Single Nucleotide Polymorphisms of NUCB2 and Their Genetic Associations with Milk Production Traits in Dairy Cows
Abstract
1. Introduction
2. Materials and Methods
2.1. Animals
2.2. Phenotypic Data Collection
2.3. DNA Extraction
2.4. SNP Identification and Genotyping
2.5. Estimation of Linkage Disequilibrium (LD) and Association Analyses
2.6. Transcription Factor Binding Sites (TFBSs) Prediction
2.7. NUCB2 Gene Expression Analysis
3. Results
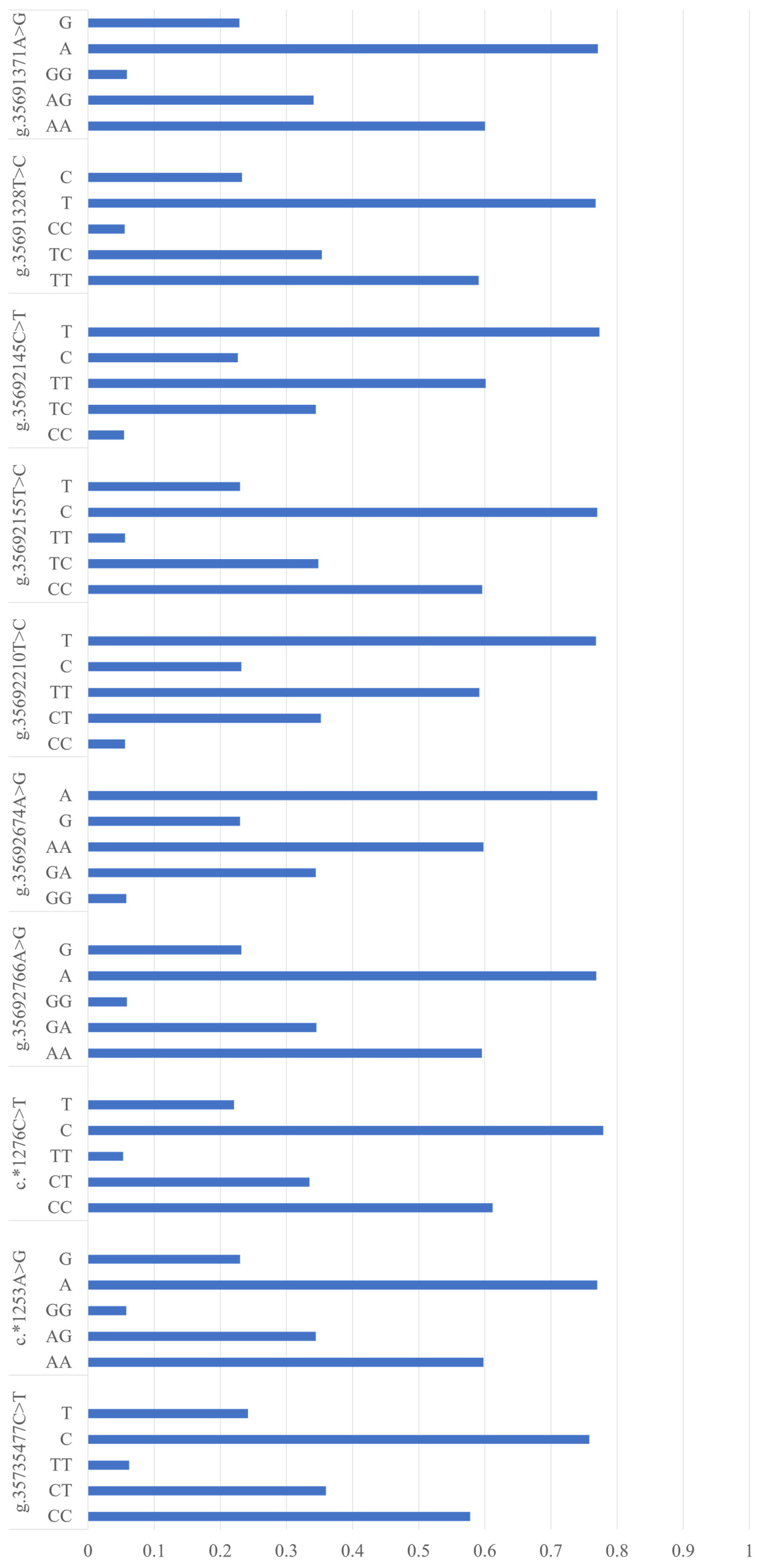
3.1. SNP Identification in NUCB2 Gene
3.2. Associations between Ten SNPs and Five Milk Production Traits
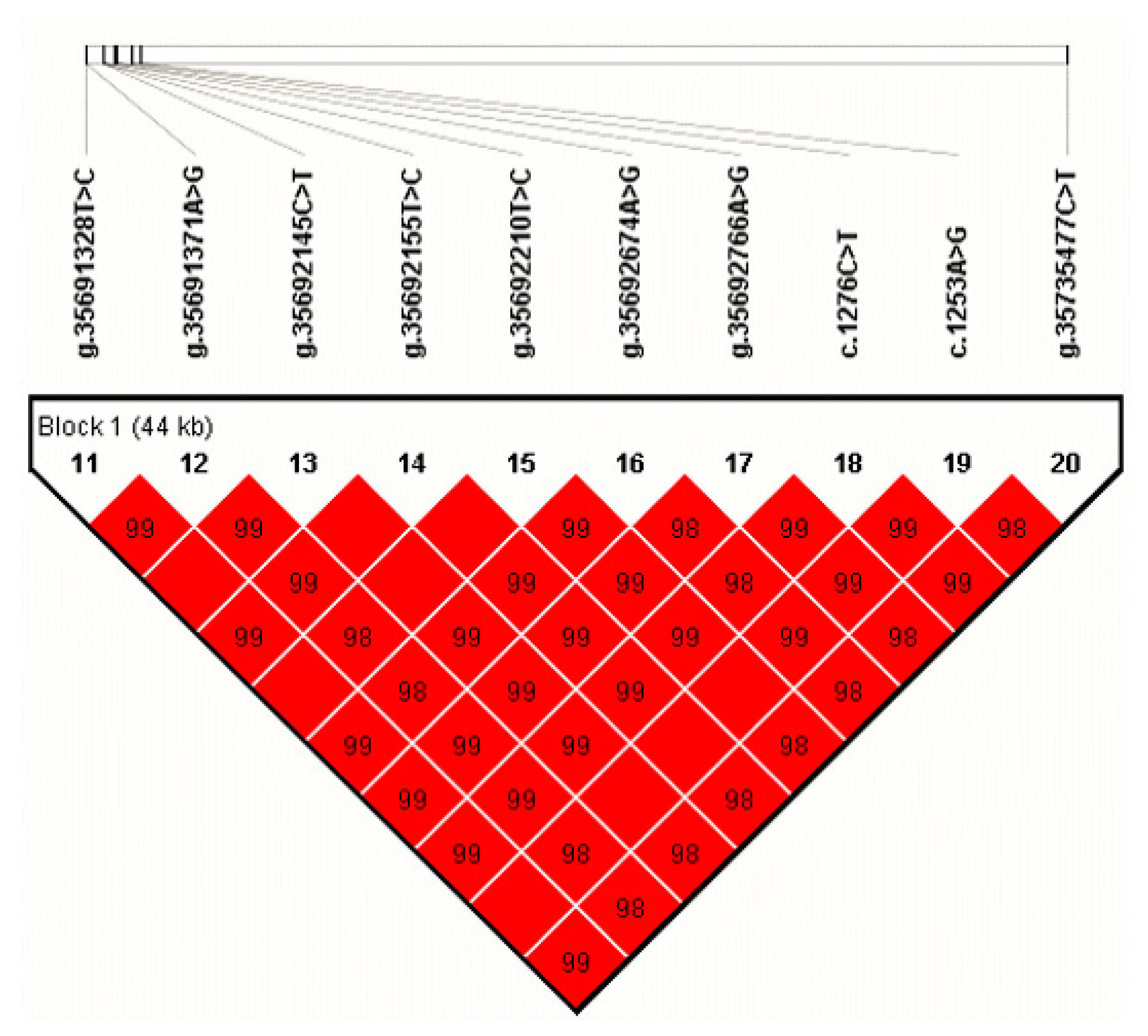
3.3. Associations between Haplotype Combinations with Five Milk Traits
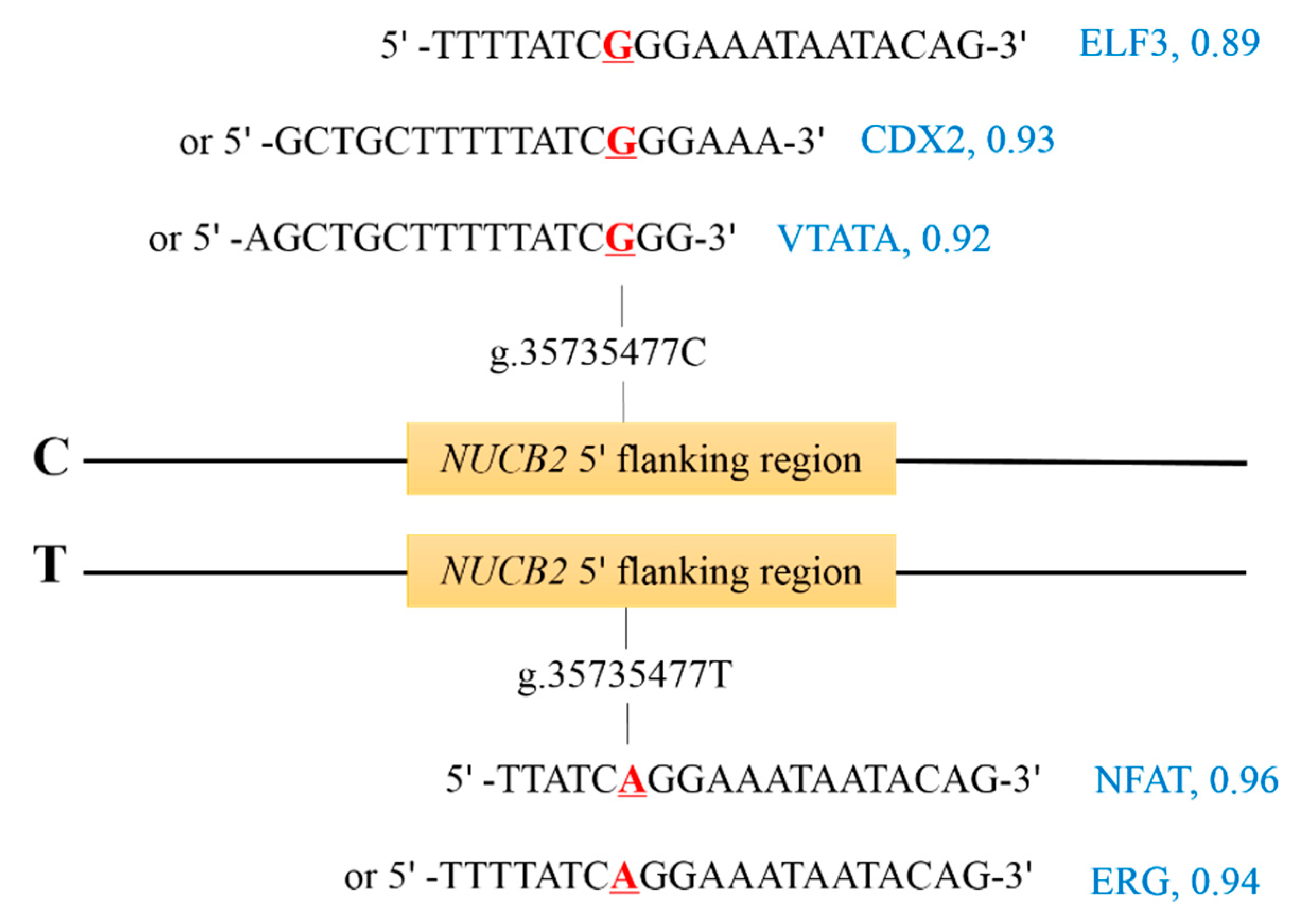
3.4. TFBSs Changed by g.35735477C>T of NUCB2
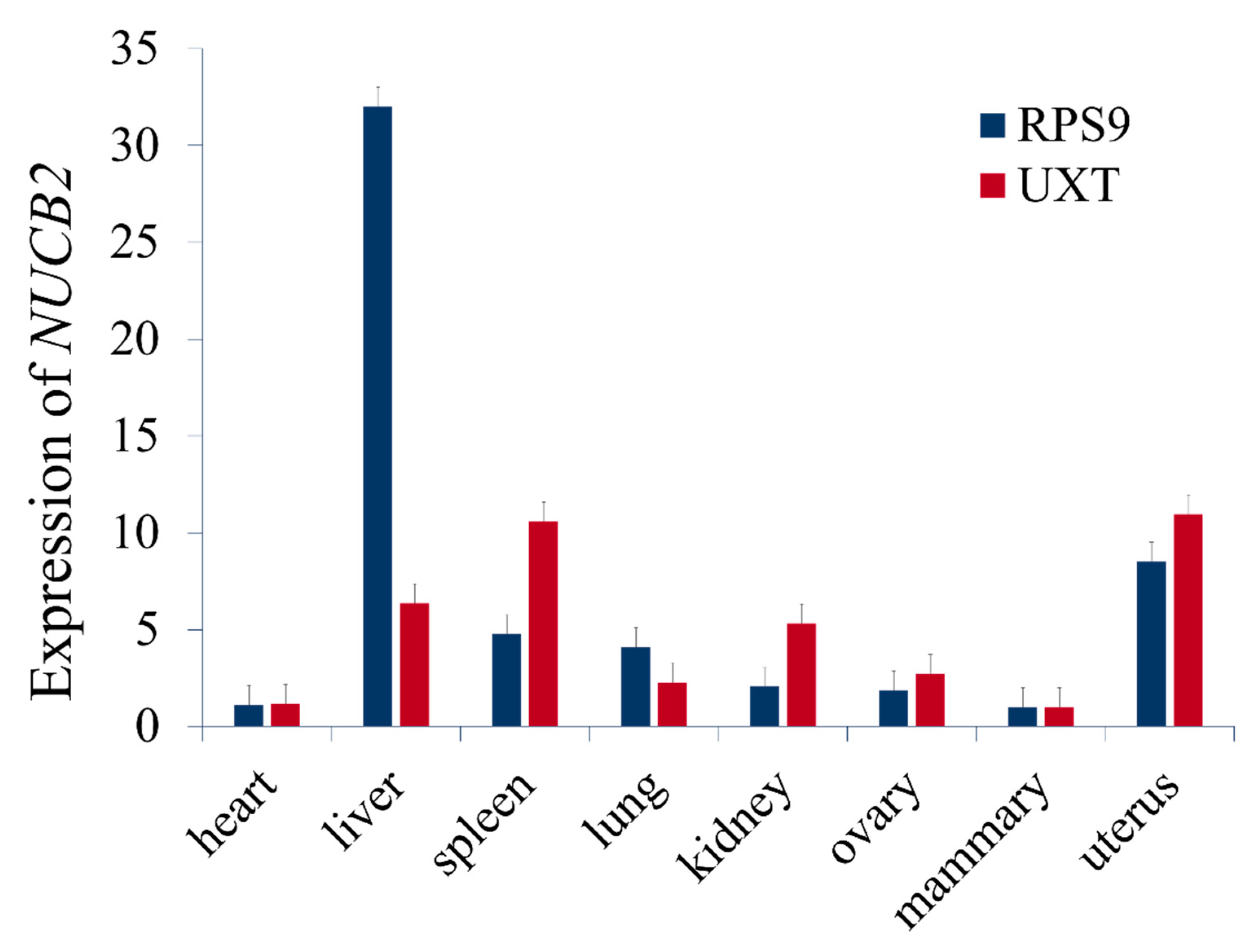
3.5. Expression of NUCB2 in Tissues
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tunick, M.H.; Van Hekken, D.L. Dairy Products and Health: Recent Insights. J. Agric. Food Chem. 2015, 63, 9381–9388. [Google Scholar] [CrossRef]
- Spelman, R.J.; Coppieters, W.; Karim, L.; van Arendonk, J.A.; Bovenhuis, H. Quantitative trait loci analysis for five milk production traits on chromosome six in the Dutch Holstein-Friesian population. Genetics 1996, 144, 1799–1808. [Google Scholar]
- Denoeud, F.; Aury, J.M.; Da Silva, C.; Noel, B.; Rogier, O.; Delledonne, M.; Morgante, M.; Valle, G.; Wincker, P.; Scarpelli, C.; et al. Annotating genomes with massive-scale RNA sequencing. Genome Biol. 2008, 9, R175. [Google Scholar] [CrossRef]
- Liang, R.; Han, B.; Li, Q.; Yuan, Y.; Li, J.; Sun, D. Using RNA sequencing to identify putative competing endogenous RNAs (ceRNAs) potentially regulating fat metabolism in bovine liver. Sci. Rep. 2017, 7, 6396. [Google Scholar] [CrossRef]
- Oh, S.; Shimizu, H.; Satoh, T.; Okada, S.; Adachi, S.; Inoue, K.; Eguchi, H.; Yamamoto, M.; Imaki, T.; Hashimoto, K.; et al. Identification of nesfatin-1 as a satiety molecule in the hypothalamus. Nature 2006, 443, 709–712. [Google Scholar] [CrossRef]
- Stengel, A.; Tache, Y. Minireview: Nesfatin-1-An Emerging New Player in the Brain-Gut, Endocrine, and Metabolic Axis. Endocrinology 2011, 152, 4033–4038. [Google Scholar] [CrossRef]
- Ramesh, N.; Gawli, K.; Pasupulleti, V.K.; Unniappan, S. Metabolic and Cardiovascular Actions of Nesfatin-1: Implications in Health and Disease. Curr. Pharm. Des. 2017, 23, 1453–1464. [Google Scholar] [CrossRef]
- Mohan, H.; Unniappan, S. Phylogenetic aspects of nucleobindin-2/nesfatin-1. Curr. Pharm. Des. 2013, 19, 6929–6934. [Google Scholar] [CrossRef]
- Sun, S.; Yang, H. Tissue-Specific Localization NUCB2/nesfatin-1 in the Liver and Heart of Mouse Fetus. Dev. Reprod. 2018, 22, 331–339. [Google Scholar] [CrossRef]
- Suzuki, S.; Takagi, K.; Miki, Y.; Onodera, Y.; Akahira, J.; Ebata, A.; Ishida, T.; Watanabe, M.; Sasano, H.; Suzuki, T. Nucleobindin 2 in human breast carcinoma as a potent prognostic factor. Cancer Sci. 2012, 103, 136–143. [Google Scholar] [CrossRef]
- Boutinaud, M.; Galio, L.; Lollivier, V.; Finot, L.; Wiart, S.; Esquerre, D.; Devinoy, E. Unilateral once daily milking locally induces differential gene expression in both mammary tissue and milk epithelial cells revealing mammary remodeling. Physiol. Genom. 2013, 45, 973–985. [Google Scholar] [CrossRef]
- Schopen, G.C.; Koks, P.D.; van Arendonk, J.A.; Bovenhuis, H.; Visker, M.H. Whole genome scan to detect quantitative trait loci for bovine milk protein composition. Anim. Genet. 2009, 40, 524–537. [Google Scholar] [CrossRef]
- Daetwyler, H.D.; Schenkel, F.S.; Sargolzaei, M.; Robinson, J.A.B. A genome scan to detect quantitative trait loci for economically important traits in Holstein cattle using two methods and a dense single nucleotide polymorphism map. J. Dairy Sci. 2008, 91, 3225–3236. [Google Scholar] [CrossRef]
- Cole, J.B.; Wiggans, G.R.; Ma, L.; Sonstegard, T.S.; Lawlor, T.J., Jr.; Crooker, B.A.; Van Tassell, C.P.; Yang, J.; Wang, S.; Matukumalli, L.K.; et al. Genome-wide association analysis of thirty one production, health, reproduction and body conformation traits in contemporary U.S. Holstein cows. BMC Genom. 2011, 12, 408. [Google Scholar] [CrossRef]
- Hagenblad, J.; Tang, C.; Molitor, J.; Werner, J.; Zhao, K.; Zheng, H.; Marjoram, P.; Weigel, D.; Nordborg, M. Haplotype structure and phenotypic associations in the chromosomal regions surrounding two Arabidopsis thaliana flowering time loci. Genetics 2004, 168, 1627–1638. [Google Scholar] [CrossRef][Green Version]
- Nothnagel, M.; Rohde, K. The effect of single-nucleotide polymorphism marker selection on patterns of haplotype blocks and haplotype frequency estimates. Am. J. Hum. Genet. 2005, 77, 988–998. [Google Scholar] [CrossRef]
- Patil, N.; Berno, A.J.; Hinds, D.A.; Barrett, W.A.; Doshi, J.M.; Hacker, C.R.; Kautzer, C.R.; Lee, D.H.; Marjoribanks, C.; McDonough, D.P.; et al. Blocks of limited haplotype diversity revealed by high-resolution scanning of human chromosome 21. Science 2001, 294, 1719–1723. [Google Scholar] [CrossRef]
- Wiggans, G.R.; Cole, J.B.; Hubbard, S.M.; Sonstegard, T.S. Genomic Selection in Dairy Cattle: The USDA Experience. Annu. Rev. Anim. Biosci. 2017, 5, 309–327. [Google Scholar] [CrossRef]
- Zhang, Z.; Ober, U.; Erbe, M.; Zhang, H.; Gao, N.; He, J.; Li, J.; Simianer, H. Improving the accuracy of whole genome prediction for complex traits using the results of genome wide association studies. PLoS ONE 2014, 9, e93017. [Google Scholar] [CrossRef]
- Bouwman, A.C.; Bovenhuis, H.; Visker, M.H.; van Arendonk, J.A. Genome-wide association of milk fatty acids in Dutch dairy cattle. BMC Genet. 2011, 12, 43. [Google Scholar] [CrossRef]
- Matsumoto, H.; Sasaki, K.; Bessho, T.; Kobayashi, E.; Abe, T.; Sasazaki, S.; Oyama, K.; Mannen, H. The SNPs in the ACACA gene are effective on fatty acid composition in Holstein milk. Mol. Biol. Rep. 2012, 39, 8637–8644. [Google Scholar] [CrossRef]
- Bhattarai, D.; Chen, X.; Ur Rehman, Z.; Hao, X.; Ullah, F.; Dad, R.; Talpur, H.S.; Kadariya, I.; Cui, L.; Fan, M.; et al. Association of MAP4K4 gene single nucleotide polymorphism with mastitis and milk traits in Chinese Holstein cattle. J. Dairy. Res. 2017, 84, 76–79. [Google Scholar] [CrossRef]
- Han, B.; Liang, W.; Liu, L.; Li, Y.; Sun, D. Determination of genetic effects of ATF3 and CDKN1A genes on milk yield and compositions in Chinese Holstein population. BMC Genet. 2017, 18, 47. [Google Scholar] [CrossRef]
- Han, B.; Liang, W.; Liu, L.; Li, Y.; Sun, D. Genetic association of the ACACB gene with milk yield and composition traits in dairy cattle. Anim. Genet. 2018. [Google Scholar] [CrossRef]
- Han, B.; Yuan, Y.; Liang, R.; Li, Y.; Liu, L.; Sun, D. Genetic Effects of LPIN1 Polymorphisms on Milk Production Traits in Dairy Cattle. Genes (Basel) 2019, 10, 265. [Google Scholar] [CrossRef]
- Bell, A.W. Regulation of organic nutrient metabolism during transition from late pregnancy to early lactation. J. Anim. Sci. 1995, 73, 2804–2819. [Google Scholar] [CrossRef]
- Rawson, P.; Stockum, C.; Peng, L.F.; Manivannan, B.; Lehnert, K.; Ward, H.E.; Berry, S.D.; Davis, S.R.; Snell, R.G.; McLauchlan, D.; et al. Metabolic proteomics of the liver and mammary gland during lactation. J. Proteom. 2012, 75, 4429–4435. [Google Scholar] [CrossRef]
- Moyes, K.M.; Bendixen, E.; Codrea, M.C.; Ingvartsen, K.L. Identification of hepatic biomarkers for physiological imbalance of dairy cows in early and mid lactation using proteomic technology. J. Dairy Sci. 2013, 96, 3599–3610. [Google Scholar] [CrossRef]
- Tugrul, M.; Paixao, T.; Barton, N.H.; Tkacik, G. Dynamics of Transcription Factor Binding Site Evolution. PLoS Genet. 2015, 11, e1005639. [Google Scholar] [CrossRef]
- Talebzadeh, M.; Zare-Mirakabad, F. Transcription factor binding sites prediction based on modified nucleosomes. PLoS ONE 2014, 9, e89226. [Google Scholar] [CrossRef]
- Oettgen, P.; Alani, R.M.; Barcinski, M.A.; Brown, L.; Akbarali, Y.; Boltax, J.; Kunsch, C.; Munger, K.; Libermann, T.A. Isolation and characterization of a novel epithelium-specific transcription factor, ESE-1, a member of the ETS family. Mol. Cell Biol. 1997, 17, 4419–4433. [Google Scholar] [CrossRef]
- Yoshida, N.; Yoshida, S.; Araie, M.; Handa, H.; Nabeshima, Y. Ets family transcription factor ESE-1 is expressed in corneal epithelial cells and is involved in their differentiation. Mech. Deve. 2000, 97, 27–34. [Google Scholar] [CrossRef]
- Zheng, L.; Xu, M.; Xu, J.; Wu, K.; Fang, Q.; Liang, Y.; Zhou, S.; Cen, D.; Ji, L.; Han, W.; et al. ELF3 promotes epithelial-mesenchymal transition by protecting ZEB1 from miR-141-3p-mediated silencing in hepatocellular carcinoma. Cell Death Dis. 2018, 9, 387. [Google Scholar] [CrossRef]
- Gajulapalli, V.N.R.; Samanthapudi, V.S.K.; Pulaganti, M.; Khumukcham, S.S.; Malisetty, V.L.; Guruprasad, L.; Chitta, S.K.; Manavathi, B. A transcriptional repressive role for epithelial-specific ETS factor ELF3 on oestrogen receptor α in breast cancer cells. Biochem. J. 2016, 473, 1047–1061. [Google Scholar] [CrossRef]
- Zhu, R.; Wong, K.F.; Lee, N.P.; Lee, K.F.; Luk, J.M. HNF1alpha and CDX2 transcriptional factors bind to cadherin-17 (CDH17) gene promoter and modulate its expression in hepatocellular carcinoma. J. Cell Biochem. 2010, 111, 618–626. [Google Scholar] [CrossRef]
- Yu, J.H.; Liu, D.; Sun, X.J.; Yang, K.; Yao, J.F.; Cheng, C.; Wang, C.B.; Zheng, J.B. CDX2 inhibits the proliferation and tumor formation of colon cancer cells by suppressing Wnt/β-catenin signaling via transactivation of GSK-3 β and Axin2 expression. Cell Death Dis. 2019, 10, 26. [Google Scholar] [CrossRef]
- Kumar, N.; Tsai, Y.H.; Chen, L.; Zhou, A.; Banerjee, K.K.; Saxena, M.; Huang, S.; Toke, N.H.; Xing, J.; Shivdasani, R.A.; et al. The lineage-specific transcription factor CDX2 navigates dynamic chromatin to control distinct stages of intestine development. Development 2019, 146. [Google Scholar] [CrossRef]
- Ross, D.G.F.; Bowles, J.; Koopman, P.; Lehnert, S. New insights into SRY regulation through identification of 5 conserved sequences. BMC Mol. Biol. 2008, 9. [Google Scholar] [CrossRef]
- Shaw, J.P.; Utz, P.J.; Durand, D.B.; Toole, J.J.; Emmel, E.A.; Crabtree, G.R. Identification of a putative regulator of early T cell activation genes. Science 1988, 241, 202–205. [Google Scholar] [CrossRef]
- Mognol, G.P.; Carneiro, F.R.; Robbs, B.K.; Faget, D.V.; Viola, J.P. Cell cycle and apoptosis regulation by NFAT transcription factors: New roles for an old player. Cell Death Dis. 2016, 7, e2199. [Google Scholar] [CrossRef]
- Dufton, N.P.; Peghaire, C.R.; Osuna-Almagro, L.; Raimondi, C.; Kalna, V.; Chuahan, A.; Webb, G.; Yang, Y.W.; Birdsey, G.M.; Lalor, P.; et al. Dynamic regulation of canonical TGF β signalling by endothelial transcription factor ERG protects from liver fibrogenesis. Nat. Commun. 2017, 8. [Google Scholar] [CrossRef]
- Kalna, V.; Yang, Y.; Peghaire, C.; Frudd, K.; Hannah, R.; Shah, A.V.; Osuna Almagro, L.; Boyle, J.J.; Gottgens, B.; Ferrer, J.; et al. The Transcription Factor ERG Regulates Super-Enhancers Associated with an Endothelial-Specific Gene Expression Program. Circ. Res. 2019. [Google Scholar] [CrossRef]
| SNP Name | GenBank No. | Location | Position (UMD3.1) | Mutation |
|---|---|---|---|---|
| g.35735477C>T | rs135882628 | 5′flanking region | chr15:35735477 | C/T |
| c.*1253A>G | rs207689152 | 3′UTR | chr15:35693831 | A/G |
| c.*1276C>T | rs208861167 | 3′UTR | chr15:35693399 | C/T |
| g.35692766A>G | rs109930800 | 3′flanking region | chr15:35692766 | A/G |
| g.35692674A>G | rs109930800 | 3′flanking region | chr15:35692674 | A/G |
| g.35692210T>C | rs208420141 | 3′flanking region | chr15:35692210 | T/C |
| g.35692155T>C | rs210564196 | 3′flanking region | chr15:35692155 | T/C |
| g.35692145C>T | rs209421884 | 3′flanking region | chr15:35692145 | C/T |
| g.35691328T>C | rs210746263 | 3′flanking region | chr15:35691328 | T/C |
| g.35691371A>G | rs207595426 | 3′flanking region | chr15:35691371 | A/G |
| SNP | Lactation | Genotype (No.) | Milk Yield (kg) | Fat Yield (kg) | Fat Percentage (%) | Protein Yield (kg) | Protein Percentage (%) | No. of Significant SNPs |
|---|---|---|---|---|---|---|---|---|
| g.35735477C>T | 1 | CC (604) | 10326 ± 63.16 | 343.09 ± 2.81 | 3.33 ± 0.03 Aa | 306 ± 2.05 | 2.96 ± 0.01 a | 2 |
| CT (376) | 10309 ± 68.49 | 340.27 ± 3.02 | 3.31 ± 0.03 ab | 305.52 ± 2.2 | 2.96 ± 0.01 a | |||
| TT (65) | 10456 ± 111.13 | 338.57 ± 4.65 | 3.21 ± 0.05 Bb | 306.19 ± 3.39 | 2.92 ± 0.02 b | |||
| p value | 0.3565 | 0.285 | 0.0122 | 0.9428 | 0.0327 | |||
| 2 | CC (415) | 10617 ± 67.14 Aa | 385.18 ± 2.99 | 3.65 ± 0.03 a | 315.66 ± 2.18 b | 2.99 ± 0.01 Aa | 4 | |
| CT (258) | 10871 ± 72.95 B | 387.94 ± 3.21 | 3.59 ± 0.03 b | 320.54 ± 2.33 a | 2.96 ± 0.01 Bb | |||
| TT (49) | 10475 ± 126.69 Aa | 383.05 ± 5.25 | 3.67 ± 0.05 ab | 309.98 ± 3.83b | 2.97 ± 0.02 ab | |||
| p value | <0.0001 | 0.4542 | 0.0257 | 0.0036 | 0.0107 | |||
| c.*1253A>G | 1 | AA (622) | 10377 ± 62.96 | 346.91 ± 2.81 a | 3.35 ± 0.03 a | 308.01 ± 2.05 | 2.96 ± 0.01 a | 3 |
| AG (358) | 10330 ± 67.85 | 343.52 ± 2.99 ab | 3.33 ± 0.03 ab | 306.64 ± 2.17 | 2.96 ± 0.01 a | |||
| GG (60) | 10411 ± 113.36 | 335.92 ± 4.73 b | 3.23 ± 0.05 b | 304.51 ± 3.44 | 2.92 ± 0.02 b | |||
| p value | 0.5823 | 0.0178 | 0.0167 | 0.4162 | 0.0334 | |||
| 2 | AA (434) | 10739 ± 65.59 Aa | 389.07 ± 3.02 a | 3.63 ± 0.03 | 319.03 ± 2.12 a | 2.98 ± 0.01 | 3 | |
| AG (248) | 10934 ± 73.57 B | 396.99 ± 3.26 b | 3.6 ± 0.03 | 322.97 ± 2.34 Aa | 2.96 ± 0.01 | |||
| GG (45) | 10468 ± 130.21 Aa | 386.53 ± 5.54 ab | 3.66 ± 0.05 | 308.67 ± 3.93 Bb | 2.96 ± 0.02 | |||
| p value | 0.0002 | 0.0073 | 0.3202 | 0.0006 | 0.089 | |||
| c.*1276C>T | 1 | CC (632) | 10332 ± 62.27 | 346.1 ± 2.78 | 3.36 ± 0.03 a | 305.38 ± 2.03 | 2.96 ± 0.01 a | 3 |
| CT (346) | 10320 ± 68.73 | 343.57 ± 3.02 | 3.34 ± 0.03 ab | 305.67 ± 2.2 | 2.96 ± 0.01 a | |||
| TT (55) | 10392 ± 113.98 | 336.79 ± 4.74 | 3.25 ± 0.05 b | 302.85 ± 3.46 | 2.92 ± 0.02 b | |||
| p value | 0.7939 | 0.0661 | 0.0278 | 0.6665 | 0.0217 | |||
| 2 | CC (442) | 10687 ± 65.62 a | 386.11 ± 2.93 | 3.63 ± 0.03 | 316.61 ± 2.13 ab | 2.98 ± 0.01 a | 3 | |
| CT (238) | 10850 ± 74 b | 388.08 ± 3.23 | 3.59 ± 0.03 | 319.51 ± 2.36 a | 2.96 ± 0.01 b | |||
| TT (42) | 10521 ± 130.66 a | 384.93±5.4 | 3.67 ± 0.05 | 309.97 ± 3.94 b | 2.95 ± 0.02 ab | |||
| p value | 0.0089 | 0.7061 | 0.1478 | 0.0347 | 0.0357 | |||
| g.35692766A>G | 1 | AA (626) | 10418 ± 63.52 | 347.12±2.84 | 3.35 ± 0.03a | 307.67 ± 2.07 | 2.96 ± 0.01 | 1 |
| GA (363) | 10345 ± 68.1 | 343.28±3 | 3.33 ± 0.03ab | 305.79 ± 2.19 | 2.96 ± 0.01 | |||
| GG (62) | 10510 ± 112.05 | 339.43±4.68 | 3.24 ± 0.05b | 306.98 ± 3.41 | 2.92 ± 0.02 | |||
| p value | 0.1772 | 0.0623 | 0.0382 | 0.4738 | 0.0638 | |||
| 2 | AA (434) | 10713 ± 65.44 Aa | 386.64±2.91 | 3.63 ± 0.03 | 318.28 ± 2.12 b | 2.98 ± 0.01 a | 4 | |
| GA (249) | 10930 ± 72.31 B | 392.91±3.16 | 3.61 ± 0.03 | 323.49 ± 2.3 Aa | 2.96 ± 0.01 ab | |||
| GG (46) | 10498 ± 130.83 Aa | 381.61±5.43 | 3.66 ± 0.05 | 308.07 ± 3.96 Ba | 2.94 ± 0.02 b | |||
| p value | 0.0002 | 0.018 | 0.4651 | <0.0001 | 0.0467 | |||
| g.35692674A>G | 1 | CC (60) | 10345 ± 113.56 | 332.97±4.74Aa | 3.23 ± 0.05a | 301.01 ± 3.45 | 2.92 ± 0.02 a | 3 |
| CT (358) | 10330 ± 67.9 | 343.02±2.99ab | 3.33 ± 0.03ab | 305.25 ± 2.18 | 2.96 ± 0.01 b | |||
| TT (622) | 10348 ± 62.26 | 345.78±2.78Bb | 3.36 ± 0.03b | 305.7 ± 2.02 | 2.96 ± 0.01 b | |||
| p value | 0.9433 | 0.0081 | 0.0116 | 0.3135 | 0.0273 | |||
| 2 | CC (45) | 10599 ± 129.91 b | 389.01±5.38ab | 3.68 ± 0.05 | 313.59 ± 3.92 Aa | 2.96 ± 0.02 | 3 | |
| CT (248) | 10973 ± 73.35 Aa | 395.5±3.21a | 3.61 ± 0.03 | 324.92 ± 2.34 B | 2.96 ± 0.01 | |||
| TT (434) | 10733 ± 65.85 Bb | 388.86±2.93b | 3.63 ± 0.03 | 318.92 ± 2.13 Aa | 2.98 ± 0.01 | |||
| p value | 0.0002 | 0.0383 | 0.3137 | 0.0009 | 0.1688 | |||
| g.35692210T>C | 1 | CC (59) | 10441 ± 113.16 | 337.01 ± 4.71 | 3.24 ± 0.05 | 305.33 ± 3.43 | 2.93 ± 0.02 | 0 |
| CT (372) | 10325 ± 68.18 | 340.45 ± 3.01 | 3.31 ± 0.03 | 305.34 ± 2.19 | 2.96 ± 0.01 | |||
| TT (625) | 10373 ± 63.73 | 344.35 ± 2.85 | 3.34 ± 0.03 | 306.62 ± 2.08 | 2.96 ± 0.01 | |||
| p value | 0.4509 | 0.0683 | 0.0533 | 0.6822 | 0.0824 | |||
| 2 | CC (44) | 10443 ± 132.87 Aa | 377.54 ± 5.51 a | 3.64 ± 0.05 | 307.55 ± 4.02 Aa | 2.95 ± 0.02 ab | 4 | |
| CT (254) | 10916 ± 73.07 B | 388.6 ± 3.21 b | 3.58 ± 0.03 | 321.67 ± 2.33 Bb | 2.96 ± 0.01 a | |||
| TT (434) | 10688 ± 66.14 Aa | 383.07 ± 2.95 a | 3.61 ± 0.03 | 316.51 ± 2.15 a | 2.98 ± 0.01b | |||
| p value | <0.0001 | 0.0331 | 0.3355 | 0.0003 | 0.0143 | |||
| g.35692155T>C | 1 | CC (620) | 10297 ± 63.03 | 344.52 ± 2.83 Aa | 3.35 ± 0.03 a | 304.35 ± 2.05 | 2.96 ± 0.01 a | 3 |
| TC (362) | 10284 ± 67.63 | 341.51 ± 3 a | 3.32 ± 0.03 ab | 304.36 ± 2.17 | 2.96 ± 0.01 a | |||
| TT (58) | 10378 ± 114.2 | 328.5 ± 4.92 Bb | 3.24 ± 0.05 b | 302.2 ± 3.46 | 2.92 ± 0.02 B | |||
| p value | 0.6782 | 0.0012 | 0.0144 | 0.7741 | 0.0138 | |||
| 2 | CC (432) | 10690 ± 65.94 Aa | 386.06 ± 2.94 | 3.63 ± 0.03 | 317.22 ± 2.14 ab | 2.98 ± 0.01 | 2 | |
| TC (249) | 10889 ± 73.19 B | 390.76 ± 3.2 | 3.6 ± 0.03 | 321.46 ± 2.33 Aa | 2.96 ± 0.01 | |||
| TT (44) | 10474 ± 131.19 Aa | 382.27 ± 5.43 | 3.66 ± 0.05 | 308.69 ± 3.96 Bb | 2.95 ± 0.02 | |||
| p value | 0.0006 | 0.109 | 0.3315 | 0.0018 | 0.0989 | |||
| g.35692145C>T | 1 | CC (57) | 10447 ± 113.95 | 338.63 ± 4.74 | 3.26 ± 0.05 a | 303.9 ± 3.45 | 2.91 ± 0.02 a | 2 |
| TC (359) | 10342 ± 68.09 | 343.23 ± 3 | 3.33 ± 0.03 ab | 305.98 ± 2.18 | 2.96 ± 0.01 b | |||
| TT (627) | 10366 ± 63.35 | 346.42 ± 2.83 | 3.36 ± 0.03 b | 306.41 ± 2.06 | 2.96 ± 0.01 b | |||
| p value | 0.6061 | 0.0937 | 0.0388 | 0.7116 | 0.0133 | |||
| 2 | CC (43) | 10461 ± 131.5 Aa | 383.01 ± 5.44ab | 3.67 ± 0.05 | 309.35 ± 3.97 Aa | 2.95 ± 0.02 ab | 3 | |
| TC (247) | 10948 ± 72.69 B | 392.58 ± 3.18a | 3.59 ± 0.03 | 323.82 ± 2.31 Bb | 2.96 ± 0.01 a | |||
| TT (439) | 10692 ± 65.5 Aa | 386.85 ± 2.92b | 3.64 ± 0.03 | 318.16 ± 2.13 a | 2.99 ± 0.01 b | |||
| p value | <0.0001 | 0.0466 | 0.1716 | 0.0002 | 0.0185 | |||
| g.35691328T>C | 1 | AA (626) | 10310 ± 63.52 | 347.08 ± 2.81Aa | 3.36 ± 0.03 a | 304.24 ± 2.07 | 2.96 ± 0.01 a | 3 |
| AG (375) | 10289 ± 68.03 | 344.2 ± 2.97a | 3.33 ± 0.03 ab | 303.88 ± 2.18 | 2.96 ± 0.01 a | |||
| GG (59) | 10389 ± 112.86 | 331.3 ± 4.95Bb | 3.25 ± 0.05 b | 302.57 ± 3.42 | 2.92 ± 0.02 b | |||
| p value | 0.6345 | 0.0014 | 0.0248 | 0.8566 | 0.0302 | |||
| 2 | AA (436) | 10631 ± 65.52 Aa | 384.07 ± 2.92 | 3.64 ± 0.03 | 316.66 ± 2.12 Aa | 2.99 ± 0.01 a | 3 | |
| AG (254) | 10910 ± 72.91 B | 388.98 ± 3.2 | 3.58 ± 0.03 | 322.6 ± 2.33 B | 2.96 ± 0.01 b | |||
| GG (44) | 10395 ± 131.93 Aa | 378.08 ± 5.47 | 3.66 ± 0.05 | 307.3 ± 3.99 Ab | 2.96 ± 0.02 ab | |||
| p value | <.0001 | 0.0505 | 0.0698 | <.0001 | 0.0175 | |||
| g.35691371A>G | 1 | AA (604) | 10378 ± 62.51 | 345.22 ± 2.79 Aa | 3.34 ± 0.03 Aa | 306.3 ± 2.03 | 2.95 ± 0.01 Aa | 3 |
| AG (343) | 10336 ± 68.31 | 341.23 ± 3.01 ab | 3.31 ± 0.03 a | 305.12 ± 2.19 | 2.95 ± 0.01 a | |||
| GG (59) | 10417 ± 112.77 | 332.6 ± 4.7 Bb | 3.2 ± 0.05B b | 302.27 ± 3.43 | 2.91 ± 0.02 Bb | |||
| p value | 0.6244 | 0.0048 | 0.0035 | 0.3689 | 0.0075 | |||
| 2 | AA (435) | 10755 ± 66.43 b | 388.73 ± 2.96 | 3.63 ± 0.03 | 320.29 ± 2.16 ab | 2.98 ± 0.01 | 2 | |
| AG (246) | 10938 ± 73.48 Aa | 391.8 ± 3.22 | 3.59 ± 0.03 | 324.21 ± 2.34 Aa | 2.96 ± 0.01 | |||
| GG (45) | 10506 ± 130.52 Bb | 385.76 ± 5.41 | 3.68 ± 0.05 | 311.74 ± 3.94Bb | 2.96 ± 0.02 | |||
| p value | 0.0007 | 0.359 | 0.1113 | 0.0026 | 0.1356 | |||
| No. of significant SNPs | 1 | 0 | 5 | 9 | 0 | 8 | ||
| 2 | 10 | 5 | 1 | 10 | 6 |
| Lactation | Haplotype Combination (No.) | Milk Yield (kg) | Fat Yield (kg) | Fat Percentage (%) | Protein Yield (kg) | Protein Percentage (%) |
|---|---|---|---|---|---|---|
| 1 | H1H1(603) | 10420 ± 65.07 | 347.91 ± 2.9 | 3.35 ± 0.03 | 309.3 ± 2.11 | 2.96 ± 0.01 a |
| H1H3(350) | 10363 ± 70.17 | 343.55 ± 3.09 | 3.33 ± 0.03 | 307.37 ± 2.25 | 2.96 ± 0.01 a | |
| H3H3(59) | 10495 ± 114.84 | 341.13 ± 4.79 | 3.26 ± 0.05 | 307.19 ± 3.49 | 2.93 ± 0.02 b | |
| p value | 0.358 | 0.066 | 0.0819 | 0.4332 | 0.0393 | |
| 2 | H1H1(416) | 10654 ± 67.2 A | 384.36 ± 3 | 3.63 ± 0.03a | 316.81 ± 2.18 Aa | 2.98 ± 0.01 a |
| H1H3(238) | 10875 ± 74.21 B | 386.56 ± 3.24 | 3.57 ± 0.03b | 321.19 ± 2.36 ab | 2.96 ± 0.01 b | |
| H3H3(44) | 10416 ± 133.65 A | 379.5 ± 5.54 | 3.66 ± 0.05ab | 307.95 ± 4.04 Bb | 2.96 ± 0.02 ab | |
| p value | 0.0002 | 0.3786 | 0.047 | 0.0015 | 0.0386 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Han, B.; Yuan, Y.; Li, Y.; Liu, L.; Sun, D. Single Nucleotide Polymorphisms of NUCB2 and Their Genetic Associations with Milk Production Traits in Dairy Cows. Genes 2019, 10, 449. https://doi.org/10.3390/genes10060449
Han B, Yuan Y, Li Y, Liu L, Sun D. Single Nucleotide Polymorphisms of NUCB2 and Their Genetic Associations with Milk Production Traits in Dairy Cows. Genes. 2019; 10(6):449. https://doi.org/10.3390/genes10060449
Chicago/Turabian StyleHan, Bo, Yuwei Yuan, Yanhua Li, Lin Liu, and Dongxiao Sun. 2019. "Single Nucleotide Polymorphisms of NUCB2 and Their Genetic Associations with Milk Production Traits in Dairy Cows" Genes 10, no. 6: 449. https://doi.org/10.3390/genes10060449
APA StyleHan, B., Yuan, Y., Li, Y., Liu, L., & Sun, D. (2019). Single Nucleotide Polymorphisms of NUCB2 and Their Genetic Associations with Milk Production Traits in Dairy Cows. Genes, 10(6), 449. https://doi.org/10.3390/genes10060449





