; Ioannidis, J.P.A.; Collins, G.S.; Maruthappu, M. Artificial intelligence versus clinicians: Systematic review of design, reporting standards, and claims of deep learning studies. BMJ 2020, 368, m689. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.