Abstract
Latent growth modeling (LGM) is widely used to examine group differences in developmental trajectories in educational research. With multiple outcomes of interest, a multiple-domain latent growth model (MDLGM) can be applied to account for associations among growth trajectories from the outcomes. While the MDLGM is conceived as a more powerful multivariate analysis technique when compared with univariate LGM, the examination of its methodological performance is very limited. Henceforth, this study compared the statistical power of the MDLGM and separate univariate LGMs via a simulation study with a two-group, two-domain design. Results indicated that the MDLGM and the set of LGMs showed largely comparable power, with a small advantage for the MDLGM under conditions of weak inter-domain correlation and small group differences and showed reduced power as missingness increased. These findings provide educational researchers with a guideline for model selection in longitudinal studies with multiple outcomes.
MSC:
62D10; 62H12; 62H25
1. Introduction
In longitudinal data analysis, missing data (e.g., attrition) are commonly encountered in educational and psychological settings, and many studies have discussed the treatment and mechanisms of missing data in longitudinal research [1,2]. In longitudinal research, the latent growth model (LGM) [3] within the structural equation modeling (SEM) framework is widely used and can be extended to incorporate multiple growth trajectories simultaneously. The resulting model is referred to as a multiple-domain latent growth model (MDLGM) or a cross-domain analysis of change model [4]. The MDLGM allows researchers to examine individual differences in developmental change both within individual domains and across multiple domains over time. By jointly modeling correlated growth processes, the MDLGM provides a flexible analytic approach for addressing diverse research questions in correlational longitudinal and developmental research.
Although the MDLGM has been applied in educational and psychological research [5,6], the literature indicates that methodological knowledge regarding its performance in terms of statistical power remains limited. Some studies have examined the statistical power of the MDLGM [7,8] and investigated the use of MDLGM for mediation and causal inference in longitudinal analyses [9,10]. Also, recent simulation studies extended MDLGMs to accommodate non-normal outcome distributions, such as count or ordinal data, via the Bayesian estimation [11]. However, empirical evaluations of their comparative statistical performance under controlled simulation conditions remain limited. The present study provides a structured assessment of the relative statistical power and Type I error rates of these existing models across varying design parameters, including missingness in outcome, as MDLGM’s statistical power for examining group differences in several growth trajectories in the presence of missing data has yet to be investigated. Further, it is well established that conducting multivariate analyses, rather than multiple univariate analyses, can reduce inflation of Type I error rates and improve statistical power when examining multiple correlated outcome variables simultaneously [12,13]. Accordingly, the MDLGM may yield greater power than separate univariate LGMs because it incorporates additional information through correlations among latent growth factors across domains. Therefore, the purpose of this study was to compare the statistical power of the MDLGM with that of a set of univariate LGMs for detecting group differences in longitudinal data with missingness.
1.1. Univariate Latent Growth Modeling
The LGM is a restricted confirmatory factor analysis (CFA) [14] model with a mean structure. The LGM detects both intra-individual differences and inter-individual differences across time using the intercept and slope factors (latent variables). In this framework, individual change over time is assumed to follow a constant rate of increase or decrease, represented by a straight line across measurement occasions. As seen in Figure 1, for instance, the two latent variables (i.e., the intercept and slope factors) are indicated by three repeated outcome measures, for the ith individual at time point t. The intercept factor describes the level in the outcome measures at a reference time point (e.g., the first measurement occasion), whereas the slope factor indicates the rate of change in the outcome measure over time [15,16]. The linear LGM in Figure 1 can be expressed as follows:
where is a 3 × 1 vector of the three observed outcome measures for the ith individual, is a 3 × 1 vector of the intercepts of the three observed measures, is a 3 × 2 matrix of factor loadings for the intercept and slope factors, is a 2 × 1 vector of the intercept and slope factor scores for the ith individual, and a 3 × 1 vector that refers to the residuals in the observed outcome measures that are not explained by the latent growth model for the ith individual that assumed to follow a multivariate normal distribution with a mean vector of zeros and a variance/covariance matrix of . The intercepts of the observed measures () are typically constrained to zero in the LGM specification due to a model identification issue. Specifically, the components of are fixed to known constants in which the first column presents ones, and the second column captures ( = t − 1) at each time point, t.
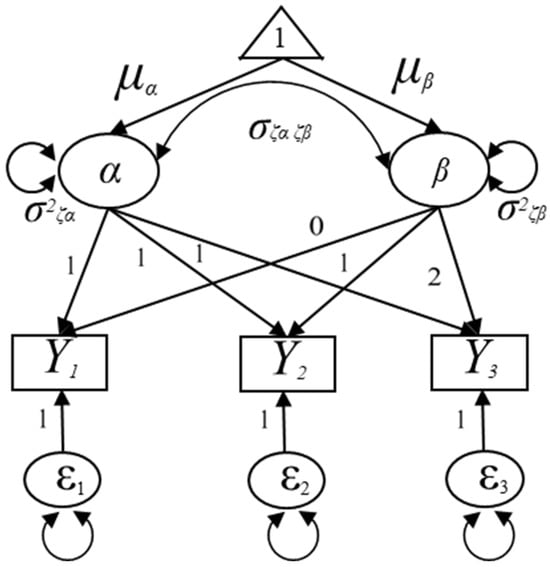
Figure 1.
Specification of the unconditional univariate latent growth model.
Further, the intercept and slope factor vector () for the ith individual can be decomposed into fixed and random effects as follows:
where is a 2 × 1 vector of means of the intercept and slope factors, and is a 2 × 1 vector of the intercept and slope factors’ disturbances for the ith individual that follows a multivariate normal distribution with a mean of zeros and variance/covariance matrix of Ψ.
1.2. Multiple-Domain Latent Growth Model
Numerous studies have investigated individual differences in developmental trajectories across multiple outcomes or domains [6,17,18]. The latent growth model (LGM) described in Equations (1) and (2) can be readily extended to assess associations among growth factors across domains, thereby allowing the estimation of intercorrelations among latent growth parameters across multiple-domains [4,19]. The resulting model is referred to as a multiple-domain latent growth model (MDLGM) [8,20,21]. This model has also been described as an associative latent growth model [19], a parallel process model [22,23,24], a cross-domain latent growth model [4], and a bivariate latent growth model [15]. For the purposes of this study, the model will henceforth be referred to as the multiple-domain latent growth model (MDLGM).
The MDLGM allows researchers to simultaneously examine the relationships among multiple growth trajectories. For example, researchers may seek to assess how individuals’ academic achievement evolves concurrently across domains such as reading and mathematics and to determine whether growth processes in these domains are interrelated [4]. In contrast, analyzing multiple outcomes using a series of separate univariate LGMs does not capture interrelations across domains regarding growth factors (e.g., intercept and slope factors), which can be directly estimated using the MDLGM.
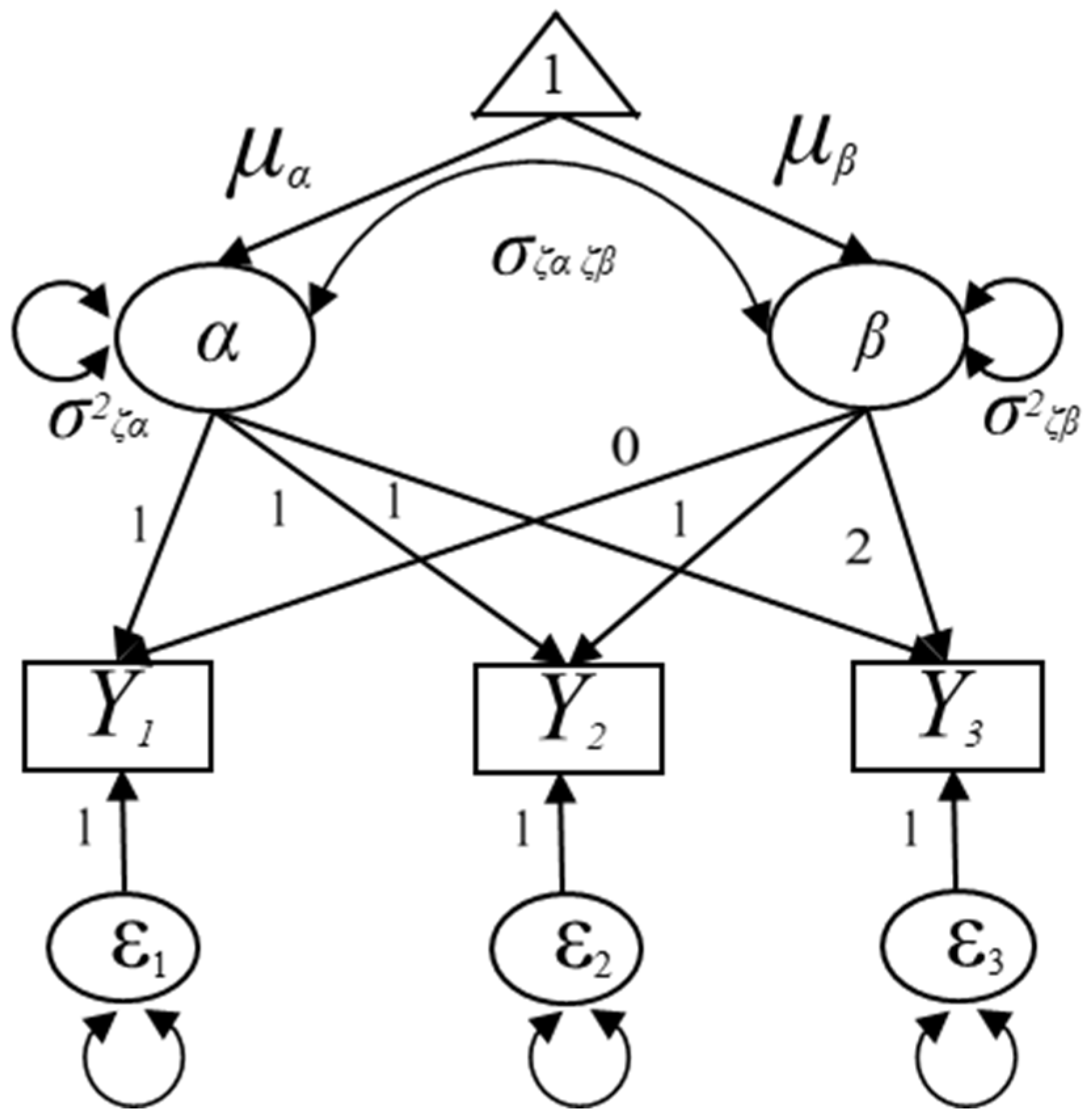
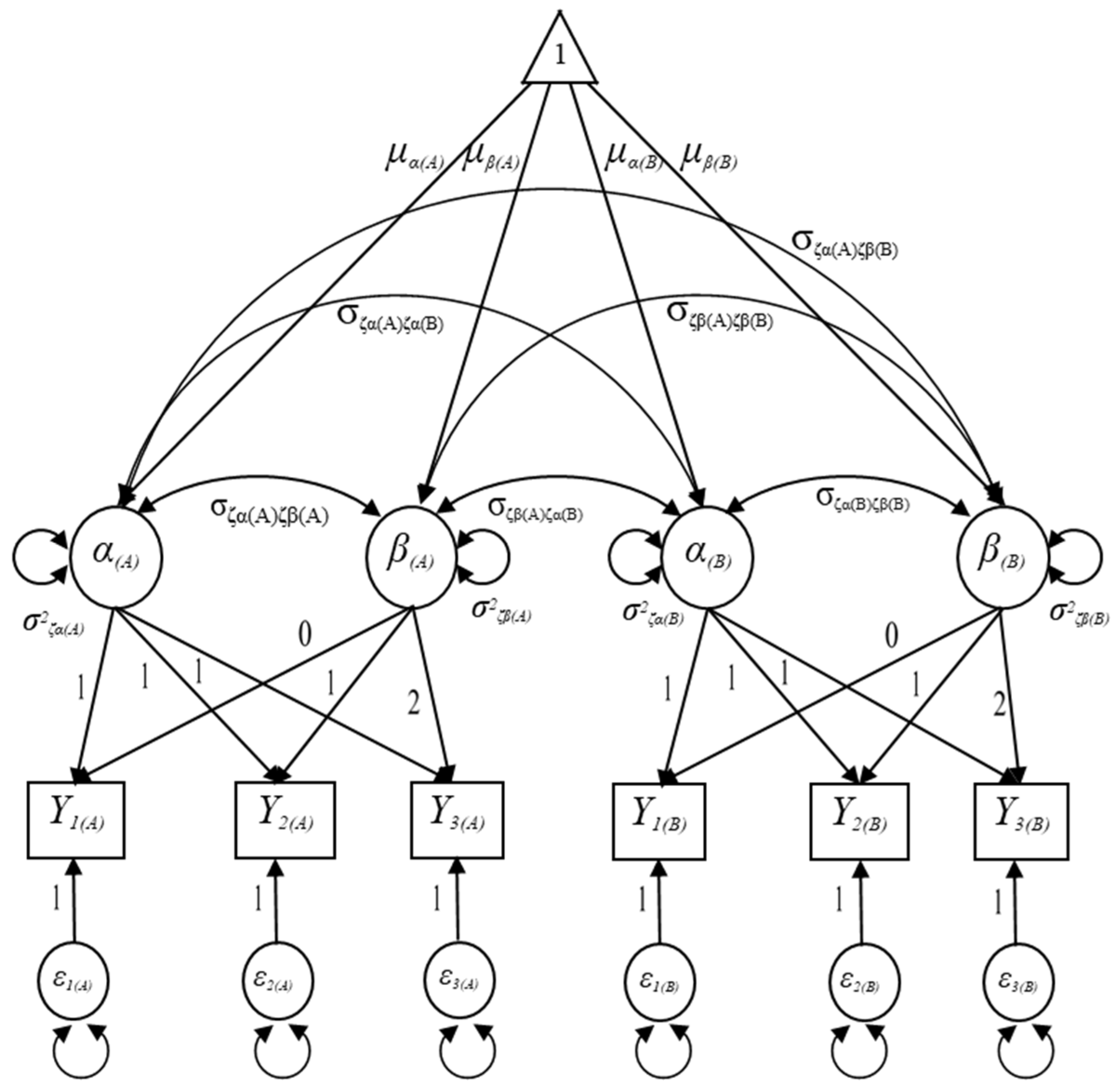
Let denote the observed outcome measure in domain A for the ith individual at time point t and represent the observed measure in domain B for the ith individual at time point t as seen in Figure 2. Given that the functional forms of the growth trajectories in both domain A and B are linear in Figure 2, the MDLGM can be expressed as follows:
where is a 6 × 1 vector of the observed outcome measures for the ith individual in the jth domain, is a 6 × 1 vector of the intercepts of outcome measures for the ith individual in the jth domain that are constrained to zeros, is a 6 × 4 matrix of the growth factors’ factor loading matrix loadings load on observed outcome measures, is a 4 × 1 vector of the growth factors, and is a 6 × 1 vector of the residuals in the observed outcome measures that are not explained by the latent growth model that assumed to follow the multivariate normal distribution with a mean vector of zeros and a variance/covariance matrix of .
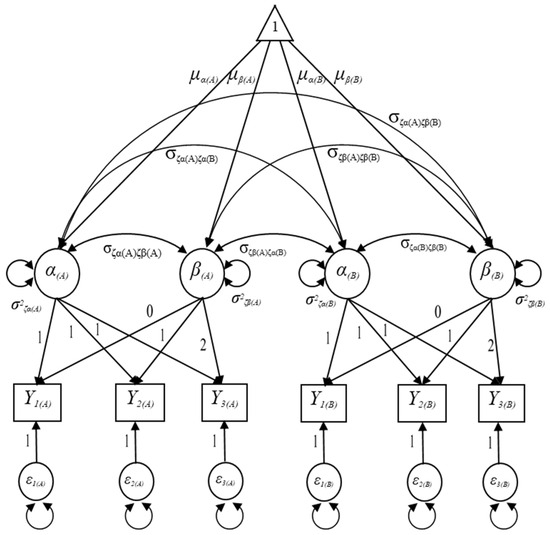
Figure 2.
Specification of the unconditional multiple-domain latent growth model.
Again, the intercept and slope factor vector () in Equation (3) for the ith individual can be decomposed into fixed and random effects as follows:
where is a 4 × 1 vector of means of the intercept and slope factors in domain A and domain B, and is a 2 × 1 vector of the intercept and slope factors’ disturbances for the ith individual in domain A and domain B that follows a multivariate normal distribution with a mean of zeros and variance/covariance matrix of Ψ.
In addition, the association between the two outcome measures is represented by the covariance structure among the latent intercept and slope factors [15]. The covariance structure of the latent growth factors is captured by the elements of the matrix, which is the covariance structure of latent residuals () in Equation (4) as follows:
The matrix includes the domain-specific variances and the covariances between domains. The diagonal entries correspond to the variances of the latent growth factors within each domain, while the off-diagonal entries represent the covariances among growth factors both within and across domains.
It is important to first evaluate the growth pattern within each domain to determine the appropriate functional form for the domain-specific univariate latent growth model [4,25]. For example, this preliminary assessment may involve visual inspection of observed scores across measurement occasions. Accordingly, separate univariate LGMs should be fitted for each domain prior to estimating the multiple-domain latent growth model (MDLGM) [4,26].
1.3. Conditional MDLGM
In the presence of heterogeneity in the intercept and slope factors, it is possible to examine whether variability in growth processes across domains is explained by time-invariant predictors such as group membership (e.g., gender) [4]. Inclusion of a binary time-invariant predictor enables the estimation of group-based differences in latent growth parameters within the MDLGM framework through dummy-coded variable specification.
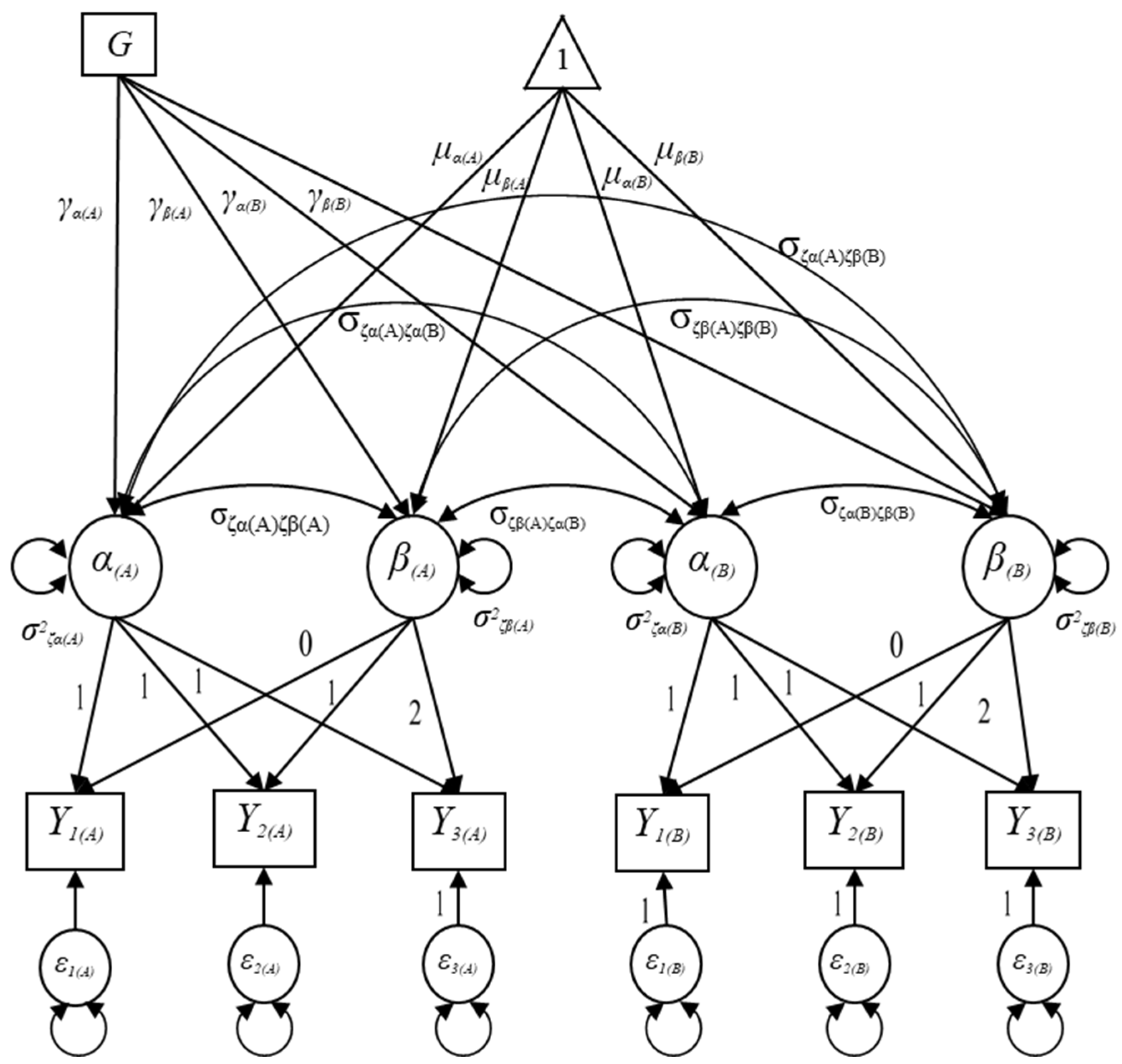
Let denote a binary predictor indicating a group membership for the ith individual (as shown in Figure 3) and let , , , and represent the path coefficients regressed on the intercept and slope factors in domains A and B; that is, the path coefficients represent the group differences in initial status and growth rate in the outcome measure in domains A and B. It is known that the inclusion of a time-invariant predictor affects the structural component of the MDLGM so that the growth factors () for the ith individual in the conditional MDLGM can be expressed as follows:
where is a 4 × 1 vector for the ith individual containing the intercept and slope factors for domains A and B; is a 4 × 1 vector of the means of the intercept and slope factors in domains A and B; is a 4 × 1 vector including the regression coefficients’ weights that quantifies linear associations between the predictor and the growth factors in domains A and B; and is a 4 × 1 vector representing the latent disturbances for the ith individual. The latent disturbances in the conditional MDLGM can be interpreted as the variability that remains in the growth factors (i.e., intercept and slope factors) after taking into account the impact of the predictor [27].
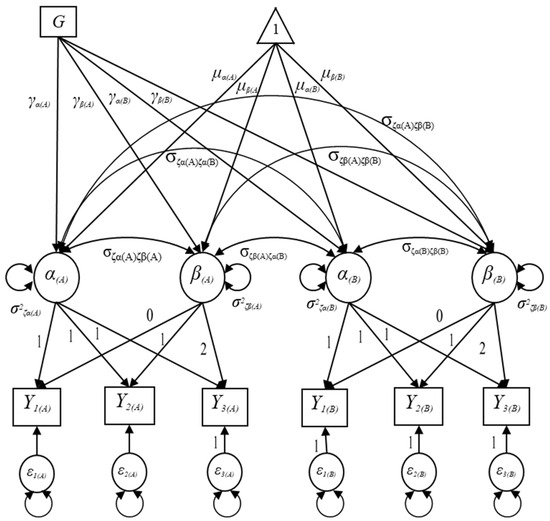
Figure 3.
Specification of the conditional multiple-domain latent growth model with a binary grouping variable.
1.4. Missing Data Mechanism and Maximum Likelihood Method
Missing data are commonly encountered in educational and psychological research, particularly in longitudinal studies [2]. Rubin [28] defined three missing data mechanisms, including missing completely at random (MCAR), missing at random (MAR), and missing not at random (MNAR). Longitudinal missing data are generally considered to be either MAR or MNAR [29]. Although various methods are available for handling parameter estimation with missing data, maximum likelihood (ML), also known as full information maximum likelihood (FIML), is a model-based estimation method that is widely used when data are MAR [2]. Under ML estimation, observations with a larger number of observed data points are weighted more heavily than those with fewer observed data points when estimating latent growth models [30]. In other words, ML estimation is advantageous because it makes inferences based on the likelihood function derived from the observed data and provides model fit indices [1].
An alternative approach for handling missing data under the MAR assumption is a multiple imputation method. In this approach, missing data points are imputed multiple times (e.g., typically five to ten imputations) using information from the observed data. The latent growth model is then fitted separately to each imputed dataset, and the results are subsequently pooled across analyses [30].
Enders [1] noted that the assumptions required for models designed to handle MNAR data (e.g., selection models and pattern-mixture models) are difficult to satisfy in practice. Furthermore, because the present study represents an initial examination of the performance of the MDLGM with incomplete data, the missing data mechanism was assumed to be MAR.
1.5. Purpose of the Present Study
Despite the substantial use of LGMs, relatively few studies have examined their statistical power [7,31,32,33,34,35,36]. Fan [32] investigated the statistical power of linear LGMs for detecting group differences in latent growth parameters by regressing intercept and slope factors on a dummy-coded predictor while varying study conditions such as group difference effect sizes and sample size. The results indicated that Type I error rates remained acceptable (i.e., approximately 0.05) across a wide range of sample sizes when no group differences in intercept and/or slope factors were present. Moreover, statistical power to detect group differences in intercept and slope factors was acceptable (e.g., 0.80) when group differences were large (d = 0.80), even with relatively small sample sizes (e.g., N = 100). In contrast, when group differences were small (d = 0.20), much larger sample sizes (e.g., N ≥ 800) were required to achieve sufficient statistical power (i.e., 0.80 or higher).
Prior methodological work has argued that multivariate latent growth modeling may yield benefits compared to estimating separate univariate growth models [21]. In a related study, Lee and Whittaker [8] found that the use of a multiple-domain latent growth model (MDLGM) to detect group differences in slope factors generally yielded power comparable to that obtained using two separate univariate LGMs. However, they also reported that under certain conditions, when relatively strong inter-domain correlations were combined with relatively small group differences in both domains, the MDLGM provided a slight power advantage over the set of univariate LGMs (e.g., a group difference of 0.0 in Domain A and 0.2 in Domain B with an inter-domain correlation of 0.50). This advantage was attributed to the additional variance components captured by random effects in the conditional MDLGM, which contributed to smaller standard errors for slope coefficients regressed on the exogenous grouping variable [8]. These findings suggest that the MDLGM may be more advantageous than a set of univariate LGMs for detecting group differences, particularly when certain degrees of missingness are present in observed outcome measures.
To our knowledge, prior research has not directly investigated the statistical power of the MDLGM for identifying group differences under conditions of incomplete data. The present study, therefore, aimed to evaluate the power of the MDLGM to identify differences in slope factors and to compare its performance with that of a set of univariate LGMs in the presence of missing data. A Monte Carlo simulation design was implemented to determine whether the MDLGM provides greater statistical power than separate univariate LGMs when testing for group differences. Five design factors were manipulated: (a) sample size, (b) group difference effect sizes, (c) the magnitude of intercorrelations among latent growth factors across domains, (d) the degree of missingness in the observed data, and (e) the number of measurement occasions, resulting in a total of 576 simulation conditions.
2. Materials and Methods
2.1. Simulation Study Conditions
In this simulation study, there was a total of 576 simulation conditions, including a combination of three sample sizes (i.e., N = 200, 400, and 800), two number of measurement occasions (i.e., three/four measurement occasions), four intercorrelations among latent factors between two domains (i.e., ρ = 0.0, 0.1, 0.3, and 0.5), four group difference effect sizes (i.e., d = 0.0, d = 0.2, d = 0.5, d = 0.8), and three different degrees of missingness (i.e., no missingness, moderate missingness, and large missingness).
2.1.1. Sample Size
It is known that a minimum sample size for employing SEM is N = 200 [37]. Also, some researchers have found that the most commonly seen sample sizes in applied studies in SEM were 200, 500, 1000, and 2000 [38]. Thus, three different sample sizes were used in this simulation study (N = 200, 400, and 800) to represent small, moderate, and large sample sizes, respectively.
2.1.2. Group Difference Effect Size
It is well established that the statistical power of a model to detect group differences depends on the magnitude of the group difference effect size [39]. The group difference effect size differed by the Cohen’s effect size measure criteria [40]. There were four group difference effect sizes, indicating null effect (d = 0.0), small group difference (d = 0.2), moderate group difference (d = 0.5), and large group difference (d = 0.8).
2.1.3. Intercorrelation Between Domains
Frane [41] demonstrated that the influence of effect sizes on the power to detect group differences is contingent upon the degree of correlation among response variables. In line with this finding, the intercorrelations among latent growth factors were varied to be zero (0.0), small (0.1), moderate (0.3), and large (0.5) correlations [40].
2.1.4. The Degree of Missingness
Assuming the degree of missingness in the observed measures increased across time, the first measurement occasion data were missing completely at random (MCAR), and the observed outcome measures at the rest of the measurement occasions were missing at random (MAR) [42]. Accordingly, data were generated with no missing (0%), moderate missingness (30% missing data at the last time point), and large missingness (50% missing data at the last time point).
2.1.5. The Number of Measurement Occasions
It is known that a minimum of three repeated measurement occasions is required to estimate a latent growth model (LGM). Additionally, Muthén and Curran [31] found that there were substantial differences in statistical power for detecting group differences when comparing models with three versus four measurement occasions. Accordingly, we consider the scenarios under three- and four-wave designs.
2.2. Data Generation
Data were generated using the conditional MDLGM with a binary predictor (dummy coded) representing group membership (Figure 3) via the Monte Carlo feature in Mplus (version 7.4) [43]. Data were generated to follow a multivariate normal distribution. Additionally, the group assignment in the present study was assumed to be random and balanced so that the variance of the binary grouping variable (G) was set to 0.25. Based on the previous literature, the intercept and slope factors’ mean values were set to zero and 0.1, respectively [22,44], and the two growth factors’ variance values were set to 1, representing a standardized group difference effect size (i.e., Cohen’s d). By doing so, the regression coefficients, , , , and , can be interpreted as Cohen’s d effect size. Further, data were generated via the effect size, indicating group difference on the two domains’ intercept factors were fixed to zero. This represents that the individuals’ initial statuses on the two domains are the same across individuals. Once the data were generated, the generated parameter estimates were checked by descriptive statistics (e.g., mean, median, standard deviation) to ensure the data generation process.
Specifically, the correlations between the intercept and slope factors within each domain were fixed at 0.20, and the correlations among growth factors across domains were also set to 0.20, consistent with values commonly reported in educational research [45]. The residual variances of the outcome variables () in each domain were all set to 0.50, and these residual variances are not correlated with each other across measurement occasions and with the growth factors [42].
2.3. Model Estimation
A total of 1000 randomly generated datasets were analyzed using Mplus 7.4 [43] with the maximum likelihood (ML) estimation method. For each of the 576 simulation conditions, the data were fitted to four models: the full MDLGM, restricted MDLGM, a set of full univariate LGMs, and a set of restricted univariate LGMs. With the full MDLGM, the direct effects of the grouping variable on the two domains’ slope factors were estimated simultaneously. With the set of full univariate LGMs, the direct effect of the grouping variable on each of the two domains’ slope factors was estimated separately. Subsequently, a restricted MDLGM was estimated in which the regression coefficients from the grouping variable to the slope factors in both domains [ and ] were constrained to zero. Similarly, restricted univariate LGMs were estimated for each domain by fixing the corresponding regression coefficient to zero. Throughout this study, the MDLGMs/LGMs in which the coefficients of and were freely estimated were referred to as full models, whereas the MDLGMs/LGMs with constraints of and equal to zero were referred to as restricted models.
2.4. Data Analysis
Upon completion of data generation, the 1000 generated datasets were fitted to four models (i.e., the full MDLGM/set of LGMs and the restricted MDLGM/set of LGMs). Thereafter, null hypotheses were examined by both MDLGMs and the set of LGMs using R 3.4.0 (R Core Team, 2017) [46].
The overall multivariate null hypothesis tested by the MDLGMs is:
where and represent the regression coefficients of the domain A and domain B’s slope factors on the grouping variable, respectively. The overall multivariate null hypothesis was utilized as a likelihood ratio test with 2 degrees of freedom (df) difference. Specifically, the likelihood ratio test was conducted by comparing the model test statistic associated with the full MDLGM to that of the restricted MDLGM. The likelihood ratio test’s null hypothesis was rejected with an occasion of statistically significant Δ between the full model and the restricted model at an α-level of 0.05.
Likewise, the overall multivariate null hypothesis tested by a set of LGMs is as follows:
The overall multivariate test for the set of univariate LGMs also utilized a set of likelihood ratio tests. The likelihood ratio tests were conducted by comparing the test statistics associated with the full LGM to those of the restricted LGM. The null hypothesis of the likelihood ratio tests was rejected when at least one of the two Δ tests between the full LGM and the restricted LGM was statistically significant at a Bonferroni-adjusted α-level of 0.025 (0.05/2 = 0.025).
Empirical statistical power was evaluated under conditions in which the regression effect of the grouping variable on the slope factor was nonzero. Power was defined as the proportion of the 1000 replications in which the overall multivariate null hypothesis for the MDLGM or the set of LGMs was rejected when a true group difference in the slope parameter was present. Consistent with Cohen’s recommendation [40], power values of 0.80 or greater were regarded as adequate. In conditions where the regression effect from the grouping variable to the slope factor was fixed at zero, empirical Type I error rates were computed as the proportion of replications in which the overall multivariate null hypothesis was incorrectly rejected. Type I error rates were evaluated by Bradley’s cutoff criterion [47] of robustness (α ± α/2). Accordingly, with a nominal significance level of α = 0.05, acceptable Type I error rates ranged from 0.025 to 0.075.
3. Results
A total of 1000 successful replications were obtained in each of the simulation conditions. When non-convergent cases occurred, those replications were automatically rerun and replaced with converged solutions to ensure a total of 1000 valid replications. Table 1 represents the Type I error rates for both the MDLGM and the set of LGMs with three repeated measures as a function of group difference effect size, the degree of missingness in the outcome measures, and intercorrelation between domains.

Table 1.
Type 1 error rates for the overall multivariate test across three repeated measures.
The empirical Type I error rates for the MDLGM and the set of LGMs ranged between 0.034 and 0.064 and between 0.033 and 0.062, respectively, indicating that the two approaches showed acceptable Type I error rates based on Bradley’s criteria [47]. Similarly, the empirical Type I error rates for the two approaches with four measurement occasions ranged from 0.043 to 0.063 and from 0.039 to 0.066, respectively, as shown in Table 2.

Table 2.
Type 1 error rates for the overall multivariate test across four repeated measures.
As seen in Table 3, the MDLGM and the set of LGMs demonstrated comparable power rates across many of the simulation conditions. However, a relatively small power advantage of the MDLGM emerged when the intercorrelation between domains was relatively low (i.e., ρ = 0.0, 0.1) and relatively low group difference effect sizes (d = 0.2). When there was a relatively large group difference effect size (d = 0.5) with a relatively high degree of intercorrelation between domains (ρ = 0.3, 0.5), the power rates from the set of LGMs tend to be similar to the power rates of the MDLGM. In the present study, power rates were considered “similar” when the absolute difference between the two models did not exceed 0.01. As expected, both modeling approaches yielded higher power under complete data conditions compared to moderate and high missingness conditions.

Table 3.
Power rates of MDLGM for the overall multivariate test across three repeated measures.
Across all conditions, empirical power increased as a function of both sample size and group difference effect size. When the degree of group difference effect size becomes 0.5 or greater, the power rates of both MDLGM and the set of LGMs reach 0.8, indicating a sufficient power rate. In contrast, when effect sizes were small (d = 0.2), adequate power was attained only under the largest sample size condition (N = 800) for both modeling approaches. Specifically, sample size had a substantial influence on power rates for both modeling approaches. Empirical power rates increased monotonically with sample size, reflecting reductions in standard errors, resulting in enhancing the stability in parameter estimates.
In Table 4, the empirical power rate patterns for both modeling approaches under four repeated measures were consistent with those obtained under three measurement occasions.

Table 4.
Power rates of MDLGM for the overall multivariate test across four repeated measures.
Overall, both the MDLGM and the set of LGMs demonstrated slightly higher power when four repeated measures were included compared to three, as expected. Again, the power rates of the two modeling approaches were comparable. However, the MDLGM showed a power advantage when inter-domain correlations were low (ρ = 0.0, 0.1) and group difference effect sizes were small (d = 0.2). In contrast, when effect sizes were moderate to large (d = 0.5) and inter-domain correlations were higher (ρ = 0.3, 0.5), the power of the set of LGMs was indistinguishable from that of the MDLGM. Furthermore, as expected, both modeling approaches achieved higher power under complete data conditions relative to situations involving a moderate-to-high degree of missingness.
4. Discussion
The present study examined the relative performance of the MDLGM and a set of univariate LGMs in terms of their statistical power to detect group differences in growth rates under incomplete data conditions. The results show that both MDLGM and the set of LGMs provided comparable power rates across most simulation conditions. However, notably, in scenarios where there is relatively low intercorrelation between domains (ρ = 0.0, 0.1) paired with relatively low group difference effect sizes (d = 0.2), the empirical power rates of the MDLGM outperformed those of the set of LGMs. This advantage arises because inter-domain correlations influence the standard errors of the slope coefficients regressed on the exogenous grouping variable. Specifically, the MDLGM utilizes the joint distribution of multiple domains to reduce the standard errors of slope coefficients under weak cross-domain dependence. This finding indicates that joint modeling improves estimation efficiency, even with a relatively small sample size, a benefit particularly salient when associations between constructs are relatively weak between domains [48]. In addition, empirical power for both the MDLGM and the set of LGMs under high and moderate missingness conditions was lower than that observed under non-missingness conditions. Nonetheless, power estimates under missing data conditions were generally comparable to those obtained with complete data for both modeling approaches. This finding is attributable to the use of full information maximum likelihood (FIML) estimation, which appropriately handles missing data assumed to satisfy missing completely at random (MCAR) or missing at random (MAR) conditions [1]. Furthermore, the empirical Type I error rates of the MDLGM and the set of LGMs (with Bonferroni correction) were comparable across all simulation conditions.
Consistent with previous research, sample size and the number of measurement occasions were positively associated with empirical power for both the MDLGM and the set of LGMs [32,33]. Specifically, adequate statistical power (i.e., 0.80) was achieved when group difference effect sizes in both domains were 0.50 or larger across all conditions for both modeling approaches. In contrast, when group difference effect sizes were small (d = 0.20), sample size had a substantially greater influence on power. For example, acceptable power levels were attained only with the largest sample size (N = 800) when group differences in both domains were small.
These results suggest that researchers may reasonably expect to detect statistically significant group differences with moderate sample sizes (e.g., N = 200) when effect sizes are moderate (e.g., d = 0.50), regardless of whether the MDLGM or a set of univariate LGMs is used. However, recent methodological advancements emphasize that the choice of modeling approach should also consider the complexity of the measurement model and the potential for time-varying covariates, which can further shift the power dynamics in longitudinal designs [49].
This study addresses a gap in the methodological literature concerning the modeling of multiple latent growth trajectories. While the current study has addressed the methodological gap related to comparing two different growth modeling approaches, there are still a few limitations. The limitations include that this study’s assumption associated with LGM includes uncorrelated measurement errors across time points with homoscedastic error variance, and the functional form of the growth trajectory is linear for both domains. These assumptions may not fully reflect the complexity of longitudinal data observed in practice. Accordingly, future research should extend this work by examining nonlinear growth forms and more flexible error structures to evaluate the robustness and statistical power of growth models under more realistic conditions. Also, future research may extend this work to include real-data analysis using a real-world longitudinal dataset to further the practical implications of the results from the simulation study.
The findings from this study may be useful for researchers who plan to collect data to detect group differences using the MDLGM. Although comparable statistical power was observed for the MDLGM and the set of univariate LGMs under certain conditions, it should be noted that the MDLGM explicitly models cross-domain covariance through joint estimation of correlated growth processes [11]. In contrast, the set of LGMs estimates each domain separately without incorporating inter-domain associations. While statistical power was comparable under certain conditions, the MDLGM offers a coherent multivariate framework that explicitly accounts for cross-domain associations and preserves the rigor of inference when growth processes are correlated. We encourage researchers to aim for a minimum of 800 participants if small-group effect sizes are anticipated. Moreover, researchers should prioritize the interpretation of practical significance; even when a statistically significant group difference is identified through a large sample size, the magnitude of the effect must be evaluated within its specific educational or psychological context.
Funding
The work was supported by a 2-Year Research Grant of Pusan National University.
Data Availability Statement
The data supporting the findings of this study are available from the corresponding author upon request.
Conflicts of Interest
The author declares no conflicts of interest.
References
- Enders, C.K. Applied Missing Data Analysis; Guilford Press: New York, NY, USA, 2010. [Google Scholar]
- Allison, P.D. Missing Data Techniques for Structural Equation Modeling. J. Abnorm. Psychol. 2003, 112, 545–557. [Google Scholar] [CrossRef] [PubMed]
- Meredith, W.; Tisak, J. Latent Curve Analysis. Psychometrika 1990, 55, 107–122. [Google Scholar] [CrossRef]
- Willett, J.B.; Sayer, A.G. Cross-Domain Analyses of Change over Time: Combining Growth Modeling and Covariance Structure Analysis. In Advanced Structural Equation Modeling: Issues and Techniques; Marcoulides, G.A., Schumacker, R.E., Eds.; Psychology Press: Hove, UK, 1996; pp. 125–157. [Google Scholar]
- Byrne, B.M.; Lam, W.W.T.; Fielding, R. Measuring patterns of change in personality assessments: An annotated application of latent growth curve modeling. J. Personal. Assess. 2008, 90, 536–546. [Google Scholar] [CrossRef]
- Keiley, M.K.; Bates, J.E.; Dodge, K.A.; Pettit, G.S. A Cross-Domain Growth Analysis: Externalizing and Internalizing Behaviors During 8 Years of Childhood. J. Abnorm. Child Psychol. 2000, 28, 161–179. [Google Scholar] [CrossRef]
- Hertzog, C.; Lindenberger, U.; Ghisletta, P.; Von Oertzen, T. On the Power of Multivariate Latent Growth Curve Models to Detect Correlated Change. Psychol. Methods 2006, 11, 244–252. [Google Scholar] [CrossRef] [PubMed]
- Lee, K.; Whittaker, T.A. Statistical Power of the Multiple Domain Latent Growth Model for Detecting Group Differences. Struct. Equ. Modeling 2018, 25, 700–714. [Google Scholar] [CrossRef]
- Liu, X.; Wang, L. Causal Mediation Analysis with the Parallel Process Latent Growth Curve Mediation Model. Struct. Equ. Modeling 2024, 31, 983–1004. [Google Scholar] [CrossRef]
- Liu, X.; Zhang, Z.; Valentino, K.; Wang, L. The Impact of Omitting Confounders in Parallel Process Latent Growth Curve Mediation Models: Three Sensitivity Analysis Approaches. Struct. Equ. Modeling 2024, 31, 132–150. [Google Scholar] [CrossRef]
- Bendler, J.; Reinecke, J. A tutorial on bayesian multiple-group comparisons of latent growth curve models with count distributed variables. Behav. Res. Methods 2025, 57, 139. [Google Scholar] [CrossRef] [PubMed]
- Stevens, J.P. Applied Multivariate Statistics for the Social Sciences, 5th ed.; Routledge: New York, NY, USA, 2012. [Google Scholar]
- Snijders, T.A.B.; Bosker, R. Multilevel Analysis: An Introduction to Basic and Advanced Multilevel Modeling, 2nd ed.; SAGE Publications Ltd.: London, UK, 2012. [Google Scholar]
- Jöreskog, K.G. A General Approach to Confirmatory Maximum Likelihood Factor Analysis. Psychometrika 1969, 34, 183–202. [Google Scholar] [CrossRef]
- Bollen, K.A.; Curran, P.J. Latent Growth Curve Models: A Structural Equation Perspective; John Wiley & Sons: Hoboken, NJ, USA, 2006; Volume 467. [Google Scholar] [CrossRef]
- Preacher, K.J.; Wichman, A.L.; MacCallum, R.C.; Briggs, N.E. Latent Growth Curve Modeling; SAGE Publications: Thousand Oaks, CA, USA, 2008. [Google Scholar]
- Sayer, A.G.; Willett, J.B. A Cross-Domain Model for Growth in Adolescent Alcohol Expectancies. Multivariate Behav. Res. 1998, 33, 509–543. [Google Scholar] [CrossRef]
- Stoel, R.D.; Peetsma, T.T.D.; Roeleveld, J. Relations between the development of school investment, self-confidence, and language achievement in elementary education: A multivariate latent growth curve approach. Learn. Individ. Differ. 2003, 13, 313–333. [Google Scholar] [CrossRef]
- Tisak, J.; Meredith, W. Descriptive and Associative Developmental Models. In Statistical Methods in Longitudinal Research; Academic Press: Cambridge, MA, USA, 1990; Volume 2, pp. 387–406. [Google Scholar]
- Byrne, B.M.; Crombie, G. Modeling and Testing Change: An Introduction to the Latent Growth Curve Model. Underst. Stat. 2003, 2, 177–203. [Google Scholar] [CrossRef]
- Whittaker, T.A.; Pituch, K.A.; McDougall, G.J. Latent Growth Modeling with Domain-Specific Outcomes Comprised of Mixed Response Types in Intervention Studies. J. Consult. Clin. Psychol. 2014, 82, 746–759. [Google Scholar] [CrossRef]
- Koo, N.; Leite, W.L.; Algina, J. Mediated Effects with the Parallel Process Latent Growth Model: An Evaluation of Methods for Testing Mediation in the Presence of Nonnormal Data. Struct. Equ. Modeling 2015, 23, 32–44. [Google Scholar] [CrossRef]
- Cheong, J.; MacKinnon, D.P.; Khoo, S.T. Investigation of Mediational Processes Using Parallel Process Latent Growth Curve Modeling. Struct. Equ. Modeling 2003, 10, 238–262. [Google Scholar] [CrossRef]
- Hancock, G.R.; Harring, J.R.; Lawrence, F.R. Using latent growth models to evaluate longitudinal change. In Structural Equation Modeling: A Second Course; Information Age: Charlotte, NC, USA, 2013; pp. 309–342. [Google Scholar]
- MacCallum, R.C.; Kim, C.; Malarkey, W.B.; Kiecolt-Glaser, J.K. Studying multivariate change using multilevel models and latent curve models. Multivariate Behav. Res. 1997, 32, 215–253. [Google Scholar] [CrossRef]
- Duncan, T.E.; Duncan, S.C.; Strycker, L.A. An Introduction to Latent Variable Growth Curve Modeling: Concepts, Issues, and Application, 2nd ed.; Routledge Academic: New York, NY, USA, 2013. [Google Scholar]
- Willett, J.B.; Keiley, M.K. Using Covariance Structure Analysis to Model Change Over Time. In Handbook of Applied Multivariate Statistics and Mathematical Modeling Humanities; Academic Press: San Diego, CA, USA, 2000; pp. 665–694. [Google Scholar]
- Rubin, D.B. Inference and Missing Data. Biometrika 1976, 63, 581–592. [Google Scholar] [CrossRef]
- Li, M.; Chen, N.; Cui, Y.; Liu, H. Comparison of different LGM-based methods with MAR and MNAR dropout data. Front. Psychol. 2017, 8, 722. [Google Scholar] [CrossRef]
- Curran, P.J.; Obeidat, K.; Losardo, D. Twelve Frequently Asked Questions about Growth Curve Modeling. J. Cogn. Dev. 2010, 11, 121–136. [Google Scholar] [CrossRef] [PubMed]
- Muthén, B.O.; Curran, P.J. General longitudinal modeling of individual differences in experimental designs: A latent variable framework for analysis and power estimation. Psychol. Methods 1997, 2, 371–402. [Google Scholar] [CrossRef]
- Fan, X. Power of Latent Growth Modeling for Detecting Group Differences in Linear Growth Trajectory Parameters. Struct. Equ. Modeling 2003, 10, 380–400. [Google Scholar] [CrossRef]
- Fan, X. Power of Latent Growth Modeling for Detecting Linear Growth: Number of Measurements and Comparison with Other Analytic Approaches. J. Exp. Educ. 2005, 73, 121–139. [Google Scholar] [CrossRef]
- Hertzog, C.; von Oertzen, T.; Ghisletta, P.; Lindenberger, U. Evaluating the Power of Latent Growth Curve Models to Detect Individual Differences in Change. Struct. Equ. Model. 2008, 15, 541–563. [Google Scholar] [CrossRef]
- Von Oertzen, T.; Ghisletta, P.; Lindenberger, U. Simulating Statistical Power in Latent Growth Curve Modeling: A Strategy for Evaluating Age-based Changes in Cognitive Resources. In Resource-Adaptive Cognitive Processes; Springer: Berlin/Heidelberg, Germany, 2010; pp. 95–117. [Google Scholar]
- Rast, P.; Hofer, S.M. Longitudinal design considerations to optimize power to detect variances and covariances among rates of change: Simulation results based on actual longitudinal studies. Psychol. Methods 2014, 19, 133–154. [Google Scholar] [CrossRef]
- Kline, R.B. Principles and Practice of Structural Equation Modeling, 3rd ed.; Guilford Publications: New York, NY, USA, 2010. [Google Scholar]
- Cheong, J. Accuracy of Estimates and Statistical Power for Testing Meditation in Latent Growth Curve Modeling. Struct. Equ. Modeling 2011, 18, 195–211. [Google Scholar] [CrossRef] [PubMed]
- Raudenbush, S.W.; Liu, X.F. Effects of Study Duration, Frequency of Observation, and Sample Size on Power in Studies of Group Differences in Polynomial Change. Psychol. Methods 2001, 6, 387–401. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J. Statistical Power Analysis for the Behavioral Sciences, 2nd ed.; Lawrence Erlbaum Associates: Hillsdale, NJ, USA, 1988. [Google Scholar]
- Frane, A.V. Power and Type I Error Control for Univariate Comparisons in Multivariate Two-Group Designs. Multivariate Behav. Res. 2015, 50, 233–247. [Google Scholar] [CrossRef]
- Muthén, L.K.; Muthén, B.O. How to Use a Monte Carlo Study to Decide on Sample Size and Determine Power. Struct. Equ. Modeling 2002, 9, 599–620. [Google Scholar] [CrossRef]
- Muthén, L.; Muthén, B. Mplus User’s Guide (Version 7.4); Muthén & Muthén: Los Angeles, CA, USA, 2012. [Google Scholar] [CrossRef]
- Thoemmes, F.; MacKinnon, D.P.; Reiser, M.R. Power Analysis for Complex Mediational Designs Using Monte Carlo Methods. Struct. Equ. Modeling 2010, 17, 510–534. [Google Scholar] [CrossRef]
- Diallo, T.M.O.; Morin, A.J.S.; Parker, P.D. Statistical power of latent growth curve models to detect quadratic growth. Behav. Res. Methods 2013, 46, 357–371. [Google Scholar] [CrossRef] [PubMed]
- R Core Team. R: A Language and Environment for Statistical Computing (Version 3.4.0) [Computer Software]. R Foundation for Statistical Computing. 2017. Available online: https://www.R-project.org/ (accessed on 31 December 2025).
- Bradley, J.V. Robustness? Br. J. Math. Stat. Psychol. 1978, 31, 144–152. [Google Scholar] [CrossRef]
- Little, T.D. Longitudinal Structural Equation Modeling; Guilford Press: New York, NY, USA, 2013. [Google Scholar]
- Grimm, K.J.; Ram, N.; Estabrook, R. Growth Modeling: Structural Equation and Multilevel Modeling Approaches; Guilford Press: New York, NY, USA, 2016. [Google Scholar]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.