Methods for Studying Magnaporthiopsis maydis, the Maize Late Wilt Causal Agent
Abstract
1. Introduction
2. Materials and Methods
2.1. Fungal Isolates and Growth Conditions
2.2. Magnaporthiopsis maydis Isolation
2.3. Magnaporthiopsis maydis Detection and Identification
2.4. Magnaporthiopsis maydis Growth Behavior Tests
2.5. Magnaporthiopsis maydis Virulence Assays under Controlled Conditions
2.6. Magnaporthiopsis maydis Infested Field Experiments
2.7. Statistical Analyses
3. Results
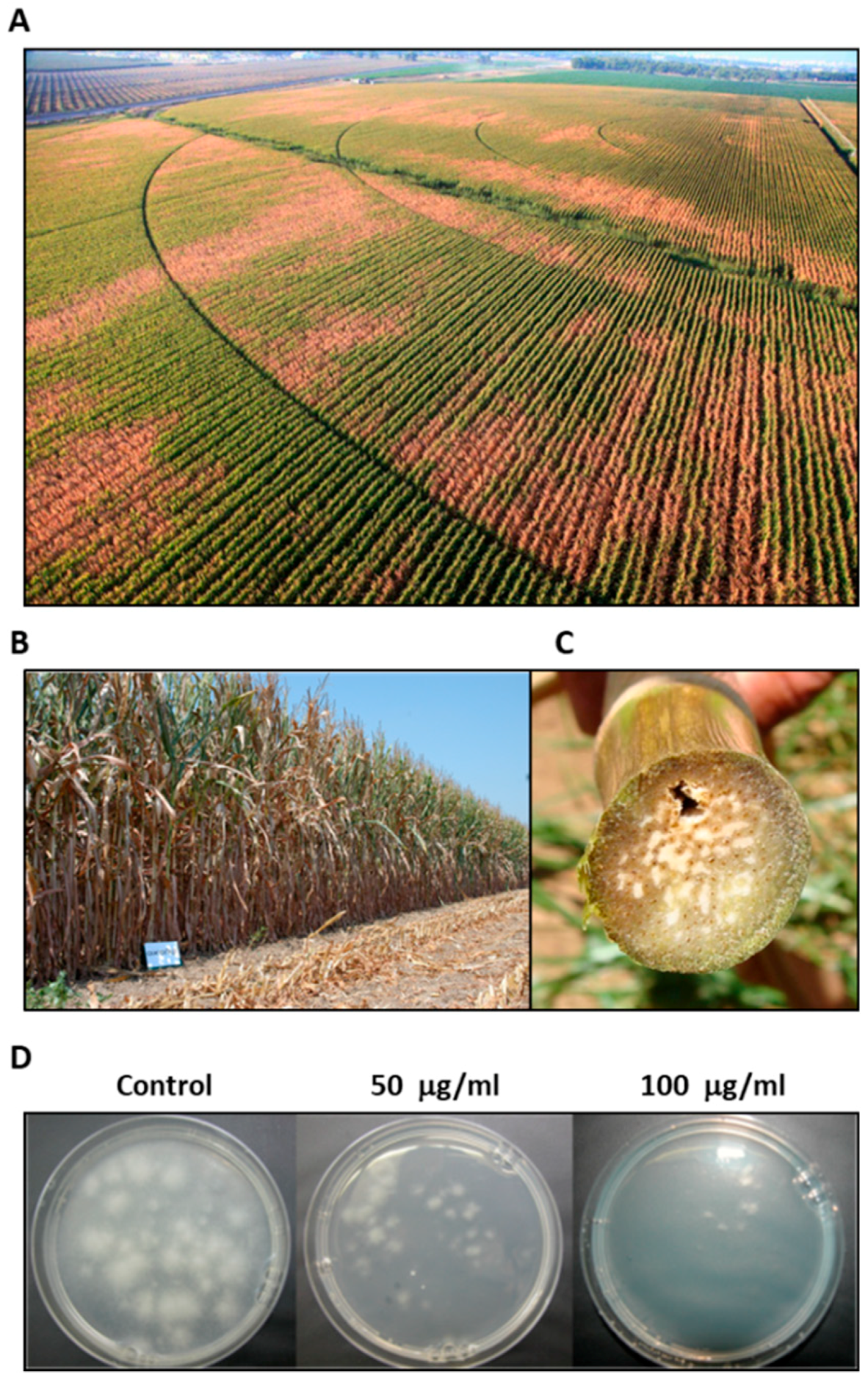
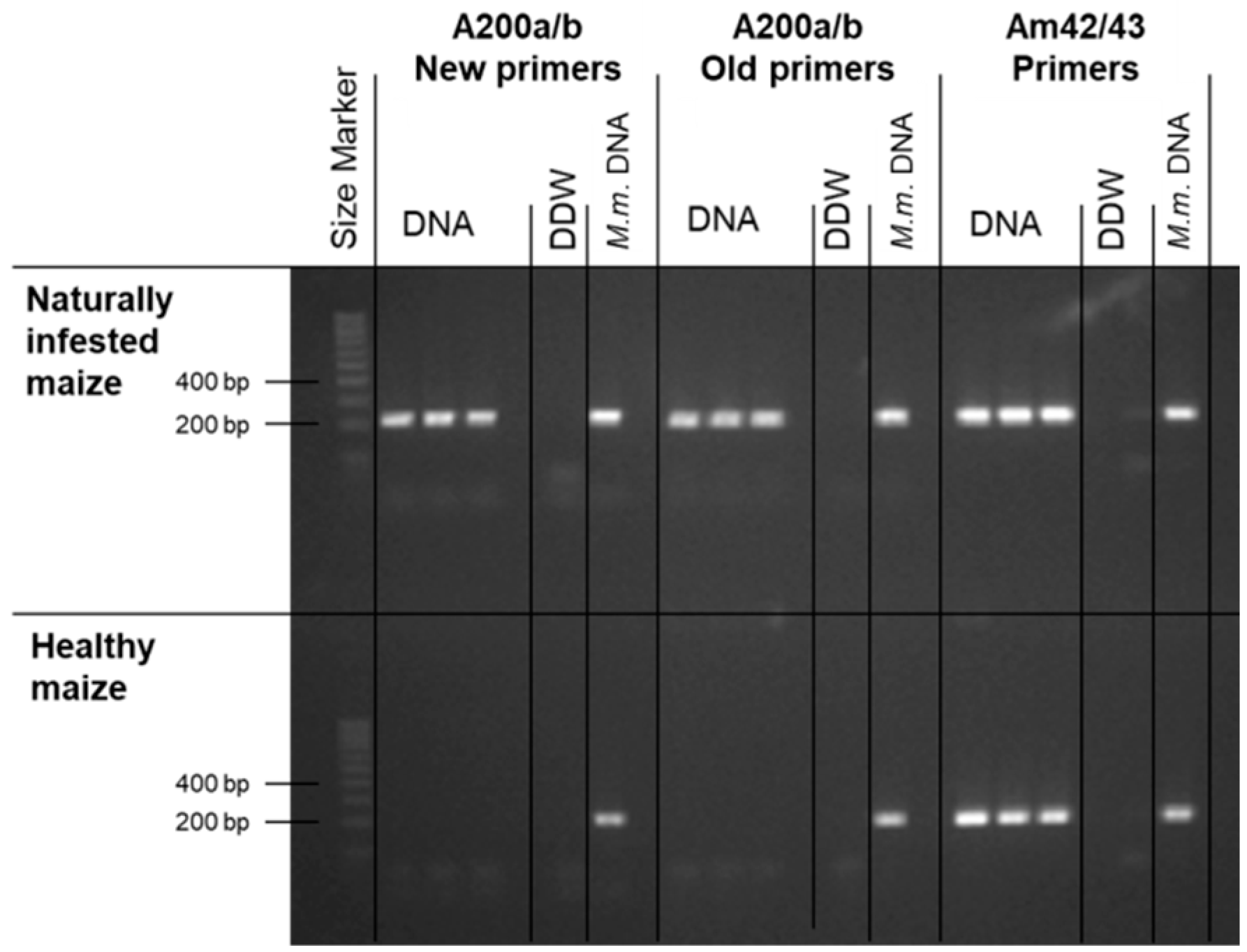
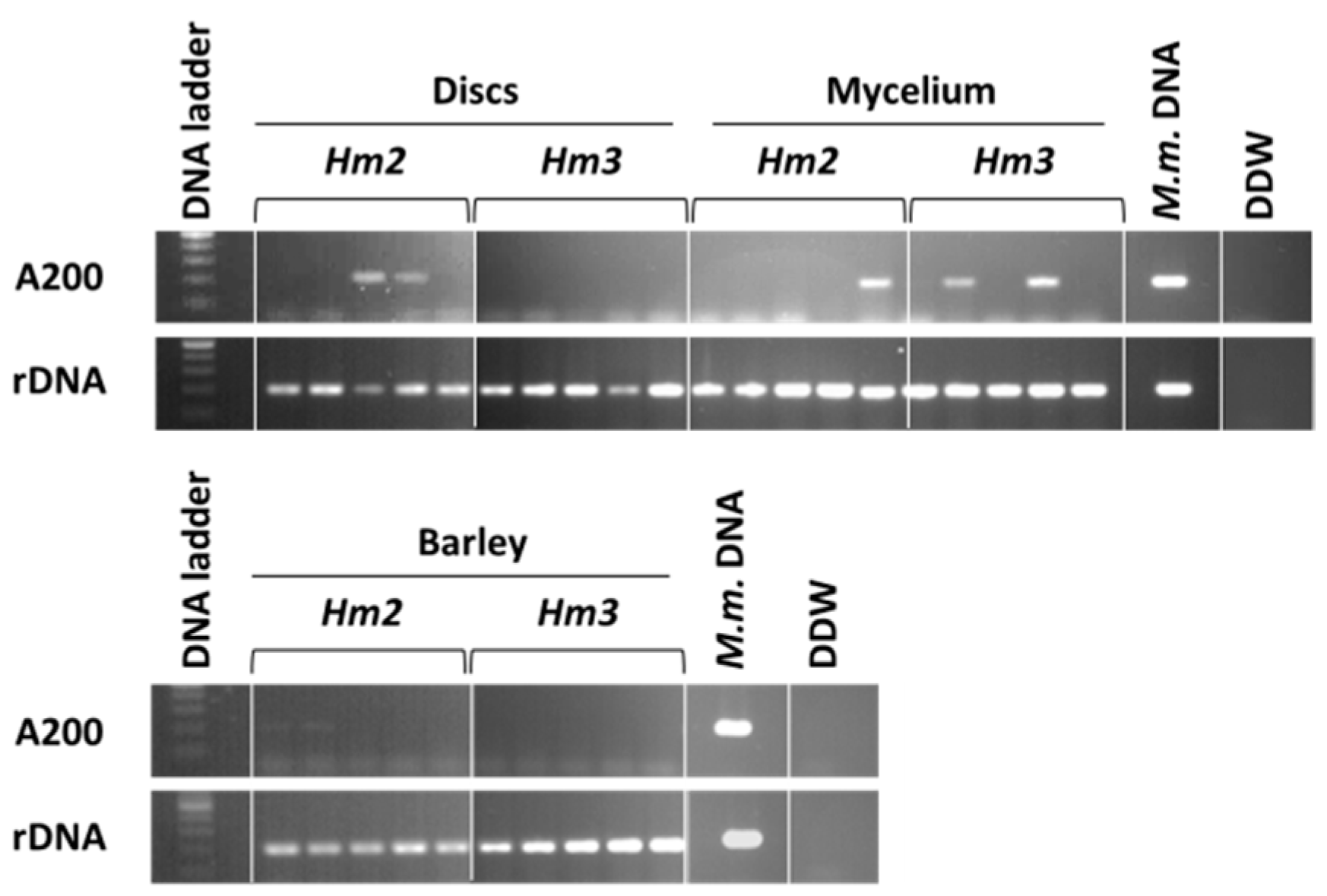
3.1. Magnaporthiopsis maydis Isolation and Detection
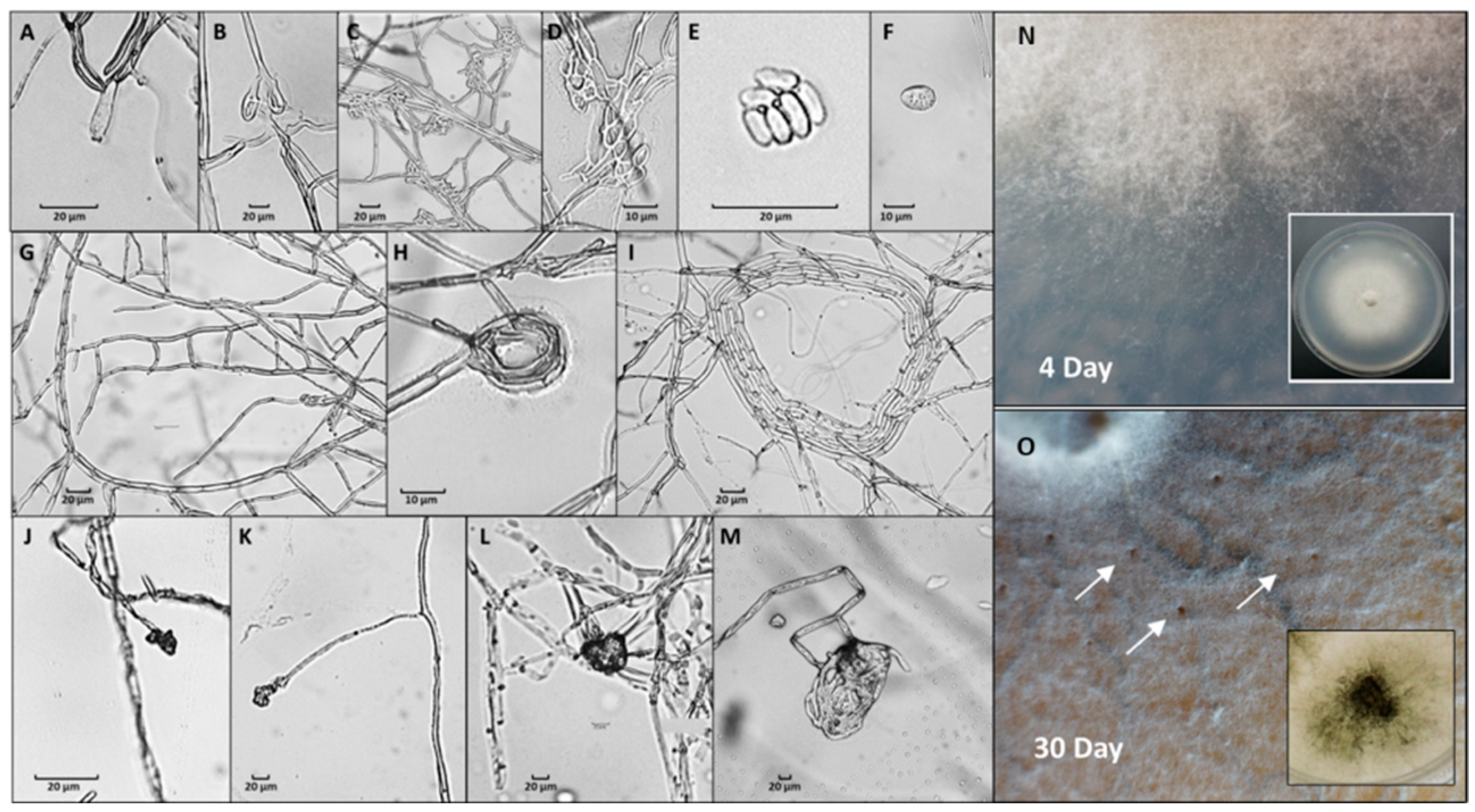
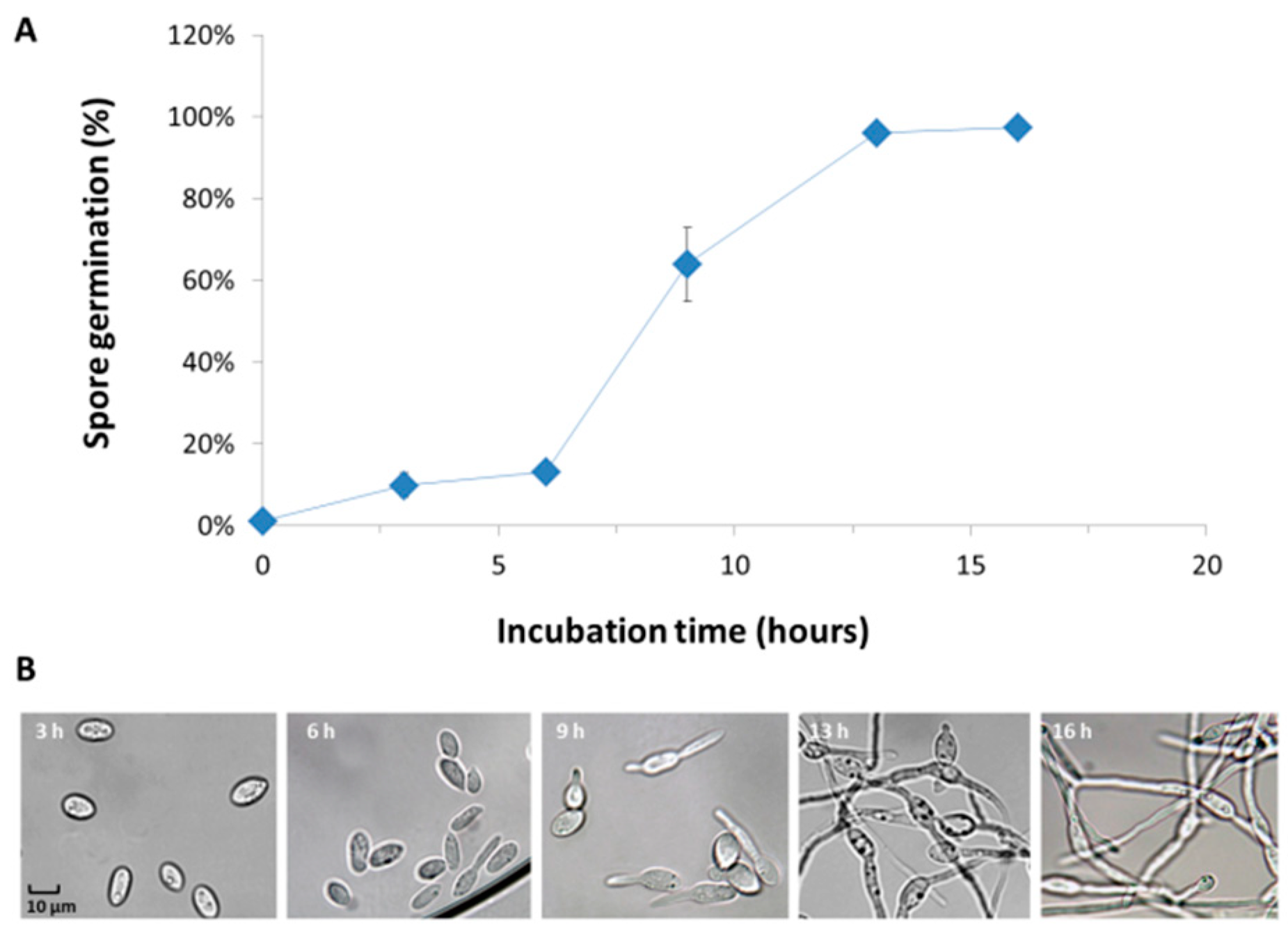
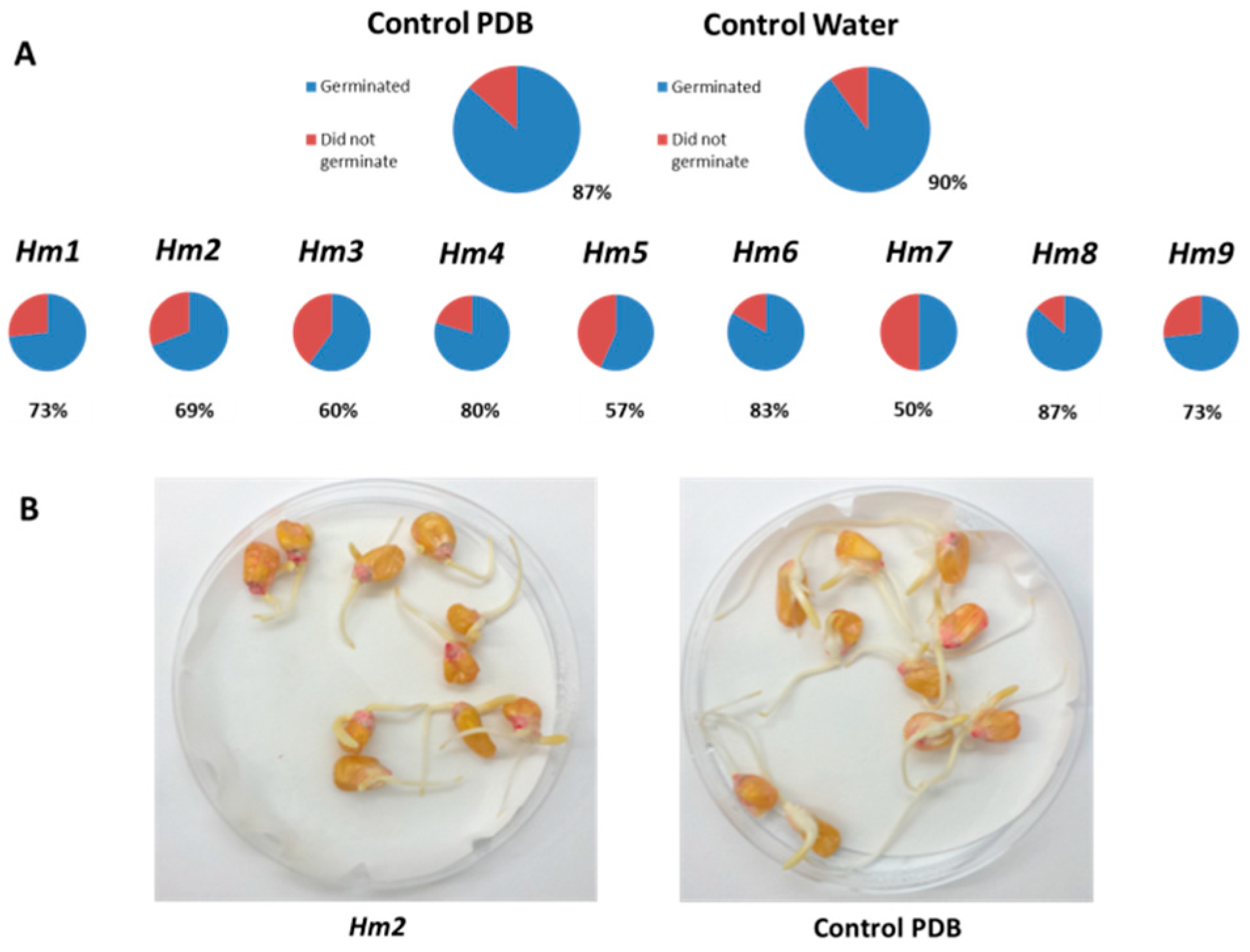
3.2. Magnaporthiopsis maydis Growth Behavior Tests
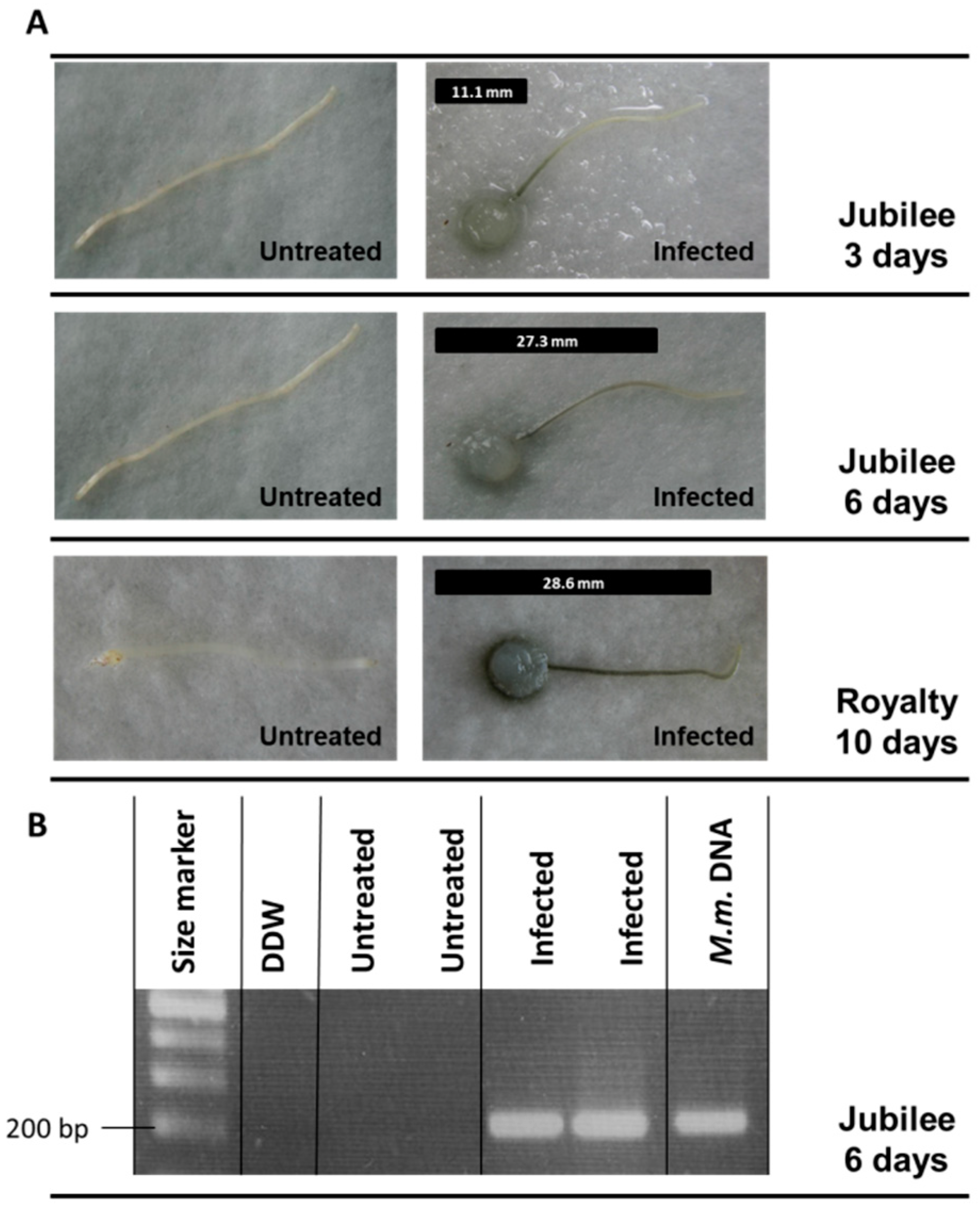
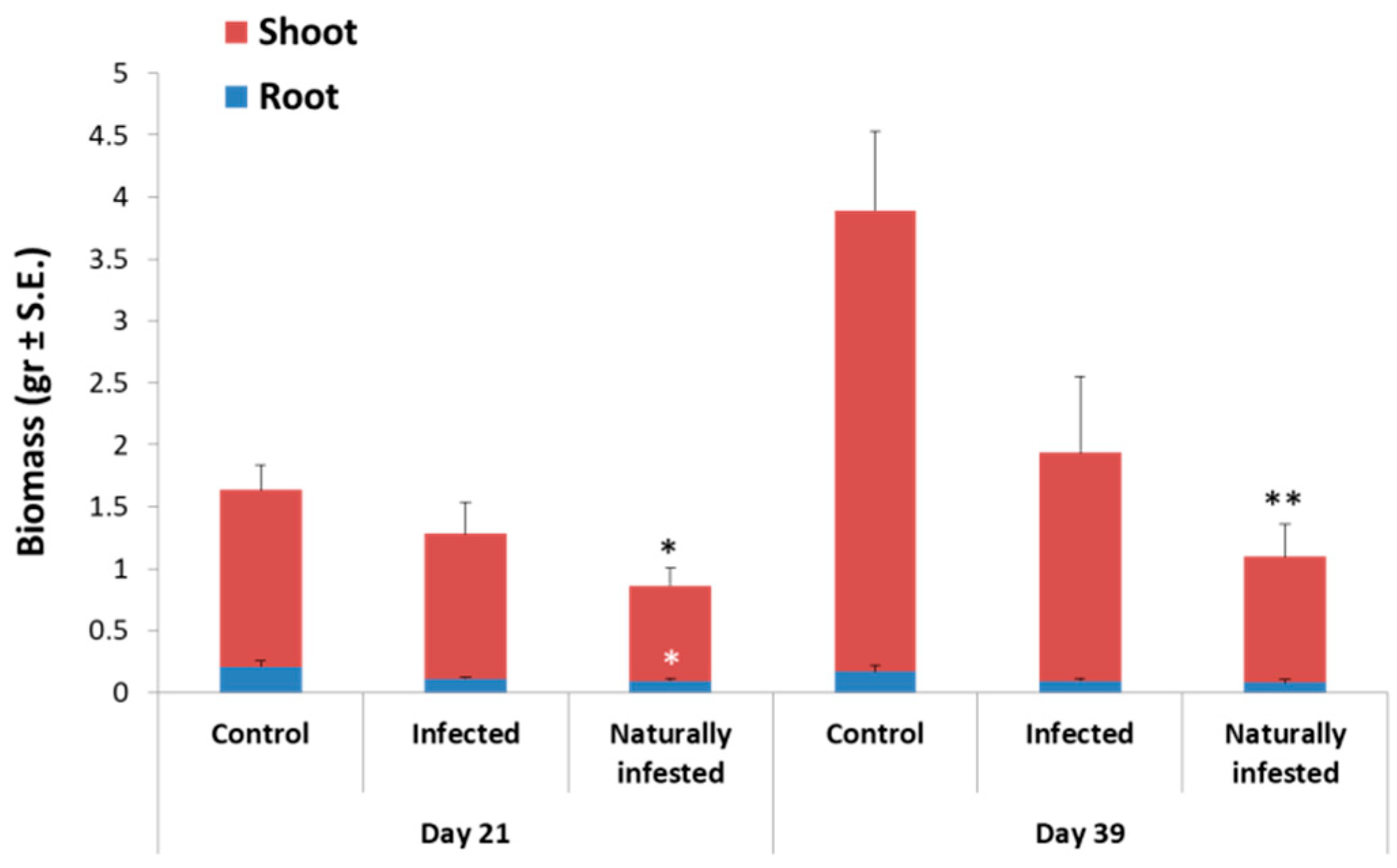
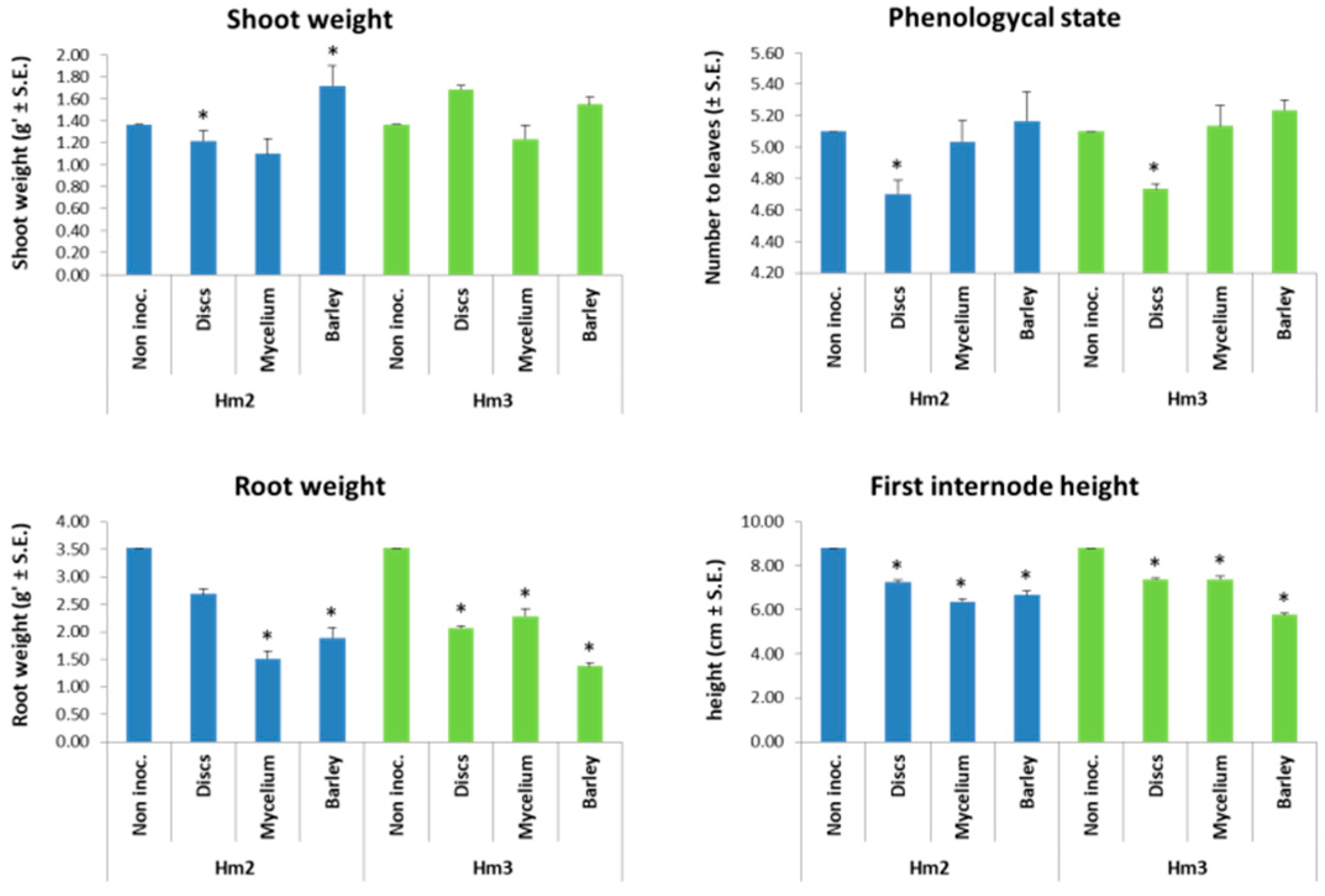
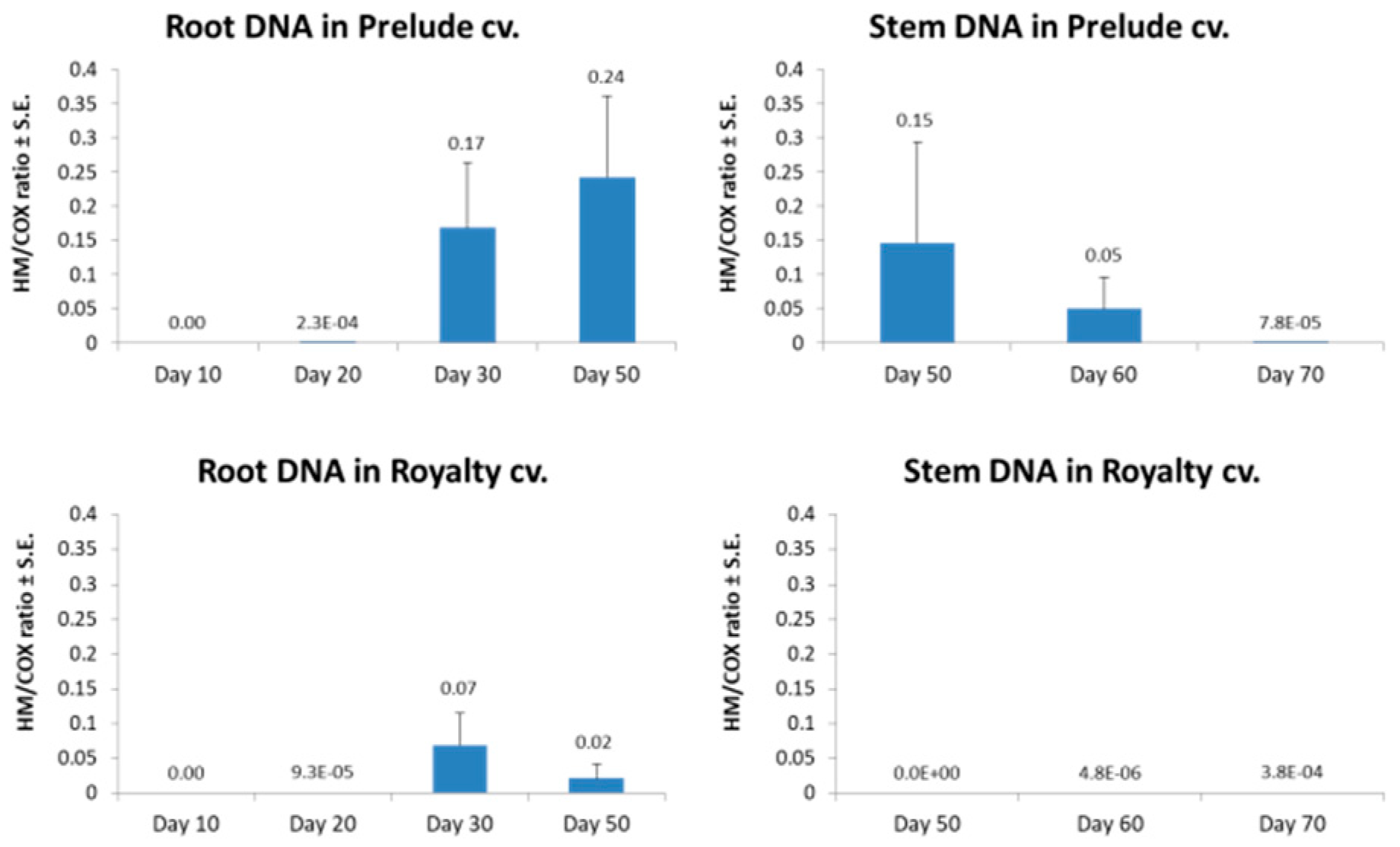
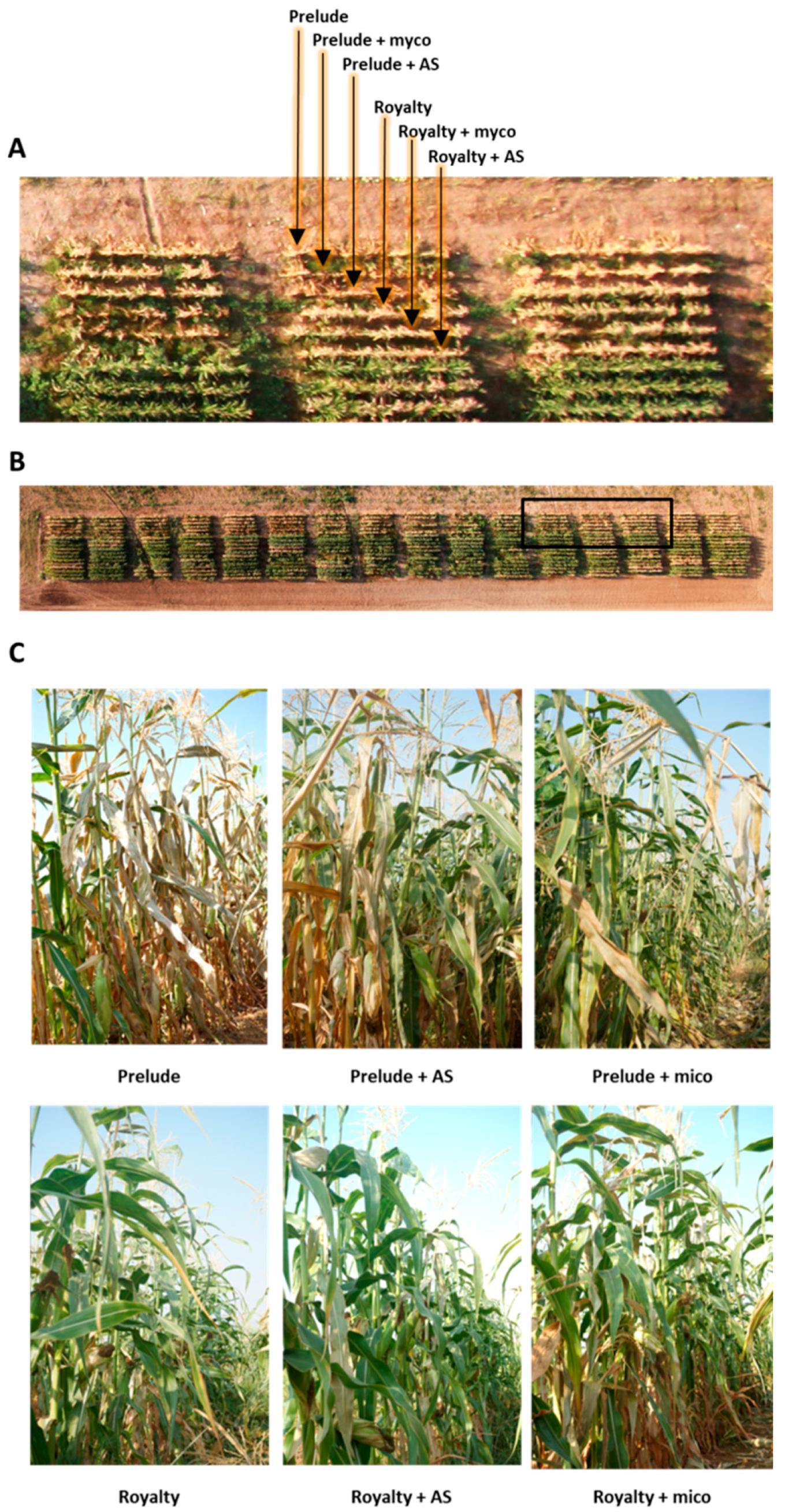
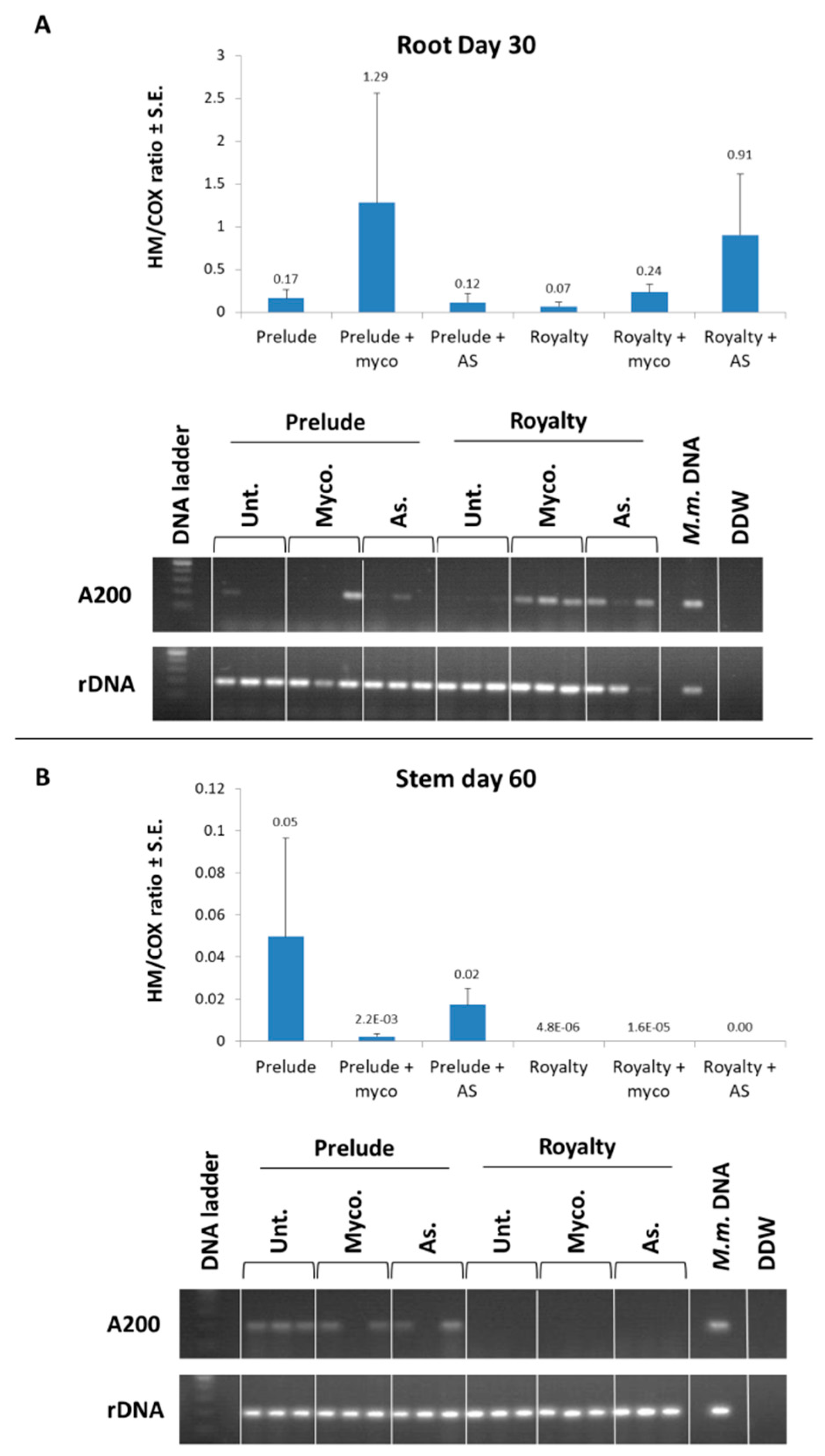
3.3. Magnaporthiopsis maydis Virulence Assays
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- Klaubauf, S.; Tharreau, D.; Fournier, E.; Groenewald, J.Z.; Crous, P.W.; de Vries, R.P.; Lebrun, M.H. Resolving the Polyphyletic Nature of Pyricularia (Pyriculariaceae). Stud. Mycol. 2014, 79, 85–120. [Google Scholar] [CrossRef] [PubMed]
- Gams, W. Phialophora and some similar morphologically little-differentiated anamorphs of divergent ascomycetes. Stud. Mycol. 2000, 45, 187–200. [Google Scholar]
- Ward, E.; Bateman, G.L. Comparison of Gaeumannomyces- and Phialophora-like fungal pathogens from maize and other plants using DNA methods. New Phytol. 1999, 141, 323–331. [Google Scholar] [CrossRef]
- Yuan, Z.-L.; Lin, F.-C.; Zhang, C.-L.; Kubicek, C.P. A new species of Harpophora (magnaporthaceae) recovered from healthy wild rice (oryza granulata) roots, representing a novel member of a beneficial dark septate endophyte. FEMS Microbiol. Lett. 2010, 307, 94–101. [Google Scholar] [CrossRef] [PubMed]
- Saleh, A.A.; Leslie, J.F. Cephalosporium Maydis is a Distinct Species in the Gaeumannomyces-Harpophora Species Complex. Mycologia 2004, 96, 1294–1305. [Google Scholar] [CrossRef] [PubMed]
- Sabet, K.A.; Samra, A.S.; Hingorani, M.K.; Mansour, I.M. Stalk and Root Rots of Maize in the United Arab Republic. FAO Plant Prot. Bull. 1961, 9, 121–125. [Google Scholar]
- Payak, M.M.; Lal, S.; Lilaramani, J.; Renfro, B.L. Cephalosporium Maydis—A New Threat to Maize in India. Indian Phytopathol. 1970, 23, 562–569. [Google Scholar]
- Molinero-Ruiz, M.L.; Melero-Vara, J.M.; Mateos, A. Cephalosporium Maydis, the Cause of Late Wilt in Maize, a Pathogen New to Portugal and Spain. Plant Dis. 2011, 94, 379. [Google Scholar] [CrossRef]
- Drori, R.; Sharon, A.; Goldberg, D.; Rabinovitz, O.; Levy, M.; Degani, O. Molecular Diagnosis for Harpophora Maydis, the Cause of Maize Late Wilt in Israel. Phytopathol. Mediterr. 2013, 52, 16–29. [Google Scholar]
- Degani, O.; Movshowitz, D.; Dor, S.; Meerson, A.; Goldblat, Y.; Rabinovitz, O. Evaluating Azoxystrobin Seed Coating against Maize Late Wilt Disease Using a Sensitive qPCR-based Method. Plant Dis. 2019, 103, 238–248. [Google Scholar] [CrossRef] [PubMed]
- Elshahawy, I.E.; El-Sayed, A.E.-K.B. Maximizing the Efficacy of Trichoderma to Control Cephalosporium maydis, Causing Maize Late Wilt Disease, Using Freshwater Microalgae Extracts. Egypt. J. Biol. Pest Control 2018, 28, 48. [Google Scholar] [CrossRef]
- El-Hosary, A.A.A.; EL-Fiki, I.A.I. Diallel Cross Analysis for Earliness, Yield, its Components and Resistance to Late Wilt in Maize. Int. J. Agric. Sci. Res. 2015, 5, 199–210. [Google Scholar]
- Warren, H.L. Potential Disease Problems: Late Wilt of Maize. Phytopathology 1983, 73, 782. [Google Scholar]
- El-Assiuty, E.M.; Ismael, A.M.; Zeller, K.A.; Leslie, J.F. Relative Colonization Ability of Greenhouse-Grown Maize by Four Lineages of Cephalosporium Maydis from Egypt. Phytopathology 1999, 89, S23. [Google Scholar]
- Saleh, A.A.; Zeller, K.A.; Ismael, A.S.; Fahmy, Z.M.; El-Assiuty, E.M.; Leslie, J.F. Amplified Fragment Length Polymorphism Diversity in Cephalosporium Maydis from Egypt. Phytopathology 2003, 93, 853–859. [Google Scholar] [CrossRef] [PubMed]
- Zeller, K.A.; Abou-Serie, M.I.; El-Assuity, E.M.; Fahmy, Z.M.; Bekheet, F.M.; Leslie, J.F. Relative Competitiveness and Virulence of Four Clonal Lineages of Cephalosporium maydis from Egypt toward Greenhouse-Grown Maize. Plant Dis. 2002, 86, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Zeller, K.A.; Jurgenson, J.E.; El-Assiuty, E.M.; Leslie, J.F. Isozyme and Amplified Fragment Length Polymorphisms from Cephalosporium Maydis in Egypt. Phytoparasitica 2000, 28, 121–130. [Google Scholar] [CrossRef]
- García-Carneros, A.; Girón, I.; Molinero-Ruiz, L. Aggressiveness of Cephalosporium Maydis Causing Late Wilt of Maize in Spain. Commun. Agric. Appl. Biol. Sci. 2011, 77, 173–179. [Google Scholar]
- Ortiz-Bustos, C.M.; Testi, L.; García-Carneros, A.B.; Molinero-Ruiz, L. Geographic Distribution and Aggressiveness of Harpophora Maydis in the Iberian Peninsula, and Thermal Detection of Maize Late Wilt. Eur. J. Plant Pathol. 2016, 144, 383–397. [Google Scholar] [CrossRef]
- Abendroth, L.J.; Elmore, R.W.; Boyer, M.J.; Marlay, S.K. Corn Growth and Development; Iowa State University Extension: Ames, IA, USA, 2011. [Google Scholar]
- Tej, R.; Rodríguez-Mallol, C.; Rodríguez-Arcos, R.; Karray-Bouraoui, N.; Molinero-Ruiz, L. Inhibitory Effect of Lycium Europaeum Extracts on Phytopathogenic Soil-Borne Fungi and the Reduction of Late Wilt in Maize. Eur. J. Plant Pathol. 2018, 152, 249–265. [Google Scholar] [CrossRef]
- Sabet, K.; Samra, A.; Mansour, I. Interaction between Fusarium Oxysporum F. Vasinfectum and Cephalosporium Maydis on Cotton and Maize. Ann. Appl. Biol. 1966, 58, 93–101. [Google Scholar] [CrossRef]
- Sabet, K.A.; Zaher, A.M.; Samra, A.S.; Mansour, I.M. Pathogenic Behaviour of Cephalosporium Maydis and C. Acremonium. Ann. Appl. Biol. 1970, 66, 257–263. [Google Scholar] [CrossRef]
- Degani, O.; Cernica, G. Diagnosis and Control of Harpophora Maydis, the Cause of Late Wilt in Maize. Adv. Microbiol. 2014, 4, 94–105. [Google Scholar] [CrossRef]
- El-Shafey, H.A.; Claflin, L.E. Late Wilt; APS Press: St. Paul, MN, USA, 1999; pp. 43–44. [Google Scholar]
- Samra, A.S.; Sabet, K.A.; Hingorani, M.K. A New Wilt Disease of Maize in Egypt. Plant Dis. Rep. 1962, 46, 481–483. [Google Scholar]
- Shehata, F.A. The Inheritance of Resistence to Late Wilt Caused by Cephalosporium maydis in Some Corn Lines. Master’s Thesis, Al-Azhar University, Cairo, Egypt, 1976. [Google Scholar]
- Degani, O.; Dor, S.; Movshowitz, D.; Fraidman, E.; Rabinowitz, O.; Graph, S. Effective Chemical Protection against the Maize Late Wilt Causal Agent, Harpophora Maydis, in the Field. PLoS ONE 2018, 13, e0208353. [Google Scholar] [CrossRef] [PubMed]
- Degani, O.; Goldblat, Y. Ambient Stresses Regulate the Development of the Maize Late Wilt Causing Agent, Harpophora Maydis. Agric. Sci. 2014, 5, 571–582. [Google Scholar]
- Singh, S.D.; Siradhana, B.S. Date of Sowing in Relation to Late Wilt Disease of Maize. Indian Phytopathol. 1988, 41, 489–491. [Google Scholar]
- Sabet, K.A.; Samra, A.S.; Mansour, I.M. Saprophytic Behaviour of Cephalosporium Maydis and C. Acremonium. Ann. Appl. Biol. 1970, 66, 265–271. [Google Scholar] [CrossRef]
- Michail, S.H.; Abou-Elseoud, M.S.; Nour Eldin, M.S. Seed Health Testing of Corn for Cephalosporium Maydis. Acta Phytopathol. Entomol. Hung. 1999, 34, 35–42. [Google Scholar]
- Sahab, A.F.; Osman, A.R.; Soleman, N.K.; Mikhail, M.S. Studies on Root-Rot of Lupin in Egypt and its Control. Egypt. J. Phytopathol. 1985, 17, 23–35. [Google Scholar]
- Degani, O.; Dor, S.; Graph, S.; Dafny-Yelin, M.; Rabinovitz, O. Uncovering host Range for the Maize Pathogen Harpophora Maydis. In Proceedings of the International Congress of Plant Pathology (ICPP) 2018: Plant Health in A Global Economy, Boston, MA, USA, 29 July–1 August 2018. [Google Scholar]
- Abd El-Rahim, M.F.; Fahmy, G.M.; Fahmy, Z.M. Alterations in transpiration and stem vascular tissues of two maize cultivars under conditions of water stress and late wilt disease. Plant Pathol. 1998, 47, 216–223. [Google Scholar] [CrossRef]
- Muhammad, S.; Amusa, N.A. In Vitro Inhibition of Growth of Some Seedling Blight Inducing Pathogens by Compost-Inhabiting Microbes. Afr. J. Biotechnol. 2003, 2, 161–164. [Google Scholar]
- El-Assiuty, E.M.; El-Hamahmy, A.A.; El-Sharkawy, A.Y. Bacillus subtilis, Pseudomonas Fluorescens and Verticillium Tricorpus as Biological Agents against Late-Wilt of Maize. Egypt. J. Appl. Sci. 1991, 6, 8245–8829. [Google Scholar]
- El-Mehalowy, A.A.; Hassanein, N.M.; Khater, H.M.; Daram El-Din, E.A.; Youssef, Y.A. Influence of Maize Root Colonization by Rhizosphere Actinomycetes and Yeast Fungi on Plant Growth and on the Biological Control of Late Wilt Disease. Inter. J. Agric. Biol. 2004, 6, 599–605. [Google Scholar]
- Abd-el-Rahim, M.F.; Sabet, K.A.; El-Shafey, H.A.; El-Assiuty, E.M. Chemical Control of the Late-Wilt Disease of Maize Caused by Cephalosporium Maydis. Agric. Res. Rev. 1982, 60, 31–49. [Google Scholar]
- Degani, O.; Weinberg, T.; Graph, S. Chemical Control of Maize Late Wilt in the Field. Phytoparasitica 2014, 42, 559–570. [Google Scholar] [CrossRef]
- Costanzo, S. New Pest response guidelines. In Late Wilt of Corn (Harpophora maydis); Michalak, P.S., Ed.; United States Department of Agriculture (USDA), Animal and Plant Health Inspection Service (APHIS), Plant Protection and Quarantine (PPQ): Washington, DC, USA, 2011; Volume 1, pp. 1–102. [Google Scholar]
- El-Naggarr, A.A.A.; Sabryr, A.M.; Yassin, M.A. Impact of Late Wilt Disease Caused by Harpophora maydis on Maize Yield. J. Biol. Chem. Environ. Sci. 2015, 10, 577–595. [Google Scholar]
- Sabet, K.A.; Samra, A.S.; Abdel-Rahim, M.F. Systemic Action of Benomyl against late-Wilt Disease of Maize. Ann. Appl. Biol. 1972, 71, 211–218. [Google Scholar] [CrossRef]
- Harris, J.L. Modified Method for Fungal Slide Culture. J. Clin. Microbiol. 1986, 24, 460–461. [Google Scholar]
- Murray, M.; Thompson, W.F. Rapid Isolation of High Molecular Weight Plant DNA. Nucleic Acids Res. 1980, 8, 4321–4326. [Google Scholar] [CrossRef]
- Fromont-Racine, M.; Senger, B.; Saveanu, C.; Fasiolo, F. Ribosome Assembly in Eukaryotes. Gene 2003, 313, 17–42. [Google Scholar] [CrossRef]
- Weller, S.; Elphinstone, J.; Smith, N.; Boonham, N.; Stead, D. Detection of Ralstonia Solanacearum Strains with a Quantitative, Multiplex, Real-Time, Fluorogenic PCR (Taqman) Assay. Appl. Environ. Microbiol. 2000, 66, 2853–2858. [Google Scholar] [CrossRef] [PubMed]
- Yuan, J.S.; Reed, A.; Chen, F.; Stewart, N.C. Statistical Analysis of Real-Time PCR Data. BMC Bioinform. 2006, 7, 85. [Google Scholar] [CrossRef] [PubMed]
- Degani, O.; Drori, R.; Goldblat, Y. Plant Growth Hormones Suppress the Development of Harpophora Maydis, the Cause of Late Wilt in Maize. Physiol. Mol. Biol. Plants 2015, 21, 137–149. [Google Scholar] [CrossRef][Green Version]
- El-Gremi, S.M.A.; Belal, E.B.A.; Ghazy, N.A. Cephalosporium Maydis as Affected by Maize Root Exudates and Role of the Fungal Metabolites in Pathogenesis. J. Agric. Sci. Mansoura Univ. 2007, 32, 7605–7615. [Google Scholar]
- Degani, O. Pathogenicity Assay for Cochliobolus Heterostrophus G-Protein and MAPK Signaling Deficiency Strains. Am. J. Plant Sci. 2014, 5, 1318–1328. [Google Scholar] [CrossRef][Green Version]
- Degani, O.; Maor, R.; Hadar, R.; Sharon, A.; Horwitz, B.A. Host Physiology and Pathogenic Variation of Cochliobolus heterostrophus Strains with Mutations in the G Protein α Subunit, Cga1. Appl. Environ. Microbiol. 2004, 70, 5005–5009. [Google Scholar] [CrossRef]
- Campos, M.D.; Patanita, M.; Campos, C.; Materatski, P.; Varanda, C.M.; Brito, I.; do Rosário Félix, M. Detection and Quantification of Fusarium spp. (F. oxysporum, F. verticillioides, F. graminearum) and Magnaporthiopsis maydis in Maize Using Real-Time PCR Targeting the ITS Region. Agronomy 2019, 9, 45. [Google Scholar] [CrossRef]
- Khokhar, M.K.; Hooda, K.S.; Sharma, S.S.; Singh, V. Post Flowering Stalk Rot Complex of Maize-Present Status and Future Prospects. Maydica 2014, 59, 226–242. [Google Scholar]
- Payak, M.; Sharma, R. Disease Rating Scales in Maize in India. Tech. Scoring Resist. Import. Dis. Maize 1983, 1–4. [Google Scholar]
- Rakesh, B.; Gangappa, E.; Sonali, G.; Gowda, R.P.V.; Swamy, S.D.; Ramesh, S.; Hemareddy, H.B.; Nagaraju, N. Modified Method of Screening Maize Inbred Lines to Late Wilt Disease Caused by Harpophora Maydis. Mysore J. Agric. Sci. 2016, 50, 684–690. [Google Scholar]
- Ghazy, N.A.; El-Gremi, S.; Belal, E.-S. Chemical and Histological Differences of Corn (Zea Mays L.) Responsive to Harpophora maydis Infection. Environ. Biodivers. Soil Secur. 2017, 1, 191–201. [Google Scholar] [CrossRef]


| Pairs | Primer | Sequence | Uses | Amplification | References |
|---|---|---|---|---|---|
| Pair 1 (old primers) | A200a-for | 5′-CCGACGCCTAAAATACAGGA-3′ | PCR and qPCR | M. maydis AFLP-derived specific fragment | [9] |
| A200a-rev | 5′-GGGCTTTTTAGGGCCTTTTT-3′ | ||||
| Pair 2 (new) primers) | A200a-for | 5′-CCTAGTAGTCCCGACTGTTAGG-3′ | PCR | M. maydis AFLP-derived specific fragment | [24] |
| A200a-rev | 5′-TTGGTTCACCGTCTTTTGTAGG-3′ | ||||
| Pair 3 | Am42 | 5′-CAACTACGAGCTTTTTAACTGC-3′ | PCR Control | Eukaryotic ribosomal DNA 18S rRNA gene product, rDNA | [46] |
| Am43 | 5′-CAAATTACCCAATCCCGACAC-3′ | ||||
| Pair 4 | Cox-F | 5′-GTATGCCACGTCGCATTCCAGA-3′ | qPCR Control | cytochrome c oxidase (COX) gene product | [47] |
| Cox-R | 5′-CAACTACGGATATATAAGRRCCRRAACTG-3′ 1 |
| Assessment | Days after Sowing | Prelude cv. | Royalty cv. | ||||
|---|---|---|---|---|---|---|---|
| NT 3 | AS | Myco | NT | AS | Myco | ||
| Root Wet Weight (gr ± S.E.) | 10 | 0.53 ± 0.16 | 0.62 ± 0.12 | 0.58 ± 0.03 | 0.62 ± 0.06 | 0.52 ± 0.09 | 0.51 ± 0.07 |
| Shoot Wet Weight (gr ± S.E.) | 10 | 1.77 ± 0.10 | 2.44 ± 0.29 4 | 1.93 ± 0.23 | 2.57 ± 0.15 | 2.35 ± 0.56 | 2.06 ± 0.33 |
| Wilting (%) 1 | 69 | 60 | 48.6 | 40.3 4 | 3.7 | 1.2 | 1.7 |
| 75 | 100 | 96.3 | 97.1 | 26.7 | 41.9 4 | 9.3 | |
| Yield (kg/m2) 2 | 77 | Less than 0.5 | Less than 0.5 | Less than 0.5 | 1.7 ± 0.10 | 1.7 ± 0.13 | 1.6 ± 0.10 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Degani, O.; Dor, S.; Movshovitz, D.; Rabinovitz, O. Methods for Studying Magnaporthiopsis maydis, the Maize Late Wilt Causal Agent. Agronomy 2019, 9, 181. https://doi.org/10.3390/agronomy9040181
Degani O, Dor S, Movshovitz D, Rabinovitz O. Methods for Studying Magnaporthiopsis maydis, the Maize Late Wilt Causal Agent. Agronomy. 2019; 9(4):181. https://doi.org/10.3390/agronomy9040181
Chicago/Turabian StyleDegani, Ofir, Shlomit Dor, Daniel Movshovitz, and Onn Rabinovitz. 2019. "Methods for Studying Magnaporthiopsis maydis, the Maize Late Wilt Causal Agent" Agronomy 9, no. 4: 181. https://doi.org/10.3390/agronomy9040181
APA StyleDegani, O., Dor, S., Movshovitz, D., & Rabinovitz, O. (2019). Methods for Studying Magnaporthiopsis maydis, the Maize Late Wilt Causal Agent. Agronomy, 9(4), 181. https://doi.org/10.3390/agronomy9040181






