Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR?
Abstract
1. Introduction
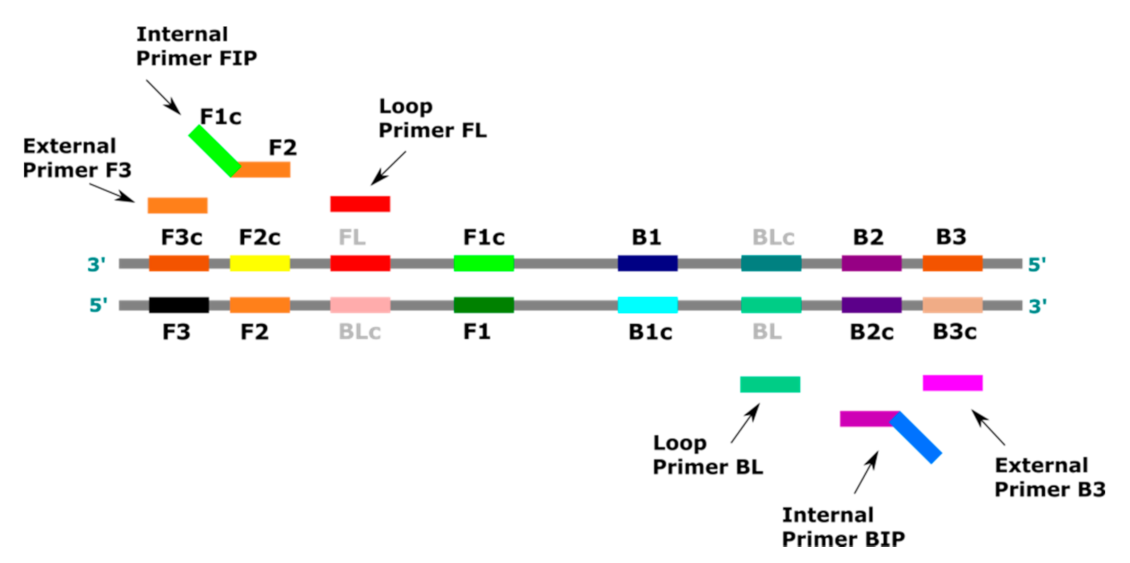
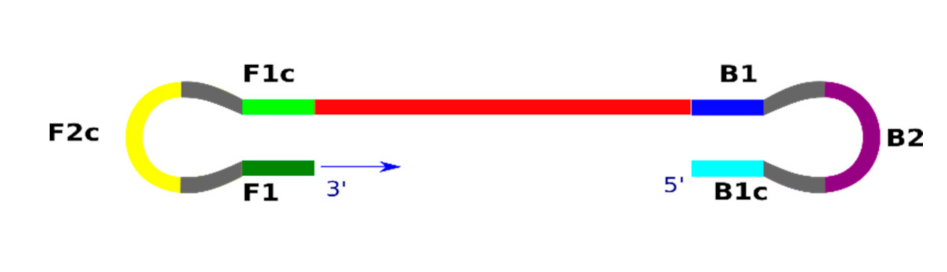
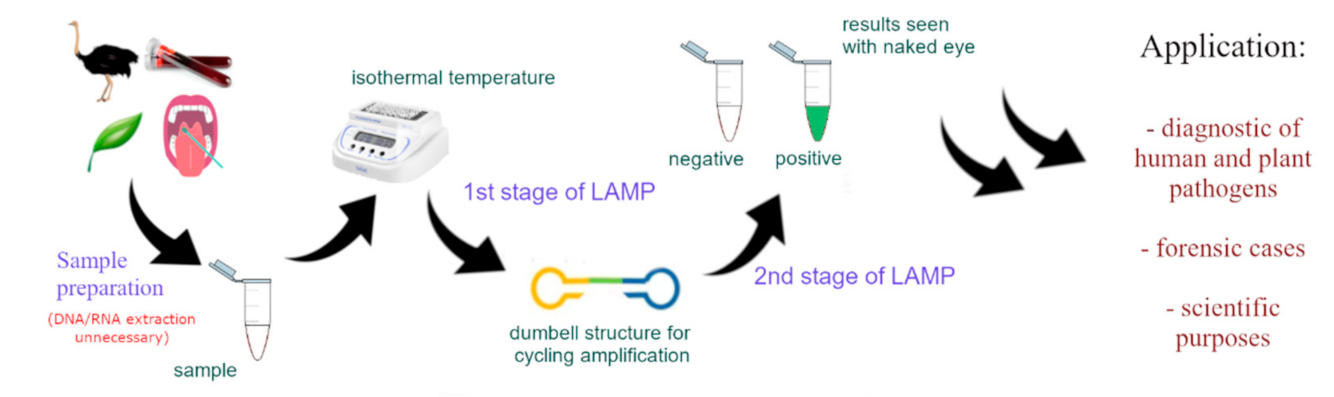
2. LAMP Method
3. Detection of LAMP Products
4. LAMP Application
4.1. Detection of Plant Pathogens
4.2. Detection of Animal Pathogens
4.3. Application of LAMP in Forensics
4.4. Detection of Human Pathogens
5. LAMP for SARS-CoV-2 Detection
6. Limitations of the LAMP Method
7. Summary
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Decorte, R.; Cassiman, J.J. Forensic medicine and the polymerase chain reaction technique. J. Med. Genet. 1993, 30, 625–633. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Noble, D. Forensic PCR. Primed, amplified, and ready for court. Anal. Chem. 1995, 67, 613A–615A. [Google Scholar] [CrossRef]
- Valasek, M.A.; Repa, J.J. The power of real-time PCR. Adv. Physiol. Educ. 2005, 29, 151–159. [Google Scholar] [CrossRef]
- Garibyan, L.; Avashia, N. Polymerase chain reaction. J. Investig. Dermatol. 2013, 133, 1–4. [Google Scholar] [CrossRef]
- Keller, M.; Naue, J.; Zengerle, R.; Stetten, F.; Schmidt, U. Automated forensic animal family identification by nested PCR and melt curve analysis on an off-the-shelf thermocycler augmented with a centrifugal microfluidic disk segment. PLoS ONE 2015, 10, e0131845. [Google Scholar] [CrossRef] [PubMed]
- Kermekchiev, M.B.; Kirilovam, L.I.; Vail, E.E.; Barnes, W.M. Mutants of Taq DNA polymerase resistant to PCR inhibitors allow DNA amplification from whole blood and crude soil samples. Nucleic Acids Res. 2009, 37, e40. [Google Scholar] [CrossRef] [PubMed]
- Cai, D.; Behrmann, O.; Hufert, F.; Dame, G.; Urban, G. Capacity of rTth polymerase to detect RNA in the presence of various inhibitors. PLoS ONE 2018, 13, e0190041. [Google Scholar] [CrossRef] [PubMed]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, E63. [Google Scholar] [CrossRef] [PubMed]
- Parida, M.; Sannarangaiah, S.; Dash, P.K.; Rao, P.V.; Morita, K. Loop mediated isothermal amplification (LAMP): A new generation of innovative gene amplification technique; perspectives in clinical diagnosis of infectious diseases. Rev. Med. Virol. 2008, 18, 407–421. [Google Scholar] [CrossRef]
- Nagamine, K.; Hase, T.; Notomi, T. Accelerated reaction by loop-mediated isothermal amplification using loop primers. Mol. Cell. Probes 2002, 16, 223–229. [Google Scholar] [CrossRef]
- Nagamine, K.; Watanabe, K.; Ohtsuka, K.; Hase, T.; Notomi, T. Loop-mediated isothermal amplification reaction using a nondenatured template. Clin. Chem. 2001, 47, 1742–1743. [Google Scholar] [CrossRef]
- Mori, Y.; Kitao, M.; Tomita, N.; Notomi, T. Real-time turbidimetry of LAMP reaction for quantifying template DNA. J. Biochem. Biophys. Methods 2004, 59, 145–157. [Google Scholar] [CrossRef]
- Chander, Y.; Koelbl, J.; Puckett, J.; Moser, M.J.; Klingele, A.J.; Liles, M.R.; Carrias, A.; Mead, D.A.; Schoenfeld, T.W. A novel thermostable polymerase for RNA and DNA loop-mediated isothermal amplification (LAMP). Front. Microbiol. 2014, 5, 395. [Google Scholar] [CrossRef]
- Paik, I.; Ngo, P.H.T.; Shroff, R.; Maranhao, A.C.; Walker, D.J.F.; Bhadra, S.; Ellington, A.D. Multi-modal engineering of Bst DNA polymerase for thermostability in ultra-fast LAMP reactions. bioRxiv 2021. [Google Scholar] [CrossRef]
- Budziszewska, M.; Wieczorek, P.; Obrępalska-Stęplowska, A. One-step reverse transcription loop-mediated isothermal amplification (RT-LAMP) for detection of tomato torrado virus. Arch. Virol. 2016, 161, 1359–1364. [Google Scholar] [CrossRef]
- Sahoo, P.R.; Sethy, K.; Mohapatra, S.; Panda, D. Loop mediated isothermal amplification: An innovative gene amplification technique for animal diseases. Vet. World 2016, 9, 465–469. [Google Scholar] [CrossRef]
- Yu, L.; Wu, S.; Hao, X.; Dong, X.; Mao, L.; Pelechano, V.; Chen, W.-H.; Yin, X. Rapid colorimetric detection of COVID-19 coronavirus using a reverse transcriptional loop-mediated isothermal amplification (RT-LAMP) diagnostic platform. Clin. Chem. 2020, 66, 975–977. [Google Scholar] [CrossRef]
- Mahony, J.; Chong, S.; Bulir, D.; Ruyter, A.; Mwawasi, K.; Waltho, D. Multiplex loop-mediated isothermal amplification (M-LAMP) assay for the detection of influenza A/H1, A/H3 and influenza B can provide a specimen-to-result diagnosis in 40 min with single genome copy sensitivity. J. Clin. Virol. 2013, 58, 127–131. [Google Scholar] [CrossRef]
- Da Silva, S.J.R.; Pardee, K.; Pena, L. Loop-Mediated Isothermal Amplification (LAMP) for the diagnosis of Zika Virus: A review. Viruses 2020, 12, 19. [Google Scholar] [CrossRef]
- Foo, P.C.; Chan, Y.Y.; Mohamed, M.; Wong, W.K.; Nurul Najian, A.B.; Lim, H. Development of a thermostabilised triplex LAMP assay with dry-reagent four target lateral flow dipstick for detection of Entamoeba histolytica and non-pathogenic Entamoeba spp. Anal. Chim. Acta 2017, 966, 71–80. [Google Scholar] [CrossRef]
- Mori, Y.; Nagamine, K.; Tomita, N.; Notomi, T. Detection of loop-mediated isothermal amplification reaction by turbidity derived from magnesium pyrophosphate formation. Biochem. Biophys. Res. Commun. 2001, 289, 150–154. [Google Scholar] [CrossRef]
- Tian, B.; Qiu, Z.; Ma, J.; de la Torre, T.Z.G.; Johansson, C.; Svedlindh, P.; Stromberg, M. Attomolar Zika virus oligonucleotide detection based on loop-mediated isothermal amplification and AC susceptometry. Biosens. Bioelectron. 2016, 86, 420–425. [Google Scholar] [CrossRef] [PubMed]
- Dhama, K.; Karthik, K.; Chakraborty, S.; Tiwari, R.; Kapoor, S.; Kumar, A.; Thomas, P. Loop-mediated isothermal amplification of DNA (LAMP): A new diagnostic tool lights the world of diagnosis of animal and human pathogens: A review. Pak. J. Biol. Sci. 2014, 17, 151–166. [Google Scholar] [CrossRef] [PubMed]
- Imai, M.; Ninomiya, A.; Minekawa, H.; Notomi, T.; Ishizaki, T.; van Tu, P.; Tien, N.T.; Tashiro, M.; Odagiri, T. Rapid diagnosis of H5N1 avian influenza virus infection by newly developed influenza H5 hemagglutinin gene-specific loop-mediated isothermal amplification method. J. Virol. Methods 2007, 141, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Cai, T.; Lou, G.; Yang, J.; Xu, D.; Meng, Z. Development and evaluation of real-time loop-mediated isothermal amplification for hepatitis B virus DNA quantification: A new tool for HBV management. J. Clin. Virol. 2008, 41, 270–276. [Google Scholar] [CrossRef] [PubMed]
- Przybylska, A.; Fiedler, Ż.; Kucharczyk, H.; Obrępalska-Stęplowska, A. Detection of the quarantine species Thrips palmi by loop-mediated isothermal amplification. PLoS ONE 2015, 10, e0122033. [Google Scholar] [CrossRef]
- Thompson, D.; Lei, Y. Mini review: Recent progress in RT-LAMP enabled COVID-19 detection. Sens. Actuators Rep. 2020, 2, 100017. [Google Scholar] [CrossRef]
- Fukuta, S.; Iidia, T.; Mizukami, Y.; Ishida, A.; Ueda, J.; Kanbee, M.; Ishimoto, Y. Detection of Japanese yam mosaic virus by RT-LAMP. Arch. Virol. 2003, 148, 1713–1720. [Google Scholar] [CrossRef] [PubMed]
- Boubourakas, I.N.; Fukuta, S.; Kyriakopoulou, P.E. Sensitive and rapid detection of peach latent mosaic viroid by the reverse transcription loop-mediated isothermal amplification. J. Virol. Methods 2009, 160, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Moradi, A.; Nasiri, J.; Abdollahi, H.; Almasi, M. Development and evaluation of a loop-mediated isothermal amplification assay for detection of Erwinia amylovora based on chromosomal DNA. Eur. J. Plant Pathol. 2012, 133, 609–620. [Google Scholar] [CrossRef]
- Przewodowska, A.; Zacharzewska, B.; Chołuj, J.; Treder, K. A one step, real-time reverse transcription loop mediated isothermal amplification assay to detect Potato virus Y. Am. J. Potato Res. 2015, 92, 303–311. [Google Scholar] [CrossRef][Green Version]
- Miyake, N.; Nagai, H.; Kato, S.; Matsusaki, M.; Fukuta, S.; Takahashi, R.; Suzuki, R.; Ishiguro, Y. Practical method combining loop-mediated isothermal amplification and bait trap to detect Pythium helicoides from hydroponic culture solutions. J. Gen. Plant Pathol. 2017, 83, 1–6. [Google Scholar] [CrossRef]
- Feng, W.; Hieno, A.; Kusunoki, M.; Suga, H.; Kageyama, K. LAMP Detection of four plant-pathogenic oomycetes and its application in lettuce fields. Plant Dis. 2019, 103, 298–307. [Google Scholar] [CrossRef] [PubMed]
- Panno, S.; Matić, S.; Tiberini, A.; Caruso, A.G.; Bella, P.; Torta, L.; Stassi, R.; Davino, S. Loop mediated isothermal amplification: Principles and applications in plant virology. Plants 2020, 9, 461. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Lee, S.Y.; Back, C.G.; Ten, L.N.; Jung, H.Y. Loop-mediated isothermal amplification for the detection of Xanthomonas arboricola pv. pruni in peaches. Plant Pathol. J. 2019, 35, 635–643. [Google Scholar] [CrossRef] [PubMed]
- Harper, S.J.; Ward, L.; Clover, G. Development of LAMP and real-time PCR methods for the rapid detection of Xylella fastidiosa for quarantine and field applications. Phytopatholgy 2010, 100, 1282–1288. [Google Scholar] [CrossRef] [PubMed]
- Fukuta, S.; Kato, S.; Yoshida, K.; Mizukami, Y.; Ishida, A.; Ueda, J.; Michio, K.; Ishimoto, Y. Detection of tomato yellow leaf curl virus by loop-mediated isothermal amplification reaction. J. Virol. Methods 2003, 112, 35–40. [Google Scholar] [CrossRef]
- Fukuta, S.; Ohishi, K.; Yoshida, K.; Mizukami, Y.; Ishida, A.; Kanbe, M. Development of immunocapture reverse transcription loop-mediated isothermal amplification for the detection of tomato spotted wilt virus from chrysanthemum. J. Virol. Methods 2004, 121, 49–55. [Google Scholar] [CrossRef]
- Fukuta, S.; Nimi, Y.; Ohishi, K.; Yoshimura, Y.; Anai, N.; Hotta, M.; Fukaya, M.; Kato, T.; Oya, T.; Kanbe, M. Development of reverse transcription loop-mediated isothermal amplification (RT-LAMP) method for detection of two viruses and chrysanthemum stunt viroid. Ann. Rep. Kansai Plant Prot. Soc. 2005, 47, 31–36. [Google Scholar] [CrossRef]
- Kaneko, H.; Kawana, T.; Fukushima, E.; Suzutani, T. Toleranceof loop-mediated isothermal amplification to a culturemedium and biological substances. J. Biochem. Biophys. Methods 2007, 70, 499–501. [Google Scholar] [CrossRef]
- Tomlinson, J.; Boonham, N. Potential of LAMP for detection of plant pathogens. CAB Rev. Perspect. Agric. Vet. Sci. Nutr. Nat. Resour. 2008, 3, 1–7. [Google Scholar] [CrossRef]
- Lee, D.; la Mura, M.; Allnutt, T.R.; Powell, W. Detection of genetically modified organisms (GMOs) using isothermal amplification of target DNA sequences. BMC Biotechnol. 2009, 9, 7. [Google Scholar] [CrossRef]
- Lee, D.; la Mura, M.; Allnutt, T.R.; Powell, W.; Reenland, A.G. Isothermal amplification of genetically modified DNA sequences directly from plant tissues lowers the barriers to high-throughput and field-based genotyping. J. Agric. Food Chem. 2009, 5, 9400–9402. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, Y.; Komatsu, K.; Nagumo, S. Rapid detection of Panax ginseng by loop-mediated isothermal amplification and its application to authentication of Ginseng. Biol. Pharm. Bull. 2008, 31, 1806–1808. [Google Scholar] [CrossRef]
- Chaudhary, A.A.; Heman, T.; Mohsin, M.; Ahmad, A. Application of loop-mediated isothermal amplification (LAMP)-based technology for authentication of Catharanthus roseus (L) G. Don. Protoplasma 2011, 249, 417–722. [Google Scholar] [CrossRef] [PubMed]
- Soliman, H.; El-Matbouli, M. Immunocapture and direct binding loop mediated isothermal amplification simplify molecular diagnosis of Cyprinid herpesvirus-3. J. Virol. Methods 2009, 162, 91–95. [Google Scholar] [CrossRef]
- Chen, H.T.; Zhang, J.; Ma, Y.P.; Ma, L.N.; Ding, Y.Z.; Liu, X.T.; Cai, X.P.; Ma, L.Q.; Zhang, Y.G.; Liu, Y.S. Reverse transcription loop-mediated isothermal amplification for the rapid detection of infectious bronchitis virus in infected chicken tissues. Mol. Cell. Probes 2010, 24, 104–106. [Google Scholar] [CrossRef]
- Bao, H.; Wang, X.; Zhao, Y.; Sun, X.; Li, Y.; Xiong, Y.; Chen, H. Development of a reverse transcription loop-mediated isothermal amplification method for the rapid detection of avian influenza virus subtype H7. J. Virol. Methods 2012, 179, 33–37. [Google Scholar] [CrossRef]
- Khunthong, S.; Jaroenram, W.; Arunrut, N.; Suebsing, R.; Mungsantisuk, I.; Kiatpathomchai, W. Rapid and sensitive detection of shrimp yellow head virus by loop-mediated isothermal amplification combined with a lateral flow dipstick. J. Virol. Methods 2013, 188, 51–56. [Google Scholar] [CrossRef]
- Li, X.; Zhou, Y.; Ji, H.; Xu, Z.; Zhu, L. One-step reverse transcription-loop-mediated isothermal amplification assay for sensitive and rapid detection of porcine kobuvirus. J. Virol. Methods 2014, 207, 1–5. [Google Scholar] [CrossRef]
- Food and Agriculture Organization (FAO). The State of World Fisheries and Aquaculture 2020; World Review; Food and Agriculture Organization (FAO): Rome, Italy, 2020. [Google Scholar] [CrossRef]
- Soliman, H.; Midtlyng, P.; El-Matbouli, M. Sensitive and rapid detection of infectious pancreatic necrosis virus by reverse transcription loop mediated isothermal amplification. J. Virol. Methods 2009, 158, 77–78. [Google Scholar] [CrossRef]
- Soliman, H.; Saleh, M.; El-Matbouli, M. Detection of fish pathogens by Loop-Mediated Isothermal Amplification (LAMP) technique. In Veterinary Infection Biology: Molecular Diagnostics and High-Throughput Strategies; Methods in Molecular Biology (Methods and Protocols); Cunha, M., Inácio, J., Eds.; Humana Press: New York, NY, USA, 2015; pp. 163–173. [Google Scholar] [CrossRef]
- Arzt, J.; Juleff, N.; Zhang, Z.; Rodriguez, L.L. The pathogenesis of foot-and-mouth disease I: Viral pathways in cattle. Transbound. Emerg. Dis. 2011, 58, 291–304. [Google Scholar] [CrossRef]
- Dukes, J.P.; King, D.P.; Alexandersen, S. Novel reverse transcription loop-mediated isothermal amplification for rapid detection of foot-and-mouth disease virus. Arch. Virol. 2006, 151, 1093–1106. [Google Scholar] [CrossRef] [PubMed]
- Bucher, B.J.; Muchaamba, G.; Kamber, T.; Kronenberg, P.A.; Abdykerimov, K.K.; Isaev, M.; Deplazes, P.; Alvarez Rojas, C.A. LAMP assay for the detection of Echinococcus multilocularis eggs isolated from canine faeces by a cost-effective NaOH-based DNA extraction method. Pathogens 2021, 10, 847. [Google Scholar] [CrossRef]
- Arenas, M.; Pereira, F.; Oliveira, M.; Pinto, N.; Lopes, A.M.; Gomes, V.; Carracedo, A.; Amorim, A. Forensic genetics and genomics: Much more than just a human affair. PLoS Genet. 2017, 13, e1006960. [Google Scholar] [CrossRef]
- Watthanapanpituck, K.; Kiatpathomchai, W.; Chu, E.; Panvisavas, N. Identification of human DNA in forensic evidence by loop-mediated isothermal amplification combined with a colorimetric gold nanoparticle hybridization probe. Int. J. Legal Med. 2014, 128, 923–931. [Google Scholar] [CrossRef]
- Johnston, S.; Newman, J.; Frappier, R. Validation study of the Abacus Diagnostics ABAcard® Hematrace® membrane test for the forensic identification of human blood. Can. Soc. Forensic Sci. J. 2003, 36, 173–183. [Google Scholar] [CrossRef]
- Misencik, A.; Laux, D.L. Validation study of the Seratec HemDirect hemoglobin assay for the forensic identification of human blood. MAFS Newslett. 2007, 36, 18–26. [Google Scholar]
- Özpınar, H.; Tezmen, G.; Gökçe, I.; Tekiner, İ.H. Detection of animal species in some meat and meat products by comparatively using DNA microarray and real-time PCR Methods. Kafkas Univ. Vet. Fak. Derg. 2013, 19, 245–252. [Google Scholar] [CrossRef]
- Dooley, J.J.; Paine, K.E.; Garrett, S.D.; Brown, H.M. Detection of meat species using TaqMan real-time PCR assays. Meat Sci. 2004, 68, 431–438. [Google Scholar] [CrossRef]
- Kumari, S.; Kumar, R.R.; Mendiratta, S.K.; Kumar, D.; Rana, P.; Kumar, D.; Jawla, J. Species-specific loop-mediated isothermal amplification (LAMP) assay for identification of tissue of cattle origin by targeting mitochondrial gene sequences. 3 Biotech 2019, 9, 69. [Google Scholar] [CrossRef] [PubMed]
- Kanchanaphum, P.; Maneenin, S.; Chaiyana, W. Analysis of pork meat using loop mediated isothermal amplification (LAMP) to confirm halal status. Int. J. Biosci. 2014, 4, 62–68. [Google Scholar] [CrossRef]
- Yang, L.; Fu, S.; Peng, X.; Li, L.; Song, T.; Li, L. Identification of pork in meat products using real-time loop-mediated isothermal amplification. Biotechnol. Biotechnol. Equip. 2014, 28, 882–888. [Google Scholar] [CrossRef]
- Lee, S.-Y.; Kim, M.-J.; Hong, Y.; Kim, H.-Y. Development of a rapid on-site detection method for pork in processed meat products using real-time loop-mediated isothermal amplification. Food Control. 2016, 66, 53–61. [Google Scholar] [CrossRef]
- Girish, P.S.; Barbuddhe, S.B.; Kumari, A.; Rawool, D.B.; Karabasanavar, N.S.; Muthukumar, M.; Vaithiyanathan, S. Rapid detection of pork using alkaline lysis- Loop Mediated Isothermal Amplification (AL-LAMP) technique. Food Control. 2020, 110, 107015. [Google Scholar] [CrossRef]
- Thangsunan, P.; Temisak, S.; Morris, P.; Rios-Solis, L.; Suree, N. Combination of loop-mediated isothermal amplification and AuNP-oligoprobe colourimetric assay for pork authentication in processed meat products. Food Anal. Methods 2020, 14, 568–580. [Google Scholar] [CrossRef]
- Abdullahi, U.F.; Igwenagu, E.; Aliyu, S.; Mu’azu, A.; Naim, R.; Wan-Taib, W.R. A rapid and sensitive Loop-mediated isothermal amplification assay for detection of pork DNA based on porcine tRNAlys and ATPase 8 genes. Int. Food Res. J. 2017, 24, 1357–1361. [Google Scholar]
- Ran, G.; Ren, L.; Han, X.; Liu, X.; Li, Z.; Pang, D. Development of a rapid method for the visible detection of pork DNA in halal products by loop-mediated isothermal amplification. Food Anal. Methods 2016, 9, 565–570. [Google Scholar] [CrossRef]
- Abdulmawjood, A.; Grabowski, N.; Fohler, S.; Kittler, S.; Nagengast, H.; Klein, G. Development of Loop-Mediated Isothermal Amplification (LAMP) assay for rapid and sensitive identification of strus meat. PLoS ONE 2014, 9, e100717. [Google Scholar] [CrossRef]
- Wang, D.; Wang, Y.; Zhu, K.; Shi, L.; Zhang, M.; Yu, J.; Liu, Y. Detection of cassava component in sweet potato noodles by Real-time Loop-mediated Isothermal Amplification (Real-time LAMP) Method. Molecules 2019, 24, 2043. [Google Scholar] [CrossRef]
- Jackson, K.R.; Layne, T.; Dent, D.A.; Tsuei, A.; Li, J.; Haverstick, D.M.; Landers, J.P. A novel loop-mediated isothermal amplification method for identification of four body fluids with smartphone detection. Forensic Sci. Int. Genet. 2019, 45, 102195. [Google Scholar] [CrossRef]
- Layne, T.; Jackson, K.; Scott, A.; Tanner, N.A.; Piland, A.; Haverstick, D.M.; Landers, J.P. Optimization of novel loop-mediated isothermal amplification with colorimetric image analysis for forensic body fluid identification. J. Forensic Sci. 2021, 66, 1033–1041. [Google Scholar] [CrossRef] [PubMed]
- Nogami, H.; Tsutsumi, H.; Komuro, T.; Mukoyama, R. Rapid and simple sex determination method from dental pulp by loop-mediated isothermal amplification. Forensic Sci. Int. Gen. 2008, 2, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Kanchanaphum, P. Time course of detection of human male DNA from stained blood sample on various surfaces by Loop Mediated Isothermal Amplification and polymerase chain reaction. BioMed Res. Int. 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Slots, J.; Genco, R.J. Black-pigmented Bacteroides species, Capnocytophaga species, and Actinobacillus actinomycetemcomitans in human periodontal disease: Virulence factors in colonization, survival, and tissue destruction. J. Dent. Res. 1984, 63, 412–421. [Google Scholar] [CrossRef]
- Gersdorf, H.; Meissner, A.; Pelz, K.; Krekeler, G.; Göbel, U.B. Identification of Bacteroides forsythus in subgingival plaque from patients with advanced periodontitis. J. Clin. Microbiol. 1993, 31, 941–946. [Google Scholar] [CrossRef]
- Kasuga, Y.; Ishihara, K.; Okuda, K. Significance of detection of Porphyromonas gingivalis, Bacteroides forsythus and Treponema denticola in periodontal pockets. Bull. Tokyo Dent. Coll. 2000, 41, 109–117. [Google Scholar] [CrossRef][Green Version]
- Kazor, C.E.; Mitchell, P.M.; Lee, A.M.; Stokes, L.N.; Loesche, W.J.; Dewhirst, F.E.; Paster, B.J. Diversity of bacterial populations on the tongue dorsa of patients with halitosis and healthy patients. J. Clin. Microbiol. 2003, 41, 558–563. [Google Scholar] [CrossRef]
- Kuramitsu, H.K.; Qi, M.; Kang, I.C.; Chen, W. Role for periodontal bacteria in cardiovascular diseases. Ann. Periodontol. 2001, 61, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Kuramitsu, H.K.; Kang, I.C.; Qi, M. Interactions of Porphyromonas gingivalis with host cells: Implications for cardiovascular diseases. J. Periodontol. 2003, 74, 85–89. [Google Scholar] [CrossRef] [PubMed]
- Ishihara, K.; Nabuchi, A.; Ito, R.; Miyachi, K.; Kuramitsu, H.K.; Okuda, K. Correlation between detection rates of periodontopathic bacterial DNA in coronary stenotic artery plaque [corrected] and in dental plaque samples. J. Clin. Microbiol. 2004, 42, 1313–1315. [Google Scholar] [CrossRef]
- Carroll, N.M.; Jaeger, E.E.; Choudhury, S.; Dunlop, A.A.; Matheson, M.M.; Adamson, P.; Okhravi, N.; Lightman, S. Detection of and discrimination between gram-positive and gram-negative bacteria in intraocular samples by using nested PCR. J. Clin. Microbiol. 2000, 38, 1753–1757. [Google Scholar] [CrossRef]
- Yoshida, A.; Nagashima, S.; Ansai, T.; Tachibana, M.; Kato, H.; Watari, H.; Notomi, T.; Takehara, T. Loop-mediated isothermal amplification method for rapid detection of the periodontopathic bacteria Porphyromonas gingivalis, Tannerella forsythia, and Treponema denticola. J. Clin. Microbiol. 2005, 43, 2418–2424. [Google Scholar] [CrossRef]
- Nakauchi, M.; Takayama, I.; Takahashi, H.; Tashiro, M.; Kageyama, T. Development of a reverse transcription loop-mediated isothermal amplification assay for the rapid diagnosis of avian influenza A (H7N9) virus infection. J. Virol. Methods 2014, 204, 101–104. [Google Scholar] [CrossRef]
- Bao, H.; Feng, X.; Ma, Y.; Shi, J.; Zhao, Y.; Gu, L.; Wang, X.; Chen, H. Rapid detection of subtype H10N8 influenza virus by one-step reverse transcription-loop-mediated isothermal amplification Methods. J. Clin. Microbiol. 2015, 53, 3884–3887. [Google Scholar] [CrossRef]
- Kim, E.M.; Jeon, H.S.; Kim, J.J.; Shin, Y.K.; Lee, Y.J.; Yeo, S.G.; Park, C.K. Uracil-DNA glycosylase-treated reverse transcription loop-mediated isothermal amplification for rapid detection of avian influenza virus preventing carry-over contamination. J. Vet. Sci. 2016, 17, 421–425. [Google Scholar] [CrossRef]
- Hu, S.F.; Li, M.; Zhong, L.L.; Lu, S.M.; Liu, Z.X.; Pu, J.Y.; Wen, J.S.; Huang, X. Development of reverse-transcription loop-mediated isothermal amplification assay for rapid detection and differentiation of dengue virus serotypes 1–4. BMC Microbiol. 2015, 15, 265. [Google Scholar] [CrossRef] [PubMed]
- Lau, Y.L.; Lai, M.Y.; Teoh, B.T.; Abd-Jamil, J.; Johari, J.; Sam, S.S.; Tan, K.K.; Abubakar, S. Colorimetric detection of dengue by single tube reverse-transcription-loop-mediated isothermal amplification. PLoS ONE 2015, 10, e0138694. [Google Scholar] [CrossRef] [PubMed]
- Hoos, J.; Peters, R.M.; Tabatabai, J.; Grulich-Henn, J.; Schnitzler, P.; Pfeil, J. Reverse-transcription loop-mediated isothermal amplification for rapid detection of respiratory syncytial virus directly from nasopharyngeal swabs. J. Virol. Methods 2017, 242, 53–57. [Google Scholar] [CrossRef]
- Nyan, D.C.; Swinson, K.L. A method for rapid detection and genotype identification of hepatitis C virus 1-6 by one-step reverse transcription loop-mediated isothermal amplification. Int. J. Infect. Dis. 2016, 43, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Zhao, N.; Liu, J.; Sun, D. Detection of HCV genotypes 1b and 2a by a reverse transcription loop-mediated isothermal amplification assay. J. Med. Virol. 2017, 89, 1048–1054. [Google Scholar] [CrossRef]
- Oloniniyi, O.K.; Kurosaki, Y.; Miyamoto, H.; Takada, A.; Yasuda, J. Rapid detection of all known ebolavirus species by reverse transcription-loop-mediated isothermal amplification (RT-LAMP). J. Virol. Methods 2017, 246, 8–14. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Yin, F.; Bi, Y.; Cheng, G.; Li, J.; Hou, L.; Li, Y.; Yang, B.; Liu, W.; Yang, L. Rapid and sensitive detection of Zika virus by reverse transcription loop-mediated isothermal amplification. J. Virol. Methods 2016, 238, 86–93. [Google Scholar] [CrossRef]
- Chotiwan, N.; Brewster, C.D.; Magalhaes, T.; Weger-Lucarelli, J.; Duggal, N.K.; Rückert, C.; Nguyen, C.; Garcia Luna, S.M.; Fauver, J.R.; Andre, B.; et al. Rapid and specific detection of Asian- and African-lineage Zika viruses. Sci. Transl. Med. 2017, 3, 9. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Nie, K.; Zhang, D.; Li, X.; Wang, Y.; Tan, W.; Ma, X. Detection of middle east respiratory syndrome coronavirus by reverse-transcription loop-mediated isothermal amplification. Chin. J. Virol. 2015, 31, 269–275. [Google Scholar]
- Lee, S.H.; Baek, Y.H.; Kim, Y.H.; Choi, Y.K.; Song, M.S.; Ahn, J.Y. One-pot reverse transcriptional loop-mediated isothermal amplification (RT-LAMP) for detecting MERS-CoV. Front. Microbiol. 2016, 7, 2166. [Google Scholar] [CrossRef]
- Ocwieja, K.E.; Sherrill-Mix, S.; Liu, C.; Song, J.; Bau, H.; Bushman, F.D. A reverse transcription loop-mediated isothermal amplification assay optimized to detect multiple HIV subtypes. PLoS ONE 2015, 10, e0117852. [Google Scholar] [CrossRef]
- Odari, E.O.; Maiyo, A.; Lwembe, R.; Gurtler, L.; Eberle, J.; Nitschko, H. Establishment and evaluation of a loop-mediated isothermal amplification (LAMP) assay for the semi-quantitative detection of HIV-1 group M virus. J. Virol. Methods 2015, 212, 30–38. [Google Scholar] [CrossRef]
- Damhorst, G.L.; Duarte-Guevara, C.; Chen, W.; Ghonge, T.; Cunningham, B.T.; Bashir, R. Smartphone-imaged HIV-1 reverse-transcription loop-mediated isothermal amplification (RT-LAMP) on a chip from whole blood. Engineering 2015, 1, 324–335. [Google Scholar] [CrossRef]
- Thi, V.L.D.; Herbst, K.; Boerner, K.; Meurer, M.; Kremer, L.; Kirrmaier, D.; Freistaedter, A.; Papagiannidis, D.; Galmozzi, C.; Stanifer, M.L.; et al. A colorimetric RT-LAMP assay and LAMP-sequencing for detecting SARS-CoV-2 RNA in clinical samples. Sci. Transl. Med. 2020, 12, eabc7075. [Google Scholar] [CrossRef]
- Lamb, L.E.; Bartolone, S.N.; Ward, E.; Chancellor, M.B. Rapid detection of novel coronavirus/Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) by reverse transcription-loop-mediated isothermal amplification. PLoS ONE 2020, 15, e0234682. [Google Scholar] [CrossRef]
- Pang, B.; Xu, J.; Liu, Y.; Peng, H.; Feng, W.; Cao, Y.; Wu, J.; Xiao, H.; Pabbaraju, K.; Tipples, G.; et al. Isothermal amplification and ambient visualization in a single tube for the detection of SARS-CoV-2 using Loop-Mediated amplification and CRISPR technology. Anal. Chem. 2020, 92, 16204–16212. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Hozumi, Y.; Yin, C.; Wei, G.-W. Decoding SARS-CoV-2 transmission and evolution and ramifications for COVID-19 diagnosis, vaccine, and medicine. J. Chem. Inf. Model. 2020, 60, 5853–5865. [Google Scholar] [CrossRef] [PubMed]
- Priya, A.; Ranjit, V. Role of SARS-CoV-2 and ACE2 variations in COVID-19. Biomed. J. 2021. In Press. [Google Scholar] [CrossRef]
- Corman, V.M.; Landt, O.; Kaiser, M.; Molenkamp, R.; Meijer, A.; Chu, D.K.; Bleicker, T.; Brünink, S. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR. Eur. Surveill. 2020, 25, 2000045. [Google Scholar] [CrossRef]
- Cagliani, R.; Forni, D.; Clerici, M.; Sironi, M. Coding potential and sequence conservation of SARS-CoV-2 and related animal viruses. Infect. Genet. Evol. 2020, 83, 104353. [Google Scholar] [CrossRef] [PubMed]
- Bai, Z.; Cao, Y.; Liu, W.; Li, J. The SARS-CoV-2 nucleocapsid protein and its role in viral structure, biological functions, and a potential target for drug or vaccine mitigation. Viruses 2021, 13, 1115. [Google Scholar] [CrossRef]
- Schmid-Burgk, J.L.; Li, D.; Feldman, D.; Slabicki, M.; Borrajo, J.; Strecker, J.; Cleary, B.; Regev, A.; Zhang, F. LAMP-Seq: Population-scale COVID-19 diagnostics using a compressed barcode space. bioRxiv 2020. [Google Scholar] [CrossRef]
- El-Tholoth, M.; Bau, H.H.; Song, J. A single and two-stage, closed-tube, molecular test for the 2019 novel coronavirus (COVID-19) at home, clinic, and points of entry. ChemRxiv 2020. Preprint. [Google Scholar] [CrossRef]
- Broughton, J.P.; Deng, X.; Yu, G.; Fasching, C.L.; Servellita, V.; Singh, J.; Miao, X.; Streithorst, J.A.; Granados, A.; Sotomayor-Gonzalez, A.; et al. CRISPR-Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 2020, 38, 870–874. [Google Scholar] [CrossRef]
- Jang, W.S.; Lim, D.H.; Yoon, J.; Kim, A.; Lim, M.; Nam, J.; Yanagihara, R.; Ryu, S.W.; Jung, B.K.; Ryoo, N.H.; et al. Development of a multiplex Loop-Mediated Isothermal Amplification (LAMP) assay for on-site diagnosis of SARS CoV-2. PLoS ONE 2021, 16, e0248042. [Google Scholar] [CrossRef]
- Lei, Y. Kitchen range oven enabled one-tube RT-LAMP for RNA detection at home—A potential solution for large-scale screening of COVID-19. engrXiv 2020. [Google Scholar] [CrossRef]
- Bektaş, A.; Covington, M.F.; Aidelberg, G.; Arce, A.; Matute, T.; Núñez, I.; Walsh, J.; Boutboul, D.; Lindner, A.B.; Federici, F.; et al. Accessible LAMP-Enabled Rapid Test (ALERT) for detecting SARS-CoV-2. medRxiv 2021. [Google Scholar] [CrossRef]
- Huang, W.E.; Lim, B.; Hsu, C.C.; Xiong, D.; Wu, W.; Yu, Y.; Jia, H.; Wang, Y.; Zeng, Y.; Ji, M.; et al. RT-LAMP for rapid diagnosis of coronavirus SARS-CoV-2. Microb. Biotechnol. 2020, 13, 950–961. [Google Scholar] [CrossRef] [PubMed]
- Chow, F.W.N.; Chan, T.T.Y.; Tam, A.R.; Zhao, S.; Yao, W.; Fung, J.; Cheng, F.K.K.; Lo, G.C.S.; Chu, S.; Aw-Yong, K.L.; et al. A rapid, simple, inexpensive, and mobile colorimetric assay COVID-19-LAMP for mass on-site screening of COVID-19. Int. J. Mol. Sci. 2020, 21, 5380. [Google Scholar] [CrossRef] [PubMed]
- Rabe, B.A.; Cepko, C. SARS-CoV-2 detection using an isothermal amplification reaction and a rapid, inexpensive protocol for sample inactivation and purification. Proc. Natl. Acad. Sci. USA 2020, 111, 24450–24458. [Google Scholar] [CrossRef]
- Wang, D. One-pot detection of COVID-19 with real-time reverse-transcription loop-mediated isothermal amplification (RT-LAMP) assay and visual RT-LAMP assay. bioRxiv 2020. [Google Scholar] [CrossRef]
- Iacobucci, G.; Coombes, R. Covid-19: Government plans to spend £100bn on expanding testing to 10 million a day. BMJ 2020, 370, m3520. [Google Scholar] [CrossRef]
- Iacobucci, G. Covid-19: Rapid test missed over 50% of positive cases in Manchester pilot. BMJ 2020, 371, m4323. [Google Scholar] [CrossRef]
- Halliday, J.; Boseley, S. Operation moonshot: Rapid Covid test missed over 50% of cases in pilot. Guardian, 5 November 2020. [Google Scholar]
- Gill, M.; Gray, M. Mass testing for covid-19 in the UK. BMJ 2020, 371, m4436. [Google Scholar] [CrossRef]
- Cox, D. The Big Problem with Operation Moonshot? False Positives. 2020. Available online: www.wired.co.uk (accessed on 17 May 2021).
- Mendick, R.; Donnelly, L. Moonshot testing plan could send 28 million into needless self-isolation. The Telegraph, 10 September 2020. [Google Scholar]
- Smyth, C.; McCall, B. Coronavirus: Experts criticise Johnson over ‘dangerous’ moonshot mass testing plan. The Times, 11 September 2020. [Google Scholar]
- Lovett, S. Coronavirus: Operation Moonshot ‘not feasible’ and ‘massive gamble’, health experts warn. The Independent, 12 September 2020. [Google Scholar]
- Department of Health and Social Care. Trial of Rapid Coronavirus Test Launched in Hampshire. 2020. Available online: www.gov.uk/government/news/trial-of-rapid-coronavirus-test-launched-in-hampshire (accessed on 31 January 2021).
- GENOMTEC. The Word’s First Genetic Diagnostic Test for the Detection of SARS-CoV-2 Directly from a Saliva Sample. Available online: https://genomtec.com/2021/05/12/genomtec-podsumowuje-osiagniecia-i-postep-prac-w-pierwszym-kwartale-2021-roku/ (accessed on 6 July 2021).
- Byrnes, S.A.; Gallagher, R.; Steadman, A.; Bennett, C.; Rivera, R.; Ortega, C.; Motley, S.T.; Jain, P.; Weigl, B.H.; Connelly, J.T. Multiplexed and extraction-free amplification for simplified SARS-CoV-2 RT-PCR Tests. Anal. Chem. 2021, 93, 4160–4165. [Google Scholar] [CrossRef] [PubMed]
- Chaouch, M. Loop-mediated isothermal amplification (LAMP): An effective molecular point-of-care technique for the rapid diagnosis of coronavirus SARS-CoV-2. Rev. Med. Virol. 2021, e2215. [Google Scholar] [CrossRef] [PubMed]
| Authors | Gene Target | Type of Samples | Number of Samples | Detection of Results | Limit of Detection (LoD) | Sample Preparation | Sensitivity/Specifity |
|---|---|---|---|---|---|---|---|
| Lamb et al. [103] | ORF1ab | Nasopharyngeal swab | 60 | Fluorescence detection, SYBR Green | 3 copies/µL | RNA extraction | — |
| Dao Thi et al. [102] | ORF1a, N | Nasopharyngeal swab | 95 | WarmStart Colorimetric LAMP | — | 5 min of hot swab–to–RT-LAMP assay | 92–99.7% |
| Pang et al. [104] | N, E | Nasopharyngeal swab | 100 | Fluorescence detection, SYBR Green, GeneFinder | 30 copies/µL | — | 94% |
| Yu et al. [17] | ORF1ab | Nasopharyngeal swab | 43 | Fluorescence detection | 10–100 copies/µL | RNA extraction | 97.6–100% |
| Schmid-Burgk et al. [110] | ORF1a, N | Nasopharyngeal swab | 28 | Deep sequencing | 100 copies/µL | Unpurified or lysed swab sample | — |
| El-Tholoth et al. [111] | ORF1ab | Synthetised | — | Fluorescence or colorimetric detection, LCV dye | 7 copies/µL | Eluting swab into water | 100% |
| Broughton et al. [112] | N, E | Nasopharyngeal swab | 78 | Lateral flow assay (LFA) | 10 copies/µL | RNA extraction | 95–100% |
| Jang et al. [113] | RdRP, N, E | Nasopharyngeal and oropharyngeal swabs, sputum, saliva and urine | 292 | Fluorescence detection | 10 copies/µL for N and RdRP genes: 100 copies/µL for E gene | RNA extraction | RdRP: 93.9%, N: 94.6%, RdRP/N: 96.9% |
| Lei et al. [114] | — | Synthetised | — | Turbidity and fluorescence detection, SYBR Green | 48 copies/µL | Eluting swab into water | — |
| Bektaş et al. [115] | N | Nasopharyngeal swabs, nasal mid-turbinate swabs, nasopharynx flush through | — | Fluorescence detection, SYBR Green | 0.1–2 copies/µL | Quick RNA extraction | >97% |
| Huang et al. [116] | ORF1ab, N, S | Nasopharyngeal swab | 16 | WarmStart Colorimetric LAMP | 0.8 copies/µL | Eluting swab into water | 100% |
| Chow et al. [117] | ORF3a, E | Nasopharyngeal swab, sputum, throat swab | 223 | WarmStart Colorimetric LAMP | ~2 copies/µL | RNA extraction | 95.07–98.21% |
| Rabe et al. [118] | ORF1a, N | Nasopharyngealswab, saliva | — | WarmStart Colorimetric LAMP | 1 copie/µL | Simple inactivation/ lyse step | 85% |
| Wang et al. [119] | N | Synthetised | — | Fluorescence detection, EvaGreen | 6 copies/µL | Extraction step omitted | — |
| LAMP | PCR | |
|---|---|---|
| Temperature | Isothermal reaction (60 to 65 °C) | Thermal cycling (multiple heating from 45 °C to 98 °C) |
| Reaction time | <1 h | ~2 h |
| DNA extrac-tion | Not required | Required |
| Primers | 4–6 primers recognize to 6–8 targets, extra looping primers increases sensitivity and effectiveness | 2 primers recognize 2 targets |
| Equipment | Dry block heater/water bath | Thermocycler |
| Modifica-tions | Real-time LAMP, MP-LAMP, RT-LAMP | Real-time PCR, MP-PCR, RT-PCR, nested PCR, nano-PCR, long PCR, RFLP-PCR |
| Sensitivity | 100× higher than standard PCR 100× lower than nested PCR | Up to modification |
| Products de-tection | With naked eye: turbidimetric analysis, fluorescent detection, electrophoresis, real-time protocol | Electrophoresis, real-time protocol |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Soroka, M.; Wasowicz, B.; Rymaszewska, A. Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR? Cells 2021, 10, 1931. https://doi.org/10.3390/cells10081931
Soroka M, Wasowicz B, Rymaszewska A. Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR? Cells. 2021; 10(8):1931. https://doi.org/10.3390/cells10081931
Chicago/Turabian StyleSoroka, Marianna, Barbara Wasowicz, and Anna Rymaszewska. 2021. "Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR?" Cells 10, no. 8: 1931. https://doi.org/10.3390/cells10081931
APA StyleSoroka, M., Wasowicz, B., & Rymaszewska, A. (2021). Loop-Mediated Isothermal Amplification (LAMP): The Better Sibling of PCR? Cells, 10(8), 1931. https://doi.org/10.3390/cells10081931






