Transcription Factors Responding to Pb Stress in Maize
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples and RNA Isolation
2.2. Gene Annotation, Differentially Expressed TFs, Enrichment Analysis and Cluster Analysis
2.3. Real-Time PCR
2.4. Identification of Arabidopsis Mutant Pb Tolerance
2.5. Intracellular Localization of Transcription Factors
3. Results and Discussion
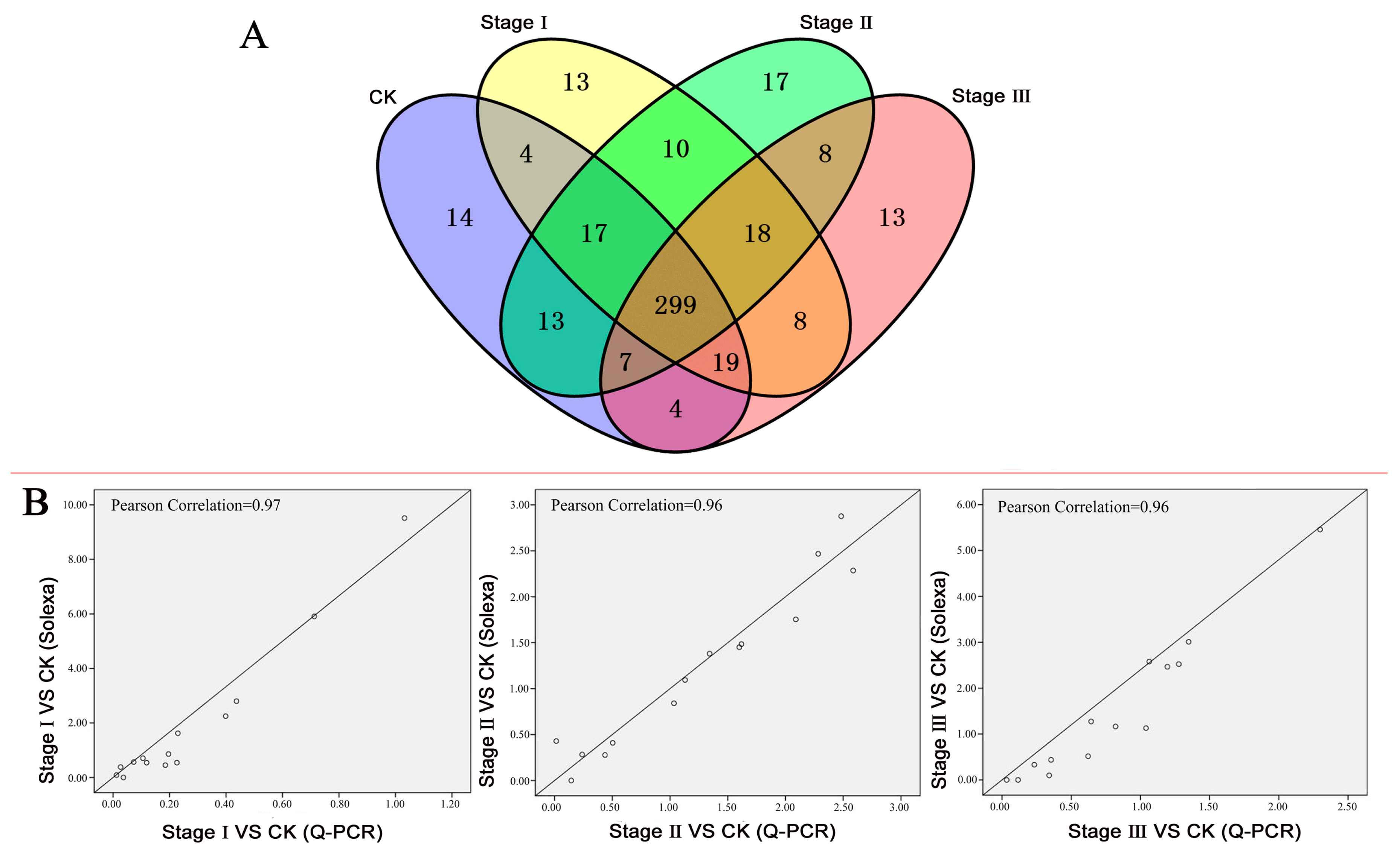
3.1. The TFs Regulated by Pb: A Summary
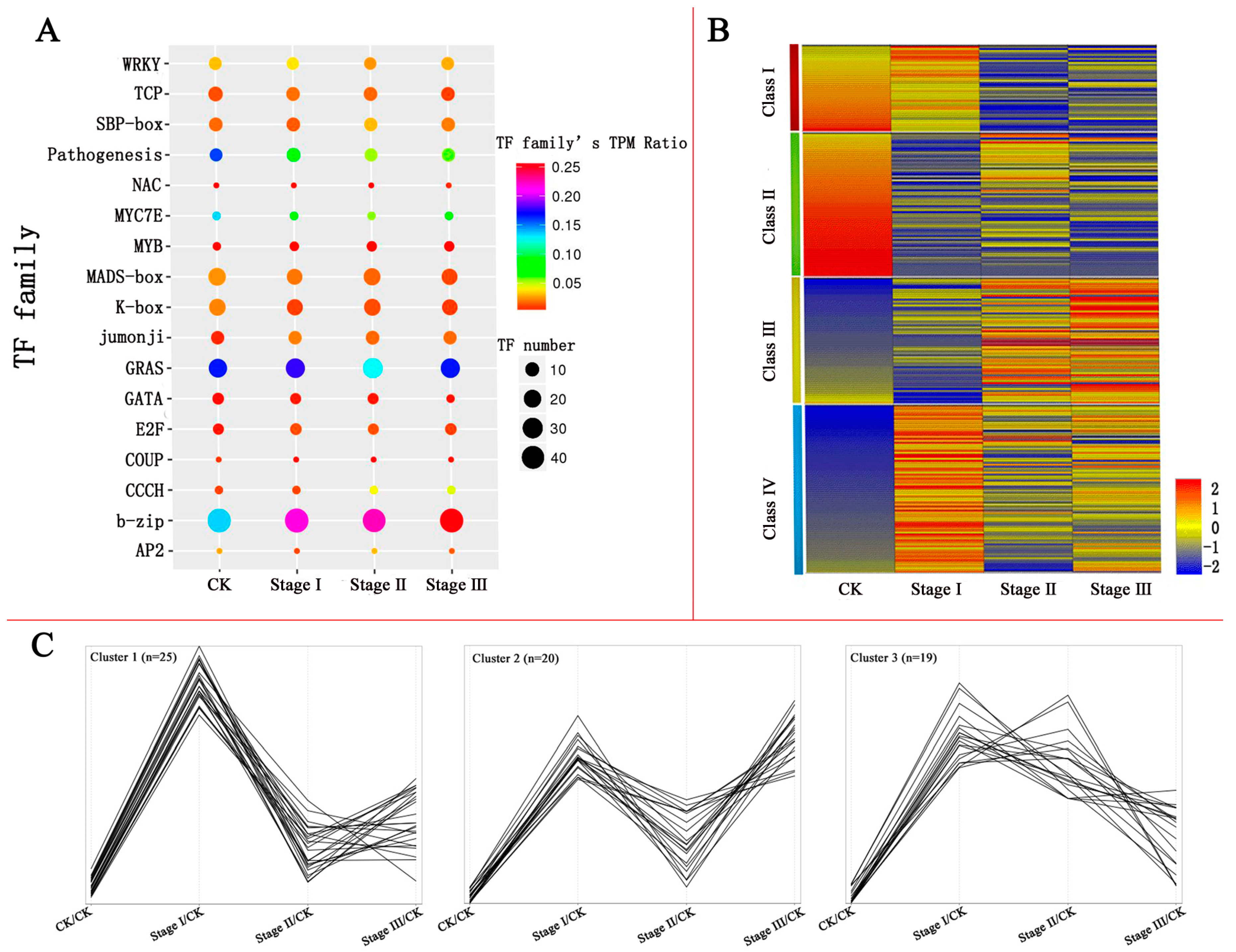
3.2. Differentially Expressed TFs (DETs) and TF Families
3.3. Validation of the DET Expression Profiles by Quantitative RT-PCR
3.4. Clustering of DETs Based on Their Expression Patterns after Pb Treatment
3.5. Pathways Regulated by DETs under Pb Treatment
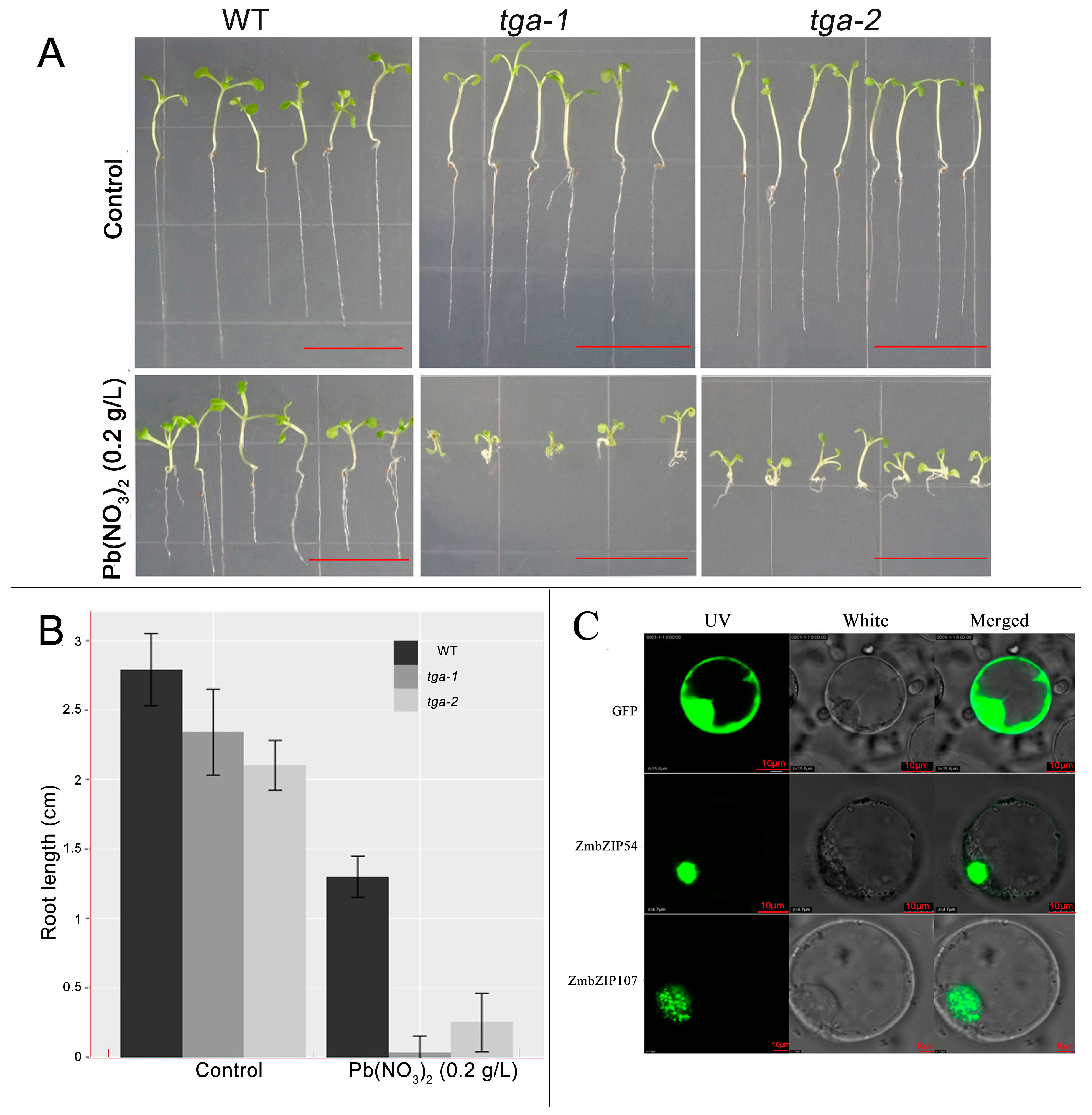
3.6. Potential Function of bZIP Genes in Pb Tolerance
3.7. ZmbZIP54-GFP and ZmbZIP107-GFP Fusion Proteins Are Both Located in the Nucleus
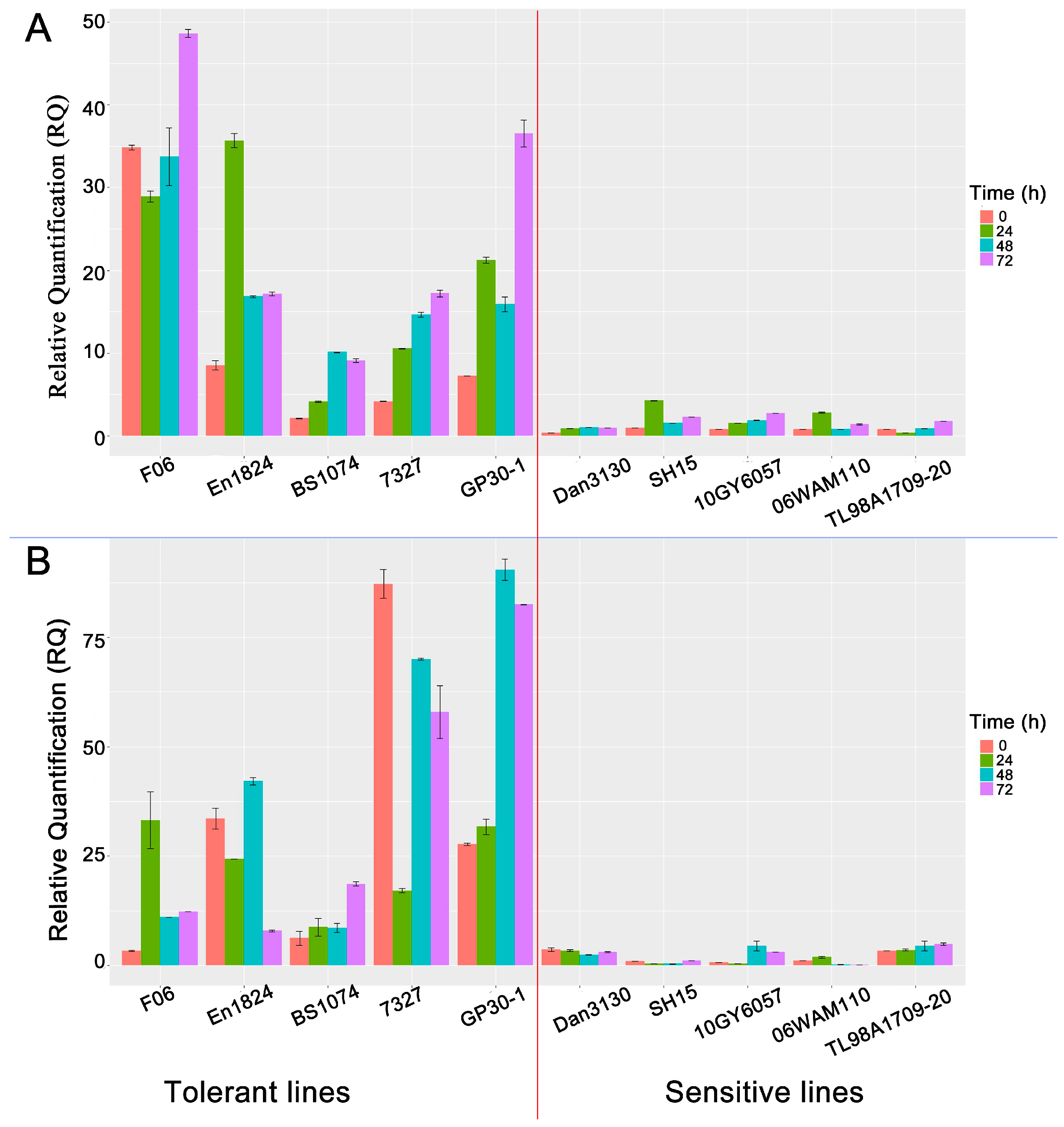
3.8. Disparity in bZIP Expression Levels Accounts for Pb-Tolerance Difference among Maize Lines
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Dey, S.K.; Dey, J.; Patra, S.; Pothal, D. Changes in the antioxidative enzyme activities and lipid peroxidation in wheat seedlings exposed to cadmium and lead stress. Braz. J. Plant Physiol. 2007, 19, 53–60. [Google Scholar] [CrossRef]
- Lan-Min, A.N.; Niu, Y.J.; Bing, X.U.; Liu, N. Influence of lead on the apoptosis and the expression of fos,jun,p53 and nitric oxide synthase in rat brain. Carcinog. Teratog. Mutagen. 2006, 18, 359–362. [Google Scholar]
- Latchman, D.S. Transcription factors: An overview. Int. J. Biochem. Cell Biol. 1993, 74, 417–422. [Google Scholar] [CrossRef]
- Yusuf, D.; Butland, S.L.; Swanson, M.I.; Bolotin, E.; Ticoll, A.; Cheung, W.A.; Zhang, X.Y.C.; Dickman, C.T.; Fulton, D.L.; Lim, J.S. The transcription factor encyclopedia. Genome Biol. 2012, 13, R24. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, P.K.; Jha, B. Transcription factors in plants and aba dependent and independent abiotic stress signaling. Biol. Plant. 2010, 54, 201–212. [Google Scholar] [CrossRef]
- Babitha, K.C.; Ramu, S.V.; Pruthvi, V.; Mahesh, P.; Nataraja, K.N.; Udayakumar, M. Co-expression of at bHLH17 and at WRKY 28 confers resistance to abiotic stress in Arabidopsis. Transgenic Res. 2013, 22, 327–341. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Zhang, D.; Li, H.; Wang, Y.; Zhang, Y.; Wood, A.J. EsDREB2B, a novel truncated DREB2-type transcription factor in the desert legumeeremosparton songoricum, enhances tolerance to multiple abiotic stresses in yeast and transgenic tobacco. BMC Plant Biol. 2014, 14, 44. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Zhang, L.; Xia, C.; Zhao, G.; Liu, J.; Jia, J.; Kong, X. A novel wheat bZIP transcription factor, TabZIP60, confers multiple abiotic stress tolerances in transgenic Arabidopsis. Physiol. Plant. 2015, 153, 538–554. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Zhou, J.; Jie, Y.; Xing, H.; Zhong, Y.; She, W.; Wei, G.; Yu, W.; Ma, Y. A ramie (boehmeria nivea) bzip transcription factor BnbZIP3 positively regulates drought, salinity and heavy metal tolerance. Mol. Breed. 2016, 36, 1–15. [Google Scholar] [CrossRef]
- Ying, S.; Zhang, D.F.; Fu, J.; Shi, Y.S.; Song, Y.C.; Wang, T.Y.; Li, Y. Cloning and characterization of a maize bZIP transcription factor, ZmbZIP72, confers drought and salt tolerance in transgenic arabidopsis. Planta 2012, 235, 253–266. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Jin, F.; Wang, C.; Luo, J.; Lin, H.; Xiang, K.; Liu, L.; Zhao, M.; Zhang, Y.; Ding, H. Difference between pb and cd accumulation in 19 elite maize inbred lines and application prospects. BioMed Res. Int. 2012, 2012, 271485. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Zhang, Y.; Chen, J.; Lin, H.; Zhao, M.; Peng, H.; Liu, L.; Yuan, G.; Zhang, S.; Zhang, Z. Genome expression profile analysis reveals important transcripts in maize roots responding to the stress of heavy metal pb. Physiol. Plant. 2013, 147, 270–282. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.L.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T. Gene Ontology: Tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef] [PubMed]
- Koonin, E.V.; Fedorova, N.D.; Jackson, J.D.; Jacobs, A.R.; Krylov, D.M.; Makarova, K.S.; Mazumder, R.; Mekhedov, S.L.; Nikolskaya, A.N.; Rao, B.S. A comprehensive evolutionary classification of proteins encoded in complete eukaryotic genomes. Genome Biol. 2004, 5, R7. [Google Scholar] [CrossRef] [PubMed]
- Xie, C.; Mao, X.; Huang, J.; Ding, Y.; Wu, J.; Dong, S.; Kong, L.; Gao, G.; Li, C.Y.; Wei, L. KOBAS 2.0: A web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res. 2011, 39, 316–322. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.; Jiang, Z.; Yao, X.; Zhang, Z.; Lin, H.; Zhao, M.; Liu, H.; Peng, H.; Li, S.; Pan, G. Genome expression profile analysis of the immature maize embryo during dedifferentiation. PLoS ONE 2012, 7, e32237. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Su, J.; Duan, S.; Ao, Y.; Dai, J.; Liu, J.; Wang, P.; Li, Y.; Liu, B.; Feng, D. A highly efficient rice green tissue protoplast system for transient gene expression and studying light/chloroplast-related processes. Plant Methods 2011, 7, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Yu, H.Q.; Yong, T.M.; Li, H.J.; Liu, Y.P.; Zhou, S.F.; Fu, F.L.; Li, W.C. Overexpression of a phospholipase Dα gene from Ammopiptanthus nanus enhances salt tolerance of phospholipase Dα1-deficient Arabidopsis mutant. Planta 2015, 242, 1495–1509. [Google Scholar] [CrossRef] [PubMed]
- Ge, F.; Luo, X.; Huang, X.; Zhang, Y.; He, X.; Liu, M.; Lin, H.; Peng, H.; Li, L.; Zhang, Z. Genome-wide analysis of transcription factors involved in maize embryonic callus formation. Physiol. Plant. 2016, 158, 452–462. [Google Scholar] [CrossRef] [PubMed]
- Zvi, P.; Eduardo, B. Hormone balance and abiotic stress tolerance in crop plants. Curr. Opin. Plant Biol. 2011, 14, 290. [Google Scholar]
- Sunkar, R.; Chinnusamy, V.; Zhu, J.; Zhu, J.K. Small rnas as big players in plant abiotic stress responses and nutrient deprivation. Trends Plant Sci. 2007, 12, 301–309. [Google Scholar] [CrossRef] [PubMed]
- Finkelstein, R.R.; Lynch, T.J. The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 2000, 12, 599. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Molina, L.; Mongrand, S.; Chua, N.H. A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc. Natl. Acad. Sci. USA 2001, 98, 4782–4787. [Google Scholar] [CrossRef] [PubMed]
- Uno, Y.; Furihata, T.; Abe, H.; Yoshida, R.; Shinozaki, K.; Yamaguchishinozaki, K. Arabidopsis basic leucine zipper transcription factors involved in an abscisic acid-dependent signal transduction pathway under drought and high-salinity conditions. Proc. Natl. Acad. Sci. USA 2000, 97, 11632–11637. [Google Scholar] [CrossRef] [PubMed]
- Michalak, A. Phenolic compounds and their antioxidant activity in plants growing under heavy metal stress. Polish J. Environ. Stud. 2006, 15, 523–530. [Google Scholar]
- Ware, D.; Jaiswal, P.; Ni, J.; Pan, X.; Chang, K.; Clark, K.; Teytelman, L.; Schmidt, S.; Zhao, W.; Cartinhour, S. Gramene: A resource for comparative grass genomics. Nucleic Acids Res. 2002, 30, 103–105. [Google Scholar] [CrossRef] [PubMed]
- Swarbreck, D.; Wilks, C.; Lamesch, P.; Berardini, T.Z.; GarciaHernandez, M.; Foerster, H.; Li, D.; Meyer, T.; Muller, R.; Ploetz, L. The arabidopsis information resource (TAIR): Gene structure and function annotation. Nucleic Acids Res. 2008, 36, 1009–1014. [Google Scholar] [CrossRef] [PubMed]
- Meshi, T.; Iwabuchi, M. Plant transcription factors. Plant Cell Physiol. 1995, 36, 1405. [Google Scholar] [PubMed]
- Stotz, H.U.; Mueller, S.; Zoeller, M.; Mueller, M.J.; Berger, S. TGA transcription factors and jasmonate-independent COI1 signalling regulate specific plant responses to reactive oxylipins. J. Exp. Botany 2013, 64, 963. [Google Scholar] [CrossRef] [PubMed]
- Devoto, A.; Ellis, C.; Magusin, A.; Chang, H.S.; Chilcott, C.; Zhu, T.; Turner, J.G. Expression profiling reveals COI1 to be a key regulator of genes involved in wound- and methyl jasmonate-induced secondary metabolism, defence, and hormone interactions. Plant Mol. Biol. 2005, 58, 497–513. [Google Scholar] [CrossRef] [PubMed]
- Maksymiec, W.; Wianowska, D.; Dawidowicz, A.L.; Radkiewicz, S.; Mardarowicz, M.; Krupa, Z. The level of jasmonic acid in Arabidopsis thaliana and Phaseolus coccineus plants under heavy metal stress. J. Plant Physiol. 2005, 162, 1338. [Google Scholar] [CrossRef] [PubMed]
- Izawa, T.; Foster, R.; Chua, N.H. Plant bZIP protein DNA binding specificity. J. Mol. Biol. 1993, 230, 1131–1144. [Google Scholar] [CrossRef] [PubMed]
- Gietz, R.D.; Woods, R.A. Yeast transformation by the Liac/SS carrier DNA/PEG method. Methods Mol. Biol. 2006, 313, 107. [Google Scholar] [PubMed]
- Gao, J.; Zhang, Y.; Lu, C.; Peng, H.; Luo, M.; Li, G.; Shen, Y.; Ding, H.; Zhang, Z.; Pan, G. The development dynamics of the maize root transcriptome responsive to heavy metal pb pollution. Biochem. Biophys. Res. Commun. 2015, 458, 287–293. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.H.; Zheng, W.Y.; Zhang, X.K. Principal component analysis and comprehensive evaluation on morphological and agronomic traits of drought tolerance in rapeseed (brassica napus L.). Scientia Agric. Sinica 2011, 44, 1775–1787. [Google Scholar]
- Liu, J.H.; Huang, X.S.; Chen, X.J. Overexpression of PtrABF gene, a bZIP transcription factor isolated from Poncirus trifoliata, enhances dehydration and drought tolerance in tobacco via scavenging ros and modulating expression of stress-responsive genes. BMC Plant Biol. 2010, 10, 230. [Google Scholar]
- Lee, S.C.; Choi, H.W.; Hwang, I.S.; Du, S.C.; Hwang, B.K. Functional roles of the pepper pathogen-induced bZIP transcription factor, CAbZIP1, in enhanced resistance to pathogen infection and environmental stresses. Planta 2006, 224, 1209. [Google Scholar] [CrossRef] [PubMed]
| TFs | Gene Identifier | log2 (Stage I/CK) | log2 (Stage II/CK) | log2 (Stage III/CK) |
|---|---|---|---|---|
| bZIP | GRMZM2G381991 | 6.38 | −6.64 | 8.33 |
| bZIP115 | GRMZM2G157177 | 8.89 | 9.32 | 9.47 |
| bZIP55 | GRMZM2G386273 | 8.18 | 5.32 | 7.01 |
| bZIP58 | GRMZM2G149040 | 7.18 | 8.13 | 7.59 |
| bZIP71 | GRMZM2G019106 | −6.64 | −6.64 | 7.88 |
| bZIP83 | GRMZM2G000171 | 7.18 | 6.64 | 8.74 |
| CCAAT-binding | GRMZM2G033245 | 7.18 | 6.32 | −6.64 |
| e2f4 | AC233850.1_FG005 | 7.54 | −6.64 | 5.43 |
| e2f5 | GRMZM2G050590 | 7.18 | 7.32 | 6.43 |
| ereb126 | GRMZM2G169654 | 5.36 | −6.64 | 6.74 |
| glk34 | GRMZM2G081671 | 7.18 | 8.32 | 8.59 |
| gras11 | GRMZM2G097456 | −6.64 | 5.32 | 7.01 |
| gras13 | GRMZM2G140094 | −6.64 | 6.64 | 6.43 |
| gras14 | GRMZM2G070371 | −6.64 | 5.91 | 7.42 |
| gras38 | GRMZM2G098517 | 7.18 | 6.32 | 6.00 |
| gras63 | GRMZM2G418899 | 6.69 | 5.91 | 6.43 |
| hb102 | GRMZM2G139963 | 5.36 | −6.64 | 5.43 |
| hb8 | GRMZM2G135447 | −6.64 | −6.64 | 7.23 |
| jmj5 | GRMZM2G070885 | −6.64 | 6.91 | −6.64 |
| K-box | GRMZM2G133568 | 7.69 | −6.64 | 6.43 |
| K-box | GRMZM2G052123 | 6.95 | 5.32 | 5.43 |
| K-box | GRMZM2G033093 | 6.95 | 5.32 | 7.23 |
| K-box | GRMZM2G003514 | −6.64 | −6.64 | −6.64 |
| K-box | GRMZM2G148693 | −6.64 | −2.01 | −6.64 |
| K-box | GRMZM2G137510 | −6.64 | −5.13 | −6.64 |
| K-box | GRMZM2G147716 | −2.44 | −1.16 | −6.64 |
| K-box | GRMZM2G046885 | −6.64 | 0.14 | −6.64 |
| K-box | GRMZM2G069370 | −6.64 | −1.85 | −6.64 |
| knox1 | GRMZM2G159431 | 6.95 | 5.32 | 7.42 |
| mads41 | GRMZM2G018589 | 8.28 | 6.64 | 6.74 |
| obf4 | GRMZM2G125243 | 9.62 | 9.13 | 8.81 |
| Pathogenesis | GRMZM2G056729 | 6.38 | 6.91 | −6.64 |
| TFIIF | GRMZM2G060284 | 9.37 | 7.78 | 8.67 |
| thx23 | GRMZM2G156348 | −6.64 | 6.32 | 8.01 |
| tsh4 | GRMZM2G307588 | 7.18 | −6.64 | 6.00 |
| Pathway | Gene ID | Arabidopsis Homologues | CK (TPM) | Stage I (TPM) | Stage II (TPM) | Stage III (TPM) |
|---|---|---|---|---|---|---|
| DETs annotated in plant basal transcription factors | GRMZM2G017831 | AT4G36650 | 9.52 | 1.86 | 11.2 | 12.22 |
| GRMZM2G091586 | AT2G41630, AT3G10330 | 18.03 | 42.84 | 25.59 | 31.72 | |
| GRMZM2G045668 | AT1G03280, AT4G20340, AT4G20810 | 2.23 | 0.41 | 1.2 | 1.29 | |
| GRMZM2G064390 | AT1G03280, AT4G20340, AT4G20810 | 36.06 | 72.43 | 69.78 | 82.73 | |
| GRMZM2G114584 | AT1G17070, AT2G42330 | 2.84 | 11.38 | 8.2 | 4.07 | |
| GRMZM2G060284 | AT1G75510, AT3G52270 | 0 | 6.62 | 2.2 | 4.07 | |
| GRMZM2G049091 | AT1G75510, AT3G52270 | 32.62 | 75.33 | 33.99 | 42.44 | |
| GRMZM2G125259 | AT1G312240 | 7.7 | 2.07 | 2.4 | 0.43 | |
| GRMZM2G151717 | AT3G02160, AT4G34340, AT5G15570 | 1.82 | 1.86 | 0 | 0.64 | |
| GRMZM2G173309 | AT1G05055 | 1.01 | 7.45 | 6.2 | 4.93 | |
| GRMZM2G161418 | NA | 1.01 | 0 | 1.8 | 1.29 | |
| GRMZM2G110076 | AT1G54140 | 3.04 | 3.31 | 7.8 | 4.07 | |
| DETs annotated in plant hormone signal transduction | GRMZM2G133331 | AT1G68640 | 3.04 | 0 | 1.6 | 0 |
| GRMZM2G381991 | NA | 0 | 0.83 | 0 | 3.22 | |
| GRMZM2G030877 | AT1G08320 | 3.65 | 8.28 | 5 | 3.22 | |
| GRMZM2G132868 | NA | 4.05 | 13.24 | 2.8 | 3.86 | |
| GRMZM2G125243 | AT3G12250, AT5G06950, AT5G06960 | 0 | 7.86 | 5.6 | 4.5 | |
| GRMZM2G002075 | AT1G45249, AT1G49720, AT3G19290, AT4G34000, AT5G42910 | 5.06 | 6 | 11.4 | 7.72 | |
| GRMZM2G159134 | AT2G41070, AT3G56850 | 2.84 | 2.07 | 5.2 | 10.29 | |
| GRMZM2G161009 | AT2G41070, AT3G56850 | 5.67 | 16.97 | 12.4 | 12 | |
| GRMZM2G060290 | AT5G06839 | 3.85 | 36.63 | 8.8 | 21.01 | |
| GRMZM2G361847 | AT3G12250, AT5G06950, AT5G06960 | 61.59 | 81.75 | 117.76 | 147.25 | |
| GRMZM2G094352 | AT1G22070, AT1G77920, AT5G10030, AT5G65210 | 98.05 | 168.04 | 115.16 | 215.19 | |
| GRMZM2G131961 | AT1G22070, AT1G77920, AT5G10030, AT5G65210 | 113.05 | 95.4 | 77.37 | 237.7 | |
| GRMZM2G024973 | AT1G14920, AT1G66350, AT2G01570, AT3G03450, AT5G17490 | 160.05 | 63.74 | 42.79 | 55.08 | |
| GRMZM2G330012 | AT3G20220 | 1.82 | 1.03 | 4 | 2.79 | |
| GRMZM2G049229 | AT1G32640, AT4G17880, AT5G46760, AT5G46830 | 134.52 | 125.2 | 56.38 | 117.67 | |
| GRMZM2G001930 | AT1G32640, AT4G17880, AT5G46760, AT5G46830 | 528.97 | 287.87 | 149.55 | 230.84 | |
| GRMZM2G030280 | AT3G12250, AT5G06950, AT5G06960 | 13.98 | 31.46 | 11.4 | 23.15 | |
| DETs annotated in plant base excision repair | GRMZM2G012654 | AT2G27470 | 7.09 | 14.49 | 12.4 | 10.72 |
| GRMZM2G139031 | AT1G21710 | 2.03 | 1.24 | 5.8 | 2.36 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, Y.; Ge, F.; Hou, F.; Sun, W.; Zheng, Q.; Zhang, X.; Ma, L.; Fu, J.; He, X.; Peng, H.; et al. Transcription Factors Responding to Pb Stress in Maize. Genes 2017, 8, 231. https://doi.org/10.3390/genes8090231
Zhang Y, Ge F, Hou F, Sun W, Zheng Q, Zhang X, Ma L, Fu J, He X, Peng H, et al. Transcription Factors Responding to Pb Stress in Maize. Genes. 2017; 8(9):231. https://doi.org/10.3390/genes8090231
Chicago/Turabian StyleZhang, Yanling, Fei Ge, Fengxia Hou, Wenting Sun, Qi Zheng, Xiaoxiang Zhang, Langlang Ma, Jun Fu, Xiujing He, Huanwei Peng, and et al. 2017. "Transcription Factors Responding to Pb Stress in Maize" Genes 8, no. 9: 231. https://doi.org/10.3390/genes8090231
APA StyleZhang, Y., Ge, F., Hou, F., Sun, W., Zheng, Q., Zhang, X., Ma, L., Fu, J., He, X., Peng, H., Pan, G., & Shen, Y. (2017). Transcription Factors Responding to Pb Stress in Maize. Genes, 8(9), 231. https://doi.org/10.3390/genes8090231




