A Generative Network Model of the Human Brain Normal Aging Process
Abstract
1. Introduction
1.1. Brain Ageing
1.2. Brain Network Modelling
1.3. Contributions of the Research
2. Materials and Methods
2.1. Data Acquisition and Processing
2.2. Construction of fMRI Brain Networks
2.3. Topological Measures in Functional MRI Brain Networks
3. Generative Network Model of Functional Brain Network
3.1. Generative Scheme of Artificial Brain Network
- Initialization: Brain network extraction is started by image pre-processing. Experimental brain fMRI datasets of participants across all age groups (group Young, group Mid-age and group Old) are processed, and the relevant initial fMRI brain networks Gy = (gy1, gy2, …, gyn), Gm = (gm1, gm2, …, gmn) and Go = (go1, go2, …, gon) are obtained. Each initial network consists of a fixed number of nodes │V│ = 90. Edge sets of participants in groups young, mid-age and old are represented as Ey = (ey1, ey2, …, eyn), Em = (em1, em2, …, emn), Eo = (eo1, eo2, …, eon), with the threshold θ value is being set to 0.5.
- Growth: After initialization, artificial brain networks start to evolve from the status of young age. For each brain network, one interested node v out of 90 ROIs is selected according to probability pv-see Equation (1) below—and the degree of node v is requested no less than 2. A connection is expected to be established between node v′s two neighbour nodes i and j. Node i is selected firstly from v′s neighbour nodes set |Γ(v)|. Moreover, node i is not connected with at least one node in |Γ(v)|. Otherwise, a new interested node v must be selected again based on pv. The network evolution process will not progress until the new qualified node v is found. Connection probability pi,j of node i and the remaining neighbours (which are necessarily unconnected to node i) is calculated by the connection likelihood model. A link will then be established according to the highest connection likelihood, in between node i and j. This step continues to the next step when each brain network within the group succeeds in adding a link.
- Two-sample t-test: After a link is established in each network, the t-test is executed to test the difference between the artificial brain networks and the fMRI brain networks (group Mid-age and group Old, respectively), in comparing the number of edges, which is used to evaluate how significant the difference is.
- Process of iterations: Artificial brain networks formation is a process of iterations. Step 2 and step 3 constitute one iteration, which adds one link to the network and performs a two-sample t-test. As new links are being added to artificial brain networks, the p-value of t-test increases. Therefore, the formation process ends when p-value stops increasing. The artificial brain network with the greatest p-value at the current iteration is considered as the final artificial brain network.The selection probability pv of selecting an interested node v is expressed by Equation (1):where kv represents the degree of node v, the pv is positive correlated to the degree of node v. This strategy allows nodes with larger degrees to get a larger probability of being selected.
3.2. Connection Likelihood Model-LNBE
4. Statistical Evaluation of Generative Network Models
5. Results
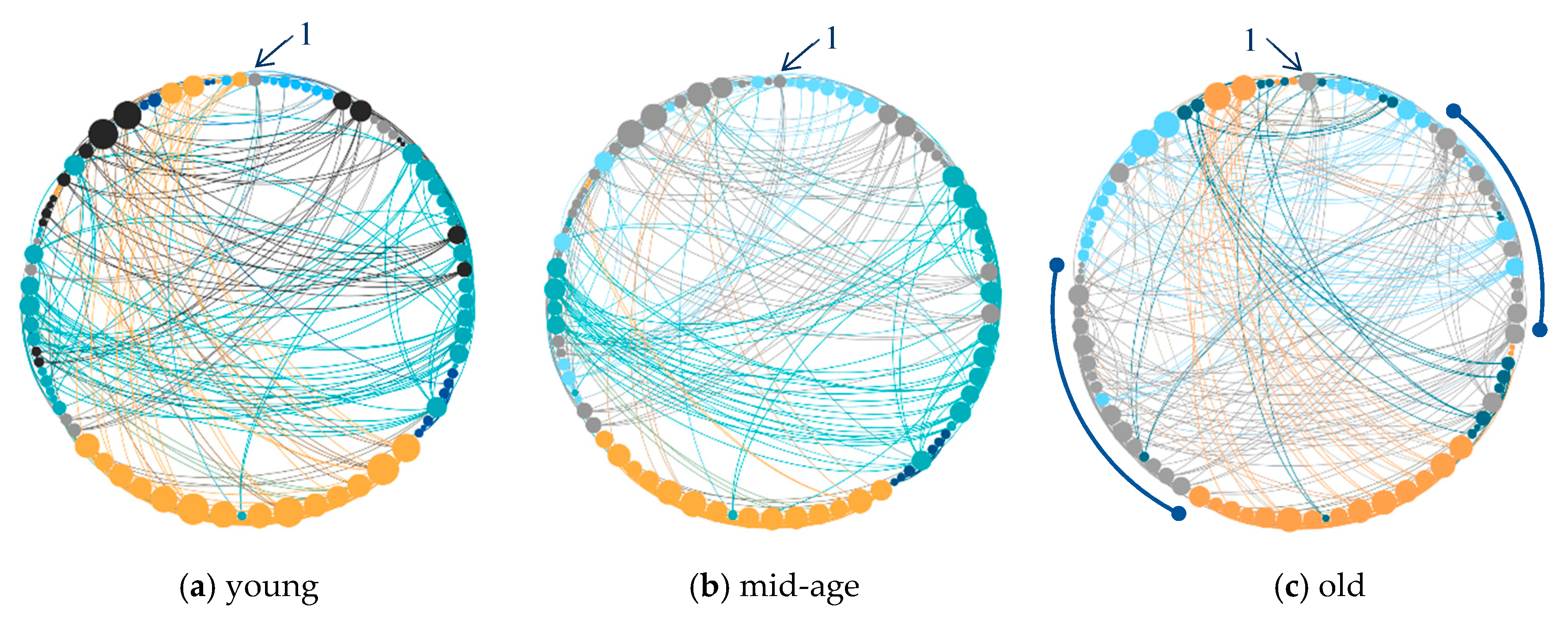
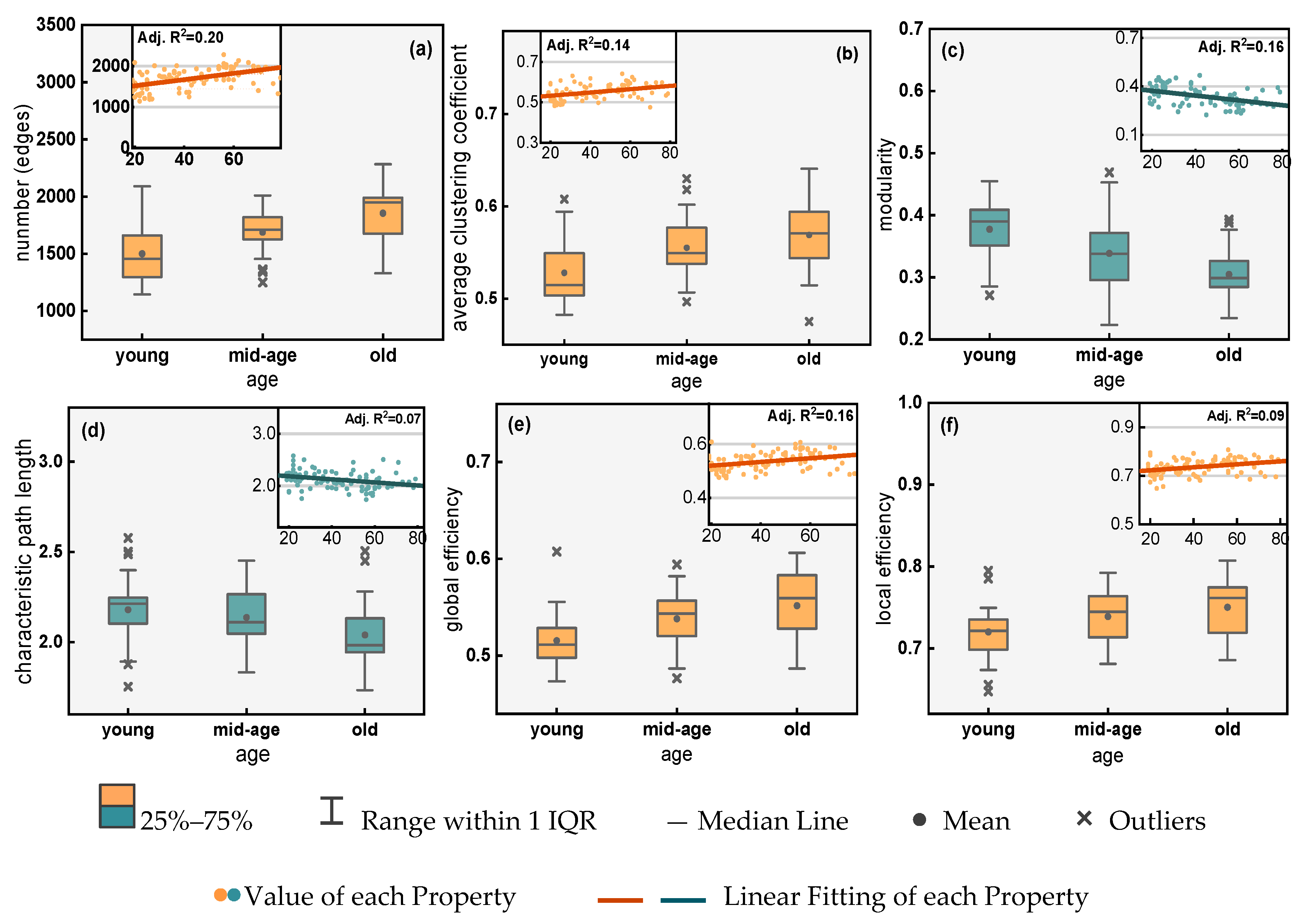
5.1. Ageing of the Functional Brain Network
5.2. Evaluation Results—Comparison of the LNBE Model Versus Mechanistic Generative Network Models
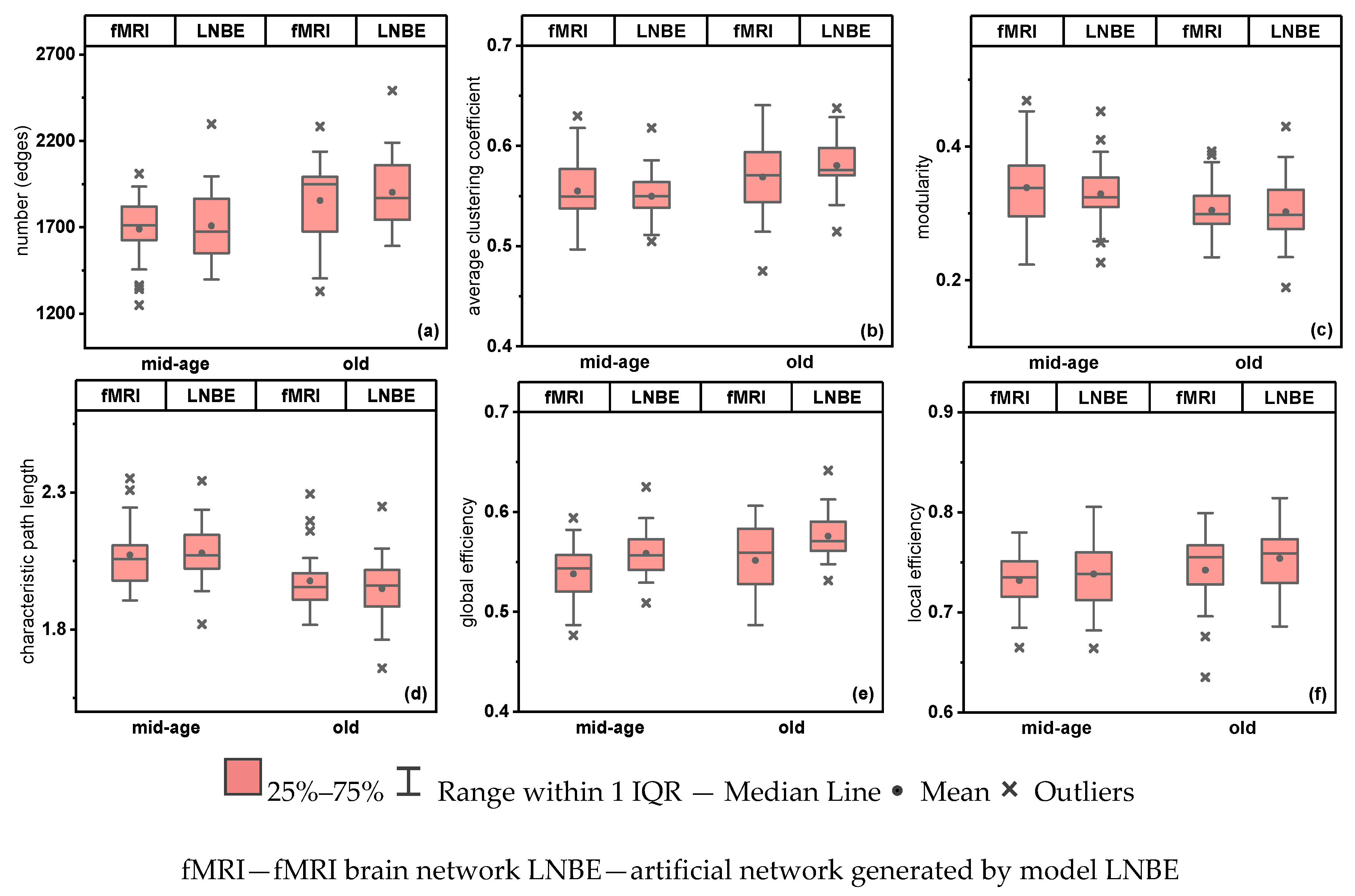
5.3. Model LNBE—In Comparison with Functional Brain Networks
6. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Isaev, N.K.; Genrikhs, E.E.; Oborina, M.V.; Stelmashook, E.V. Accelerated aging and aging process in the brain. Rev. Neurosci. 2018, 29, 233–240. [Google Scholar] [CrossRef]
- Martin, G.M. Genetic syndromes in man with potential relevance to the pathobiology of aging. Birth Defects Orig Artic. Ser. 1978, 14, 5–39. [Google Scholar] [PubMed]
- Martin, G.M. Genetics and aging; the Werner syndrome as a segmental progeroid syndrome. Adv. Exp. Med. Biol. 1985, 190, 161. [Google Scholar] [PubMed]
- He, Y.; Chen, Z.; Evans, A. Structural Insights into Aberrant Topological Patterns of Large-Scale Cortical Networks in Alzheimer’s Disease. J. Neurosci. 2008, 28, 4756–4766. [Google Scholar] [CrossRef] [PubMed]
- Isaev, N.K.; Stelmashook, E.V.; Stelmashook, N.N.; Sharonova, I.N.; Skrebitsky, V.G. Brain aging and mitochondria-targeted plastoquinone antioxidants of SkQ-type. Biochemistry 2013, 78, 295–300. [Google Scholar] [CrossRef]
- Xekardaki, A.; Eniko, K.; Gold, G. Neuropathological changes in aging brain. Adv. Exp. Med. Biol. 2015, 821, 11. [Google Scholar]
- Pannese, E. Morphological changes in nerve cells during normal aging. Brain Struct. Funct. 2011, 216, 85–89. [Google Scholar] [CrossRef]
- Sporns, O.; Tononi, G.; Kötter, R. The human connectome: A structural description of the human brain. PLoS Comput. Biol. 2005, 1, e42. [Google Scholar] [CrossRef]
- Hagmann, P. From Diffusion MRI to Brain Connectomics. Ph.D. Thesis, Ecole Polytechnique Fédérale de Lausanne, Lausanne, Switzerland, 2005. [Google Scholar]
- Yap, P.T.; Wu, G.; Shen, D. Human Brain Connectomics: Networks, Techniques, and Applications [Life Sciences]. IEEE Signal Process. Mag. 2010, 27, 131–134. [Google Scholar] [CrossRef]
- Barthélemy, M. Spatial networks. Phys. Rep. 2011, 499, 1–101. [Google Scholar] [CrossRef]
- Vértes, P.E.; Alexander-Bloch, A.F.; Gogtay, N.; Giedd, J.N.; Rapoport, J.L.; Bullmore, E.T. Simple models of human brain functional networks. Proc. Natl. Acad. Sci. USA 2012, 109, 5868–5873. [Google Scholar] [CrossRef] [PubMed]
- Friston, K.J. Functional and effective connectivity, Neuroimaging: A synthesis. Hum. Brain Map. 1994, 2, 56–78. [Google Scholar] [CrossRef]
- Mesulam, M. Large-scale neurocognitive networks and distributed processing for attention, language, and memory. Ann. Neurol. 1900, 28, 597–613. [Google Scholar] [CrossRef] [PubMed]
- McIntosh, A.R. Towards a network theory of cognition. Neural Netw. 2000, 13, 861–870. [Google Scholar] [CrossRef]
- Bressler, S.L.; Menon, V. Large-scale brain networks in cognition: Emerging methods and principles. Trends Cogn. Sci. 2010, 14, 277–290. [Google Scholar] [CrossRef]
- Sporns, O. Contributions and challenges for network models in cognitive neuroscience. Nat. Neurosci. 2014, 17, 652–660. [Google Scholar] [CrossRef]
- Medaglia, J.D.; Lynall, M.-E.; Bassett, D.S. Cognitive network neuroscience. J. Cogn. Neurosci. 2015, 27, 1471–1491. [Google Scholar] [CrossRef]
- Stillman, P.E.; Wilson, J.D.; Denny, M.J.; Desmarais, B.A.; Bhamidi, S.; Cranmer, S.J.; Lu, Z.L. Statistical Modeling of the Default Mode Brain Network Reveals a Segregated Highway Structure. Sci. Rep. 2017, 7, 11694. [Google Scholar] [CrossRef]
- Kaiser, M. Modelling the development of cortical networks. Neurocomputing 2004, 58–60, 297–302. [Google Scholar] [CrossRef]
- Zhao, Q.-B.; Feng, H.-B.; Tang, Y.-Y. Modelling human cortical network in real brain space. Chin. Phys. Lett. 2007, 24, 3582. [Google Scholar]
- Cheng, C.; Chen, J.; Cao, X.; Guo, H. Comparison of local information indices applied in resting state functional brain network connectivity prediction. Front. Neurosci. 2016, 10, 585–599. [Google Scholar] [CrossRef] [PubMed]
- Betzel, R.F.; Avena-Koenigsberger, A.; Goñi, J. Generative models of the human connectome. Neuroimage 2016, 124, 1054–1064. [Google Scholar] [CrossRef] [PubMed]
- Liben-Nowell, D.; Kleinberg, J. The Link Prediction Problem for Social Networks. J. Am. Soc. Inf. Sci. Technol. 2007, 58, 1019–1031. [Google Scholar] [CrossRef]
- Lü, L.; Zhou, T. Link prediction in complex networks: A survey. Phys. A Stat. Mech. Appl. 2011, 390, 1150–1170. [Google Scholar] [CrossRef]
- Liu, Z.; Zhang, Q.M.; Lü, L. Link prediction in complex networks: A local naïve Bayes model. Europhys. Lett. 2011, 96, 48007. [Google Scholar] [CrossRef]
- Available online: http://fcon_1000.projects.nitrc.org/ (accessed on 1 January 2019).
- Biswal, B.B.; Mennes, M.; Zuo, X.N. Toward discovery science of human brain function. Proc. Natl. Acad. Sci. USA 2010, 107, 4734–4739. [Google Scholar] [CrossRef]
- Yan, C.; Zang, Y. DPARSF: A MATLAB toolbox for “pipeline” data analysis of resting-state fMRI. Front. Syst. Neurosci. 2010, 4, 13. [Google Scholar] [CrossRef]
- Tzourio-Mazoyer, N.; Landeau, B.; Papathanassiou, D.; Crivello, F.; Etard, O.; Delcroix, N.; Mazyoer, B.; Joliot, M. Automated anatomical labeling of activations in SPM using a macroscopic anatomical parcellation of the MNI MRI single-subject brain. Neuroimage 2002, 15, 273–289. [Google Scholar] [CrossRef]
- Betzel, R.F.; Fukushima, M.; He, Y. Dynamic fluctuations coincide with periods of high and low modularity in resting-state functional brain networks. NeuroImage 2016, 127, 287–297. [Google Scholar] [CrossRef]
- Hayasaka, S.; Laurienti, P.J. Comparison of characteristics between region-and voxel-based network analyses in resting-state fMRI data. Neuroimage 2009, 50, 499–508. [Google Scholar] [CrossRef]
- Garrison, K.A.; Scheinost, D.; Finn, E.S.; Shen, X.; Constable, R.T. The (in)stability of functional brain network measures across thresholds. Neuroimage 2015, 118, 651–661. [Google Scholar] [CrossRef] [PubMed]
- Rubinov, M.; Sporns, O. Complex network measures of brain connectivity: Uses and interpretations. Neuroimage 2010, 52, 1059–1069. [Google Scholar] [CrossRef] [PubMed]
- Newman, M. Modularity and community structure in networks. Proc. Natl. Acad. Sci. USA 2006, 103, 8577–8582. [Google Scholar] [CrossRef] [PubMed]
- Newman, M.E.J. Fast algorithm for detecting community structure in networks. Phys. Rev. E 2004, 69, 66133. [Google Scholar] [CrossRef]
- Watts, D.J.; Strogatz, S.H. Collective dynamics of “small-world” networks. Nature 1998, 393, 440–442. [Google Scholar] [CrossRef]
- Telesford, Q.K.; Joyce, K.E.; Hayasaka, S. The ubiquity of small-world networks. Brain Connect. 2011, 1, 367–375. [Google Scholar] [CrossRef]
- Ajilore, O.; Lamar, M.; Kumar, A. Association of brain network efficiency with aging, depression, and cognition. Am. J. Geriatr. Psychiatry 2013, 22, 102–110. [Google Scholar] [CrossRef]
- Wen, W.; Zhu, W.; He, Y. Discrete neuroanatomical networks are associated with specific cognitive abilities in old age. J. Neurosci. 2011, 31, 1204–1212. [Google Scholar] [CrossRef]
- Barabási, A.-L.; Albert, R. Emergence of scaling in random networks. Science 1999, 286, 509–512. [Google Scholar] [CrossRef]
- Newman, M. Networks: An Introduction; Oxford University Press: Oxford, UK, 2010; p. 8. [Google Scholar]
- Newman, M.E.J. Clustering and preferential attachment in growing networks. Phys. Rev. E 2001, 64, 025102. [Google Scholar] [CrossRef]
- Jaccard, P. Distribution de la flore alpine dans le bassin des Dranses et dans quelques régions voisines. Bull. Soc. Vaud. Sci. Nat. 1901, 37, 241–272. [Google Scholar]
- Cannistraci, C.V.; Alanis-Lobato, G.; Ravasi, T. From link-prediction in brain connectomes and protein interactomes to the local-community-paradigm in complex networks. Sci. Rep. 2013, 3, 1613. [Google Scholar] [CrossRef] [PubMed]
| Group Classification | Number of Participants | Sex Ratio (M/F) | Age Range | Mean Age |
|---|---|---|---|---|
| Young | 30 | 13/17 | 19–30 | 22.7 |
| Mid-age | 30 | 16/14 | 31–53 | 42.6 |
| Old | 30 | 12/18 | 54–79 | 61.3 |
| Comparision | Model | E | C | Q | L | Eglbl | Elcl | Energy |
|---|---|---|---|---|---|---|---|---|
| (Group Mid-age) artificial brain network vs. fMRI brain network | LNBE | 0.9863 | 0.3766 | 0.8346 | 0.7501 | 0.9953 | 0.5365 | 4.4794 |
| CN | 0.9863 | 0.1694 | 0.7246 | 0.7338 | 0.9866 | 0.4997 | 4.1004 | |
| CAR | 0.9863 | 0.2761 | 0.4875 | 0.0590 | 0.9158 | 0.5279 | 3.2525 | |
| CRA | 0.9863 | 0.2761 | 0.3229 | 0.9799 | 0.8923 | 0.0041 | 3.4617 | |
| BA | 0.9863 | 0.4812 | 0.4770 | 0.0092 | 0.8800 | 0.1510 | 2.9847 | |
| JC | 0.9863 | 0.3902 | 0.4371 | 0.0119 | 0.2643 | 0.1430 | 2.2328 | |
| Random | 0.9863 | 0.1274 | 0.6284 | 0.0152 | 0.0382 | 0.0965 | 1.8920 | |
| (Group Old) artificial brain network vs. fMRI brain network | LNBE | 0.9927 | 0.0215 | 0.5624 | 0.6437 | 0.8775 | 0.3043 | 3.4021 |
| CN | 0.9927 | 0.0104 | 0.4046 | 0.5923 | 0.8316 | 0.1836 | 3.0132 | |
| CAR | 0.9863 | 0.7095 | 0.0442 | 0.9599 | 0.2632 | 0.0162 | 2.9794 | |
| CRA | 0.9863 | 0.5151 | 0.1492 | 0.5650 | 0.9346 | 0.0298 | 3.1800 | |
| BA | 0.9863 | 0.0496 | 0.0017 | 0.2778 | 0.0756 | 0.1008 | 1.4919 | |
| JC | 0.9863 | 0.4574 | 0.0321 | 0.2294 | 0.0223 | 0.5938 | 2.3212 | |
| Random | 0.9927 | 0.0022 | 0.1502 | 0.0051 | 0.0407 | 0.4003 | 1.5912 |
 range of values of the index of comparison among group mid-age (lowest–highest).
range of values of the index of comparison among group mid-age (lowest–highest).  range of values of the index of comparison group among old (lowest–highest).
range of values of the index of comparison group among old (lowest–highest).© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Si, S.; Hu, B.; Zhao, H.; Zhu, J. A Generative Network Model of the Human Brain Normal Aging Process. Symmetry 2020, 12, 91. https://doi.org/10.3390/sym12010091
Liu X, Si S, Hu B, Zhao H, Zhu J. A Generative Network Model of the Human Brain Normal Aging Process. Symmetry. 2020; 12(1):91. https://doi.org/10.3390/sym12010091
Chicago/Turabian StyleLiu, Xiao, Shuaizong Si, Bo Hu, Hai Zhao, and Jian Zhu. 2020. "A Generative Network Model of the Human Brain Normal Aging Process" Symmetry 12, no. 1: 91. https://doi.org/10.3390/sym12010091
APA StyleLiu, X., Si, S., Hu, B., Zhao, H., & Zhu, J. (2020). A Generative Network Model of the Human Brain Normal Aging Process. Symmetry, 12(1), 91. https://doi.org/10.3390/sym12010091






